Project
avrg: GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
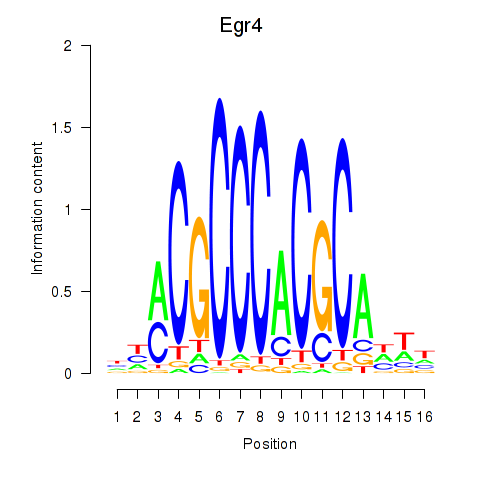
Results for Egr4
Z-value: 0.50
Transcription factors associated with Egr4
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Egr4
|
ENSMUSG00000071341.3 | early growth response 4 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Egr4 | mm10_v2_chr6_-_85513586_85513591 | -0.26 | 1.3e-01 | Click! |
Activity profile of Egr4 motif
Sorted Z-values of Egr4 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chrX_+_49470555 | 0.77 |
ENSMUST00000042444.6
|
Arhgap36
|
Rho GTPase activating protein 36 |
| chrX_+_49470450 | 0.76 |
ENSMUST00000114904.3
|
Arhgap36
|
Rho GTPase activating protein 36 |
| chr11_-_53470479 | 0.51 |
ENSMUST00000057722.2
|
Gm9837
|
predicted gene 9837 |
| chr4_-_43046196 | 0.45 |
ENSMUST00000036462.5
|
Fam214b
|
family with sequence similarity 214, member B |
| chr8_-_92356103 | 0.40 |
ENSMUST00000034183.3
|
4933436C20Rik
|
RIKEN cDNA 4933436C20 gene |
| chr6_+_127149389 | 0.36 |
ENSMUST00000180811.1
|
9330179D12Rik
|
RIKEN cDNA 9330179D12 gene |
| chr12_+_76370266 | 0.35 |
ENSMUST00000042779.3
|
Zbtb1
|
zinc finger and BTB domain containing 1 |
| chr7_+_27653906 | 0.34 |
ENSMUST00000008088.7
|
Ttc9b
|
tetratricopeptide repeat domain 9B |
| chr8_-_92355764 | 0.34 |
ENSMUST00000180102.1
ENSMUST00000179421.1 ENSMUST00000179222.1 ENSMUST00000179029.1 |
4933436C20Rik
|
RIKEN cDNA 4933436C20 gene |
| chr14_-_79481268 | 0.32 |
ENSMUST00000022601.5
|
Wbp4
|
WW domain binding protein 4 |
| chr2_+_105127200 | 0.32 |
ENSMUST00000139585.1
|
Wt1
|
Wilms tumor 1 homolog |
| chr5_+_137288273 | 0.32 |
ENSMUST00000024099.4
ENSMUST00000085934.3 |
Ache
|
acetylcholinesterase |
| chr16_+_44173271 | 0.28 |
ENSMUST00000088356.4
ENSMUST00000169582.1 |
Gm608
|
predicted gene 608 |
| chrX_-_143393893 | 0.27 |
ENSMUST00000166406.2
|
Chrdl1
|
chordin-like 1 |
| chr14_+_79481164 | 0.27 |
ENSMUST00000040131.5
|
Elf1
|
E74-like factor 1 |
| chr15_-_36608959 | 0.26 |
ENSMUST00000001809.8
|
Pabpc1
|
poly(A) binding protein, cytoplasmic 1 |
| chr2_-_9883993 | 0.26 |
ENSMUST00000114915.2
|
9230102O04Rik
|
RIKEN cDNA 9230102O04 gene |
| chr10_-_116473418 | 0.24 |
ENSMUST00000087965.4
ENSMUST00000164271.1 |
Kcnmb4
|
potassium large conductance calcium-activated channel, subfamily M, beta member 4 |
| chrX_+_36195968 | 0.23 |
ENSMUST00000115256.1
|
Zcchc12
|
zinc finger, CCHC domain containing 12 |
| chr16_-_36071515 | 0.21 |
ENSMUST00000004057.7
|
Fam162a
|
family with sequence similarity 162, member A |
| chr3_-_89089955 | 0.20 |
ENSMUST00000166687.1
|
Rusc1
|
RUN and SH3 domain containing 1 |
| chr11_-_102407455 | 0.20 |
ENSMUST00000107098.1
ENSMUST00000018821.2 |
Slc25a39
|
solute carrier family 25, member 39 |
| chr1_+_135132693 | 0.20 |
ENSMUST00000049449.4
|
Ptpn7
|
protein tyrosine phosphatase, non-receptor type 7 |
| chrX_+_36195950 | 0.20 |
ENSMUST00000115257.1
|
Zcchc12
|
zinc finger, CCHC domain containing 12 |
| chr16_+_44173239 | 0.19 |
ENSMUST00000119746.1
|
Gm608
|
predicted gene 608 |
| chr19_-_50678642 | 0.19 |
ENSMUST00000072685.6
ENSMUST00000164039.2 |
Sorcs1
|
VPS10 domain receptor protein SORCS 1 |
| chr1_-_97661668 | 0.19 |
ENSMUST00000153115.1
ENSMUST00000142234.1 |
D1Ertd622e
|
DNA segment, Chr 1, ERATO Doi 622, expressed |
| chrX_+_36195904 | 0.19 |
ENSMUST00000115258.2
|
Zcchc12
|
zinc finger, CCHC domain containing 12 |
| chr17_+_24470393 | 0.18 |
ENSMUST00000053024.6
|
Pgp
|
phosphoglycolate phosphatase |
| chr5_-_149051604 | 0.18 |
ENSMUST00000093196.4
|
Hmgb1
|
high mobility group box 1 |
| chr15_+_102966794 | 0.17 |
ENSMUST00000001699.7
|
Hoxc10
|
homeobox C10 |
| chr4_+_99656299 | 0.17 |
ENSMUST00000087285.3
|
Foxd3
|
forkhead box D3 |
| chrX_+_36195938 | 0.17 |
ENSMUST00000048067.3
|
Zcchc12
|
zinc finger, CCHC domain containing 12 |
| chr8_+_109868586 | 0.17 |
ENSMUST00000179721.1
ENSMUST00000034175.4 |
Phlpp2
|
PH domain and leucine rich repeat protein phosphatase 2 |
| chr6_-_28261907 | 0.17 |
ENSMUST00000115320.1
ENSMUST00000123098.1 ENSMUST00000115321.2 ENSMUST00000155494.1 |
Zfp800
|
zinc finger protein 800 |
| chr11_-_102407899 | 0.16 |
ENSMUST00000124755.1
|
Slc25a39
|
solute carrier family 25, member 39 |
| chr11_-_102407315 | 0.16 |
ENSMUST00000149777.1
ENSMUST00000154001.1 |
Slc25a39
|
solute carrier family 25, member 39 |
| chr1_+_153425162 | 0.15 |
ENSMUST00000042373.5
|
Shcbp1l
|
Shc SH2-domain binding protein 1-like |
| chr6_-_85961631 | 0.15 |
ENSMUST00000032071.9
|
Dusp11
|
dual specificity phosphatase 11 (RNA/RNP complex 1-interacting) |
| chr14_+_56668242 | 0.14 |
ENSMUST00000116468.1
|
Mphosph8
|
M-phase phosphoprotein 8 |
| chr16_+_8830093 | 0.14 |
ENSMUST00000023150.5
|
1810013L24Rik
|
RIKEN cDNA 1810013L24 gene |
| chr6_-_3968357 | 0.14 |
ENSMUST00000031674.8
|
Tfpi2
|
tissue factor pathway inhibitor 2 |
| chr7_-_4778141 | 0.14 |
ENSMUST00000094892.5
|
Il11
|
interleukin 11 |
| chr19_-_50678485 | 0.13 |
ENSMUST00000111756.3
|
Sorcs1
|
VPS10 domain receptor protein SORCS 1 |
| chr9_+_21032038 | 0.13 |
ENSMUST00000019616.4
|
Icam5
|
intercellular adhesion molecule 5, telencephalin |
| chr2_+_180589245 | 0.13 |
ENSMUST00000029087.3
|
Ogfr
|
opioid growth factor receptor |
| chr3_+_144570687 | 0.13 |
ENSMUST00000106211.1
|
Sep15
|
selenoprotein |
| chr1_-_36939521 | 0.13 |
ENSMUST00000027290.5
|
Tmem131
|
transmembrane protein 131 |
| chr12_+_49382791 | 0.13 |
ENSMUST00000179669.1
|
Foxg1
|
forkhead box G1 |
| chr17_-_31658729 | 0.13 |
ENSMUST00000166526.1
ENSMUST00000014684.4 |
U2af1
|
U2 small nuclear ribonucleoprotein auxiliary factor (U2AF) 1 |
| chr16_+_36071624 | 0.12 |
ENSMUST00000164916.1
ENSMUST00000163352.1 |
Ccdc58
|
coiled-coil domain containing 58 |
| chr17_+_26138661 | 0.12 |
ENSMUST00000074370.3
ENSMUST00000118904.2 ENSMUST00000163421.1 |
Axin1
|
axin 1 |
| chr16_-_4213404 | 0.12 |
ENSMUST00000023165.6
|
Crebbp
|
CREB binding protein |
| chr2_+_91922178 | 0.12 |
ENSMUST00000170432.1
|
Chrm4
|
cholinergic receptor, muscarinic 4 |
| chr13_-_111490028 | 0.12 |
ENSMUST00000091236.4
|
Gpbp1
|
GC-rich promoter binding protein 1 |
| chr17_+_69383024 | 0.11 |
ENSMUST00000112674.1
|
Zbtb14
|
zinc finger and BTB domain containing 14 |
| chr14_+_75845093 | 0.11 |
ENSMUST00000110894.2
|
Tpt1
|
tumor protein, translationally-controlled 1 |
| chr13_+_55445301 | 0.11 |
ENSMUST00000001115.8
ENSMUST00000099482.3 |
Grk6
|
G protein-coupled receptor kinase 6 |
| chr2_-_121355284 | 0.11 |
ENSMUST00000134796.1
ENSMUST00000110628.1 ENSMUST00000110627.1 ENSMUST00000110625.1 |
Ppip5k1
|
diphosphoinositol pentakisphosphate kinase 1 |
| chr9_+_40269430 | 0.10 |
ENSMUST00000171835.2
|
Scn3b
|
sodium channel, voltage-gated, type III, beta |
| chrX_+_7842056 | 0.09 |
ENSMUST00000115667.3
ENSMUST00000115668.3 ENSMUST00000115665.1 |
Otud5
|
OTU domain containing 5 |
| chr4_+_86748526 | 0.09 |
ENSMUST00000082026.7
ENSMUST00000045512.8 |
Dennd4c
|
DENN/MADD domain containing 4C |
| chr16_+_93832121 | 0.09 |
ENSMUST00000044068.6
|
Morc3
|
microrchidia 3 |
| chr13_-_111490111 | 0.08 |
ENSMUST00000047627.7
|
Gpbp1
|
GC-rich promoter binding protein 1 |
| chr12_+_49383007 | 0.08 |
ENSMUST00000021333.3
|
Foxg1
|
forkhead box G1 |
| chr2_-_48949206 | 0.08 |
ENSMUST00000090976.3
ENSMUST00000149679.1 ENSMUST00000028098.4 |
Orc4
|
origin recognition complex, subunit 4 |
| chr2_+_164785823 | 0.08 |
ENSMUST00000174070.1
ENSMUST00000172577.1 ENSMUST00000056181.6 |
Snx21
|
sorting nexin family member 21 |
| chr4_+_127169131 | 0.08 |
ENSMUST00000046659.7
|
Dlgap3
|
discs, large (Drosophila) homolog-associated protein 3 |
| chr2_+_164785994 | 0.07 |
ENSMUST00000152471.1
|
Snx21
|
sorting nexin family member 21 |
| chr11_-_86757483 | 0.07 |
ENSMUST00000060766.9
ENSMUST00000103186.4 |
Cltc
|
clathrin, heavy polypeptide (Hc) |
| chr9_+_65908967 | 0.07 |
ENSMUST00000034949.3
ENSMUST00000154589.1 |
Csnk1g1
|
casein kinase 1, gamma 1 |
| chr9_-_44440868 | 0.06 |
ENSMUST00000098837.1
|
Foxr1
|
forkhead box R1 |
| chr5_-_145084030 | 0.06 |
ENSMUST00000151196.1
|
Kpna7
|
karyopherin alpha 7 (importin alpha 8) |
| chr6_+_85451488 | 0.06 |
ENSMUST00000032078.6
|
Cct7
|
chaperonin containing Tcp1, subunit 7 (eta) |
| chr15_-_38078842 | 0.06 |
ENSMUST00000110336.2
|
Ubr5
|
ubiquitin protein ligase E3 component n-recognin 5 |
| chr5_-_96161742 | 0.06 |
ENSMUST00000129646.1
ENSMUST00000113005.2 ENSMUST00000154500.1 ENSMUST00000141383.1 |
Cnot6l
|
CCR4-NOT transcription complex, subunit 6-like |
| chr9_+_114688782 | 0.05 |
ENSMUST00000047404.6
|
Dync1li1
|
dynein cytoplasmic 1 light intermediate chain 1 |
| chr8_+_69088646 | 0.05 |
ENSMUST00000006435.7
|
Atp6v1b2
|
ATPase, H+ transporting, lysosomal V1 subunit B2 |
| chr10_+_93160824 | 0.04 |
ENSMUST00000069965.7
|
Cdk17
|
cyclin-dependent kinase 17 |
| chr15_+_58141397 | 0.04 |
ENSMUST00000067563.7
|
Wdyhv1
|
WDYHV motif containing 1 |
| chr1_-_177258182 | 0.04 |
ENSMUST00000111159.1
|
Akt3
|
thymoma viral proto-oncogene 3 |
| chr4_+_99955715 | 0.04 |
ENSMUST00000102783.4
|
Pgm2
|
phosphoglucomutase 2 |
| chr19_-_8786245 | 0.04 |
ENSMUST00000177216.1
|
Taf6l
|
TAF6-like RNA polymerase II, p300/CBP-associated factor (PCAF)-associated factor |
| chr19_-_8786272 | 0.04 |
ENSMUST00000176610.1
ENSMUST00000177056.1 |
Taf6l
|
TAF6-like RNA polymerase II, p300/CBP-associated factor (PCAF)-associated factor |
| chr6_-_85451248 | 0.03 |
ENSMUST00000113770.1
ENSMUST00000032080.2 |
Pradc1
|
protease-associated domain containing 1 |
| chr17_-_26841565 | 0.03 |
ENSMUST00000015723.4
|
Nkx2-5
|
NK2 homeobox 5 |
| chr9_-_95511897 | 0.03 |
ENSMUST00000079659.5
ENSMUST00000078374.6 |
U2surp
|
U2 snRNP-associated SURP domain containing |
| chr4_+_128883549 | 0.03 |
ENSMUST00000035667.8
|
Trim62
|
tripartite motif-containing 62 |
| chr18_+_86394952 | 0.03 |
ENSMUST00000058829.2
|
Neto1
|
neuropilin (NRP) and tolloid (TLL)-like 1 |
| chr11_-_86993682 | 0.02 |
ENSMUST00000018571.4
|
Ypel2
|
yippee-like 2 (Drosophila) |
| chr14_+_84443553 | 0.02 |
ENSMUST00000071370.5
|
Pcdh17
|
protocadherin 17 |
| chr12_+_17690793 | 0.02 |
ENSMUST00000071858.3
|
Hpcal1
|
hippocalcin-like 1 |
| chr2_+_92184106 | 0.02 |
ENSMUST00000111294.1
ENSMUST00000111293.2 ENSMUST00000162146.1 ENSMUST00000111292.1 ENSMUST00000162497.1 |
Phf21a
|
PHD finger protein 21A |
| chr19_+_38264761 | 0.02 |
ENSMUST00000087252.5
|
Lgi1
|
leucine-rich repeat LGI family, member 1 |
| chr19_-_8786408 | 0.02 |
ENSMUST00000176496.1
|
Taf6l
|
TAF6-like RNA polymerase II, p300/CBP-associated factor (PCAF)-associated factor |
| chr10_-_81350305 | 0.02 |
ENSMUST00000167481.1
|
Hmg20b
|
high mobility group 20B |
| chr2_+_48949495 | 0.02 |
ENSMUST00000112745.1
|
Mbd5
|
methyl-CpG binding domain protein 5 |
| chr10_-_81350389 | 0.02 |
ENSMUST00000020454.4
ENSMUST00000105324.2 ENSMUST00000154609.2 ENSMUST00000105323.1 |
Hmg20b
|
high mobility group 20B |
| chr16_+_64851991 | 0.01 |
ENSMUST00000067744.7
|
Cggbp1
|
CGG triplet repeat binding protein 1 |
| chr10_-_81350191 | 0.01 |
ENSMUST00000122993.1
|
Hmg20b
|
high mobility group 20B |
| chr19_-_21652714 | 0.01 |
ENSMUST00000177577.1
|
1110059E24Rik
|
RIKEN cDNA 1110059E24 gene |
| chr9_+_108306205 | 0.01 |
ENSMUST00000007959.8
|
Rhoa
|
ras homolog gene family, member A |
| chr5_+_145084100 | 0.01 |
ENSMUST00000124379.1
|
Arpc1a
|
actin related protein 2/3 complex, subunit 1A |
| chr2_-_66634952 | 0.01 |
ENSMUST00000100064.2
ENSMUST00000100063.2 |
Scn9a
|
sodium channel, voltage-gated, type IX, alpha |
| chr1_+_167001457 | 0.01 |
ENSMUST00000126198.1
|
Fam78b
|
family with sequence similarity 78, member B |
| chr9_+_65630552 | 0.01 |
ENSMUST00000055844.8
|
Rbpms2
|
RNA binding protein with multiple splicing 2 |
| chr9_+_108347827 | 0.01 |
ENSMUST00000035237.6
|
Usp4
|
ubiquitin specific peptidase 4 (proto-oncogene) |
| chr6_+_4747306 | 0.01 |
ENSMUST00000175823.1
ENSMUST00000176204.1 ENSMUST00000166678.1 |
Peg10
|
paternally expressed 10 |
| chr9_+_118926453 | 0.01 |
ENSMUST00000073109.5
|
Ctdspl
|
CTD (carboxy-terminal domain, RNA polymerase II, polypeptide A) small phosphatase-like |
| chr11_-_96777492 | 0.00 |
ENSMUST00000127375.1
ENSMUST00000021246.2 ENSMUST00000107661.3 |
Snx11
|
sorting nexin 11 |
| chr7_-_63938862 | 0.00 |
ENSMUST00000063694.8
|
Klf13
|
Kruppel-like factor 13 |
| chr17_+_34647128 | 0.00 |
ENSMUST00000015605.8
ENSMUST00000182587.1 |
Atf6b
|
activating transcription factor 6 beta |
| chr1_-_52500679 | 0.00 |
ENSMUST00000069792.7
|
Nab1
|
Ngfi-A binding protein 1 |
| chr13_-_55513427 | 0.00 |
ENSMUST00000069929.6
ENSMUST00000069968.6 ENSMUST00000131306.1 ENSMUST00000046246.6 |
Pdlim7
|
PDZ and LIM domain 7 |
Network of associatons between targets according to the STRING database.
First level regulatory network of Egr4
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.8 | GO:2000327 | regulation of ligand-dependent nuclear receptor transcription coactivator activity(GO:2000325) positive regulation of ligand-dependent nuclear receptor transcription coactivator activity(GO:2000327) |
| 0.1 | 0.3 | GO:2001076 | negative regulation of metanephric glomerulus development(GO:0072299) negative regulation of metanephric glomerular mesangial cell proliferation(GO:0072302) regulation of metanephric ureteric bud development(GO:2001074) positive regulation of metanephric ureteric bud development(GO:2001076) |
| 0.1 | 0.3 | GO:0002572 | pro-T cell differentiation(GO:0002572) |
| 0.1 | 0.2 | GO:0021852 | pyramidal neuron migration(GO:0021852) |
| 0.1 | 0.3 | GO:0008291 | acetylcholine metabolic process(GO:0008291) acetate ester metabolic process(GO:1900619) |
| 0.1 | 0.2 | GO:0002270 | plasmacytoid dendritic cell activation(GO:0002270) |
| 0.1 | 0.3 | GO:2000623 | regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000622) negative regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000623) |
| 0.1 | 0.2 | GO:0098501 | polynucleotide dephosphorylation(GO:0098501) |
| 0.0 | 0.2 | GO:0006114 | glycerol biosynthetic process(GO:0006114) |
| 0.0 | 0.1 | GO:0048320 | axial mesoderm formation(GO:0048320) |
| 0.0 | 0.1 | GO:1905077 | negative regulation of interleukin-17 secretion(GO:1905077) |
| 0.0 | 0.1 | GO:0045358 | negative regulation of interferon-beta biosynthetic process(GO:0045358) |
| 0.0 | 0.1 | GO:2000384 | regulation of ectoderm development(GO:2000383) negative regulation of ectoderm development(GO:2000384) |
| 0.0 | 0.1 | GO:0010360 | negative regulation of anion channel activity(GO:0010360) |
| 0.0 | 0.1 | GO:1900126 | negative regulation of hyaluronan biosynthetic process(GO:1900126) |
| 0.0 | 0.1 | GO:0007197 | adenylate cyclase-inhibiting G-protein coupled acetylcholine receptor signaling pathway(GO:0007197) phospholipase C-activating G-protein coupled acetylcholine receptor signaling pathway(GO:0007207) |
| 0.0 | 0.3 | GO:0045292 | mRNA cis splicing, via spliceosome(GO:0045292) |
| 0.0 | 0.2 | GO:0010764 | negative regulation of fibroblast migration(GO:0010764) |
| 0.0 | 0.5 | GO:0006783 | heme biosynthetic process(GO:0006783) |
| 0.0 | 0.3 | GO:0050860 | negative regulation of T cell receptor signaling pathway(GO:0050860) |
| 0.0 | 0.0 | GO:0003285 | septum secundum development(GO:0003285) atrial cardiac muscle cell differentiation(GO:0055011) atrial cardiac muscle cell development(GO:0055014) |
| 0.0 | 0.1 | GO:0060373 | regulation of atrial cardiac muscle cell membrane depolarization(GO:0060371) regulation of ventricular cardiac muscle cell membrane depolarization(GO:0060373) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | GO:0043083 | synaptic cleft(GO:0043083) |
| 0.0 | 0.2 | GO:0019907 | cyclin-dependent protein kinase activating kinase holoenzyme complex(GO:0019907) |
| 0.0 | 0.1 | GO:0031523 | Myb complex(GO:0031523) |
| 0.0 | 0.1 | GO:0089701 | U2AF(GO:0089701) |
| 0.0 | 0.2 | GO:0072687 | meiotic spindle(GO:0072687) |
| 0.0 | 0.2 | GO:0042622 | photoreceptor outer segment membrane(GO:0042622) |
| 0.0 | 0.0 | GO:0002111 | BRCA2-BRAF35 complex(GO:0002111) |
| 0.0 | 0.1 | GO:0045298 | tubulin complex(GO:0045298) |
| 0.0 | 0.3 | GO:1990124 | messenger ribonucleoprotein complex(GO:1990124) |
| 0.0 | 0.1 | GO:1990909 | Wnt signalosome(GO:1990909) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0004104 | cholinesterase activity(GO:0004104) |
| 0.1 | 0.2 | GO:0000402 | open form four-way junction DNA binding(GO:0000401) crossed form four-way junction DNA binding(GO:0000402) |
| 0.1 | 0.3 | GO:0098519 | nucleotide phosphatase activity, acting on free nucleotides(GO:0098519) |
| 0.0 | 0.3 | GO:0044729 | hemi-methylated DNA-binding(GO:0044729) |
| 0.0 | 0.1 | GO:0004703 | G-protein coupled receptor kinase activity(GO:0004703) |
| 0.0 | 0.8 | GO:0032183 | SUMO binding(GO:0032183) |
| 0.0 | 0.1 | GO:0016907 | G-protein coupled acetylcholine receptor activity(GO:0016907) |
| 0.0 | 0.1 | GO:0000828 | inositol hexakisphosphate kinase activity(GO:0000828) inositol hexakisphosphate 5-kinase activity(GO:0000832) inositol hexakisphosphate 1-kinase activity(GO:0052723) inositol hexakisphosphate 3-kinase activity(GO:0052724) |
| 0.0 | 0.1 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.0 | 0.1 | GO:0001093 | TFIIB-class transcription factor binding(GO:0001093) |
| 0.0 | 0.1 | GO:0086006 | voltage-gated sodium channel activity involved in cardiac muscle cell action potential(GO:0086006) |
| 0.0 | 0.3 | GO:0070530 | K63-linked polyubiquitin binding(GO:0070530) |
| 0.0 | 0.1 | GO:0050733 | RS domain binding(GO:0050733) |
| 0.0 | 0.3 | GO:0070064 | proline-rich region binding(GO:0070064) |
| 0.0 | 0.3 | GO:0008143 | poly(A) binding(GO:0008143) |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | REACTOME APOPTOSIS INDUCED DNA FRAGMENTATION | Genes involved in Apoptosis induced DNA fragmentation |
| 0.0 | 0.3 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |