Project
avrg: GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
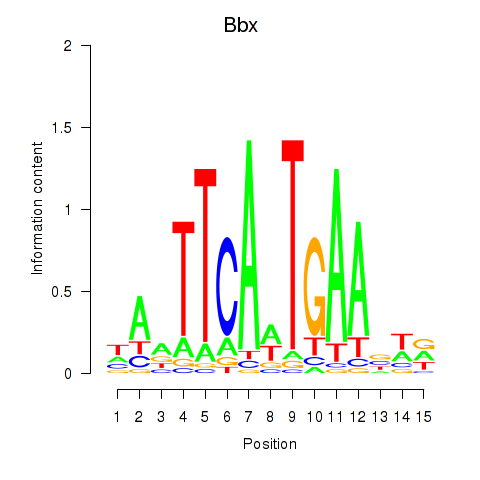
Results for Bbx
Z-value: 0.41
Transcription factors associated with Bbx
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Bbx
|
ENSMUSG00000022641.9 | bobby sox HMG box containing |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Bbx | mm10_v2_chr16_-_50330987_50331017 | -0.22 | 1.9e-01 | Click! |
Activity profile of Bbx motif
Sorted Z-values of Bbx motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr19_-_7966000 | 1.94 |
ENSMUST00000182102.1
ENSMUST00000075619.4 |
Slc22a27
|
solute carrier family 22, member 27 |
| chr19_-_11081088 | 1.35 |
ENSMUST00000025636.6
|
Ms4a8a
|
membrane-spanning 4-domains, subfamily A, member 8A |
| chr8_+_45658666 | 1.06 |
ENSMUST00000134675.1
ENSMUST00000139869.1 ENSMUST00000126067.1 |
Sorbs2
|
sorbin and SH3 domain containing 2 |
| chr2_+_58754910 | 1.03 |
ENSMUST00000059102.6
|
Upp2
|
uridine phosphorylase 2 |
| chr6_+_124304646 | 1.02 |
ENSMUST00000112541.2
ENSMUST00000032234.2 |
Cd163
|
CD163 antigen |
| chr1_-_13660476 | 0.97 |
ENSMUST00000027071.5
|
Lactb2
|
lactamase, beta 2 |
| chr15_-_3303521 | 0.95 |
ENSMUST00000165386.1
|
Ccdc152
|
coiled-coil domain containing 152 |
| chr8_+_45658273 | 0.91 |
ENSMUST00000153798.1
|
Sorbs2
|
sorbin and SH3 domain containing 2 |
| chr7_+_51880312 | 0.77 |
ENSMUST00000145049.1
|
Gas2
|
growth arrest specific 2 |
| chr5_-_108675569 | 0.70 |
ENSMUST00000051757.7
|
Slc26a1
|
solute carrier family 26 (sulfate transporter), member 1 |
| chr8_+_45658731 | 0.68 |
ENSMUST00000143820.1
ENSMUST00000132139.1 |
Sorbs2
|
sorbin and SH3 domain containing 2 |
| chr18_+_37421418 | 0.60 |
ENSMUST00000053073.4
|
Pcdhb11
|
protocadherin beta 11 |
| chr7_+_127800604 | 0.57 |
ENSMUST00000046863.5
ENSMUST00000106272.1 ENSMUST00000139068.1 |
Hsd3b7
|
hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 7 |
| chr2_-_134554348 | 0.48 |
ENSMUST00000028704.2
|
Hao1
|
hydroxyacid oxidase 1, liver |
| chr8_-_41054771 | 0.43 |
ENSMUST00000093534.4
|
Mtus1
|
mitochondrial tumor suppressor 1 |
| chr7_-_127708886 | 0.38 |
ENSMUST00000061468.8
|
Bcl7c
|
B cell CLL/lymphoma 7C |
| chr15_-_101892916 | 0.36 |
ENSMUST00000100179.1
|
Krt76
|
keratin 76 |
| chr3_+_16183177 | 0.35 |
ENSMUST00000108345.2
ENSMUST00000108346.2 |
Ythdf3
|
YTH domain family 3 |
| chrX_-_75161620 | 0.34 |
ENSMUST00000165080.1
|
Smim9
|
small integral membrane protein 9 |
| chr6_+_139621888 | 0.28 |
ENSMUST00000032353.8
|
Pik3c2g
|
phosphatidylinositol 3-kinase, C2 domain containing, gamma polypeptide |
| chr11_+_105146893 | 0.27 |
ENSMUST00000100338.1
|
Gm10842
|
predicted gene 10842 |
| chr10_-_107486077 | 0.25 |
ENSMUST00000000445.1
|
Myf5
|
myogenic factor 5 |
| chr4_+_80910646 | 0.24 |
ENSMUST00000055922.3
|
Lurap1l
|
leucine rich adaptor protein 1-like |
| chr7_-_127708738 | 0.23 |
ENSMUST00000106282.1
|
Bcl7c
|
B cell CLL/lymphoma 7C |
| chr2_-_155592567 | 0.22 |
ENSMUST00000155347.1
ENSMUST00000130881.1 ENSMUST00000079691.6 |
Gss
|
glutathione synthetase |
| chr1_-_156032948 | 0.19 |
ENSMUST00000136397.1
|
Tor1aip1
|
torsin A interacting protein 1 |
| chrX_-_107816238 | 0.16 |
ENSMUST00000120722.1
|
2610002M06Rik
|
RIKEN cDNA 2610002M06 gene |
| chr16_+_88728828 | 0.16 |
ENSMUST00000060494.6
|
Krtap13-1
|
keratin associated protein 13-1 |
| chr12_+_88027004 | 0.16 |
ENSMUST00000178852.1
|
Gm2056
|
predicted gene 2056 |
| chr12_-_87919857 | 0.15 |
ENSMUST00000180053.1
|
Gm2035
|
predicted pseudogene 2035 |
| chr5_-_146585239 | 0.14 |
ENSMUST00000036211.6
|
Gpr12
|
G-protein coupled receptor 12 |
| chr2_+_136892168 | 0.12 |
ENSMUST00000099311.2
|
Slx4ip
|
SLX4 interacting protein |
| chr1_-_163725123 | 0.12 |
ENSMUST00000159679.1
|
Mettl11b
|
methyltransferase like 11B |
| chr5_+_104202609 | 0.10 |
ENSMUST00000066708.5
|
Dmp1
|
dentin matrix protein 1 |
| chr13_-_91807658 | 0.09 |
ENSMUST00000022121.6
|
Zcchc9
|
zinc finger, CCHC domain containing 9 |
| chr11_+_59306920 | 0.08 |
ENSMUST00000000128.3
ENSMUST00000108783.3 |
Wnt9a
|
wingless-type MMTV integration site 9A |
| chr13_+_49608030 | 0.08 |
ENSMUST00000021822.5
|
Ogn
|
osteoglycin |
| chr3_-_37232565 | 0.08 |
ENSMUST00000161015.1
ENSMUST00000029273.1 |
Il21
|
interleukin 21 |
| chr1_+_180111339 | 0.04 |
ENSMUST00000145181.1
|
Cdc42bpa
|
CDC42 binding protein kinase alpha |
| chr12_+_88329254 | 0.04 |
ENSMUST00000091715.1
|
Gm10264
|
predicted gene 10264 |
| chr18_-_65970178 | 0.03 |
ENSMUST00000025397.5
|
Cplx4
|
complexin 4 |
Network of associatons between targets according to the STRING database.
First level regulatory network of Bbx
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.9 | GO:0015747 | urate transport(GO:0015747) |
| 0.1 | 0.5 | GO:0009441 | glycolate metabolic process(GO:0009441) |
| 0.1 | 0.7 | GO:0019532 | oxalate transport(GO:0019532) |
| 0.1 | 0.6 | GO:0035754 | B cell chemotaxis(GO:0035754) |
| 0.0 | 2.7 | GO:0061049 | physiological muscle hypertrophy(GO:0003298) physiological cardiac muscle hypertrophy(GO:0003301) cell growth involved in cardiac muscle cell development(GO:0061049) |
| 0.0 | 1.0 | GO:0090502 | RNA phosphodiester bond hydrolysis, endonucleolytic(GO:0090502) |
| 0.0 | 0.1 | GO:0070173 | regulation of enamel mineralization(GO:0070173) |
| 0.0 | 0.2 | GO:0048743 | positive regulation of skeletal muscle fiber development(GO:0048743) |
| 0.0 | 0.1 | GO:0002729 | positive regulation of natural killer cell cytokine production(GO:0002729) |
| 0.0 | 0.4 | GO:0048733 | sebaceous gland development(GO:0048733) |
| 0.0 | 0.2 | GO:0071763 | nuclear membrane organization(GO:0071763) |
| 0.0 | 1.0 | GO:0006953 | acute-phase response(GO:0006953) |
| 0.0 | 0.3 | GO:0045948 | positive regulation of translational initiation(GO:0045948) |
| 0.0 | 0.4 | GO:0010758 | regulation of macrophage chemotaxis(GO:0010758) |
| 0.0 | 0.3 | GO:0039703 | viral RNA genome replication(GO:0039694) RNA replication(GO:0039703) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.0 | GO:0045098 | type III intermediate filament(GO:0045098) |
| 0.0 | 0.4 | GO:0045095 | keratin filament(GO:0045095) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.0 | GO:0004850 | uridine phosphorylase activity(GO:0004850) |
| 0.1 | 1.9 | GO:1901702 | urate transmembrane transporter activity(GO:0015143) salt transmembrane transporter activity(GO:1901702) |
| 0.1 | 0.5 | GO:0016899 | oxidoreductase activity, acting on the CH-OH group of donors, oxygen as acceptor(GO:0016899) |
| 0.0 | 0.6 | GO:0003854 | 3-beta-hydroxy-delta5-steroid dehydrogenase activity(GO:0003854) |
| 0.0 | 0.7 | GO:0019531 | oxalate transmembrane transporter activity(GO:0019531) |
| 0.0 | 0.3 | GO:1990247 | N6-methyladenosine-containing RNA binding(GO:1990247) |
| 0.0 | 1.0 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.0 | 0.3 | GO:0035005 | 1-phosphatidylinositol-4-phosphate 3-kinase activity(GO:0035005) |
| 0.0 | 1.0 | GO:0004521 | endoribonuclease activity(GO:0004521) |
| 0.0 | 0.1 | GO:0005134 | interleukin-2 receptor binding(GO:0005134) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.8 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.0 | REACTOME PYRIMIDINE CATABOLISM | Genes involved in Pyrimidine catabolism |
| 0.0 | 0.6 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 24 HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 24-hydroxycholesterol |
| 0.0 | 0.8 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.0 | 0.3 | REACTOME SYNTHESIS OF PIPS AT THE GOLGI MEMBRANE | Genes involved in Synthesis of PIPs at the Golgi membrane |