Project
GSE53960: rat RNA-Seq transcriptomic Bodymap
Navigation
Downloads
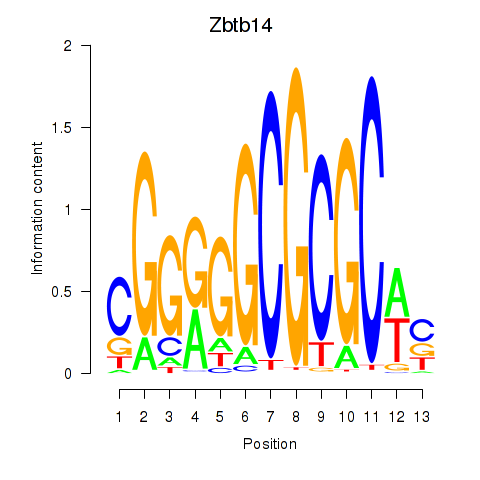
Results for Zbtb14
Z-value: 1.26
Transcription factors associated with Zbtb14
| Gene Symbol | Gene ID | Gene Info |
|---|
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Zfp161 | rn6_v1_chr9_+_117737611_117737611 | 0.24 | 1.0e-05 | Click! |
Activity profile of Zbtb14 motif
Sorted Z-values of Zbtb14 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr10_-_14056169 | 61.11 |
ENSRNOT00000017833
|
Syngr3
|
synaptogyrin 3 |
| chr13_-_49522415 | 57.84 |
ENSRNOT00000060871
ENSRNOT00000051512 |
Nfasc
|
neurofascin |
| chr8_-_116965396 | 56.89 |
ENSRNOT00000042528
|
Bsn
|
bassoon (presynaptic cytomatrix protein) |
| chr10_-_92008082 | 50.31 |
ENSRNOT00000006361
|
Nsf
|
N-ethylmaleimide sensitive factor, vesicle fusing ATPase |
| chr4_+_49941304 | 49.90 |
ENSRNOT00000008719
|
Ptprz1
|
protein tyrosine phosphatase, receptor type Z1 |
| chr2_-_119537837 | 46.51 |
ENSRNOT00000015200
|
Pex5l
|
peroxisomal biogenesis factor 5-like |
| chr3_-_2534375 | 45.98 |
ENSRNOT00000037725
|
Grin1
|
glutamate ionotropic receptor NMDA type subunit 1 |
| chr19_+_41482728 | 44.95 |
ENSRNOT00000022943
|
Calb2
|
calbindin 2 |
| chr9_+_38297322 | 44.66 |
ENSRNOT00000078157
ENSRNOT00000088824 |
Bend6
|
BEN domain containing 6 |
| chr8_+_39305128 | 43.40 |
ENSRNOT00000008285
|
Fez1
|
fasciculation and elongation protein zeta 1 |
| chr10_+_92288910 | 38.87 |
ENSRNOT00000006947
ENSRNOT00000045127 |
Mapt
|
microtubule-associated protein tau |
| chr4_-_169999873 | 38.81 |
ENSRNOT00000011697
|
Grin2b
|
glutamate ionotropic receptor NMDA type subunit 2B |
| chr13_+_101181994 | 37.99 |
ENSRNOT00000052407
|
Susd4
|
sushi domain containing 4 |
| chr19_-_11326139 | 37.55 |
ENSRNOT00000025669
|
Mt3
|
metallothionein 3 |
| chr14_-_82048251 | 37.13 |
ENSRNOT00000074734
|
Nat8l
|
N-acetyltransferase 8-like |
| chr10_+_92289107 | 36.91 |
ENSRNOT00000050070
|
Mapt
|
microtubule-associated protein tau |
| chr8_-_23063041 | 36.46 |
ENSRNOT00000018416
|
Elavl3
|
ELAV like RNA binding protein 3 |
| chr15_+_108908607 | 35.86 |
ENSRNOT00000089455
|
Zic2
|
Zic family member 2 |
| chr2_-_188645196 | 35.72 |
ENSRNOT00000083793
|
Efna3
|
ephrin A3 |
| chr19_-_11669578 | 35.59 |
ENSRNOT00000026373
|
Gnao1
|
G protein subunit alpha o1 |
| chr7_-_130827152 | 35.56 |
ENSRNOT00000019406
|
Syt10
|
synaptotagmin 10 |
| chr15_+_52220578 | 35.13 |
ENSRNOT00000015104
|
Lgi3
|
leucine-rich repeat LGI family, member 3 |
| chr8_+_119566509 | 33.62 |
ENSRNOT00000028633
|
Trank1
|
tetratricopeptide repeat and ankyrin repeat containing 1 |
| chr10_-_108691367 | 33.61 |
ENSRNOT00000005067
|
Nptx1
|
neuronal pentraxin 1 |
| chr5_+_166533181 | 33.57 |
ENSRNOT00000045063
|
Clstn1
|
calsyntenin 1 |
| chr14_-_86297623 | 33.54 |
ENSRNOT00000067162
ENSRNOT00000081607 ENSRNOT00000085265 |
Camk2b
|
calcium/calmodulin-dependent protein kinase II beta |
| chrX_+_106523278 | 33.37 |
ENSRNOT00000070802
|
MGC109340
|
similar to Microsomal signal peptidase 23 kDa subunit (SPase 22 kDa subunit) (SPC22/23) |
| chr1_+_82174451 | 33.18 |
ENSRNOT00000027783
|
Tmem145
|
transmembrane protein 145 |
| chr5_-_58019836 | 32.71 |
ENSRNOT00000066977
|
Enho
|
energy homeostasis associated |
| chr14_+_87312203 | 32.57 |
ENSRNOT00000088032
|
Adcy1
|
adenylate cyclase 1 |
| chr7_-_116607674 | 32.22 |
ENSRNOT00000076014
|
Ly6h
|
lymphocyte antigen 6 complex, locus H |
| chr3_+_137618898 | 31.91 |
ENSRNOT00000007249
|
Pcsk2
|
proprotein convertase subtilisin/kexin type 2 |
| chr19_-_37245217 | 31.88 |
ENSRNOT00000020927
|
MGC116202
|
hypothetical protein LOC688735 |
| chr6_-_122545899 | 31.72 |
ENSRNOT00000005175
|
Kcnk10
|
potassium two pore domain channel subfamily K member 10 |
| chr7_-_116607408 | 31.47 |
ENSRNOT00000076009
ENSRNOT00000056554 |
Ly6h
|
lymphocyte antigen 6 complex, locus H |
| chr9_+_70133969 | 30.70 |
ENSRNOT00000040525
|
Adam23
|
ADAM metallopeptidase domain 23 |
| chr8_-_2045817 | 30.36 |
ENSRNOT00000009542
ENSRNOT00000081171 ENSRNOT00000078765 |
Gria4
|
glutamate ionotropic receptor AMPA type subunit 4 |
| chrX_+_71199491 | 29.58 |
ENSRNOT00000076168
ENSRNOT00000005077 ENSRNOT00000005102 |
Nlgn3
|
neuroligin 3 |
| chr4_+_88441067 | 29.47 |
ENSRNOT00000008885
|
Vopp1
|
vesicular, overexpressed in cancer, prosurvival protein 1 |
| chr3_-_2534663 | 29.24 |
ENSRNOT00000049297
ENSRNOT00000044246 |
Grin1
|
glutamate ionotropic receptor NMDA type subunit 1 |
| chr7_-_70552897 | 29.18 |
ENSRNOT00000080594
|
Kif5a
|
kinesin family member 5A |
| chr11_+_14005732 | 29.04 |
ENSRNOT00000047240
|
Rbm11
|
RNA binding motif protein 11 |
| chr1_-_124039196 | 28.91 |
ENSRNOT00000051883
|
Chrna7
|
cholinergic receptor nicotinic alpha 7 subunit |
| chr8_-_94564525 | 28.31 |
ENSRNOT00000084437
|
Snap91
|
synaptosomal-associated protein 91 |
| chr6_+_126018841 | 28.20 |
ENSRNOT00000089840
ENSRNOT00000088089 |
Slc24a4
|
solute carrier family 24 member 4 |
| chr8_+_128972311 | 27.82 |
ENSRNOT00000025460
|
Myrip
|
myosin VIIA and Rab interacting protein |
| chr2_+_3400977 | 27.79 |
ENSRNOT00000093593
|
Mctp1
|
multiple C2 and transmembrane domain containing 1 |
| chr16_-_20439206 | 26.81 |
ENSRNOT00000026391
|
Rab3a
|
RAB3A, member RAS oncogene family |
| chr4_-_145147397 | 26.10 |
ENSRNOT00000010347
|
Lhfpl4
|
lipoma HMGIC fusion partner-like 4 |
| chrX_-_106558366 | 25.97 |
ENSRNOT00000042126
|
Bex2
|
brain expressed X-linked 2 |
| chr1_+_173607101 | 25.88 |
ENSRNOT00000074636
|
Tub
|
tubby bipartite transcription factor |
| chr1_-_146556171 | 25.74 |
ENSRNOT00000017636
|
Arnt2
|
aryl hydrocarbon receptor nuclear translocator 2 |
| chr3_-_5693147 | 25.59 |
ENSRNOT00000008660
|
Fam163b
|
family with sequence similarity 163, member B |
| chr8_+_111600532 | 25.39 |
ENSRNOT00000081952
|
Rab6b
|
RAB6B, member RAS oncogene family |
| chr1_-_212622537 | 24.71 |
ENSRNOT00000025609
|
Sprn
|
shadow of prion protein homolog (zebrafish) |
| chr1_-_195096460 | 24.20 |
ENSRNOT00000077253
|
Snrpn
|
small nuclear ribonucleoprotein polypeptide N |
| chr5_-_39215102 | 24.18 |
ENSRNOT00000050653
|
Klhl32
|
kelch-like family member 32 |
| chr1_-_195096694 | 24.11 |
ENSRNOT00000088874
|
Snurf
|
SNRPN upstream reading frame |
| chr12_-_12025549 | 23.30 |
ENSRNOT00000001331
|
Nptx2
|
neuronal pentraxin 2 |
| chr4_-_120817377 | 23.29 |
ENSRNOT00000021826
|
Podxl2
|
podocalyxin-like 2 |
| chr17_+_13670520 | 23.27 |
ENSRNOT00000019442
|
Shc3
|
SHC adaptor protein 3 |
| chr13_+_31081804 | 23.18 |
ENSRNOT00000041413
|
Cdh7
|
cadherin 7 |
| chr16_-_81945127 | 23.06 |
ENSRNOT00000023352
|
Mcf2l
|
MCF.2 cell line derived transforming sequence-like |
| chr16_-_20860767 | 23.01 |
ENSRNOT00000027275
ENSRNOT00000092417 |
Gdf1
Cers1
|
growth differentiation factor 1 ceramide synthase 1 |
| chrX_+_39711201 | 22.56 |
ENSRNOT00000080512
ENSRNOT00000009802 |
Cnksr2
|
connector enhancer of kinase suppressor of Ras 2 |
| chr14_-_78902063 | 22.55 |
ENSRNOT00000088469
|
Ppp2r2c
|
protein phosphatase 2, regulatory subunit B, gamma |
| chr15_-_48670257 | 22.11 |
ENSRNOT00000071464
|
Fzd3
|
frizzled class receptor 3 |
| chr8_-_61290240 | 22.06 |
ENSRNOT00000023084
|
Lingo1
|
leucine rich repeat and Ig domain containing 1 |
| chr14_+_39154529 | 21.43 |
ENSRNOT00000003191
|
Gabra4
|
gamma-aminobutyric acid type A receptor alpha4 subunit |
| chr10_-_88266210 | 21.19 |
ENSRNOT00000090702
ENSRNOT00000020603 |
Hap1
|
huntingtin-associated protein 1 |
| chr20_+_19318250 | 21.14 |
ENSRNOT00000000299
|
Phyhipl
|
phytanoyl-CoA 2-hydroxylase interacting protein-like |
| chr9_-_28973246 | 21.12 |
ENSRNOT00000091865
ENSRNOT00000015453 |
Rims1
|
regulating synaptic membrane exocytosis 1 |
| chrX_+_106487870 | 21.11 |
ENSRNOT00000075521
|
Arxes2
|
adipocyte-related X-chromosome expressed sequence 2 |
| chr5_+_140870140 | 21.00 |
ENSRNOT00000074347
|
Hpcal4
|
hippocalcin-like 4 |
| chr3_+_97721668 | 20.74 |
ENSRNOT00000065168
|
Mpped2
|
metallophosphoesterase domain containing 2 |
| chr5_-_58078545 | 20.68 |
ENSRNOT00000075777
|
Cntfr
|
ciliary neurotrophic factor receptor |
| chr1_-_103256823 | 20.63 |
ENSRNOT00000018860
|
Ptpn5
|
protein tyrosine phosphatase, non-receptor type 5 |
| chr14_+_81725513 | 20.51 |
ENSRNOT00000020154
|
Zfyve28
|
zinc finger FYVE-type containing 28 |
| chrX_+_157150655 | 20.38 |
ENSRNOT00000090795
|
Pnck
|
pregnancy up-regulated nonubiquitous CaM kinase |
| chr1_+_266953139 | 20.34 |
ENSRNOT00000054696
|
Neurl1
|
neuralized E3 ubiquitin protein ligase 1 |
| chr8_+_27777179 | 20.33 |
ENSRNOT00000009825
|
B3gat1
|
beta-1,3-glucuronyltransferase 1 |
| chr1_-_89483988 | 20.20 |
ENSRNOT00000028603
|
Fxyd7
|
FXYD domain-containing ion transport regulator 7 |
| chr7_+_70364813 | 19.94 |
ENSRNOT00000084012
ENSRNOT00000031230 |
Agap2
|
ArfGAP with GTPase domain, ankyrin repeat and PH domain 2 |
| chr5_+_31568419 | 19.92 |
ENSRNOT00000050137
|
Mmp16
|
matrix metallopeptidase 16 |
| chr2_-_24923128 | 19.90 |
ENSRNOT00000044087
|
Pde8b
|
phosphodiesterase 8B |
| chr12_+_10254951 | 19.80 |
ENSRNOT00000061129
|
Gpr12
|
G protein-coupled receptor 12 |
| chr19_+_54553419 | 19.64 |
ENSRNOT00000025392
|
Jph3
|
junctophilin 3 |
| chr1_-_261669584 | 19.57 |
ENSRNOT00000020568
ENSRNOT00000076555 |
Crtac1
|
cartilage acidic protein 1 |
| chr3_-_109044420 | 19.53 |
ENSRNOT00000007687
|
Rasgrp1
|
RAS guanyl releasing protein 1 |
| chr1_-_112947162 | 19.41 |
ENSRNOT00000014573
ENSRNOT00000083894 |
Gabra5
|
gamma-aminobutyric acid type A receptor alpha 5 subunit |
| chr1_-_211923929 | 19.27 |
ENSRNOT00000054887
|
Nkx6-2
|
NK6 homeobox 2 |
| chr3_+_168345152 | 19.20 |
ENSRNOT00000017654
|
Dok5
|
docking protein 5 |
| chr10_-_74679858 | 19.17 |
ENSRNOT00000003859
|
Ppm1e
|
protein phosphatase, Mg2+/Mn2+ dependent, 1E |
| chr6_-_28464118 | 18.57 |
ENSRNOT00000068214
|
Efr3b
|
EFR3 homolog B |
| chr6_-_122897997 | 18.49 |
ENSRNOT00000057601
|
Eml5
|
echinoderm microtubule associated protein like 5 |
| chr1_+_266952561 | 18.31 |
ENSRNOT00000076452
|
Neurl1
|
neuralized E3 ubiquitin protein ligase 1 |
| chrX_-_140303686 | 18.26 |
ENSRNOT00000033219
|
Gpr101
|
G protein-coupled receptor 101 |
| chr12_+_10255416 | 18.16 |
ENSRNOT00000092720
|
Gpr12
|
G protein-coupled receptor 12 |
| chr14_-_84106997 | 18.08 |
ENSRNOT00000065501
|
Osbp2
|
oxysterol binding protein 2 |
| chr3_+_8534440 | 18.07 |
ENSRNOT00000045827
ENSRNOT00000082672 |
Sptan1
|
spectrin, alpha, non-erythrocytic 1 |
| chr1_-_220480132 | 18.06 |
ENSRNOT00000027421
|
Cnih2
|
cornichon family AMPA receptor auxiliary protein 2 |
| chr8_-_77089097 | 17.89 |
ENSRNOT00000078685
|
Fam63b
|
family with sequence similarity 63, member B |
| chr11_+_34598492 | 17.88 |
ENSRNOT00000065600
|
Ttc3
|
tetratricopeptide repeat domain 3 |
| chr17_+_42133076 | 17.85 |
ENSRNOT00000031384
|
Aldh5a1
|
aldehyde dehydrogenase 5 family, member A1 |
| chr5_-_28737719 | 17.69 |
ENSRNOT00000009718
|
Necab1
|
N-terminal EF-hand calcium binding protein 1 |
| chr1_-_242441247 | 17.59 |
ENSRNOT00000068645
|
Pip5k1b
|
phosphatidylinositol-4-phosphate 5-kinase type 1 beta |
| chr14_+_100217289 | 17.59 |
ENSRNOT00000077171
ENSRNOT00000007104 |
Cnrip1
|
cannabinoid receptor interacting protein 1 |
| chr1_-_242440885 | 17.57 |
ENSRNOT00000076537
|
Pip5k1b
|
phosphatidylinositol-4-phosphate 5-kinase type 1 beta |
| chr8_+_94821247 | 17.48 |
ENSRNOT00000081848
|
Mrap2
|
melanocortin 2 receptor accessory protein 2 |
| chr7_-_145154131 | 17.42 |
ENSRNOT00000055271
|
Ppp1r1a
|
protein phosphatase 1, regulatory (inhibitor) subunit 1A |
| chrX_+_156963870 | 17.40 |
ENSRNOT00000077109
|
Pdzd4
|
PDZ domain containing 4 |
| chr10_-_109224370 | 17.26 |
ENSRNOT00000005839
|
Aatk
|
apoptosis-associated tyrosine kinase |
| chr1_-_84254645 | 17.23 |
ENSRNOT00000077651
|
Sptbn4
|
spectrin, beta, non-erythrocytic 4 |
| chr4_+_118655728 | 16.83 |
ENSRNOT00000043082
|
Aak1
|
AP2 associated kinase 1 |
| chr5_+_148193710 | 16.70 |
ENSRNOT00000088568
|
Adgrb2
|
adhesion G protein-coupled receptor B2 |
| chr8_+_116154736 | 16.62 |
ENSRNOT00000021218
|
Cacna2d2
|
calcium voltage-gated channel auxiliary subunit alpha2delta 2 |
| chr6_+_41917132 | 16.39 |
ENSRNOT00000071387
|
Ntsr2
|
neurotensin receptor 2 |
| chr6_-_124735741 | 16.21 |
ENSRNOT00000064716
ENSRNOT00000091693 |
Rps6ka5
|
ribosomal protein S6 kinase A5 |
| chr13_-_82758004 | 16.10 |
ENSRNOT00000003932
|
Atp1b1
|
ATPase Na+/K+ transporting subunit beta 1 |
| chr4_+_99618622 | 16.00 |
ENSRNOT00000064772
|
Reep1
|
receptor accessory protein 1 |
| chr1_+_263186235 | 15.82 |
ENSRNOT00000021876
|
Cnnm1
|
cyclin and CBS domain divalent metal cation transport mediator 1 |
| chr4_+_57204944 | 15.74 |
ENSRNOT00000085027
|
Strip2
|
striatin interacting protein 2 |
| chr10_-_20622600 | 15.44 |
ENSRNOT00000020554
|
Fbll1
|
fibrillarin-like 1 |
| chr6_-_39363367 | 15.28 |
ENSRNOT00000088687
ENSRNOT00000065531 |
Fam84a
|
family with sequence similarity 84, member A |
| chr4_-_123040609 | 15.24 |
ENSRNOT00000070832
|
Wnt7a
|
wingless-type MMTV integration site family, member 7A |
| chr1_+_202432366 | 15.16 |
ENSRNOT00000027681
|
Plpp4
|
phospholipid phosphatase 4 |
| chr17_+_21382455 | 15.15 |
ENSRNOT00000019756
|
Elovl2
|
ELOVL fatty acid elongase 2 |
| chr9_-_28972835 | 15.14 |
ENSRNOT00000086967
ENSRNOT00000079684 |
Rims1
|
regulating synaptic membrane exocytosis 1 |
| chr3_+_145032200 | 15.10 |
ENSRNOT00000068210
|
Syndig1
|
synapse differentiation inducing 1 |
| chr3_+_177013604 | 15.07 |
ENSRNOT00000020581
|
Dnajc5
|
DnaJ heat shock protein family (Hsp40) member C5 |
| chr6_+_83083740 | 14.96 |
ENSRNOT00000007600
|
Lrfn5
|
leucine rich repeat and fibronectin type III domain containing 5 |
| chr7_+_73588163 | 14.96 |
ENSRNOT00000015707
|
Kcns2
|
potassium voltage-gated channel, modifier subfamily S, member 2 |
| chr9_-_114619711 | 14.92 |
ENSRNOT00000050012
|
Mtcl1
|
microtubule crosslinking factor 1 |
| chr3_+_154043873 | 14.82 |
ENSRNOT00000072502
ENSRNOT00000034166 |
Nnat
|
neuronatin |
| chr18_-_56331991 | 14.73 |
ENSRNOT00000085841
ENSRNOT00000091220 |
Slc6a7
|
solute carrier family 6 member 7 |
| chr15_-_88036354 | 14.66 |
ENSRNOT00000014747
|
Ednrb
|
endothelin receptor type B |
| chr6_-_132958546 | 14.61 |
ENSRNOT00000041903
|
Begain
|
brain-enriched guanylate kinase-associated |
| chr1_-_78851719 | 14.50 |
ENSRNOT00000022603
|
Calm2
|
calmodulin 2 |
| chr15_-_28313841 | 14.47 |
ENSRNOT00000085897
|
Ndrg2
|
NDRG family member 2 |
| chr8_-_59077690 | 14.37 |
ENSRNOT00000066197
|
Dmxl2
|
Dmx-like 2 |
| chrX_+_106823491 | 14.31 |
ENSRNOT00000045997
|
Bex3
|
brain expressed X-linked 3 |
| chr10_+_47282208 | 13.96 |
ENSRNOT00000057953
|
Kcnj12
|
potassium voltage-gated channel subfamily J member 12 |
| chr7_+_128500011 | 13.82 |
ENSRNOT00000074625
|
Fam19a5
|
family with sequence similarity 19 member A5, C-C motif chemokine like |
| chr17_-_78962476 | 13.73 |
ENSRNOT00000030228
|
Nmt2
|
N-myristoyltransferase 2 |
| chr3_+_103726238 | 13.69 |
ENSRNOT00000006776
|
Lpcat4
|
lysophosphatidylcholine acyltransferase 4 |
| chr7_-_59514939 | 13.68 |
ENSRNOT00000085579
|
Kcnmb4
|
potassium calcium-activated channel subfamily M regulatory beta subunit 4 |
| chr9_-_61134963 | 13.64 |
ENSRNOT00000017967
|
Pgap1
|
post-GPI attachment to proteins 1 |
| chr18_-_63357194 | 13.60 |
ENSRNOT00000089408
ENSRNOT00000066103 |
Spire1
|
spire-type actin nucleation factor 1 |
| chr3_-_153893847 | 13.57 |
ENSRNOT00000085938
|
LOC100911769
|
band 4.1-like protein 1-like |
| chr5_+_74649765 | 13.33 |
ENSRNOT00000075952
|
Palm2
|
paralemmin 2 |
| chr2_-_252691886 | 13.31 |
ENSRNOT00000068739
|
Prkacb
|
protein kinase cAMP-activated catalytic subunit beta |
| chr12_+_50270189 | 13.24 |
ENSRNOT00000086782
|
Asphd2
|
aspartate beta-hydroxylase domain containing 2 |
| chrX_+_15035569 | 13.20 |
ENSRNOT00000006491
ENSRNOT00000078392 |
Porcn
|
porcupine homolog (Drosophila) |
| chr10_-_83332851 | 13.17 |
ENSRNOT00000007133
|
Nxph3
|
neurexophilin 3 |
| chr19_-_6031655 | 13.14 |
ENSRNOT00000092016
|
AABR07042733.2
|
|
| chr13_-_98772968 | 13.14 |
ENSRNOT00000057735
|
Stum
|
stum, mechanosensory transduction mediator homolog |
| chr19_-_55183557 | 12.98 |
ENSRNOT00000017317
|
Mlnr
|
motilin receptor |
| chr14_-_43072843 | 12.97 |
ENSRNOT00000064263
|
Limch1
|
LIM and calponin homology domains 1 |
| chr10_-_98644800 | 12.96 |
ENSRNOT00000005848
ENSRNOT00000075561 |
Abca5
|
ATP binding cassette subfamily A member 5 |
| chr6_-_132972511 | 12.92 |
ENSRNOT00000082216
|
Begain
|
brain-enriched guanylate kinase-associated |
| chrX_+_14358224 | 12.89 |
ENSRNOT00000005042
|
Lancl3
|
LanC like 3 |
| chr5_+_67708676 | 12.88 |
ENSRNOT00000075061
|
AABR07048211.1
|
|
| chr6_-_108415093 | 12.80 |
ENSRNOT00000031650
|
Syndig1l
|
synapse differentiation inducing 1-like |
| chr6_+_44009872 | 12.79 |
ENSRNOT00000082657
|
Mboat2
|
membrane bound O-acyltransferase domain containing 2 |
| chr4_+_168976859 | 12.70 |
ENSRNOT00000068663
|
Fam234b
|
family with sequence similarity 234, member B |
| chr6_-_49324450 | 12.62 |
ENSRNOT00000066904
|
Sntg2
|
syntrophin, gamma 2 |
| chr10_+_74436208 | 12.60 |
ENSRNOT00000090921
ENSRNOT00000091163 |
Trim37
|
tripartite motif-containing 37 |
| chr12_+_22026075 | 12.58 |
ENSRNOT00000029041
|
LOC100910838
|
neuronal tyrosine-phosphorylated phosphoinositide-3-kinase adapter 1-like |
| chr10_+_55927223 | 12.45 |
ENSRNOT00000011523
|
Kcnab3
|
potassium voltage-gated channel subfamily A regulatory beta subunit 3 |
| chr1_+_15620653 | 12.29 |
ENSRNOT00000017401
|
Map7
|
microtubule-associated protein 7 |
| chr5_+_143500441 | 12.20 |
ENSRNOT00000045513
|
Grik3
|
glutamate ionotropic receptor kainate type subunit 3 |
| chr1_-_265573117 | 12.13 |
ENSRNOT00000044195
ENSRNOT00000055915 |
LOC100911951
|
Kv channel-interacting protein 2-like |
| chr13_-_15395617 | 12.12 |
ENSRNOT00000046060
|
LOC304725
|
similar to contactin associated protein-like 5 isoform 1 |
| chr14_-_84978255 | 12.04 |
ENSRNOT00000078179
|
Cabp7
|
calcium binding protein 7 |
| chr10_-_85517683 | 11.89 |
ENSRNOT00000016070
|
Srcin1
|
SRC kinase signaling inhibitor 1 |
| chr10_+_74436534 | 11.86 |
ENSRNOT00000008298
|
Trim37
|
tripartite motif-containing 37 |
| chr16_+_54765325 | 11.83 |
ENSRNOT00000065327
ENSRNOT00000086899 |
Mtmr7
|
myotubularin related protein 7 |
| chr5_+_173183990 | 11.82 |
ENSRNOT00000068062
|
Tmem240
|
transmembrane protein 240 |
| chr4_+_438668 | 11.76 |
ENSRNOT00000008951
|
En2
|
engrailed homeobox 2 |
| chr4_+_13405136 | 11.67 |
ENSRNOT00000091004
|
Gnai1
|
G protein subunit alpha i1 |
| chr10_+_35392762 | 11.63 |
ENSRNOT00000059277
|
Rasgef1c
|
RasGEF domain family, member 1C |
| chr3_+_58632476 | 11.51 |
ENSRNOT00000010630
|
Rapgef4
|
Rap guanine nucleotide exchange factor 4 |
| chr9_+_27333956 | 11.50 |
ENSRNOT00000037489
|
Tmem14a
|
transmembrane protein 14A |
| chr10_-_47857326 | 11.46 |
ENSRNOT00000085703
ENSRNOT00000091963 |
Epn2
|
epsin 2 |
| chr18_-_27159693 | 11.45 |
ENSRNOT00000027345
|
Reep5
|
receptor accessory protein 5 |
| chr10_-_64642292 | 11.43 |
ENSRNOT00000084670
|
Abr
|
active BCR-related |
| chr6_+_107596782 | 11.42 |
ENSRNOT00000076882
|
Dnal1
|
dynein, axonemal, light chain 1 |
| chr10_-_13107771 | 11.37 |
ENSRNOT00000005879
|
Flywch1
|
FLYWCH-type zinc finger 1 |
| chr18_-_5314511 | 11.32 |
ENSRNOT00000022637
ENSRNOT00000079682 |
Zfp521
|
zinc finger protein 521 |
| chr4_+_25825567 | 11.26 |
ENSRNOT00000009417
|
Cdk14
|
cyclin-dependent kinase 14 |
| chr20_-_45815940 | 11.23 |
ENSRNOT00000073276
|
Gpr6
|
G protein-coupled receptor 6 |
| chr1_+_165237847 | 11.16 |
ENSRNOT00000022963
|
Pgm2l1
|
phosphoglucomutase 2-like 1 |
| chr7_-_70476340 | 11.13 |
ENSRNOT00000006800
|
Arhgef25
|
Rho guanine nucleotide exchange factor 25 |
| chr13_-_51076165 | 11.00 |
ENSRNOT00000004602
|
Adora1
|
adenosine A1 receptor |
| chr18_+_30954609 | 10.99 |
ENSRNOT00000086893
|
Pcdhgc3
|
protocadherin gamma subfamily C, 3 |
| chr10_-_18443934 | 10.76 |
ENSRNOT00000059895
ENSRNOT00000080021 |
Ranbp17
|
RAN binding protein 17 |
| chr14_-_80732010 | 10.69 |
ENSRNOT00000012322
|
Adra2c
|
adrenoceptor alpha 2C |
| chr3_+_81498022 | 10.60 |
ENSRNOT00000010510
|
Chst1
|
carbohydrate sulfotransferase 1 |
| chr20_+_40236437 | 10.55 |
ENSRNOT00000000743
|
Hs3st5
|
heparan sulfate-glucosamine 3-sulfotransferase 5 |
| chr11_+_84321078 | 10.55 |
ENSRNOT00000058043
|
Map6d1
|
MAP6 domain containing 1 |
Network of associatons between targets according to the STRING database.
First level regulatory network of Zbtb14
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 25.1 | 75.2 | GO:1900673 | olefin metabolic process(GO:1900673) |
| 12.7 | 38.0 | GO:0045959 | negative regulation of complement activation, classical pathway(GO:0045959) |
| 12.6 | 63.1 | GO:0031630 | regulation of synaptic vesicle fusion to presynaptic membrane(GO:0031630) |
| 12.6 | 50.3 | GO:0035494 | SNARE complex disassembly(GO:0035494) |
| 12.5 | 37.5 | GO:0097213 | cadmium ion homeostasis(GO:0055073) regulation of lysosomal membrane permeability(GO:0097213) positive regulation of lysosomal membrane permeability(GO:0097214) regulation of hydrogen peroxide catabolic process(GO:2000295) |
| 12.5 | 49.9 | GO:1900147 | Schwann cell migration(GO:0036135) regulation of Schwann cell migration(GO:1900147) |
| 11.2 | 33.5 | GO:0032430 | positive regulation of phospholipase A2 activity(GO:0032430) |
| 10.5 | 105.4 | GO:1902474 | positive regulation of protein localization to synapse(GO:1902474) |
| 9.7 | 29.2 | GO:0098971 | anterograde dendritic transport of neurotransmitter receptor complex(GO:0098971) |
| 9.7 | 38.8 | GO:0098989 | NMDA selective glutamate receptor signaling pathway(GO:0098989) |
| 9.6 | 57.8 | GO:0071205 | protein localization to juxtaparanode region of axon(GO:0071205) |
| 9.5 | 57.0 | GO:0098928 | presynaptic signal transduction(GO:0098928) presynapse to nucleus signaling pathway(GO:0099526) |
| 9.3 | 37.1 | GO:0051944 | positive regulation of neurotransmitter uptake(GO:0051582) positive regulation of dopamine uptake involved in synaptic transmission(GO:0051586) positive regulation of catecholamine uptake involved in synaptic transmission(GO:0051944) |
| 9.0 | 18.1 | GO:1902473 | regulation of protein localization to synapse(GO:1902473) |
| 8.7 | 26.1 | GO:0001982 | baroreceptor response to decreased systemic arterial blood pressure(GO:0001982) |
| 8.5 | 25.6 | GO:1900453 | negative regulation of long term synaptic depression(GO:1900453) |
| 7.7 | 23.0 | GO:0072720 | response to dithiothreitol(GO:0072720) |
| 7.2 | 50.2 | GO:0051461 | positive regulation of corticotropin secretion(GO:0051461) |
| 7.1 | 21.2 | GO:0031587 | positive regulation of inositol 1,4,5-trisphosphate-sensitive calcium-release channel activity(GO:0031587) |
| 7.0 | 28.1 | GO:1900224 | positive regulation of nodal signaling pathway involved in determination of lateral mesoderm left/right asymmetry(GO:1900224) |
| 5.8 | 29.2 | GO:0030070 | insulin processing(GO:0030070) |
| 5.4 | 16.1 | GO:1903281 | positive regulation of calcium:sodium antiporter activity(GO:1903281) |
| 5.3 | 10.5 | GO:0006498 | N-terminal protein lipidation(GO:0006498) |
| 5.1 | 15.4 | GO:0018364 | peptidyl-glutamine methylation(GO:0018364) |
| 5.1 | 20.6 | GO:0045448 | regulation of mitotic cell cycle, embryonic(GO:0009794) mitotic cell cycle, embryonic(GO:0045448) |
| 4.6 | 13.7 | GO:0018201 | peptidyl-glycine modification(GO:0018201) |
| 4.6 | 22.8 | GO:0014045 | establishment of endothelial blood-brain barrier(GO:0014045) central nervous system vasculogenesis(GO:0022009) |
| 4.5 | 13.6 | GO:0070649 | formin-nucleated actin cable assembly(GO:0070649) |
| 4.4 | 13.2 | GO:0034486 | vacuolar transmembrane transport(GO:0034486) chaperone-mediated protein transport involved in chaperone-mediated autophagy(GO:0061741) |
| 4.4 | 48.1 | GO:0006465 | signal peptide processing(GO:0006465) |
| 4.4 | 13.1 | GO:0006083 | acetate metabolic process(GO:0006083) |
| 4.3 | 25.9 | GO:0097500 | receptor localization to nonmotile primary cilium(GO:0097500) |
| 4.1 | 16.2 | GO:0043987 | histone H3-S10 phosphorylation(GO:0043987) histone H3-S28 phosphorylation(GO:0043988) histone H2A phosphorylation(GO:1990164) |
| 4.0 | 28.3 | GO:0016185 | synaptic vesicle budding from presynaptic endocytic zone membrane(GO:0016185) |
| 4.0 | 20.2 | GO:1901621 | negative regulation of smoothened signaling pathway involved in dorsal/ventral neural tube patterning(GO:1901621) |
| 4.0 | 19.9 | GO:0035106 | operant conditioning(GO:0035106) |
| 3.9 | 15.6 | GO:0036353 | histone H2A-K119 monoubiquitination(GO:0036353) |
| 3.9 | 19.3 | GO:0021913 | regulation of transcription from RNA polymerase II promoter involved in spinal cord motor neuron fate specification(GO:0021912) regulation of transcription from RNA polymerase II promoter involved in ventral spinal cord interneuron specification(GO:0021913) |
| 3.8 | 11.5 | GO:1904457 | positive regulation of neuronal action potential(GO:1904457) |
| 3.8 | 11.4 | GO:0043314 | negative regulation of neutrophil degranulation(GO:0043314) |
| 3.8 | 34.2 | GO:2001023 | regulation of response to drug(GO:2001023) |
| 3.7 | 14.7 | GO:0014826 | vein smooth muscle contraction(GO:0014826) |
| 3.6 | 14.5 | GO:0090360 | platelet-derived growth factor production(GO:0090360) regulation of platelet-derived growth factor production(GO:0090361) |
| 3.5 | 49.3 | GO:0045741 | positive regulation of epidermal growth factor-activated receptor activity(GO:0045741) |
| 3.5 | 10.5 | GO:0015015 | heparan sulfate proteoglycan biosynthetic process, enzymatic modification(GO:0015015) |
| 3.4 | 6.7 | GO:0050893 | sensory processing(GO:0050893) |
| 3.3 | 9.9 | GO:0021849 | neuroblast division in subventricular zone(GO:0021849) |
| 3.3 | 9.8 | GO:0090427 | activation of meiosis(GO:0090427) |
| 3.1 | 33.6 | GO:0060385 | axonogenesis involved in innervation(GO:0060385) |
| 3.1 | 9.2 | GO:0046078 | dUMP metabolic process(GO:0046078) |
| 3.0 | 15.1 | GO:0034626 | fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 3.0 | 14.9 | GO:2000574 | regulation of microtubule motor activity(GO:2000574) |
| 2.8 | 8.4 | GO:0032377 | regulation of intracellular lipid transport(GO:0032377) regulation of intracellular sterol transport(GO:0032380) regulation of intracellular cholesterol transport(GO:0032383) |
| 2.8 | 19.5 | GO:0032252 | secretory granule localization(GO:0032252) |
| 2.8 | 8.3 | GO:0038028 | insulin receptor signaling pathway via phosphatidylinositol 3-kinase(GO:0038028) |
| 2.7 | 10.6 | GO:0042339 | keratan sulfate metabolic process(GO:0042339) |
| 2.6 | 7.8 | GO:1902276 | pancreatic amylase secretion(GO:0036395) regulation of pancreatic amylase secretion(GO:1902276) |
| 2.5 | 2.5 | GO:0045715 | negative regulation of low-density lipoprotein particle receptor biosynthetic process(GO:0045715) |
| 2.5 | 17.2 | GO:0045162 | clustering of voltage-gated sodium channels(GO:0045162) |
| 2.4 | 7.3 | GO:0002035 | brain renin-angiotensin system(GO:0002035) |
| 2.4 | 14.5 | GO:0099566 | regulation of postsynaptic cytosolic calcium ion concentration(GO:0099566) |
| 2.4 | 9.5 | GO:2000277 | positive regulation of oxidative phosphorylation uncoupler activity(GO:2000277) |
| 2.4 | 16.6 | GO:0060024 | rhythmic synaptic transmission(GO:0060024) |
| 2.3 | 20.7 | GO:0003360 | brainstem development(GO:0003360) |
| 2.2 | 63.0 | GO:1903831 | acetylcholine receptor signaling pathway(GO:0095500) signal transduction involved in cellular response to ammonium ion(GO:1903831) response to acetylcholine(GO:1905144) cellular response to acetylcholine(GO:1905145) |
| 2.2 | 6.5 | GO:0007181 | transforming growth factor beta receptor complex assembly(GO:0007181) |
| 2.1 | 8.6 | GO:1901910 | diadenosine polyphosphate catabolic process(GO:0015961) diphosphoinositol polyphosphate metabolic process(GO:0071543) diadenosine pentaphosphate metabolic process(GO:1901906) diadenosine pentaphosphate catabolic process(GO:1901907) diadenosine hexaphosphate metabolic process(GO:1901908) diadenosine hexaphosphate catabolic process(GO:1901909) adenosine 5'-(hexahydrogen pentaphosphate) metabolic process(GO:1901910) adenosine 5'-(hexahydrogen pentaphosphate) catabolic process(GO:1901911) |
| 2.1 | 8.5 | GO:0021636 | trigeminal nerve morphogenesis(GO:0021636) trigeminal nerve structural organization(GO:0021637) semaphorin-plexin signaling pathway involved in axon guidance(GO:1902287) |
| 2.1 | 8.5 | GO:0070650 | actin filament bundle distribution(GO:0070650) |
| 2.0 | 33.2 | GO:0051386 | regulation of neurotrophin TRK receptor signaling pathway(GO:0051386) |
| 1.9 | 23.3 | GO:1902902 | negative regulation of autophagosome assembly(GO:1902902) |
| 1.9 | 28.9 | GO:0035970 | peptidyl-threonine dephosphorylation(GO:0035970) |
| 1.9 | 15.1 | GO:0097091 | synaptic vesicle clustering(GO:0097091) |
| 1.9 | 3.7 | GO:0016094 | polyprenol biosynthetic process(GO:0016094) |
| 1.9 | 7.5 | GO:0010615 | positive regulation of cardiac muscle adaptation(GO:0010615) positive regulation of cardiac muscle hypertrophy in response to stress(GO:1903244) regulation of connective tissue replacement(GO:1905203) |
| 1.8 | 47.6 | GO:0035235 | ionotropic glutamate receptor signaling pathway(GO:0035235) |
| 1.8 | 5.4 | GO:0060128 | positive regulation of cell fate specification(GO:0042660) corticotropin hormone secreting cell differentiation(GO:0060128) |
| 1.8 | 24.5 | GO:0032482 | Rab protein signal transduction(GO:0032482) |
| 1.7 | 5.0 | GO:2000474 | regulation of opioid receptor signaling pathway(GO:2000474) |
| 1.6 | 4.9 | GO:0098749 | cerebellar neuron development(GO:0098749) |
| 1.6 | 4.9 | GO:1900748 | positive regulation of vascular endothelial growth factor signaling pathway(GO:1900748) |
| 1.6 | 4.8 | GO:0042489 | negative regulation of odontogenesis of dentin-containing tooth(GO:0042489) |
| 1.6 | 55.6 | GO:0045956 | positive regulation of calcium ion-dependent exocytosis(GO:0045956) |
| 1.6 | 7.8 | GO:0042780 | tRNA 3'-end processing(GO:0042780) |
| 1.5 | 13.6 | GO:0015791 | polyol transport(GO:0015791) |
| 1.5 | 19.6 | GO:1900121 | negative regulation of receptor binding(GO:1900121) |
| 1.5 | 14.7 | GO:0035524 | proline transmembrane transport(GO:0035524) |
| 1.5 | 13.2 | GO:0098596 | vocal learning(GO:0042297) imitative learning(GO:0098596) |
| 1.4 | 13.0 | GO:0010745 | negative regulation of macrophage derived foam cell differentiation(GO:0010745) |
| 1.4 | 7.2 | GO:0001579 | medium-chain fatty acid transport(GO:0001579) |
| 1.4 | 19.9 | GO:0035988 | chondrocyte proliferation(GO:0035988) |
| 1.4 | 24.2 | GO:0070262 | peptidyl-serine dephosphorylation(GO:0070262) |
| 1.4 | 13.9 | GO:0046855 | inositol phosphate dephosphorylation(GO:0046855) |
| 1.4 | 4.2 | GO:0099541 | diacylglycerol catabolic process(GO:0046340) retrograde trans-synaptic signaling(GO:0098917) trans-synaptic signaling by lipid(GO:0099541) trans-synaptic signaling by endocannabinoid(GO:0099542) |
| 1.4 | 22.1 | GO:0048715 | negative regulation of oligodendrocyte differentiation(GO:0048715) |
| 1.4 | 9.5 | GO:2000467 | positive regulation of glycogen (starch) synthase activity(GO:2000467) |
| 1.3 | 37.5 | GO:0045746 | negative regulation of Notch signaling pathway(GO:0045746) |
| 1.3 | 46.3 | GO:0045747 | positive regulation of Notch signaling pathway(GO:0045747) |
| 1.3 | 2.6 | GO:0071579 | regulation of zinc ion transport(GO:0071579) |
| 1.3 | 9.1 | GO:0050915 | sensory perception of sour taste(GO:0050915) |
| 1.3 | 56.6 | GO:0046854 | phosphatidylinositol phosphorylation(GO:0046854) |
| 1.2 | 6.2 | GO:0038026 | reelin-mediated signaling pathway(GO:0038026) |
| 1.2 | 3.7 | GO:0000448 | cleavage in ITS2 between 5.8S rRNA and LSU-rRNA of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000448) |
| 1.2 | 3.6 | GO:0030505 | inorganic diphosphate transport(GO:0030505) |
| 1.2 | 11.7 | GO:1904321 | response to forskolin(GO:1904321) cellular response to forskolin(GO:1904322) |
| 1.2 | 3.5 | GO:1902303 | negative regulation of potassium ion export(GO:1902303) |
| 1.1 | 51.2 | GO:0021762 | substantia nigra development(GO:0021762) |
| 1.1 | 14.4 | GO:0070072 | vacuolar proton-transporting V-type ATPase complex assembly(GO:0070072) |
| 1.1 | 3.3 | GO:0090370 | negative regulation of cholesterol efflux(GO:0090370) |
| 1.1 | 13.0 | GO:0007175 | negative regulation of epidermal growth factor-activated receptor activity(GO:0007175) |
| 1.1 | 16.0 | GO:0071786 | endoplasmic reticulum tubular network organization(GO:0071786) |
| 1.0 | 12.5 | GO:0033601 | positive regulation of mammary gland epithelial cell proliferation(GO:0033601) |
| 1.0 | 9.4 | GO:0035330 | regulation of hippo signaling(GO:0035330) |
| 1.0 | 3.1 | GO:0032763 | regulation of mast cell cytokine production(GO:0032763) |
| 1.0 | 5.1 | GO:0060633 | negative regulation of transcription initiation from RNA polymerase II promoter(GO:0060633) |
| 1.0 | 3.0 | GO:1903721 | regulation of I-kappaB phosphorylation(GO:1903719) positive regulation of I-kappaB phosphorylation(GO:1903721) |
| 1.0 | 19.6 | GO:0060314 | regulation of ryanodine-sensitive calcium-release channel activity(GO:0060314) |
| 1.0 | 3.9 | GO:1901491 | negative regulation of lymphangiogenesis(GO:1901491) |
| 1.0 | 10.6 | GO:0045634 | regulation of melanocyte differentiation(GO:0045634) |
| 1.0 | 32.6 | GO:0010226 | response to lithium ion(GO:0010226) |
| 0.9 | 5.5 | GO:0046520 | sphingoid biosynthetic process(GO:0046520) |
| 0.9 | 9.1 | GO:0010944 | negative regulation of transcription by competitive promoter binding(GO:0010944) |
| 0.8 | 20.2 | GO:0060384 | innervation(GO:0060384) |
| 0.8 | 25.2 | GO:0007212 | dopamine receptor signaling pathway(GO:0007212) |
| 0.8 | 4.2 | GO:0006432 | phenylalanyl-tRNA aminoacylation(GO:0006432) |
| 0.8 | 5.0 | GO:0071386 | cellular response to corticosterone stimulus(GO:0071386) |
| 0.8 | 3.3 | GO:0086053 | AV node cell to bundle of His cell communication by electrical coupling(GO:0086053) |
| 0.8 | 15.6 | GO:1902259 | regulation of delayed rectifier potassium channel activity(GO:1902259) |
| 0.8 | 23.3 | GO:0050901 | leukocyte tethering or rolling(GO:0050901) |
| 0.8 | 1.6 | GO:0071895 | odontoblast differentiation(GO:0071895) |
| 0.8 | 6.3 | GO:0097210 | response to gonadotropin-releasing hormone(GO:0097210) cellular response to gonadotropin-releasing hormone(GO:0097211) |
| 0.8 | 4.7 | GO:0009169 | ribonucleoside monophosphate catabolic process(GO:0009158) purine ribonucleoside monophosphate catabolic process(GO:0009169) |
| 0.8 | 8.6 | GO:0015693 | magnesium ion transport(GO:0015693) |
| 0.7 | 9.0 | GO:0045217 | cell-cell junction maintenance(GO:0045217) |
| 0.7 | 23.9 | GO:0006890 | retrograde vesicle-mediated transport, Golgi to ER(GO:0006890) |
| 0.7 | 6.5 | GO:1903690 | negative regulation of wound healing, spreading of epidermal cells(GO:1903690) |
| 0.7 | 15.7 | GO:0061098 | positive regulation of protein tyrosine kinase activity(GO:0061098) |
| 0.7 | 2.1 | GO:0006121 | mitochondrial electron transport, succinate to ubiquinone(GO:0006121) |
| 0.7 | 4.3 | GO:1904668 | positive regulation of ubiquitin protein ligase activity(GO:1904668) |
| 0.7 | 1.4 | GO:1902951 | negative regulation of dendritic spine maintenance(GO:1902951) |
| 0.7 | 3.5 | GO:0015842 | aminergic neurotransmitter loading into synaptic vesicle(GO:0015842) |
| 0.7 | 4.1 | GO:0098881 | exocytic insertion of neurotransmitter receptor to plasma membrane(GO:0098881) exocytic insertion of neurotransmitter receptor to postsynaptic membrane(GO:0098967) |
| 0.7 | 13.8 | GO:0071880 | adenylate cyclase-activating adrenergic receptor signaling pathway(GO:0071880) |
| 0.7 | 3.4 | GO:1902162 | regulation of DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator(GO:1902162) |
| 0.7 | 2.0 | GO:0072061 | inner medullary collecting duct development(GO:0072061) |
| 0.7 | 9.5 | GO:0009312 | oligosaccharide biosynthetic process(GO:0009312) |
| 0.7 | 4.6 | GO:0010724 | regulation of definitive erythrocyte differentiation(GO:0010724) |
| 0.6 | 3.8 | GO:0048096 | chromatin-mediated maintenance of transcription(GO:0048096) |
| 0.6 | 2.5 | GO:0070086 | ubiquitin-dependent endocytosis(GO:0070086) |
| 0.6 | 6.9 | GO:0006388 | tRNA splicing, via endonucleolytic cleavage and ligation(GO:0006388) |
| 0.6 | 2.4 | GO:0017187 | peptidyl-glutamic acid carboxylation(GO:0017187) protein carboxylation(GO:0018214) |
| 0.6 | 4.3 | GO:0021796 | cerebral cortex regionalization(GO:0021796) |
| 0.6 | 2.3 | GO:2001168 | regulation of histone H2B ubiquitination(GO:2001166) positive regulation of histone H2B ubiquitination(GO:2001168) |
| 0.6 | 18.4 | GO:0035025 | positive regulation of Rho protein signal transduction(GO:0035025) |
| 0.6 | 8.5 | GO:0003376 | sphingosine-1-phosphate signaling pathway(GO:0003376) |
| 0.5 | 1.1 | GO:0015959 | diadenosine polyphosphate metabolic process(GO:0015959) |
| 0.5 | 6.6 | GO:1990403 | embryonic brain development(GO:1990403) |
| 0.5 | 9.6 | GO:0001964 | startle response(GO:0001964) |
| 0.5 | 2.1 | GO:1903336 | negative regulation of vacuolar transport(GO:1903336) |
| 0.5 | 1.9 | GO:0036476 | neuron death in response to hydrogen peroxide(GO:0036476) regulation of hydrogen peroxide-induced neuron death(GO:1903207) negative regulation of hydrogen peroxide-induced neuron death(GO:1903208) |
| 0.5 | 14.0 | GO:0010107 | potassium ion import(GO:0010107) |
| 0.5 | 9.6 | GO:0033327 | Leydig cell differentiation(GO:0033327) |
| 0.5 | 1.9 | GO:0016081 | synaptic vesicle docking(GO:0016081) |
| 0.5 | 0.5 | GO:1901069 | guanosine-containing compound catabolic process(GO:1901069) |
| 0.5 | 0.9 | GO:0050975 | sensory perception of touch(GO:0050975) |
| 0.5 | 1.4 | GO:0035790 | platelet-derived growth factor receptor-alpha signaling pathway(GO:0035790) regulation of platelet-derived growth factor receptor-alpha signaling pathway(GO:2000583) negative regulation of platelet-derived growth factor receptor-alpha signaling pathway(GO:2000584) |
| 0.4 | 1.3 | GO:0055129 | L-proline biosynthetic process(GO:0055129) |
| 0.4 | 4.0 | GO:0048712 | negative regulation of astrocyte differentiation(GO:0048712) |
| 0.4 | 1.3 | GO:0006244 | pyrimidine nucleotide catabolic process(GO:0006244) |
| 0.4 | 1.3 | GO:0019287 | isopentenyl diphosphate biosynthetic process, mevalonate pathway(GO:0019287) |
| 0.4 | 1.3 | GO:0000189 | MAPK import into nucleus(GO:0000189) |
| 0.4 | 2.1 | GO:0010032 | meiotic chromosome condensation(GO:0010032) |
| 0.4 | 3.0 | GO:0050427 | sulfate assimilation(GO:0000103) purine ribonucleoside bisphosphate metabolic process(GO:0034035) 3'-phosphoadenosine 5'-phosphosulfate metabolic process(GO:0050427) |
| 0.4 | 5.1 | GO:0007614 | short-term memory(GO:0007614) |
| 0.4 | 1.2 | GO:1901165 | positive regulation of trophoblast cell migration(GO:1901165) |
| 0.4 | 11.1 | GO:0030866 | cortical actin cytoskeleton organization(GO:0030866) |
| 0.4 | 5.4 | GO:0060850 | regulation of transcription involved in cell fate commitment(GO:0060850) |
| 0.4 | 29.2 | GO:0051965 | positive regulation of synapse assembly(GO:0051965) |
| 0.4 | 24.9 | GO:0032024 | positive regulation of insulin secretion(GO:0032024) |
| 0.4 | 1.2 | GO:0071387 | cellular response to cortisol stimulus(GO:0071387) |
| 0.4 | 2.8 | GO:0016926 | protein desumoylation(GO:0016926) |
| 0.4 | 1.2 | GO:0070084 | protein initiator methionine removal(GO:0070084) |
| 0.4 | 4.1 | GO:0002098 | tRNA wobble uridine modification(GO:0002098) |
| 0.4 | 4.1 | GO:0006013 | mannose metabolic process(GO:0006013) |
| 0.4 | 0.7 | GO:0097187 | dentinogenesis(GO:0097187) |
| 0.4 | 3.3 | GO:0090042 | tubulin deacetylation(GO:0090042) regulation of tubulin deacetylation(GO:0090043) |
| 0.4 | 2.2 | GO:0016199 | axon midline choice point recognition(GO:0016199) |
| 0.4 | 32.8 | GO:0007613 | memory(GO:0007613) |
| 0.4 | 1.1 | GO:0046604 | positive regulation of mitotic centrosome separation(GO:0046604) |
| 0.4 | 20.0 | GO:0035249 | synaptic transmission, glutamatergic(GO:0035249) |
| 0.4 | 3.5 | GO:0006527 | arginine catabolic process(GO:0006527) |
| 0.4 | 4.9 | GO:0046689 | response to mercury ion(GO:0046689) |
| 0.3 | 39.5 | GO:0035725 | sodium ion transmembrane transport(GO:0035725) |
| 0.3 | 3.1 | GO:0031145 | anaphase-promoting complex-dependent catabolic process(GO:0031145) |
| 0.3 | 4.3 | GO:0046498 | S-adenosylhomocysteine metabolic process(GO:0046498) |
| 0.3 | 19.4 | GO:0007200 | phospholipase C-activating G-protein coupled receptor signaling pathway(GO:0007200) |
| 0.3 | 1.9 | GO:0070459 | prolactin secretion(GO:0070459) |
| 0.3 | 4.1 | GO:1901673 | regulation of mitotic spindle assembly(GO:1901673) |
| 0.3 | 17.5 | GO:0030819 | positive regulation of cAMP biosynthetic process(GO:0030819) |
| 0.3 | 1.2 | GO:1904431 | positive regulation of t-circle formation(GO:1904431) |
| 0.3 | 10.0 | GO:0048013 | ephrin receptor signaling pathway(GO:0048013) |
| 0.3 | 3.9 | GO:0048733 | sebaceous gland development(GO:0048733) |
| 0.3 | 3.0 | GO:0050774 | negative regulation of dendrite morphogenesis(GO:0050774) |
| 0.3 | 13.8 | GO:0008306 | associative learning(GO:0008306) |
| 0.3 | 2.4 | GO:0000395 | mRNA 5'-splice site recognition(GO:0000395) |
| 0.3 | 1.4 | GO:0002536 | respiratory burst involved in inflammatory response(GO:0002536) |
| 0.3 | 7.0 | GO:0061462 | protein localization to lysosome(GO:0061462) |
| 0.3 | 1.6 | GO:0017196 | N-terminal peptidyl-methionine acetylation(GO:0017196) |
| 0.3 | 0.5 | GO:0070366 | regulation of hepatocyte differentiation(GO:0070366) |
| 0.3 | 0.8 | GO:0016598 | protein arginylation(GO:0016598) |
| 0.3 | 4.3 | GO:0045475 | locomotor rhythm(GO:0045475) |
| 0.3 | 3.8 | GO:0048714 | positive regulation of oligodendrocyte differentiation(GO:0048714) |
| 0.2 | 6.1 | GO:0031571 | mitotic G1 DNA damage checkpoint(GO:0031571) |
| 0.2 | 11.0 | GO:0000381 | regulation of alternative mRNA splicing, via spliceosome(GO:0000381) |
| 0.2 | 6.0 | GO:0019228 | neuronal action potential(GO:0019228) |
| 0.2 | 1.2 | GO:0048172 | regulation of short-term neuronal synaptic plasticity(GO:0048172) |
| 0.2 | 34.9 | GO:0017157 | regulation of exocytosis(GO:0017157) |
| 0.2 | 3.1 | GO:0032401 | establishment of melanosome localization(GO:0032401) melanosome transport(GO:0032402) |
| 0.2 | 4.5 | GO:0015721 | bile acid and bile salt transport(GO:0015721) |
