Project
GSE53960: rat RNA-Seq transcriptomic Bodymap
Navigation
Downloads
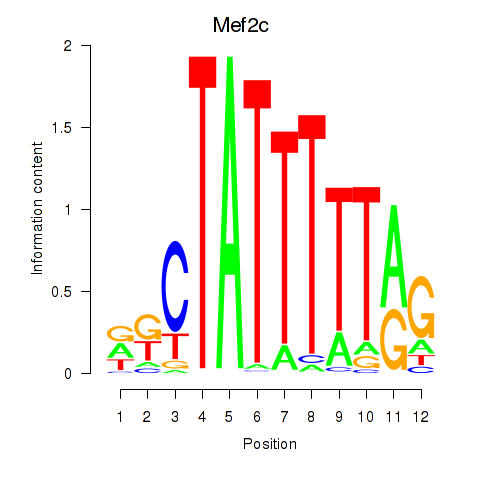
Results for Mef2c
Z-value: 5.08
Transcription factors associated with Mef2c
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Mef2c
|
ENSRNOG00000033134 | myocyte enhancer factor 2C |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Mef2c | rn6_v1_chr2_+_11658568_11658568 | 0.60 | 2.0e-32 | Click! |
Activity profile of Mef2c motif
Sorted Z-values of Mef2c motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr10_+_86337728 | 584.02 |
ENSRNOT00000085408
|
Tcap
|
titin-cap |
| chr4_-_41212072 | 515.82 |
ENSRNOT00000085596
|
Ppp1r3a
|
protein phosphatase 1, regulatory subunit 3A |
| chr1_-_104166367 | 500.33 |
ENSRNOT00000092211
|
Csrp3
|
cysteine and glycine rich protein 3 |
| chr7_-_118108864 | 498.21 |
ENSRNOT00000006184
|
Mb
|
myoglobin |
| chr8_+_119030875 | 446.26 |
ENSRNOT00000028458
|
Myl3
|
myosin light chain 3 |
| chr1_-_199624783 | 445.63 |
ENSRNOT00000026908
|
Cox6a2
|
cytochrome c oxidase subunit 6A2 |
| chr2_-_227207584 | 436.63 |
ENSRNOT00000065361
ENSRNOT00000080215 |
Myoz2
|
myozenin 2 |
| chr3_+_53563194 | 423.58 |
ENSRNOT00000048300
|
Xirp2
|
xin actin-binding repeat containing 2 |
| chr10_+_54352270 | 404.75 |
ENSRNOT00000036752
|
Dhrs7c
|
dehydrogenase/reductase 7C |
| chr1_-_25839198 | 404.65 |
ENSRNOT00000090388
ENSRNOT00000092757 ENSRNOT00000042072 |
Trdn
|
triadin |
| chr15_+_4064706 | 368.15 |
ENSRNOT00000011956
|
Synpo2l
|
synaptopodin 2-like |
| chr10_-_56558487 | 356.01 |
ENSRNOT00000023256
|
Slc2a4
|
solute carrier family 2 member 4 |
| chr9_+_47536824 | 340.33 |
ENSRNOT00000049349
|
Tmem182
|
transmembrane protein 182 |
| chr9_-_73948583 | 339.92 |
ENSRNOT00000018097
|
Myl1
|
myosin, light chain 1 |
| chr12_+_49761120 | 337.44 |
ENSRNOT00000070961
|
Myo18b
|
myosin XVIIIb |
| chr1_+_221756286 | 327.11 |
ENSRNOT00000028636
|
Pygm
|
glycogen phosphorylase, muscle associated |
| chrX_-_31780425 | 318.84 |
ENSRNOT00000004693
|
Asb9
|
ankyrin repeat and SOCS box-containing 9 |
| chr13_+_52624878 | 312.10 |
ENSRNOT00000076054
ENSRNOT00000076299 |
Tnni1
|
troponin I1, slow skeletal type |
| chr1_+_31531143 | 309.58 |
ENSRNOT00000034704
|
Lrrc14b
|
leucine rich repeat containing 14B |
| chr3_+_55910177 | 307.57 |
ENSRNOT00000009969
|
Klhl41
|
kelch-like family member 41 |
| chr1_+_215609036 | 303.72 |
ENSRNOT00000076187
|
Tnni2
|
troponin I2, fast skeletal type |
| chr1_+_215666628 | 302.31 |
ENSRNOT00000040598
ENSRNOT00000066135 ENSRNOT00000051425 ENSRNOT00000080339 ENSRNOT00000066896 ENSRNOT00000063918 |
Tnnt3
|
troponin T3, fast skeletal type |
| chr2_+_204512302 | 299.49 |
ENSRNOT00000021846
|
Casq2
|
calsequestrin 2 |
| chr2_-_45518502 | 288.57 |
ENSRNOT00000014627
|
Hspb3
|
heat shock protein family B (small) member 3 |
| chr1_-_277181345 | 282.74 |
ENSRNOT00000038017
ENSRNOT00000038038 |
Nrap
|
nebulin-related anchoring protein |
| chrX_+_159158194 | 282.54 |
ENSRNOT00000043820
ENSRNOT00000001169 ENSRNOT00000083502 |
Fhl1
|
four and a half LIM domains 1 |
| chr16_-_10941414 | 278.78 |
ENSRNOT00000086627
ENSRNOT00000085414 ENSRNOT00000081631 ENSRNOT00000087521 ENSRNOT00000083623 |
Ldb3
|
LIM domain binding 3 |
| chr1_-_143398093 | 276.13 |
ENSRNOT00000078916
|
Fsd2
|
fibronectin type III and SPRY domain containing 2 |
| chr13_-_90602365 | 272.85 |
ENSRNOT00000009344
|
Casq1
|
calsequestrin 1 |
| chr1_+_215609645 | 264.02 |
ENSRNOT00000076140
ENSRNOT00000027487 |
Tnni2
|
troponin I2, fast skeletal type |
| chr2_+_205568935 | 241.75 |
ENSRNOT00000025248
|
Ampd1
|
adenosine monophosphate deaminase 1 |
| chrX_-_64726210 | 240.70 |
ENSRNOT00000076012
ENSRNOT00000086265 |
Asb12
|
ankyrin repeat and SOCS box-containing 12 |
| chr10_+_110138586 | 234.19 |
ENSRNOT00000086096
|
Slc16a3
|
solute carrier family 16 member 3 |
| chr5_+_152533349 | 234.04 |
ENSRNOT00000067524
|
Trim63
|
tripartite motif containing 63 |
| chr7_-_98197087 | 233.84 |
ENSRNOT00000010484
ENSRNOT00000079961 |
Klhl38
|
kelch-like family member 38 |
| chr2_+_198655437 | 233.12 |
ENSRNOT00000028781
|
Hfe2
|
hemochromatosis type 2 (juvenile) |
| chr10_+_53778662 | 232.68 |
ENSRNOT00000045718
|
Myh2
|
myosin heavy chain 2 |
| chr4_-_99125111 | 229.27 |
ENSRNOT00000009184
|
Smyd1
|
SET and MYND domain containing 1 |
| chr9_+_98490608 | 222.36 |
ENSRNOT00000027232
|
Klhl30
|
kelch-like family member 30 |
| chr8_+_110982777 | 221.27 |
ENSRNOT00000010992
|
Ky
|
kyphoscoliosis peptidase |
| chr9_-_97151832 | 218.38 |
ENSRNOT00000040169
|
Asb18
|
ankyrin repeat and SOCS box-containing 18 |
| chr4_+_144192989 | 214.86 |
ENSRNOT00000007523
|
Lmcd1
|
LIM and cysteine-rich domains 1 |
| chr5_+_148528725 | 210.82 |
ENSRNOT00000017325
|
Fabp3
|
fatty acid binding protein 3 |
| chr5_-_22769907 | 207.36 |
ENSRNOT00000047805
ENSRNOT00000076167 ENSRNOT00000076507 ENSRNOT00000076113 ENSRNOT00000083779 |
Asph
|
aspartate-beta-hydroxylase |
| chr16_-_64806050 | 206.25 |
ENSRNOT00000015725
|
Dusp26
|
dual specificity phosphatase 26 |
| chr15_-_27819376 | 202.56 |
ENSRNOT00000067400
|
A930018M24Rik
|
RIKEN cDNA A930018M24 gene |
| chr9_-_50762082 | 186.86 |
ENSRNOT00000015492
|
Mettl21c
|
methyltransferase like 21C |
| chr7_+_70807867 | 185.08 |
ENSRNOT00000010639
|
Stac3
|
SH3 and cysteine rich domain 3 |
| chr12_-_23841049 | 179.73 |
ENSRNOT00000031555
|
Hspb1
|
heat shock protein family B (small) member 1 |
| chr4_+_7158448 | 176.53 |
ENSRNOT00000076953
|
Asb10
|
ankyrin repeat and SOCS box-containing 10 |
| chr10_-_82785142 | 174.90 |
ENSRNOT00000005381
|
Sgca
|
sarcoglycan, alpha |
| chr8_+_130416355 | 173.33 |
ENSRNOT00000026234
|
Klhl40
|
kelch-like family member 40 |
| chr2_+_119197239 | 172.10 |
ENSRNOT00000048030
|
Usp13
|
ubiquitin specific peptidase 13 |
| chr10_-_62699723 | 169.64 |
ENSRNOT00000086706
|
Coro6
|
coronin 6 |
| chr3_+_79940561 | 169.17 |
ENSRNOT00000016652
|
Mybpc3
|
myosin binding protein C, cardiac |
| chr5_+_90338795 | 168.55 |
ENSRNOT00000077864
ENSRNOT00000058882 |
LOC298139
|
similar to RIKEN cDNA 2310003M01 |
| chr6_+_73553210 | 167.91 |
ENSRNOT00000006562
|
Akap6
|
A-kinase anchoring protein 6 |
| chr1_+_101397828 | 163.80 |
ENSRNOT00000028189
|
Kcna7
|
potassium voltage-gated channel subfamily A member 7 |
| chr10_-_8654892 | 163.02 |
ENSRNOT00000066534
|
Rbfox1
|
RNA binding protein, fox-1 homolog 1 |
| chr14_-_17171580 | 160.14 |
ENSRNOT00000085525
|
Art3
|
ADP-ribosyltransferase 3 |
| chr2_+_219598162 | 159.52 |
ENSRNOT00000020297
|
Lrrc39
|
leucine rich repeat containing 39 |
| chr4_+_71675383 | 157.03 |
ENSRNOT00000051265
|
Clcn1
|
chloride voltage-gated channel 1 |
| chr5_+_154598758 | 154.38 |
ENSRNOT00000015776
|
Tcea3
|
transcription elongation factor A3 |
| chr3_+_112228919 | 154.03 |
ENSRNOT00000011761
|
Capn3
|
calpain 3 |
| chr17_-_78735324 | 148.91 |
ENSRNOT00000036299
|
Cdnf
|
cerebral dopamine neurotrophic factor |
| chr17_-_55346279 | 139.80 |
ENSRNOT00000025037
|
Svil
|
supervillin |
| chr8_-_87419564 | 135.77 |
ENSRNOT00000015365
|
Filip1
|
filamin A interacting protein 1 |
| chr7_-_49741540 | 134.57 |
ENSRNOT00000006523
|
Myf6
|
myogenic factor 6 |
| chrX_-_157013443 | 133.18 |
ENSRNOT00000082711
|
Srpk3
|
SRSF protein kinase 3 |
| chr7_+_11383116 | 131.44 |
ENSRNOT00000066348
|
Nmrk2
|
nicotinamide riboside kinase 2 |
| chr3_+_112228720 | 130.18 |
ENSRNOT00000079079
|
Capn3
|
calpain 3 |
| chr4_+_14001761 | 128.01 |
ENSRNOT00000076519
|
Cd36
|
CD36 molecule |
| chr13_+_52662996 | 123.41 |
ENSRNOT00000047682
|
Tnnt2
|
troponin T2, cardiac type |
| chr1_-_198233215 | 122.11 |
ENSRNOT00000087928
|
Aldoa
|
aldolase, fructose-bisphosphate A |
| chr13_+_51126459 | 119.98 |
ENSRNOT00000042046
ENSRNOT00000087240 |
Myog
|
myogenin |
| chr1_-_198233588 | 119.01 |
ENSRNOT00000088473
|
Aldoa
|
aldolase, fructose-bisphosphate A |
| chr9_-_19372673 | 118.33 |
ENSRNOT00000073667
ENSRNOT00000079517 |
Clic5
|
chloride intracellular channel 5 |
| chr3_+_58965552 | 117.27 |
ENSRNOT00000002068
|
Map3k20
|
mitogen-activated protein kinase kinase kinase 20 |
| chr14_-_43072843 | 114.72 |
ENSRNOT00000064263
|
Limch1
|
LIM and calponin homology domains 1 |
| chr10_+_39655455 | 113.48 |
ENSRNOT00000058817
|
Acsl6
|
acyl-CoA synthetase long-chain family member 6 |
| chr2_-_172361779 | 111.42 |
ENSRNOT00000085876
|
Schip1
|
schwannomin interacting protein 1 |
| chr1_+_219250265 | 99.36 |
ENSRNOT00000024353
|
Doc2g
|
double C2-like domains, gamma |
| chr9_+_82571269 | 96.20 |
ENSRNOT00000026941
|
Speg
|
SPEG complex locus |
| chr2_-_261337163 | 90.68 |
ENSRNOT00000030341
|
Tnni3k
|
TNNI3 interacting kinase |
| chr19_+_52313795 | 88.13 |
ENSRNOT00000021630
|
Wfdc1
|
WAP four-disulfide core domain 1 |
| chr15_+_2766710 | 87.50 |
ENSRNOT00000017483
|
Dupd1
|
dual specificity phosphatase and pro isomerase domain containing 1 |
| chr3_-_51054378 | 83.73 |
ENSRNOT00000089243
|
Grb14
|
growth factor receptor bound protein 14 |
| chr2_-_104958034 | 82.56 |
ENSRNOT00000080699
|
Gyg1
|
glycogenin 1 |
| chr20_+_20378861 | 72.28 |
ENSRNOT00000091044
|
Ank3
|
ankyrin 3 |
| chr16_-_73827488 | 70.67 |
ENSRNOT00000064070
|
Ank1
|
ankyrin 1 |
| chr2_-_247988462 | 69.90 |
ENSRNOT00000022387
|
Pdlim5
|
PDZ and LIM domain 5 |
| chr3_+_151126591 | 67.21 |
ENSRNOT00000025859
|
Myh7b
|
myosin heavy chain 7B |
| chr8_-_72204730 | 64.73 |
ENSRNOT00000023810
|
Fbxl22
|
F-box and leucine-rich repeat protein 22 |
| chr3_-_64024205 | 56.97 |
ENSRNOT00000037015
|
Ccdc141
|
coiled-coil domain containing 141 |
| chr4_+_85551502 | 56.16 |
ENSRNOT00000087191
ENSRNOT00000015692 |
Aqp1
|
aquaporin 1 |
| chr2_+_242634399 | 54.71 |
ENSRNOT00000035700
|
Emcn
|
endomucin |
| chr9_+_111028575 | 52.48 |
ENSRNOT00000043451
|
Pam
|
peptidylglycine alpha-amidating monooxygenase |
| chr9_-_9702306 | 51.80 |
ENSRNOT00000082341
|
Trip10
|
thyroid hormone receptor interactor 10 |
| chr9_+_111028824 | 50.38 |
ENSRNOT00000041418
ENSRNOT00000056457 |
Pam
|
peptidylglycine alpha-amidating monooxygenase |
| chr1_-_56683731 | 50.23 |
ENSRNOT00000014552
|
Thbs2
|
thrombospondin 2 |
| chr3_-_161299024 | 48.71 |
ENSRNOT00000021216
|
Neurl2
|
neuralized E3 ubiquitin protein ligase 2 |
| chr4_+_148782479 | 47.56 |
ENSRNOT00000018133
|
LOC500300
|
similar to hypothetical protein MGC6835 |
| chr12_+_17734133 | 41.28 |
ENSRNOT00000042117
|
Pdgfa
|
platelet derived growth factor subunit A |
| chrX_+_14498119 | 40.54 |
ENSRNOT00000051135
|
Xk
|
X-linked Kx blood group |
| chr13_-_81214494 | 38.53 |
ENSRNOT00000004950
ENSRNOT00000082385 |
Prrx1
|
paired related homeobox 1 |
| chr20_-_9291610 | 38.25 |
ENSRNOT00000000650
|
Glo1
|
glyoxalase 1 |
| chr6_-_137733026 | 37.04 |
ENSRNOT00000019213
|
Jag2
|
jagged 2 |
| chr20_+_20236151 | 31.21 |
ENSRNOT00000079630
|
Ank3
|
ankyrin 3 |
| chr10_-_86688730 | 30.94 |
ENSRNOT00000055333
|
Nr1d1
|
nuclear receptor subfamily 1, group D, member 1 |
| chr3_+_60026747 | 29.18 |
ENSRNOT00000081881
|
Scrn3
|
secernin 3 |
| chr5_-_160219812 | 26.86 |
ENSRNOT00000016333
|
Plekhm2
|
pleckstrin homology and RUN domain containing M2 |
| chr4_+_89079014 | 23.78 |
ENSRNOT00000087451
|
Herc3
|
HECT and RLD domain containing E3 ubiquitin protein ligase 3 |
| chr17_+_36334147 | 22.99 |
ENSRNOT00000050261
|
E2f3
|
E2F transcription factor 3 |
| chr14_+_63095720 | 22.11 |
ENSRNOT00000006071
|
Ppargc1a
|
PPARG coactivator 1 alpha |
| chr10_-_76039964 | 19.04 |
ENSRNOT00000003164
|
Msi2
|
musashi RNA-binding protein 2 |
| chr6_+_64252020 | 18.54 |
ENSRNOT00000047296
ENSRNOT00000082105 |
Pnpla8
|
patatin-like phospholipase domain containing 8 |
| chr4_+_159563798 | 18.18 |
ENSRNOT00000081426
|
Fgf6
|
fibroblast growth factor 6 |
| chr7_+_132857628 | 17.64 |
ENSRNOT00000005438
|
Lrrk2
|
leucine-rich repeat kinase 2 |
| chr18_-_58423196 | 11.28 |
ENSRNOT00000025556
|
Piezo2
|
piezo-type mechanosensitive ion channel component 2 |
| chr6_-_136550371 | 10.64 |
ENSRNOT00000065971
|
Rd3l
|
retinal degeneration 3-like |
| chr4_-_85329362 | 9.94 |
ENSRNOT00000014925
|
Crhr2
|
corticotropin releasing hormone receptor 2 |
| chr5_-_139748489 | 9.79 |
ENSRNOT00000078741
|
Nfyc
|
nuclear transcription factor Y subunit gamma |
| chr17_+_39510936 | 9.16 |
ENSRNOT00000087330
|
Prl8a7
|
prolactin family 8, subfamily a, member 7 |
| chr11_-_74315248 | 8.52 |
ENSRNOT00000002346
|
Hes1
|
hes family bHLH transcription factor 1 |
| chr19_-_11669578 | 7.68 |
ENSRNOT00000026373
|
Gnao1
|
G protein subunit alpha o1 |
| chr2_-_187854363 | 6.42 |
ENSRNOT00000092993
|
Lmna
|
lamin A/C |
| chr1_-_221370322 | 4.43 |
ENSRNOT00000028431
|
Capn1
|
calpain 1 |
| chr3_+_75412372 | 2.79 |
ENSRNOT00000057483
|
Olr560
|
olfactory receptor 560 |
| chr11_+_86520992 | 2.69 |
ENSRNOT00000040954
|
Gp1bb
|
glycoprotein Ib platelet beta subunit |
Network of associatons between targets according to the STRING database.
First level regulatory network of Mef2c
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 271.1 | 1084.4 | GO:0035995 | detection of muscle stretch(GO:0035995) |
| 166.1 | 498.2 | GO:0031444 | slow-twitch skeletal muscle fiber contraction(GO:0031444) |
| 134.9 | 404.6 | GO:0090158 | endoplasmic reticulum membrane organization(GO:0090158) |
| 94.7 | 284.2 | GO:1990091 | sodium-dependent self proteolysis(GO:1990091) |
| 75.2 | 376.1 | GO:0014878 | response to electrical stimulus involved in regulation of muscle adaptation(GO:0014878) |
| 70.3 | 210.8 | GO:2001245 | regulation of phosphatidylcholine biosynthetic process(GO:2001245) |
| 68.8 | 206.3 | GO:1902310 | positive regulation of peptidyl-serine dephosphorylation(GO:1902310) |
| 68.2 | 272.9 | GO:0014809 | regulation of skeletal muscle contraction by regulation of release of sequestered calcium ion(GO:0014809) |
| 59.9 | 179.7 | GO:1903544 | cellular response to interleukin-11(GO:0071348) response to butyrate(GO:1903544) |
| 49.9 | 299.5 | GO:0071313 | cellular response to caffeine(GO:0071313) |
| 43.0 | 172.1 | GO:1904378 | protein K29-linked deubiquitination(GO:0035523) maintenance of unfolded protein(GO:0036506) protein K6-linked deubiquitination(GO:0044313) maintenance of unfolded protein involved in ERAD pathway(GO:1904378) |
| 42.0 | 167.9 | GO:1902261 | positive regulation of delayed rectifier potassium channel activity(GO:1902261) |
| 41.5 | 207.4 | GO:0042264 | peptidyl-aspartic acid hydroxylation(GO:0042264) |
| 41.1 | 123.4 | GO:0032972 | regulation of muscle filament sliding speed(GO:0032972) |
| 40.3 | 241.8 | GO:0032264 | IMP salvage(GO:0032264) |
| 37.8 | 113.5 | GO:0010747 | positive regulation of plasma membrane long-chain fatty acid transport(GO:0010747) |
| 37.1 | 445.6 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
| 34.7 | 173.3 | GO:0098528 | skeletal muscle fiber differentiation(GO:0098528) |
| 33.3 | 1367.2 | GO:0003009 | skeletal muscle contraction(GO:0003009) |
| 30.7 | 337.4 | GO:0048739 | cardiac muscle fiber development(GO:0048739) |
| 27.7 | 470.6 | GO:2001014 | regulation of skeletal muscle cell differentiation(GO:2001014) |
| 26.8 | 401.8 | GO:0031034 | myosin filament assembly(GO:0031034) |
| 26.0 | 234.2 | GO:0035879 | plasma membrane lactate transport(GO:0035879) |
| 25.9 | 103.5 | GO:0010650 | positive regulation of cell communication by electrical coupling(GO:0010650) positive regulation of membrane depolarization during cardiac muscle cell action potential(GO:1900827) |
| 25.7 | 102.9 | GO:0031179 | peptide modification(GO:0031179) |
| 25.6 | 128.0 | GO:0072564 | blood microparticle formation(GO:0072564) regulation of blood microparticle formation(GO:2000332) positive regulation of blood microparticle formation(GO:2000334) |
| 22.4 | 134.6 | GO:0048743 | positive regulation of skeletal muscle fiber development(GO:0048743) |
| 21.3 | 404.7 | GO:0010880 | regulation of release of sequestered calcium ion into cytosol by sarcoplasmic reticulum(GO:0010880) |
| 18.7 | 56.2 | GO:0072237 | metanephric proximal tubule development(GO:0072237) |
| 18.2 | 327.1 | GO:0044247 | glycogen catabolic process(GO:0005980) glucan catabolic process(GO:0009251) cellular polysaccharide catabolic process(GO:0044247) |
| 17.9 | 214.9 | GO:0070886 | positive regulation of calcineurin-NFAT signaling cascade(GO:0070886) |
| 17.2 | 241.1 | GO:0030388 | fructose 1,6-bisphosphate metabolic process(GO:0030388) |
| 17.2 | 446.3 | GO:0002026 | regulation of the force of heart contraction(GO:0002026) |
| 16.7 | 83.7 | GO:0060283 | negative regulation of oocyte development(GO:0060283) negative regulation of oocyte maturation(GO:1900194) |
| 16.1 | 726.2 | GO:0030239 | myofibril assembly(GO:0030239) |
| 15.9 | 174.9 | GO:0014894 | response to muscle inactivity involved in regulation of muscle adaptation(GO:0014877) response to denervation involved in regulation of muscle adaptation(GO:0014894) |
| 14.8 | 237.5 | GO:0016540 | protein autoprocessing(GO:0016540) |
| 13.8 | 41.3 | GO:1990401 | regulation of branching involved in salivary gland morphogenesis by epithelial-mesenchymal signaling(GO:0060683) embryonic lung development(GO:1990401) |
| 13.2 | 356.0 | GO:0044381 | glucose import in response to insulin stimulus(GO:0044381) |
| 13.1 | 157.0 | GO:0098870 | neuronal action potential propagation(GO:0019227) action potential propagation(GO:0098870) |
| 11.5 | 229.3 | GO:0045663 | positive regulation of myoblast differentiation(GO:0045663) |
| 10.3 | 30.9 | GO:0060086 | circadian temperature homeostasis(GO:0060086) |
| 9.1 | 45.6 | GO:0060120 | auditory receptor cell fate commitment(GO:0009912) inner ear receptor cell fate commitment(GO:0060120) |
| 9.1 | 118.3 | GO:0002024 | diet induced thermogenesis(GO:0002024) |
| 7.6 | 423.6 | GO:0055008 | cardiac muscle tissue morphogenesis(GO:0055008) |
| 6.8 | 40.5 | GO:0010961 | cellular magnesium ion homeostasis(GO:0010961) |
| 6.5 | 90.7 | GO:1903779 | regulation of cardiac conduction(GO:1903779) |
| 6.4 | 160.1 | GO:0006471 | protein ADP-ribosylation(GO:0006471) |
| 6.4 | 300.2 | GO:0003254 | regulation of membrane depolarization(GO:0003254) |
| 6.0 | 131.4 | GO:0045662 | negative regulation of myoblast differentiation(GO:0045662) |
| 5.6 | 111.4 | GO:0008210 | estrogen metabolic process(GO:0008210) |
| 5.5 | 38.5 | GO:0048664 | neuron fate determination(GO:0048664) |
| 5.2 | 344.0 | GO:0032233 | positive regulation of actin filament bundle assembly(GO:0032233) |
| 5.1 | 121.7 | GO:0003301 | physiological muscle hypertrophy(GO:0003298) physiological cardiac muscle hypertrophy(GO:0003301) cell growth involved in cardiac muscle cell development(GO:0061049) |
| 4.7 | 102.6 | GO:0055013 | cardiac muscle cell development(GO:0055013) |
| 4.6 | 18.5 | GO:0046338 | phosphatidylethanolamine catabolic process(GO:0046338) |
| 4.5 | 26.9 | GO:1903527 | positive regulation of membrane tubulation(GO:1903527) |
| 4.4 | 287.3 | GO:0060048 | cardiac muscle contraction(GO:0060048) |
| 4.4 | 219.9 | GO:0007528 | neuromuscular junction development(GO:0007528) |
| 4.1 | 148.9 | GO:0071542 | dopaminergic neuron differentiation(GO:0071542) |
| 3.8 | 38.2 | GO:0009438 | methylglyoxal metabolic process(GO:0009438) |
| 3.3 | 117.3 | GO:0007257 | activation of JUN kinase activity(GO:0007257) |
| 3.2 | 283.3 | GO:0031032 | actomyosin structure organization(GO:0031032) |
| 2.8 | 133.2 | GO:0000245 | spliceosomal complex assembly(GO:0000245) |
| 2.5 | 9.9 | GO:2000293 | gastric motility(GO:0035482) regulation of defecation(GO:2000292) negative regulation of defecation(GO:2000293) |
| 2.4 | 154.4 | GO:0006414 | translational elongation(GO:0006414) |
| 2.4 | 70.7 | GO:0072661 | protein targeting to plasma membrane(GO:0072661) |
| 2.0 | 17.7 | GO:0070345 | negative regulation of fat cell proliferation(GO:0070345) |
| 1.9 | 57.0 | GO:0051642 | centrosome localization(GO:0051642) |
| 1.8 | 1363.8 | GO:0016567 | protein ubiquitination(GO:0016567) |
| 1.8 | 99.4 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
| 1.4 | 186.9 | GO:0018022 | peptidyl-lysine methylation(GO:0018022) |
| 1.3 | 64.2 | GO:0061045 | negative regulation of wound healing(GO:0061045) |
| 1.1 | 18.2 | GO:0001502 | cartilage condensation(GO:0001502) |
| 0.8 | 26.4 | GO:0005978 | glycogen biosynthetic process(GO:0005978) glucan biosynthetic process(GO:0009250) |
| 0.8 | 87.5 | GO:0035335 | peptidyl-tyrosine dephosphorylation(GO:0035335) |
| 0.7 | 50.2 | GO:0016525 | negative regulation of angiogenesis(GO:0016525) |
| 0.4 | 124.4 | GO:0051260 | protein homooligomerization(GO:0051260) |
| 0.3 | 11.3 | GO:0050974 | detection of mechanical stimulus involved in sensory perception(GO:0050974) |
| 0.2 | 7.7 | GO:0051926 | negative regulation of calcium ion transport(GO:0051926) |
| 0.1 | 89.1 | GO:0007010 | cytoskeleton organization(GO:0007010) |
| 0.1 | 18.6 | GO:0006887 | exocytosis(GO:0006887) |
| 0.0 | 2.7 | GO:0007596 | blood coagulation(GO:0007596) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 190.9 | 572.6 | GO:0014701 | junctional sarcoplasmic reticulum membrane(GO:0014701) |
| 143.1 | 572.3 | GO:0033018 | sarcoplasmic reticulum lumen(GO:0033018) |
| 134.9 | 404.7 | GO:0014801 | longitudinal sarcoplasmic reticulum(GO:0014801) |
| 108.8 | 1305.6 | GO:0005861 | troponin complex(GO:0005861) |
| 69.4 | 416.1 | GO:0097512 | cardiac myofibril(GO:0097512) |
| 66.3 | 331.5 | GO:0005927 | muscle tendon junction(GO:0005927) |
| 38.9 | 233.1 | GO:1990712 | HFE-transferrin receptor complex(GO:1990712) |
| 35.6 | 356.0 | GO:0032593 | insulin-responsive compartment(GO:0032593) |
| 35.1 | 316.1 | GO:0016461 | unconventional myosin complex(GO:0016461) |
| 32.0 | 1631.1 | GO:0031672 | A band(GO:0031672) |
| 29.1 | 174.9 | GO:0016012 | sarcoglycan complex(GO:0016012) |
| 27.9 | 445.6 | GO:0005751 | mitochondrial respiratory chain complex IV(GO:0005751) |
| 25.3 | 3335.1 | GO:0030018 | Z disc(GO:0030018) |
| 18.9 | 207.4 | GO:0032541 | cortical endoplasmic reticulum(GO:0032541) |
| 14.0 | 56.2 | GO:0020005 | symbiont-containing vacuole membrane(GO:0020005) |
| 13.4 | 537.9 | GO:0016529 | sarcoplasmic reticulum(GO:0016529) |
| 7.9 | 339.9 | GO:0016459 | myosin complex(GO:0016459) |
| 6.4 | 139.8 | GO:0043034 | costamere(GO:0043034) |
| 6.1 | 456.2 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 4.8 | 275.4 | GO:0034707 | chloride channel complex(GO:0034707) |
| 4.4 | 17.6 | GO:0097487 | multivesicular body, internal vesicle(GO:0097487) |
| 2.2 | 50.2 | GO:0031091 | platelet alpha granule(GO:0031091) |
| 1.9 | 51.8 | GO:0001891 | phagocytic cup(GO:0001891) |
| 1.5 | 163.8 | GO:0008076 | voltage-gated potassium channel complex(GO:0008076) |
| 1.4 | 128.0 | GO:0031526 | brush border membrane(GO:0031526) |
| 1.4 | 9.8 | GO:0016602 | CCAAT-binding factor complex(GO:0016602) |
| 1.3 | 255.3 | GO:0005802 | trans-Golgi network(GO:0005802) |
| 0.8 | 22.1 | GO:0097440 | apical dendrite(GO:0097440) |
| 0.8 | 6.4 | GO:0005638 | lamin filament(GO:0005638) |
| 0.8 | 305.4 | GO:0015629 | actin cytoskeleton(GO:0015629) |
| 0.7 | 113.5 | GO:0005741 | mitochondrial outer membrane(GO:0005741) |
| 0.7 | 9.9 | GO:0070852 | cell body fiber(GO:0070852) |
| 0.6 | 145.5 | GO:0000151 | ubiquitin ligase complex(GO:0000151) |
| 0.5 | 179.3 | GO:0005924 | cell-substrate adherens junction(GO:0005924) focal adhesion(GO:0005925) |
| 0.5 | 38.4 | GO:0005902 | microvillus(GO:0005902) |
| 0.5 | 99.4 | GO:0070382 | exocytic vesicle(GO:0070382) |
| 0.5 | 110.6 | GO:0010008 | endosome membrane(GO:0010008) |
| 0.4 | 19.0 | GO:0005844 | polysome(GO:0005844) |
| 0.4 | 214.9 | GO:0031012 | extracellular matrix(GO:0031012) |
| 0.4 | 129.0 | GO:0016607 | nuclear speck(GO:0016607) |
| 0.3 | 18.2 | GO:0042383 | sarcolemma(GO:0042383) |
| 0.2 | 7.7 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.1 | 30.9 | GO:0016604 | nuclear body(GO:0016604) |
| 0.1 | 67.5 | GO:0005794 | Golgi apparatus(GO:0005794) |
| 0.1 | 70.2 | GO:0005856 | cytoskeleton(GO:0005856) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 204.1 | 1020.6 | GO:0051373 | FATZ binding(GO:0051373) |
| 125.1 | 500.3 | GO:0031433 | telethonin binding(GO:0031433) |
| 124.3 | 870.1 | GO:0031014 | troponin T binding(GO:0031014) |
| 109.0 | 327.1 | GO:0008184 | glycogen phosphorylase activity(GO:0008184) |
| 68.8 | 206.3 | GO:0004647 | phosphoserine phosphatase activity(GO:0004647) |
| 61.7 | 123.4 | GO:0030172 | troponin C binding(GO:0030172) |
| 60.4 | 241.8 | GO:0003876 | AMP deaminase activity(GO:0003876) adenosine-phosphate deaminase activity(GO:0047623) |
| 56.5 | 338.8 | GO:0070538 | oleic acid binding(GO:0070538) |
| 45.7 | 730.7 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) |
| 45.3 | 90.7 | GO:0031013 | troponin I binding(GO:0031013) |
| 43.2 | 518.3 | GO:0031432 | titin binding(GO:0031432) |
| 43.0 | 172.1 | GO:1904288 | BAT3 complex binding(GO:1904288) |
| 39.6 | 356.0 | GO:0055056 | D-glucose transmembrane transporter activity(GO:0055056) |
| 38.9 | 233.1 | GO:0098821 | BMP receptor activity(GO:0098821) |
| 34.4 | 241.1 | GO:0004332 | fructose-bisphosphate aldolase activity(GO:0004332) |
| 33.2 | 498.2 | GO:0005344 | oxygen transporter activity(GO:0005344) |
| 29.3 | 234.2 | GO:0015129 | lactate transmembrane transporter activity(GO:0015129) |
| 25.7 | 102.9 | GO:0016842 | amidine-lyase activity(GO:0016842) |
| 20.0 | 160.1 | GO:0003956 | NAD(P)+-protein-arginine ADP-ribosyltransferase activity(GO:0003956) |
| 18.7 | 167.9 | GO:0043495 | protein anchor(GO:0043495) |
| 14.9 | 446.3 | GO:0003785 | actin monomer binding(GO:0003785) |
| 14.9 | 445.6 | GO:0004129 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 14.6 | 423.6 | GO:0051393 | alpha-actinin binding(GO:0051393) |
| 14.3 | 314.0 | GO:0004745 | retinol dehydrogenase activity(GO:0004745) |
| 14.1 | 70.7 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 11.8 | 82.6 | GO:0035251 | UDP-glucosyltransferase activity(GO:0035251) |
| 11.2 | 56.2 | GO:0015166 | polyol transmembrane transporter activity(GO:0015166) glycerol transmembrane transporter activity(GO:0015168) |
| 9.5 | 113.5 | GO:0102391 | decanoate--CoA ligase activity(GO:0102391) |
| 8.6 | 549.1 | GO:0005080 | protein kinase C binding(GO:0005080) |
| 8.3 | 157.0 | GO:0005247 | voltage-gated chloride channel activity(GO:0005247) |
| 7.0 | 154.4 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 5.9 | 17.6 | GO:0034211 | GTP-dependent protein kinase activity(GO:0034211) |
| 4.5 | 117.3 | GO:0004709 | MAP kinase kinase kinase activity(GO:0004709) |
| 4.2 | 680.8 | GO:0044325 | ion channel binding(GO:0044325) |
| 3.3 | 1337.6 | GO:0003779 | actin binding(GO:0003779) |
| 3.2 | 38.2 | GO:0016846 | carbon-sulfur lyase activity(GO:0016846) |
| 3.2 | 337.4 | GO:0003774 | motor activity(GO:0003774) |
| 3.1 | 83.7 | GO:0005070 | SH3/SH2 adaptor activity(GO:0005070) |
| 2.9 | 168.6 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 2.9 | 186.9 | GO:0016279 | lysine N-methyltransferase activity(GO:0016278) protein-lysine N-methyltransferase activity(GO:0016279) |
| 2.7 | 38.4 | GO:0048407 | platelet-derived growth factor binding(GO:0048407) |
| 2.7 | 441.7 | GO:0003714 | transcription corepressor activity(GO:0003714) |
| 2.6 | 128.5 | GO:0070888 | E-box binding(GO:0070888) |
| 2.5 | 9.9 | GO:0015056 | corticotrophin-releasing factor receptor activity(GO:0015056) corticotropin-releasing hormone receptor activity(GO:0043404) |
| 2.5 | 1199.1 | GO:0004842 | ubiquitin-protein transferase activity(GO:0004842) |
| 2.3 | 18.5 | GO:0047499 | calcium-independent phospholipase A2 activity(GO:0047499) |
| 2.2 | 29.2 | GO:0016805 | dipeptidase activity(GO:0016805) |
| 2.1 | 30.9 | GO:0001222 | transcription corepressor binding(GO:0001222) |
| 1.7 | 87.5 | GO:0008138 | protein tyrosine/serine/threonine phosphatase activity(GO:0008138) |
| 1.5 | 99.4 | GO:0005544 | calcium-dependent phospholipid binding(GO:0005544) |
| 1.4 | 11.3 | GO:0008381 | mechanically-gated ion channel activity(GO:0008381) mechanically gated channel activity(GO:0022833) |
| 1.4 | 109.0 | GO:0005254 | chloride channel activity(GO:0005254) |
| 1.3 | 826.2 | GO:0005509 | calcium ion binding(GO:0005509) |
| 1.3 | 7.7 | GO:0031852 | mu-type opioid receptor binding(GO:0031852) corticotropin-releasing hormone receptor 1 binding(GO:0051430) |
| 1.2 | 22.1 | GO:0016922 | ligand-dependent nuclear receptor binding(GO:0016922) alpha-tubulin binding(GO:0043014) |
| 1.1 | 167.1 | GO:0008083 | growth factor activity(GO:0008083) |
| 0.4 | 6.4 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.2 | 117.4 | GO:0004674 | protein serine/threonine kinase activity(GO:0004674) |
| 0.2 | 77.8 | GO:0016773 | phosphotransferase activity, alcohol group as acceptor(GO:0016773) |
| 0.2 | 40.4 | GO:0008022 | protein C-terminus binding(GO:0008022) |
| 0.1 | 2.3 | GO:0071837 | HMG box domain binding(GO:0071837) |
| 0.0 | 5.0 | GO:0005179 | hormone activity(GO:0005179) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 26.3 | 813.8 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
| 10.0 | 179.7 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 6.6 | 282.9 | PID INTEGRIN A4B1 PATHWAY | Alpha4 beta1 integrin signaling events |
| 4.5 | 233.1 | PID BMP PATHWAY | BMP receptor signaling |
| 3.1 | 232.1 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 1.8 | 30.9 | PID CIRCADIAN PATHWAY | Circadian rhythm pathway |
| 1.7 | 83.7 | PID INSULIN PATHWAY | Insulin Pathway |
| 0.8 | 22.1 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.8 | 62.1 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.8 | 58.7 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.8 | 41.3 | PID CERAMIDE PATHWAY | Ceramide signaling pathway |
| 0.7 | 37.0 | PID NOTCH PATHWAY | Notch signaling pathway |
| 0.6 | 7.7 | PID S1P S1P4 PATHWAY | S1P4 pathway |
| 0.4 | 18.2 | PID FGF PATHWAY | FGF signaling pathway |
| 0.3 | 17.7 | PID RB 1PATHWAY | Regulation of retinoblastoma protein |
| 0.3 | 69.6 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.3 | 45.0 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.2 | 9.8 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
| 0.1 | 6.4 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.1 | 4.4 | PID PTP1B PATHWAY | Signaling events mediated by PTP1B |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 60.9 | 2373.5 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 22.5 | 179.7 | REACTOME DESTABILIZATION OF MRNA BY AUF1 HNRNP D0 | Genes involved in Destabilization of mRNA by AUF1 (hnRNP D0) |
| 20.9 | 356.0 | REACTOME FACILITATIVE NA INDEPENDENT GLUCOSE TRANSPORTERS | Genes involved in Facilitative Na+-independent glucose transporters |
| 20.5 | 409.7 | REACTOME GLYCOGEN BREAKDOWN GLYCOGENOLYSIS | Genes involved in Glycogen breakdown (glycogenolysis) |
| 16.7 | 234.2 | REACTOME BILE SALT AND ORGANIC ANION SLC TRANSPORTERS | Genes involved in Bile salt and organic anion SLC transporters |
| 15.1 | 241.8 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 6.3 | 113.5 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 6.2 | 241.1 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 6.1 | 219.5 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 5.0 | 174.2 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 4.7 | 56.2 | REACTOME PASSIVE TRANSPORT BY AQUAPORINS | Genes involved in Passive Transport by Aquaporins |
| 4.1 | 37.0 | REACTOME RECEPTOR LIGAND BINDING INITIATES THE SECOND PROTEOLYTIC CLEAVAGE OF NOTCH RECEPTOR | Genes involved in Receptor-ligand binding initiates the second proteolytic cleavage of Notch receptor |
| 4.0 | 83.7 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 3.3 | 163.8 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 3.2 | 882.5 | REACTOME CLASS I MHC MEDIATED ANTIGEN PROCESSING PRESENTATION | Genes involved in Class I MHC mediated antigen processing & presentation |
| 2.3 | 121.6 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 2.0 | 47.4 | REACTOME SIGNALING BY PDGF | Genes involved in Signaling by PDGF |
| 2.0 | 53.1 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |
| 1.4 | 18.2 | REACTOME SIGNALING BY ACTIVATED POINT MUTANTS OF FGFR1 | Genes involved in Signaling by activated point mutants of FGFR1 |
| 0.7 | 18.5 | REACTOME ACYL CHAIN REMODELLING OF PE | Genes involved in Acyl chain remodelling of PE |
| 0.6 | 17.7 | REACTOME G1 PHASE | Genes involved in G1 Phase |
| 0.3 | 7.7 | REACTOME G PROTEIN ACTIVATION | Genes involved in G-protein activation |
| 0.2 | 8.5 | REACTOME REGULATION OF BETA CELL DEVELOPMENT | Genes involved in Regulation of beta-cell development |
| 0.2 | 2.7 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 0.2 | 17.3 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.1 | 6.4 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |