Project
GSE53960: rat RNA-Seq transcriptomic Bodymap
Navigation
Downloads
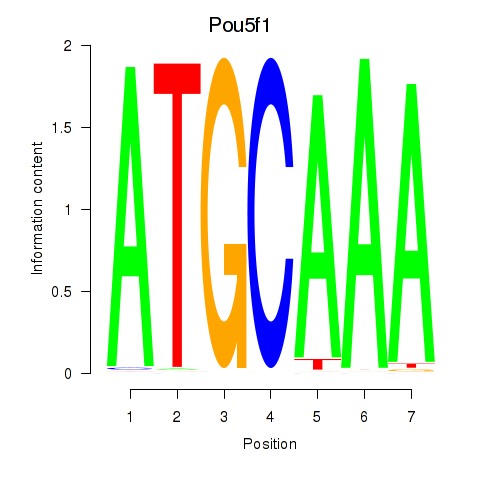
Results for Pou5f1
Z-value: 4.67
Transcription factors associated with Pou5f1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Pou5f1
|
ENSRNOG00000046487 | POU class 5 homeobox 1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Pou5f1 | rn6_v1_chr20_-_3751994_3751994 | 0.07 | 2.3e-01 | Click! |
Activity profile of Pou5f1 motif
Sorted Z-values of Pou5f1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr4_+_98370797 | 270.58 |
ENSRNOT00000031991
|
AABR07060872.1
|
|
| chr10_-_94500591 | 235.84 |
ENSRNOT00000015976
|
Cd79b
|
CD79b molecule |
| chr11_+_85532526 | 229.38 |
ENSRNOT00000036565
|
AABR07034730.2
|
|
| chr4_+_102489916 | 225.54 |
ENSRNOT00000082031
|
AABR07061001.1
|
|
| chr3_+_16413080 | 224.36 |
ENSRNOT00000040386
|
LOC100912707
|
Ig kappa chain V19-17-like |
| chr3_+_17546566 | 218.75 |
ENSRNOT00000050825
|
AABR07051583.1
|
|
| chr3_+_18315320 | 217.00 |
ENSRNOT00000006954
|
AABR07051626.2
|
|
| chr6_-_138662365 | 216.80 |
ENSRNOT00000066209
ENSRNOT00000084892 |
Ighm
|
immunoglobulin heavy constant mu |
| chr3_-_16537433 | 215.43 |
ENSRNOT00000048523
|
AABR07051533.2
|
|
| chr3_+_18706988 | 212.95 |
ENSRNOT00000074650
|
AABR07051652.1
|
|
| chr6_-_140102325 | 205.26 |
ENSRNOT00000072238
|
AABR07065750.2
|
|
| chr17_-_31780120 | 203.18 |
ENSRNOT00000058388
|
AABR07027450.1
|
|
| chr3_+_16846412 | 196.08 |
ENSRNOT00000074266
|
AABR07051551.1
|
|
| chr11_+_85508300 | 194.76 |
ENSRNOT00000038646
|
AABR07034730.3
|
|
| chr4_-_103258134 | 194.30 |
ENSRNOT00000086827
|
AABR07061052.1
|
|
| chr3_-_17081510 | 193.60 |
ENSRNOT00000063862
|
AABR07051562.1
|
|
| chrX_+_96991658 | 192.59 |
ENSRNOT00000049969
|
AABR07040288.1
|
|
| chr4_+_106323089 | 192.41 |
ENSRNOT00000091402
|
AABR07061134.1
|
|
| chr4_+_98481520 | 192.09 |
ENSRNOT00000078381
ENSRNOT00000048493 |
AABR07060886.1
|
|
| chr6_-_138632159 | 187.49 |
ENSRNOT00000082921
ENSRNOT00000040702 |
Ighm
|
immunoglobulin heavy constant mu |
| chr6_+_139158334 | 183.11 |
ENSRNOT00000089227
|
AABR07065673.1
|
|
| chr6_-_138744480 | 181.50 |
ENSRNOT00000089387
|
AABR07065651.5
|
|
| chr3_+_17107861 | 179.71 |
ENSRNOT00000043097
|
AABR07051563.1
|
|
| chr4_+_102351036 | 177.89 |
ENSRNOT00000079277
|
AABR07060994.1
|
|
| chr6_+_139551751 | 176.87 |
ENSRNOT00000081684
|
AABR07065699.2
|
|
| chr3_-_20419417 | 175.56 |
ENSRNOT00000077772
|
AABR07051733.3
|
|
| chr3_+_20375699 | 175.03 |
ENSRNOT00000088492
|
AABR07051731.1
|
|
| chr6_-_141147264 | 174.72 |
ENSRNOT00000042900
|
LOC100361105
|
Igh protein-like |
| chr6_+_139405966 | 174.69 |
ENSRNOT00000088974
|
AABR07065693.3
|
|
| chr3_+_16610086 | 171.71 |
ENSRNOT00000046231
|
LOC100361009
|
rCG64257-like |
| chr4_+_93791054 | 170.89 |
ENSRNOT00000042300
|
AABR07060788.1
|
|
| chr3_+_17889972 | 170.49 |
ENSRNOT00000073021
|
AABR07051611.1
|
|
| chr6_-_140880070 | 169.60 |
ENSRNOT00000073779
|
LOC691828
|
uncharacterized LOC691828 |
| chr6_-_139102378 | 169.38 |
ENSRNOT00000086423
|
AABR07065656.5
|
|
| chr6_-_141321108 | 158.47 |
ENSRNOT00000040556
|
AABR07065789.3
|
|
| chr11_+_85618714 | 157.82 |
ENSRNOT00000074614
|
AC109901.1
|
|
| chr2_+_223121410 | 157.12 |
ENSRNOT00000087559
|
AABR07013116.1
|
|
| chr3_-_19320915 | 155.74 |
ENSRNOT00000043673
|
RGD1565617
|
similar to Ig variable region, light chain |
| chr19_+_3325893 | 155.45 |
ENSRNOT00000048879
|
RGD1565617
|
similar to Ig variable region, light chain |
| chr1_-_17378047 | 154.59 |
ENSRNOT00000020102
|
Themis
|
thymocyte selection associated |
| chr6_+_139428999 | 153.21 |
ENSRNOT00000084482
|
AABR07065693.2
|
|
| chr6_-_138565245 | 153.04 |
ENSRNOT00000070980
|
AABR07065645.2
|
|
| chr11_+_85561460 | 152.84 |
ENSRNOT00000075455
|
AABR07072262.1
|
|
| chr4_+_102665529 | 152.59 |
ENSRNOT00000082333
|
AABR07061005.1
|
|
| chr6_-_140642221 | 150.84 |
ENSRNOT00000081996
|
AABR07065772.2
|
|
| chr3_-_20479999 | 150.65 |
ENSRNOT00000050573
|
AABR07051733.2
|
|
| chr17_+_43627930 | 147.68 |
ENSRNOT00000081719
|
LOC102549061
|
histone H2B type 1-N-like |