| 0.2 | 3.4 | GO:0045576 | mast cell activation(GO:0045576) |
| 0.2 | 0.6 | GO:0097037 | heme export(GO:0097037) |
| 0.2 | 1.8 | GO:0034244 | negative regulation of transcription elongation from RNA polymerase II promoter(GO:0034244) |
| 0.2 | 12.1 | GO:0005977 | glycogen metabolic process(GO:0005977) cellular glucan metabolic process(GO:0006073) glucan metabolic process(GO:0044042) |
| 0.2 | 0.7 | GO:0044211 | CTP salvage(GO:0044211) |
| 0.2 | 7.5 | GO:1903146 | regulation of mitophagy(GO:1903146) |
| 0.2 | 0.5 | GO:0055005 | ventricular cardiac myofibril assembly(GO:0055005) |
| 0.2 | 0.3 | GO:0071431 | tRNA export from nucleus(GO:0006409) tRNA-containing ribonucleoprotein complex export from nucleus(GO:0071431) |
| 0.2 | 4.4 | GO:0046839 | phospholipid dephosphorylation(GO:0046839) |
| 0.2 | 1.1 | GO:0006152 | purine nucleoside catabolic process(GO:0006152) purine ribonucleoside catabolic process(GO:0046130) |
| 0.2 | 5.9 | GO:0048278 | vesicle docking(GO:0048278) |
| 0.2 | 1.4 | GO:1902031 | regulation of NADP metabolic process(GO:1902031) |
| 0.2 | 0.6 | GO:0045347 | negative regulation of MHC class II biosynthetic process(GO:0045347) |
| 0.1 | 0.9 | GO:0035726 | common myeloid progenitor cell proliferation(GO:0035726) |
| 0.1 | 2.6 | GO:0050650 | chondroitin sulfate proteoglycan biosynthetic process(GO:0050650) |
| 0.1 | 0.4 | GO:0044565 | dendritic cell proliferation(GO:0044565) |
| 0.1 | 2.8 | GO:0051931 | regulation of sensory perception of pain(GO:0051930) regulation of sensory perception(GO:0051931) |
| 0.1 | 0.4 | GO:0032364 | oxygen homeostasis(GO:0032364) |
| 0.1 | 5.0 | GO:1902475 | L-alpha-amino acid transmembrane transport(GO:1902475) |
| 0.1 | 9.0 | GO:0030316 | osteoclast differentiation(GO:0030316) |
| 0.1 | 9.5 | GO:0018107 | peptidyl-threonine phosphorylation(GO:0018107) |
| 0.1 | 14.7 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
| 0.1 | 4.8 | GO:0042775 | mitochondrial ATP synthesis coupled electron transport(GO:0042775) |
| 0.1 | 0.5 | GO:0031508 | pericentric heterochromatin assembly(GO:0031508) |
| 0.1 | 1.9 | GO:0010257 | NADH dehydrogenase complex assembly(GO:0010257) mitochondrial respiratory chain complex I assembly(GO:0032981) mitochondrial respiratory chain complex I biogenesis(GO:0097031) |
| 0.1 | 1.6 | GO:0045022 | early endosome to late endosome transport(GO:0045022) |
| 0.1 | 1.3 | GO:0022027 | interkinetic nuclear migration(GO:0022027) |
| 0.1 | 0.5 | GO:0035735 | intraciliary transport involved in cilium morphogenesis(GO:0035735) |
| 0.1 | 1.5 | GO:0045540 | regulation of cholesterol biosynthetic process(GO:0045540) |
| 0.1 | 0.7 | GO:0006384 | transcription initiation from RNA polymerase III promoter(GO:0006384) |
| 0.1 | 1.6 | GO:1902187 | negative regulation of viral release from host cell(GO:1902187) |
| 0.1 | 23.2 | GO:0055074 | calcium ion homeostasis(GO:0055074) |
| 0.1 | 4.0 | GO:0006835 | dicarboxylic acid transport(GO:0006835) |
| 0.1 | 1.0 | GO:2000400 | positive regulation of T cell differentiation in thymus(GO:0033089) positive regulation of thymocyte aggregation(GO:2000400) |
| 0.1 | 2.0 | GO:0006024 | glycosaminoglycan biosynthetic process(GO:0006024) |
| 0.1 | 0.2 | GO:0060467 | peptidyl-tyrosine sulfation(GO:0006478) negative regulation of fertilization(GO:0060467) prevention of polyspermy(GO:0060468) |
| 0.1 | 0.6 | GO:0097354 | protein prenylation(GO:0018342) prenylation(GO:0097354) |
| 0.1 | 0.5 | GO:0031087 | deadenylation-independent decapping of nuclear-transcribed mRNA(GO:0031087) |
| 0.1 | 0.2 | GO:0071494 | cellular response to UV-C(GO:0071494) |
| 0.1 | 5.2 | GO:0035023 | regulation of Rho protein signal transduction(GO:0035023) |
| 0.1 | 3.9 | GO:0014065 | phosphatidylinositol 3-kinase signaling(GO:0014065) |
| 0.1 | 3.3 | GO:0090502 | RNA phosphodiester bond hydrolysis, endonucleolytic(GO:0090502) |
| 0.1 | 2.2 | GO:0045669 | positive regulation of osteoblast differentiation(GO:0045669) |
| 0.0 | 0.8 | GO:0000188 | inactivation of MAPK activity(GO:0000188) |
| 0.0 | 1.6 | GO:1901379 | regulation of potassium ion transmembrane transport(GO:1901379) |
| 0.0 | 0.9 | GO:0072503 | cellular divalent inorganic cation homeostasis(GO:0072503) |
| 0.0 | 5.1 | GO:0009566 | fertilization(GO:0009566) |
| 0.0 | 1.6 | GO:0030032 | lamellipodium assembly(GO:0030032) |
| 0.0 | 0.5 | GO:0099517 | anterograde synaptic vesicle transport(GO:0048490) synaptic vesicle cytoskeletal transport(GO:0099514) synaptic vesicle transport along microtubule(GO:0099517) |
| 0.0 | 3.8 | GO:0008360 | regulation of cell shape(GO:0008360) |
| 0.0 | 2.2 | GO:0008625 | extrinsic apoptotic signaling pathway via death domain receptors(GO:0008625) |
| 0.0 | 2.1 | GO:0043491 | protein kinase B signaling(GO:0043491) |
| 0.0 | 0.1 | GO:1904380 | endoplasmic reticulum mannose trimming(GO:1904380) |
| 0.0 | 0.4 | GO:0061037 | negative regulation of chondrocyte differentiation(GO:0032331) negative regulation of cartilage development(GO:0061037) |
| 0.0 | 1.3 | GO:0045766 | positive regulation of angiogenesis(GO:0045766) |
| 0.0 | 0.9 | GO:0006887 | exocytosis(GO:0006887) |
| 0.0 | 0.4 | GO:0051693 | actin filament capping(GO:0051693) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 19.0 | 56.9 | GO:1990257 | piccolo-bassoon transport vesicle(GO:1990257) |
| 18.9 | 75.8 | GO:0045298 | tubulin complex(GO:0045298) |
| 18.8 | 75.2 | GO:0044307 | dendritic branch(GO:0044307) |
| 11.0 | 55.1 | GO:0097454 | Schwann cell microvillus(GO:0097454) |
| 10.0 | 49.9 | GO:0072534 | perineuronal net(GO:0072534) |
| 9.4 | 28.3 | GO:0098830 | presynaptic endosome(GO:0098830) presynaptic endocytic zone(GO:0098833) presynaptic endocytic zone membrane(GO:0098835) |
| 8.0 | 48.1 | GO:0005787 | signal peptidase complex(GO:0005787) |
| 6.9 | 20.7 | GO:0070110 | ciliary neurotrophic factor receptor complex(GO:0070110) |
| 6.0 | 36.3 | GO:0098831 | presynaptic active zone cytoplasmic component(GO:0098831) |
| 5.8 | 11.5 | GO:0044316 | cone cell pedicle(GO:0044316) |
| 5.1 | 30.4 | GO:0032983 | kainate selective glutamate receptor complex(GO:0032983) |
| 5.0 | 49.5 | GO:0051286 | cell tip(GO:0051286) |
| 4.7 | 37.5 | GO:0097450 | astrocyte end-foot(GO:0097450) |
| 3.9 | 38.8 | GO:0098839 | postsynaptic density membrane(GO:0098839) |
| 3.8 | 15.3 | GO:1990812 | growth cone filopodium(GO:1990812) |
| 3.5 | 35.3 | GO:0008091 | spectrin(GO:0008091) |
| 3.2 | 35.2 | GO:0001931 | uropod(GO:0001931) cell trailing edge(GO:0031254) |
| 2.9 | 11.8 | GO:0097427 | microtubule bundle(GO:0097427) |
| 2.8 | 44.8 | GO:0032279 | asymmetric synapse(GO:0032279) |
| 2.7 | 5.4 | GO:0098888 | extrinsic component of presynaptic membrane(GO:0098888) |
| 2.4 | 37.7 | GO:0031045 | dense core granule(GO:0031045) |
| 2.3 | 72.5 | GO:0097440 | apical dendrite(GO:0097440) |
| 2.2 | 19.6 | GO:0030314 | junctional membrane complex(GO:0030314) |
| 2.0 | 24.2 | GO:0005687 | U4 snRNP(GO:0005687) |
| 1.9 | 15.4 | GO:0031428 | box C/D snoRNP complex(GO:0031428) |
| 1.8 | 21.5 | GO:0005796 | Golgi lumen(GO:0005796) |
| 1.8 | 40.8 | GO:1902711 | GABA-A receptor complex(GO:1902711) |
| 1.7 | 16.8 | GO:0071439 | clathrin complex(GO:0071439) |
| 1.7 | 5.0 | GO:0044326 | dendritic spine neck(GO:0044326) |
| 1.6 | 48.2 | GO:0005921 | gap junction(GO:0005921) |
| 1.5 | 4.6 | GO:0098837 | postsynaptic recycling endosome(GO:0098837) |
| 1.4 | 7.1 | GO:0070552 | BRISC complex(GO:0070552) |
| 1.4 | 4.2 | GO:0009328 | phenylalanine-tRNA ligase complex(GO:0009328) |
| 1.3 | 99.9 | GO:0043198 | dendritic shaft(GO:0043198) |
| 1.3 | 16.1 | GO:0005890 | sodium:potassium-exchanging ATPase complex(GO:0005890) |
| 1.3 | 4.0 | GO:0036194 | muscle cell projection(GO:0036194) muscle cell projection membrane(GO:0036195) |
| 1.2 | 7.5 | GO:0005955 | calcineurin complex(GO:0005955) |
| 1.2 | 29.6 | GO:0060077 | inhibitory synapse(GO:0060077) |
| 1.2 | 4.6 | GO:0097196 | Shu complex(GO:0097196) |
| 1.1 | 91.8 | GO:0099501 | synaptic vesicle membrane(GO:0030672) exocytic vesicle membrane(GO:0099501) |
| 1.1 | 30.5 | GO:0097546 | ciliary base(GO:0097546) |
| 1.1 | 14.6 | GO:0032433 | filopodium tip(GO:0032433) |
| 1.1 | 28.0 | GO:0008074 | guanylate cyclase complex, soluble(GO:0008074) |
| 1.1 | 3.3 | GO:0032937 | SREBP-SCAP-Insig complex(GO:0032937) |
| 1.1 | 31.0 | GO:0030867 | rough endoplasmic reticulum membrane(GO:0030867) |
| 1.1 | 157.5 | GO:0070382 | exocytic vesicle(GO:0070382) |
| 1.0 | 34.1 | GO:0051233 | spindle midzone(GO:0051233) |
| 1.0 | 1.9 | GO:1990037 | Lewy body core(GO:1990037) |
| 1.0 | 36.5 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.9 | 4.5 | GO:0033588 | Elongator holoenzyme complex(GO:0033588) |
| 0.9 | 6.3 | GO:0097524 | sperm plasma membrane(GO:0097524) |
| 0.9 | 32.3 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.8 | 3.3 | GO:0032299 | ribonuclease H2 complex(GO:0032299) |
| 0.8 | 2.5 | GO:0035867 | integrin alphav-beta8 complex(GO:0034686) alphav-beta3 integrin-IGF-1-IGF1R complex(GO:0035867) |
| 0.7 | 5.9 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.7 | 17.8 | GO:0042629 | mast cell granule(GO:0042629) |
| 0.7 | 14.4 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.6 | 5.0 | GO:0032591 | dendritic spine membrane(GO:0032591) |
| 0.6 | 14.7 | GO:0032839 | dendrite cytoplasm(GO:0032839) |
| 0.6 | 2.3 | GO:0033503 | HULC complex(GO:0033503) |
| 0.6 | 2.3 | GO:0032044 | DSIF complex(GO:0032044) |
| 0.5 | 12.3 | GO:0098827 | endoplasmic reticulum tubular network(GO:0071782) endoplasmic reticulum subcompartment(GO:0098827) |
| 0.5 | 5.9 | GO:0030128 | clathrin coat of endocytic vesicle(GO:0030128) |
| 0.5 | 44.5 | GO:0008076 | voltage-gated potassium channel complex(GO:0008076) |
| 0.5 | 20.9 | GO:0016235 | aggresome(GO:0016235) |
| 0.5 | 17.1 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.5 | 2.8 | GO:0030877 | beta-catenin destruction complex(GO:0030877) |
| 0.5 | 2.3 | GO:0000214 | tRNA-intron endonuclease complex(GO:0000214) |
| 0.4 | 1.3 | GO:0031310 | integral component of vacuolar membrane(GO:0031166) intrinsic component of vacuolar membrane(GO:0031310) |
| 0.4 | 2.1 | GO:0031315 | extrinsic component of mitochondrial outer membrane(GO:0031315) |
| 0.4 | 2.1 | GO:0030130 | clathrin coat of trans-Golgi network vesicle(GO:0030130) |
| 0.4 | 1.2 | GO:0033557 | Slx1-Slx4 complex(GO:0033557) |
| 0.4 | 56.1 | GO:0030133 | transport vesicle(GO:0030133) |
| 0.4 | 27.9 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.4 | 3.4 | GO:0017146 | NMDA selective glutamate receptor complex(GO:0017146) |
| 0.4 | 3.0 | GO:0072669 | tRNA-splicing ligase complex(GO:0072669) |
| 0.4 | 8.2 | GO:0032588 | trans-Golgi network membrane(GO:0032588) |
| 0.3 | 51.2 | GO:0045211 | postsynaptic membrane(GO:0045211) |
| 0.3 | 13.9 | GO:0097060 | synaptic membrane(GO:0097060) |
| 0.3 | 1.6 | GO:0070695 | FHF complex(GO:0070695) |
| 0.3 | 11.8 | GO:0097610 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.3 | 2.1 | GO:0000796 | condensin complex(GO:0000796) |
| 0.3 | 1.5 | GO:0000125 | PCAF complex(GO:0000125) |
| 0.3 | 24.7 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.3 | 10.5 | GO:0005801 | cis-Golgi network(GO:0005801) |
| 0.2 | 1.2 | GO:0048179 | activin receptor complex(GO:0048179) |
| 0.2 | 4.9 | GO:0002080 | acrosomal membrane(GO:0002080) |
| 0.2 | 3.7 | GO:0005751 | mitochondrial respiratory chain complex IV(GO:0005751) |
| 0.2 | 6.3 | GO:0031307 | integral component of mitochondrial outer membrane(GO:0031307) |
| 0.2 | 14.5 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.2 | 2.0 | GO:0005797 | Golgi medial cisterna(GO:0005797) |
| 0.2 | 0.8 | GO:0045098 | type III intermediate filament(GO:0045098) |
| 0.2 | 1.4 | GO:0031211 | serine C-palmitoyltransferase complex(GO:0017059) endoplasmic reticulum palmitoyltransferase complex(GO:0031211) |
| 0.2 | 7.4 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.2 | 7.4 | GO:0045271 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.1 | 5.1 | GO:0016323 | basolateral plasma membrane(GO:0016323) |
| 0.1 | 4.4 | GO:0043679 | axon terminus(GO:0043679) |
| 0.1 | 7.0 | GO:0030315 | T-tubule(GO:0030315) |
| 0.1 | 31.0 | GO:0030425 | dendrite(GO:0030425) |
| 0.1 | 6.1 | GO:0030864 | cortical actin cytoskeleton(GO:0030864) |
| 0.1 | 4.3 | GO:0030136 | clathrin-coated vesicle(GO:0030136) |
| 0.1 | 12.6 | GO:0030426 | growth cone(GO:0030426) |
| 0.1 | 19.4 | GO:0045121 | membrane raft(GO:0045121) membrane microdomain(GO:0098857) |
| 0.1 | 12.5 | GO:0030659 | cytoplasmic vesicle membrane(GO:0030659) |
| 0.1 | 5.6 | GO:0005901 | caveola(GO:0005901) |
| 0.1 | 2.1 | GO:0055038 | recycling endosome membrane(GO:0055038) |
| 0.1 | 2.0 | GO:0000780 | condensed nuclear chromosome, centromeric region(GO:0000780) |
| 0.1 | 0.9 | GO:0042587 | glycogen granule(GO:0042587) |
| 0.1 | 5.1 | GO:0005762 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.1 | 0.4 | GO:0097361 | CIA complex(GO:0097361) |
| 0.1 | 1.2 | GO:0000930 | gamma-tubulin complex(GO:0000930) |
| 0.1 | 1.3 | GO:0000145 | exocyst(GO:0000145) |
| 0.1 | 5.5 | GO:0098793 | presynapse(GO:0098793) |
| 0.1 | 1.2 | GO:0005782 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.1 | 16.1 | GO:0005840 | ribosome(GO:0005840) |
| 0.1 | 0.5 | GO:0030123 | AP-3 adaptor complex(GO:0030123) |
| 0.1 | 0.5 | GO:0000815 | ESCRT III complex(GO:0000815) |
| 0.1 | 2.6 | GO:0031234 | extrinsic component of cytoplasmic side of plasma membrane(GO:0031234) |
| 0.1 | 9.0 | GO:0098852 | lysosomal membrane(GO:0005765) lytic vacuole membrane(GO:0098852) |
| 0.1 | 36.8 | GO:0005887 | integral component of plasma membrane(GO:0005887) |
| 0.0 | 0.2 | GO:0070545 | PeBoW complex(GO:0070545) |
| 0.0 | 0.9 | GO:0045335 | phagocytic vesicle(GO:0045335) |
| 0.0 | 199.4 | GO:0016021 | integral component of membrane(GO:0016021) |
| 0.0 | 0.7 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.0 | 1.4 | GO:0005776 | autophagosome(GO:0005776) |
| 0.0 | 0.5 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.0 | 0.1 | GO:0000124 | SAGA complex(GO:0000124) |
| 0.0 | 0.3 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 19.0 | 114.0 | GO:0099529 | neurotransmitter receptor activity involved in regulation of postsynaptic membrane potential(GO:0099529) |
| 18.9 | 75.8 | GO:0099609 | microtubule lateral binding(GO:0099609) |
| 10.1 | 30.4 | GO:0004971 | AMPA glutamate receptor activity(GO:0004971) |
| 9.3 | 46.5 | GO:0005052 | peroxisome matrix targeting signal-1 binding(GO:0005052) |
| 8.5 | 59.5 | GO:0045503 | dynein light chain binding(GO:0045503) |
| 7.5 | 52.8 | GO:0031821 | G-protein coupled serotonin receptor binding(GO:0031821) |
| 6.8 | 20.3 | GO:0015018 | galactosylgalactosylxylosylprotein 3-beta-glucuronosyltransferase activity(GO:0015018) |
| 6.4 | 57.8 | GO:0086080 | protein binding involved in heterotypic cell-cell adhesion(GO:0086080) |
| 5.8 | 17.5 | GO:0031780 | corticotropin hormone receptor binding(GO:0031780) type 5 melanocortin receptor binding(GO:0031783) |
| 5.3 | 10.7 | GO:0004938 | alpha2-adrenergic receptor activity(GO:0004938) |
| 4.8 | 14.5 | GO:0031997 | N-terminal myristoylation domain binding(GO:0031997) calcium ion binding involved in regulation of postsynaptic cytosolic calcium ion concentration(GO:0099567) |
| 4.7 | 37.6 | GO:0016308 | 1-phosphatidylinositol-4-phosphate 5-kinase activity(GO:0016308) |
| 4.7 | 28.0 | GO:0008294 | calcium- and calmodulin-responsive adenylate cyclase activity(GO:0008294) |
| 4.2 | 21.2 | GO:0048403 | brain-derived neurotrophic factor binding(GO:0048403) |
| 4.1 | 16.4 | GO:0016492 | G-protein coupled neurotensin receptor activity(GO:0016492) |
| 4.1 | 56.7 | GO:0030550 | acetylcholine receptor inhibitor activity(GO:0030550) |
| 4.0 | 28.2 | GO:0008273 | calcium, potassium:sodium antiporter activity(GO:0008273) |
| 3.7 | 18.7 | GO:0032795 | heterotrimeric G-protein binding(GO:0032795) |
| 3.7 | 33.3 | GO:0046870 | cadmium ion binding(GO:0046870) |
| 3.6 | 14.3 | GO:0005163 | nerve growth factor receptor binding(GO:0005163) |
| 3.4 | 60.8 | GO:0043008 | ATP-dependent protein binding(GO:0043008) |
| 3.3 | 13.3 | GO:0004691 | cAMP-dependent protein kinase activity(GO:0004691) |
| 3.2 | 16.0 | GO:0031849 | olfactory receptor binding(GO:0031849) |
| 3.1 | 6.3 | GO:0032551 | UTP binding(GO:0002134) sulfonylurea receptor binding(GO:0017098) pyrimidine ribonucleoside binding(GO:0032551) |
| 3.1 | 28.3 | GO:0032050 | clathrin heavy chain binding(GO:0032050) |
| 3.0 | 3.0 | GO:0035651 | AP-3 adaptor complex binding(GO:0035651) |
| 2.9 | 14.7 | GO:0004962 | endothelin receptor activity(GO:0004962) |
| 2.9 | 11.5 | GO:0008499 | UDP-galactose:beta-N-acetylglucosamine beta-1,3-galactosyltransferase activity(GO:0008499) |
| 2.7 | 5.4 | GO:0098748 | clathrin adaptor activity(GO:0035615) endocytic adaptor activity(GO:0098748) |
| 2.6 | 20.7 | GO:0050291 | sphingosine N-acyltransferase activity(GO:0050291) |
| 2.5 | 7.4 | GO:0004724 | magnesium-dependent protein serine/threonine phosphatase activity(GO:0004724) |
| 2.3 | 52.2 | GO:0031489 | myosin V binding(GO:0031489) |
| 2.2 | 15.7 | GO:0097109 | neuroligin family protein binding(GO:0097109) |
| 2.2 | 17.4 | GO:0004865 | protein serine/threonine phosphatase inhibitor activity(GO:0004865) |
| 2.2 | 19.5 | GO:0019992 | diacylglycerol binding(GO:0019992) |
| 2.1 | 8.6 | GO:0034431 | endopolyphosphatase activity(GO:0000298) diphosphoinositol-polyphosphate diphosphatase activity(GO:0008486) bis(5'-adenosyl)-hexaphosphatase activity(GO:0034431) bis(5'-adenosyl)-pentaphosphatase activity(GO:0034432) |
| 2.1 | 25.7 | GO:0017162 | aryl hydrocarbon receptor binding(GO:0017162) |
| 2.1 | 44.6 | GO:0008266 | poly(U) RNA binding(GO:0008266) |
| 2.1 | 29.6 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 2.1 | 8.4 | GO:0051425 | PTB domain binding(GO:0051425) |
| 2.0 | 12.2 | GO:0015277 | kainate selective glutamate receptor activity(GO:0015277) |
| 2.0 | 26.1 | GO:0008179 | adenylate cyclase binding(GO:0008179) |
| 1.9 | 29.2 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 1.9 | 7.5 | GO:0033192 | calcium-dependent protein serine/threonine phosphatase activity(GO:0004723) calmodulin-dependent protein phosphatase activity(GO:0033192) |
| 1.8 | 7.3 | GO:0004711 | ribosomal protein S6 kinase activity(GO:0004711) |
| 1.8 | 5.4 | GO:0008142 | oxysterol binding(GO:0008142) |
| 1.8 | 10.6 | GO:0001517 | N-acetylglucosamine 6-O-sulfotransferase activity(GO:0001517) |
| 1.8 | 10.5 | GO:0008467 | [heparan sulfate]-glucosamine 3-sulfotransferase 1 activity(GO:0008467) |
| 1.7 | 6.9 | GO:0030144 | alpha-1,6-mannosylglycoprotein 6-beta-N-acetylglucosaminyltransferase activity(GO:0030144) |
| 1.7 | 38.0 | GO:0031210 | phosphatidylcholine binding(GO:0031210) |
| 1.6 | 40.8 | GO:0004890 | GABA-A receptor activity(GO:0004890) |
| 1.6 | 9.8 | GO:0008508 | bile acid:sodium symporter activity(GO:0008508) |
| 1.6 | 53.9 | GO:0004683 | calmodulin-dependent protein kinase activity(GO:0004683) |
| 1.5 | 15.1 | GO:0009922 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 1.5 | 14.7 | GO:0015193 | L-proline transmembrane transporter activity(GO:0015193) |
| 1.5 | 92.1 | GO:0005544 | calcium-dependent phospholipid binding(GO:0005544) |
| 1.4 | 18.3 | GO:0004935 | adrenergic receptor activity(GO:0004935) |
| 1.4 | 37.5 | GO:0017091 | AU-rich element binding(GO:0017091) |
| 1.4 | 4.1 | GO:0016312 | inositol bisphosphate phosphatase activity(GO:0016312) |
| 1.4 | 4.1 | GO:0008607 | phosphorylase kinase regulator activity(GO:0008607) |
| 1.3 | 10.8 | GO:0005049 | nuclear export signal receptor activity(GO:0005049) |
| 1.3 | 16.1 | GO:0030955 | potassium ion binding(GO:0030955) |
| 1.3 | 46.9 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 1.3 | 38.9 | GO:0017134 | fibroblast growth factor binding(GO:0017134) |
| 1.3 | 9.0 | GO:0051998 | carboxyl-O-methyltransferase activity(GO:0010340) protein carboxyl O-methyltransferase activity(GO:0051998) |
| 1.3 | 5.1 | GO:0008597 | calcium-dependent protein serine/threonine phosphatase regulator activity(GO:0008597) |
| 1.3 | 5.0 | GO:0031749 | D2 dopamine receptor binding(GO:0031749) |
| 1.2 | 3.7 | GO:0016453 | C-acetyltransferase activity(GO:0016453) |
| 1.2 | 6.2 | GO:0030229 | very-low-density lipoprotein particle receptor activity(GO:0030229) |
| 1.2 | 6.1 | GO:0005315 | inorganic phosphate transmembrane transporter activity(GO:0005315) |
| 1.2 | 23.1 | GO:0005545 | 1-phosphatidylinositol binding(GO:0005545) |
| 1.2 | 55.2 | GO:0042169 | SH2 domain binding(GO:0042169) |
| 1.2 | 15.4 | GO:0008649 | rRNA methyltransferase activity(GO:0008649) |
| 1.2 | 4.7 | GO:0047623 | AMP deaminase activity(GO:0003876) adenosine-phosphate deaminase activity(GO:0047623) |
| 1.2 | 41.5 | GO:0001106 | RNA polymerase II transcription corepressor activity(GO:0001106) |
| 1.2 | 3.5 | GO:0016403 | dimethylargininase activity(GO:0016403) |
| 1.2 | 3.5 | GO:0015222 | serotonin transmembrane transporter activity(GO:0015222) |
| 1.1 | 6.9 | GO:0030235 | nitric-oxide synthase regulator activity(GO:0030235) |
| 1.1 | 11.2 | GO:0038036 | sphingosine-1-phosphate receptor activity(GO:0038036) |
| 1.1 | 9.5 | GO:0015217 | ATP transmembrane transporter activity(GO:0005347) ADP transmembrane transporter activity(GO:0015217) |
| 1.0 | 11.3 | GO:0035612 | AP-2 adaptor complex binding(GO:0035612) |
| 1.0 | 9.1 | GO:0015280 | ligand-gated sodium channel activity(GO:0015280) |
| 1.0 | 3.0 | GO:0004020 | adenylylsulfate kinase activity(GO:0004020) sulfate adenylyltransferase activity(GO:0004779) sulfate adenylyltransferase (ATP) activity(GO:0004781) |
| 1.0 | 28.5 | GO:0017147 | Wnt-protein binding(GO:0017147) |
| 0.9 | 34.7 | GO:0001784 | phosphotyrosine binding(GO:0001784) |
| 0.9 | 19.6 | GO:0015278 | calcium-release channel activity(GO:0015278) ligand-gated calcium channel activity(GO:0099604) |
| 0.9 | 11.2 | GO:0016868 | intramolecular transferase activity, phosphotransferases(GO:0016868) |
| 0.9 | 38.6 | GO:0045182 | translation regulator activity(GO:0045182) |
| 0.9 | 21.1 | GO:0070006 | metalloaminopeptidase activity(GO:0070006) |
| 0.9 | 6.4 | GO:0030160 | GKAP/Homer scaffold activity(GO:0030160) |
| 0.9 | 15.2 | GO:0008195 | phosphatidate phosphatase activity(GO:0008195) |
| 0.9 | 3.5 | GO:0051731 | polynucleotide 5'-hydroxyl-kinase activity(GO:0051731) |
| 0.9 | 3.5 | GO:0004897 | ciliary neurotrophic factor receptor activity(GO:0004897) |
| 0.9 | 4.3 | GO:0010997 | anaphase-promoting complex binding(GO:0010997) |
| 0.8 | 3.4 | GO:0050659 | chondroitin sulfotransferase activity(GO:0034481) chondroitin 4-sulfotransferase activity(GO:0047756) N-acetylgalactosamine 4-sulfate 6-O-sulfotransferase activity(GO:0050659) |
| 0.8 | 17.4 | GO:0005242 | inward rectifier potassium channel activity(GO:0005242) |
| 0.8 | 3.3 | GO:0086077 | gap junction channel activity involved in cardiac conduction electrical coupling(GO:0086075) gap junction channel activity involved in AV node cell-bundle of His cell electrical coupling(GO:0086077) |
| 0.8 | 2.4 | GO:0016900 | oxidoreductase activity, acting on the CH-OH group of donors, disulfide as acceptor(GO:0016900) vitamin-K-epoxide reductase (warfarin-sensitive) activity(GO:0047057) |
| 0.8 | 35.5 | GO:0030507 | spectrin binding(GO:0030507) |
| 0.8 | 11.0 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.8 | 13.2 | GO:0048018 | receptor agonist activity(GO:0048018) |
| 0.7 | 27.4 | GO:0005158 | insulin receptor binding(GO:0005158) |
| 0.7 | 21.6 | GO:0046875 | ephrin receptor binding(GO:0046875) |
| 0.7 | 6.3 | GO:0001727 | lipid kinase activity(GO:0001727) |
| 0.7 | 11.8 | GO:0071617 | lysophospholipid acyltransferase activity(GO:0071617) |
| 0.7 | 4.2 | GO:0004826 | phenylalanine-tRNA ligase activity(GO:0004826) |
| 0.7 | 2.0 | GO:0001075 | transcription factor activity, RNA polymerase II core promoter sequence-specific binding involved in preinitiation complex assembly(GO:0001075) |
| 0.7 | 21.7 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.7 | 3.3 | GO:0004523 | RNA-DNA hybrid ribonuclease activity(GO:0004523) |
| 0.7 | 2.0 | GO:0003696 | satellite DNA binding(GO:0003696) |
| 0.6 | 22.8 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.6 | 13.9 | GO:0052866 | phosphatidylinositol phosphate phosphatase activity(GO:0052866) |
| 0.6 | 19.4 | GO:0032266 | phosphatidylinositol-3-phosphate binding(GO:0032266) |
| 0.6 | 8.9 | GO:0005092 | GDP-dissociation inhibitor activity(GO:0005092) |
| 0.6 | 28.1 | GO:0017022 | myosin binding(GO:0017022) |
| 0.6 | 9.1 | GO:0070300 | phosphatidic acid binding(GO:0070300) |
| 0.6 | 18.5 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.5 | 22.0 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.5 | 8.9 | GO:0008253 | 5'-nucleotidase activity(GO:0008253) |
| 0.5 | 28.2 | GO:0004722 | protein serine/threonine phosphatase activity(GO:0004722) |
| 0.5 | 14.2 | GO:0004693 | cyclin-dependent protein serine/threonine kinase activity(GO:0004693) |
| 0.5 | 4.0 | GO:0015556 | C4-dicarboxylate transmembrane transporter activity(GO:0015556) |
| 0.5 | 65.8 | GO:0005267 | potassium channel activity(GO:0005267) |
| 0.5 | 1.9 | GO:0004427 | inorganic diphosphatase activity(GO:0004427) |
| 0.5 | 2.3 | GO:0000213 | tRNA-intron endonuclease activity(GO:0000213) |
| 0.5 | 16.1 | GO:0015485 | cholesterol binding(GO:0015485) |
| 0.5 | 1.8 | GO:0008449 | N-acetylglucosamine-6-sulfatase activity(GO:0008449) |
| 0.4 | 1.3 | GO:0004735 | pyrroline-5-carboxylate reductase activity(GO:0004735) |
| 0.4 | 15.0 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.4 | 8.3 | GO:0016891 | endoribonuclease activity, producing 5'-phosphomonoesters(GO:0016891) |
| 0.4 | 17.8 | GO:0016620 | oxidoreductase activity, acting on the aldehyde or oxo group of donors, NAD or NADP as acceptor(GO:0016620) |
| 0.4 | 6.5 | GO:0002162 | dystroglycan binding(GO:0002162) |
| 0.4 | 5.7 | GO:0004115 | 3',5'-cyclic-AMP phosphodiesterase activity(GO:0004115) |
| 0.4 | 42.4 | GO:0016410 | N-acyltransferase activity(GO:0016410) |
| 0.4 | 15.4 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.4 | 1.4 | GO:0008379 | thioredoxin peroxidase activity(GO:0008379) |
| 0.3 | 9.4 | GO:0050136 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.3 | 1.0 | GO:1902379 | chemoattractant activity involved in axon guidance(GO:1902379) |
| 0.3 | 1.3 | GO:0045134 | uridine-diphosphatase activity(GO:0045134) |
| 0.3 | 1.9 | GO:0043426 | MRF binding(GO:0043426) |
| 0.3 | 0.9 | GO:0003976 | UDP-N-acetylglucosamine-lysosomal-enzyme N-acetylglucosaminephosphotransferase activity(GO:0003976) |
| 0.3 | 2.8 | GO:0016929 | SUMO-specific protease activity(GO:0016929) |
| 0.3 | 5.0 | GO:0005388 | calcium-transporting ATPase activity(GO:0005388) |
| 0.3 | 1.4 | GO:0016454 | serine C-palmitoyltransferase activity(GO:0004758) C-palmitoyltransferase activity(GO:0016454) |
| 0.2 | 4.9 | GO:0005537 | mannose binding(GO:0005537) |
| 0.2 | 8.8 | GO:0005484 | SNAP receptor activity(GO:0005484) |
| 0.2 | 4.8 | GO:0005031 | tumor necrosis factor-activated receptor activity(GO:0005031) death receptor activity(GO:0005035) |
| 0.2 | 2.8 | GO:0003747 | translation release factor activity(GO:0003747) translation termination factor activity(GO:0008079) |
| 0.2 | 1.4 | GO:0004473 | malic enzyme activity(GO:0004470) malate dehydrogenase (decarboxylating) (NAD+) activity(GO:0004471) malate dehydrogenase (decarboxylating) (NADP+) activity(GO:0004473) |
| 0.2 | 2.2 | GO:0004551 | nucleotide diphosphatase activity(GO:0004551) |
| 0.2 | 4.2 | GO:0004559 | alpha-mannosidase activity(GO:0004559) mannosidase activity(GO:0015923) |
| 0.2 | 27.6 | GO:0017137 | Rab GTPase binding(GO:0017137) |
| 0.2 | 1.2 | GO:0016361 | activin receptor activity, type I(GO:0016361) |
| 0.2 | 1.2 | GO:0017108 | 5'-flap endonuclease activity(GO:0017108) |
| 0.2 | 3.3 | GO:0016863 | intramolecular oxidoreductase activity, transposing C=C bonds(GO:0016863) |
| 0.2 | 2.0 | GO:0070097 | delta-catenin binding(GO:0070097) |
| 0.2 | 13.9 | GO:0005089 | Rho guanyl-nucleotide exchange factor activity(GO:0005089) |
| 0.2 | 0.7 | GO:0000995 | transcription factor activity, core RNA polymerase III binding(GO:0000995) |
| 0.2 | 3.8 | GO:0042056 | chemoattractant activity(GO:0042056) |
| 0.2 | 4.1 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.2 | 0.6 | GO:0008318 | protein prenyltransferase activity(GO:0008318) |
| 0.2 | 0.5 | GO:0044715 | 8-oxo-dGDP phosphatase activity(GO:0044715) |
| 0.1 | 0.9 | GO:0048039 | ubiquinone binding(GO:0048039) |
| 0.1 | 2.3 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) |
| 0.1 | 0.7 | GO:0004849 | uridine kinase activity(GO:0004849) |
| 0.1 | 3.5 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.1 | 10.0 | GO:0051117 | ATPase binding(GO:0051117) |
| 0.1 | 1.6 | GO:0043325 | phosphatidylinositol-3,4-bisphosphate binding(GO:0043325) |
| 0.1 | 1.6 | GO:0030552 | cAMP binding(GO:0030552) |
| 0.1 | 5.2 | GO:0035254 | glutamate receptor binding(GO:0035254) |
| 0.1 | 3.0 | GO:0004190 | aspartic-type endopeptidase activity(GO:0004190) |
| 0.1 | 0.9 | GO:0005173 | stem cell factor receptor binding(GO:0005173) |
| 0.1 | 2.4 | GO:0015106 | bicarbonate transmembrane transporter activity(GO:0015106) |
| 0.1 | 0.6 | GO:0042289 | MHC class II protein binding(GO:0042289) |
| 0.1 | 3.7 | GO:0015002 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.1 | 2.6 | GO:0030332 | cyclin binding(GO:0030332) |
| 0.1 | 5.0 | GO:0042605 | peptide antigen binding(GO:0042605) |
| 0.1 | 3.7 | GO:0071855 | neuropeptide receptor binding(GO:0071855) |
| 0.1 | 17.3 | GO:0005096 | GTPase activator activity(GO:0005096) |
| 0.1 | 0.9 | GO:0032794 | GTPase activating protein binding(GO:0032794) |
| 0.1 | 12.8 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.1 | 8.8 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.1 | 2.5 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.1 | 0.6 | GO:0044388 | small protein activating enzyme binding(GO:0044388) |
| 0.1 | 13.1 | GO:0016746 | transferase activity, transferring acyl groups(GO:0016746) |
| 0.1 | 5.7 | GO:0008081 | phosphoric diester hydrolase activity(GO:0008081) |
| 0.1 | 1.2 | GO:0034237 | protein kinase A regulatory subunit binding(GO:0034237) |
| 0.1 | 17.5 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.1 | 1.4 | GO:0097602 | cullin family protein binding(GO:0097602) |
| 0.1 | 0.5 | GO:0086059 | voltage-gated calcium channel activity involved SA node cell action potential(GO:0086059) |
| 0.1 | 0.2 | GO:0008476 | protein-tyrosine sulfotransferase activity(GO:0008476) |
| 0.1 | 1.3 | GO:0017160 | Ral GTPase binding(GO:0017160) |
| 0.1 | 0.9 | GO:0050750 | low-density lipoprotein particle receptor binding(GO:0050750) |
| 0.1 | 1.3 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.1 | 2.7 | GO:0008188 | neuropeptide receptor activity(GO:0008188) |
| 0.1 | 6.3 | GO:0042626 | ATPase activity, coupled to transmembrane movement of substances(GO:0042626) |
| 0.1 | 0.6 | GO:0015232 | heme transporter activity(GO:0015232) |
| 0.1 | 0.3 | GO:0030346 | protein phosphatase 2B binding(GO:0030346) |
| 0.1 | 2.8 | GO:0032947 | protein complex scaffold(GO:0032947) |
| 0.0 | 22.8 | GO:0005509 | calcium ion binding(GO:0005509) |
| 0.0 | 1.5 | GO:0019894 | kinesin binding(GO:0019894) |
| 0.0 | 0.7 | GO:0016594 | glycine binding(GO:0016594) |
| 0.0 | 1.3 | GO:0030544 | Hsp70 protein binding(GO:0030544) |
| 0.0 | 0.2 | GO:0010385 | double-stranded methylated DNA binding(GO:0010385) |
| 0.0 | 8.9 | GO:0008233 | peptidase activity(GO:0008233) |
| 0.0 | 0.4 | GO:0005522 | profilin binding(GO:0005522) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.8 | 75.8 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 2.2 | 8.8 | SA CASPASE CASCADE | Apoptosis is mediated by caspases, cysteine proteases arranged in a proteolytic cascade. |
| 2.1 | 32.2 | PID S1P S1P4 PATHWAY | S1P4 pathway |
| 2.1 | 44.2 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 1.4 | 5.6 | ST PAC1 RECEPTOR PATHWAY | PAC1 Receptor Pathway |
| 1.3 | 33.5 | ST WNT CA2 CYCLIC GMP PATHWAY | Wnt/Ca2+/cyclic GMP signaling. |
| 1.0 | 32.5 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.9 | 33.7 | PID REELIN PATHWAY | Reelin signaling pathway |
| 0.8 | 3.1 | SIG PIP3 SIGNALING IN B LYMPHOCYTES | Genes related to PIP3 signaling in B lymphocytes |
| 0.7 | 15.4 | PID WNT CANONICAL PATHWAY | Canonical Wnt signaling pathway |
| 0.7 | 6.3 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.6 | 30.0 | PID P75 NTR PATHWAY | p75(NTR)-mediated signaling |
| 0.6 | 17.1 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
| 0.5 | 11.4 | PID RAC1 REG PATHWAY | Regulation of RAC1 activity |
| 0.5 | 13.8 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.5 | 14.9 | PID EPHA FWDPATHWAY | EPHA forward signaling |
| 0.5 | 28.3 | PID TRKR PATHWAY | Neurotrophic factor-mediated Trk receptor signaling |
| 0.5 | 30.4 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.5 | 1.4 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.5 | 19.4 | PID ERBB1 INTERNALIZATION PATHWAY | Internalization of ErbB1 |
| 0.4 | 3.1 | PID PDGFRA PATHWAY | PDGFR-alpha signaling pathway |
| 0.4 | 12.9 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
| 0.4 | 12.7 | PID ENDOTHELIN PATHWAY | Endothelins |
| 0.4 | 7.6 | PID RET PATHWAY | Signaling events regulated by Ret tyrosine kinase |
| 0.4 | 32.7 | WNT SIGNALING | Genes related to Wnt-mediated signal transduction |
| 0.3 | 6.2 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.3 | 7.3 | ST ERK1 ERK2 MAPK PATHWAY | ERK1/ERK2 MAPK Pathway |
| 0.3 | 4.1 | PID PRL SIGNALING EVENTS PATHWAY | Signaling events mediated by PRL |
| 0.2 | 7.8 | PID P38 ALPHA BETA DOWNSTREAM PATHWAY | Signaling mediated by p38-alpha and p38-beta |
| 0.2 | 1.3 | SA PTEN PATHWAY | PTEN is a tumor suppressor that dephosphorylates the lipid messenger phosphatidylinositol triphosphate. |
| 0.2 | 3.8 | PID ERBB2 ERBB3 PATHWAY | ErbB2/ErbB3 signaling events |
| 0.2 | 5.4 | PID SYNDECAN 4 PATHWAY | Syndecan-4-mediated signaling events |
| 0.2 | 44.9 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.2 | 3.0 | SIG CD40PATHWAYMAP | Genes related to CD40 signaling |
| 0.2 | 2.0 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 0.1 | 1.0 | ST DIFFERENTIATION PATHWAY IN PC12 CELLS | Differentiation Pathway in PC12 Cells; this is a specific case of PAC1 Receptor Pathway. |
| 0.1 | 2.0 | PID TGFBR PATHWAY | TGF-beta receptor signaling |
| 0.1 | 1.2 | PID P73PATHWAY | p73 transcription factor network |
| 0.1 | 0.6 | PID RANBP2 PATHWAY | Sumoylation by RanBP2 regulates transcriptional repression |
| 0.0 | 1.4 | PID ECADHERIN NASCENT AJ PATHWAY | E-cadherin signaling in the nascent adherens junction |
| 0.0 | 1.5 | PID ATF2 PATHWAY | ATF-2 transcription factor network |
| 0.0 | 1.3 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.0 | 1.2 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.0 | 0.3 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 9.7 | 173.9 | REACTOME UNBLOCKING OF NMDA RECEPTOR GLUTAMATE BINDING AND ACTIVATION | Genes involved in Unblocking of NMDA receptor, glutamate binding and activation |
| 4.8 | 66.5 | REACTOME NOREPINEPHRINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Norepinephrine Neurotransmitter Release Cycle |
| 3.5 | 49.3 | REACTOME TRAFFICKING OF GLUR2 CONTAINING AMPA RECEPTORS | Genes involved in Trafficking of GluR2-containing AMPA receptors |
| 3.2 | 93.8 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 2.9 | 14.5 | REACTOME CREB PHOSPHORYLATION THROUGH THE ACTIVATION OF CAMKII | Genes involved in CREB phosphorylation through the activation of CaMKII |
| 2.5 | 50.4 | REACTOME ACTIVATION OF NMDA RECEPTOR UPON GLUTAMATE BINDING AND POSTSYNAPTIC EVENTS | Genes involved in Activation of NMDA receptor upon glutamate binding and postsynaptic events |
| 2.4 | 31.7 | REACTOME TANDEM PORE DOMAIN POTASSIUM CHANNELS | Genes involved in Tandem pore domain potassium channels |
| 2.4 | 40.8 | REACTOME GABA A RECEPTOR ACTIVATION | Genes involved in GABA A receptor activation |
| 2.2 | 32.9 | REACTOME GABA SYNTHESIS RELEASE REUPTAKE AND DEGRADATION | Genes involved in GABA synthesis, release, reuptake and degradation |
| 2.1 | 6.2 | REACTOME PLATELET SENSITIZATION BY LDL | Genes involved in Platelet sensitization by LDL |
| 2.0 | 26.1 | REACTOME HIGHLY CALCIUM PERMEABLE POSTSYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | Genes involved in Highly calcium permeable postsynaptic nicotinic acetylcholine receptors |
| 1.7 | 26.2 | REACTOME RAP1 SIGNALLING | Genes involved in Rap1 signalling |
| 1.6 | 24.5 | REACTOME SIGNAL ATTENUATION | Genes involved in Signal attenuation |
| 1.6 | 47.3 | REACTOME G PROTEIN ACTIVATION | Genes involved in G-protein activation |
| 1.5 | 13.7 | REACTOME MEMBRANE BINDING AND TARGETTING OF GAG PROTEINS | Genes involved in Membrane binding and targetting of GAG proteins |
| 1.5 | 33.2 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 1.1 | 11.8 | REACTOME SYNTHESIS OF PIPS AT THE LATE ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the late endosome membrane |
| 1.0 | 26.0 | REACTOME ACYL CHAIN REMODELLING OF PE | Genes involved in Acyl chain remodelling of PE |
| 1.0 | 29.2 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.9 | 12.2 | REACTOME IONOTROPIC ACTIVITY OF KAINATE RECEPTORS | Genes involved in Ionotropic activity of Kainate Receptors |
| 0.9 | 14.0 | REACTOME INHIBITION OF VOLTAGE GATED CA2 CHANNELS VIA GBETA GAMMA SUBUNITS | Genes involved in Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits |
| 0.9 | 5.6 | REACTOME ADP SIGNALLING THROUGH P2RY1 | Genes involved in ADP signalling through P2Y purinoceptor 1 |
| 0.9 | 26.0 | REACTOME BASIGIN INTERACTIONS | Genes involved in Basigin interactions |
| 0.9 | 13.5 | REACTOME RECYCLING PATHWAY OF L1 | Genes involved in Recycling pathway of L1 |
| 0.8 | 6.3 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |
| 0.8 | 3.0 | REACTOME SYNTHESIS OF PIPS AT THE EARLY ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the early endosome membrane |
| 0.7 | 6.0 | REACTOME NEGATIVE REGULATION OF THE PI3K AKT NETWORK | Genes involved in Negative regulation of the PI3K/AKT network |
| 0.7 | 19.9 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.7 | 22.2 | REACTOME SYNTHESIS OF PIPS AT THE PLASMA MEMBRANE | Genes involved in Synthesis of PIPs at the plasma membrane |
| 0.7 | 9.2 | REACTOME PYRIMIDINE CATABOLISM | Genes involved in Pyrimidine catabolism |
| 0.7 | 8.4 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 0.6 | 17.2 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.6 | 10.6 | REACTOME NUCLEOTIDE LIKE PURINERGIC RECEPTORS | Genes involved in Nucleotide-like (purinergic) receptors |
| 0.6 | 16.9 | REACTOME A TETRASACCHARIDE LINKER SEQUENCE IS REQUIRED FOR GAG SYNTHESIS | Genes involved in A tetrasaccharide linker sequence is required for GAG synthesis |
| 0.6 | 6.1 | REACTOME ALPHA LINOLENIC ACID ALA METABOLISM | Genes involved in alpha-linolenic acid (ALA) metabolism |
| 0.6 | 16.6 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.6 | 30.5 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.6 | 10.5 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
| 0.5 | 13.7 | REACTOME POTASSIUM CHANNELS | Genes involved in Potassium Channels |
| 0.5 | 7.0 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.5 | 8.5 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.5 | 10.4 | REACTOME SIGNALING BY NODAL | Genes involved in Signaling by NODAL |
| 0.5 | 7.4 | REACTOME REGULATION OF PYRUVATE DEHYDROGENASE PDH COMPLEX | Genes involved in Regulation of pyruvate dehydrogenase (PDH) complex |
| 0.5 | 8.3 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.4 | 12.8 | REACTOME KERATAN SULFATE BIOSYNTHESIS | Genes involved in Keratan sulfate biosynthesis |
| 0.3 | 27.0 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.3 | 8.8 | REACTOME FRS2 MEDIATED CASCADE | Genes involved in FRS2-mediated cascade |
| 0.3 | 4.6 | REACTOME PROTEOLYTIC CLEAVAGE OF SNARE COMPLEX PROTEINS | Genes involved in Proteolytic cleavage of SNARE complex proteins |
| 0.3 | 4.7 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.3 | 45.7 | REACTOME G ALPHA Q SIGNALLING EVENTS | Genes involved in G alpha (q) signalling events |
| 0.3 | 3.7 | REACTOME ABORTIVE ELONGATION OF HIV1 TRANSCRIPT IN THE ABSENCE OF TAT | Genes involved in Abortive elongation of HIV-1 transcript in the absence of Tat |
| 0.3 | 2.6 | REACTOME N GLYCAN TRIMMING IN THE ER AND CALNEXIN CALRETICULIN CYCLE | Genes involved in N-glycan trimming in the ER and Calnexin/Calreticulin cycle |
| 0.2 | 4.5 | REACTOME POST CHAPERONIN TUBULIN FOLDING PATHWAY | Genes involved in Post-chaperonin tubulin folding pathway |
| 0.2 | 1.4 | REACTOME SEMA3A PAK DEPENDENT AXON REPULSION | Genes involved in Sema3A PAK dependent Axon repulsion |
| 0.2 | 0.9 | REACTOME REGULATION OF KIT SIGNALING | Genes involved in Regulation of KIT signaling |
| 0.2 | 4.2 | REACTOME ACTIVATED NOTCH1 TRANSMITS SIGNAL TO THE NUCLEUS | Genes involved in Activated NOTCH1 Transmits Signal to the Nucleus |
| 0.2 | 7.3 | REACTOME TRANSPORT OF VITAMINS NUCLEOSIDES AND RELATED MOLECULES | Genes involved in Transport of vitamins, nucleosides, and related molecules |
| 0.2 | 3.4 | REACTOME CHONDROITIN SULFATE BIOSYNTHESIS | Genes involved in Chondroitin sulfate biosynthesis |
| 0.2 | 13.3 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.2 | 3.4 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.2 | 3.9 | REACTOME MITOCHONDRIAL TRNA AMINOACYLATION | Genes involved in Mitochondrial tRNA aminoacylation |
| 0.2 | 8.9 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.2 | 2.5 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.2 | 14.5 | REACTOME P75 NTR RECEPTOR MEDIATED SIGNALLING | Genes involved in p75 NTR receptor-mediated signalling |
| 0.1 | 1.7 | REACTOME BMAL1 CLOCK NPAS2 ACTIVATES CIRCADIAN EXPRESSION | Genes involved in BMAL1:CLOCK/NPAS2 Activates Circadian Expression |
| 0.1 | 1.2 | REACTOME RECRUITMENT OF NUMA TO MITOTIC CENTROSOMES | Genes involved in Recruitment of NuMA to mitotic centrosomes |
| 0.1 | 0.7 | REACTOME OPSINS | Genes involved in Opsins |
| 0.1 | 1.6 | REACTOME DCC MEDIATED ATTRACTIVE SIGNALING | Genes involved in DCC mediated attractive signaling |
| 0.1 | 1.5 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.1 | 2.0 | REACTOME INTERACTIONS OF VPR WITH HOST CELLULAR PROTEINS | Genes involved in Interactions of Vpr with host cellular proteins |
| 0.1 | 1.3 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.1 | 0.7 | REACTOME PYRIMIDINE METABOLISM | Genes involved in Pyrimidine metabolism |
| 0.0 | 0.4 | REACTOME ACTIVATION OF IRF3 IRF7 MEDIATED BY TBK1 IKK EPSILON | Genes involved in Activation of IRF3/IRF7 mediated by TBK1/IKK epsilon |
| 0.0 | 2.5 | REACTOME ANTIGEN PROCESSING CROSS PRESENTATION | Genes involved in Antigen processing-Cross presentation |
| 0.0 | 0.5 | REACTOME ENDOSOMAL SORTING COMPLEX REQUIRED FOR TRANSPORT ESCRT | Genes involved in Endosomal Sorting Complex Required For Transport (ESCRT) |
| 0.0 | 2.8 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 0.5 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.0 | 0.4 | REACTOME GENERATION OF SECOND MESSENGER MOLECULES | Genes involved in Generation of second messenger molecules |
| 0.0 | 1.1 | REACTOME G ALPHA S SIGNALLING EVENTS | Genes involved in G alpha (s) signalling events |
| 0.0 | 1.6 | REACTOME NONSENSE MEDIATED DECAY ENHANCED BY THE EXON JUNCTION COMPLEX | Genes involved in Nonsense Mediated Decay Enhanced by the Exon Junction Complex |
| 0.0 | 0.6 | REACTOME IRON UPTAKE AND TRANSPORT | Genes involved in Iron uptake and transport |