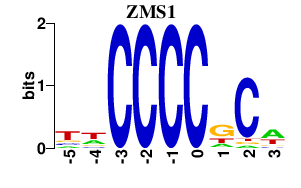
Results for ZMS1
Z-value: 0.76
Transcription factors associated with ZMS1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
RSF2
|
S000003888 | Zinc-finger protein |
Activity-expression correlation:
Activity profile of ZMS1 motif
Sorted Z-values of ZMS1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| YKL163W | 13.40 |
PIR3
|
O-glycosylated covalently-bound cell wall protein required for cell wall stability; expression is cell cycle regulated, peaking in M/G1 and also subject to regulation by the cell integrity pathway |
|
| YGR067C | 11.87 |
Putative protein of unknown function; contains a zinc finger motif similar to that of Adr1p |
||
| YDR342C | 11.10 |
HXT7
|
High-affinity glucose transporter of the major facilitator superfamily, nearly identical to Hxt6p, expressed at high basal levels relative to other HXTs, expression repressed by high glucose levels |
|
| YOR100C | 10.87 |
CRC1
|
Mitochondrial inner membrane carnitine transporter, required for carnitine-dependent transport of acetyl-CoA from peroxisomes to mitochondria during fatty acid beta-oxidation |
|
| YDR277C | 10.53 |
MTH1
|
Negative regulator of the glucose-sensing signal transduction pathway, required for repression of transcription by Rgt1p; interacts with Rgt1p and the Snf3p and Rgt2p glucose sensors; phosphorylated by Yck1p, triggering Mth1p degradation |
|
| YKL217W | 9.85 |
JEN1
|
Lactate transporter, required for uptake of lactate and pyruvate; phosphorylated; expression is derepressed by transcriptional activator Cat8p during respiratory growth, and repressed in the presence of glucose, fructose, and mannose |
|
| YLR327C | 9.53 |
TMA10
|
Protein of unknown function that associates with ribosomes |
|
| YGR243W | 9.16 |
FMP43
|
Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies |
|
| YFR053C | 8.84 |
HXK1
|
Hexokinase isoenzyme 1, a cytosolic protein that catalyzes phosphorylation of glucose during glucose metabolism; expression is highest during growth on non-glucose carbon sources; glucose-induced repression involves the hexokinase Hxk2p |
|
| YDR536W | 8.55 |
STL1
|
Glycerol proton symporter of the plasma membrane, subject to glucose-induced inactivation, strongly but transiently induced when cells are subjected to osmotic shock |
|
| YFL052W | 7.31 |
Putative zinc cluster protein that contains a DNA binding domain; null mutant sensitive to calcofluor white, low osmolarity and heat, suggesting a role for YFL052Wp in cell wall integrity |
||
| YOR382W | 7.03 |
FIT2
|
Mannoprotein that is incorporated into the cell wall via a glycosylphosphatidylinositol (GPI) anchor, involved in the retention of siderophore-iron in the cell wall |
|
| YOR348C | 6.87 |
PUT4
|
Proline permease, required for high-affinity transport of proline; also transports the toxic proline analog azetidine-2-carboxylate (AzC); PUT4 transcription is repressed in ammonia-grown cells |
|
| YMR107W | 6.84 |
SPG4
|
Protein required for survival at high temperature during stationary phase; not required for growth on nonfermentable carbon sources |
|
| YJR115W | 6.57 |
Putative protein of unknown function |
||
| YKR097W | 6.45 |
PCK1
|
Phosphoenolpyruvate carboxykinase, key enzyme in gluconeogenesis, catalyzes early reaction in carbohydrate biosynthesis, glucose represses transcription and accelerates mRNA degradation, regulated by Mcm1p and Cat8p, located in the cytosol |
|
| YKL026C | 6.27 |
GPX1
|
Phospholipid hydroperoxide glutathione peroxidase induced by glucose starvation that protects cells from phospholipid hydroperoxides and nonphospholipid peroxides during oxidative stress |
|
| YOR178C | 6.16 |
GAC1
|
Regulatory subunit for Glc7p type-1 protein phosphatase (PP1), tethers Glc7p to Gsy2p glycogen synthase, binds Hsf1p heat shock transcription factor, required for induction of some HSF-regulated genes under heat shock |
|
| YDR343C | 6.04 |
HXT6
|
High-affinity glucose transporter of the major facilitator superfamily, nearly identical to Hxt7p, expressed at high basal levels relative to other HXTs, repression of expression by high glucose requires SNF3 |
|
| YML042W | 5.96 |
CAT2
|
Carnitine acetyl-CoA transferase present in both mitochondria and peroxisomes, transfers activated acetyl groups to carnitine to form acetylcarnitine which can be shuttled across membranes |
|
| YJR095W | 5.83 |
SFC1
|
Mitochondrial succinate-fumarate transporter, transports succinate into and fumarate out of the mitochondrion; required for ethanol and acetate utilization |
|
| YDR119W-A | 5.68 |
Putative protein of unknown function |
||
| YPL054W | 5.65 |
LEE1
|
Zinc-finger protein of unknown function |
|
| YOL052C-A | 5.61 |
DDR2
|
Multistress response protein, expression is activated by a variety of xenobiotic agents and environmental or physiological stresses |
|
| YNR073C | 5.52 |
Putative mannitol dehydrogenase |
||
| YEL039C | 5.29 |
CYC7
|
Cytochrome c isoform 2, expressed under hypoxic conditions; electron carrier of the mitochondrial intermembrane space that transfers electrons from ubiquinone-cytochrome c oxidoreductase to cytochrome c oxidase during cellular respiration |
|
| YEL011W | 5.11 |
GLC3
|
Glycogen branching enzyme, involved in glycogen accumulation; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern |
|
| YMR175W | 5.01 |
SIP18
|
Protein of unknown function whose expression is induced by osmotic stress |
|
| YIL057C | 5.00 |
Putative protein of unknown function; expression induced under carbon limitation and repressed under high glucose |
||
| YPL230W | 4.99 |
USV1
|
Putative transcription factor containing a C2H2 zinc finger; mutation affects transcriptional regulation of genes involved in protein folding, ATP binding, and cell wall biosynthesis |
|
| YOR383C | 4.96 |
FIT3
|
Mannoprotein that is incorporated into the cell wall via a glycosylphosphatidylinositol (GPI) anchor, involved in the retention of siderophore-iron in the cell wall |
|
| YMR081C | 4.92 |
ISF1
|
Serine-rich, hydrophilic protein with similarity to Mbr1p; overexpression suppresses growth defects of hap2, hap3, and hap4 mutants; expression is under glucose control; cotranscribed with NAM7 in a cyp1 mutant |
|
| YLR312C | 4.69 |
QNQ1
|
Protein of unknown function; null mutant quiescent and non-quiescent cells exhibit reduced reproductive capacity |
|
| YGR032W | 4.64 |
GSC2
|
Catalytic subunit of 1,3-beta-glucan synthase, involved in formation of the inner layer of the spore wall; activity positively regulated by Rho1p and negatively by Smk1p; has similarity to an alternate catalytic subunit, Fks1p (Gsc1p) |
|
| YFR017C | 4.64 |
Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and is induced in response to the DNA-damaging agent MMS; YFR017C is not an essential gene |
||
| YMR194C-B | 4.51 |
Putative protein of unknown function |
||
| YHR096C | 4.49 |
HXT5
|
Hexose transporter with moderate affinity for glucose, induced in the presence of non-fermentable carbon sources, induced by a decrease in growth rate, contains an extended N-terminal domain relative to other HXTs |
|
| YOR374W | 4.35 |
ALD4
|
Mitochondrial aldehyde dehydrogenase, required for growth on ethanol and conversion of acetaldehyde to acetate; phosphorylated; activity is K+ dependent; utilizes NADP+ or NAD+ equally as coenzymes; expression is glucose repressed |
|
| YOR120W | 4.17 |
GCY1
|
Putative NADP(+) coupled glycerol dehydrogenase, proposed to be involved in an alternative pathway for glycerol catabolism; member of the aldo-keto reductase (AKR) family |
|
| YGR183C | 4.12 |
QCR9
|
Subunit 9 of the ubiquinol cytochrome-c reductase complex, which is a component of the mitochondrial inner membrane electron transport chain; required for electron transfer at the ubiquinol oxidase site of the complex |
|
| YKL109W | 3.99 |
HAP4
|
Subunit of the heme-activated, glucose-repressed Hap2p/3p/4p/5p CCAAT-binding complex, a transcriptional activator and global regulator of respiratory gene expression; provides the principal activation function of the complex |
|
| YHR217C | 3.94 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; located in the telomeric region TEL08R. |
||
| YKL044W | 3.92 |
Dubious open reading frame unlikely to encode a functional protein, based on available experimental and comparative sequence data |
||
| YDL222C | 3.92 |
FMP45
|
Integral membrane protein localized to mitochondria (untagged protein) and eisosomes, immobile patches at the cortex associated with endocytosis; sporulation and sphingolipid content are altered in mutants; has homologs SUR7 and YNL194C |
|
| YNR002C | 3.88 |
ATO2
|
Putative transmembrane protein involved in export of ammonia, a starvation signal that promotes cell death in aging colonies; phosphorylated in mitochondria; member of the TC 9.B.33 YaaH family; homolog of Ady2p and Y. lipolytica Gpr1p |
|
| YLR023C | 3.87 |
IZH3
|
Membrane protein involved in zinc metabolism, member of the four-protein IZH family, expression induced by zinc deficiency; deletion reduces sensitivity to elevated zinc and shortens lag phase, overexpression reduces Zap1p activity |
|
| YPR184W | 3.72 |
GDB1
|
Glycogen debranching enzyme containing glucanotranferase and alpha-1,6-amyloglucosidase activities, required for glycogen degradation; phosphorylated in mitochondria |
|
| YNR034W-A | 3.66 |
Putative protein of unknown function; expression is regulated by Msn2p/Msn4p |
||
| YGL062W | 3.61 |
PYC1
|
Pyruvate carboxylase isoform, cytoplasmic enzyme that converts pyruvate to oxaloacetate; highly similar to isoform Pyc2p but differentially regulated; mutations in the human homolog are associated with lactic acidosis |
|
| YIL162W | 3.59 |
SUC2
|
Invertase, sucrose hydrolyzing enzyme; a secreted, glycosylated form is regulated by glucose repression, and an intracellular, nonglycosylated enzyme is produced constitutively |
|
| YBR105C | 3.56 |
VID24
|
Peripheral membrane protein located at Vid (vacuole import and degradation) vesicles; regulates fructose-1,6-bisphosphatase (FBPase) targeting to the vacuole; involved in proteasome-dependent catabolite degradation of FBPase |
|
| YLR377C | 3.41 |
FBP1
|
Fructose-1,6-bisphosphatase, key regulatory enzyme in the gluconeogenesis pathway, required for glucose metabolism |
|
| YJL045W | 3.34 |
Minor succinate dehydrogenase isozyme; homologous to Sdh1p, the major isozyme reponsible for the oxidation of succinate and transfer of electrons to ubiquinone; induced during the diauxic shift in a Cat8p-dependent manner |
||
| YGR043C | 3.31 |
NQM1
|
Protein of unknown function; transcription is repressed by Mot1p and induced by alpha-factor and during diauxic shift; null mutant non-quiescent cells exhibit reduced reproductive capacity |
|
| YDR096W | 3.23 |
GIS1
|
JmjC domain-containing histone demethylase; transcription factor involved in the expression of genes during nutrient limitation; also involved in the negative regulation of DPP1 and PHR1 |
|
| YOR343C | 3.23 |
Dubious open reading frame, unlikely to encode a functional protein; based on available experimental and comparative sequence data |
||
| YIL155C | 3.12 |
GUT2
|
Mitochondrial glycerol-3-phosphate dehydrogenase; expression is repressed by both glucose and cAMP and derepressed by non-fermentable carbon sources in a Snf1p, Rsf1p, Hap2/3/4/5 complex dependent manner |
|
| YDL194W | 3.12 |
SNF3
|
Plasma membrane glucose sensor that regulates glucose transport; has 12 predicted transmembrane segments; long cytoplasmic C-terminal tail is required for low glucose induction of hexose transporter genes HXT2 and HXT4 |
|
| YML089C | 3.12 |
Dubious open reading frame unlikely to encode a functional protein, based on available experimental and comparative sequence data; expression induced by calcium shortage |
||
| YHR092C | 3.10 |
HXT4
|
High-affinity glucose transporter of the major facilitator superfamily, expression is induced by low levels of glucose and repressed by high levels of glucose |
|
| YKL043W | 3.06 |
PHD1
|
Transcriptional activator that enhances pseudohyphal growth; regulates expression of FLO11, an adhesin required for pseudohyphal filament formation; similar to StuA, an A. nidulans developmental regulator; potential Cdc28p substrate |
|
| YPR030W | 3.04 |
CSR2
|
Nuclear protein with a potential regulatory role in utilization of galactose and nonfermentable carbon sources; overproduction suppresses the lethality at high temperature of a chs5 spa2 double null mutation; potential Cdc28p substrate |
|
| YFL011W | 3.03 |
HXT10
|
Putative hexose transporter, expressed at low levels and expression is repressed by glucose |
|
| YPL119C | 2.99 |
DBP1
|
Putative ATP-dependent RNA helicase of the DEAD-box protein family; mutants show reduced stability of the 40S ribosomal subunit scanning through 5' untranslated regions of mRNAs |
|
| YMR272C | 2.86 |
SCS7
|
Sphingolipid alpha-hydroxylase, functions in the alpha-hydroxylation of sphingolipid-associated very long chain fatty acids, has both cytochrome b5-like and hydroxylase/desaturase domains, not essential for growth |
|
| YOL084W | 2.79 |
PHM7
|
Protein of unknown function, expression is regulated by phosphate levels; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery and vacuole |
|
| YMR195W | 2.73 |
ICY1
|
Protein of unknown function, required for viability in rich media of cells lacking mitochondrial DNA; mutants have an invasive growth defect with elongated morphology; induced by amino acid starvation |
|
| YPL061W | 2.72 |
ALD6
|
Cytosolic aldehyde dehydrogenase, activated by Mg2+ and utilizes NADP+ as the preferred coenzyme; required for conversion of acetaldehyde to acetate; constitutively expressed; locates to the mitochondrial outer surface upon oxidative stress |
|
| YBR033W | 2.71 |
EDS1
|
Putative zinc cluster protein; YBR033W is not an essential gene |
|
| YHL032C | 2.71 |
GUT1
|
Glycerol kinase, converts glycerol to glycerol-3-phosphate; glucose repression of expression is mediated by Adr1p and Ino2p-Ino4p; derepression of expression on non-fermentable carbon sources is mediated by Opi1p and Rsf1p |
|
| YDR010C | 2.70 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data |
||
| YER084W | 2.68 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data |
||
| YCR021C | 2.67 |
HSP30
|
Hydrophobic plasma membrane localized, stress-responsive protein that negatively regulates the H(+)-ATPase Pma1p; induced by heat shock, ethanol treatment, weak organic acid, glucose limitation, and entry into stationary phase |
|
| YAR053W | 2.57 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data |
||
| YNL117W | 2.55 |
MLS1
|
Malate synthase, enzyme of the glyoxylate cycle, involved in utilization of non-fermentable carbon sources; expression is subject to carbon catabolite repression; localizes in peroxisomes during growth in oleic acid medium |
|
| YOL051W | 2.51 |
GAL11
|
Subunit of the RNA polymerase II mediator complex; associates with core polymerase subunits to form the RNA polymerase II holoenzyme; affects transcription by acting as target of activators and repressors |
|
| YNL144C | 2.49 |
Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; YNL144C is not an essential gene |
||
| YHR218W | 2.49 |
Helicase-like protein encoded within the telomeric Y' element |
||
| YMR017W | 2.45 |
SPO20
|
Meiosis-specific subunit of the t-SNARE complex, required for prospore membrane formation during sporulation; similar to but not functionally redundant with Sec9p; SNAP-25 homolog |
|
| YDR216W | 2.45 |
ADR1
|
Carbon source-responsive zinc-finger transcription factor, required for transcription of the glucose-repressed gene ADH2, of peroxisomal protein genes, and of genes required for ethanol, glycerol, and fatty acid utilization |
|
| YIL055C | 2.44 |
Putative protein of unknown function |
||
| YPR005C | 2.38 |
HAL1
|
Cytoplasmic protein involved in halotolerance; decreases intracellular Na+ (via Ena1p) and increases intracellular K+ by decreasing efflux; expression repressed by Ssn6p-Tup1p and Sko1p and induced by NaCl, KCl, and sorbitol through Gcn4p |
|
| YOL154W | 2.36 |
ZPS1
|
Putative GPI-anchored protein; transcription is induced under low-zinc conditions, as mediated by the Zap1p transcription factor, and at alkaline pH |
|
| YPL262W | 2.35 |
FUM1
|
Fumarase, converts fumaric acid to L-malic acid in the TCA cycle; cytosolic and mitochondrial localization determined by the N-terminal mitochondrial targeting sequence and protein conformation; phosphorylated in mitochondria |
|
| YLL053C | 2.33 |
Putative protein; in the Sigma 1278B strain background YLL053C is contiguous with AQY2 which encodes an aquaporin |
||
| YDR178W | 2.33 |
SDH4
|
Membrane anchor subunit of succinate dehydrogenase (Sdh1p, Sdh2p, Sdh3p, Sdh4p), which couples the oxidation of succinate to the transfer of electrons to ubiquinone |
|
| YJL103C | 2.28 |
GSM1
|
Putative zinc cluster protein of unknown function; proposed to be involved in the regulation of energy metabolism, based on patterns of expression and sequence analysis |
|
| YPR026W | 2.25 |
ATH1
|
Acid trehalase required for utilization of extracellular trehalose |
|
| YIL136W | 2.25 |
OM45
|
Protein of unknown function, major constituent of the mitochondrial outer membrane; located on the outer (cytosolic) face of the outer membrane |
|
| YJL199C | 2.24 |
MBB1
|
Dubious open reading frame, unlikely to encode a protein; not conserved in closely related Saccharomyces species; protein detected in large-scale protein-protein interaction studies |
|
| YHL021C | 2.23 |
AIM17
|
Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; null mutant displays decreased frequency of mitochondrial genome loss (petite formation) |
|
| YDL079C | 2.22 |
MRK1
|
Glycogen synthase kinase 3 (GSK-3) homolog; one of four GSK-3 homologs in S. cerevisiae that function to activate Msn2p-dependent transcription of stress responsive genes and that function in protein degradation |
|
| YBR051W | 2.22 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; partiallly overlaps the REG2/YBR050C regulatory subunit of the Glc7p type-1 protein phosphatase |
||
| YER150W | 2.21 |
SPI1
|
GPI-anchored cell wall protein involved in weak acid resistance; basal expression requires Msn2p/Msn4p; expression is induced under conditions of stress and during the diauxic shift; similar to Sed1p |
|
| YNL036W | 2.21 |
NCE103
|
Carbonic anhydrase; poorly transcribed under aerobic conditions and at an undetectable level under anaerobic conditions; involved in non-classical protein export pathway |
|
| YBR117C | 2.20 |
TKL2
|
Transketolase, similar to Tkl1p; catalyzes conversion of xylulose-5-phosphate and ribose-5-phosphate to sedoheptulose-7-phosphate and glyceraldehyde-3-phosphate in the pentose phosphate pathway; needed for synthesis of aromatic amino acids |
|
| YPL186C | 2.20 |
UIP4
|
Protein that interacts with Ulp1p, a Ubl (ubiquitin-like protein)-specific protease for Smt3p protein conjugates; detected in a phosphorylated state in the mitochondrial outer membrane; also detected in ER and nuclear envelope |
|
| YAR035W | 2.20 |
YAT1
|
Outer mitochondrial carnitine acetyltransferase, minor ethanol-inducible enzyme involved in transport of activated acyl groups from the cytoplasm into the mitochondrial matrix; phosphorylated |
|
| YOL060C | 2.19 |
MAM3
|
Protein required for normal mitochondrial morphology, has similarity to hemolysins |
|
| YLR174W | 2.18 |
IDP2
|
Cytosolic NADP-specific isocitrate dehydrogenase, catalyzes oxidation of isocitrate to alpha-ketoglutarate; levels are elevated during growth on non-fermentable carbon sources and reduced during growth on glucose |
|
| YKL093W | 2.17 |
MBR1
|
Protein involved in mitochondrial functions and stress response; overexpression suppresses growth defects of hap2, hap3, and hap4 mutants |
|
| YMR280C | 2.17 |
CAT8
|
Zinc cluster transcriptional activator necessary for derepression of a variety of genes under non-fermentative growth conditions, active after diauxic shift, binds carbon source responsive elements |
|
| YCR007C | 2.17 |
Putative integral membrane protein, member of DUP240 gene family; YCR007C is not an essential gene |
||
| YAL039C | 2.15 |
CYC3
|
Cytochrome c heme lyase (holocytochrome c synthase), attaches heme to apo-cytochrome c (Cyc1p or Cyc7p) in the mitochondrial intermembrane space; human ortholog may have a role in microphthalmia with linear skin defects (MLS) |
|
| YBR050C | 2.15 |
REG2
|
Regulatory subunit of the Glc7p type-1 protein phosphatase; involved with Reg1p, Glc7p, and Snf1p in regulation of glucose-repressible genes, also involved in glucose-induced proteolysis of maltose permease |
|
| YGL146C | 2.15 |
Putative protein of unknown function, contains two putative transmembrane spans, shows no significant homology to any other known protein sequence, YGL146C is not an essential gene |
||
| YPL024W | 2.14 |
RMI1
|
Involved in response to DNA damage; null mutants have increased rates of recombination and delayed S phase; interacts physically and genetically with Sgs1p (RecQ family member) and Top3p (topoisomerase III) |
|
| YEL012W | 2.14 |
UBC8
|
Ubiquitin-conjugating enzyme that negatively regulates gluconeogenesis by mediating the glucose-induced ubiquitination of fructose-1,6-bisphosphatase (FBPase); cytoplasmic enzyme that catalyzes the ubiquitination of histones in vitro |
|
| YPL201C | 2.12 |
YIG1
|
Protein that interacts with glycerol 3-phosphatase and plays a role in anaerobic glycerol production; localizes to the nucleus and cytosol |
|
| YFL064C | 2.09 |
Putative protein of unknown function |
||
| YLL052C | 2.09 |
AQY2
|
Water channel that mediates the transport of water across cell membranes, only expressed in proliferating cells, controlled by osmotic signals, may be involved in freeze tolerance; disrupted by a stop codon in many S. cerevisiae strains |
|
| YER121W | 2.08 |
Putative protein of unknown function; may be involved in phosphatase regulation and/or generation of precursor metabolites and energy |
||
| YKL031W | 2.08 |
Dubious open reading frame, unlikely to encode a protein; not conserved in closely related Saccharomyces species |
||
| YPR015C | 2.08 |
Putative protein of unknown function |
||
| YLR438W | 2.07 |
CAR2
|
L-ornithine transaminase (OTAse), catalyzes the second step of arginine degradation, expression is dually-regulated by allophanate induction and a specific arginine induction process; not nitrogen catabolite repression sensitive |
|
| YFL063W | 2.07 |
Dubious open reading frame, based on available experimental and comparative sequence data |
||
| YEL070W | 2.05 |
DSF1
|
Deletion suppressor of mpt5 mutation |
|
| YGL045W | 2.05 |
RIM8
|
Protein of unknown function, involved in the proteolytic activation of Rim101p in response to alkaline pH; has similarity to A. nidulans PalF |
|
| YPR151C | 2.05 |
SUE1
|
Mitochondrial protein required for degradation of unstable forms of cytochrome c |
|
| YBR298C | 2.01 |
MAL31
|
Maltose permease, high-affinity maltose transporter (alpha-glucoside transporter); encoded in the MAL3 complex locus; member of the 12 transmembrane domain superfamily of sugar transporters; functional in genomic reference strain S288C |
|
| YHR211W | 2.01 |
FLO5
|
Lectin-like cell wall protein (flocculin) involved in flocculation, binds to mannose chains on the surface of other cells, confers floc-forming ability that is chymotrypsin resistant but heat labile; similar to Flo1p |
|
| YLR122C | 2.00 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; partially overlaps the dubious ORF YLR123C |
||
| YMR174C | 1.98 |
PAI3
|
Cytoplasmic proteinase A (Pep4p) inhibitor, dependent on Pbs2p and Hog1p protein kinases for osmotic induction; intrinsically unstructured, N-terminal half becomes ordered in the active site of proteinase A upon contact |
|
| YBR203W | 1.97 |
COS111
|
Protein required for resistance to the antifungal drug ciclopirox olamine; not related to the subtelomerically-encoded COS family; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies |
|
| YDL085W | 1.97 |
NDE2
|
Mitochondrial external NADH dehydrogenase, catalyzes the oxidation of cytosolic NADH; Nde1p and Nde2p are involved in providing the cytosolic NADH to the mitochondrial respiratory chain |
|
| YOR031W | 1.97 |
CRS5
|
Copper-binding metallothionein, required for wild-type copper resistance |
|
| YER187W | 1.97 |
Putative protein of unknown function; induced in respiratory-deficient cells |
||
| YOR376W | 1.97 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; YOR376W is not an essential gene. |
||
| YML090W | 1.96 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; partially overlaps the dubious ORF YML089C; exhibits growth defect on a non-fermentable (respiratory) carbon source |
||
| YDR186C | 1.96 |
Putative protein of unknown function; may interact with ribosomes, based on co-purification experiments; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm |
||
| YJL133C-A | 1.96 |
Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies |
||
| YBR132C | 1.95 |
AGP2
|
High affinity polyamine permease, preferentially uses spermidine over putrescine; expression is down-regulated by osmotic stress; plasma membrane carnitine transporter, also functions as a low-affinity amino acid permease |
|
| YLR332W | 1.93 |
MID2
|
O-glycosylated plasma membrane protein that acts as a sensor for cell wall integrity signaling and activates the pathway; interacts with Rom2p, a guanine nucleotide exchange factor for Rho1p, and with cell integrity pathway protein Zeo1p |
|
| YMR105C | 1.92 |
PGM2
|
Phosphoglucomutase, catalyzes the conversion from glucose-1-phosphate to glucose-6-phosphate, which is a key step in hexose metabolism; functions as the acceptor for a Glc-phosphotransferase |
|
| YAR060C | 1.92 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data |
||
| YNL142W | 1.91 |
MEP2
|
Ammonium permease involved in regulation of pseudohyphal growth; belongs to a ubiquitous family of cytoplasmic membrane proteins that transport only ammonium (NH4+); expression is under the nitrogen catabolite repression regulation |
|
| YJR096W | 1.91 |
Putative xylose and arabinose reductase; member of the aldo-keto reductase (AKR) family; GFP-fusion protein is induced in response to the DNA-damaging agent MMS |
||
| YGR144W | 1.90 |
THI4
|
Thiazole synthase, catalyzes formation of the thiazole moiety of thiamin pyrophosphate; required for thiamine biosynthesis and for mitochondrial genome stability |
|
| YBR201C-A | 1.89 |
Putative protein of unknown function |
||
| YPL185W | 1.88 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; partially overlaps the verified gene UIP4/YPL186C |
||
| YPR006C | 1.84 |
ICL2
|
2-methylisocitrate lyase of the mitochondrial matrix, functions in the methylcitrate cycle to catalyze the conversion of 2-methylisocitrate to succinate and pyruvate; ICL2 transcription is repressed by glucose and induced by ethanol |
|
| YOL081W | 1.84 |
IRA2
|
GTPase-activating protein that negatively regulates RAS by converting it from the GTP- to the GDP-bound inactive form, required for reducing cAMP levels under nutrient limiting conditions, has similarity to Ira1p and human neurofibromin |
|
| YLR331C | 1.82 |
JIP3
|
Dubious open reading frame, unlikely to encode a protein; not conserved in closely related Saccharomyces species; 98% of ORF overlaps the verified gene MID2 |
|
| YGR023W | 1.80 |
MTL1
|
Protein with both structural and functional similarity to Mid2p, which is a plasma membrane sensor required for cell integrity signaling during pheromone-induced morphogenesis; suppresses rgd1 null mutations |
|
| YDL244W | 1.79 |
THI13
|
Protein involved in synthesis of the thiamine precursor hydroxymethylpyrimidine (HMP); member of a subtelomeric gene family including THI5, THI11, THI12, and THI13 |
|
| YOR065W | 1.78 |
CYT1
|
Cytochrome c1, component of the mitochondrial respiratory chain; expression is regulated by the heme-activated, glucose-repressed Hap2p/3p/4p/5p CCAAT-binding complex |
|
| YCR010C | 1.76 |
ADY2
|
Acetate transporter required for normal sporulation; phosphorylated in mitochondria |
|
| YLR437C-A | 1.76 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; partially overlaps the verified ORF CAR2/YLR438W |
||
| YDR259C | 1.75 |
YAP6
|
Putative basic leucine zipper (bZIP) transcription factor; overexpression increases sodium and lithium tolerance |
|
| YJR151C | 1.74 |
DAN4
|
Cell wall mannoprotein with similarity to Tir1p, Tir2p, Tir3p, and Tir4p; expressed under anaerobic conditions, completely repressed during aerobic growth |
|
| YHR033W | 1.72 |
Putative protein of unknown function; epitope-tagged protein localizes to the cytoplasm |
||
| YPR150W | 1.71 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; partially overlaps the verified gene SUE1/YPR151C |
||
| YDR406W | 1.71 |
PDR15
|
Plasma membrane ATP binding cassette (ABC) transporter, multidrug transporter and general stress response factor implicated in cellular detoxification; regulated by Pdr1p, Pdr3p and Pdr8p; promoter contains a PDR responsive element |
|
| YCL025C | 1.70 |
AGP1
|
Low-affinity amino acid permease with broad substrate range, involved in uptake of asparagine, glutamine, and other amino acids; expression is regulated by the SPS plasma membrane amino acid sensor system (Ssy1p-Ptr3p-Ssy5p) |
|
| YGR022C | 1.70 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; overlaps almost completely with the verified ORF MTL1/YGR023W |
||
| YFL062W | 1.69 |
COS4
|
Protein of unknown function, member of the DUP380 subfamily of conserved, often subtelomerically-encoded proteins |
|
| YGR258C | 1.69 |
RAD2
|
Single-stranded DNA endonuclease, cleaves single-stranded DNA during nucleotide excision repair to excise damaged DNA; subunit of Nucleotide Excision Repair Factor 3 (NEF3); homolog of human XPG protein |
|
| YLR258W | 1.68 |
GSY2
|
Glycogen synthase, similar to Gsy1p; expression induced by glucose limitation, nitrogen starvation, heat shock, and stationary phase; activity regulated by cAMP-dependent, Snf1p and Pho85p kinases as well as by the Gac1p-Glc7p phosphatase |
|
| YIL099W | 1.68 |
SGA1
|
Intracellular sporulation-specific glucoamylase involved in glycogen degradation; induced during starvation of a/a diploids late in sporulation, but dispensable for sporulation |
|
| YAL061W | 1.66 |
BDH2
|
Putative medium-chain alcohol dehydrogenase with similarity to BDH1; transcription induced by constitutively active PDR1 and PDR3; BDH2 is an essential gene |
|
| YDR247W | 1.64 |
VHS1
|
Cytoplasmic serine/threonine protein kinase; identified as a high-copy suppressor of the synthetic lethality of a sis2 sit4 double mutant, suggesting a role in G1/S phase progression; homolog of Sks1p |
|
| YLR123C | 1.64 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; partially overlaps the dubious ORF YLR122C; contains characteristic aminoacyl-tRNA motif |
||
| YPL026C | 1.63 |
SKS1
|
Putative serine/threonine protein kinase; involved in the adaptation to low concentrations of glucose independent of the SNF3 regulated pathway |
|
| YKL085W | 1.62 |
MDH1
|
Mitochondrial malate dehydrogenase, catalyzes interconversion of malate and oxaloacetate; involved in the tricarboxylic acid (TCA) cycle; phosphorylated |
|
| YBR047W | 1.60 |
FMP23
|
Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies |
|
| YOR328W | 1.59 |
PDR10
|
ATP-binding cassette (ABC) transporter, multidrug transporter involved in the pleiotropic drug resistance network; regulated by Pdr1p and Pdr3p |
|
| YJL102W | 1.59 |
MEF2
|
Mitochondrial elongation factor involved in translational elongation |
|
| YPL057C | 1.58 |
SUR1
|
Probable catalytic subunit of a mannosylinositol phosphorylceramide (MIPC) synthase, forms a complex with probable regulatory subunit Csg2p; function in sphingolipid biosynthesis is overlapping with that of Csh1p |
|
| YOR235W | 1.58 |
IRC13
|
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; null mutant displays increased levels of spontaneous Rad52 foci |
|
| YDL223C | 1.56 |
HBT1
|
Substrate of the Hub1p ubiquitin-like protein that localizes to the shmoo tip (mating projection); mutants are defective for mating projection formation, thereby implicating Hbt1p in polarized cell morphogenesis |
|
| YAL062W | 1.55 |
GDH3
|
NADP(+)-dependent glutamate dehydrogenase, synthesizes glutamate from ammonia and alpha-ketoglutarate; rate of alpha-ketoglutarate utilization differs from Gdh1p; expression regulated by nitrogen and carbon sources |
|
| YLR149C | 1.55 |
Putative protein of unknown function; YLR149C is not an essential gene |
||
| YPR160W | 1.54 |
GPH1
|
Non-essential glycogen phosphorylase required for the mobilization of glycogen, activity is regulated by cyclic AMP-mediated phosphorylation, expression is regulated by stress-response elements and by the HOG MAP kinase pathway |
|
| YGR256W | 1.52 |
GND2
|
6-phosphogluconate dehydrogenase (decarboxylating), catalyzes an NADPH regenerating reaction in the pentose phosphate pathway; required for growth on D-glucono-delta-lactone |
|
| YJL220W | 1.52 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; partially overlaps the verified gene YJL221C/FSP2 |
||
| YML128C | 1.50 |
MSC1
|
Protein of unknown function; mutant is defective in directing meiotic recombination events to homologous chromatids; the authentic, non-tagged protein is detected in highly purified mitochondria and is phosphorylated |
|
| YGL163C | 1.49 |
RAD54
|
DNA-dependent ATPase, stimulates strand exchange by modifying the topology of double-stranded DNA; involved in the recombinational repair of double-strand breaks in DNA during vegetative growth and meiosis; member of the SWI/SNF family |
|
| YLR279W | 1.49 |
Dubious open reading frame unlikely to encode a functional protein, based on available experimental and comparative sequence data |
||
| YML091C | 1.49 |
RPM2
|
Protein subunit of mitochondrial RNase P, has roles in nuclear transcription, cytoplasmic and mitochondrial RNA processing, and mitochondrial translation; distributed to mitochondria, cytoplasmic processing bodies, and the nucleus |
|
| YML131W | 1.48 |
Putative protein of unknown function with similarity to oxidoreductases; HOG1 and SKO1-dependent mRNA expression is induced after osmotic shock; GFP-fusion protein localizes to the cytoplasm and is induced by the DNA-damaging agent MMS |
||
| YDL214C | 1.48 |
PRR2
|
Serine/threonine protein kinase that inhibits pheromone induced signalling downstream of MAPK, possibly at the level of the Ste12p transcription factor |
|
| YBL033C | 1.48 |
RIB1
|
GTP cyclohydrolase II; catalyzes the first step of the riboflavin biosynthesis pathway |
|
| YGL117W | 1.48 |
Putative protein of unknown function |
||
| YPR154W | 1.47 |
PIN3
|
Protein that induces appearance of [PIN+] prion when overproduced |
|
| YHR198C | 1.46 |
AIM18
|
Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; null mutant displays increased frequency of mitochondrial genome loss (petite formation) |
|
| YOR211C | 1.44 |
MGM1
|
Mitochondrial GTPase related to dynamin, present in a complex containing Ugo1p and Fzo1p; required for normal morphology of cristae and for stability of Tim11p; homolog of human OPA1 involved in autosomal dominant optic atrophy |
|
| YPR036W-A | 1.44 |
Protein of unknown function; transcription is regulated by Pdr1p |
||
| YKL141W | 1.44 |
SDH3
|
Cytochrome b subunit of succinate dehydrogenase (Sdh1p, Sdh2p, Sdh3p, Sdh4p), which couples the oxidation of succinate to the transfer of electrons to ubiquinone |
|
| YIL045W | 1.43 |
PIG2
|
Putative type-1 protein phosphatase targeting subunit that tethers Glc7p type-1 protein phosphatase to Gsy2p glycogen synthase |
|
| YPL017C | 1.42 |
IRC15
|
Putative S-adenosylmethionine-dependent methyltransferase of the seven beta-strand family, required for accurate meiotic chromosome segregation; null mutant displays increased levels of spontaneous Rad52 foci |
|
| YHR212C | 1.42 |
Dubious open reading frame unlikely to encode a functional protein, based on available experimental and comparative sequence data |
||
| YGL096W | 1.42 |
TOS8
|
Homeodomain-containing transcription factor; SBF regulated target gene that in turn regulates expression of genes involved in G1/S phase events such as bud site selection, bud emergence and cell cycle progression; similarity to Cup9p |
|
| YJR120W | 1.41 |
Protein of unknown function; essential for growth under anaerobic conditions; mutation causes decreased expression of ATP2, impaired respiration, defective sterol uptake, and altered levels/localization of ABC transporters Aus1p and Pdr11p |
||
| YOR152C | 1.41 |
Putative protein of unknown function; has no similarity to any known protein; YOR152C is not an essential gene |
||
| YAL034C | 1.40 |
FUN19
|
Non-essential protein of unknown function |
|
| YCR079W | 1.40 |
PTC6
|
Mitochondrial protein phosphatase of type 2C with similarity to mammalian PP1Ks; involved in mitophagy; null mutant is sensitive to rapamycin and has decreased phosphorylation of the Pda1 subunit of pyruvate dehydrogenase |
|
| YDR306C | 1.39 |
F-box protein of unknown function; interacts with Sgt1p via a Leucine-Rich Repeat (LRR) domain |
||
| YPL250C | 1.39 |
ICY2
|
Protein of unknown function; mobilized into polysomes upon a shift from a fermentable to nonfermentable carbon source; potential Cdc28p substrate |
|
| YPR013C | 1.39 |
Putative zinc finger protein; YPR013C is not an essential gene |
||
| YER067W | 1.38 |
Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; YER067W is not an essential gene |
Network of associatons between targets according to the STRING database.
First level regulatory network of ZMS1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.5 | 18.0 | GO:0006848 | pyruvate transport(GO:0006848) |
| 3.7 | 11.0 | GO:0015755 | fructose transport(GO:0015755) |
| 2.7 | 8.2 | GO:0006577 | amino-acid betaine metabolic process(GO:0006577) carnitine metabolic process(GO:0009437) |
| 2.4 | 2.4 | GO:0034310 | ethanol catabolic process(GO:0006068) primary alcohol catabolic process(GO:0034310) |
| 2.2 | 6.6 | GO:0019413 | acetate biosynthetic process(GO:0019413) |
| 2.0 | 7.9 | GO:0015804 | neutral amino acid transport(GO:0015804) |
| 1.7 | 5.1 | GO:0015740 | C4-dicarboxylate transport(GO:0015740) |
| 1.7 | 6.6 | GO:0019405 | alditol catabolic process(GO:0019405) glycerol catabolic process(GO:0019563) |
| 1.5 | 7.4 | GO:0015793 | glycerol transport(GO:0015793) |
| 1.5 | 14.5 | GO:0006122 | mitochondrial electron transport, ubiquinol to cytochrome c(GO:0006122) |
| 1.4 | 4.2 | GO:0042843 | D-xylose catabolic process(GO:0042843) |
| 1.4 | 6.9 | GO:0005980 | glycogen catabolic process(GO:0005980) |
| 1.4 | 4.1 | GO:0032780 | negative regulation of ATPase activity(GO:0032780) |
| 1.3 | 10.7 | GO:0015891 | siderophore transport(GO:0015891) |
| 1.3 | 5.3 | GO:0006075 | (1->3)-beta-D-glucan biosynthetic process(GO:0006075) |
| 1.3 | 27.5 | GO:0015749 | hexose transport(GO:0008645) monosaccharide transport(GO:0015749) |
| 1.2 | 3.7 | GO:0061416 | regulation of transcription from RNA polymerase II promoter in response to salt stress(GO:0061416) |
| 1.2 | 3.6 | GO:0051469 | vesicle fusion with vacuole(GO:0051469) |
| 1.1 | 3.3 | GO:0019541 | propionate metabolic process(GO:0019541) propionate catabolic process(GO:0019543) short-chain fatty acid catabolic process(GO:0019626) propionate catabolic process, 2-methylcitrate cycle(GO:0019629) |
| 1.1 | 3.2 | GO:0072488 | ammonium transmembrane transport(GO:0072488) |
| 1.1 | 18.0 | GO:0005978 | glycogen biosynthetic process(GO:0005978) |
| 1.0 | 3.1 | GO:0052547 | negative regulation of peptidase activity(GO:0010466) regulation of peptidase activity(GO:0052547) |
| 1.0 | 3.1 | GO:0005993 | trehalose catabolic process(GO:0005993) |
| 1.0 | 8.7 | GO:0006097 | glyoxylate cycle(GO:0006097) |
| 0.9 | 6.6 | GO:0000023 | maltose metabolic process(GO:0000023) |
| 0.9 | 1.8 | GO:0045980 | negative regulation of nucleotide metabolic process(GO:0045980) |
| 0.9 | 3.5 | GO:0006482 | protein demethylation(GO:0006482) protein dealkylation(GO:0008214) histone demethylation(GO:0016577) demethylation(GO:0070988) |
| 0.9 | 3.4 | GO:0006527 | arginine catabolic process(GO:0006527) |
| 0.8 | 4.9 | GO:0000501 | flocculation(GO:0000128) flocculation via cell wall protein-carbohydrate interaction(GO:0000501) |
| 0.8 | 3.2 | GO:0009415 | response to water(GO:0009415) cellular response to water stimulus(GO:0071462) |
| 0.8 | 1.6 | GO:0006108 | malate metabolic process(GO:0006108) |
| 0.8 | 3.1 | GO:0071467 | cellular response to pH(GO:0071467) |
| 0.8 | 0.8 | GO:0046487 | glyoxylate metabolic process(GO:0046487) |
| 0.8 | 3.8 | GO:0006121 | mitochondrial electron transport, succinate to ubiquinone(GO:0006121) |
| 0.7 | 2.2 | GO:0017006 | protein-heme linkage(GO:0017003) protein-tetrapyrrole linkage(GO:0017006) cytochrome c-heme linkage(GO:0018063) |
| 0.6 | 3.2 | GO:0006673 | inositolphosphoceramide metabolic process(GO:0006673) |
| 0.6 | 2.5 | GO:0051101 | positive regulation of DNA binding(GO:0043388) regulation of DNA binding(GO:0051101) |
| 0.6 | 0.6 | GO:0071456 | cellular response to hypoxia(GO:0071456) |
| 0.6 | 1.2 | GO:0006102 | isocitrate metabolic process(GO:0006102) |
| 0.6 | 8.7 | GO:0051156 | glucose 6-phosphate metabolic process(GO:0051156) |
| 0.6 | 2.4 | GO:0015976 | carbon utilization(GO:0015976) |
| 0.6 | 17.6 | GO:0006094 | gluconeogenesis(GO:0006094) |
| 0.6 | 3.5 | GO:0005987 | sucrose metabolic process(GO:0005985) sucrose catabolic process(GO:0005987) |
| 0.6 | 2.3 | GO:0006083 | acetate metabolic process(GO:0006083) |
| 0.5 | 1.6 | GO:0033499 | galactose catabolic process via UDP-galactose(GO:0033499) |
| 0.5 | 1.0 | GO:0097201 | negative regulation of transcription from RNA polymerase II promoter in response to stress(GO:0097201) |
| 0.5 | 5.4 | GO:0000436 | carbon catabolite activation of transcription from RNA polymerase II promoter(GO:0000436) |
| 0.5 | 1.9 | GO:0070054 | mRNA splicing via endonucleolytic cleavage and ligation involved in unfolded protein response(GO:0030969) IRE1-mediated unfolded protein response(GO:0036498) mRNA splicing, via endonucleolytic cleavage and ligation(GO:0070054) |
| 0.5 | 1.8 | GO:0006103 | 2-oxoglutarate metabolic process(GO:0006103) |
| 0.4 | 3.6 | GO:0070086 | ubiquitin-dependent endocytosis(GO:0070086) |
| 0.4 | 0.4 | GO:0043902 | positive regulation of multi-organism process(GO:0043902) |
| 0.4 | 1.7 | GO:0032079 | positive regulation of deoxyribonuclease activity(GO:0032077) positive regulation of endodeoxyribonuclease activity(GO:0032079) |
| 0.4 | 1.7 | GO:0010688 | negative regulation of ribosomal protein gene transcription from RNA polymerase II promoter(GO:0010688) |
| 0.4 | 1.2 | GO:0001324 | age-dependent general metabolic decline involved in chronological cell aging(GO:0001323) age-dependent response to oxidative stress involved in chronological cell aging(GO:0001324) |
| 0.4 | 1.6 | GO:0045141 | meiotic telomere clustering(GO:0045141) establishment of chromosome localization(GO:0051303) chromosome localization to nuclear envelope involved in homologous chromosome segregation(GO:0090220) |
| 0.4 | 1.9 | GO:0000255 | allantoin metabolic process(GO:0000255) allantoin catabolic process(GO:0000256) |
| 0.4 | 3.8 | GO:0055069 | cellular zinc ion homeostasis(GO:0006882) zinc ion homeostasis(GO:0055069) |
| 0.4 | 0.4 | GO:0032070 | regulation of deoxyribonuclease activity(GO:0032070) regulation of endodeoxyribonuclease activity(GO:0032071) |
| 0.4 | 1.1 | GO:0042743 | hydrogen peroxide metabolic process(GO:0042743) hydrogen peroxide catabolic process(GO:0042744) |
| 0.4 | 1.1 | GO:0006390 | transcription from mitochondrial promoter(GO:0006390) |
| 0.4 | 1.1 | GO:0070199 | establishment of protein localization to chromosome(GO:0070199) establishment of protein localization to chromatin(GO:0071169) |
| 0.3 | 1.4 | GO:0034316 | negative regulation of Arp2/3 complex-mediated actin nucleation(GO:0034316) negative regulation of actin nucleation(GO:0051126) |
| 0.3 | 1.4 | GO:0015855 | pyrimidine nucleobase transport(GO:0015855) |
| 0.3 | 1.4 | GO:0031138 | negative regulation of conjugation(GO:0031135) negative regulation of conjugation with cellular fusion(GO:0031138) negative regulation of multi-organism process(GO:0043901) |
| 0.3 | 5.4 | GO:0019740 | nitrogen utilization(GO:0019740) |
| 0.3 | 1.7 | GO:0033683 | nucleotide-excision repair, DNA incision(GO:0033683) |
| 0.3 | 0.6 | GO:0009190 | cAMP biosynthetic process(GO:0006171) cyclic nucleotide biosynthetic process(GO:0009190) cyclic purine nucleotide metabolic process(GO:0052652) |
| 0.3 | 0.9 | GO:0006545 | glycine biosynthetic process(GO:0006545) |
| 0.3 | 0.8 | GO:0006598 | polyamine catabolic process(GO:0006598) |
| 0.3 | 1.4 | GO:0008053 | mitochondrial fusion(GO:0008053) |
| 0.3 | 0.8 | GO:0019748 | secondary metabolic process(GO:0019748) |
| 0.3 | 1.8 | GO:0030026 | cellular manganese ion homeostasis(GO:0030026) manganese ion homeostasis(GO:0055071) |
| 0.3 | 1.0 | GO:0007039 | protein catabolic process in the vacuole(GO:0007039) |
| 0.2 | 4.2 | GO:0009228 | thiamine biosynthetic process(GO:0009228) thiamine-containing compound biosynthetic process(GO:0042724) |
| 0.2 | 0.7 | GO:0008216 | spermidine metabolic process(GO:0008216) |
| 0.2 | 0.7 | GO:0043068 | positive regulation of cell death(GO:0010942) positive regulation of apoptotic process(GO:0043065) positive regulation of programmed cell death(GO:0043068) |
| 0.2 | 0.9 | GO:0006112 | glycogen metabolic process(GO:0005977) energy reserve metabolic process(GO:0006112) |
| 0.2 | 3.7 | GO:0006754 | ATP biosynthetic process(GO:0006754) |
| 0.2 | 3.2 | GO:0015718 | monocarboxylic acid transport(GO:0015718) |
| 0.2 | 1.4 | GO:0006855 | drug transmembrane transport(GO:0006855) |
| 0.2 | 1.8 | GO:0000321 | re-entry into mitotic cell cycle after pheromone arrest(GO:0000321) |
| 0.2 | 0.9 | GO:0006285 | base-excision repair, AP site formation(GO:0006285) |
| 0.2 | 0.7 | GO:2000284 | positive regulation of cellular amino acid biosynthetic process(GO:2000284) |
| 0.2 | 0.6 | GO:0044209 | AMP salvage(GO:0044209) |
| 0.2 | 0.2 | GO:0046185 | aldehyde catabolic process(GO:0046185) |
| 0.2 | 1.0 | GO:0031507 | heterochromatin assembly(GO:0031507) |
| 0.2 | 1.2 | GO:0006099 | tricarboxylic acid cycle(GO:0006099) citrate metabolic process(GO:0006101) |
| 0.2 | 0.6 | GO:1903137 | inactivation of MAPK activity involved in cell wall organization or biogenesis(GO:0000200) regulation of MAPK cascade involved in cell wall organization or biogenesis(GO:1903137) negative regulation of MAPK cascade involved in cell wall organization or biogenesis(GO:1903138) negative regulation of cell wall organization or biogenesis(GO:1903339) |
| 0.2 | 3.3 | GO:0097034 | mitochondrial respiratory chain complex IV biogenesis(GO:0097034) |
| 0.2 | 4.7 | GO:0000422 | mitophagy(GO:0000422) mitochondrion disassembly(GO:0061726) |
| 0.2 | 0.6 | GO:0034090 | maintenance of meiotic sister chromatid cohesion(GO:0034090) |
| 0.2 | 0.8 | GO:0016024 | CDP-diacylglycerol biosynthetic process(GO:0016024) CDP-diacylglycerol metabolic process(GO:0046341) |
| 0.2 | 7.1 | GO:0009060 | aerobic respiration(GO:0009060) |
| 0.2 | 1.1 | GO:0071322 | cellular response to carbohydrate stimulus(GO:0071322) |
| 0.2 | 1.8 | GO:0007135 | meiosis II(GO:0007135) meiotic sister chromatid segregation(GO:0045144) |
| 0.2 | 0.2 | GO:0051098 | regulation of binding(GO:0051098) |
| 0.2 | 0.4 | GO:0008272 | sulfate transport(GO:0008272) |
| 0.2 | 0.2 | GO:0042542 | response to hydrogen peroxide(GO:0042542) |
| 0.2 | 0.9 | GO:0044070 | regulation of anion transport(GO:0044070) |
| 0.2 | 0.5 | GO:0001113 | transcriptional open complex formation at RNA polymerase II promoter(GO:0001113) |
| 0.2 | 0.5 | GO:0055078 | cellular sodium ion homeostasis(GO:0006883) sodium ion homeostasis(GO:0055078) |
| 0.2 | 0.6 | GO:0032102 | negative regulation of macroautophagy(GO:0016242) negative regulation of response to external stimulus(GO:0032102) negative regulation of response to extracellular stimulus(GO:0032105) negative regulation of response to nutrient levels(GO:0032108) |
| 0.2 | 0.8 | GO:0000722 | telomere maintenance via recombination(GO:0000722) |
| 0.2 | 1.5 | GO:0046686 | response to cadmium ion(GO:0046686) |
| 0.2 | 0.6 | GO:0072367 | positive regulation of lipid transport(GO:0032370) regulation of sterol transport(GO:0032371) positive regulation of sterol transport(GO:0032373) positive regulation of transmembrane transport(GO:0034764) regulation of sterol import by regulation of transcription from RNA polymerase II promoter(GO:0035968) positive regulation of sterol import by positive regulation of transcription from RNA polymerase II promoter(GO:0035969) regulation of lipid transport by regulation of transcription from RNA polymerase II promoter(GO:0072367) regulation of lipid transport by positive regulation of transcription from RNA polymerase II promoter(GO:0072369) regulation of sterol import(GO:2000909) positive regulation of sterol import(GO:2000911) |
| 0.2 | 0.2 | GO:0034497 | protein localization to pre-autophagosomal structure(GO:0034497) |
| 0.2 | 0.3 | GO:0000304 | response to singlet oxygen(GO:0000304) |
| 0.2 | 0.6 | GO:0000715 | nucleotide-excision repair, DNA damage recognition(GO:0000715) |
| 0.2 | 0.9 | GO:0006771 | riboflavin metabolic process(GO:0006771) riboflavin biosynthetic process(GO:0009231) |
| 0.2 | 0.9 | GO:0045117 | azole transport(GO:0045117) |
| 0.1 | 0.3 | GO:0071931 | positive regulation of transcription involved in G1/S transition of mitotic cell cycle(GO:0071931) |
| 0.1 | 2.0 | GO:0001101 | response to acid chemical(GO:0001101) |
| 0.1 | 0.4 | GO:0036257 | multivesicular body organization(GO:0036257) |
| 0.1 | 0.3 | GO:0043335 | protein unfolding(GO:0043335) |
| 0.1 | 0.7 | GO:0072348 | sulfur compound transport(GO:0072348) |
| 0.1 | 0.4 | GO:0046466 | sphingolipid catabolic process(GO:0030149) membrane lipid catabolic process(GO:0046466) |
| 0.1 | 0.4 | GO:0070304 | positive regulation of stress-activated MAPK cascade(GO:0032874) positive regulation of stress-activated protein kinase signaling cascade(GO:0070304) |
| 0.1 | 1.0 | GO:0018345 | protein palmitoylation(GO:0018345) |
| 0.1 | 0.1 | GO:2000142 | regulation of transcription initiation from RNA polymerase II promoter(GO:0060260) regulation of DNA-templated transcription, initiation(GO:2000142) |
| 0.1 | 0.6 | GO:0070900 | mitochondrial tRNA modification(GO:0070900) mitochondrial RNA modification(GO:1900864) |
| 0.1 | 0.6 | GO:0018904 | glycerol ether metabolic process(GO:0006662) ether metabolic process(GO:0018904) |
| 0.1 | 2.5 | GO:0016579 | protein deubiquitination(GO:0016579) |
| 0.1 | 0.8 | GO:0034440 | fatty acid beta-oxidation(GO:0006635) fatty acid oxidation(GO:0019395) lipid oxidation(GO:0034440) |
| 0.1 | 0.1 | GO:2000219 | positive regulation of invasive growth in response to glucose limitation by positive regulation of transcription from RNA polymerase II promoter(GO:0036095) positive regulation of invasive growth in response to glucose limitation(GO:2000219) |
| 0.1 | 0.2 | GO:0015851 | nucleobase transport(GO:0015851) |
| 0.1 | 0.3 | GO:0010039 | response to iron ion(GO:0010039) regulation of transcription from RNA polymerase II promoter in response to iron(GO:0034395) cellular response to iron ion(GO:0071281) |
| 0.1 | 0.1 | GO:0051222 | positive regulation of protein transport(GO:0051222) positive regulation of establishment of protein localization(GO:1904951) |
| 0.1 | 0.3 | GO:0051058 | negative regulation of Ras protein signal transduction(GO:0046580) negative regulation of small GTPase mediated signal transduction(GO:0051058) |
| 0.1 | 0.5 | GO:0019722 | calcium-mediated signaling(GO:0019722) |
| 0.1 | 0.8 | GO:0006119 | oxidative phosphorylation(GO:0006119) |
| 0.1 | 5.7 | GO:0007031 | peroxisome organization(GO:0007031) |
| 0.1 | 0.5 | GO:0006829 | zinc II ion transport(GO:0006829) |
| 0.1 | 0.1 | GO:0070726 | cell wall assembly(GO:0070726) |
| 0.1 | 0.4 | GO:0007234 | osmosensory signaling via phosphorelay pathway(GO:0007234) |
| 0.1 | 0.9 | GO:0016575 | histone deacetylation(GO:0016575) |
| 0.1 | 0.2 | GO:0071466 | cellular response to xenobiotic stimulus(GO:0071466) |
| 0.1 | 0.3 | GO:0072414 | cellular response to biotic stimulus(GO:0071216) response to cell cycle checkpoint signaling(GO:0072396) response to mitotic cell cycle checkpoint signaling(GO:0072414) response to spindle checkpoint signaling(GO:0072417) response to mitotic spindle checkpoint signaling(GO:0072476) response to mitotic cell cycle spindle assembly checkpoint signaling(GO:0072479) response to spindle assembly checkpoint signaling(GO:0072485) negative regulation of protein import into nucleus during spindle assembly checkpoint(GO:1901925) |
| 0.1 | 0.3 | GO:0071050 | snoRNA polyadenylation(GO:0071050) |
| 0.1 | 1.1 | GO:0036003 | positive regulation of transcription from RNA polymerase II promoter in response to stress(GO:0036003) |
| 0.1 | 0.5 | GO:0006089 | lactate metabolic process(GO:0006089) |
| 0.1 | 0.2 | GO:0090527 | actin filament reorganization involved in cell cycle(GO:0030037) actin filament reorganization(GO:0090527) |
| 0.1 | 1.2 | GO:0000767 | cell morphogenesis involved in conjugation with cellular fusion(GO:0000753) cell morphogenesis involved in conjugation(GO:0000767) |
| 0.1 | 0.7 | GO:0006616 | SRP-dependent cotranslational protein targeting to membrane, translocation(GO:0006616) |
| 0.1 | 0.6 | GO:0000114 | obsolete regulation of transcription involved in G1 phase of mitotic cell cycle(GO:0000114) |
| 0.1 | 0.5 | GO:0006279 | premeiotic DNA replication(GO:0006279) |
| 0.1 | 1.1 | GO:0031321 | ascospore-type prospore assembly(GO:0031321) |
| 0.1 | 0.4 | GO:0072329 | monocarboxylic acid catabolic process(GO:0072329) |
| 0.1 | 0.4 | GO:0016241 | regulation of macroautophagy(GO:0016241) |
| 0.1 | 1.0 | GO:0046854 | lipid phosphorylation(GO:0046834) phosphatidylinositol phosphorylation(GO:0046854) |
| 0.1 | 2.7 | GO:0051123 | RNA polymerase II transcriptional preinitiation complex assembly(GO:0051123) |
| 0.1 | 0.1 | GO:1905038 | regulation of sphingolipid biosynthetic process(GO:0090153) negative regulation of sphingolipid biosynthetic process(GO:0090155) regulation of membrane lipid metabolic process(GO:1905038) |
| 0.1 | 0.6 | GO:0051181 | cofactor transport(GO:0051181) |
| 0.1 | 0.4 | GO:0000387 | spliceosomal snRNP assembly(GO:0000387) |
| 0.1 | 0.9 | GO:0007129 | synapsis(GO:0007129) |
| 0.1 | 0.3 | GO:0000196 | MAPK cascade involved in cell wall organization or biogenesis(GO:0000196) |
| 0.1 | 1.1 | GO:0007266 | Rho protein signal transduction(GO:0007266) |
| 0.1 | 0.3 | GO:0032979 | protein insertion into membrane from inner side(GO:0032978) protein insertion into mitochondrial membrane from inner side(GO:0032979) |
| 0.1 | 0.1 | GO:0031297 | replication fork processing(GO:0031297) |
| 0.1 | 0.3 | GO:0060237 | regulation of fungal-type cell wall organization(GO:0060237) |
| 0.1 | 0.2 | GO:0097577 | intracellular sequestering of iron ion(GO:0006880) sequestering of metal ion(GO:0051238) sequestering of iron ion(GO:0097577) |
| 0.1 | 0.4 | GO:1901658 | nucleoside catabolic process(GO:0009164) glycosyl compound catabolic process(GO:1901658) |
| 0.1 | 0.2 | GO:0046685 | response to arsenic-containing substance(GO:0046685) |
| 0.1 | 0.2 | GO:0043954 | cellular component maintenance(GO:0043954) |
| 0.1 | 0.2 | GO:0046464 | triglyceride catabolic process(GO:0019433) neutral lipid catabolic process(GO:0046461) acylglycerol catabolic process(GO:0046464) |
| 0.1 | 0.3 | GO:0006573 | valine metabolic process(GO:0006573) valine biosynthetic process(GO:0009099) |
| 0.1 | 3.7 | GO:0048468 | ascospore formation(GO:0030437) cell development(GO:0048468) |
| 0.1 | 0.5 | GO:0019878 | lysine biosynthetic process via aminoadipic acid(GO:0019878) |
| 0.1 | 0.1 | GO:2000058 | regulation of protein ubiquitination involved in ubiquitin-dependent protein catabolic process(GO:2000058) |
| 0.1 | 0.4 | GO:0006620 | posttranslational protein targeting to membrane(GO:0006620) |
| 0.1 | 0.2 | GO:0007534 | gene conversion at mating-type locus(GO:0007534) |
| 0.1 | 0.8 | GO:0022900 | electron transport chain(GO:0022900) |
| 0.1 | 0.3 | GO:0043328 | protein targeting to vacuole involved in ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway(GO:0043328) |
| 0.0 | 2.0 | GO:0035825 | reciprocal meiotic recombination(GO:0007131) reciprocal DNA recombination(GO:0035825) |
| 0.0 | 1.6 | GO:0006631 | fatty acid metabolic process(GO:0006631) |
| 0.0 | 0.8 | GO:0042493 | response to drug(GO:0042493) |
| 0.0 | 0.1 | GO:0006655 | phosphatidylglycerol biosynthetic process(GO:0006655) |
| 0.0 | 0.2 | GO:0006078 | (1->6)-beta-D-glucan biosynthetic process(GO:0006078) |
| 0.0 | 0.2 | GO:0048279 | vesicle fusion with endoplasmic reticulum(GO:0048279) |
| 0.0 | 0.2 | GO:0051292 | pore complex assembly(GO:0046931) nuclear pore complex assembly(GO:0051292) |
| 0.0 | 0.2 | GO:0070058 | tRNA gene clustering(GO:0070058) |
| 0.0 | 0.5 | GO:0016925 | protein sumoylation(GO:0016925) |
| 0.0 | 0.3 | GO:0006312 | mitotic recombination(GO:0006312) |
| 0.0 | 0.5 | GO:0070127 | tRNA aminoacylation for mitochondrial protein translation(GO:0070127) |
| 0.0 | 0.1 | GO:0055088 | lipid homeostasis(GO:0055088) |
| 0.0 | 0.4 | GO:0015802 | basic amino acid transport(GO:0015802) |
| 0.0 | 0.3 | GO:0006826 | iron ion transport(GO:0006826) |
| 0.0 | 0.5 | GO:0006986 | response to unfolded protein(GO:0006986) |
| 0.0 | 0.1 | GO:0017062 | respiratory chain complex III assembly(GO:0017062) mitochondrial respiratory chain complex III assembly(GO:0034551) |
| 0.0 | 0.1 | GO:0031335 | regulation of sulfur amino acid metabolic process(GO:0031335) regulation of sulfur metabolic process(GO:0042762) |
| 0.0 | 0.2 | GO:0031167 | rRNA methylation(GO:0031167) |
| 0.0 | 0.3 | GO:0009092 | homoserine metabolic process(GO:0009092) |
| 0.0 | 0.5 | GO:0034629 | cellular protein complex localization(GO:0034629) |
| 0.0 | 0.2 | GO:0000338 | protein deneddylation(GO:0000338) cullin deneddylation(GO:0010388) |
| 0.0 | 0.5 | GO:0043248 | proteasome assembly(GO:0043248) |
| 0.0 | 0.0 | GO:1900544 | positive regulation of nucleotide metabolic process(GO:0045981) positive regulation of purine nucleotide metabolic process(GO:1900544) |
| 0.0 | 0.0 | GO:0097271 | protein localization to bud neck(GO:0097271) |
| 0.0 | 0.1 | GO:0034354 | 'de novo' NAD biosynthetic process from tryptophan(GO:0034354) 'de novo' NAD biosynthetic process(GO:0034627) |
| 0.0 | 0.1 | GO:0051260 | protein homooligomerization(GO:0051260) |
| 0.0 | 0.2 | GO:0070987 | error-free translesion synthesis(GO:0070987) |
| 0.0 | 0.1 | GO:0045332 | phospholipid translocation(GO:0045332) |
| 0.0 | 0.0 | GO:0019255 | glucose 1-phosphate metabolic process(GO:0019255) |
| 0.0 | 0.6 | GO:0042176 | regulation of protein catabolic process(GO:0042176) |
| 0.0 | 0.1 | GO:0000161 | MAPK cascade involved in osmosensory signaling pathway(GO:0000161) |
| 0.0 | 0.1 | GO:0071629 | cytoplasm-associated proteasomal ubiquitin-dependent protein catabolic process(GO:0071629) |
| 0.0 | 0.0 | GO:0010513 | positive regulation of phosphatidylinositol biosynthetic process(GO:0010513) |
| 0.0 | 0.0 | GO:0032365 | intracellular lipid transport(GO:0032365) |
| 0.0 | 0.1 | GO:0009410 | response to xenobiotic stimulus(GO:0009410) |
| 0.0 | 0.1 | GO:0043461 | mitochondrial proton-transporting ATP synthase complex assembly(GO:0033615) proton-transporting ATP synthase complex assembly(GO:0043461) |
| 0.0 | 0.0 | GO:0006269 | DNA replication, synthesis of RNA primer(GO:0006269) |
| 0.0 | 0.1 | GO:0000018 | regulation of DNA recombination(GO:0000018) |
| 0.0 | 0.3 | GO:0016573 | histone acetylation(GO:0016573) |
| 0.0 | 0.0 | GO:0030100 | regulation of endocytosis(GO:0030100) |
| 0.0 | 0.0 | GO:0017004 | cytochrome complex assembly(GO:0017004) |
| 0.0 | 0.1 | GO:0051568 | histone H3-K4 methylation(GO:0051568) |
| 0.0 | 0.0 | GO:0050667 | homocysteine metabolic process(GO:0050667) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.6 | 6.3 | GO:0031315 | extrinsic component of mitochondrial outer membrane(GO:0031315) |
| 1.3 | 5.3 | GO:0000148 | 1,3-beta-D-glucan synthase complex(GO:0000148) |
| 1.1 | 9.1 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 1.1 | 4.2 | GO:0016602 | CCAAT-binding factor complex(GO:0016602) |
| 1.0 | 9.1 | GO:0005750 | mitochondrial respiratory chain complex III(GO:0005750) respiratory chain complex III(GO:0045275) |
| 0.8 | 2.4 | GO:0031422 | RecQ helicase-Topo III complex(GO:0031422) |
| 0.8 | 4.8 | GO:0045239 | tricarboxylic acid cycle enzyme complex(GO:0045239) |
| 0.8 | 5.4 | GO:0034657 | GID complex(GO:0034657) |
| 0.8 | 3.8 | GO:0045283 | mitochondrial respiratory chain complex II, succinate dehydrogenase complex (ubiquinone)(GO:0005749) succinate dehydrogenase complex (ubiquinone)(GO:0045257) respiratory chain complex II(GO:0045273) succinate dehydrogenase complex(GO:0045281) fumarate reductase complex(GO:0045283) |
| 0.7 | 1.4 | GO:0030061 | mitochondrial crista(GO:0030061) |
| 0.6 | 4.8 | GO:0042597 | periplasmic space(GO:0042597) |
| 0.5 | 1.8 | GO:0031235 | intrinsic component of the cytoplasmic side of the plasma membrane(GO:0031235) |
| 0.4 | 7.3 | GO:0031307 | intrinsic component of mitochondrial outer membrane(GO:0031306) integral component of mitochondrial outer membrane(GO:0031307) |
| 0.4 | 0.4 | GO:0005775 | vacuolar lumen(GO:0005775) |
| 0.4 | 1.2 | GO:0000015 | phosphopyruvate hydratase complex(GO:0000015) |
| 0.4 | 4.2 | GO:0031907 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.4 | 4.7 | GO:0070469 | respiratory chain(GO:0070469) |
| 0.4 | 1.1 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.3 | 1.4 | GO:0000275 | mitochondrial proton-transporting ATP synthase complex, catalytic core F(1)(GO:0000275) proton-transporting ATP synthase complex, catalytic core F(1)(GO:0045261) |
| 0.3 | 2.5 | GO:0000328 | fungal-type vacuole lumen(GO:0000328) |
| 0.3 | 20.3 | GO:0005576 | extracellular region(GO:0005576) |
| 0.3 | 1.1 | GO:0032116 | SMC loading complex(GO:0032116) |
| 0.3 | 5.6 | GO:0009295 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.2 | 1.7 | GO:0000112 | nucleotide-excision repair factor 3 complex(GO:0000112) |
| 0.2 | 39.5 | GO:0005743 | mitochondrial inner membrane(GO:0005743) |
| 0.2 | 7.2 | GO:0005758 | mitochondrial intermembrane space(GO:0005758) |
| 0.2 | 1.3 | GO:0034967 | Set3 complex(GO:0034967) |
| 0.2 | 0.6 | GO:0000113 | nucleotide-excision repair factor 4 complex(GO:0000113) |
| 0.2 | 1.6 | GO:0045121 | membrane raft(GO:0045121) membrane microdomain(GO:0098857) |
| 0.2 | 1.4 | GO:0005671 | Ada2/Gcn5/Ada3 transcription activator complex(GO:0005671) |
| 0.1 | 3.0 | GO:0070847 | core mediator complex(GO:0070847) |
| 0.1 | 3.4 | GO:0005887 | integral component of plasma membrane(GO:0005887) |
| 0.1 | 0.3 | GO:0032126 | eisosome(GO:0032126) |
| 0.1 | 0.5 | GO:0097038 | perinuclear endoplasmic reticulum(GO:0097038) |
| 0.1 | 0.5 | GO:0042406 | extrinsic component of endoplasmic reticulum membrane(GO:0042406) |
| 0.1 | 0.4 | GO:0030870 | Mre11 complex(GO:0030870) |
| 0.1 | 1.4 | GO:0009898 | cytoplasmic side of plasma membrane(GO:0009898) |
| 0.1 | 0.6 | GO:0005640 | nuclear outer membrane(GO:0005640) |
| 0.1 | 0.3 | GO:1990316 | ATG1/ULK1 kinase complex(GO:1990316) |
| 0.1 | 0.2 | GO:0001400 | mating projection base(GO:0001400) |
| 0.1 | 0.2 | GO:0000111 | nucleotide-excision repair factor 2 complex(GO:0000111) |
| 0.1 | 0.2 | GO:0070985 | TFIIK complex(GO:0070985) |
| 0.1 | 0.9 | GO:0000974 | Prp19 complex(GO:0000974) |
| 0.1 | 0.3 | GO:0072380 | ER membrane insertion complex(GO:0072379) TRC complex(GO:0072380) |
| 0.1 | 1.7 | GO:0042763 | prospore membrane(GO:0005628) intracellular immature spore(GO:0042763) ascospore-type prospore(GO:0042764) |
| 0.1 | 0.3 | GO:0005825 | half bridge of spindle pole body(GO:0005825) |
| 0.1 | 1.1 | GO:0000812 | Swr1 complex(GO:0000812) |
| 0.1 | 0.3 | GO:0033100 | NuA3 histone acetyltransferase complex(GO:0033100) H3 histone acetyltransferase complex(GO:0070775) |
| 0.1 | 0.8 | GO:0098827 | endoplasmic reticulum tubular network(GO:0071782) endoplasmic reticulum subcompartment(GO:0098827) |
| 0.1 | 39.2 | GO:0005739 | mitochondrion(GO:0005739) |
| 0.0 | 0.9 | GO:0005811 | lipid particle(GO:0005811) |
| 0.0 | 0.6 | GO:0030014 | CCR4-NOT complex(GO:0030014) |
| 0.0 | 0.8 | GO:0070822 | Sin3-type complex(GO:0070822) |
| 0.0 | 0.1 | GO:0032585 | multivesicular body membrane(GO:0032585) |
| 0.0 | 0.5 | GO:0000124 | SAGA complex(GO:0000124) |
| 0.0 | 0.1 | GO:0005796 | Golgi lumen(GO:0005796) |
| 0.0 | 0.2 | GO:0000808 | origin recognition complex(GO:0000808) nuclear origin of replication recognition complex(GO:0005664) |
| 0.0 | 0.4 | GO:0008540 | proteasome regulatory particle, base subcomplex(GO:0008540) |
| 0.0 | 0.1 | GO:0033551 | monopolin complex(GO:0033551) |
| 0.0 | 0.3 | GO:0000942 | condensed chromosome outer kinetochore(GO:0000940) condensed nuclear chromosome outer kinetochore(GO:0000942) |
| 0.0 | 0.1 | GO:0000817 | COMA complex(GO:0000817) |
| 0.0 | 0.4 | GO:0005847 | mRNA cleavage and polyadenylation specificity factor complex(GO:0005847) |
| 0.0 | 0.6 | GO:0009277 | fungal-type cell wall(GO:0009277) |
| 0.0 | 0.2 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.0 | 1.1 | GO:0019898 | extrinsic component of membrane(GO:0019898) |
| 0.0 | 0.1 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.0 | 0.1 | GO:0031499 | TRAMP complex(GO:0031499) |
| 0.0 | 0.1 | GO:0005952 | cAMP-dependent protein kinase complex(GO:0005952) |
| 0.0 | 0.0 | GO:0005955 | calcineurin complex(GO:0005955) |
| 0.0 | 0.1 | GO:0097346 | Ino80 complex(GO:0031011) INO80-type complex(GO:0097346) |
| 0.0 | 0.0 | GO:0033309 | SBF transcription complex(GO:0033309) |
| 0.0 | 0.2 | GO:0005682 | U5 snRNP(GO:0005682) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.0 | 19.9 | GO:0015295 | solute:proton symporter activity(GO:0015295) |
| 2.7 | 8.2 | GO:0016406 | carnitine O-acetyltransferase activity(GO:0004092) carnitine O-acyltransferase activity(GO:0016406) |
| 2.7 | 8.1 | GO:0050833 | pyruvate transmembrane transporter activity(GO:0050833) |
| 2.2 | 8.8 | GO:0004396 | hexokinase activity(GO:0004396) |
| 2.1 | 6.3 | GO:0047066 | phospholipid-hydroperoxide glutathione peroxidase activity(GO:0047066) |
| 1.8 | 5.3 | GO:0003843 | 1,3-beta-D-glucan synthase activity(GO:0003843) |
| 1.6 | 6.4 | GO:0015193 | L-proline transmembrane transporter activity(GO:0015193) |
| 1.4 | 4.3 | GO:0004575 | beta-fructofuranosidase activity(GO:0004564) sucrose alpha-glucosidase activity(GO:0004575) |
| 1.4 | 2.8 | GO:0042947 | alpha-glucoside transmembrane transporter activity(GO:0015151) glucoside transmembrane transporter activity(GO:0042947) |
| 1.3 | 5.1 | GO:0015556 | C4-dicarboxylate transmembrane transporter activity(GO:0015556) |
| 1.3 | 3.8 | GO:0048038 | quinone binding(GO:0048038) |
| 1.2 | 7.5 | GO:0004029 | aldehyde dehydrogenase (NAD) activity(GO:0004029) aldehyde dehydrogenase [NAD(P)+] activity(GO:0004030) |
| 1.2 | 3.7 | GO:0090599 | alpha-glucosidase activity(GO:0090599) |
| 1.2 | 1.2 | GO:0030976 | thiamine pyrophosphate binding(GO:0030976) |
| 1.1 | 3.4 | GO:0003954 | NADH dehydrogenase activity(GO:0003954) NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 1.1 | 3.4 | GO:0005536 | glucose binding(GO:0005536) |
| 1.1 | 14.9 | GO:0005353 | fructose transmembrane transporter activity(GO:0005353) mannose transmembrane transporter activity(GO:0015578) |
| 1.1 | 3.2 | GO:0051864 | histone demethylase activity (H3-K36 specific)(GO:0051864) |
| 1.0 | 3.1 | GO:0030414 | endopeptidase inhibitor activity(GO:0004866) peptidase inhibitor activity(GO:0030414) |
| 1.0 | 4.1 | GO:0005537 | mannose binding(GO:0005537) |
| 1.0 | 3.1 | GO:0015927 | alpha,alpha-trehalase activity(GO:0004555) trehalase activity(GO:0015927) |
| 1.0 | 4.0 | GO:0004032 | alditol:NADP+ 1-oxidoreductase activity(GO:0004032) |
| 0.9 | 3.8 | GO:0001094 | TFIID-class transcription factor binding(GO:0001094) |
| 0.9 | 8.5 | GO:0008519 | ammonium transmembrane transporter activity(GO:0008519) |
| 0.9 | 2.8 | GO:0036440 | citrate (Si)-synthase activity(GO:0004108) citrate synthase activity(GO:0036440) |
| 0.9 | 7.3 | GO:0008121 | ubiquinol-cytochrome-c reductase activity(GO:0008121) oxidoreductase activity, acting on diphenols and related substances as donors(GO:0016679) oxidoreductase activity, acting on diphenols and related substances as donors, cytochrome as acceptor(GO:0016681) |
| 0.8 | 4.8 | GO:0016744 | transferase activity, transferring aldehyde or ketonic groups(GO:0016744) |
| 0.7 | 2.2 | GO:0004450 | isocitrate dehydrogenase (NADP+) activity(GO:0004450) |
| 0.7 | 8.5 | GO:0008599 | protein phosphatase type 1 regulator activity(GO:0008599) |
| 0.7 | 11.1 | GO:0020037 | heme binding(GO:0020037) tetrapyrrole binding(GO:0046906) |
| 0.6 | 3.2 | GO:0050308 | sugar-phosphatase activity(GO:0050308) |
| 0.6 | 2.5 | GO:0001097 | TFIIH-class transcription factor binding(GO:0001097) |
| 0.6 | 1.2 | GO:0004448 | isocitrate dehydrogenase activity(GO:0004448) |
| 0.6 | 1.8 | GO:0030060 | L-malate dehydrogenase activity(GO:0030060) |
| 0.6 | 2.3 | GO:0016885 | ligase activity, forming carbon-carbon bonds(GO:0016885) |
| 0.5 | 2.2 | GO:0016405 | CoA-ligase activity(GO:0016405) acid-thiol ligase activity(GO:0016878) |
| 0.5 | 2.7 | GO:0008198 | ferrous iron binding(GO:0008198) |
| 0.5 | 1.0 | GO:0016655 | oxidoreductase activity, acting on NAD(P)H, quinone or similar compound as acceptor(GO:0016655) |
| 0.5 | 0.5 | GO:0051185 | coenzyme transporter activity(GO:0051185) |
| 0.5 | 0.5 | GO:0004075 | biotin carboxylase activity(GO:0004075) |
| 0.4 | 3.0 | GO:0046912 | transferase activity, transferring acyl groups, acyl groups converted into alkyl on transfer(GO:0046912) |
| 0.4 | 2.4 | GO:0072349 | modified amino acid transmembrane transporter activity(GO:0072349) |
| 0.4 | 1.6 | GO:0015294 | solute:cation symporter activity(GO:0015294) |
| 0.4 | 2.4 | GO:0016833 | oxo-acid-lyase activity(GO:0016833) |
| 0.4 | 1.2 | GO:0004634 | phosphopyruvate hydratase activity(GO:0004634) |
| 0.4 | 4.5 | GO:0005199 | structural constituent of cell wall(GO:0005199) |
| 0.4 | 1.5 | GO:0004614 | phosphoglucomutase activity(GO:0004614) |
| 0.4 | 7.2 | GO:0016831 | carboxy-lyase activity(GO:0016831) |
| 0.3 | 1.0 | GO:0000099 | sulfur amino acid transmembrane transporter activity(GO:0000099) |
| 0.3 | 2.1 | GO:0000400 | four-way junction DNA binding(GO:0000400) |
| 0.3 | 5.5 | GO:0015297 | antiporter activity(GO:0015297) |
| 0.3 | 0.9 | GO:0016289 | CoA hydrolase activity(GO:0016289) |
| 0.3 | 0.3 | GO:0032451 | demethylase activity(GO:0032451) histone demethylase activity(GO:0032452) |
| 0.3 | 1.8 | GO:0015293 | symporter activity(GO:0015293) |
| 0.3 | 0.9 | GO:0005253 | anion channel activity(GO:0005253) voltage-gated anion channel activity(GO:0008308) porin activity(GO:0015288) wide pore channel activity(GO:0022829) |
| 0.3 | 3.6 | GO:0044389 | ubiquitin protein ligase binding(GO:0031625) ubiquitin-like protein ligase binding(GO:0044389) |
| 0.3 | 2.7 | GO:0035251 | UDP-glucosyltransferase activity(GO:0035251) |
| 0.3 | 1.3 | GO:0016888 | endodeoxyribonuclease activity, producing 5'-phosphomonoesters(GO:0016888) |
| 0.3 | 1.0 | GO:0017057 | 6-phosphogluconolactonase activity(GO:0017057) |
| 0.2 | 1.0 | GO:0005310 | dicarboxylic acid transmembrane transporter activity(GO:0005310) |
| 0.2 | 4.1 | GO:0001102 | RNA polymerase II activating transcription factor binding(GO:0001102) |
| 0.2 | 1.7 | GO:0000014 | single-stranded DNA endodeoxyribonuclease activity(GO:0000014) |
| 0.2 | 0.9 | GO:0000156 | phosphorelay response regulator activity(GO:0000156) |
| 0.2 | 0.7 | GO:0070569 | uridylyltransferase activity(GO:0070569) |
| 0.2 | 0.7 | GO:0004862 | cAMP-dependent protein kinase inhibitor activity(GO:0004862) |
| 0.2 | 2.4 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 0.2 | 0.6 | GO:0016813 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amidines(GO:0016813) |
| 0.2 | 2.1 | GO:0004872 | receptor activity(GO:0004872) molecular transducer activity(GO:0060089) |
| 0.2 | 0.6 | GO:1901682 | sulfur compound transmembrane transporter activity(GO:1901682) |
| 0.2 | 0.6 | GO:1901474 | azole transmembrane transporter activity(GO:1901474) |
| 0.2 | 0.7 | GO:0000978 | RNA polymerase II core promoter proximal region sequence-specific DNA binding(GO:0000978) |
| 0.2 | 1.6 | GO:0000293 | ferric-chelate reductase activity(GO:0000293) oxidoreductase activity, oxidizing metal ions, NAD or NADP as acceptor(GO:0016723) |
| 0.2 | 0.9 | GO:0019104 | DNA N-glycosylase activity(GO:0019104) |
| 0.2 | 1.5 | GO:0016668 | oxidoreductase activity, acting on a sulfur group of donors, NAD(P) as acceptor(GO:0016668) |
| 0.2 | 0.5 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.2 | 0.5 | GO:0001135 | transcription factor activity, RNA polymerase II transcription factor recruiting(GO:0001135) |
| 0.2 | 0.8 | GO:0051213 | dioxygenase activity(GO:0051213) |
| 0.2 | 0.8 | GO:0004559 | alpha-mannosidase activity(GO:0004559) mannosidase activity(GO:0015923) |
| 0.2 | 0.6 | GO:0005337 | nucleoside transmembrane transporter activity(GO:0005337) |
| 0.1 | 1.2 | GO:0001083 | transcription factor activity, RNA polymerase II basal transcription factor binding(GO:0001083) |
| 0.1 | 1.3 | GO:0016413 | O-acetyltransferase activity(GO:0016413) |
| 0.1 | 0.6 | GO:0030246 | carbohydrate binding(GO:0030246) |
| 0.1 | 1.7 | GO:0045182 | translation regulator activity(GO:0045182) |
| 0.1 | 0.3 | GO:0015037 | peptide disulfide oxidoreductase activity(GO:0015037) glutathione disulfide oxidoreductase activity(GO:0015038) |
| 0.1 | 0.6 | GO:0017022 | myosin binding(GO:0017022) |
| 0.1 | 0.4 | GO:0043812 | phosphatidylinositol phosphate 4-phosphatase activity(GO:0034596) phosphatidylinositol-4-phosphate phosphatase activity(GO:0043812) |
| 0.1 | 0.6 | GO:0004806 | triglyceride lipase activity(GO:0004806) retinyl-palmitate esterase activity(GO:0050253) |
| 0.1 | 0.4 | GO:0003691 | double-stranded telomeric DNA binding(GO:0003691) |
| 0.1 | 0.4 | GO:0016832 | aldehyde-lyase activity(GO:0016832) |
| 0.1 | 2.2 | GO:0036459 | obsolete ubiquitin thiolesterase activity(GO:0004221) thiol-dependent ubiquitin-specific protease activity(GO:0004843) thiol-dependent ubiquitinyl hydrolase activity(GO:0036459) ubiquitinyl hydrolase activity(GO:0101005) |
| 0.1 | 0.3 | GO:0016898 | lactate dehydrogenase activity(GO:0004457) D-lactate dehydrogenase (cytochrome) activity(GO:0004458) oxidoreductase activity, acting on the CH-OH group of donors, cytochrome as acceptor(GO:0016898) |
| 0.1 | 1.9 | GO:0050661 | NADP binding(GO:0050661) |
| 0.1 | 0.4 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.1 | 0.8 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.1 | 0.7 | GO:0019238 | cyclohydrolase activity(GO:0019238) |
| 0.1 | 0.3 | GO:0004128 | cytochrome-b5 reductase activity, acting on NAD(P)H(GO:0004128) |
| 0.1 | 6.6 | GO:0016758 | transferase activity, transferring hexosyl groups(GO:0016758) |
| 0.1 | 1.9 | GO:0005507 | copper ion binding(GO:0005507) |
| 0.1 | 1.1 | GO:0001085 | RNA polymerase II transcription factor binding(GO:0001085) |
| 0.1 | 0.3 | GO:0004619 | phosphoglycerate mutase activity(GO:0004619) |
| 0.1 | 0.4 | GO:0004430 | 1-phosphatidylinositol 4-kinase activity(GO:0004430) |
| 0.1 | 0.2 | GO:0001026 | transcription factor activity, core RNA polymerase III binding(GO:0000995) TFIIIB-type transcription factor activity(GO:0001026) |
| 0.1 | 1.5 | GO:0008081 | phosphoric diester hydrolase activity(GO:0008081) |
| 0.1 | 0.3 | GO:0008172 | S-methyltransferase activity(GO:0008172) |
| 0.1 | 0.9 | GO:0000384 | first spliceosomal transesterification activity(GO:0000384) |
| 0.1 | 2.9 | GO:0001228 | transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001077) transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001228) |
| 0.1 | 0.2 | GO:0047429 | nucleoside-triphosphate diphosphatase activity(GO:0047429) |
| 0.1 | 0.6 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.1 | 0.4 | GO:0016620 | oxidoreductase activity, acting on the aldehyde or oxo group of donors, NAD or NADP as acceptor(GO:0016620) |
| 0.1 | 1.1 | GO:0008483 | transaminase activity(GO:0008483) transferase activity, transferring nitrogenous groups(GO:0016769) |
| 0.1 | 0.2 | GO:0098518 | polynucleotide phosphatase activity(GO:0098518) |
| 0.1 | 0.2 | GO:0003840 | gamma-glutamyltransferase activity(GO:0003840) |
| 0.1 | 0.6 | GO:0008234 | cysteine-type peptidase activity(GO:0008234) |
| 0.1 | 0.2 | GO:0003916 | DNA topoisomerase activity(GO:0003916) |
| 0.1 | 0.3 | GO:0004713 | protein tyrosine kinase activity(GO:0004713) |
| 0.1 | 0.2 | GO:0004724 | magnesium-dependent protein serine/threonine phosphatase activity(GO:0004724) |
| 0.1 | 0.2 | GO:0008169 | C-methyltransferase activity(GO:0008169) |
| 0.1 | 0.6 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.1 | 0.4 | GO:0015174 | basic amino acid transmembrane transporter activity(GO:0015174) |
| 0.1 | 0.3 | GO:0004652 | polynucleotide adenylyltransferase activity(GO:0004652) |
| 0.1 | 0.2 | GO:0008902 | hydroxymethylpyrimidine kinase activity(GO:0008902) phosphomethylpyrimidine kinase activity(GO:0008972) |
| 0.1 | 0.4 | GO:0004012 | phospholipid-translocating ATPase activity(GO:0004012) |
| 0.1 | 1.6 | GO:0008237 | metallopeptidase activity(GO:0008237) |
| 0.0 | 0.4 | GO:0016651 | oxidoreductase activity, acting on NAD(P)H(GO:0016651) |
| 0.0 | 0.2 | GO:0008171 | O-methyltransferase activity(GO:0008171) |
| 0.0 | 0.6 | GO:0004702 | receptor signaling protein serine/threonine kinase activity(GO:0004702) |
| 0.0 | 1.1 | GO:0016881 | acid-amino acid ligase activity(GO:0016881) |
| 0.0 | 0.1 | GO:0004180 | carboxypeptidase activity(GO:0004180) |
| 0.0 | 0.4 | GO:0070001 | aspartic-type endopeptidase activity(GO:0004190) aspartic-type peptidase activity(GO:0070001) |
| 0.0 | 0.2 | GO:0097372 | NAD-dependent histone deacetylase activity (H3-K18 specific)(GO:0097372) |
| 0.0 | 0.2 | GO:0005315 | inorganic phosphate transmembrane transporter activity(GO:0005315) |
| 0.0 | 0.2 | GO:0002161 | aminoacyl-tRNA editing activity(GO:0002161) |
| 0.0 | 0.2 | GO:0003993 | acid phosphatase activity(GO:0003993) |
| 0.0 | 0.2 | GO:0017048 | Rho GTPase binding(GO:0017048) |
| 0.0 | 0.2 | GO:0016782 | transferase activity, transferring sulfur-containing groups(GO:0016782) |
| 0.0 | 0.6 | GO:0005509 | calcium ion binding(GO:0005509) |
| 0.0 | 0.5 | GO:0004857 | enzyme inhibitor activity(GO:0004857) |
| 0.0 | 0.2 | GO:0051184 | cofactor transporter activity(GO:0051184) |
| 0.0 | 0.4 | GO:0032947 | protein complex scaffold(GO:0032947) |
| 0.0 | 0.0 | GO:0004723 | calcium-dependent protein serine/threonine phosphatase activity(GO:0004723) |
| 0.0 | 0.3 | GO:0004725 | protein tyrosine phosphatase activity(GO:0004725) |
| 0.0 | 0.1 | GO:0005102 | receptor binding(GO:0005102) |
| 0.0 | 0.4 | GO:0061733 | histone acetyltransferase activity(GO:0004402) peptide-lysine-N-acetyltransferase activity(GO:0061733) |
| 0.0 | 1.0 | GO:0000981 | RNA polymerase II transcription factor activity, sequence-specific DNA binding(GO:0000981) |
| 0.0 | 0.1 | GO:0001139 | transcription factor activity, core RNA polymerase II recruiting(GO:0001139) |
| 0.0 | 0.3 | GO:0008276 | protein methyltransferase activity(GO:0008276) |
| 0.0 | 0.1 | GO:0008142 | oxysterol binding(GO:0008142) |
| 0.0 | 0.1 | GO:0019888 | protein phosphatase regulator activity(GO:0019888) |
| 0.0 | 0.1 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.0 | 0.1 | GO:0000774 | adenyl-nucleotide exchange factor activity(GO:0000774) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.2 | 6.6 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.9 | 2.7 | PID IL4 2PATHWAY | IL4-mediated signaling events |
| 0.6 | 0.6 | PID AR PATHWAY | Coregulation of Androgen receptor activity |
| 0.4 | 1.1 | SA G1 AND S PHASES | Cdk2, 4, and 6 bind cyclin D in G1, while cdk2/cyclin E promotes the G1/S transition. |
| 0.3 | 1.0 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.3 | 2.2 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 0.3 | 1.1 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.2 | 1.2 | PID ERA GENOMIC PATHWAY | Validated nuclear estrogen receptor alpha network |
| 0.2 | 0.6 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.2 | 1.1 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.2 | 0.3 | PID TCPTP PATHWAY | Signaling events mediated by TCPTP |
| 0.0 | 0.1 | PID FOXO PATHWAY | FoxO family signaling |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 10.7 | REACTOME GLUCONEOGENESIS | Genes involved in Gluconeogenesis |
| 0.7 | 1.5 | REACTOME GLUCOSE METABOLISM | Genes involved in Glucose metabolism |
| 0.6 | 1.9 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |
| 0.6 | 1.7 | REACTOME MEIOTIC SYNAPSIS | Genes involved in Meiotic Synapsis |
| 0.3 | 1.3 | REACTOME DOUBLE STRAND BREAK REPAIR | Genes involved in Double-Strand Break Repair |
| 0.3 | 0.5 | REACTOME SHC1 EVENTS IN ERBB4 SIGNALING | Genes involved in SHC1 events in ERBB4 signaling |
| 0.1 | 0.1 | REACTOME TCA CYCLE AND RESPIRATORY ELECTRON TRANSPORT | Genes involved in The citric acid (TCA) cycle and respiratory electron transport |
| 0.0 | 0.2 | REACTOME E2F ENABLED INHIBITION OF PRE REPLICATION COMPLEX FORMATION | Genes involved in E2F-enabled inhibition of pre-replication complex formation |
| 0.0 | 0.2 | REACTOME METABOLISM OF AMINO ACIDS AND DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |
| 0.0 | 0.2 | REACTOME MEIOTIC RECOMBINATION | Genes involved in Meiotic Recombination |
| 0.0 | 0.0 | REACTOME SYNTHESIS OF SUBSTRATES IN N GLYCAN BIOSYTHESIS | Genes involved in Synthesis of substrates in N-glycan biosythesis |
| 0.0 | 0.1 | REACTOME HEMOSTASIS | Genes involved in Hemostasis |
| 0.0 | 0.2 | REACTOME TRANSCRIPTION | Genes involved in Transcription |