| chr6_-_142353308 | 145.26 |
ENSRNOT00000066416
|
AABR07065814.2
|
|
| chr6_-_141062581 | 144.90 |
ENSRNOT00000073446
|
AABR07065778.3
|
|
| chr6_+_139560028 | 144.70 |
ENSRNOT00000072633
|
LOC100360581
|
rCG58847-like |
| chr6_-_138736203 | 143.38 |
ENSRNOT00000052021
|
LOC100360169
|
rCG21044-like |
| chr1_+_192233910 | 142.62 |
ENSRNOT00000016418
ENSRNOT00000016442 |
Prkcb
|
protein kinase C, beta |
| chr4_+_102147211 | 140.69 |
ENSRNOT00000083239
|
AABR07060980.1
|
|
| chr6_+_139345486 | 140.51 |
ENSRNOT00000081540
|
AABR07065688.1
|
|
| chr6_-_139911839 | 140.03 |
ENSRNOT00000077113
ENSRNOT00000084547 |
AABR07065714.1
|
|
| chr3_+_17180411 | 138.44 |
ENSRNOT00000058260
|
AABR07051565.1
|
|
| chr3_+_16817051 | 136.94 |
ENSRNOT00000071666
|
AABR07051550.1
|
|
| chr4_+_101639641 | 136.02 |
ENSRNOT00000058282
|
AABR07060952.1
|
|
| chr3_+_16571602 | 135.40 |
ENSRNOT00000048351
|
LOC100361705
|
rCG64259-like |
| chr3_+_17139670 | 134.72 |
ENSRNOT00000073316
|
AABR07051564.1
|
|
| chr6_+_139486775 | 133.81 |
ENSRNOT00000077771
|
AABR07065699.3
|
|
| chr6_-_138565404 | 133.11 |
ENSRNOT00000079420
|
AABR07065645.2
|
|
| chr2_+_32820322 | 132.96 |
ENSRNOT00000013768
|
Cd180
|
CD180 molecule |
| chr6_-_141008427 | 132.59 |
ENSRNOT00000074472
|
AABR07065778.2
|
|
| chr3_+_16495748 | 132.45 |
ENSRNOT00000045492
|
AABR07051533.1
|
|
| chr4_-_102124609 | 132.21 |
ENSRNOT00000048263
|
AABR07060979.1
|
|
| chr3_-_20457554 | 130.66 |
ENSRNOT00000074237
|
AABR07051733.1
|
|
| chr6_-_138536321 | 128.68 |
ENSRNOT00000077743
|
AABR07065643.1
|
|
| chr6_-_139710905 | 128.60 |
ENSRNOT00000077430
|
AABR07065705.2
|
|
| chr6_-_138772894 | 127.66 |
ENSRNOT00000080779
|
AABR07065651.1
|
|
| chr7_-_18793289 | 126.88 |
ENSRNOT00000036375
|
AABR07056026.1
|
|
| chr4_+_93888502 | 125.98 |
ENSRNOT00000090783
|
AABR07060792.1
|
|
| chr6_-_139004980 | 124.43 |
ENSRNOT00000085427
ENSRNOT00000087351 |
AABR07065656.4
|
|
| chr3_-_16441030 | 123.67 |
ENSRNOT00000047784
|
AABR07051532.1
|
|
| chr6_-_138536162 | 122.76 |
ENSRNOT00000083031
|
AABR07065643.1
|
|
| chr3_-_16753987 | 121.99 |
ENSRNOT00000091257
|
AABR07051548.1
|
|
| chr6_-_138954741 | 121.90 |
ENSRNOT00000083278
|
AABR07065656.7
|
|
| chr1_+_87938042 | 119.03 |
ENSRNOT00000027837
|
Map4k1
|
mitogen activated protein kinase kinase kinase kinase 1 |
| chr6_-_138640187 | 118.96 |
ENSRNOT00000087983
|
AABR07065651.6
|
|
| chr6_-_138954577 | 117.07 |
ENSRNOT00000042728
|
AABR07065656.7
|
|
| chr4_-_103569159 | 116.81 |
ENSRNOT00000084036
|
AABR07061072.1
|
|
| chr9_-_23493081 | 116.02 |
ENSRNOT00000072144
|
Rhag
|
Rh-associated glycoprotein |
| chr6_-_143131118 | 115.53 |
ENSRNOT00000074930
|
AABR07065834.1
|
|
| chr11_+_85042348 | 115.40 |
ENSRNOT00000042220
|
AABR07034718.1
|
|
| chr6_-_139660666 | 114.83 |
ENSRNOT00000086098
|
AABR07065705.1
|
|
| chr6_-_139719323 | 113.31 |
ENSRNOT00000090133
|
AABR07065705.4
|
|
| chr11_+_85992696 | 111.84 |
ENSRNOT00000084433
|
AABR07034736.1
|
|
| chr6_-_138631997 | 110.38 |
ENSRNOT00000073304
|
AABR07065651.3
|
|
| chr8_-_117932518 | 109.53 |
ENSRNOT00000028130
|
Camp
|
cathelicidin antimicrobial peptide |
| chr6_-_142060032 | 109.41 |
ENSRNOT00000064717
|
AABR07065811.1
|
|
| chr6_-_140485913 | 107.44 |
ENSRNOT00000048463
|
AABR07065768.1
|
|
| chr12_+_9360672 | 107.40 |
ENSRNOT00000088957
|
Flt3
|
fms-related tyrosine kinase 3 |
| chr7_-_107634287 | 107.33 |
ENSRNOT00000093672
ENSRNOT00000087116 |
Sla
|
src-like adaptor |
| chr6_-_140715174 | 106.60 |
ENSRNOT00000085345
|
AABR07065773.1
|
|
| chr6_-_141198565 | 105.22 |
ENSRNOT00000064361
|
AABR07065782.1
|
|
| chr10_-_104628676 | 102.89 |
ENSRNOT00000010466
|
Unc13d
|
unc-13 homolog D |
| chr4_+_102068556 | 101.91 |
ENSRNOT00000077412
|
AABR07060971.1
|
|
| chr6_+_139293294 | 101.17 |
ENSRNOT00000050297
ENSRNOT00000081817 |
LOC100359993
|
Ighg protein-like |
| chr4_-_119327822 | 101.17 |
ENSRNOT00000012645
|
Arhgap25
|
Rho GTPase activating protein 25 |
| chr6_-_138931429 | 99.90 |
ENSRNOT00000090584
|
LOC100359993
|
Ighg protein-like |
| chr12_+_38160464 | 98.08 |
ENSRNOT00000032249
|
Hcar2
|
hydroxycarboxylic acid receptor 2 |
| chr6_-_139637187 | 95.56 |
ENSRNOT00000089454
|
LOC100359993
|
Ighg protein-like |
| chr3_+_16590244 | 94.18 |
ENSRNOT00000073229
|
AABR07051535.1
|
|
| chr2_-_139528162 | 94.11 |
ENSRNOT00000014317
|
Slc7a11
|
solute carrier family 7 member 11 |
| chr2_+_55835151 | 94.11 |
ENSRNOT00000018634
|
Fyb
|
FYN binding protein |
| chr17_+_43423111 | 93.14 |
ENSRNOT00000022630
|
Hist1h2ba
|
histone cluster 1 H2B family member a |
| chr6_-_139637354 | 92.77 |
ENSRNOT00000072900
|
LOC100359993
|
Ighg protein-like |
| chr6_+_139523337 | 89.15 |
ENSRNOT00000090711
|
AABR07065699.4
|
|
| chr7_-_107391184 | 88.60 |
ENSRNOT00000056793
|
Tmem71
|
transmembrane protein 71 |
| chr4_+_102290338 | 87.62 |
ENSRNOT00000011067
|
AABR07060992.1
|
|
| chr11_+_85633243 | 86.32 |
ENSRNOT00000045807
|
LOC682352
|
Ig lambda chain V-VI region AR-like |
| chr3_+_19174027 | 85.04 |
ENSRNOT00000074445
|
AABR07051678.1
|
|
| chr5_-_169658875 | 85.02 |
ENSRNOT00000015840
|
Kcnab2
|
potassium voltage-gated channel subfamily A regulatory beta subunit 2 |
| chr6_-_139654508 | 83.60 |
ENSRNOT00000082576
|
AABR07065705.5
|
|
| chr20_+_7788084 | 83.48 |
ENSRNOT00000000597
|
Def6
|
DEF6 guanine nucleotide exchange factor |
| chr6_+_139531330 | 83.27 |
ENSRNOT00000083025
|
AABR07065699.1
|
|
| chr17_-_31916553 | 83.26 |
ENSRNOT00000074220
|
LOC100911032
|
uncharacterized LOC100911032 |
| chr1_-_260254600 | 83.01 |
ENSRNOT00000019014
|
Blnk
|
B-cell linker |
| chr3_+_17009089 | 82.76 |
ENSRNOT00000048829
|
LOC100361052
|
rCG64257-like |
| chr6_+_139293130 | 82.57 |
ENSRNOT00000071714
|
LOC100359993
|
Ighg protein-like |
| chr6_+_139523495 | 82.34 |
ENSRNOT00000075467
|
AABR07065699.4
|
|
| chr4_+_101909389 | 82.03 |
ENSRNOT00000086458
|
AABR07060963.2
|
|
| chr3_+_79918969 | 81.93 |
ENSRNOT00000016306
|
Spi1
|
Spi-1 proto-oncogene |
| chr6_-_140418831 | 80.50 |
ENSRNOT00000086301
|
AABR07065768.2
|
|
| chr17_+_43632397 | 80.43 |
ENSRNOT00000013790
|
Hist1h2ah
|
histone cluster 1, H2ah |
| chr6_-_141365198 | 79.93 |
ENSRNOT00000040523
|
AABR07065789.2
|
|
| chr17_+_44738643 | 78.83 |
ENSRNOT00000087643
|
LOC100910554
|
histone H2A type 1-like |
| chr17_-_43627629 | 78.62 |
ENSRNOT00000022965
|
Hist1h2af
|
histone cluster 1, H2af |
| chr6_-_139033819 | 78.35 |
ENSRNOT00000091961
|
AABR07065656.6
|
|
| chr2_+_198418691 | 78.33 |
ENSRNOT00000089409
|
Hist1h2bk
|
histone cluster 1, H2bk |
| chr6_-_138685656 | 77.01 |
ENSRNOT00000041706
|
AABR07065651.7
|
|
| chr6_-_138550417 | 75.51 |
ENSRNOT00000071945
|
AABR07065645.1
|
|
| chr17_-_44738330 | 74.34 |
ENSRNOT00000072195
|
LOC100364835
|
histone cluster 1, H2bd-like |
| chr17_+_44528125 | 74.06 |
ENSRNOT00000084538
|
LOC680322
|
similar to Histone H2A type 1 |
| chr6_-_141831632 | 71.14 |
ENSRNOT00000075526
|
AABR07065802.1
|
|
| chr4_-_103761881 | 70.92 |
ENSRNOT00000084103
|
AABR07061087.1
|
|
| chr1_-_57327379 | 68.53 |
ENSRNOT00000080429
|
Dll1
|
delta like canonical Notch ligand 1 |
| chr6_-_142178771 | 67.61 |
ENSRNOT00000071577
|
Ighv12-3
|
immunoglobulin heavy variable V12-3 |
| chr9_-_54457753 | 63.45 |
ENSRNOT00000020032
|
Stat1
|
signal transducer and activator of transcription 1 |
| chr10_-_45297385 | 62.36 |
ENSRNOT00000041187
|
Hist3h2bb
|
histone cluster 3 H2B family member b |
| chr17_-_44527801 | 61.29 |
ENSRNOT00000089643
|
Hist1h2bk
|
histone cluster 1 H2B family member k |
| chr15_-_95514259 | 59.19 |
ENSRNOT00000038433
|
Slitrk6
|
SLIT and NTRK-like family, member 6 |
| chr8_+_54993859 | 58.89 |
ENSRNOT00000013093
|
Il18
|
interleukin 18 |
| chr2_+_212247451 | 58.34 |
ENSRNOT00000027813
|
Vav3
|
vav guanine nucleotide exchange factor 3 |
| chrX_+_96863891 | 57.48 |
ENSRNOT00000085665
|
AABR07040284.1
|
|
| chr18_-_12640716 | 56.16 |
ENSRNOT00000020697
|
Klhl14
|
kelch-like family member 14 |
| chr17_+_44520537 | 53.74 |
ENSRNOT00000077985
|
hist1h2ail2
|
histone cluster 1, H2ai-like2 |
| chr3_+_124088157 | 52.28 |
ENSRNOT00000028876
|
Smox
|
spermine oxidase |
| chr1_-_118826977 | 51.46 |
ENSRNOT00000079274
|
AABR07003833.1
|
|
| chr5_+_16526058 | 48.92 |
ENSRNOT00000011130
|
Lyn
|
LYN proto-oncogene, Src family tyrosine kinase |
| chr3_+_19071980 | 48.73 |
ENSRNOT00000079487
|
Igkv4-81
|
immunoglobulin kappa variable 4-81 |
| chr6_-_141957537 | 47.95 |
ENSRNOT00000090358
|
Ighv13-2
|
immunoglobulin heavy variable 13-2 |
| chr6_-_140805551 | 47.08 |
ENSRNOT00000080018
|
AABR07065774.1
|
|
| chr17_-_44520240 | 46.79 |
ENSRNOT00000086538
|
Hist1h2bh
|
histone cluster 1 H2B family member h |
| chr12_+_37544918 | 46.06 |
ENSRNOT00000001403
|
Rilpl2
|
Rab interacting lysosomal protein-like 2 |
| chr4_+_147333056 | 45.98 |
ENSRNOT00000012137
|
Pparg
|
peroxisome proliferator-activated receptor gamma |
| chr2_-_250778269 | 44.64 |
ENSRNOT00000085670
ENSRNOT00000083535 ENSRNOT00000017824 |
Clca5
|
chloride channel accessory 5 |
| chr6_-_55524436 | 43.96 |
ENSRNOT00000006752
|
Tspan13
|
tetraspanin 13 |
| chr10_-_108196217 | 42.81 |
ENSRNOT00000075440
|
Cbx4
|
chromobox 4 |
| chr10_-_37311625 | 41.70 |
ENSRNOT00000043343
|
Jade2
|
jade family PHD finger 2 |
| chr17_-_67904674 | 38.46 |
ENSRNOT00000078532
|
Klf6
|
Kruppel-like factor 6 |
| chr13_-_26769374 | 36.82 |
ENSRNOT00000003768
|
Bcl2
|
BCL2, apoptosis regulator |
| chr3_-_2411544 | 36.29 |
ENSRNOT00000012406
|
Tor4a
|
torsin family 4, member A |
| chr8_-_78397123 | 35.47 |
ENSRNOT00000087270
ENSRNOT00000084925 |
Tcf12
|
transcription factor 12 |
| chr2_-_222935973 | 35.38 |
ENSRNOT00000071934
|
AABR07013095.1
|
|
| chr6_-_141472746 | 35.31 |
ENSRNOT00000048010
|
AABR07065792.2
|
|
| chr18_+_44737154 | 34.68 |
ENSRNOT00000021972
|
Tnfaip8
|
TNF alpha induced protein 8 |
| chr2_-_198360678 | 33.75 |
ENSRNOT00000051917
|
Hist2h2ac
|
histone cluster 2 H2A family member c |
| chr7_-_104541392 | 33.49 |
ENSRNOT00000078116
|
Fam49b
|
family with sequence similarity 49, member B |
| chr10_-_50402616 | 33.39 |
ENSRNOT00000004546
|
Hs3st3b1
|
heparan sulfate-glucosamine 3-sulfotransferase 3B1 |
| chr6_-_125723732 | 32.43 |
ENSRNOT00000084815
|
Fbln5
|
fibulin 5 |
| chr20_-_13657943 | 31.96 |
ENSRNOT00000032290
|
Vpreb3
|
pre-B lymphocyte 3 |
| chr17_+_23116661 | 31.77 |
ENSRNOT00000067374
|
Nedd9
|
neural precursor cell expressed, developmentally down-regulated 9 |
| chr6_-_141866756 | 31.26 |
ENSRNOT00000068561
|
AABR07065804.1
|
|
| chr18_-_86279680 | 30.76 |
ENSRNOT00000006169
|
LOC689166
|
hypothetical protein LOC689166 |
| chr6_-_140587251 | 29.99 |
ENSRNOT00000090704
|
AABR07065772.4
|
|
| chr13_-_89874008 | 29.72 |
ENSRNOT00000051368
|
Ptma
|
prothymosin alpha |
| chr6_-_125723944 | 28.80 |
ENSRNOT00000007004
|
Fbln5
|
fibulin 5 |
| chr17_+_15749978 | 28.72 |
ENSRNOT00000067311
|
Fgd3
|
FYVE, RhoGEF and PH domain containing 3 |
| chr20_-_22459025 | 27.64 |
ENSRNOT00000000792
|
Egr2
|
early growth response 2 |
| chr14_-_86078954 | 26.51 |
ENSRNOT00000018319
|
Polm
|
DNA polymerase mu |
| chr9_+_93545396 | 26.35 |
ENSRNOT00000025093
|
LOC100359583
|
hypothetical protein LOC100359583 |
| chr6_-_141715846 | 25.97 |
ENSRNOT00000044966
|
AABR07065798.1
|
|
| chr14_-_45165207 | 25.61 |
ENSRNOT00000002960
|
Klf3
|
Kruppel like factor 3 |
| chr14_+_48740190 | 24.96 |
ENSRNOT00000031638
|
Arap2
|
ArfGAP with RhoGAP domain, ankyrin repeat and PH domain 2 |
| chr6_-_141384305 | 23.47 |
ENSRNOT00000045320
|
AABR07065790.1
|
|
| chr14_-_59980586 | 21.48 |
ENSRNOT00000064713
|
LOC100361259
|
60S ribosomal protein L13-like |
| chr16_-_21017163 | 20.76 |
ENSRNOT00000027661
|
Mef2b
|
myocyte enhancer factor 2B |
| chr20_+_47596575 | 19.73 |
ENSRNOT00000087230
ENSRNOT00000064905 |
Scml4
|
sex comb on midleg-like 4 (Drosophila) |
| chr4_-_183424449 | 19.71 |
ENSRNOT00000071930
|
Fam60a
|
family with sequence similarity 60, member A |
| chr3_-_152179193 | 19.52 |
ENSRNOT00000026700
|
Rbm12
|
RNA binding motif protein 12 |
| chr3_-_18244535 | 18.58 |
ENSRNOT00000040689
|
AABR07051626.1
|
|
| chr17_+_52938980 | 17.40 |
ENSRNOT00000081052
|
Gabpb1l
|
GA binding protein transcription factor, beta subunit 1-like |
| chr15_+_12827707 | 15.59 |
ENSRNOT00000012452
|
Fezf2
|
Fez family zinc finger 2 |
| chr4_+_106289816 | 14.80 |
ENSRNOT00000040075
|
AABR07061131.1
|
|
| chr10_-_78993045 | 13.00 |
ENSRNOT00000043005
|
LOC100363469
|
ribosomal protein S24-like |
| chr13_-_60849094 | 12.97 |
ENSRNOT00000005156
|
Rgs2
|
regulator of G-protein signaling 2 |
| chr9_-_94495333 | 12.96 |
ENSRNOT00000021507
|
Kcnj13
|
potassium voltage-gated channel subfamily J member 13 |
| chr8_+_114916122 | 12.88 |
ENSRNOT00000074194
|
Tlr9
|
toll-like receptor 9 |
| chr8_-_112648880 | 12.81 |
ENSRNOT00000015265
|
Ackr4
|
atypical chemokine receptor 4 |
Network of associatons between targets according to the STRING database.
First level regulatory network of Pou5f1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 47.5 | 142.6 | GO:0035408 | histone H3-T6 phosphorylation(GO:0035408) |
| 35.8 | 107.4 | GO:0002572 | pro-T cell differentiation(GO:0002572) |
| 34.3 | 102.9 | GO:0002432 | granuloma formation(GO:0002432) |
| 26.6 | 133.0 | GO:0031666 | positive regulation of lipopolysaccharide-mediated signaling pathway(GO:0031666) |
| 25.5 | 76.4 | GO:0045608 | negative regulation of auditory receptor cell differentiation(GO:0045608) |
| 24.5 | 48.9 | GO:0034136 | negative regulation of toll-like receptor 2 signaling pathway(GO:0034136) |
| 20.5 | 81.9 | GO:0045347 | negative regulation of MHC class II biosynthetic process(GO:0045347) |
| 18.3 | 109.5 | GO:0071224 | cellular response to peptidoglycan(GO:0071224) |
| 16.6 | 116.0 | GO:0015670 | carbon dioxide transport(GO:0015670) |
| 15.9 | 63.4 | GO:0034240 | negative regulation of macrophage fusion(GO:0034240) |
| 15.5 | 93.1 | GO:0035093 | spermatogenesis, exchange of chromosomal proteins(GO:0035093) |
| 15.4 | 46.1 | GO:1903445 | protein transport from ciliary membrane to plasma membrane(GO:1903445) |
| 15.3 | 46.0 | GO:2000230 | negative regulation of pancreatic stellate cell proliferation(GO:2000230) |
| 14.7 | 58.9 | GO:0051140 | regulation of NK T cell proliferation(GO:0051140) positive regulation of NK T cell proliferation(GO:0051142) |
| 12.3 | 36.8 | GO:0046671 | regulation of nitrogen utilization(GO:0006808) nitrogen utilization(GO:0019740) negative regulation of retinal cell programmed cell death(GO:0046671) |
| 12.3 | 98.1 | GO:0070163 | positive regulation of neutrophil apoptotic process(GO:0033031) adiponectin secretion(GO:0070162) regulation of adiponectin secretion(GO:0070163) |
| 11.8 | 59.2 | GO:0060005 | vestibular reflex(GO:0060005) |
| 11.2 | 33.5 | GO:2000567 | memory T cell activation(GO:0035709) regulation of memory T cell activation(GO:2000567) positive regulation of memory T cell activation(GO:2000568) |
| 10.2 | 152.3 | GO:0002227 | innate immune response in mucosa(GO:0002227) |
| 9.2 | 27.6 | GO:0021569 | rhombomere 3 development(GO:0021569) |
| 8.7 | 52.3 | GO:0046208 | spermine catabolic process(GO:0046208) |
| 8.5 | 85.0 | GO:0070995 | NADPH oxidation(GO:0070995) |
| 8.1 | 154.6 | GO:0043383 | negative T cell selection(GO:0043383) |
| 6.4 | 12.9 | GO:0032741 | positive regulation of interleukin-18 production(GO:0032741) |
| 6.1 | 61.2 | GO:0048251 | elastic fiber assembly(GO:0048251) |
| 5.5 | 94.1 | GO:0089711 | L-glutamate transmembrane transport(GO:0089711) |
| 4.9 | 227.0 | GO:0050853 | B cell receptor signaling pathway(GO:0050853) |
| 4.8 | 38.5 | GO:0071409 | cellular response to cycloheximide(GO:0071409) |
| 4.8 | 33.4 | GO:0006477 | protein sulfation(GO:0006477) |
| 3.9 | 15.6 | GO:0021797 | forebrain anterior/posterior pattern specification(GO:0021797) |
| 3.8 | 41.7 | GO:0043983 | histone H4-K12 acetylation(GO:0043983) |
| 3.3 | 325.4 | GO:0006342 | chromatin silencing(GO:0006342) |
| 3.2 | 13.0 | GO:0010519 | negative regulation of phospholipase activity(GO:0010519) |
| 2.3 | 29.7 | GO:0043486 | histone exchange(GO:0043486) |
| 2.2 | 26.5 | GO:0016446 | somatic hypermutation of immunoglobulin genes(GO:0016446) |
| 1.9 | 5.6 | GO:0061152 | trachea submucosa development(GO:0061152) trachea gland development(GO:0061153) |
| 1.6 | 6.5 | GO:0097167 | circadian regulation of translation(GO:0097167) |
| 1.6 | 107.3 | GO:0038083 | peptidyl-tyrosine autophosphorylation(GO:0038083) |
| 1.3 | 94.1 | GO:0045576 | mast cell activation(GO:0045576) |
| 1.2 | 30.4 | GO:0002377 | immunoglobulin production(GO:0002377) |
| 1.0 | 8.6 | GO:0043402 | glucocorticoid mediated signaling pathway(GO:0043402) |
| 1.0 | 3.8 | GO:1902202 | regulation of hepatocyte growth factor receptor signaling pathway(GO:1902202) |
| 0.9 | 8.3 | GO:2000774 | positive regulation of cellular senescence(GO:2000774) |
| 0.8 | 69.6 | GO:0006334 | nucleosome assembly(GO:0006334) |
| 0.7 | 34.1 | GO:0016925 | protein sumoylation(GO:0016925) |
| 0.6 | 3.5 | GO:0021633 | optic nerve structural organization(GO:0021633) |
| 0.6 | 8.6 | GO:0042118 | endothelial cell activation(GO:0042118) |
| 0.5 | 3.3 | GO:0035331 | negative regulation of hippo signaling(GO:0035331) |
| 0.4 | 25.7 | GO:0032611 | interleukin-1 beta production(GO:0032611) |
| 0.4 | 83.0 | GO:0042113 | B cell activation(GO:0042113) |
| 0.3 | 66.4 | GO:0046777 | protein autophosphorylation(GO:0046777) |
| 0.3 | 26.4 | GO:0051092 | positive regulation of NF-kappaB transcription factor activity(GO:0051092) |
| 0.3 | 13.0 | GO:0000462 | maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000462) |
| 0.3 | 2.7 | GO:0002043 | blood vessel endothelial cell proliferation involved in sprouting angiogenesis(GO:0002043) |
| 0.3 | 34.3 | GO:1903169 | regulation of calcium ion transmembrane transport(GO:1903169) |
| 0.2 | 12.8 | GO:0070098 | chemokine-mediated signaling pathway(GO:0070098) |
| 0.2 | 124.7 | GO:0043547 | positive regulation of GTPase activity(GO:0043547) |
| 0.2 | 7.6 | GO:0006378 | mRNA polyadenylation(GO:0006378) |
| 0.2 | 4.6 | GO:0016574 | histone ubiquitination(GO:0016574) |
| 0.1 | 0.6 | GO:0038203 | TORC2 signaling(GO:0038203) |
| 0.1 | 0.9 | GO:0043517 | positive regulation of DNA damage response, signal transduction by p53 class mediator(GO:0043517) |
| 0.0 | 0.5 | GO:1903830 | magnesium ion transmembrane transport(GO:1903830) |
| 0.0 | 0.3 | GO:0014912 | negative regulation of smooth muscle cell migration(GO:0014912) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 59.0 | 235.8 | GO:0019815 | B cell receptor complex(GO:0019815) |
| 17.1 | 102.9 | GO:0033093 | Weibel-Palade body(GO:0033093) |
| 16.3 | 48.9 | GO:0034666 | integrin alpha2-beta1 complex(GO:0034666) |
| 12.2 | 61.2 | GO:0071953 | elastic fiber(GO:0071953) |
| 6.5 | 640.4 | GO:0000786 | nucleosome(GO:0000786) |
| 6.4 | 109.5 | GO:0042581 | specific granule(GO:0042581) |
| 5.5 | 88.5 | GO:0044224 | juxtaparanode region of axon(GO:0044224) |
| 4.0 | 154.6 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 3.2 | 47.4 | GO:0035102 | PRC1 complex(GO:0035102) |
| 2.0 | 7.9 | GO:0002193 | MAML1-RBP-Jkappa- ICN1 complex(GO:0002193) |
| 1.8 | 26.9 | GO:0016580 | Sin3 complex(GO:0016580) |
| 1.6 | 142.6 | GO:0031526 | brush border membrane(GO:0031526) |
| 1.3 | 12.9 | GO:0032009 | early phagosome(GO:0032009) |
| 1.1 | 56.2 | GO:0016235 | aggresome(GO:0016235) |
| 1.1 | 7.6 | GO:0042382 | paraspeckles(GO:0042382) |
| 1.0 | 36.8 | GO:0046930 | pore complex(GO:0046930) |
| 0.8 | 94.1 | GO:0005791 | rough endoplasmic reticulum(GO:0005791) |
| 0.7 | 91.5 | GO:0031234 | extrinsic component of cytoplasmic side of plasma membrane(GO:0031234) |
| 0.5 | 41.7 | GO:0000123 | histone acetyltransferase complex(GO:0000123) |
| 0.5 | 64.6 | GO:0090575 | RNA polymerase II transcription factor complex(GO:0090575) |
| 0.4 | 51.5 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.3 | 8.9 | GO:0030131 | clathrin adaptor complex(GO:0030131) |
| 0.2 | 77.7 | GO:0016324 | apical plasma membrane(GO:0016324) |
| 0.2 | 21.5 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.2 | 13.0 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.2 | 13.0 | GO:0009898 | cytoplasmic side of plasma membrane(GO:0009898) |
| 0.2 | 11.7 | GO:0015934 | large ribosomal subunit(GO:0015934) |
| 0.2 | 42.9 | GO:0009897 | external side of plasma membrane(GO:0009897) |
| 0.2 | 41.5 | GO:0000790 | nuclear chromatin(GO:0000790) |
| 0.1 | 0.6 | GO:0061700 | GATOR2 complex(GO:0061700) |
| 0.1 | 16.0 | GO:0031965 | nuclear membrane(GO:0031965) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 47.5 | 142.6 | GO:0035403 | histone kinase activity (H3-T6 specific)(GO:0035403) |
| 31.4 | 94.1 | GO:0005294 | neutral L-amino acid secondary active transmembrane transporter activity(GO:0005294) |
| 19.8 | 119.0 | GO:0008349 | MAP kinase kinase kinase kinase activity(GO:0008349) |
| 17.4 | 52.3 | GO:0046592 | polyamine oxidase activity(GO:0046592) |
| 14.5 | 116.0 | GO:0022840 | leak channel activity(GO:0022840) narrow pore channel activity(GO:0022842) |
| 11.2 | 33.5 | GO:0023025 | MHC class Ib protein complex binding(GO:0023025) MHC class Ib protein binding, via antigen binding groove(GO:0023030) |
| 10.4 | 83.0 | GO:0005068 | transmembrane receptor protein tyrosine kinase adaptor activity(GO:0005068) |
| 10.2 | 81.9 | GO:0051525 | NFAT protein binding(GO:0051525) |
| 7.6 | 68.5 | GO:0030957 | Tat protein binding(GO:0030957) |
| 6.6 | 46.0 | GO:0050692 | DBD domain binding(GO:0050692) |
| 6.1 | 36.8 | GO:0051434 | BH3 domain binding(GO:0051434) |
| 5.6 | 33.4 | GO:0008467 | [heparan sulfate]-glucosamine 3-sulfotransferase 1 activity(GO:0008467) |
| 5.4 | 48.9 | GO:0043208 | glycosphingolipid binding(GO:0043208) |
| 5.3 | 63.4 | GO:0031730 | CCR5 chemokine receptor binding(GO:0031730) |
| 4.6 | 46.1 | GO:0051959 | dynein light intermediate chain binding(GO:0051959) |
| 4.5 | 107.4 | GO:0035259 | glucocorticoid receptor binding(GO:0035259) |
| 4.5 | 98.1 | GO:0001614 | purinergic nucleotide receptor activity(GO:0001614) nucleotide receptor activity(GO:0016502) |
| 3.9 | 85.0 | GO:0004033 | aldo-keto reductase (NADP) activity(GO:0004033) |
| 2.9 | 8.6 | GO:0001087 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
| 2.7 | 58.3 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
| 2.6 | 12.9 | GO:0045322 | unmethylated CpG binding(GO:0045322) |
| 2.4 | 42.8 | GO:0019789 | SUMO transferase activity(GO:0019789) |
| 2.2 | 35.5 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 1.7 | 8.6 | GO:0043423 | 3-phosphoinositide-dependent protein kinase binding(GO:0043423) |
| 1.5 | 91.5 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 1.3 | 3.8 | GO:0005171 | hepatocyte growth factor receptor binding(GO:0005171) |
| 1.2 | 34.7 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) |
| 0.9 | 26.5 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.7 | 57.4 | GO:0033613 | activating transcription factor binding(GO:0033613) |
| 0.6 | 13.0 | GO:0005242 | inward rectifier potassium channel activity(GO:0005242) |
| 0.5 | 25.0 | GO:0005547 | phosphatidylinositol-3,4,5-trisphosphate binding(GO:0005547) |
| 0.5 | 12.8 | GO:0019956 | chemokine binding(GO:0019956) |
| 0.5 | 28.7 | GO:0005089 | Rho guanyl-nucleotide exchange factor activity(GO:0005089) |
| 0.4 | 48.9 | GO:0005178 | integrin binding(GO:0005178) |
| 0.4 | 111.6 | GO:0005096 | GTPase activator activity(GO:0005096) |
| 0.4 | 11.3 | GO:0035035 | histone acetyltransferase binding(GO:0035035) |
| 0.3 | 12.9 | GO:0001205 | transcriptional activator activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001205) |
| 0.2 | 3.5 | GO:0015271 | outward rectifier potassium channel activity(GO:0015271) |
| 0.2 | 46.2 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.2 | 7.9 | GO:0001190 | transcriptional activator activity, RNA polymerase II transcription factor binding(GO:0001190) transcriptional repressor activity, RNA polymerase II activating transcription factor binding(GO:0098811) |
| 0.1 | 3.6 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.1 | 4.6 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.0 | 21.3 | GO:0004842 | ubiquitin-protein transferase activity(GO:0004842) |
| 0.0 | 0.5 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.0 | 0.2 | GO:0015119 | hexose phosphate transmembrane transporter activity(GO:0015119) organophosphate:inorganic phosphate antiporter activity(GO:0015315) hexose-phosphate:inorganic phosphate antiporter activity(GO:0015526) glucose 6-phosphate:inorganic phosphate antiporter activity(GO:0061513) |
| 0.0 | 0.9 | GO:0030291 | protein serine/threonine kinase inhibitor activity(GO:0030291) |
| 0.0 | 2.9 | GO:0017137 | Rab GTPase binding(GO:0017137) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 18.7 | 261.7 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
| 15.6 | 109.5 | ST PAC1 RECEPTOR PATHWAY | PAC1 Receptor Pathway |
| 6.3 | 190.3 | SA B CELL RECEPTOR COMPLEXES | Antigen binding to B cell receptors activates protein tyrosine kinases, such as the Src family, which ultimate activate MAP kinases. |
| 4.5 | 63.4 | ST INTERFERON GAMMA PATHWAY | Interferon gamma pathway. |
| 4.2 | 235.8 | PID BCR 5PATHWAY | BCR signaling pathway |
| 3.3 | 36.8 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 3.1 | 58.9 | PID IL27 PATHWAY | IL27-mediated signaling events |
| 1.9 | 83.5 | PID RAC1 REG PATHWAY | Regulation of RAC1 activity |
| 1.8 | 109.6 | PID IL4 2PATHWAY | IL4-mediated signaling events |
| 1.8 | 84.4 | PID TCR PATHWAY | TCR signaling in naïve CD4+ T cells |
| 1.5 | 26.5 | PID DNA PK PATHWAY | DNA-PK pathway in nonhomologous end joining |
| 1.5 | 66.7 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 1.4 | 38.7 | ST ERK1 ERK2 MAPK PATHWAY | ERK1/ERK2 MAPK Pathway |
| 1.2 | 46.0 | PID WNT NONCANONICAL PATHWAY | Noncanonical Wnt signaling pathway |
| 1.1 | 42.2 | PID PS1 PATHWAY | Presenilin action in Notch and Wnt signaling |
| 0.6 | 25.0 | PID ARF6 PATHWAY | Arf6 signaling events |
| 0.6 | 42.8 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.3 | 8.6 | PID PI3KCI PATHWAY | Class I PI3K signaling events |
| 0.1 | 28.3 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 0.9 | PID KIT PATHWAY | Signaling events mediated by Stem cell factor receptor (c-Kit) |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 14.9 | 388.3 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |
| 8.8 | 342.4 | REACTOME ANTIGEN ACTIVATES B CELL RECEPTOR LEADING TO GENERATION OF SECOND MESSENGERS | Genes involved in Antigen Activates B Cell Receptor Leading to Generation of Second Messengers |
| 7.2 | 137.7 | REACTOME TRAFFICKING OF GLUR2 CONTAINING AMPA RECEPTORS | Genes involved in Trafficking of GluR2-containing AMPA receptors |
| 6.7 | 94.1 | REACTOME SIGNAL REGULATORY PROTEIN SIRP FAMILY INTERACTIONS | Genes involved in Signal regulatory protein (SIRP) family interactions |
| 5.7 | 68.5 | REACTOME SIGNALING BY NOTCH3 | Genes involved in Signaling by NOTCH3 |
| 5.3 | 63.4 | REACTOME IL 6 SIGNALING | Genes involved in Interleukin-6 signaling |
| 3.1 | 109.5 | REACTOME LATENT INFECTION OF HOMO SAPIENS WITH MYCOBACTERIUM TUBERCULOSIS | Genes involved in Latent infection of Homo sapiens with Mycobacterium tuberculosis |
| 3.1 | 116.0 | REACTOME AMINE COMPOUND SLC TRANSPORTERS | Genes involved in Amine compound SLC transporters |
| 2.9 | 94.1 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 2.9 | 52.3 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |
| 1.9 | 36.8 | REACTOME INFLAMMASOMES | Genes involved in Inflammasomes |
| 1.9 | 58.3 | REACTOME GPVI MEDIATED ACTIVATION CASCADE | Genes involved in GPVI-mediated activation cascade |
| 1.4 | 12.9 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION IN TLR7 8 OR 9 SIGNALING | Genes involved in TRAF6 mediated IRF7 activation in TLR7/8 or 9 signaling |
| 1.1 | 154.9 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 1.1 | 31.7 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
| 1.0 | 28.2 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.9 | 58.9 | REACTOME SIGNALING BY ILS | Genes involved in Signaling by Interleukins |
| 0.9 | 107.0 | REACTOME TOLL RECEPTOR CASCADES | Genes involved in Toll Receptor Cascades |
| 0.7 | 7.6 | REACTOME SIGNALING BY FGFR1 FUSION MUTANTS | Genes involved in Signaling by FGFR1 fusion mutants |
| 0.6 | 6.5 | REACTOME BMAL1 CLOCK NPAS2 ACTIVATES CIRCADIAN EXPRESSION | Genes involved in BMAL1:CLOCK/NPAS2 Activates Circadian Expression |
| 0.5 | 98.1 | REACTOME G ALPHA I SIGNALLING EVENTS | Genes involved in G alpha (i) signalling events |
| 0.4 | 9.2 | REACTOME PROCESSING OF INTRONLESS PRE MRNAS | Genes involved in Processing of Intronless Pre-mRNAs |
| 0.1 | 7.5 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.0 | 0.3 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |