Project
avrg: GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
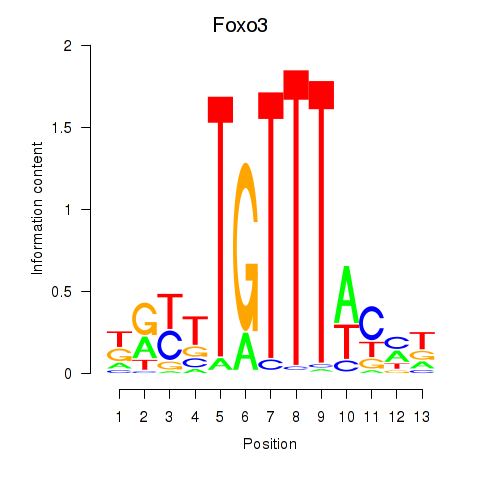
Results for Foxo3
Z-value: 0.62
Transcription factors associated with Foxo3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Foxo3
|
ENSMUSG00000048756.5 | forkhead box O3 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Foxo3 | mm10_v2_chr10_-_42276744_42276763 | -0.19 | 2.8e-01 | Click! |
Activity profile of Foxo3 motif
Sorted Z-values of Foxo3 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr3_+_107230608 | 3.05 |
ENSMUST00000179399.1
|
A630076J17Rik
|
RIKEN cDNA A630076J17 gene |
| chr11_+_101367542 | 1.81 |
ENSMUST00000019469.2
|
G6pc
|
glucose-6-phosphatase, catalytic |
| chr11_-_68386974 | 1.23 |
ENSMUST00000135141.1
|
Ntn1
|
netrin 1 |
| chr3_+_63295815 | 1.20 |
ENSMUST00000029400.1
|
Mme
|
membrane metallo endopeptidase |
| chr14_-_51913393 | 1.10 |
ENSMUST00000004673.7
ENSMUST00000111632.3 |
Ndrg2
|
N-myc downstream regulated gene 2 |
| chr3_-_27896360 | 1.00 |
ENSMUST00000058077.3
|
Tmem212
|
transmembrane protein 212 |
| chr9_-_71163224 | 0.91 |
ENSMUST00000074465.2
|
Aqp9
|
aquaporin 9 |
| chr4_-_49549523 | 0.81 |
ENSMUST00000029987.9
|
Aldob
|
aldolase B, fructose-bisphosphate |
| chr15_-_78468620 | 0.80 |
ENSMUST00000017086.3
|
Tmprss6
|
transmembrane serine protease 6 |
| chr8_-_25038875 | 0.74 |
ENSMUST00000084031.4
|
Htra4
|
HtrA serine peptidase 4 |
| chr11_-_68386821 | 0.71 |
ENSMUST00000021284.3
|
Ntn1
|
netrin 1 |
| chr4_-_119415494 | 0.68 |
ENSMUST00000063642.2
|
Ccdc30
|
coiled-coil domain containing 30 |
| chr15_-_89379246 | 0.67 |
ENSMUST00000049968.7
|
Odf3b
|
outer dense fiber of sperm tails 3B |
| chr1_+_172698046 | 0.62 |
ENSMUST00000038495.3
|
Crp
|
C-reactive protein, pentraxin-related |
| chr1_-_140183283 | 0.61 |
ENSMUST00000111977.1
|
Cfh
|
complement component factor h |
| chr4_+_105157339 | 0.60 |
ENSMUST00000064139.7
|
Ppap2b
|
phosphatidic acid phosphatase type 2B |
| chr1_-_140183404 | 0.58 |
ENSMUST00000066859.6
ENSMUST00000111976.2 |
Cfh
|
complement component factor h |
| chr19_-_7802578 | 0.56 |
ENSMUST00000120522.1
ENSMUST00000065634.7 |
Slc22a26
|
solute carrier family 22 (organic cation transporter), member 26 |
| chr8_+_105269788 | 0.56 |
ENSMUST00000036127.2
ENSMUST00000163734.2 |
Hsf4
|
heat shock transcription factor 4 |
| chr9_-_48835932 | 0.55 |
ENSMUST00000093852.3
|
Zbtb16
|
zinc finger and BTB domain containing 16 |
| chr10_-_89533550 | 0.52 |
ENSMUST00000105297.1
|
Nr1h4
|
nuclear receptor subfamily 1, group H, member 4 |
| chr11_-_100354040 | 0.51 |
ENSMUST00000173630.1
|
Hap1
|
huntingtin-associated protein 1 |
| chr7_-_34655500 | 0.50 |
ENSMUST00000032709.1
|
Kctd15
|
potassium channel tetramerisation domain containing 15 |
| chr19_-_58455903 | 0.48 |
ENSMUST00000131877.1
|
Gfra1
|
glial cell line derived neurotrophic factor family receptor alpha 1 |
| chr11_+_3332426 | 0.45 |
ENSMUST00000136474.1
|
Pik3ip1
|
phosphoinositide-3-kinase interacting protein 1 |
| chr14_+_30879257 | 0.45 |
ENSMUST00000040715.6
|
Mustn1
|
musculoskeletal, embryonic nuclear protein 1 |
| chr15_+_102966794 | 0.44 |
ENSMUST00000001699.7
|
Hoxc10
|
homeobox C10 |
| chr5_-_151190154 | 0.44 |
ENSMUST00000062015.8
ENSMUST00000110483.2 |
Stard13
|
StAR-related lipid transfer (START) domain containing 13 |
| chr19_-_34879452 | 0.40 |
ENSMUST00000036584.5
|
Pank1
|
pantothenate kinase 1 |
| chr7_+_130692532 | 0.38 |
ENSMUST00000033141.6
|
Tacc2
|
transforming, acidic coiled-coil containing protein 2 |
| chr5_+_65131184 | 0.36 |
ENSMUST00000031089.5
ENSMUST00000101191.3 |
Klhl5
|
kelch-like 5 |
| chr2_-_164389095 | 0.35 |
ENSMUST00000167427.1
|
Slpi
|
secretory leukocyte peptidase inhibitor |
| chr11_-_60352869 | 0.33 |
ENSMUST00000095254.5
ENSMUST00000102683.4 ENSMUST00000093048.6 ENSMUST00000093046.6 ENSMUST00000064019.8 ENSMUST00000102682.4 |
Tom1l2
|
target of myb1-like 2 (chicken) |
| chr1_+_24005505 | 0.32 |
ENSMUST00000181961.1
|
Gm26524
|
predicted gene, 26524 |
| chr9_+_114731177 | 0.32 |
ENSMUST00000035007.8
|
Cmtm6
|
CKLF-like MARVEL transmembrane domain containing 6 |
| chr11_-_101785252 | 0.31 |
ENSMUST00000164750.1
ENSMUST00000107176.1 ENSMUST00000017868.6 |
Etv4
|
ets variant gene 4 (E1A enhancer binding protein, E1AF) |
| chr13_-_95478655 | 0.30 |
ENSMUST00000022186.3
|
S100z
|
S100 calcium binding protein, zeta |
| chr1_+_171225054 | 0.29 |
ENSMUST00000111321.1
ENSMUST00000005824.5 ENSMUST00000111320.1 ENSMUST00000111319.1 |
Apoa2
|
apolipoprotein A-II |
| chr2_+_4718145 | 0.29 |
ENSMUST00000056914.6
|
Bend7
|
BEN domain containing 7 |
| chr2_-_148046896 | 0.29 |
ENSMUST00000172928.1
ENSMUST00000047315.3 |
Foxa2
|
forkhead box A2 |
| chr13_+_60602182 | 0.28 |
ENSMUST00000044083.7
|
Dapk1
|
death associated protein kinase 1 |
| chr6_+_14901440 | 0.28 |
ENSMUST00000128567.1
|
Foxp2
|
forkhead box P2 |
| chrX_-_48208870 | 0.27 |
ENSMUST00000088935.3
|
Zdhhc9
|
zinc finger, DHHC domain containing 9 |
| chr4_-_155345696 | 0.27 |
ENSMUST00000103178.4
|
Prkcz
|
protein kinase C, zeta |
| chr3_-_145649970 | 0.27 |
ENSMUST00000029846.3
|
Cyr61
|
cysteine rich protein 61 |
| chr2_-_91710519 | 0.26 |
ENSMUST00000028678.8
ENSMUST00000076803.5 |
Atg13
|
autophagy related 13 |
| chr2_+_10370494 | 0.26 |
ENSMUST00000114862.1
ENSMUST00000114861.1 |
Sfmbt2
|
Scm-like with four mbt domains 2 |
| chr11_+_7197780 | 0.26 |
ENSMUST00000020704.7
|
Igfbp1
|
insulin-like growth factor binding protein 1 |
| chr6_+_24733241 | 0.25 |
ENSMUST00000031690.5
|
Hyal6
|
hyaluronoglucosaminidase 6 |
| chr2_+_155517948 | 0.25 |
ENSMUST00000029135.8
ENSMUST00000065973.2 ENSMUST00000103142.5 |
Acss2
|
acyl-CoA synthetase short-chain family member 2 |
| chr13_+_119606650 | 0.25 |
ENSMUST00000178948.1
|
Gm21967
|
predicted gene, 21967 |
| chr7_+_82175156 | 0.24 |
ENSMUST00000180243.1
|
Sh3gl3
|
SH3-domain GRB2-like 3 |
| chr7_-_90129339 | 0.23 |
ENSMUST00000181189.1
|
2310010J17Rik
|
RIKEN cDNA 2310010J17 gene |
| chr10_+_96616998 | 0.23 |
ENSMUST00000038377.7
|
Btg1
|
B cell translocation gene 1, anti-proliferative |
| chr7_+_29309429 | 0.23 |
ENSMUST00000137848.1
|
Dpf1
|
D4, zinc and double PHD fingers family 1 |
| chr2_+_124610573 | 0.23 |
ENSMUST00000103239.3
ENSMUST00000103240.2 |
Sema6d
|
sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6D |
| chr18_-_74961252 | 0.23 |
ENSMUST00000066532.4
|
Lipg
|
lipase, endothelial |
| chr11_+_82911253 | 0.22 |
ENSMUST00000164945.1
ENSMUST00000018989.7 |
Unc45b
|
unc-45 homolog B (C. elegans) |
| chr8_+_71568866 | 0.22 |
ENSMUST00000034267.4
|
Slc27a1
|
solute carrier family 27 (fatty acid transporter), member 1 |
| chr8_-_84773381 | 0.21 |
ENSMUST00000109764.1
|
Nfix
|
nuclear factor I/X |
| chr14_-_121698417 | 0.21 |
ENSMUST00000040700.7
|
Dock9
|
dedicator of cytokinesis 9 |
| chr7_+_30291941 | 0.21 |
ENSMUST00000144508.1
|
Clip3
|
CAP-GLY domain containing linker protein 3 |
| chr8_-_41016749 | 0.20 |
ENSMUST00000117735.1
|
Mtus1
|
mitochondrial tumor suppressor 1 |
| chr2_+_76675265 | 0.20 |
ENSMUST00000111920.1
|
Plekha3
|
pleckstrin homology domain-containing, family A (phosphoinositide binding specific) member 3 |
| chr17_+_24840108 | 0.19 |
ENSMUST00000164251.1
|
Hagh
|
hydroxyacyl glutathione hydrolase |
| chr7_-_109616548 | 0.18 |
ENSMUST00000077909.1
ENSMUST00000084738.3 |
St5
|
suppression of tumorigenicity 5 |
| chr18_-_62756275 | 0.18 |
ENSMUST00000067450.1
ENSMUST00000048109.5 |
2700046A07Rik
|
RIKEN cDNA 2700046A07 gene |
| chr7_-_65371210 | 0.18 |
ENSMUST00000102592.3
|
Tjp1
|
tight junction protein 1 |
| chr17_+_43389436 | 0.16 |
ENSMUST00000113599.1
|
Gpr116
|
G protein-coupled receptor 116 |
| chr2_+_136713444 | 0.16 |
ENSMUST00000028727.4
ENSMUST00000110098.3 |
Snap25
|
synaptosomal-associated protein 25 |
| chr2_+_91710852 | 0.16 |
ENSMUST00000128140.1
ENSMUST00000140183.1 |
Harbi1
|
harbinger transposase derived 1 |
| chr1_+_75142775 | 0.16 |
ENSMUST00000097694.4
|
Fam134a
|
family with sequence similarity 134, member A |
| chr2_+_127854628 | 0.16 |
ENSMUST00000028859.1
|
Acoxl
|
acyl-Coenzyme A oxidase-like |
| chr2_+_91710900 | 0.15 |
ENSMUST00000142692.1
ENSMUST00000090608.5 |
Harbi1
|
harbinger transposase derived 1 |
| chr2_+_164486856 | 0.15 |
ENSMUST00000109349.2
|
Dbndd2
|
dysbindin (dystrobrevin binding protein 1) domain containing 2 |
| chr2_+_164486455 | 0.14 |
ENSMUST00000069385.8
ENSMUST00000143690.1 |
Dbndd2
|
dysbindin (dystrobrevin binding protein 1) domain containing 2 |
| chr3_+_5218589 | 0.14 |
ENSMUST00000177488.1
|
Zfhx4
|
zinc finger homeodomain 4 |
| chr16_+_93683184 | 0.14 |
ENSMUST00000039620.6
|
Cbr3
|
carbonyl reductase 3 |
| chr2_+_143546144 | 0.13 |
ENSMUST00000028905.9
|
Pcsk2
|
proprotein convertase subtilisin/kexin type 2 |
| chr1_+_74284930 | 0.13 |
ENSMUST00000113805.1
ENSMUST00000027370.6 ENSMUST00000087226.4 |
Pnkd
|
paroxysmal nonkinesiogenic dyskinesia |
| chr12_+_71016658 | 0.13 |
ENSMUST00000125125.1
|
Arid4a
|
AT rich interactive domain 4A (RBP1-like) |
| chr19_+_5877794 | 0.13 |
ENSMUST00000145200.1
ENSMUST00000025732.7 ENSMUST00000125114.1 ENSMUST00000155697.1 |
Slc25a45
|
solute carrier family 25, member 45 |
| chr1_+_36511867 | 0.12 |
ENSMUST00000001166.7
ENSMUST00000097776.3 |
Cnnm3
|
cyclin M3 |
| chr10_+_88201158 | 0.12 |
ENSMUST00000171151.2
|
Ccdc53
|
coiled-coil domain containing 53 |
| chr11_+_60353324 | 0.11 |
ENSMUST00000070805.6
ENSMUST00000094140.2 ENSMUST00000108723.2 ENSMUST00000108722.4 |
Lrrc48
|
leucine rich repeat containing 48 |
| chr8_-_41016295 | 0.11 |
ENSMUST00000131965.1
|
Mtus1
|
mitochondrial tumor suppressor 1 |
| chr4_-_91376433 | 0.10 |
ENSMUST00000107109.2
ENSMUST00000107111.2 ENSMUST00000107120.1 |
Elavl2
|
ELAV (embryonic lethal, abnormal vision, Drosophila)-like 2 (Hu antigen B) |
| chr19_+_22692613 | 0.10 |
ENSMUST00000099564.2
ENSMUST00000099566.3 |
Trpm3
|
transient receptor potential cation channel, subfamily M, member 3 |
| chr2_-_104493690 | 0.10 |
ENSMUST00000111124.1
|
Hipk3
|
homeodomain interacting protein kinase 3 |
| chr4_-_137785371 | 0.10 |
ENSMUST00000133473.1
|
Alpl
|
alkaline phosphatase, liver/bone/kidney |
| chr3_+_5218516 | 0.10 |
ENSMUST00000175866.1
|
Zfhx4
|
zinc finger homeodomain 4 |
| chr7_-_65370908 | 0.10 |
ENSMUST00000032729.6
|
Tjp1
|
tight junction protein 1 |
| chr10_+_88201223 | 0.09 |
ENSMUST00000182619.1
|
Ccdc53
|
coiled-coil domain containing 53 |
| chr18_+_37513652 | 0.09 |
ENSMUST00000061405.4
|
Pcdhb21
|
protocadherin beta 21 |
| chr2_+_124610278 | 0.09 |
ENSMUST00000051419.8
ENSMUST00000077847.5 ENSMUST00000078621.5 ENSMUST00000076335.5 |
Sema6d
|
sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6D |
| chr4_-_91376490 | 0.09 |
ENSMUST00000107124.3
|
Elavl2
|
ELAV (embryonic lethal, abnormal vision, Drosophila)-like 2 (Hu antigen B) |
| chr3_+_5218546 | 0.08 |
ENSMUST00000026284.6
|
Zfhx4
|
zinc finger homeodomain 4 |
| chrX_+_7878298 | 0.08 |
ENSMUST00000033495.8
|
Pim2
|
proviral integration site 2 |
| chr10_-_93310963 | 0.08 |
ENSMUST00000151153.1
|
Elk3
|
ELK3, member of ETS oncogene family |
| chr3_-_146770218 | 0.08 |
ENSMUST00000106137.1
|
Prkacb
|
protein kinase, cAMP dependent, catalytic, beta |
| chr8_-_25785154 | 0.08 |
ENSMUST00000038498.8
|
Bag4
|
BCL2-associated athanogene 4 |
| chr9_-_58741543 | 0.08 |
ENSMUST00000098674.4
|
2410076I21Rik
|
RIKEN cDNA 2410076I21 gene |
| chr19_-_43986552 | 0.08 |
ENSMUST00000026210.4
|
Cpn1
|
carboxypeptidase N, polypeptide 1 |
| chr7_+_30458280 | 0.08 |
ENSMUST00000126297.1
|
Nphs1
|
nephrosis 1, nephrin |
| chr7_-_70360593 | 0.07 |
ENSMUST00000032768.7
|
Nr2f2
|
nuclear receptor subfamily 2, group F, member 2 |
| chr1_-_58695944 | 0.06 |
ENSMUST00000055313.7
|
Als2cr12
|
amyotrophic lateral sclerosis 2 (juvenile) chromosome region, candidate 12 (human) |
| chr11_-_99422252 | 0.06 |
ENSMUST00000017741.3
|
Krt12
|
keratin 12 |
| chr11_+_85353156 | 0.06 |
ENSMUST00000108061.1
ENSMUST00000108062.1 ENSMUST00000108056.1 ENSMUST00000138423.1 ENSMUST00000074875.4 ENSMUST00000092821.3 |
Bcas3
|
breast carcinoma amplified sequence 3 |
| chr10_-_93311073 | 0.06 |
ENSMUST00000008542.5
|
Elk3
|
ELK3, member of ETS oncogene family |
| chr2_+_143915273 | 0.05 |
ENSMUST00000103172.3
|
Dstn
|
destrin |
| chr6_-_142278836 | 0.05 |
ENSMUST00000111825.3
|
Slco1a5
|
solute carrier organic anion transporter family, member 1a5 |
| chr16_-_56712825 | 0.05 |
ENSMUST00000136394.1
|
Tfg
|
Trk-fused gene |
| chr2_-_51973219 | 0.05 |
ENSMUST00000028314.2
|
Nmi
|
N-myc (and STAT) interactor |
| chr16_-_22439719 | 0.05 |
ENSMUST00000079601.6
|
Etv5
|
ets variant gene 5 |
| chr4_-_134915010 | 0.05 |
ENSMUST00000105863.1
ENSMUST00000030626.5 |
Tmem50a
|
transmembrane protein 50A |
| chr11_+_96133786 | 0.05 |
ENSMUST00000167258.1
|
Ttll6
|
tubulin tyrosine ligase-like family, member 6 |
| chr5_-_137531204 | 0.04 |
ENSMUST00000150063.2
|
Gnb2
|
guanine nucleotide binding protein (G protein), beta 2 |
| chr9_+_65361049 | 0.04 |
ENSMUST00000147185.1
|
Gm514
|
predicted gene 514 |
| chr11_+_19924403 | 0.04 |
ENSMUST00000093298.5
|
Spred2
|
sprouty-related, EVH1 domain containing 2 |
| chr11_+_98446826 | 0.04 |
ENSMUST00000019456.4
|
Grb7
|
growth factor receptor bound protein 7 |
| chr18_-_39489880 | 0.04 |
ENSMUST00000152853.1
|
Nr3c1
|
nuclear receptor subfamily 3, group C, member 1 |
| chr16_-_38433145 | 0.04 |
ENSMUST00000002926.6
|
Pla1a
|
phospholipase A1 member A |
| chr6_-_83831736 | 0.03 |
ENSMUST00000058383.8
|
Paip2b
|
poly(A) binding protein interacting protein 2B |
| chr11_-_48946148 | 0.03 |
ENSMUST00000104958.1
|
Psme2b
|
protease (prosome, macropain) activator subunit 2B |
| chr10_-_53647080 | 0.03 |
ENSMUST00000169866.1
|
Fam184a
|
family with sequence similarity 184, member A |
| chr18_-_66022580 | 0.03 |
ENSMUST00000143990.1
|
Lman1
|
lectin, mannose-binding, 1 |
| chr1_-_163403627 | 0.03 |
ENSMUST00000045138.4
|
Gorab
|
golgin, RAB6-interacting |
| chr8_-_122476036 | 0.03 |
ENSMUST00000014614.3
|
Rnf166
|
ring finger protein 166 |
| chr10_+_40349265 | 0.03 |
ENSMUST00000044672.4
ENSMUST00000095743.2 |
Cdk19
|
cyclin-dependent kinase 19 |
| chr3_-_30793549 | 0.02 |
ENSMUST00000180833.1
|
4933429H19Rik
|
RIKEN cDNA 4933429H19 gene |
| chr6_+_34863130 | 0.02 |
ENSMUST00000074949.3
|
Tmem140
|
transmembrane protein 140 |
| chr5_-_17888884 | 0.02 |
ENSMUST00000169095.1
|
Cd36
|
CD36 antigen |
| chr14_-_57133585 | 0.02 |
ENSMUST00000039380.8
|
Gjb6
|
gap junction protein, beta 6 |
| chr10_-_5069044 | 0.02 |
ENSMUST00000095899.3
|
Syne1
|
spectrin repeat containing, nuclear envelope 1 |
| chr5_+_66968559 | 0.02 |
ENSMUST00000127184.1
|
Limch1
|
LIM and calponin homology domains 1 |
| chr11_-_99543158 | 0.01 |
ENSMUST00000107443.1
ENSMUST00000074253.3 |
Krt40
|
keratin 40 |
| chr19_-_19111181 | 0.01 |
ENSMUST00000112832.1
|
Rorb
|
RAR-related orphan receptor beta |
| chr12_+_72761211 | 0.01 |
ENSMUST00000021514.8
|
Ppm1a
|
protein phosphatase 1A, magnesium dependent, alpha isoform |
| chr7_+_141476374 | 0.01 |
ENSMUST00000117634.1
|
Tspan4
|
tetraspanin 4 |
| chr15_+_25752860 | 0.01 |
ENSMUST00000022882.5
ENSMUST00000135173.1 |
Myo10
|
myosin X |
| chr6_+_15196949 | 0.01 |
ENSMUST00000151301.1
ENSMUST00000131414.1 ENSMUST00000140557.1 ENSMUST00000115469.1 |
Foxp2
|
forkhead box P2 |
Network of associatons between targets according to the STRING database.
First level regulatory network of Foxo3
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.2 | GO:0071492 | cellular response to UV-A(GO:0071492) |
| 0.3 | 1.8 | GO:0015712 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 0.3 | 1.1 | GO:0090360 | platelet-derived growth factor production(GO:0090360) regulation of platelet-derived growth factor production(GO:0090361) |
| 0.2 | 1.9 | GO:0033564 | anterior/posterior axon guidance(GO:0033564) |
| 0.2 | 0.9 | GO:0015851 | nucleobase transport(GO:0015851) pyrimidine nucleobase transport(GO:0015855) |
| 0.2 | 0.8 | GO:0006116 | fructose catabolic process(GO:0006001) NADH oxidation(GO:0006116) fructose catabolic process to hydroxyacetone phosphate and glyceraldehyde-3-phosphate(GO:0061624) glycolytic process through fructose-1-phosphate(GO:0061625) |
| 0.2 | 0.5 | GO:0034971 | histone H3-R17 methylation(GO:0034971) |
| 0.1 | 1.2 | GO:0045919 | positive regulation of cytolysis(GO:0045919) |
| 0.1 | 0.4 | GO:0097498 | endothelial tube lumen extension(GO:0097498) |
| 0.1 | 0.6 | GO:0032929 | negative regulation of superoxide anion generation(GO:0032929) |
| 0.1 | 0.5 | GO:0032901 | positive regulation of neurotrophin production(GO:0032901) |
| 0.1 | 0.3 | GO:0045013 | carbon catabolite repression of transcription(GO:0045013) negative regulation of transcription by glucose(GO:0045014) |
| 0.1 | 0.2 | GO:0010982 | regulation of high-density lipoprotein particle clearance(GO:0010982) positive regulation of lipoprotein particle clearance(GO:0010986) |
| 0.1 | 0.4 | GO:1902732 | positive regulation of chondrocyte proliferation(GO:1902732) |
| 0.1 | 0.6 | GO:0098728 | germ-line stem cell division(GO:0042078) male germ-line stem cell asymmetric division(GO:0048133) germline stem cell asymmetric division(GO:0098728) |
| 0.1 | 0.3 | GO:0010901 | regulation of very-low-density lipoprotein particle remodeling(GO:0010901) diacylglycerol catabolic process(GO:0046340) |
| 0.1 | 0.5 | GO:0043553 | negative regulation of phosphatidylinositol 3-kinase activity(GO:0043553) |
| 0.1 | 0.8 | GO:0097264 | self proteolysis(GO:0097264) |
| 0.1 | 0.2 | GO:0006659 | phosphatidylserine biosynthetic process(GO:0006659) |
| 0.1 | 0.2 | GO:0030070 | insulin processing(GO:0030070) |
| 0.1 | 0.3 | GO:0060762 | regulation of branching involved in mammary gland duct morphogenesis(GO:0060762) |
| 0.0 | 0.3 | GO:2000304 | positive regulation of sphingolipid biosynthetic process(GO:0090154) positive regulation of ceramide biosynthetic process(GO:2000304) |
| 0.0 | 0.1 | GO:0042376 | phylloquinone metabolic process(GO:0042374) phylloquinone catabolic process(GO:0042376) quinone catabolic process(GO:1901662) |
| 0.0 | 0.2 | GO:0006083 | acetate metabolic process(GO:0006083) short-chain fatty acid biosynthetic process(GO:0051790) |
| 0.0 | 0.6 | GO:0015747 | urate transport(GO:0015747) |
| 0.0 | 0.3 | GO:2000667 | positive regulation of interleukin-5 secretion(GO:2000664) positive regulation of interleukin-13 secretion(GO:2000667) |
| 0.0 | 0.2 | GO:2000271 | positive regulation of fibroblast apoptotic process(GO:2000271) |
| 0.0 | 0.3 | GO:0061727 | methylglyoxal catabolic process to D-lactate via S-lactoyl-glutathione(GO:0019243) methylglyoxal catabolic process(GO:0051596) methylglyoxal catabolic process to lactate(GO:0061727) |
| 0.0 | 0.6 | GO:0033169 | histone H3-K9 demethylation(GO:0033169) |
| 0.0 | 0.4 | GO:0015937 | coenzyme A biosynthetic process(GO:0015937) |
| 0.0 | 0.3 | GO:0090557 | establishment of endothelial intestinal barrier(GO:0090557) |
| 0.0 | 0.4 | GO:0030953 | astral microtubule organization(GO:0030953) |
| 0.0 | 0.1 | GO:0034773 | histone H4-K20 trimethylation(GO:0034773) |
| 0.0 | 0.1 | GO:0090365 | regulation of mRNA modification(GO:0090365) |
| 0.0 | 0.3 | GO:0071447 | cellular response to hydroperoxide(GO:0071447) |
| 0.0 | 0.2 | GO:0098967 | exocytic insertion of neurotransmitter receptor to plasma membrane(GO:0098881) exocytic insertion of neurotransmitter receptor to postsynaptic membrane(GO:0098967) |
| 0.0 | 0.2 | GO:0018231 | peptidyl-L-cysteine S-palmitoylation(GO:0018230) peptidyl-S-diacylglycerol-L-cysteine biosynthetic process from peptidyl-cysteine(GO:0018231) |
| 0.0 | 0.4 | GO:0021520 | spinal cord motor neuron cell fate specification(GO:0021520) |
| 0.0 | 0.2 | GO:0016191 | synaptic vesicle uncoating(GO:0016191) |
| 0.0 | 0.1 | GO:0060849 | regulation of transcription involved in lymphatic endothelial cell fate commitment(GO:0060849) |
| 0.0 | 0.2 | GO:0021707 | cerebellar granular layer formation(GO:0021684) cerebellar granule cell differentiation(GO:0021707) |
| 0.0 | 0.1 | GO:0030043 | actin filament fragmentation(GO:0030043) |
| 0.0 | 0.2 | GO:0060501 | positive regulation of epithelial cell proliferation involved in lung morphogenesis(GO:0060501) |
| 0.0 | 0.3 | GO:0098780 | response to mitochondrial depolarisation(GO:0098780) |
| 0.0 | 0.1 | GO:0061303 | cornea development in camera-type eye(GO:0061303) |
| 0.0 | 0.1 | GO:1901620 | regulation of smoothened signaling pathway involved in dorsal/ventral neural tube patterning(GO:1901620) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.8 | GO:0030868 | smooth endoplasmic reticulum membrane(GO:0030868) smooth endoplasmic reticulum part(GO:0097425) |
| 0.0 | 0.4 | GO:0019907 | cyclin-dependent protein kinase activating kinase holoenzyme complex(GO:0019907) |
| 0.0 | 0.2 | GO:0070033 | synaptobrevin 2-SNAP-25-syntaxin-1a-complexin I complex(GO:0070032) synaptobrevin 2-SNAP-25-syntaxin-1a-complexin II complex(GO:0070033) |
| 0.0 | 0.3 | GO:1990316 | ATG1/ULK1 kinase complex(GO:1990316) |
| 0.0 | 0.3 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.0 | 0.3 | GO:0002178 | palmitoyltransferase complex(GO:0002178) |
| 0.0 | 0.3 | GO:0034366 | spherical high-density lipoprotein particle(GO:0034366) |
| 0.0 | 0.3 | GO:0043203 | axon hillock(GO:0043203) apical cortex(GO:0045179) |
| 0.0 | 1.9 | GO:0005604 | basement membrane(GO:0005604) |
| 0.0 | 1.8 | GO:0030176 | integral component of endoplasmic reticulum membrane(GO:0030176) |
| 0.0 | 0.5 | GO:0005719 | nuclear euchromatin(GO:0005719) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 1.8 | GO:0050309 | glucose-6-phosphatase activity(GO:0004346) sugar-terminal-phosphatase activity(GO:0050309) |
| 0.2 | 0.9 | GO:0005350 | purine nucleobase transmembrane transporter activity(GO:0005345) pyrimidine nucleobase transmembrane transporter activity(GO:0005350) nucleobase transmembrane transporter activity(GO:0015205) glycerol channel activity(GO:0015254) |
| 0.2 | 1.2 | GO:0001851 | complement component C3b binding(GO:0001851) |
| 0.1 | 0.5 | GO:0036313 | phosphatidylinositol 3-kinase catalytic subunit binding(GO:0036313) |
| 0.1 | 0.5 | GO:0038181 | bile acid receptor activity(GO:0038181) |
| 0.1 | 0.8 | GO:0004332 | fructose-bisphosphate aldolase activity(GO:0004332) |
| 0.1 | 0.2 | GO:0003987 | acetate-CoA ligase activity(GO:0003987) |
| 0.1 | 0.4 | GO:0004594 | pantothenate kinase activity(GO:0004594) |
| 0.1 | 0.6 | GO:0001849 | complement component C1q binding(GO:0001849) |
| 0.1 | 0.3 | GO:0070653 | high-density lipoprotein particle receptor binding(GO:0070653) |
| 0.1 | 0.5 | GO:0048403 | brain-derived neurotrophic factor binding(GO:0048403) |
| 0.0 | 0.1 | GO:0000253 | 3-keto sterol reductase activity(GO:0000253) |
| 0.0 | 0.3 | GO:0008970 | phosphatidylcholine 1-acylhydrolase activity(GO:0008970) |
| 0.0 | 1.3 | GO:0005520 | insulin-like growth factor binding(GO:0005520) |
| 0.0 | 0.5 | GO:0001206 | transcriptional repressor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001206) |
| 0.0 | 0.6 | GO:1901702 | urate transmembrane transporter activity(GO:0015143) salt transmembrane transporter activity(GO:1901702) |
| 0.0 | 0.3 | GO:0004416 | hydroxyacylglutathione hydrolase activity(GO:0004416) |
| 0.0 | 0.1 | GO:0032422 | purine-rich negative regulatory element binding(GO:0032422) |
| 0.0 | 0.3 | GO:0004415 | hyalurononglucosaminidase activity(GO:0004415) |
| 0.0 | 0.3 | GO:0071253 | connexin binding(GO:0071253) |
| 0.0 | 0.1 | GO:0010698 | acetyltransferase activator activity(GO:0010698) |
| 0.0 | 0.2 | GO:0031957 | very long-chain fatty acid-CoA ligase activity(GO:0031957) |
| 0.0 | 0.3 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.0 | 0.1 | GO:0004035 | alkaline phosphatase activity(GO:0004035) |
| 0.0 | 0.2 | GO:0003997 | acyl-CoA oxidase activity(GO:0003997) |
| 0.0 | 0.3 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.0 | 0.3 | GO:0004697 | protein kinase C activity(GO:0004697) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 2.2 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
| 0.0 | 2.5 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 0.6 | PID NEPHRIN NEPH1 PATHWAY | Nephrin/Neph1 signaling in the kidney podocyte |
| 0.0 | 0.5 | PID RET PATHWAY | Signaling events regulated by Ret tyrosine kinase |
| 0.0 | 1.1 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
| 0.0 | 0.6 | PID IL6 7 PATHWAY | IL6-mediated signaling events |
| 0.0 | 0.3 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.0 | 0.5 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.9 | REACTOME ROLE OF SECOND MESSENGERS IN NETRIN1 SIGNALING | Genes involved in Role of second messengers in netrin-1 signaling |
| 0.1 | 1.2 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 0.1 | 0.6 | REACTOME CREATION OF C4 AND C2 ACTIVATORS | Genes involved in Creation of C4 and C2 activators |
| 0.1 | 0.9 | REACTOME PASSIVE TRANSPORT BY AQUAPORINS | Genes involved in Passive Transport by Aquaporins |
| 0.0 | 0.2 | REACTOME ETHANOL OXIDATION | Genes involved in Ethanol oxidation |
| 0.0 | 1.8 | REACTOME GLUCOSE TRANSPORT | Genes involved in Glucose transport |
| 0.0 | 0.4 | REACTOME VITAMIN B5 PANTOTHENATE METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |
| 0.0 | 0.3 | REACTOME ROLE OF DCC IN REGULATING APOPTOSIS | Genes involved in Role of DCC in regulating apoptosis |
| 0.0 | 0.9 | REACTOME GLUCONEOGENESIS | Genes involved in Gluconeogenesis |
| 0.0 | 0.2 | REACTOME ACETYLCHOLINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Acetylcholine Neurotransmitter Release Cycle |
| 0.0 | 0.6 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.0 | 0.3 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.0 | 0.3 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.0 | 0.3 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.0 | 0.3 | REACTOME TGF BETA RECEPTOR SIGNALING IN EMT EPITHELIAL TO MESENCHYMAL TRANSITION | Genes involved in TGF-beta receptor signaling in EMT (epithelial to mesenchymal transition) |