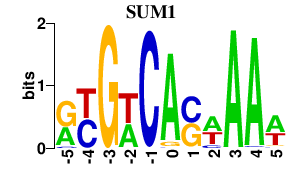
Results for SUM1
Z-value: 4.01
Transcription factors associated with SUM1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
SUM1
|
S000002718 | Transcriptional repressor that regulates middle-sporulation genes |
Activity-expression correlation:
Activity profile of SUM1 motif
Sorted Z-values of SUM1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| YLR307C-A | 62.68 |
Putative protein of unknown function |
||
| YAL062W | 46.80 |
GDH3
|
NADP(+)-dependent glutamate dehydrogenase, synthesizes glutamate from ammonia and alpha-ketoglutarate; rate of alpha-ketoglutarate utilization differs from Gdh1p; expression regulated by nitrogen and carbon sources |
|
| YCR005C | 43.23 |
CIT2
|
Citrate synthase, catalyzes the condensation of acetyl coenzyme A and oxaloacetate to form citrate, peroxisomal isozyme involved in glyoxylate cycle; expression is controlled by Rtg1p and Rtg2p transcription factors |
|
| YDL210W | 26.24 |
UGA4
|
Permease that serves as a gamma-aminobutyrate (GABA) transport protein involved in the utilization of GABA as a nitrogen source; catalyzes the transport of putrescine and delta-aminolevulinic acid (ALA); localized to the vacuolar membrane |
|
| YJL037W | 24.34 |
IRC18
|
Putative protein of unknown function; expression induced in respiratory-deficient cells and in carbon-limited chemostat cultures; similar to adjacent ORF, YJL038C; null mutant displays increased levels of spontaneous Rad52p foci |
|
| YJL133C-A | 23.82 |
Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies |
||
| YGR256W | 23.02 |
GND2
|
6-phosphogluconate dehydrogenase (decarboxylating), catalyzes an NADPH regenerating reaction in the pentose phosphate pathway; required for growth on D-glucono-delta-lactone |
|
| YPL240C | 22.50 |
HSP82
|
Hsp90 chaperone required for pheromone signaling and negative regulation of Hsf1p; docks with Tom70p for mitochondrial preprotein delivery; promotes telomerase DNA binding and nucleotide addition; interacts with Cns1p, Cpr6p, Cpr7p, Sti1p |
|
| YKL177W | 20.88 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; partially overlaps the verified gene STE3 |
||
| YAR035W | 20.01 |
YAT1
|
Outer mitochondrial carnitine acetyltransferase, minor ethanol-inducible enzyme involved in transport of activated acyl groups from the cytoplasm into the mitochondrial matrix; phosphorylated |
|
| YAR053W | 19.16 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data |
||
| YGR087C | 19.06 |
PDC6
|
Minor isoform of pyruvate decarboxylase, decarboxylates pyruvate to acetaldehyde, involved in amino acid catabolism; transcription is glucose- and ethanol-dependent, and is strongly induced during sulfur limitation |
|
| YOR072W | 18.81 |
Dubious open reading frame unlikely to encode a protein, based on experimental and comparative sequence data; partially overlaps the dubious gene YOR072W-A; diploid deletion strains are methotrexate, paraquat and wortmannin sensitive |
||
| YBR117C | 18.43 |
TKL2
|
Transketolase, similar to Tkl1p; catalyzes conversion of xylulose-5-phosphate and ribose-5-phosphate to sedoheptulose-7-phosphate and glyceraldehyde-3-phosphate in the pentose phosphate pathway; needed for synthesis of aromatic amino acids |
|
| YLR327C | 17.79 |
TMA10
|
Protein of unknown function that associates with ribosomes |
|
| YAL067C | 17.22 |
SEO1
|
Putative permease, member of the allantoate transporter subfamily of the major facilitator superfamily; mutation confers resistance to ethionine sulfoxide |
|
| YJR078W | 16.85 |
BNA2
|
Putative tryptophan 2,3-dioxygenase or indoleamine 2,3-dioxygenase, required for the de novo biosynthesis of NAD from tryptophan via kynurenine; expression regulated by Hst1p and Aft2p |
|
| YDL223C | 16.65 |
HBT1
|
Substrate of the Hub1p ubiquitin-like protein that localizes to the shmoo tip (mating projection); mutants are defective for mating projection formation, thereby implicating Hbt1p in polarized cell morphogenesis |
|
| YNL318C | 16.18 |
HXT14
|
Protein with similarity to hexose transporter family members, expression is induced in low glucose and repressed in high glucose; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies |
|
| YCL048W | 16.01 |
SPS22
|
Protein of unknown function, redundant with Sps2p for the organization of the beta-glucan layer of the spore wall |
|
| YKL178C | 15.80 |
STE3
|
Receptor for a factor receptor, transcribed in alpha cells and required for mating by alpha cells, couples to MAP kinase cascade to mediate pheromone response; ligand bound receptors are endocytosed and recycled to the plasma membrane; GPCR |
|
| YHR212C | 15.70 |
Dubious open reading frame unlikely to encode a functional protein, based on available experimental and comparative sequence data |
||
| YOR071C | 15.64 |
NRT1
|
High-affinity nicotinamide riboside transporter; also transports thiamine with low affinity; shares sequence similarity with Thi7p and Thi72p; proposed to be involved in 5-fluorocytosine sensitivity |
|
| YHR126C | 15.56 |
Putative protein of unknown function; transcription dependent upon Azf1p |
||
| YMR256C | 15.55 |
COX7
|
Subunit VII of cytochrome c oxidase, which is the terminal member of the mitochondrial inner membrane electron transport chain |
|
| YKR102W | 15.49 |
FLO10
|
Lectin-like protein with similarity to Flo1p, thought to be involved in flocculation |
|
| YER065C | 15.49 |
ICL1
|
Isocitrate lyase, catalyzes the formation of succinate and glyoxylate from isocitrate, a key reaction of the glyoxylate cycle; expression of ICL1 is induced by growth on ethanol and repressed by growth on glucose |
|
| YFR023W | 15.18 |
PES4
|
Poly(A) binding protein, suppressor of DNA polymerase epsilon mutation, similar to Mip6p |
|
| YFL011W | 15.03 |
HXT10
|
Putative hexose transporter, expressed at low levels and expression is repressed by glucose |
|
| YHR053C | 15.03 |
CUP1-1
|
Metallothionein, binds copper and mediates resistance to high concentrations of copper and cadmium; locus is variably amplified in different strains, with two copies, CUP1-1 and CUP1-2, in the genomic sequence reference strain S288C |
|
| YJL038C | 14.91 |
LOH1
|
Putative protein of unknown function; expression induced during sporulation and repressed during vegetative growth by Sum1p and Hst1p; similar to adjacent open reading frame, YJL037W |
|
| YBR296C | 14.68 |
PHO89
|
Na+/Pi cotransporter, active in early growth phase; similar to phosphate transporters of Neurospora crassa; transcription regulated by inorganic phosphate concentrations and Pho4p |
|
| YIL160C | 14.61 |
POT1
|
3-ketoacyl-CoA thiolase with broad chain length specificity, cleaves 3-ketoacyl-CoA into acyl-CoA and acetyl-CoA during beta-oxidation of fatty acids |
|
| YOR072W-A | 14.12 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; partially overlaps the uncharacterized ORF YOR072W; originally identified by fungal homology and RT-PCR |
||
| YER024W | 14.05 |
YAT2
|
Carnitine acetyltransferase; has similarity to Yat1p, which is a carnitine acetyltransferase associated with the mitochondrial outer membrane |
|
| YAL063C | 13.89 |
FLO9
|
Lectin-like protein with similarity to Flo1p, thought to be expressed and involved in flocculation |
|
| YAR060C | 13.67 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data |
||
| YER150W | 13.63 |
SPI1
|
GPI-anchored cell wall protein involved in weak acid resistance; basal expression requires Msn2p/Msn4p; expression is induced under conditions of stress and during the diauxic shift; similar to Sed1p |
|
| YHR212W-A | 13.58 |
Putative protein of unknown function; identified by gene-trapping, microarray-based expression analysis, and genome-wide homology searching |
||
| YBR076W | 13.55 |
ECM8
|
Non-essential protein of unknown function |
|
| YCL054W | 13.28 |
SPB1
|
AdoMet-dependent methyltransferase involved in rRNA processing and 60S ribosomal subunit maturation; methylates G2922 in the tRNA docking site of the large subunit rRNA and in the absence of snR52, U2921; suppressor of PAB1 mutants |
|
| YLR004C | 13.15 |
THI73
|
Putative plasma membrane permease proposed to be involved in carboxylic acid uptake and repressed by thiamine; substrate of Dbf2p/Mob1p kinase; transcription is altered if mitochondrial dysfunction occurs |
|
| YPR078C | 13.06 |
Putative protein of unknown function; possible role in DNA metabolism and/or in genome stability; expression is heat-inducible |
||
| YDR218C | 13.03 |
SPR28
|
Sporulation-specific homolog of the yeast CDC3/10/11/12 family of bud neck microfilament genes; meiotic septin expressed at high levels during meiotic divisions and ascospore formation |
|
| YGR258C | 12.96 |
RAD2
|
Single-stranded DNA endonuclease, cleaves single-stranded DNA during nucleotide excision repair to excise damaged DNA; subunit of Nucleotide Excision Repair Factor 3 (NEF3); homolog of human XPG protein |
|
| YGL187C | 12.89 |
COX4
|
Subunit IV of cytochrome c oxidase, which is the terminal member of the mitochondrial inner membrane electron transport chain; N-terminal 25 residues of precursor are cleaved during mitochondrial import; phosphorylated |
|
| YGR059W | 12.86 |
SPR3
|
Sporulation-specific homolog of the yeast CDC3/10/11/12 family of bud neck microfilament genes; septin protein involved in sporulation; regulated by ABFI |
|
| YDR536W | 12.80 |
STL1
|
Glycerol proton symporter of the plasma membrane, subject to glucose-induced inactivation, strongly but transiently induced when cells are subjected to osmotic shock |
|
| YOR211C | 12.42 |
MGM1
|
Mitochondrial GTPase related to dynamin, present in a complex containing Ugo1p and Fzo1p; required for normal morphology of cristae and for stability of Tim11p; homolog of human OPA1 involved in autosomal dominant optic atrophy |
|
| YNL202W | 12.32 |
SPS19
|
Peroxisomal 2,4-dienoyl-CoA reductase, auxiliary enzyme of fatty acid beta-oxidation; homodimeric enzyme required for growth and sporulation on petroselineate medium; expression induced during late sporulation and in the presence of oleate |
|
| YLR136C | 11.77 |
TIS11
|
mRNA-binding protein expressed during iron starvation; binds to a sequence element in the 3'-untranslated regions of specific mRNAs to mediate their degradation; involved in iron homeostasis |
|
| YBR072W | 11.74 |
HSP26
|
Small heat shock protein (sHSP) with chaperone activity; forms hollow, sphere-shaped oligomers that suppress unfolded proteins aggregation; oligomer activation requires a heat-induced conformational change; not expressed in unstressed cells |
|
| YPL271W | 11.72 |
ATP15
|
Epsilon subunit of the F1 sector of mitochondrial F1F0 ATP synthase, which is a large, evolutionarily conserved enzyme complex required for ATP synthesis; phosphorylated |
|
| YFL052W | 11.64 |
Putative zinc cluster protein that contains a DNA binding domain; null mutant sensitive to calcofluor white, low osmolarity and heat, suggesting a role for YFL052Wp in cell wall integrity |
||
| YMR056C | 11.58 |
AAC1
|
Mitochondrial inner membrane ADP/ATP translocator, exchanges cytosolic ADP for mitochondrially synthesized ATP; phosphorylated; Aac1p is a minor isoform while Pet9p is the major ADP/ATP translocator |
|
| YOR381W | 11.39 |
FRE3
|
Ferric reductase, reduces siderophore-bound iron prior to uptake by transporters; expression induced by low iron levels |
|
| YJR155W | 11.02 |
AAD10
|
Putative aryl-alcohol dehydrogenase with similarity to P. chrysosporium aryl-alcohol dehydrogenase; mutational analysis has not yet revealed a physiological role |
|
| YAR069C | 11.00 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data |
||
| YHR055C | 11.00 |
CUP1-2
|
Metallothionein, binds copper and mediates resistance to high concentrations of copper and cadmium; locus is variably amplified in different strains, with two copies, CUP1-1 and CUP1-2, in the genomic sequence reference strain S288C |
|
| YGL255W | 10.97 |
ZRT1
|
High-affinity zinc transporter of the plasma membrane, responsible for the majority of zinc uptake; transcription is induced under low-zinc conditions by the Zap1p transcription factor |
|
| YBR200W-A | 10.92 |
Putative protein of unknown function; identified by fungal homology and RT-PCR |
||
| YLR308W | 10.86 |
CDA2
|
Chitin deacetylase, together with Cda1p involved in the biosynthesis ascospore wall component, chitosan; required for proper rigidity of the ascospore wall |
|
| YHR015W | 10.76 |
MIP6
|
Putative RNA-binding protein, interacts with Mex67p, which is a component of the nuclear pore involved in nuclear mRNA export |
|
| YPL027W | 10.65 |
SMA1
|
Protein of unknown function involved in the assembly of the prospore membrane during sporulation |
|
| YAR050W | 10.52 |
FLO1
|
Lectin-like protein involved in flocculation, cell wall protein that binds to mannose chains on the surface of other cells, confers floc-forming ability that is chymotrypsin sensitive and heat resistant; similar to Flo5p |
|
| YJL043W | 10.51 |
Putative protein of unknown function; YJL043W is a non-essential gene |
||
| YAR070C | 10.47 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data |
||
| YHR054W-A | 10.44 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; partially overlaps CUP1-2 |
||
| YDL114W | 10.43 |
Putative protein of unknown function with similarity to acyl-carrier-protein reductases; YDL114W is not an essential gene |
||
| YMR279C | 10.26 |
Putative protein of unknown function; identified as a heat-induced gene in a high-throughout screen; YMR279C is not an essential gene |
||
| YGR142W | 10.24 |
BTN2
|
v-SNARE binding protein that facilitates specific protein retrieval from a late endosome to the Golgi; modulates arginine uptake, possible role in mediating pH homeostasis between the vacuole and plasma membrane H(+)-ATPase |
|
| YPL033C | 10.16 |
Putative protein of unknown function; may be involved in DNA metabolism; expression is induced by Kar4p |
||
| YEL059W | 10.11 |
Dubious open reading frame unlikely to encode a functional protein |
||
| YBR035C | 10.08 |
PDX3
|
Pyridoxine (pyridoxamine) phosphate oxidase, has homologs in E. coli and Myxococcus xanthus; transcription is under the general control of nitrogen metabolism |
|
| YKR033C | 10.02 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; partially overlaps the verified gene DAL80 |
||
| YGR088W | 9.98 |
CTT1
|
Cytosolic catalase T, has a role in protection from oxidative damage by hydrogen peroxide |
|
| YKR015C | 9.97 |
Putative protein of unknown function |
||
| YAL018C | 9.87 |
Putative protein of unknown function |
||
| YLL005C | 9.84 |
SPO75
|
Meiosis-specific protein of unknown function, required for spore wall formation during sporulation; dispensable for both nuclear divisions during meiosis |
|
| YFL012W | 9.83 |
Putative protein of unknown function; transcribed during sporulation; null mutant exhibits increased resistance to rapamycin |
||
| YOR347C | 9.53 |
PYK2
|
Pyruvate kinase that appears to be modulated by phosphorylation; PYK2 transcription is repressed by glucose, and Pyk2p may be active under low glycolytic flux |
|
| YMR141C | 9.43 |
Dubious open reading frame unlikely to encode a functional protein, based on available experimental and comparative sequence data |
||
| YDR540C | 9.40 |
IRC4
|
Putative protein of unknown function; null mutant displays increased levels of spontaneous Rad52p foci; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus |
|
| YOL118C | 9.26 |
Dubious open reading frame, unlikely to encode a functional protein; based on available experimental and comparative sequence data |
||
| YGL138C | 9.19 |
Putative protein of unknown function; has no significant sequence similarity to any known protein |
||
| YOL084W | 8.99 |
PHM7
|
Protein of unknown function, expression is regulated by phosphate levels; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery and vacuole |
|
| YER106W | 8.91 |
MAM1
|
Monopolin, kinetochore associated protein involved in chromosome attachment to meiotic spindle |
|
| YPR191W | 8.84 |
QCR2
|
Subunit 2 of the ubiquinol cytochrome-c reductase complex, which is a component of the mitochondrial inner membrane electron transport chain; phosphorylated; transcription is regulated by Hap1p, Hap2p/Hap3p, and heme |
|
| YER085C | 8.67 |
Putative protein of unknown function |
||
| YOR346W | 8.61 |
REV1
|
Deoxycytidyl transferase, forms a complex with the subunits of DNA polymerase zeta, Rev3p and Rev7p; involved in repair of abasic sites in damaged DNA |
|
| YHR211W | 8.57 |
FLO5
|
Lectin-like cell wall protein (flocculin) involved in flocculation, binds to mannose chains on the surface of other cells, confers floc-forming ability that is chymotrypsin resistant but heat labile; similar to Flo1p |
|
| YAR047C | 8.52 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data |
||
| YBR201C-A | 8.49 |
Putative protein of unknown function |
||
| YMR272C | 8.46 |
SCS7
|
Sphingolipid alpha-hydroxylase, functions in the alpha-hydroxylation of sphingolipid-associated very long chain fatty acids, has both cytochrome b5-like and hydroxylase/desaturase domains, not essential for growth |
|
| YDR310C | 8.36 |
SUM1
|
Transcriptional repressor required for mitotic repression of middle sporulation-specific genes; involved in telomere maintenance, regulated by the pachytene checkpoint |
|
| YGL170C | 8.29 |
SPO74
|
Component of the meiotic outer plaque of the spindle pole body, involved in modifying the meiotic outer plaque that is required prior to prospore membrane formation |
|
| YPR013C | 8.27 |
Putative zinc finger protein; YPR013C is not an essential gene |
||
| YOL060C | 8.16 |
MAM3
|
Protein required for normal mitochondrial morphology, has similarity to hemolysins |
|
| YOR345C | 8.15 |
Dubious ORF unlikely to encode a protein, based on available experimental and comparative sequence data; overlaps the verified gene REV1; null mutant displays increased resistance to antifungal agents gliotoxin, cycloheximide and H2O2 |
||
| YNR063W | 8.14 |
Putative zinc-cluster protein of unknown function |
||
| YDL224C | 8.13 |
WHI4
|
Putative RNA binding protein and partially redundant Whi3p homolog that regulates the cell size requirement for passage through Start and commitment to cell division |
|
| YGR236C | 8.06 |
SPG1
|
Protein required for survival at high temperature during stationary phase; not required for growth on nonfermentable carbon sources; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies |
|
| YAL066W | 8.06 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data |
||
| YNL018C | 7.89 |
Putative protein of unknown function |
||
| YJR077C | 7.84 |
MIR1
|
Mitochondrial phosphate carrier, imports inorganic phosphate into mitochondria; functionally redundant with Pic2p but more abundant than Pic2p under normal conditions; phosphorylated |
|
| YOR339C | 7.82 |
UBC11
|
Ubiquitin-conjugating enzyme most similar in sequence to Xenopus ubiquitin-conjugating enzyme E2-C, but not a true functional homolog of this E2; unlike E2-C, not required for the degradation of mitotic cyclin Clb2 |
|
| YCR007C | 7.73 |
Putative integral membrane protein, member of DUP240 gene family; YCR007C is not an essential gene |
||
| YDR022C | 7.73 |
CIS1
|
Autophagy-specific protein required for autophagosome formation; may form a complex with Atg17p and Atg29p that localizes other proteins to the pre-autophagosomal structure; high-copy suppressor of CIK1 deletion |
|
| YLR054C | 7.70 |
OSW2
|
Protein of unknown function proposed to be involved in the assembly of the spore wall |
|
| YJL213W | 7.62 |
Protein of unknown function that may interact with ribosomes; periodically expressed during the yeast metabolic cycle; phosphorylated in vitro by the mitotic exit network (MEN) kinase complex, Dbf2p/Mob1p |
||
| YOR255W | 7.61 |
OSW1
|
Protein involved in sporulation; required for the construction of the outer spore wall layers; required for proper localization of Spo14p |
|
| YKL163W | 7.55 |
PIR3
|
O-glycosylated covalently-bound cell wall protein required for cell wall stability; expression is cell cycle regulated, peaking in M/G1 and also subject to regulation by the cell integrity pathway |
|
| YJL116C | 7.43 |
NCA3
|
Protein that functions with Nca2p to regulate mitochondrial expression of subunits 6 (Atp6p) and 8 (Atp8p ) of the Fo-F1 ATP synthase; member of the SUN family |
|
| YOR173W | 7.39 |
DCS2
|
Non-essential, stress induced regulatory protein containing a HIT (histidine triad) motif; modulates m7G-oligoribonucleotide metabolism; inhibits Dcs1p; regulated by Msn2p, Msn4p, and the Ras-cAMP-cAPK signaling pathway, similar to Dcs1p. |
|
| YBR250W | 7.36 |
SPO23
|
Protein of unknown function; associates with meiosis-specific protein Spo1p |
|
| YFL030W | 7.21 |
AGX1
|
Alanine:glyoxylate aminotransferase (AGT), catalyzes the synthesis of glycine from glyoxylate, which is one of three pathways for glycine biosynthesis in yeast; has similarity to mammalian and plant alanine:glyoxylate aminotransferases |
|
| YGR191W | 7.18 |
HIP1
|
High-affinity histidine permease, also involved in the transport of manganese ions |
|
| YLR111W | 7.13 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data |
||
| YAR068W | 7.00 |
Fungal-specific protein of unknown function; induced in respiratory-deficient cells |
||
| YKL161C | 6.96 |
Protein kinase implicated in the Slt2p mitogen-activated (MAP) kinase signaling pathway; associates with Rlm1p |
||
| YLR213C | 6.87 |
CRR1
|
Putative glycoside hydrolase of the spore wall envelope; required for normal spore wall assembly, possibly for cross-linking between the glucan and chitosan layers; expressed during sporulation |
|
| YJR128W | 6.76 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; partially overlaps the verified ORF RSF2 |
||
| YBR148W | 6.67 |
YSW1
|
Protein expressed specifically in spores |
|
| YIL108W | 6.66 |
Putative metalloprotease |
||
| YIR027C | 6.61 |
DAL1
|
Allantoinase, converts allantoin to allantoate in the first step of allantoin degradation; expression sensitive to nitrogen catabolite repression |
|
| YPR054W | 6.57 |
SMK1
|
Middle sporulation-specific mitogen-activated protein kinase (MAPK) required for production of the outer spore wall layers; negatively regulates activity of the glucan synthase subunit Gsc2p |
|
| YGL180W | 6.54 |
ATG1
|
Protein serine/threonine kinase required for vesicle formation in autophagy and the cytoplasm-to-vacuole targeting (Cvt) pathway; structurally required for pre-autophagosome formation; during autophagy forms a complex with Atg13p and Atg17p |
|
| YOL047C | 6.48 |
Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern |
||
| YGL015C | 6.47 |
Hypothetical protein |
||
| YDR034C | 6.43 |
LYS14
|
Transcriptional activator involved in regulation of genes of the lysine biosynthesis pathway; requires 2-aminoadipate semialdehyde as co-inducer |
|
| YOR242C | 6.42 |
SSP2
|
Sporulation specific protein that localizes to the spore wall; required for sporulation at a point after meiosis II and during spore wall formation; SSP2 expression is induced midway in meiosis |
|
| YGR043C | 6.41 |
NQM1
|
Protein of unknown function; transcription is repressed by Mot1p and induced by alpha-factor and during diauxic shift; null mutant non-quiescent cells exhibit reduced reproductive capacity |
|
| YOR343C | 6.22 |
Dubious open reading frame, unlikely to encode a functional protein; based on available experimental and comparative sequence data |
||
| YKR016W | 6.19 |
AIM28
|
Protein of unknown function; non-tagged protein is detected in purified mitochondria in high-throughput studies; null mutant displays decreased frequency of mitochondrial genome loss and severe growth defect in minimal glycerol media |
|
| YDR281C | 6.12 |
PHM6
|
Protein of unknown function, expression is regulated by phosphate levels |
|
| YEL009C-A | 6.11 |
Dubious open reading frame unlikely to encode a functional protein, based on available experimental and comparative sequence data |
||
| YER088C | 6.10 |
DOT6
|
Protein of unknown function, involved in telomeric gene silencing and filamentation |
|
| YOL119C | 6.04 |
MCH4
|
Protein with similarity to mammalian monocarboxylate permeases, which are involved in transport of monocarboxylic acids across the plasma membrane; mutant is not deficient in monocarboxylate transport |
|
| YMR107W | 5.93 |
SPG4
|
Protein required for survival at high temperature during stationary phase; not required for growth on nonfermentable carbon sources |
|
| YOL132W | 5.88 |
GAS4
|
1,3-beta-glucanosyltransferase, involved with Gas2p in spore wall assembly; has similarity to Gas1p; localizes to the cell wall |
|
| YMR081C | 5.86 |
ISF1
|
Serine-rich, hydrophilic protein with similarity to Mbr1p; overexpression suppresses growth defects of hap2, hap3, and hap4 mutants; expression is under glucose control; cotranscribed with NAM7 in a cyp1 mutant |
|
| YJR151C | 5.85 |
DAN4
|
Cell wall mannoprotein with similarity to Tir1p, Tir2p, Tir3p, and Tir4p; expressed under anaerobic conditions, completely repressed during aerobic growth |
|
| YDR043C | 5.85 |
NRG1
|
Transcriptional repressor that recruits the Cyc8p-Tup1p complex to promoters; mediates glucose repression and negatively regulates a variety of processes including filamentous growth and alkaline pH response |
|
| YMR232W | 5.83 |
FUS2
|
Cytoplasmic protein localized to the shmoo tip; required for the alignment of parental nuclei before nuclear fusion during mating |
|
| YDR317W | 5.81 |
HIM1
|
Protein of unknown function involved in DNA repair |
|
| YGR190C | 5.80 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; overlaps the verified gene HIP1/YGR191W |
||
| YPL187W | 5.78 |
MF(ALPHA)1
|
Mating pheromone alpha-factor, made by alpha cells; interacts with mating type a cells to induce cell cycle arrest and other responses leading to mating; also encoded by MF(ALPHA)2, although MF(ALPHA)1 produces most alpha-factor |
|
| YEL074W | 5.78 |
Hypothetical protein |
||
| YDL169C | 5.72 |
UGX2
|
Protein of unknown function, transcript accumulates in response to any combination of stress conditions |
|
| YGR273C | 5.70 |
Putative protein of unknown function; deletion mutant has no readily detectable phenotype; expression downregulated by treatment with 8-methoxypsoralen plus UVA irradiation |
||
| YPR196W | 5.70 |
Putative maltose activator |
||
| YHR054C | 5.66 |
Putative protein of unknown function |
||
| YOR227W | 5.64 |
Protein of unknown function that may interact with ribosomes, based on co-purification experiments; authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies |
||
| YOR190W | 5.58 |
SPR1
|
Sporulation-specific exo-1,3-beta-glucanase; contributes to ascospore thermoresistance |
|
| YNL034W | 5.54 |
Putative protein of unknown function; YNL034W is not an essential gene |
||
| YNR034W | 5.52 |
SOL1
|
Protein with a possible role in tRNA export; shows similarity to 6-phosphogluconolactonase non-catalytic domains but does not exhibit this enzymatic activity; homologous to Sol2p, Sol3p, and Sol4p |
|
| YNR047W | 5.52 |
Putative protein kinase that, when overexpressed, interferes with pheromone-induced growth arrest; localizes to the cytoplasm; potential Cdc28p substrate |
||
| YBR180W | 5.50 |
DTR1
|
Putative dityrosine transporter, required for spore wall synthesis; expressed during sporulation; member of the major facilitator superfamily (DHA1 family) of multidrug resistance transporters |
|
| YCR011C | 5.48 |
ADP1
|
Putative ATP-dependent permease of the ABC transporter family of proteins |
|
| YBR179C | 5.41 |
FZO1
|
Mitochondrial integral membrane protein involved in mitochondrial fusion and maintenance of the mitochondrial genome; contains N-terminal GTPase domain |
|
| YER103W | 5.38 |
SSA4
|
Heat shock protein that is highly induced upon stress; plays a role in SRP-dependent cotranslational protein-membrane targeting and translocation; member of the HSP70 family; cytoplasmic protein that concentrates in nuclei upon starvation |
|
| YFR047C | 5.34 |
BNA6
|
Quinolinate phosphoribosyl transferase, required for the de novo biosynthesis of NAD from tryptophan via kynurenine; expression regulated by Hst1p |
|
| YOR065W | 5.32 |
CYT1
|
Cytochrome c1, component of the mitochondrial respiratory chain; expression is regulated by the heme-activated, glucose-repressed Hap2p/3p/4p/5p CCAAT-binding complex |
|
| YIL140W | 5.30 |
AXL2
|
Integral plasma membrane protein required for axial budding in haploid cells, localizes to the incipient bud site and bud neck; glycosylated by Pmt4p; potential Cdc28p substrate |
|
| YBR045C | 5.27 |
GIP1
|
Meiosis-specific regulatory subunit of the Glc7p protein phosphatase, regulates spore wall formation and septin organization, required for expression of some late meiotic genes and for normal localization of Glc7p |
|
| YPR015C | 5.24 |
Putative protein of unknown function |
||
| YJL130C | 5.20 |
URA2
|
Bifunctional carbamoylphosphate synthetase (CPSase)-aspartate transcarbamylase (ATCase), catalyzes the first two enzymatic steps in the de novo biosynthesis of pyrimidines; both activities are subject to feedback inhibition by UTP |
|
| YNL328C | 5.20 |
MDJ2
|
Constituent of the mitochondrial import motor associated with the presequence translocase; function overlaps with that of Pam18p; stimulates the ATPase activity of Ssc1p to drive mitochondrial import; contains a J domain |
|
| YNR034W-A | 5.18 |
Putative protein of unknown function; expression is regulated by Msn2p/Msn4p |
||
| YHR145C | 5.13 |
Dubious open reading frame unlikely to encode a functional protein, based on available experimental and comparative sequence data |
||
| YAL039C | 5.12 |
CYC3
|
Cytochrome c heme lyase (holocytochrome c synthase), attaches heme to apo-cytochrome c (Cyc1p or Cyc7p) in the mitochondrial intermembrane space; human ortholog may have a role in microphthalmia with linear skin defects (MLS) |
|
| YMR194C-B | 4.95 |
Putative protein of unknown function |
||
| YBR039W | 4.95 |
ATP3
|
Gamma subunit of the F1 sector of mitochondrial F1F0 ATP synthase, which is a large, evolutionarily conserved enzyme complex required for ATP synthesis |
|
| YMR271C | 4.94 |
URA10
|
Minor orotate phosphoribosyltransferase (OPRTase) isozyme that catalyzes the fifth enzymatic step in the de novo biosynthesis of pyrimidines, converting orotate into orotidine-5'-phosphate; major OPRTase encoded by URA5 |
|
| YHR206W | 4.92 |
SKN7
|
Nuclear response regulator and transcription factor, part of a branched two-component signaling system; required for optimal induction of heat-shock genes in response to oxidative stress; involved in osmoregulation |
|
| YEL075C | 4.87 |
Putative protein of unknown function |
||
| YFL012W-A | 4.85 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; overlaps the verified gene IES1/YFL013C |
||
| YGR260W | 4.81 |
TNA1
|
High affinity nicotinic acid plasma membrane permease, responsible for uptake of low levels of nicotinic acid; expression of the gene increases in the absence of extracellular nicotinic acid or para-aminobenzoate (PABA) |
|
| YFL013C | 4.81 |
IES1
|
Subunit of the INO80 chromatin remodeling complex |
|
| YER166W | 4.81 |
DNF1
|
Aminophospholipid translocase (flippase) that localizes primarily to the plasma membrane; contributes to endocytosis, protein transport and cell polarity; type 4 P-type ATPase |
|
| YDR273W | 4.78 |
DON1
|
Meiosis-specific component of the spindle pole body, part of the leading edge protein (LEP) coat, forms a ring-like structure at the leading edge of the prospore membrane during meiosis II |
|
| YGL117W | 4.71 |
Putative protein of unknown function |
||
| YCL038C | 4.69 |
ATG22
|
Vacuolar integral membrane protein required for efflux of amino acids during autophagic body breakdown in the vacuole; null mutation causes a gradual loss of viability during starvation |
|
| YOL085W-A | 4.66 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; partially overlaps the dubious ORF YOL085C |
||
| YER142C | 4.65 |
MAG1
|
3-methyl-adenine DNA glycosylase involved in protecting DNA against alkylating agents; initiates base excision repair by removing damaged bases to create abasic sites that are subsequently repaired |
|
| YHL016C | 4.64 |
DUR3
|
Plasma membrane transporter for both urea and polyamines, expression is highly sensitive to nitrogen catabolite repression and induced by allophanate, the last intermediate of the allantoin degradative pathway |
|
| YOR213C | 4.58 |
SAS5
|
Subunit of the SAS complex (Sas2p, Sas4p, Sas5p), which acetylates free histones and nucleosomes and regulates transcriptional silencing; stimulates Sas2p HAT activity |
|
| YOL024W | 4.53 |
Putative protein of unknown function, predicted to have thiol-disulfide oxidoreductase active site |
||
| YPL261C | 4.50 |
Dubious open reading frame unlikely to encode a protein; partially overlaps the uncharacterized ORF YPL260W |
||
| YPL021W | 4.48 |
ECM23
|
Non-essential protein of unconfirmed function; affects pre-rRNA processing, may act as a negative regulator of the transcription of genes involved in pseudohyphal growth; homologous to Srd1p |
|
| YPR005C | 4.42 |
HAL1
|
Cytoplasmic protein involved in halotolerance; decreases intracellular Na+ (via Ena1p) and increases intracellular K+ by decreasing efflux; expression repressed by Ssn6p-Tup1p and Sko1p and induced by NaCl, KCl, and sorbitol through Gcn4p |
|
| YJR107W | 4.42 |
Putative protein of unknown function; has sequence or structural similarity to lipases |
||
| YBR255C-A | 4.41 |
Putative protein of unknown function; identified by sequence comparison with hemiascomycetous yeast species |
||
| YEL010W | 4.40 |
Dubious open reading frame unlikely to encode a functional protein, based on available experimental and comparative sequence data |
||
| YLR040C | 4.40 |
Putative protein of unknown function; localizes to the cell wall; predicted to be a GPI-attached protein; upregulated by Mcm1p-Alpha1p transcription factor; partially overlaps the dubious ORF YLR041W; YLR040C is not essential |
||
| YGL072C | 4.39 |
Dubious open reading frame unlikely to encode a protein; partially overlaps the verified gene HSF1; null mutant displays increased resistance to antifungal agents gliotoxin, cycloheximide and H2O2 |
||
| YCR021C | 4.33 |
HSP30
|
Hydrophobic plasma membrane localized, stress-responsive protein that negatively regulates the H(+)-ATPase Pma1p; induced by heat shock, ethanol treatment, weak organic acid, glucose limitation, and entry into stationary phase |
|
| YNL196C | 4.22 |
SLZ1
|
Sporulation-specific protein with a leucine zipper motif |
|
| YLL053C | 4.20 |
Putative protein; in the Sigma 1278B strain background YLL053C is contiguous with AQY2 which encodes an aquaporin |
||
| YBR021W | 4.19 |
FUR4
|
Uracil permease, localized to the plasma membrane; expression is tightly regulated by uracil levels and environmental cues |
Network of associatons between targets according to the STRING database.
First level regulatory network of SUM1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 15.0 | 45.1 | GO:0019676 | ammonia assimilation cycle(GO:0019676) |
| 11.4 | 34.1 | GO:0006577 | amino-acid betaine metabolic process(GO:0006577) carnitine metabolic process(GO:0009437) |
| 8.1 | 48.5 | GO:0000501 | flocculation(GO:0000128) flocculation via cell wall protein-carbohydrate interaction(GO:0000501) |
| 7.7 | 30.9 | GO:0015847 | putrescine transport(GO:0015847) |
| 6.8 | 61.4 | GO:0006097 | glyoxylate cycle(GO:0006097) |
| 6.7 | 26.9 | GO:0046688 | response to copper ion(GO:0046688) |
| 5.2 | 15.6 | GO:0015888 | thiamine transport(GO:0015888) |
| 4.6 | 23.0 | GO:0009051 | pentose-phosphate shunt, oxidative branch(GO:0009051) |
| 4.4 | 22.0 | GO:0034627 | 'de novo' NAD biosynthetic process from tryptophan(GO:0034354) 'de novo' NAD biosynthetic process(GO:0034627) |
| 4.2 | 16.7 | GO:0000949 | aromatic amino acid family catabolic process to alcohol via Ehrlich pathway(GO:0000949) |
| 4.0 | 31.7 | GO:0045740 | positive regulation of DNA replication(GO:0045740) |
| 3.6 | 17.8 | GO:0008053 | mitochondrial fusion(GO:0008053) |
| 3.3 | 10.0 | GO:0042744 | hydrogen peroxide catabolic process(GO:0042744) |
| 3.2 | 28.4 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
| 3.0 | 30.4 | GO:0006098 | pentose-phosphate shunt(GO:0006098) glyceraldehyde-3-phosphate metabolic process(GO:0019682) |
| 2.9 | 11.6 | GO:0015886 | heme transport(GO:0015886) |
| 2.6 | 13.0 | GO:0033683 | nucleotide-excision repair, DNA incision(GO:0033683) |
| 2.4 | 101.8 | GO:0071940 | ascospore wall assembly(GO:0030476) spore wall assembly(GO:0042244) spore wall biogenesis(GO:0070590) ascospore wall biogenesis(GO:0070591) fungal-type cell wall assembly(GO:0071940) |
| 2.4 | 21.8 | GO:0000755 | cytogamy(GO:0000755) |
| 2.4 | 7.2 | GO:0006545 | glycine biosynthetic process(GO:0006545) |
| 2.4 | 12.0 | GO:0015793 | glycerol transport(GO:0015793) |
| 2.4 | 7.2 | GO:0045117 | azole transport(GO:0045117) |
| 1.9 | 5.8 | GO:1900460 | biofilm formation(GO:0042710) negative regulation of invasive growth in response to glucose limitation by negative regulation of transcription from RNA polymerase II promoter(GO:1900460) |
| 1.9 | 5.8 | GO:0032780 | negative regulation of ATPase activity(GO:0032780) |
| 1.9 | 1.9 | GO:0043902 | positive regulation of multi-organism process(GO:0043902) |
| 1.8 | 14.2 | GO:0006122 | mitochondrial electron transport, ubiquinol to cytochrome c(GO:0006122) |
| 1.7 | 13.8 | GO:0044091 | ascospore-type prospore membrane assembly(GO:0032120) membrane biogenesis(GO:0044091) membrane assembly(GO:0071709) |
| 1.7 | 5.1 | GO:0000304 | response to singlet oxygen(GO:0000304) |
| 1.7 | 5.1 | GO:0018063 | protein-heme linkage(GO:0017003) protein-tetrapyrrole linkage(GO:0017006) cytochrome c-heme linkage(GO:0018063) |
| 1.7 | 18.7 | GO:0006817 | phosphate ion transport(GO:0006817) |
| 1.7 | 8.5 | GO:0006673 | inositolphosphoceramide metabolic process(GO:0006673) |
| 1.7 | 5.0 | GO:1904951 | positive regulation of protein transport(GO:0051222) positive regulation of establishment of protein localization(GO:1904951) |
| 1.7 | 40.0 | GO:0015749 | hexose transport(GO:0008645) monosaccharide transport(GO:0015749) |
| 1.6 | 15.9 | GO:0000753 | cell morphogenesis involved in conjugation with cellular fusion(GO:0000753) |
| 1.6 | 1.6 | GO:0046487 | glyoxylate metabolic process(GO:0046487) |
| 1.6 | 4.7 | GO:0006285 | base-excision repair, AP site formation(GO:0006285) |
| 1.5 | 26.2 | GO:0009062 | fatty acid catabolic process(GO:0009062) |
| 1.5 | 13.7 | GO:0031167 | rRNA methylation(GO:0031167) |
| 1.5 | 21.3 | GO:0015985 | energy coupled proton transport, down electrochemical gradient(GO:0015985) ATP synthesis coupled proton transport(GO:0015986) |
| 1.5 | 4.4 | GO:0042148 | strand invasion(GO:0042148) |
| 1.5 | 8.8 | GO:0000256 | allantoin metabolic process(GO:0000255) allantoin catabolic process(GO:0000256) |
| 1.4 | 10.1 | GO:0008614 | pyridoxine metabolic process(GO:0008614) pyridoxine biosynthetic process(GO:0008615) vitamin B6 metabolic process(GO:0042816) vitamin B6 biosynthetic process(GO:0042819) |
| 1.4 | 5.7 | GO:0045719 | negative regulation of glycogen biosynthetic process(GO:0045719) negative regulation of glycogen metabolic process(GO:0070874) |
| 1.4 | 1.4 | GO:0000706 | meiotic DNA double-strand break processing(GO:0000706) |
| 1.3 | 6.7 | GO:0032108 | negative regulation of macroautophagy(GO:0016242) negative regulation of response to external stimulus(GO:0032102) negative regulation of response to extracellular stimulus(GO:0032105) negative regulation of response to nutrient levels(GO:0032108) |
| 1.3 | 6.4 | GO:2000284 | positive regulation of cellular amino acid biosynthetic process(GO:2000284) |
| 1.3 | 2.5 | GO:0035947 | regulation of gluconeogenesis by regulation of transcription from RNA polymerase II promoter(GO:0035947) positive regulation of gluconeogenesis by positive regulation of transcription from RNA polymerase II promoter(GO:0035948) regulation of transcription from RNA polymerase II promoter by a nonfermentable carbon source(GO:0061413) positive regulation of transcription from RNA polymerase II promoter by a nonfermentable carbon source(GO:0061414) |
| 1.2 | 9.9 | GO:0015891 | siderophore transport(GO:0015891) |
| 1.2 | 7.3 | GO:0006207 | 'de novo' pyrimidine nucleobase biosynthetic process(GO:0006207) 'de novo' UMP biosynthetic process(GO:0044205) |
| 1.2 | 10.5 | GO:0006829 | zinc II ion transport(GO:0006829) |
| 1.2 | 3.5 | GO:0045950 | negative regulation of mitotic recombination(GO:0045950) |
| 1.1 | 8.9 | GO:0051177 | meiotic sister chromatid cohesion(GO:0051177) |
| 1.1 | 4.3 | GO:0015855 | pyrimidine nucleobase transport(GO:0015855) |
| 1.0 | 4.1 | GO:0006848 | pyruvate transport(GO:0006848) |
| 1.0 | 8.7 | GO:0045332 | phospholipid translocation(GO:0045332) |
| 0.9 | 5.7 | GO:0071805 | cellular potassium ion transport(GO:0071804) potassium ion transmembrane transport(GO:0071805) |
| 0.9 | 4.7 | GO:0044746 | amino acid transmembrane export from vacuole(GO:0032974) vacuolar transmembrane transport(GO:0034486) vacuolar amino acid transmembrane transport(GO:0034487) amino acid transmembrane export(GO:0044746) |
| 0.9 | 1.8 | GO:0051352 | negative regulation of ligase activity(GO:0051352) negative regulation of ubiquitin-protein transferase activity(GO:0051444) |
| 0.9 | 10.0 | GO:0042724 | thiamine biosynthetic process(GO:0009228) thiamine-containing compound biosynthetic process(GO:0042724) |
| 0.9 | 8.0 | GO:0070987 | error-free translesion synthesis(GO:0070987) |
| 0.8 | 1.7 | GO:0010942 | positive regulation of cell death(GO:0010942) positive regulation of apoptotic process(GO:0043065) positive regulation of programmed cell death(GO:0043068) |
| 0.8 | 2.5 | GO:0072488 | ammonium transmembrane transport(GO:0072488) |
| 0.8 | 5.0 | GO:0000338 | protein deneddylation(GO:0000338) cullin deneddylation(GO:0010388) |
| 0.8 | 2.5 | GO:0043270 | positive regulation of ion transport(GO:0043270) |
| 0.8 | 1.6 | GO:0061395 | regulation of transcription from RNA polymerase II promoter in response to arsenic-containing substance(GO:0061394) positive regulation of transcription from RNA polymerase II promoter in response to arsenic-containing substance(GO:0061395) cellular response to arsenic-containing substance(GO:0071243) |
| 0.8 | 1.6 | GO:1900544 | positive regulation of nucleotide metabolic process(GO:0045981) positive regulation of purine nucleotide metabolic process(GO:1900544) |
| 0.8 | 5.5 | GO:0000409 | regulation of transcription by galactose(GO:0000409) positive regulation of transcription by galactose(GO:0000411) |
| 0.8 | 15.0 | GO:0001101 | response to acid chemical(GO:0001101) |
| 0.7 | 6.7 | GO:0006616 | SRP-dependent cotranslational protein targeting to membrane, translocation(GO:0006616) |
| 0.7 | 3.0 | GO:0006103 | 2-oxoglutarate metabolic process(GO:0006103) |
| 0.7 | 2.2 | GO:0006183 | GTP biosynthetic process(GO:0006183) GTP metabolic process(GO:0046039) |
| 0.7 | 2.1 | GO:0006089 | lactate metabolic process(GO:0006089) |
| 0.7 | 6.3 | GO:0000290 | deadenylation-dependent decapping of nuclear-transcribed mRNA(GO:0000290) |
| 0.7 | 32.3 | GO:0034293 | sexual sporulation(GO:0034293) sexual sporulation resulting in formation of a cellular spore(GO:0043935) |
| 0.7 | 2.7 | GO:1903137 | inactivation of MAPK activity involved in cell wall organization or biogenesis(GO:0000200) regulation of MAPK cascade involved in cell wall organization or biogenesis(GO:1903137) negative regulation of MAPK cascade involved in cell wall organization or biogenesis(GO:1903138) negative regulation of cell wall organization or biogenesis(GO:1903339) |
| 0.6 | 6.5 | GO:0051181 | cofactor transport(GO:0051181) |
| 0.6 | 1.9 | GO:0090110 | cargo loading into vesicle(GO:0035459) cargo loading into COPII-coated vesicle(GO:0090110) |
| 0.6 | 7.7 | GO:0007120 | axial cellular bud site selection(GO:0007120) |
| 0.6 | 1.9 | GO:0071050 | snoRNA polyadenylation(GO:0071050) |
| 0.6 | 1.8 | GO:2001021 | regulation of cell cycle checkpoint(GO:1901976) negative regulation of cell cycle checkpoint(GO:1901977) regulation of DNA damage checkpoint(GO:2000001) negative regulation of DNA damage checkpoint(GO:2000002) negative regulation of response to DNA damage stimulus(GO:2001021) |
| 0.6 | 3.0 | GO:0001308 | negative regulation of chromatin silencing involved in replicative cell aging(GO:0001308) |
| 0.6 | 8.5 | GO:0006081 | cellular aldehyde metabolic process(GO:0006081) |
| 0.6 | 1.8 | GO:0051469 | vesicle fusion with vacuole(GO:0051469) |
| 0.6 | 5.9 | GO:0006826 | iron ion transport(GO:0006826) |
| 0.6 | 2.4 | GO:0000196 | MAPK cascade involved in cell wall organization or biogenesis(GO:0000196) |
| 0.6 | 1.2 | GO:0019627 | urea metabolic process(GO:0019627) |
| 0.6 | 1.7 | GO:0042843 | D-xylose catabolic process(GO:0042843) |
| 0.6 | 14.9 | GO:0006312 | mitotic recombination(GO:0006312) |
| 0.5 | 4.9 | GO:0015802 | basic amino acid transport(GO:0015802) |
| 0.5 | 3.2 | GO:0046337 | phosphatidylethanolamine metabolic process(GO:0046337) |
| 0.5 | 1.0 | GO:0033499 | galactose catabolic process via UDP-galactose(GO:0033499) |
| 0.5 | 10.0 | GO:0036003 | positive regulation of transcription from RNA polymerase II promoter in response to stress(GO:0036003) |
| 0.5 | 7.5 | GO:0006513 | protein monoubiquitination(GO:0006513) |
| 0.5 | 1.4 | GO:0006003 | fructose 2,6-bisphosphate metabolic process(GO:0006003) |
| 0.5 | 0.5 | GO:0031571 | mitotic G1 DNA damage checkpoint(GO:0031571) mitotic G1/S transition checkpoint(GO:0044819) |
| 0.5 | 6.4 | GO:0007129 | synapsis(GO:0007129) |
| 0.4 | 3.1 | GO:0030026 | cellular manganese ion homeostasis(GO:0030026) manganese ion homeostasis(GO:0055071) |
| 0.4 | 9.1 | GO:0061726 | mitophagy(GO:0000422) mitochondrion disassembly(GO:0061726) |
| 0.4 | 1.2 | GO:0032186 | cellular bud neck septin ring organization(GO:0032186) protein autoubiquitination(GO:0051865) protein localization to bud neck(GO:0097271) |
| 0.4 | 0.4 | GO:0099515 | actin filament-based movement(GO:0030048) actin filament-based transport(GO:0099515) |
| 0.4 | 17.9 | GO:0030435 | sporulation resulting in formation of a cellular spore(GO:0030435) anatomical structure formation involved in morphogenesis(GO:0048646) |
| 0.4 | 6.0 | GO:0070816 | phosphorylation of RNA polymerase II C-terminal domain(GO:0070816) |
| 0.4 | 2.4 | GO:0000114 | obsolete regulation of transcription involved in G1 phase of mitotic cell cycle(GO:0000114) |
| 0.4 | 1.2 | GO:0051180 | vitamin transport(GO:0051180) |
| 0.4 | 2.6 | GO:0043409 | negative regulation of MAPK cascade(GO:0043409) |
| 0.4 | 0.7 | GO:0044070 | regulation of anion transport(GO:0044070) |
| 0.4 | 11.8 | GO:0006200 | obsolete ATP catabolic process(GO:0006200) |
| 0.4 | 1.5 | GO:0019346 | transsulfuration(GO:0019346) |
| 0.4 | 8.6 | GO:0006885 | regulation of pH(GO:0006885) |
| 0.4 | 1.8 | GO:0072329 | monocarboxylic acid catabolic process(GO:0072329) |
| 0.3 | 1.4 | GO:0019218 | regulation of steroid metabolic process(GO:0019218) regulation of ergosterol biosynthetic process(GO:0032443) regulation of steroid biosynthetic process(GO:0050810) |
| 0.3 | 7.1 | GO:0031503 | protein complex localization(GO:0031503) |
| 0.3 | 1.3 | GO:0035753 | maintenance of DNA trinucleotide repeats(GO:0035753) |
| 0.3 | 1.6 | GO:0045910 | negative regulation of DNA recombination(GO:0045910) |
| 0.3 | 0.6 | GO:0010513 | positive regulation of phosphatidylinositol biosynthetic process(GO:0010513) |
| 0.3 | 1.3 | GO:0070988 | protein demethylation(GO:0006482) protein dealkylation(GO:0008214) histone demethylation(GO:0016577) demethylation(GO:0070988) |
| 0.3 | 0.9 | GO:0055078 | cellular sodium ion homeostasis(GO:0006883) sodium ion homeostasis(GO:0055078) |
| 0.3 | 4.2 | GO:0000902 | cell morphogenesis(GO:0000902) |
| 0.3 | 0.9 | GO:0006598 | polyamine catabolic process(GO:0006598) |
| 0.3 | 1.1 | GO:0035337 | fatty-acyl-CoA metabolic process(GO:0035337) |
| 0.3 | 0.5 | GO:0097201 | negative regulation of transcription from RNA polymerase II promoter in response to stress(GO:0097201) |
| 0.3 | 3.9 | GO:0008535 | respiratory chain complex IV assembly(GO:0008535) |
| 0.3 | 0.5 | GO:0071456 | cellular response to hypoxia(GO:0071456) |
| 0.2 | 5.2 | GO:0030150 | protein import into mitochondrial matrix(GO:0030150) |
| 0.2 | 0.2 | GO:0044783 | G1 DNA damage checkpoint(GO:0044783) |
| 0.2 | 1.0 | GO:0046685 | response to arsenic-containing substance(GO:0046685) |
| 0.2 | 0.9 | GO:0007234 | osmosensory signaling via phosphorelay pathway(GO:0007234) |
| 0.2 | 2.8 | GO:0019740 | nitrogen utilization(GO:0019740) |
| 0.2 | 5.1 | GO:0000282 | cellular bud site selection(GO:0000282) cytokinesis, site selection(GO:0007105) mitotic cytokinesis, site selection(GO:1902408) |
| 0.2 | 0.7 | GO:1902751 | positive regulation of G2/M transition of mitotic cell cycle(GO:0010971) positive regulation of cell cycle G2/M phase transition(GO:1902751) |
| 0.2 | 2.3 | GO:0033108 | mitochondrial respiratory chain complex assembly(GO:0033108) |
| 0.2 | 0.7 | GO:0090282 | regulation of transcription involved in G2/M transition of mitotic cell cycle(GO:0000117) positive regulation of transcription involved in G2/M transition of mitotic cell cycle(GO:0090282) |
| 0.2 | 0.9 | GO:0060237 | regulation of fungal-type cell wall organization(GO:0060237) |
| 0.2 | 0.4 | GO:0070911 | global genome nucleotide-excision repair(GO:0070911) |
| 0.2 | 0.9 | GO:0071168 | protein localization to chromatin(GO:0071168) |
| 0.2 | 1.5 | GO:0097034 | mitochondrial respiratory chain complex IV biogenesis(GO:0097034) |
| 0.2 | 0.2 | GO:0042542 | response to hydrogen peroxide(GO:0042542) |
| 0.2 | 0.4 | GO:0006740 | NADPH regeneration(GO:0006740) |
| 0.2 | 3.1 | GO:0007266 | Rho protein signal transduction(GO:0007266) |
| 0.2 | 4.0 | GO:0008361 | regulation of cell size(GO:0008361) |
| 0.2 | 1.0 | GO:0070676 | intralumenal vesicle formation(GO:0070676) |
| 0.2 | 1.2 | GO:0005987 | sucrose metabolic process(GO:0005985) sucrose catabolic process(GO:0005987) |
| 0.2 | 0.8 | GO:0032581 | ER-dependent peroxisome organization(GO:0032581) peroxisome inheritance(GO:0045033) |
| 0.2 | 1.0 | GO:0000023 | maltose metabolic process(GO:0000023) |
| 0.2 | 0.4 | GO:0006279 | premeiotic DNA replication(GO:0006279) |
| 0.2 | 1.0 | GO:0006627 | protein processing involved in protein targeting to mitochondrion(GO:0006627) |
| 0.2 | 0.2 | GO:0005980 | glycogen catabolic process(GO:0005980) glucan catabolic process(GO:0009251) cellular polysaccharide catabolic process(GO:0044247) |
| 0.2 | 0.4 | GO:0031340 | positive regulation of vesicle fusion(GO:0031340) |
| 0.2 | 1.0 | GO:0009410 | response to xenobiotic stimulus(GO:0009410) |
| 0.2 | 7.1 | GO:0009060 | aerobic respiration(GO:0009060) |
| 0.2 | 2.4 | GO:0030474 | spindle pole body duplication(GO:0030474) |
| 0.2 | 0.6 | GO:0045721 | negative regulation of gluconeogenesis(GO:0045721) |
| 0.2 | 2.3 | GO:0005978 | glycogen biosynthetic process(GO:0005978) |
| 0.1 | 2.4 | GO:0043162 | ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway(GO:0043162) |
| 0.1 | 0.9 | GO:0048478 | replication fork protection(GO:0048478) |
| 0.1 | 0.7 | GO:0018904 | glycerol ether metabolic process(GO:0006662) ether metabolic process(GO:0018904) |
| 0.1 | 1.5 | GO:0031145 | anaphase-promoting complex-dependent catabolic process(GO:0031145) |
| 0.1 | 0.5 | GO:0071495 | cellular response to endogenous stimulus(GO:0071495) |
| 0.1 | 0.6 | GO:0006768 | biotin metabolic process(GO:0006768) biotin biosynthetic process(GO:0009102) |
| 0.1 | 1.5 | GO:0000147 | actin cortical patch assembly(GO:0000147) actin cortical patch organization(GO:0044396) |
| 0.1 | 0.9 | GO:0000266 | mitochondrial fission(GO:0000266) |
| 0.1 | 1.4 | GO:0034968 | histone lysine methylation(GO:0034968) |
| 0.1 | 1.0 | GO:0030847 | termination of RNA polymerase II transcription, exosome-dependent(GO:0030847) |
| 0.1 | 0.5 | GO:0001113 | transcriptional open complex formation at RNA polymerase II promoter(GO:0001113) |
| 0.1 | 0.6 | GO:0032048 | cardiolipin metabolic process(GO:0032048) |
| 0.1 | 0.6 | GO:0000390 | spliceosomal complex disassembly(GO:0000390) |
| 0.1 | 1.7 | GO:0034401 | chromatin organization involved in regulation of transcription(GO:0034401) |
| 0.1 | 0.2 | GO:0009268 | response to pH(GO:0009268) |
| 0.1 | 1.0 | GO:0060261 | positive regulation of RNA polymerase II transcriptional preinitiation complex assembly(GO:0045899) positive regulation of transcription initiation from RNA polymerase II promoter(GO:0060261) positive regulation of DNA-templated transcription, initiation(GO:2000144) |
| 0.1 | 0.3 | GO:0060211 | regulation of nuclear-transcribed mRNA poly(A) tail shortening(GO:0060211) |
| 0.1 | 0.3 | GO:0000042 | protein targeting to Golgi(GO:0000042) establishment of protein localization to Golgi(GO:0072600) |
| 0.1 | 1.8 | GO:0009267 | cellular response to starvation(GO:0009267) |
| 0.1 | 0.3 | GO:0019401 | glycerol biosynthetic process(GO:0006114) alditol biosynthetic process(GO:0019401) |
| 0.1 | 0.4 | GO:0002183 | cytoplasmic translational initiation(GO:0002183) |
| 0.1 | 0.2 | GO:0019748 | secondary metabolic process(GO:0019748) |
| 0.1 | 1.0 | GO:0000742 | karyogamy involved in conjugation with cellular fusion(GO:0000742) |
| 0.1 | 0.8 | GO:0006369 | termination of RNA polymerase II transcription(GO:0006369) |
| 0.1 | 0.1 | GO:0001120 | DNA-templated transcriptional open complex formation(GO:0001112) protein-DNA complex remodeling(GO:0001120) macromolecular complex remodeling(GO:0034367) |
| 0.1 | 0.7 | GO:0006490 | oligosaccharide-lipid intermediate biosynthetic process(GO:0006490) |
| 0.1 | 0.3 | GO:0031086 | nuclear-transcribed mRNA catabolic process, deadenylation-independent decay(GO:0031086) deadenylation-independent decapping of nuclear-transcribed mRNA(GO:0031087) |
| 0.1 | 0.1 | GO:0042732 | D-xylose metabolic process(GO:0042732) |
| 0.1 | 0.4 | GO:0000436 | carbon catabolite activation of transcription from RNA polymerase II promoter(GO:0000436) |
| 0.1 | 0.2 | GO:0034503 | protein localization to nucleolar rDNA repeats(GO:0034503) |
| 0.1 | 0.5 | GO:0042766 | nucleosome mobilization(GO:0042766) |
| 0.1 | 0.5 | GO:0006526 | arginine biosynthetic process(GO:0006526) |
| 0.1 | 0.5 | GO:0046854 | lipid phosphorylation(GO:0046834) phosphatidylinositol phosphorylation(GO:0046854) |
| 0.1 | 0.5 | GO:0030952 | establishment or maintenance of actin cytoskeleton polarity(GO:0030950) establishment or maintenance of cytoskeleton polarity(GO:0030952) |
| 0.0 | 0.3 | GO:0009113 | purine nucleobase biosynthetic process(GO:0009113) |
| 0.0 | 0.4 | GO:0046173 | polyol biosynthetic process(GO:0046173) |
| 0.0 | 0.1 | GO:0031055 | chromatin remodeling at centromere(GO:0031055) |
| 0.0 | 0.2 | GO:0010821 | regulation of mitochondrion organization(GO:0010821) |
| 0.0 | 0.4 | GO:0006749 | glutathione metabolic process(GO:0006749) |
| 0.0 | 0.1 | GO:0036257 | multivesicular body organization(GO:0036257) |
| 0.0 | 0.1 | GO:0030149 | sphingolipid catabolic process(GO:0030149) membrane lipid catabolic process(GO:0046466) |
| 0.0 | 0.3 | GO:0019878 | lysine biosynthetic process via aminoadipic acid(GO:0019878) |
| 0.0 | 0.5 | GO:0006476 | protein deacetylation(GO:0006476) protein deacylation(GO:0035601) macromolecule deacylation(GO:0098732) |
| 0.0 | 0.3 | GO:0048309 | endoplasmic reticulum inheritance(GO:0048309) |
| 0.0 | 0.9 | GO:0000082 | G1/S transition of mitotic cell cycle(GO:0000082) cell cycle G1/S phase transition(GO:0044843) |
| 0.0 | 0.2 | GO:0045039 | protein import into mitochondrial inner membrane(GO:0045039) |
| 0.0 | 0.4 | GO:0043086 | negative regulation of catalytic activity(GO:0043086) |
| 0.0 | 2.2 | GO:0032543 | mitochondrial translation(GO:0032543) |
| 0.0 | 0.3 | GO:0045727 | positive regulation of cellular amide metabolic process(GO:0034250) positive regulation of translation(GO:0045727) |
| 0.0 | 0.1 | GO:0007329 | positive regulation of transcription from RNA polymerase II promoter by pheromones(GO:0007329) positive regulation of transcription by pheromones(GO:0009371) |
| 0.0 | 0.2 | GO:0016579 | protein deubiquitination(GO:0016579) |
| 0.0 | 0.1 | GO:0006189 | 'de novo' IMP biosynthetic process(GO:0006189) |
| 0.0 | 0.0 | GO:0006529 | asparagine biosynthetic process(GO:0006529) |
| 0.0 | 0.3 | GO:0035825 | reciprocal meiotic recombination(GO:0007131) reciprocal DNA recombination(GO:0035825) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 7.1 | 21.3 | GO:0030061 | mitochondrial crista(GO:0030061) |
| 5.6 | 16.7 | GO:0005756 | mitochondrial proton-transporting ATP synthase, central stalk(GO:0005756) proton-transporting ATP synthase, central stalk(GO:0045269) |
| 4.3 | 13.0 | GO:0072687 | meiotic spindle(GO:0072687) |
| 3.7 | 44.0 | GO:0031160 | spore wall(GO:0031160) |
| 3.3 | 29.6 | GO:0045277 | mitochondrial respiratory chain complex IV(GO:0005751) respiratory chain complex IV(GO:0045277) |
| 2.7 | 19.0 | GO:0000112 | nucleotide-excision repair factor 3 complex(GO:0000112) |
| 2.3 | 9.1 | GO:0033551 | monopolin complex(GO:0033551) |
| 2.3 | 27.4 | GO:0031907 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 2.2 | 6.5 | GO:1990316 | ATG1/ULK1 kinase complex(GO:1990316) |
| 2.0 | 48.1 | GO:0042763 | prospore membrane(GO:0005628) intracellular immature spore(GO:0042763) ascospore-type prospore(GO:0042764) |
| 1.9 | 5.8 | GO:0051286 | cell tip(GO:0051286) |
| 1.6 | 4.7 | GO:0031166 | integral component of vacuolar membrane(GO:0031166) |
| 1.5 | 4.6 | GO:0033255 | SAS acetyltransferase complex(GO:0033255) |
| 1.4 | 2.8 | GO:0045261 | mitochondrial proton-transporting ATP synthase complex, catalytic core F(1)(GO:0000275) proton-transporting ATP synthase complex, catalytic core F(1)(GO:0045261) |
| 1.2 | 2.4 | GO:0032126 | eisosome(GO:0032126) |
| 1.2 | 56.8 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 1.1 | 27.3 | GO:0005887 | integral component of plasma membrane(GO:0005887) |
| 1.0 | 3.0 | GO:0045240 | mitochondrial alpha-ketoglutarate dehydrogenase complex(GO:0005947) mitochondrial oxoglutarate dehydrogenase complex(GO:0009353) dihydrolipoyl dehydrogenase complex(GO:0045240) oxoglutarate dehydrogenase complex(GO:0045252) |
| 0.9 | 3.6 | GO:0098552 | side of membrane(GO:0098552) cytoplasmic side of membrane(GO:0098562) |
| 0.9 | 2.6 | GO:0031229 | integral component of nuclear inner membrane(GO:0005639) intrinsic component of nuclear inner membrane(GO:0031229) |
| 0.9 | 16.2 | GO:0048471 | perinuclear region of cytoplasm(GO:0048471) |
| 0.8 | 5.0 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.8 | 8.6 | GO:0005769 | early endosome(GO:0005769) |
| 0.8 | 2.3 | GO:0042720 | mitochondrial inner membrane peptidase complex(GO:0042720) |
| 0.7 | 5.2 | GO:0001405 | presequence translocase-associated import motor(GO:0001405) |
| 0.7 | 2.0 | GO:0005784 | Sec61 translocon complex(GO:0005784) |
| 0.7 | 2.0 | GO:0032777 | Piccolo NuA4 histone acetyltransferase complex(GO:0032777) |
| 0.7 | 5.3 | GO:0070069 | cytochrome complex(GO:0070069) |
| 0.6 | 11.0 | GO:0031305 | intrinsic component of mitochondrial inner membrane(GO:0031304) integral component of mitochondrial inner membrane(GO:0031305) |
| 0.6 | 1.8 | GO:0000274 | mitochondrial proton-transporting ATP synthase, stator stalk(GO:0000274) proton-transporting ATP synthase, stator stalk(GO:0045265) |
| 0.6 | 32.3 | GO:0042579 | peroxisome(GO:0005777) microbody(GO:0042579) |
| 0.5 | 2.2 | GO:0034045 | pre-autophagosomal structure membrane(GO:0034045) |
| 0.5 | 1.6 | GO:0035649 | Nrd1 complex(GO:0035649) |
| 0.5 | 1.5 | GO:0097042 | extrinsic component of fungal-type vacuolar membrane(GO:0097042) |
| 0.5 | 22.8 | GO:0005758 | mitochondrial intermembrane space(GO:0005758) |
| 0.5 | 8.7 | GO:0031307 | intrinsic component of mitochondrial outer membrane(GO:0031306) integral component of mitochondrial outer membrane(GO:0031307) |
| 0.5 | 3.3 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.5 | 1.8 | GO:0033309 | SBF transcription complex(GO:0033309) |
| 0.5 | 4.1 | GO:0000795 | synaptonemal complex(GO:0000795) |
| 0.4 | 1.3 | GO:0016363 | nuclear matrix(GO:0016363) |
| 0.4 | 6.5 | GO:0033698 | Rpd3L complex(GO:0033698) |
| 0.4 | 1.2 | GO:0032177 | split septin rings(GO:0032176) cellular bud neck split septin rings(GO:0032177) |
| 0.4 | 0.7 | GO:0030478 | actin cap(GO:0030478) |
| 0.3 | 2.4 | GO:0034657 | GID complex(GO:0034657) |
| 0.3 | 2.1 | GO:0044614 | nuclear pore cytoplasmic filaments(GO:0044614) |
| 0.3 | 29.1 | GO:0000329 | fungal-type vacuole membrane(GO:0000329) lytic vacuole membrane(GO:0098852) |
| 0.3 | 1.0 | GO:0000815 | ESCRT III complex(GO:0000815) |
| 0.3 | 6.6 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.2 | 2.7 | GO:0030014 | CCR4-NOT complex(GO:0030014) |
| 0.2 | 1.5 | GO:0034967 | Set3 complex(GO:0034967) |
| 0.2 | 85.5 | GO:0005886 | plasma membrane(GO:0005886) |
| 0.2 | 1.0 | GO:0071008 | U2-type post-mRNA release spliceosomal complex(GO:0071008) |
| 0.2 | 3.0 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.2 | 2.9 | GO:0008023 | transcription elongation factor complex(GO:0008023) |
| 0.1 | 0.8 | GO:0000799 | condensin complex(GO:0000796) nuclear condensin complex(GO:0000799) |
| 0.1 | 5.1 | GO:0000323 | storage vacuole(GO:0000322) lytic vacuole(GO:0000323) fungal-type vacuole(GO:0000324) |
| 0.1 | 0.6 | GO:0000974 | Prp19 complex(GO:0000974) |
| 0.1 | 0.6 | GO:0071007 | U2-type catalytic step 2 spliceosome(GO:0071007) catalytic step 2 spliceosome(GO:0071013) |
| 0.1 | 7.6 | GO:0005815 | microtubule organizing center(GO:0005815) spindle pole body(GO:0005816) |
| 0.1 | 3.6 | GO:0000314 | organellar small ribosomal subunit(GO:0000314) mitochondrial small ribosomal subunit(GO:0005763) |
| 0.1 | 1.3 | GO:0000788 | nuclear nucleosome(GO:0000788) |
| 0.1 | 2.1 | GO:0070847 | core mediator complex(GO:0070847) |
| 0.1 | 0.3 | GO:0030869 | RENT complex(GO:0030869) |
| 0.1 | 0.4 | GO:0016602 | CCAAT-binding factor complex(GO:0016602) |
| 0.1 | 0.4 | GO:0016587 | Isw1 complex(GO:0016587) |
| 0.1 | 0.5 | GO:0016272 | prefoldin complex(GO:0016272) |
| 0.1 | 3.4 | GO:0005643 | nuclear pore(GO:0005643) |
| 0.1 | 8.5 | GO:0005743 | mitochondrial inner membrane(GO:0005743) |
| 0.1 | 2.3 | GO:0000315 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.1 | 0.4 | GO:0035097 | histone methyltransferase complex(GO:0035097) Set1C/COMPASS complex(GO:0048188) |
| 0.0 | 0.1 | GO:0000126 | transcription factor TFIIIB complex(GO:0000126) |
| 0.0 | 3.4 | GO:0031966 | mitochondrial membrane(GO:0031966) |
| 0.0 | 0.4 | GO:0005847 | mRNA cleavage and polyadenylation specificity factor complex(GO:0005847) |
| 0.0 | 0.2 | GO:0031390 | Ctf18 RFC-like complex(GO:0031390) |
| 0.0 | 13.1 | GO:0005739 | mitochondrion(GO:0005739) |
| 0.0 | 0.3 | GO:0005686 | U2 snRNP(GO:0005686) |
| 0.0 | 0.2 | GO:0043189 | NuA4 histone acetyltransferase complex(GO:0035267) H4/H2A histone acetyltransferase complex(GO:0043189) H4 histone acetyltransferase complex(GO:1902562) |
| 0.0 | 1.1 | GO:0043332 | mating projection tip(GO:0043332) |
| 0.0 | 0.2 | GO:0012507 | ER to Golgi transport vesicle membrane(GO:0012507) |
| 0.0 | 0.3 | GO:0005802 | trans-Golgi network(GO:0005802) |
| 0.0 | 0.2 | GO:0000407 | pre-autophagosomal structure(GO:0000407) |
| 0.0 | 0.0 | GO:0034518 | RNA cap binding complex(GO:0034518) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 14.4 | 43.2 | GO:0004108 | citrate (Si)-synthase activity(GO:0004108) citrate synthase activity(GO:0036440) |
| 12.1 | 48.5 | GO:0005537 | mannose binding(GO:0005537) |
| 11.4 | 34.1 | GO:0004092 | carnitine O-acetyltransferase activity(GO:0004092) carnitine O-acyltransferase activity(GO:0016406) |
| 11.3 | 45.1 | GO:0016639 | oxidoreductase activity, acting on the CH-NH2 group of donors, NAD or NADP as acceptor(GO:0016639) |
| 10.5 | 31.5 | GO:0015489 | putrescine transmembrane transporter activity(GO:0015489) |
| 6.2 | 37.0 | GO:0005354 | galactose transmembrane transporter activity(GO:0005354) |
| 5.3 | 15.8 | GO:0004930 | G-protein coupled receptor activity(GO:0004930) |
| 4.9 | 14.7 | GO:0015114 | phosphate ion transmembrane transporter activity(GO:0015114) |
| 4.3 | 26.0 | GO:0016721 | superoxide dismutase activity(GO:0004784) oxidoreductase activity, acting on superoxide radicals as acceptor(GO:0016721) |
| 4.2 | 16.7 | GO:0004737 | pyruvate decarboxylase activity(GO:0004737) |
| 4.1 | 24.9 | GO:0016744 | transferase activity, transferring aldehyde or ketonic groups(GO:0016744) |
| 3.7 | 14.8 | GO:0015295 | solute:proton symporter activity(GO:0015295) |
| 3.3 | 13.3 | GO:0016435 | rRNA (guanine) methyltransferase activity(GO:0016435) |
| 3.0 | 29.6 | GO:0015002 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors(GO:0016675) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 3.0 | 11.8 | GO:0005471 | ATP transmembrane transporter activity(GO:0005347) ATP:ADP antiporter activity(GO:0005471) ADP transmembrane transporter activity(GO:0015217) |
| 2.6 | 15.8 | GO:0005337 | nucleoside transmembrane transporter activity(GO:0005337) |
| 2.4 | 7.2 | GO:1901474 | azole transmembrane transporter activity(GO:1901474) |
| 2.2 | 6.7 | GO:0004088 | carbamoyl-phosphate synthase (glutamine-hydrolyzing) activity(GO:0004088) |
| 2.2 | 15.5 | GO:0016833 | oxo-acid-lyase activity(GO:0016833) |
| 2.2 | 10.8 | GO:0018456 | aryl-alcohol dehydrogenase (NAD+) activity(GO:0018456) |
| 2.2 | 8.6 | GO:0005315 | inorganic phosphate transmembrane transporter activity(GO:0005315) |
| 2.0 | 10.1 | GO:0016638 | oxidoreductase activity, acting on the CH-NH2 group of donors(GO:0016638) |
| 2.0 | 13.8 | GO:0000014 | single-stranded DNA endodeoxyribonuclease activity(GO:0000014) |
| 1.9 | 14.9 | GO:0016702 | oxidoreductase activity, acting on single donors with incorporation of molecular oxygen(GO:0016701) oxidoreductase activity, acting on single donors with incorporation of molecular oxygen, incorporation of two atoms of oxygen(GO:0016702) |
| 1.8 | 19.5 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 1.7 | 10.4 | GO:0016408 | C-acyltransferase activity(GO:0016408) |
| 1.7 | 15.2 | GO:0016723 | ferric-chelate reductase activity(GO:0000293) oxidoreductase activity, oxidizing metal ions, NAD or NADP as acceptor(GO:0016723) |
| 1.7 | 6.7 | GO:0000772 | mating pheromone activity(GO:0000772) pheromone activity(GO:0005186) |
| 1.6 | 6.4 | GO:0015294 | solute:cation symporter activity(GO:0015294) |
| 1.5 | 3.0 | GO:0030976 | thiamine pyrophosphate binding(GO:0030976) |
| 1.5 | 5.9 | GO:0000156 | phosphorelay response regulator activity(GO:0000156) |
| 1.4 | 4.3 | GO:0050136 | NADH dehydrogenase activity(GO:0003954) NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 1.4 | 5.5 | GO:0017057 | 6-phosphogluconolactonase activity(GO:0017057) |
| 1.4 | 11.0 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 1.3 | 4.0 | GO:0031492 | nucleosomal DNA binding(GO:0031492) |
| 1.3 | 6.6 | GO:0016888 | endodeoxyribonuclease activity, producing 5'-phosphomonoesters(GO:0016888) |
| 1.3 | 5.3 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 1.3 | 11.4 | GO:0000149 | SNARE binding(GO:0000149) |
| 1.2 | 7.2 | GO:0005089 | Rho guanyl-nucleotide exchange factor activity(GO:0005089) |
| 1.1 | 3.4 | GO:0003843 | 1,3-beta-D-glucan synthase activity(GO:0003843) |
| 1.1 | 8.8 | GO:0016681 | ubiquinol-cytochrome-c reductase activity(GO:0008121) oxidoreductase activity, acting on diphenols and related substances as donors(GO:0016679) oxidoreductase activity, acting on diphenols and related substances as donors, cytochrome as acceptor(GO:0016681) |
| 1.1 | 3.2 | GO:0015385 | sodium:proton antiporter activity(GO:0015385) |
| 1.0 | 1.0 | GO:0015151 | alpha-glucoside transmembrane transporter activity(GO:0015151) glucoside transmembrane transporter activity(GO:0042947) |
| 1.0 | 6.0 | GO:0015293 | symporter activity(GO:0015293) |
| 1.0 | 4.0 | GO:0004396 | hexokinase activity(GO:0004396) |
| 1.0 | 8.7 | GO:0016628 | oxidoreductase activity, acting on the CH-CH group of donors, NAD or NADP as acceptor(GO:0016628) |
| 0.9 | 4.7 | GO:0019104 | DNA N-glycosylase activity(GO:0019104) |
| 0.9 | 4.6 | GO:0050308 | sugar-phosphatase activity(GO:0050308) |
| 0.9 | 2.7 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) MAP kinase phosphatase activity(GO:0033549) |
| 0.9 | 16.9 | GO:0050661 | NADP binding(GO:0050661) |
| 0.8 | 6.5 | GO:0051184 | cofactor transporter activity(GO:0051184) |
| 0.8 | 2.4 | GO:0005078 | MAP-kinase scaffold activity(GO:0005078) signaling adaptor activity(GO:0035591) |
| 0.8 | 5.6 | GO:0000774 | adenyl-nucleotide exchange factor activity(GO:0000774) |
| 0.8 | 4.0 | GO:0004707 | MAP kinase activity(GO:0004707) |
| 0.8 | 3.2 | GO:0004338 | glucan exo-1,3-beta-glucosidase activity(GO:0004338) |
| 0.8 | 3.9 | GO:0042124 | glucanosyltransferase activity(GO:0042123) 1,3-beta-glucanosyltransferase activity(GO:0042124) |
| 0.8 | 6.1 | GO:0004012 | phospholipid-translocating ATPase activity(GO:0004012) |
| 0.8 | 4.5 | GO:0008106 | aldo-keto reductase (NADP) activity(GO:0004033) alcohol dehydrogenase (NADP+) activity(GO:0008106) |
| 0.7 | 11.3 | GO:0020037 | heme binding(GO:0020037) tetrapyrrole binding(GO:0046906) |
| 0.7 | 2.1 | GO:0008902 | hydroxymethylpyrimidine kinase activity(GO:0008902) phosphomethylpyrimidine kinase activity(GO:0008972) |
| 0.7 | 12.5 | GO:0019213 | deacetylase activity(GO:0019213) |
| 0.6 | 1.9 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.6 | 5.8 | GO:0001671 | ATPase activator activity(GO:0001671) |
| 0.6 | 1.9 | GO:0008138 | protein tyrosine/serine/threonine phosphatase activity(GO:0008138) |
| 0.6 | 1.7 | GO:0032183 | SUMO binding(GO:0032183) |
| 0.6 | 1.2 | GO:0051183 | vitamin transporter activity(GO:0051183) |
| 0.5 | 8.8 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 0.5 | 1.5 | GO:0008559 | xenobiotic-transporting ATPase activity(GO:0008559) xenobiotic transporter activity(GO:0042910) |
| 0.5 | 7.2 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.5 | 4.2 | GO:0008599 | protein phosphatase type 1 regulator activity(GO:0008599) |
| 0.5 | 1.4 | GO:0033592 | RNA strand annealing activity(GO:0033592) annealing activity(GO:0097617) |
| 0.5 | 1.4 | GO:0001094 | TFIID-class transcription factor binding(GO:0001094) |
| 0.4 | 1.3 | GO:0016722 | oxidoreductase activity, oxidizing metal ions(GO:0016722) |
| 0.4 | 2.9 | GO:0015174 | basic amino acid transmembrane transporter activity(GO:0015174) |
| 0.4 | 1.2 | GO:0004575 | beta-fructofuranosidase activity(GO:0004564) sucrose alpha-glucosidase activity(GO:0004575) |
| 0.4 | 1.6 | GO:0016531 | copper chaperone activity(GO:0016531) |
| 0.4 | 6.1 | GO:0016763 | transferase activity, transferring pentosyl groups(GO:0016763) |
| 0.4 | 1.9 | GO:0004652 | polynucleotide adenylyltransferase activity(GO:0004652) |
| 0.4 | 2.2 | GO:0004030 | aldehyde dehydrogenase (NAD) activity(GO:0004029) aldehyde dehydrogenase [NAD(P)+] activity(GO:0004030) |
| 0.3 | 1.4 | GO:0042927 | siderophore transmembrane transporter activity(GO:0015343) iron chelate transmembrane transporter activity(GO:0015603) siderophore transporter activity(GO:0042927) |
| 0.3 | 2.4 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.3 | 2.7 | GO:0046912 | transferase activity, transferring acyl groups, acyl groups converted into alkyl on transfer(GO:0046912) |
| 0.3 | 2.0 | GO:0008379 | thioredoxin peroxidase activity(GO:0008379) |
| 0.3 | 2.0 | GO:0005244 | voltage-gated ion channel activity(GO:0005244) voltage-gated channel activity(GO:0022832) |
| 0.3 | 0.9 | GO:0072349 | modified amino acid transmembrane transporter activity(GO:0072349) |
| 0.3 | 1.5 | GO:0017048 | Rho GTPase binding(GO:0017048) |
| 0.3 | 3.9 | GO:0044389 | ubiquitin protein ligase binding(GO:0031625) ubiquitin-like protein ligase binding(GO:0044389) |
| 0.3 | 1.8 | GO:0016620 | oxidoreductase activity, acting on the aldehyde or oxo group of donors, NAD or NADP as acceptor(GO:0016620) |
| 0.3 | 2.0 | GO:0015450 | P-P-bond-hydrolysis-driven protein transmembrane transporter activity(GO:0015450) |
| 0.3 | 7.0 | GO:0016881 | acid-amino acid ligase activity(GO:0016881) |
| 0.3 | 1.1 | GO:0016885 | ligase activity, forming carbon-carbon bonds(GO:0016885) |
| 0.3 | 0.9 | GO:0000150 | recombinase activity(GO:0000150) |
| 0.3 | 2.5 | GO:0030246 | carbohydrate binding(GO:0030246) |
| 0.3 | 3.7 | GO:0001135 | transcription factor activity, RNA polymerase II transcription factor recruiting(GO:0001135) |
| 0.3 | 4.8 | GO:0101005 | thiol-dependent ubiquitin-specific protease activity(GO:0004843) thiol-dependent ubiquitinyl hydrolase activity(GO:0036459) ubiquitinyl hydrolase activity(GO:0101005) |
| 0.3 | 0.8 | GO:0004457 | lactate dehydrogenase activity(GO:0004457) oxidoreductase activity, acting on the CH-OH group of donors, cytochrome as acceptor(GO:0016898) |
| 0.2 | 4.2 | GO:0008081 | phosphoric diester hydrolase activity(GO:0008081) |
| 0.2 | 0.5 | GO:0005102 | receptor binding(GO:0005102) |
| 0.2 | 17.0 | GO:0051082 | unfolded protein binding(GO:0051082) |
| 0.2 | 0.9 | GO:0015193 | L-proline transmembrane transporter activity(GO:0015193) |
| 0.2 | 0.6 | GO:0016813 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amidines(GO:0016813) |
| 0.2 | 4.3 | GO:0005509 | calcium ion binding(GO:0005509) |
| 0.2 | 0.6 | GO:0050839 | cell adhesion molecule binding(GO:0050839) |
| 0.2 | 0.6 | GO:0016417 | S-acyltransferase activity(GO:0016417) |
| 0.2 | 1.4 | GO:0015079 | potassium ion transmembrane transporter activity(GO:0015079) |
| 0.2 | 1.5 | GO:0010181 | FMN binding(GO:0010181) |
| 0.2 | 6.7 | GO:0015399 | primary active transmembrane transporter activity(GO:0015399) P-P-bond-hydrolysis-driven transmembrane transporter activity(GO:0015405) |
| 0.2 | 0.5 | GO:0004430 | 1-phosphatidylinositol 4-kinase activity(GO:0004430) |
| 0.2 | 2.4 | GO:0000993 | RNA polymerase II core binding(GO:0000993) |
| 0.2 | 2.2 | GO:0001102 | RNA polymerase II activating transcription factor binding(GO:0001102) |
| 0.2 | 1.8 | GO:0030983 | mismatched DNA binding(GO:0030983) |
| 0.2 | 0.6 | GO:0008195 | phosphatidate phosphatase activity(GO:0008195) |
| 0.1 | 1.3 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.1 | 16.1 | GO:0004674 | protein serine/threonine kinase activity(GO:0004674) |
| 0.1 | 7.2 | GO:0000982 | transcription factor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0000982) |
| 0.1 | 0.4 | GO:0016903 | oxidoreductase activity, acting on the aldehyde or oxo group of donors(GO:0016903) |
| 0.1 | 0.7 | GO:0008601 | protein phosphatase type 2A regulator activity(GO:0008601) |
| 0.1 | 1.6 | GO:0000384 | first spliceosomal transesterification activity(GO:0000384) |
| 0.1 | 0.4 | GO:0070336 | flap-structured DNA binding(GO:0070336) |
| 0.1 | 0.5 | GO:0008171 | O-methyltransferase activity(GO:0008171) |
| 0.1 | 0.5 | GO:0003916 | DNA topoisomerase activity(GO:0003916) |
| 0.1 | 3.0 | GO:0032266 | phosphatidylinositol-3-phosphate binding(GO:0032266) |
| 0.1 | 0.6 | GO:0016783 | sulfurtransferase activity(GO:0016783) |
| 0.1 | 0.6 | GO:0017169 | CDP-alcohol phosphatidyltransferase activity(GO:0017169) |
| 0.1 | 1.9 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.1 | 1.5 | GO:0001104 | RNA polymerase II transcription cofactor activity(GO:0001104) |
| 0.1 | 1.0 | GO:0017025 | TBP-class protein binding(GO:0017025) |
| 0.1 | 1.0 | GO:0045182 | translation regulator activity(GO:0045182) |
| 0.1 | 0.7 | GO:0018024 | histone-lysine N-methyltransferase activity(GO:0018024) |
| 0.1 | 2.0 | GO:0004722 | protein serine/threonine phosphatase activity(GO:0004722) |
| 0.1 | 0.2 | GO:0043813 | phosphatidylinositol-3,5-bisphosphate 5-phosphatase activity(GO:0043813) |
| 0.1 | 0.2 | GO:0004180 | carboxypeptidase activity(GO:0004180) |
| 0.1 | 0.2 | GO:0019789 | SUMO transferase activity(GO:0019789) |
| 0.1 | 0.2 | GO:0030943 | mitochondrion targeting sequence binding(GO:0030943) |
| 0.0 | 0.2 | GO:0034450 | ubiquitin-ubiquitin ligase activity(GO:0034450) |
| 0.0 | 0.7 | GO:0043022 | ribosome binding(GO:0043022) |
| 0.0 | 0.2 | GO:0050072 | m7G(5')pppN diphosphatase activity(GO:0050072) |
| 0.0 | 2.2 | GO:0004842 | ubiquitin-protein transferase activity(GO:0004842) |
| 0.0 | 0.1 | GO:0031683 | G-protein beta/gamma-subunit complex binding(GO:0031683) |
| 0.0 | 0.1 | GO:0070577 | lysine-acetylated histone binding(GO:0070577) |
| 0.0 | 0.2 | GO:0008028 | monocarboxylic acid transmembrane transporter activity(GO:0008028) |
| 0.0 | 0.4 | GO:0008236 | serine-type peptidase activity(GO:0008236) |
| 0.0 | 0.1 | GO:0004128 | cytochrome-b5 reductase activity, acting on NAD(P)H(GO:0004128) |
| 0.0 | 0.1 | GO:0008143 | poly(A) binding(GO:0008143) poly-purine tract binding(GO:0070717) |
| 0.0 | 0.1 | GO:0001034 | RNA polymerase III type 2 promoter sequence-specific DNA binding(GO:0001003) transcription factor activity, RNA polymerase III promoter sequence-specific binding, TFIIIB recruiting(GO:0001004) transcription factor activity, RNA polymerase III type 1 promoter sequence-specific binding, TFIIIB recruiting(GO:0001005) transcription factor activity, RNA polymerase III type 2 promoter sequence-specific binding, TFIIIB recruiting(GO:0001008) RNA polymerase III type 2 promoter DNA binding(GO:0001031) RNA polymerase III transcription factor activity, sequence-specific DNA binding(GO:0001034) |
| 0.0 | 0.1 | GO:0016742 | hydroxymethyl-, formyl- and related transferase activity(GO:0016742) |
| 0.0 | 0.1 | GO:0030276 | clathrin binding(GO:0030276) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.9 | 14.8 | PID IL4 2PATHWAY | IL4-mediated signaling events |
| 2.4 | 7.2 | PID AR PATHWAY | Coregulation of Androgen receptor activity |
| 2.0 | 6.1 | PID ERBB4 PATHWAY | ErbB4 signaling events |
| 1.2 | 2.4 | PID ANGIOPOIETIN RECEPTOR PATHWAY | Angiopoietin receptor Tie2-mediated signaling |
| 0.9 | 0.9 | PID ERBB1 RECEPTOR PROXIMAL PATHWAY | EGF receptor (ErbB1) signaling pathway |
| 0.9 | 1.8 | ST DIFFERENTIATION PATHWAY IN PC12 CELLS | Differentiation Pathway in PC12 Cells; this is a specific case of PAC1 Receptor Pathway. |
| 0.4 | 581.8 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.3 | 1.9 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.2 | 0.7 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.2 | 0.5 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.9 | 14.6 | REACTOME MEIOTIC SYNAPSIS | Genes involved in Meiotic Synapsis |
| 2.3 | 7.0 | REACTOME SHC1 EVENTS IN ERBB4 SIGNALING | Genes involved in SHC1 events in ERBB4 signaling |
| 1.4 | 1.4 | REACTOME LAGGING STRAND SYNTHESIS | Genes involved in Lagging Strand Synthesis |
| 0.9 | 2.8 | REACTOME ETHANOL OXIDATION | Genes involved in Ethanol oxidation |
| 0.9 | 0.9 | REACTOME PI3K EVENTS IN ERBB2 SIGNALING | Genes involved in PI3K events in ERBB2 signaling |
| 0.9 | 0.9 | REACTOME PHASE1 FUNCTIONALIZATION OF COMPOUNDS | Genes involved in Phase 1 - Functionalization of compounds |
| 0.4 | 1.8 | REACTOME NGF SIGNALLING VIA TRKA FROM THE PLASMA MEMBRANE | Genes involved in NGF signalling via TRKA from the plasma membrane |
| 0.4 | 1.7 | REACTOME PHOSPHORYLATION OF THE APC C | Genes involved in Phosphorylation of the APC/C |
| 0.4 | 3.9 | REACTOME DNA REPAIR | Genes involved in DNA Repair |
| 0.4 | 3.4 | REACTOME MEIOTIC RECOMBINATION | Genes involved in Meiotic Recombination |
| 0.4 | 567.2 | REACTOME CHONDROITIN SULFATE BIOSYNTHESIS | Genes involved in Chondroitin sulfate biosynthesis |
| 0.3 | 0.9 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.1 | 0.8 | REACTOME TRANSMEMBRANE TRANSPORT OF SMALL MOLECULES | Genes involved in Transmembrane transport of small molecules |
| 0.1 | 0.9 | REACTOME RNA POL II TRANSCRIPTION PRE INITIATION AND PROMOTER OPENING | Genes involved in RNA Polymerase II Transcription Pre-Initiation And Promoter Opening |
| 0.1 | 0.2 | REACTOME SYNTHESIS OF PIPS AT THE LATE ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the late endosome membrane |
| 0.1 | 0.2 | REACTOME SIGNALING BY ERBB4 | Genes involved in Signaling by ERBB4 |
| 0.0 | 0.7 | REACTOME BIOSYNTHESIS OF THE N GLYCAN PRECURSOR DOLICHOL LIPID LINKED OLIGOSACCHARIDE LLO AND TRANSFER TO A NASCENT PROTEIN | Genes involved in Biosynthesis of the N-glycan precursor (dolichol lipid-linked oligosaccharide, LLO) and transfer to a nascent protein |