Project
GSE53960: rat RNA-Seq transcriptomic Bodymap
Navigation
Downloads
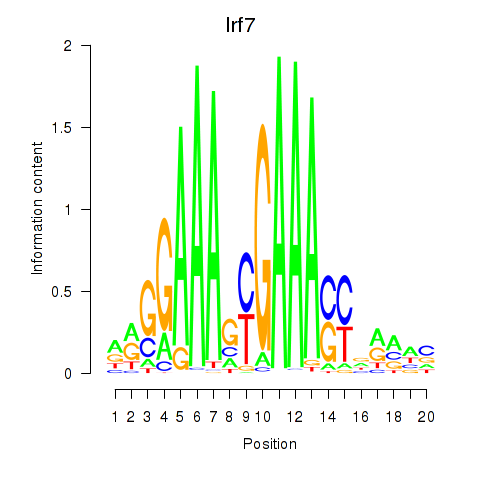
Results for Irf2_Irf1_Irf8_Irf9_Irf7
Z-value: 4.08
Transcription factors associated with Irf2_Irf1_Irf8_Irf9_Irf7
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Irf2
|
ENSRNOG00000009824 | interferon regulatory factor 2 |
|
Irf1
|
ENSRNOG00000008144 | interferon regulatory factor 1 |
|
Irf8
|
ENSRNOG00000017869 | interferon regulatory factor 8 |
|
Irf9
|
ENSRNOG00000019478 | interferon regulatory factor 9 |
|
Irf7
|
ENSRNOG00000017414 | interferon regulatory factor 7 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Irf1 | rn6_v1_chr10_+_39109522_39109522 | 0.91 | 5.2e-123 | Click! |
| Irf7 | rn6_v1_chr1_-_214252456_214252456 | 0.88 | 1.2e-105 | Click! |
| Irf9 | rn6_v1_chr15_+_34282936_34282936 | 0.85 | 6.3e-91 | Click! |
| Irf8 | rn6_v1_chr19_+_54314865_54314865 | 0.77 | 4.0e-63 | Click! |
| Irf2 | rn6_v1_chr16_-_48692476_48692476 | 0.69 | 8.2e-47 | Click! |
Activity profile of Irf2_Irf1_Irf8_Irf9_Irf7 motif
Sorted Z-values of Irf2_Irf1_Irf8_Irf9_Irf7 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr20_-_3978845 | 283.69 |
ENSRNOT00000000532
|
Psmb9
|
proteasome subunit beta 9 |
| chr4_+_153805993 | 234.55 |
ENSRNOT00000056174
|
Usp18
|
ubiquitin specific peptidase 18 |
| chr10_+_34277993 | 232.98 |
ENSRNOT00000055872
ENSRNOT00000003343 |
Ifi47
|
interferon gamma inducible protein 47 |
| chr3_-_171286413 | 207.24 |
ENSRNOT00000008365
ENSRNOT00000081036 |
Zbp1
|
Z-DNA binding protein 1 |
| chr1_-_214252456 | 187.13 |
ENSRNOT00000023504
|
Irf7
|
interferon regulatory factor 7 |
| chr16_-_19942343 | 185.56 |
ENSRNOT00000087162
ENSRNOT00000091906 |
Bst2
|
bone marrow stromal cell antigen 2 |
| chr10_+_108527740 | 184.93 |
ENSRNOT00000044983
|
Rnf213
|
ring finger protein 213 |
| chr11_-_37914983 | 173.91 |
ENSRNOT00000039876
|
Mx1
|
myxovirus (influenza virus) resistance 1 |
| chr7_-_117300662 | 173.25 |
ENSRNOT00000006376
|
Parp10
|
poly (ADP-ribose) polymerase family, member 10 |
| chr5_-_58198782 | 166.60 |
ENSRNOT00000023951
|
Ccl21
|
C-C motif chemokine ligand 21 |
| chr20_+_3990820 | 164.66 |
ENSRNOT00000000528
|
Psmb8
|
proteasome subunit beta 8 |
| chr11_+_67757928 | 164.03 |
ENSRNOT00000039215
|
Dtx3l
|
deltex E3 ubiquitin ligase 3L |
| chr20_-_27682861 | 159.60 |
ENSRNOT00000057317
|
Fam26f
|
family with sequence similarity 26, member F |
| chr5_-_173626248 | 159.33 |
ENSRNOT00000039263
|
Isg15
|
ISG15 ubiquitin-like modifier |
| chr20_+_3979035 | 151.65 |
ENSRNOT00000000529
|
Tap1
|
transporter 1, ATP binding cassette subfamily B member |
| chr19_-_37912027 | 150.35 |
ENSRNOT00000026462
|
Psmb10
|
proteasome subunit beta 10 |
| chr18_+_55576239 | 146.89 |
ENSRNOT00000050063
|
LOC100910979
|
interferon-inducible GTPase 1-like |
| chr2_+_248276709 | 145.19 |
ENSRNOT00000068683
|
Gbp2
|
guanylate binding protein 2 |
| chr5_-_163167299 | 144.67 |
ENSRNOT00000022478
|
Tnfrsf1b
|
TNF receptor superfamily member 1B |
| chr11_+_68105369 | 142.11 |
ENSRNOT00000046888
|
Parp14
|
poly (ADP-ribose) polymerase family, member 14 |
| chr2_-_256915563 | 140.98 |
ENSRNOT00000084873
ENSRNOT00000029990 |
Ifi44
|
interferon-induced protein 44 |
| chr10_+_31880918 | 140.13 |
ENSRNOT00000059448
|
Timd4
|
T-cell immunoglobulin and mucin domain containing 4 |
| chr7_+_116889572 | 129.97 |
ENSRNOT00000075917
|
Gsdmd
|
gasdermin D |
| chr1_-_167708685 | 120.60 |
ENSRNOT00000092857
ENSRNOT00000024992 |
Trim21
|
tripartite motif-containing 21 |
| chr3_-_37480984 | 118.14 |
ENSRNOT00000030373
|
Nmi
|
N-myc (and STAT) interactor |
| chr20_-_1878126 | 112.60 |
ENSRNOT00000000995
|
Ubd
|
ubiquitin D |
| chr1_-_166677026 | 112.05 |
ENSRNOT00000026644
ENSRNOT00000076714 |
Art2b
|
ADP-ribosyltransferase 2b |
| chr2_+_209097927 | 109.97 |
ENSRNOT00000023807
|
Dennd2d
|
DENN domain containing 2D |
| chr4_+_67165883 | 108.97 |
ENSRNOT00000012280
|
Rab19
|
RAB19, member RAS oncogene family |
| chr1_+_226897625 | 108.82 |
ENSRNOT00000029443
|
Slc15a3
|
solute carrier family 15 member 3 |
| chr8_+_2604962 | 104.61 |
ENSRNOT00000009993
|
Casp1
|
caspase 1 |
| chr6_-_127337791 | 102.20 |
ENSRNOT00000032015
|
Ifi27l2b
|
interferon, alpha-inducible protein 27 like 2B |
| chr20_-_3166569 | 101.78 |
ENSRNOT00000091390
|
AABR07044362.1
|
|
| chr13_+_93684437 | 101.10 |
ENSRNOT00000005005
|
Kmo
|
kynurenine 3-monooxygenase |
| chr10_-_34221928 | 100.94 |
ENSRNOT00000045545
|
Irgm
|
immunity-related GTPase M |
| chr14_-_43992587 | 99.49 |
ENSRNOT00000003425
|
Rhoh
|
ras homolog family member H |
| chr10_-_70342411 | 99.43 |
ENSRNOT00000076269
ENSRNOT00000076477 |
Slfn13
|
schlafen family member 13 |
| chr10_+_43613931 | 97.38 |
ENSRNOT00000039335
|
Igtp
|
interferon gamma induced GTPase |
| chr15_+_60084918 | 96.77 |
ENSRNOT00000012632
|
Epsti1
|
epithelial stromal interaction 1 |
| chr2_+_248178389 | 96.30 |
ENSRNOT00000037339
|
Gbp5
|
guanylate binding protein 5 |
| chrX_+_14578264 | 94.59 |
ENSRNOT00000038994
|
Cybb
|
cytochrome b-245 beta chain |
| chr3_-_160802433 | 94.05 |
ENSRNOT00000076191
|
Slpi
|
secretory leukocyte peptidase inhibitor |
| chr20_+_3995544 | 93.11 |
ENSRNOT00000000527
|
Tap2
|
transporter 2, ATP binding cassette subfamily B member |
| chr20_+_3167079 | 92.44 |
ENSRNOT00000001035
|
RT1-N3
|
RT1 class Ib, locus N3 |
| chr20_-_5476193 | 91.32 |
ENSRNOT00000044975
|
Tapbp
|
TAP binding protein |
| chr14_+_17210733 | 90.79 |
ENSRNOT00000003075
|
Cxcl10
|
C-X-C motif chemokine ligand 10 |
| chr9_-_92616165 | 90.57 |
ENSRNOT00000056995
|
Sp110
|
SP110 nuclear body protein |
| chr16_-_20383337 | 90.15 |
ENSRNOT00000025977
|
Il12rb1
|
interleukin 12 receptor subunit beta 1 |
| chr4_-_66062089 | 87.06 |
ENSRNOT00000018782
|
Zc3hav1
|
zinc finger CCCH-type containing, antiviral 1 |
| chr8_+_116754178 | 86.67 |
ENSRNOT00000068295
ENSRNOT00000084429 |
Uba7
|
ubiquitin-like modifier activating enzyme 7 |
| chr1_+_247519939 | 86.36 |
ENSRNOT00000034421
|
Cd274
|
CD274 molecule |
| chr4_-_157743199 | 85.35 |
ENSRNOT00000038178
|
Tapbpl
|
TAP binding protein-like |
| chr11_+_38035611 | 84.88 |
ENSRNOT00000087603
ENSRNOT00000002695 |
Mx2
|
MX dynamin like GTPase 2 |
| chr20_-_5212624 | 84.78 |
ENSRNOT00000074261
|
LOC103689996
|
antigen peptide transporter 2 |
| chr2_+_248249468 | 83.99 |
ENSRNOT00000022648
|
Gbp4
|
guanylate binding protein 4 |
| chr2_-_44981458 | 82.19 |
ENSRNOT00000014134
|
Gzma
|
granzyme A |
| chr1_+_169433539 | 82.02 |
ENSRNOT00000055210
|
Trim34
|
tripartite motif-containing 34 |
| chr11_+_38035450 | 81.90 |
ENSRNOT00000083067
|
Mx2
|
MX dynamin like GTPase 2 |
| chr10_+_70417108 | 80.36 |
ENSRNOT00000079325
|
Slfn4
|
schlafen 4 |
| chr1_+_277190964 | 80.27 |
ENSRNOT00000080511
|
Casp7
|
caspase 7 |
| chr10_-_107424710 | 79.85 |
ENSRNOT00000004320
|
Lgals3bp
|
galectin 3 binding protein |
| chr8_+_96551245 | 79.29 |
ENSRNOT00000039850
|
Bcl2a1
|
BCL2-related protein A1 |
| chr10_+_70411738 | 78.95 |
ENSRNOT00000078046
|
Slfn4
|
schlafen 4 |
| chr20_-_4698718 | 78.77 |
ENSRNOT00000047527
|
RT1-CE7
|
RT1 class I, locus CE7 |
| chr6_-_45669148 | 78.72 |
ENSRNOT00000010092
|
Rsad2
|
radical S-adenosyl methionine domain containing 2 |
| chr1_+_247562202 | 77.99 |
ENSRNOT00000021614
|
Pdcd1lg2
|
programmed cell death 1 ligand 2 |
| chr10_+_58860940 | 77.19 |
ENSRNOT00000056551
ENSRNOT00000074523 |
XAF1
|
XIAP associated factor-1 |
| chr4_-_66899914 | 75.75 |
ENSRNOT00000011481
|
Parp12
|
poly (ADP-ribose) polymerase family, member 12 |
| chr15_+_34256071 | 74.97 |
ENSRNOT00000025887
|
Psme1
|
proteasome activator subunit 1 |
| chr1_+_252906234 | 74.13 |
ENSRNOT00000031025
|
Ifit3
|
interferon-induced protein with tetratricopeptide repeats 3 |
| chr9_-_54327958 | 73.76 |
ENSRNOT00000019465
|
Stat1
|
signal transducer and activator of transcription 1 |
| chr1_-_101131413 | 73.48 |
ENSRNOT00000093729
|
Flt3lg
|
fms-related tyrosine kinase 3 ligand |
| chr1_+_140601791 | 73.31 |
ENSRNOT00000091588
|
Isg20
|
interferon stimulated exonuclease gene 20 |
| chr5_-_56536772 | 72.32 |
ENSRNOT00000060765
|
Ddx58
|
DEXD/H-box helicase 58 |
| chr4_+_179398621 | 71.67 |
ENSRNOT00000049474
ENSRNOT00000067506 |
Lrmp
|
lymphoid-restricted membrane protein |
| chr15_+_18710492 | 71.62 |
ENSRNOT00000012532
|
Dnase1l3
|
deoxyribonuclease 1-like 3 |
| chr7_+_118685181 | 71.49 |
ENSRNOT00000068221
|
Apol3
|
apolipoprotein L, 3 |
| chr19_-_55367353 | 71.23 |
ENSRNOT00000091139
|
Piezo1
|
piezo-type mechanosensitive ion channel component 1 |
| chr10_-_5251809 | 70.82 |
ENSRNOT00000085479
|
Ciita
|
class II, major histocompatibility complex, transactivator |
| chr17_-_32661865 | 70.15 |
ENSRNOT00000022194
|
Serpinb9
|
serpin family B member 9 |
| chr10_-_66229311 | 69.91 |
ENSRNOT00000016897
|
Lgals5
|
lectin, galactose binding, soluble 5 |
| chr10_-_57064600 | 68.72 |
ENSRNOT00000032926
|
Cxcl16
|
C-X-C motif chemokine ligand 16 |
| chr20_+_3146856 | 68.52 |
ENSRNOT00000050159
|
RT1-N2
|
RT1 class Ib, locus N2 |
| chr8_+_116857684 | 67.88 |
ENSRNOT00000026711
|
Mst1
|
macrophage stimulating 1 |
| chr10_-_5253336 | 66.45 |
ENSRNOT00000085310
|
Ciita
|
class II, major histocompatibility complex, transactivator |
| chr20_-_3793985 | 66.21 |
ENSRNOT00000049540
ENSRNOT00000086293 |
RT1-CE16
|
RT1 class I, locus CE16 |
| chr3_+_114087287 | 66.16 |
ENSRNOT00000023017
|
B2m
|
beta-2 microglobulin |
| chr5_-_58183017 | 65.88 |
ENSRNOT00000020982
|
Ccl19
|
C-C motif chemokine ligand 19 |
| chr1_+_221538104 | 64.80 |
ENSRNOT00000028527
|
Batf2
|
basic leucine zipper ATF-like transcription factor 2 |
| chr4_+_163174487 | 64.62 |
ENSRNOT00000088108
|
Clec9a
|
C-type lectin domain family 9, member A |
| chr7_+_124460358 | 62.82 |
ENSRNOT00000014089
|
Tspo
|
translocator protein |
| chrX_+_11648989 | 61.81 |
ENSRNOT00000041003
|
Bcor
|
BCL6 co-repressor |
| chr4_-_88649216 | 61.31 |
ENSRNOT00000058626
|
Herc6
|
HECT and RLD domain containing E3 ubiquitin protein ligase family member 6 |
| chr1_-_252922486 | 60.88 |
ENSRNOT00000076802
|
Ifit1bl
|
interferon-induced protein with tetratricopeptide repeats 1B-like |
| chr1_-_169463760 | 60.30 |
ENSRNOT00000023100
|
Trim30c
|
tripartite motif-containing 30C |
| chr18_+_56071478 | 58.98 |
ENSRNOT00000025344
ENSRNOT00000025354 |
Cd74
|
CD74 molecule |
| chr20_+_11436267 | 58.88 |
ENSRNOT00000001631
|
Trpm2
|
transient receptor potential cation channel, subfamily M, member 2 |
| chr16_-_48692476 | 58.65 |
ENSRNOT00000013118
|
Irf2
|
interferon regulatory factor 2 |
| chr2_-_207300854 | 58.43 |
ENSRNOT00000018061
|
Mov10
|
Mov10 RISC complex RNA helicase |
| chr4_+_162934195 | 58.39 |
ENSRNOT00000090939
|
Clec2g
|
C-type lectin domain family 2, member G |
| chr17_-_43640387 | 58.27 |
ENSRNOT00000087731
|
Hist1h1c
|
histone cluster 1 H1 family member c |
| chr7_-_28932641 | 58.13 |
ENSRNOT00000059487
|
Dram1
|
DNA-damage regulated autophagy modulator 1 |
| chr17_-_44840131 | 57.24 |
ENSRNOT00000083417
|
Hist1h3b
|
histone cluster 1, H3b |
| chr20_-_1818800 | 56.75 |
ENSRNOT00000000990
|
RT1-M3-1
|
RT1 class Ib, locus M3, gene 1 |
| chr16_+_50111306 | 56.60 |
ENSRNOT00000019302
|
Cyp4v3
|
cytochrome P450, family 4, subfamily v, polypeptide 3 |
| chr12_+_41200718 | 56.18 |
ENSRNOT00000038426
ENSRNOT00000048450 ENSRNOT00000067176 |
Oas1a
|
2'-5' oligoadenylate synthetase 1A |
| chr13_-_102643223 | 55.77 |
ENSRNOT00000003155
|
Hlx
|
H2.0-like homeobox |
| chr8_-_124399494 | 55.46 |
ENSRNOT00000037883
|
Tgfbr2
|
transforming growth factor, beta receptor 2 |
| chr15_+_36809361 | 55.17 |
ENSRNOT00000076667
|
Parp4
|
poly (ADP-ribose) polymerase family, member 4 |
| chr3_-_48604097 | 55.11 |
ENSRNOT00000009620
|
Ifih1
|
interferon induced with helicase C domain 1 |
| chr12_+_42343123 | 54.95 |
ENSRNOT00000043279
|
Oas1i
|
2 ' -5 ' oligoadenylate synthetase 1I |
| chr4_-_163403653 | 54.87 |
ENSRNOT00000088151
|
Klrk1
|
killer cell lectin like receptor K1 |
| chr2_-_241545250 | 54.54 |
ENSRNOT00000073875
|
Bank1
|
B-cell scaffold protein with ankyrin repeats 1 |
| chr10_-_110274768 | 54.52 |
ENSRNOT00000054931
|
Sectm1a
|
secreted and transmembrane 1A |
| chr1_-_227441442 | 53.99 |
ENSRNOT00000028433
|
Ms4a1
|
membrane spanning 4-domains A1 |
| chr3_+_161519743 | 53.71 |
ENSRNOT00000055148
|
Cd40
|
CD40 molecule |
| chr10_+_89358376 | 53.57 |
ENSRNOT00000028067
|
Ifi35
|
interferon-induced protein 35 |
| chr7_+_118685022 | 53.51 |
ENSRNOT00000089135
|
Apol3
|
apolipoprotein L, 3 |
| chr4_+_162493908 | 53.34 |
ENSRNOT00000072064
|
Clec2d
|
C-type lectin domain family 2, member D |
| chr12_+_41341417 | 51.22 |
ENSRNOT00000072024
ENSRNOT00000091460 |
Oas2
|
2'-5' oligoadenylate synthetase 2 |
| chr20_+_5374985 | 50.90 |
ENSRNOT00000052270
|
RT1-A2
|
RT1 class Ia, locus A2 |
| chr10_+_76343847 | 50.90 |
ENSRNOT00000055674
|
Trim25
|
tripartite motif-containing 25 |
| chr20_-_1339488 | 50.00 |
ENSRNOT00000041074
|
RT1-M2
|
RT1 class Ib, locus M2 |
| chr20_-_4935887 | 49.90 |
ENSRNOT00000064734
|
RT1-CE4
|
RT1 class I, locus CE4 |
| chr20_-_35450513 | 49.70 |
ENSRNOT00000087342
|
Man1a1
|
mannosidase, alpha, class 1A, member 1 |
| chr1_-_213811901 | 49.08 |
ENSRNOT00000020265
|
Ifitm3
|
interferon induced transmembrane protein 3 |
| chr18_+_55685613 | 48.07 |
ENSRNOT00000040116
|
MGC105567
|
similar to cDNA sequence BC023105 |
| chr20_-_4935372 | 47.82 |
ENSRNOT00000050099
ENSRNOT00000047779 |
RT1-CE3
RT1-CE4
|
RT1 class I, locus CE3 RT1 class I, locus CE4 |
| chr16_-_19349080 | 47.51 |
ENSRNOT00000038494
|
Hsh2d
|
hematopoietic SH2 domain containing |
| chr4_-_163214678 | 47.39 |
ENSRNOT00000091602
|
Clec1a
|
C-type lectin domain family 1, member A |
| chr3_-_176744377 | 47.35 |
ENSRNOT00000017787
|
Helz2
|
helicase with zinc finger 2, transcriptional coactivator |
| chr20_-_4542073 | 46.96 |
ENSRNOT00000000477
|
Cfb
|
complement factor B |
| chr1_-_101131012 | 46.86 |
ENSRNOT00000082283
ENSRNOT00000093498 ENSRNOT00000093559 |
Flt3lg
Rpl13a
|
fms-related tyrosine kinase 3 ligand ribosomal protein L13A |
| chr11_-_14304603 | 46.53 |
ENSRNOT00000040202
ENSRNOT00000082143 |
Samsn1
|
SAM domain, SH3 domain and nuclear localization signals, 1 |
| chr8_-_128026841 | 46.43 |
ENSRNOT00000018341
|
Myd88
|
myeloid differentiation primary response 88 |
| chr1_-_167093560 | 45.88 |
ENSRNOT00000027301
|
Il18bp
|
interleukin 18 binding protein |
| chr4_-_72074683 | 45.81 |
ENSRNOT00000071511
|
Fam115c
|
family with sequence similarity 115, member C |
| chr12_+_41155497 | 45.28 |
ENSRNOT00000041741
|
Oas1g
|
2'-5' oligoadenylate synthetase 1G |
| chr10_+_64737022 | 44.96 |
ENSRNOT00000017071
ENSRNOT00000093232 ENSRNOT00000017042 ENSRNOT00000093244 |
Lgals9
|
galectin 9 |
| chr16_+_72401887 | 44.40 |
ENSRNOT00000074449
|
LOC100910163
|
uncharacterized LOC100910163 |
| chr9_+_92618352 | 44.34 |
ENSRNOT00000034603
|
Sp140
|
SP140 nuclear body protein |
| chr20_-_4896970 | 44.14 |
ENSRNOT00000061114
ENSRNOT00000060980 |
RT1-CE7
RT1-CE5
|
RT1 class I, locus CE7 RT1 class I, locus CE5 |
| chr3_-_17081510 | 44.02 |
ENSRNOT00000063862
|
AABR07051562.1
|
|
| chr20_-_157861 | 44.02 |
ENSRNOT00000084461
|
RT1-CE10
|
RT1 class I, locus CE10 |
| chr9_+_65620658 | 43.71 |
ENSRNOT00000084498
|
Casp8
|
caspase 8 |
| chr20_+_5414448 | 43.51 |
ENSRNOT00000078972
ENSRNOT00000080900 |
RT1-A1
|
RT1 class Ia, locus A1 |
| chr4_+_149261044 | 43.33 |
ENSRNOT00000066670
|
Cxcl12
|
C-X-C motif chemokine ligand 12 |
| chr2_-_189254628 | 42.52 |
ENSRNOT00000028234
|
Il6r
|
interleukin 6 receptor |
| chr8_-_85803433 | 42.36 |
ENSRNOT00000081544
ENSRNOT00000073286 |
Mb21d1
|
Mab-21 domain containing 1 |
| chr2_+_1410934 | 42.13 |
ENSRNOT00000013625
ENSRNOT00000080222 |
Erap1
|
endoplasmic reticulum aminopeptidase 1 |
| chr10_-_70744315 | 41.94 |
ENSRNOT00000014865
|
Ccl5
|
C-C motif chemokine ligand 5 |
| chr10_-_88611105 | 41.78 |
ENSRNOT00000024718
|
Dhx58
|
DEXH-box helicase 58 |
| chr4_-_163049084 | 41.24 |
ENSRNOT00000091644
|
Cd69
|
Cd69 molecule |
| chr20_-_4561062 | 41.19 |
ENSRNOT00000065044
ENSRNOT00000092698 ENSRNOT00000060607 |
Cfb
C2
|
complement factor B complement C2 |
| chr20_-_157665 | 41.14 |
ENSRNOT00000048858
ENSRNOT00000079494 |
RT1-CE10
|
RT1 class I, locus CE10 |
| chr4_-_163890801 | 41.13 |
ENSRNOT00000081946
|
Ly49si1
|
immunoreceptor Ly49si1 |
| chr15_+_34552410 | 40.96 |
ENSRNOT00000027802
|
Khnyn
|
KH and NYN domain containing |
| chr3_-_119405453 | 40.67 |
ENSRNOT00000090355
|
Sppl2a
|
signal peptide peptidase-like 2A |
| chr8_-_63034226 | 40.64 |
ENSRNOT00000043434
|
Pml
|
promyelocytic leukemia |
| chr5_+_157222636 | 40.62 |
ENSRNOT00000022579
|
Pla2g2d
|
phospholipase A2, group IID |
| chr4_+_14001761 | 40.49 |
ENSRNOT00000076519
|
Cd36
|
CD36 molecule |
| chr1_-_101236065 | 39.86 |
ENSRNOT00000066834
|
Cd37
|
CD37 molecule |
| chr15_+_24141651 | 39.78 |
ENSRNOT00000082304
|
Lgals3
|
galectin 3 |
| chr15_+_57241968 | 39.56 |
ENSRNOT00000082191
|
Lcp1
|
lymphocyte cytosolic protein 1 |
| chr4_+_70689737 | 39.23 |
ENSRNOT00000018852
|
Prss2
|
protease, serine, 2 |
| chr16_+_50016857 | 39.22 |
ENSRNOT00000035781
|
Tlr3
|
toll-like receptor 3 |
| chr4_-_21920651 | 38.97 |
ENSRNOT00000066211
|
Tmem243
|
transmembrane protein 243 |
| chr1_-_227505267 | 38.92 |
ENSRNOT00000028443
|
Ms4a7
|
membrane spanning 4-domains A7 |
| chr10_+_94944436 | 38.77 |
ENSRNOT00000078968
|
Milr1
|
mast cell immunoglobulin-like receptor 1 |
| chr7_-_140401686 | 38.73 |
ENSRNOT00000083955
|
Fkbp11
|
FK506 binding protein 11 |
| chr1_-_47331412 | 38.22 |
ENSRNOT00000046746
|
Ezr
|
ezrin |
| chr16_+_72388880 | 37.64 |
ENSRNOT00000072459
|
LOC684871
|
similar to Protein C8orf4 (Thyroid cancer protein 1) (TC-1) |
| chr20_-_4921348 | 37.59 |
ENSRNOT00000082497
ENSRNOT00000041151 |
RT1-CE4
|
RT1 class I, locus CE4 |
| chr4_+_62380914 | 37.39 |
ENSRNOT00000029845
|
Tmem140
|
transmembrane protein 140 |
| chrX_+_54390733 | 37.32 |
ENSRNOT00000004977
|
RGD1565785
|
similar to chromosome X open reading frame 21 |
| chr2_-_105089659 | 37.31 |
ENSRNOT00000043381
|
Cpb1
|
carboxypeptidase B1 |
| chr1_+_140602542 | 36.90 |
ENSRNOT00000085570
|
Isg20
|
interferon stimulated exonuclease gene 20 |
| chr15_-_34269851 | 36.66 |
ENSRNOT00000026279
|
Psme2
|
proteasome activator subunit 2 |
| chr20_+_3176107 | 36.24 |
ENSRNOT00000001036
|
RT1-S3
|
RT1 class Ib, locus S3 |
| chr3_-_160738927 | 36.06 |
ENSRNOT00000043470
|
Slpil2
|
antileukoproteinase-like 2 |
| chr20_+_3155652 | 35.41 |
ENSRNOT00000042882
|
RT1-S2
|
RT1 class Ib, locus S2 |
| chr1_-_73619356 | 35.22 |
ENSRNOT00000074352
|
Lilrb3
|
leukocyte immunoglobulin like receptor B3 |
| chr6_-_1466201 | 34.95 |
ENSRNOT00000089185
|
Eif2ak2
|
eukaryotic translation initiation factor 2-alpha kinase 2 |
| chr8_-_23146689 | 34.76 |
ENSRNOT00000092200
|
Acp5
|
acid phosphatase 5, tartrate resistant |
| chr20_+_44680449 | 34.71 |
ENSRNOT00000000728
|
Traf3ip2
|
Traf3 interacting protein 2 |
| chr3_-_160739137 | 34.45 |
ENSRNOT00000075836
|
Slpil2
|
antileukoproteinase-like 2 |
| chr5_+_79382096 | 34.33 |
ENSRNOT00000085461
|
Tmem268
|
transmembrane protein 268 |
| chr9_-_14668297 | 34.20 |
ENSRNOT00000042404
|
Treml2
|
triggering receptor expressed on myeloid cells-like 2 |
| chr2_+_186776644 | 34.13 |
ENSRNOT00000046778
|
Fcrl3
|
Fc receptor-like 3 |
| chr13_+_48426820 | 34.09 |
ENSRNOT00000048391
|
Ctse
|
cathepsin E |
| chr4_+_70776046 | 34.02 |
ENSRNOT00000040403
|
Prss1
|
protease, serine 1 |
| chr20_-_5179217 | 33.65 |
ENSRNOT00000065940
ENSRNOT00000092443 |
Lst1
|
leukocyte specific transcript 1 |
| chr19_+_15033108 | 33.28 |
ENSRNOT00000021812
|
Ces1d
|
carboxylesterase 1D |
| chr17_-_31780120 | 32.81 |
ENSRNOT00000058388
|
AABR07027450.1
|
|
| chr4_+_102351036 | 32.62 |
ENSRNOT00000079277
|
AABR07060994.1
|
|
| chr10_-_4257868 | 32.36 |
ENSRNOT00000035886
|
Tnfrsf17
|
TNF receptor superfamily member 17 |
| chr9_+_65534704 | 32.35 |
ENSRNOT00000016730
|
Cflar
|
CASP8 and FADD-like apoptosis regulator |
| chr19_+_15195565 | 32.33 |
ENSRNOT00000090865
ENSRNOT00000078874 |
Ces1d
|
carboxylesterase 1D |
| chr4_+_101687327 | 31.93 |
ENSRNOT00000082501
|
AABR07060957.1
|
|
| chr2_-_210550490 | 31.65 |
ENSRNOT00000081835
ENSRNOT00000025222 ENSRNOT00000086403 |
Csf1
|
colony stimulating factor 1 |
Network of associatons between targets according to the STRING database.
First level regulatory network of Irf2_Irf1_Irf8_Irf9_Irf7
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 81.6 | 244.8 | GO:0046968 | cytosol to ER transport(GO:0046967) peptide antigen transport(GO:0046968) |
| 70.9 | 283.7 | GO:1901423 | response to benzene(GO:1901423) |
| 61.9 | 185.6 | GO:0002731 | negative regulation of dendritic cell cytokine production(GO:0002731) |
| 61.5 | 122.9 | GO:0002481 | antigen processing and presentation of exogenous peptide antigen via MHC class Ib(GO:0002477) antigen processing and presentation of exogenous protein antigen via MHC class Ib, TAP-dependent(GO:0002481) |
| 58.1 | 232.5 | GO:0097026 | dendritic cell dendrite assembly(GO:0097026) |
| 57.8 | 173.3 | GO:0070212 | regulation of chromatin assembly(GO:0010847) protein poly-ADP-ribosylation(GO:0070212) |
| 53.8 | 269.1 | GO:0019885 | antigen processing and presentation of endogenous peptide antigen via MHC class I(GO:0019885) |
| 49.2 | 246.0 | GO:0032020 | ISG15-protein conjugation(GO:0032020) |
| 49.1 | 98.1 | GO:0002589 | regulation of antigen processing and presentation of peptide antigen via MHC class I(GO:0002589) |
| 41.0 | 123.1 | GO:1901740 | negative regulation of myoblast fusion(GO:1901740) |
| 37.9 | 340.7 | GO:0003373 | dynamin polymerization involved in membrane fission(GO:0003373) dynamin polymerization involved in mitochondrial fission(GO:0003374) |
| 37.4 | 112.1 | GO:0019677 | NAD catabolic process(GO:0019677) |
| 36.7 | 110.2 | GO:0000738 | DNA catabolic process, exonucleolytic(GO:0000738) |
| 36.7 | 256.6 | GO:0070269 | pyroptosis(GO:0070269) |
| 32.9 | 394.4 | GO:0060340 | positive regulation of type I interferon-mediated signaling pathway(GO:0060340) |
| 31.8 | 127.2 | GO:0019805 | quinolinate biosynthetic process(GO:0019805) |
| 31.5 | 94.6 | GO:1904845 | response to L-glutamine(GO:1904844) cellular response to L-glutamine(GO:1904845) |
| 31.0 | 1052.9 | GO:0035458 | cellular response to interferon-beta(GO:0035458) |
| 30.1 | 120.3 | GO:0090290 | positive regulation of osteoclast proliferation(GO:0090290) |
| 30.0 | 90.1 | GO:0035722 | interleukin-12-mediated signaling pathway(GO:0035722) cellular response to interleukin-12(GO:0071349) |
| 29.5 | 118.1 | GO:0045349 | interferon-alpha biosynthetic process(GO:0045349) regulation of interferon-alpha biosynthetic process(GO:0045354) |
| 27.6 | 165.4 | GO:0045345 | MHC class I biosynthetic process(GO:0045341) regulation of MHC class I biosynthetic process(GO:0045343) positive regulation of MHC class I biosynthetic process(GO:0045345) |
| 26.8 | 214.5 | GO:0032727 | positive regulation of interferon-alpha production(GO:0032727) |
| 26.3 | 131.3 | GO:2001181 | positive regulation of interleukin-10 secretion(GO:2001181) |
| 26.2 | 78.7 | GO:0034165 | positive regulation of toll-like receptor 9 signaling pathway(GO:0034165) |
| 21.9 | 65.6 | GO:0019626 | short-chain fatty acid catabolic process(GO:0019626) |
| 20.8 | 186.8 | GO:2000051 | negative regulation of non-canonical Wnt signaling pathway(GO:2000051) |
| 18.5 | 55.5 | GO:0003430 | growth plate cartilage chondrocyte growth(GO:0003430) |
| 18.3 | 54.9 | GO:2000502 | stimulatory C-type lectin receptor signaling pathway(GO:0002223) negative regulation of natural killer cell chemotaxis(GO:2000502) |
| 18.1 | 108.8 | GO:0035672 | oligopeptide transmembrane transport(GO:0035672) |
| 18.1 | 36.2 | GO:0002428 | antigen processing and presentation of peptide antigen via MHC class Ib(GO:0002428) |
| 17.5 | 577.5 | GO:0002474 | antigen processing and presentation of peptide antigen via MHC class I(GO:0002474) |
| 16.3 | 179.6 | GO:0046598 | positive regulation of viral entry into host cell(GO:0046598) |
| 16.2 | 113.2 | GO:0033634 | positive regulation of cell-cell adhesion mediated by integrin(GO:0033634) |
| 16.1 | 80.3 | GO:0072733 | response to staurosporine(GO:0072733) cellular response to staurosporine(GO:0072734) |
| 15.5 | 46.4 | GO:0045351 | type I interferon biosynthetic process(GO:0045351) |
| 15.5 | 61.8 | GO:0000414 | regulation of histone H3-K36 methylation(GO:0000414) |
| 15.3 | 122.1 | GO:0032621 | interleukin-18 production(GO:0032621) |
| 14.8 | 73.8 | GO:0042270 | protection from natural killer cell mediated cytotoxicity(GO:0042270) |
| 14.7 | 58.9 | GO:1903223 | positive regulation of oxidative stress-induced neuron death(GO:1903223) |
| 14.1 | 112.6 | GO:0070842 | aggresome assembly(GO:0070842) |
| 13.9 | 55.8 | GO:0045627 | positive regulation of T-helper 1 cell differentiation(GO:0045627) |
| 13.3 | 279.1 | GO:0035634 | response to stilbenoid(GO:0035634) |
| 13.3 | 39.9 | GO:0030886 | negative regulation of myeloid dendritic cell activation(GO:0030886) |
| 13.3 | 39.8 | GO:1903769 | negative regulation of cell proliferation in bone marrow(GO:1903769) |
| 12.4 | 62.1 | GO:0070120 | ciliary neurotrophic factor-mediated signaling pathway(GO:0070120) |
| 12.0 | 84.0 | GO:0032074 | negative regulation of nuclease activity(GO:0032074) |
| 11.7 | 46.9 | GO:0039534 | negative regulation of MDA-5 signaling pathway(GO:0039534) |
| 11.7 | 58.4 | GO:0035279 | mRNA cleavage involved in gene silencing by miRNA(GO:0035279) mRNA cleavage involved in gene silencing(GO:0098795) |
| 11.1 | 99.5 | GO:0060763 | mammary duct terminal end bud growth(GO:0060763) |
| 11.0 | 21.9 | GO:2000520 | regulation of immunological synapse formation(GO:2000520) |
| 10.8 | 43.3 | GO:1903237 | negative regulation of leukocyte tethering or rolling(GO:1903237) response to ultrasound(GO:1990478) |
| 10.5 | 62.8 | GO:0010266 | response to vitamin B1(GO:0010266) |
| 10.5 | 83.6 | GO:0050859 | negative regulation of B cell receptor signaling pathway(GO:0050859) |
| 10.3 | 30.8 | GO:0070946 | neutrophil mediated killing of gram-positive bacterium(GO:0070946) |
| 10.3 | 30.8 | GO:0060369 | positive regulation of Fc receptor mediated stimulatory signaling pathway(GO:0060369) |
| 10.2 | 30.6 | GO:0007356 | thorax and anterior abdomen determination(GO:0007356) |
| 10.2 | 30.5 | GO:2000501 | natural killer cell chemotaxis(GO:0035747) regulation of natural killer cell chemotaxis(GO:2000501) positive regulation of natural killer cell chemotaxis(GO:2000503) |
| 9.8 | 29.4 | GO:0070175 | positive regulation of enamel mineralization(GO:0070175) |
| 9.6 | 38.2 | GO:1902896 | terminal web assembly(GO:1902896) |
| 9.5 | 19.1 | GO:0001803 | type III hypersensitivity(GO:0001802) regulation of type III hypersensitivity(GO:0001803) positive regulation of type III hypersensitivity(GO:0001805) |
| 9.5 | 47.5 | GO:0051902 | negative regulation of mitochondrial depolarization(GO:0051902) |
| 9.5 | 19.0 | GO:0002584 | negative regulation of antigen processing and presentation of peptide antigen(GO:0002584) |
| 8.9 | 44.4 | GO:1900019 | regulation of protein kinase C activity(GO:1900019) positive regulation of protein kinase C activity(GO:1900020) |
| 8.8 | 26.3 | GO:0045625 | regulation of T-helper 1 cell differentiation(GO:0045625) |
| 8.6 | 94.8 | GO:0006957 | complement activation, alternative pathway(GO:0006957) |
| 8.5 | 144.7 | GO:0051044 | positive regulation of membrane protein ectodomain proteolysis(GO:0051044) |
| 8.5 | 85.0 | GO:0071357 | type I interferon signaling pathway(GO:0060337) cellular response to type I interferon(GO:0071357) |
| 8.5 | 42.4 | GO:0071360 | cellular response to exogenous dsRNA(GO:0071360) |
| 8.4 | 33.7 | GO:0050672 | negative regulation of mononuclear cell proliferation(GO:0032945) negative regulation of lymphocyte proliferation(GO:0050672) |
| 8.3 | 8.3 | GO:0051025 | negative regulation of immunoglobulin secretion(GO:0051025) |
| 8.1 | 16.2 | GO:2000646 | positive regulation of receptor catabolic process(GO:2000646) |
| 7.8 | 31.4 | GO:0016332 | establishment or maintenance of polarity of embryonic epithelium(GO:0016332) |
| 7.8 | 15.5 | GO:0097198 | histone H3-K36 trimethylation(GO:0097198) |
| 7.7 | 30.8 | GO:1902263 | apoptotic process involved in embryonic digit morphogenesis(GO:1902263) |
| 7.7 | 145.7 | GO:0019884 | antigen processing and presentation of exogenous antigen(GO:0019884) |
| 7.5 | 60.4 | GO:0002634 | regulation of germinal center formation(GO:0002634) |
| 7.5 | 7.5 | GO:1902623 | negative regulation of neutrophil migration(GO:1902623) |
| 7.5 | 59.7 | GO:0033590 | response to cobalamin(GO:0033590) |
| 7.5 | 164.0 | GO:0010390 | histone monoubiquitination(GO:0010390) |
| 7.5 | 29.8 | GO:0030221 | basophil differentiation(GO:0030221) |
| 7.3 | 58.3 | GO:0016584 | nucleosome positioning(GO:0016584) |
| 7.2 | 86.9 | GO:0006309 | apoptotic DNA fragmentation(GO:0006309) |
| 7.0 | 21.1 | GO:0051105 | regulation of DNA ligation(GO:0051105) positive regulation of DNA ligation(GO:0051106) protein localization to site of double-strand break(GO:1990166) |
| 7.0 | 35.0 | GO:1903300 | negative regulation of glucokinase activity(GO:0033132) negative regulation of hexokinase activity(GO:1903300) |
| 6.9 | 20.7 | GO:0002149 | hypochlorous acid metabolic process(GO:0002148) hypochlorous acid biosynthetic process(GO:0002149) |
| 6.7 | 94.3 | GO:0010818 | T cell chemotaxis(GO:0010818) |
| 6.7 | 33.6 | GO:0006556 | S-adenosylmethionine biosynthetic process(GO:0006556) |
| 6.6 | 26.3 | GO:1990375 | baculum development(GO:1990375) |
| 6.4 | 51.2 | GO:0018377 | protein myristoylation(GO:0018377) |
| 6.3 | 25.3 | GO:0097527 | necroptotic signaling pathway(GO:0097527) |
| 6.2 | 18.6 | GO:0060414 | aorta smooth muscle tissue morphogenesis(GO:0060414) |
| 6.1 | 54.5 | GO:1900165 | negative regulation of interleukin-6 secretion(GO:1900165) |
| 6.0 | 229.6 | GO:0045071 | negative regulation of viral genome replication(GO:0045071) |
| 5.9 | 5.9 | GO:1905072 | cardiac jelly development(GO:1905072) |
| 5.8 | 34.8 | GO:0032929 | negative regulation of superoxide anion generation(GO:0032929) |
| 5.8 | 34.7 | GO:0002230 | positive regulation of defense response to virus by host(GO:0002230) |
| 5.7 | 62.7 | GO:0031293 | membrane protein intracellular domain proteolysis(GO:0031293) |
| 5.7 | 17.1 | GO:1905204 | response to selenite ion(GO:0072714) negative regulation of connective tissue replacement(GO:1905204) |
| 5.7 | 17.0 | GO:0036378 | calcitriol biosynthetic process from calciol(GO:0036378) |
| 5.6 | 28.0 | GO:0016098 | monoterpenoid metabolic process(GO:0016098) |
| 5.6 | 16.8 | GO:0033625 | positive regulation of integrin activation(GO:0033625) |
| 5.6 | 5.6 | GO:0018307 | enzyme active site formation(GO:0018307) |
| 5.5 | 71.3 | GO:1902187 | negative regulation of viral release from host cell(GO:1902187) |
| 5.3 | 53.2 | GO:1900115 | extracellular regulation of signal transduction(GO:1900115) extracellular negative regulation of signal transduction(GO:1900116) |
| 5.2 | 10.4 | GO:0070634 | transepithelial ammonium transport(GO:0070634) |
| 5.1 | 20.5 | GO:1903463 | regulation of mitotic cell cycle DNA replication(GO:1903463) |
| 4.8 | 24.2 | GO:2000327 | positive regulation of ligand-dependent nuclear receptor transcription coactivator activity(GO:2000327) |
| 4.8 | 28.7 | GO:0072383 | plus-end-directed vesicle transport along microtubule(GO:0072383) plus-end-directed organelle transport along microtubule(GO:0072386) |
| 4.8 | 57.3 | GO:0036462 | TRAIL-activated apoptotic signaling pathway(GO:0036462) |
| 4.6 | 13.9 | GO:0045590 | negative regulation of regulatory T cell differentiation(GO:0045590) |
| 4.6 | 23.0 | GO:0051919 | positive regulation of fibrinolysis(GO:0051919) |
| 4.6 | 18.3 | GO:0060155 | platelet dense granule organization(GO:0060155) |
| 4.6 | 18.3 | GO:0007296 | vitellogenesis(GO:0007296) |
| 4.5 | 13.5 | GO:0061357 | positive regulation of Wnt protein secretion(GO:0061357) |
| 4.4 | 13.1 | GO:1903544 | positive regulation of bicellular tight junction assembly(GO:1903348) response to butyrate(GO:1903544) |
| 4.3 | 17.4 | GO:0043987 | histone H3-S10 phosphorylation(GO:0043987) |
| 4.3 | 17.1 | GO:0001692 | histamine metabolic process(GO:0001692) positive regulation of interleukin-8 biosynthetic process(GO:0045416) |
| 4.3 | 34.1 | GO:1902916 | positive regulation of protein polyubiquitination(GO:1902916) |
| 4.0 | 16.2 | GO:0000320 | re-entry into mitotic cell cycle(GO:0000320) |
| 3.9 | 3.9 | GO:0045077 | negative regulation of interferon-gamma biosynthetic process(GO:0045077) |
| 3.9 | 11.8 | GO:0032066 | nucleolus to nucleoplasm transport(GO:0032066) |
| 3.8 | 11.4 | GO:1903719 | regulation of I-kappaB phosphorylation(GO:1903719) positive regulation of I-kappaB phosphorylation(GO:1903721) |
| 3.8 | 7.5 | GO:0003330 | regulation of extracellular matrix constituent secretion(GO:0003330) positive regulation of extracellular matrix constituent secretion(GO:0003331) |
| 3.7 | 11.2 | GO:0003249 | left ventricular cardiac muscle tissue morphogenesis(GO:0003220) cell proliferation involved in heart valve morphogenesis(GO:0003249) regulation of cell proliferation involved in heart valve morphogenesis(GO:0003250) gonad morphogenesis(GO:0035262) nephrogenic mesenchyme morphogenesis(GO:0072134) |
| 3.6 | 10.9 | GO:0042631 | cellular response to water deprivation(GO:0042631) |
| 3.6 | 25.1 | GO:0048245 | eosinophil chemotaxis(GO:0048245) |
| 3.6 | 10.7 | GO:0046947 | hydroxylysine metabolic process(GO:0046946) hydroxylysine biosynthetic process(GO:0046947) |
| 3.6 | 53.5 | GO:0042832 | defense response to protozoan(GO:0042832) |
| 3.5 | 7.0 | GO:1902202 | regulation of hepatocyte growth factor receptor signaling pathway(GO:1902202) |
| 3.5 | 31.3 | GO:0046826 | negative regulation of protein export from nucleus(GO:0046826) |
| 3.5 | 10.4 | GO:0007161 | calcium-independent cell-matrix adhesion(GO:0007161) interaction with other organism via secreted substance involved in symbiotic interaction(GO:0052047) regulation of transforming growth factor-beta secretion(GO:2001201) |
| 3.5 | 159.8 | GO:0019882 | antigen processing and presentation(GO:0019882) |
| 3.4 | 27.6 | GO:2000406 | positive regulation of T cell migration(GO:2000406) |
| 3.4 | 40.6 | GO:0002361 | CD4-positive, CD25-positive, alpha-beta regulatory T cell differentiation(GO:0002361) |
| 3.3 | 10.0 | GO:0035444 | nickel cation transport(GO:0015675) vanadium ion transport(GO:0015676) lead ion transport(GO:0015692) nickel cation transmembrane transport(GO:0035444) |
| 3.3 | 46.3 | GO:0001867 | complement activation, lectin pathway(GO:0001867) |
| 3.3 | 69.3 | GO:0006491 | N-glycan processing(GO:0006491) |
| 3.2 | 9.7 | GO:1901492 | positive regulation of lymphangiogenesis(GO:1901492) |
| 3.1 | 9.4 | GO:0000715 | nucleotide-excision repair, DNA damage recognition(GO:0000715) |
| 3.1 | 12.4 | GO:0080154 | regulation of fertilization(GO:0080154) |
| 3.0 | 12.2 | GO:0002188 | translation reinitiation(GO:0002188) |
| 3.0 | 60.0 | GO:0032689 | negative regulation of interferon-gamma production(GO:0032689) |
| 3.0 | 38.8 | GO:0033004 | negative regulation of mast cell activation(GO:0033004) |
| 3.0 | 65.5 | GO:0030574 | collagen catabolic process(GO:0030574) |
| 3.0 | 17.8 | GO:0001835 | blastocyst hatching(GO:0001835) hatching(GO:0035188) organism emergence from protective structure(GO:0071684) |
| 3.0 | 3.0 | GO:0038172 | interleukin-33-mediated signaling pathway(GO:0038172) |
| 2.9 | 14.6 | GO:0071409 | cellular response to cycloheximide(GO:0071409) |
| 2.9 | 8.6 | GO:0033184 | positive regulation of histone ubiquitination(GO:0033184) |
| 2.8 | 8.5 | GO:0090191 | negative regulation of branching involved in ureteric bud morphogenesis(GO:0090191) |
| 2.8 | 8.5 | GO:2001245 | regulation of phosphatidylcholine biosynthetic process(GO:2001245) |
| 2.8 | 11.2 | GO:0006272 | leading strand elongation(GO:0006272) |
| 2.8 | 8.3 | GO:0033306 | phytol metabolic process(GO:0033306) fatty alcohol metabolic process(GO:1903173) |
| 2.8 | 16.6 | GO:0090367 | negative regulation of mRNA modification(GO:0090367) |
| 2.7 | 5.4 | GO:1905146 | lysosomal protein catabolic process(GO:1905146) |
| 2.6 | 10.6 | GO:1990481 | mRNA pseudouridine synthesis(GO:1990481) |
| 2.6 | 2.6 | GO:0003104 | positive regulation of glomerular filtration(GO:0003104) |
| 2.6 | 7.8 | GO:0032380 | regulation of intracellular lipid transport(GO:0032377) regulation of intracellular sterol transport(GO:0032380) regulation of intracellular cholesterol transport(GO:0032383) |
| 2.6 | 10.4 | GO:0070102 | interleukin-6-mediated signaling pathway(GO:0070102) |
| 2.6 | 46.5 | GO:0002820 | negative regulation of adaptive immune response(GO:0002820) |
| 2.6 | 7.7 | GO:0090271 | positive regulation of fibroblast growth factor production(GO:0090271) |
| 2.6 | 7.7 | GO:0042706 | eye photoreceptor cell fate commitment(GO:0042706) photoreceptor cell fate commitment(GO:0046552) |
| 2.5 | 10.1 | GO:0035655 | interleukin-18-mediated signaling pathway(GO:0035655) cellular response to interleukin-18(GO:0071351) |
| 2.5 | 10.1 | GO:0045916 | negative regulation of complement activation(GO:0045916) negative regulation of protein activation cascade(GO:2000258) |
| 2.5 | 7.4 | GO:0036451 | cap mRNA methylation(GO:0036451) |
| 2.5 | 7.4 | GO:0036112 | medium-chain fatty-acyl-CoA metabolic process(GO:0036112) |
| 2.4 | 14.7 | GO:0050862 | positive regulation of T cell receptor signaling pathway(GO:0050862) |
| 2.4 | 4.8 | GO:0010637 | negative regulation of mitochondrial fusion(GO:0010637) |
| 2.4 | 36.0 | GO:0050832 | defense response to fungus(GO:0050832) |
| 2.4 | 26.3 | GO:0032897 | negative regulation of viral transcription(GO:0032897) |
| 2.3 | 7.0 | GO:0006045 | N-acetylglucosamine biosynthetic process(GO:0006045) N-acetylneuraminate biosynthetic process(GO:0046380) |
| 2.3 | 126.2 | GO:0043124 | negative regulation of I-kappaB kinase/NF-kappaB signaling(GO:0043124) |
| 2.3 | 9.0 | GO:0046098 | guanine metabolic process(GO:0046098) |
| 2.2 | 29.2 | GO:0051764 | actin crosslink formation(GO:0051764) |
| 2.2 | 6.6 | GO:0071461 | cellular response to redox state(GO:0071461) |
| 2.2 | 6.6 | GO:0090361 | platelet-derived growth factor production(GO:0090360) regulation of platelet-derived growth factor production(GO:0090361) |
| 2.2 | 21.9 | GO:0019720 | Mo-molybdopterin cofactor biosynthetic process(GO:0006777) Mo-molybdopterin cofactor metabolic process(GO:0019720) |
| 2.2 | 10.9 | GO:0051987 | positive regulation of attachment of spindle microtubules to kinetochore(GO:0051987) |
| 2.2 | 8.6 | GO:0032919 | spermine acetylation(GO:0032919) |
| 2.2 | 6.5 | GO:1902748 | positive regulation of lens fiber cell differentiation(GO:1902748) |
| 2.1 | 6.4 | GO:0046726 | positive regulation by virus of viral protein levels in host cell(GO:0046726) |
| 2.1 | 156.0 | GO:0048286 | lung alveolus development(GO:0048286) |
| 2.0 | 6.1 | GO:0097298 | regulation of nucleus size(GO:0097298) |
| 2.0 | 202.4 | GO:0051607 | defense response to virus(GO:0051607) |
| 2.0 | 8.1 | GO:0071284 | cellular response to lead ion(GO:0071284) |
| 2.0 | 8.0 | GO:0046292 | formaldehyde metabolic process(GO:0046292) |
| 2.0 | 8.0 | GO:0019477 | lysine catabolic process(GO:0006554) L-lysine catabolic process to acetyl-CoA(GO:0019474) L-lysine catabolic process(GO:0019477) L-lysine metabolic process(GO:0046440) |
| 2.0 | 17.8 | GO:0042699 | follicle-stimulating hormone signaling pathway(GO:0042699) |
| 1.9 | 17.5 | GO:0002430 | complement receptor mediated signaling pathway(GO:0002430) |
| 1.9 | 25.1 | GO:0051601 | exocyst localization(GO:0051601) |
| 1.9 | 26.8 | GO:0010838 | positive regulation of keratinocyte proliferation(GO:0010838) |
| 1.9 | 5.7 | GO:0045716 | positive regulation of low-density lipoprotein particle receptor biosynthetic process(GO:0045716) |
| 1.9 | 24.6 | GO:0016446 | somatic diversification of immune receptors via somatic mutation(GO:0002566) somatic hypermutation of immunoglobulin genes(GO:0016446) |
| 1.8 | 31.2 | GO:0048242 | epinephrine secretion(GO:0048242) |
| 1.8 | 7.3 | GO:0018214 | peptidyl-glutamic acid carboxylation(GO:0017187) protein carboxylation(GO:0018214) |
| 1.8 | 3.6 | GO:0033685 | negative regulation of luteinizing hormone secretion(GO:0033685) |
| 1.8 | 8.9 | GO:0015838 | amino-acid betaine transport(GO:0015838) |
| 1.8 | 5.3 | GO:0033386 | geranylgeranyl diphosphate metabolic process(GO:0033385) geranylgeranyl diphosphate biosynthetic process(GO:0033386) |
| 1.8 | 14.1 | GO:0042994 | cytoplasmic sequestering of transcription factor(GO:0042994) |
| 1.7 | 20.9 | GO:0035589 | G-protein coupled purinergic nucleotide receptor signaling pathway(GO:0035589) |
| 1.7 | 12.0 | GO:0002326 | B cell lineage commitment(GO:0002326) immunoglobulin V(D)J recombination(GO:0033152) |
| 1.7 | 6.8 | GO:1903385 | regulation of homophilic cell adhesion(GO:1903385) |
| 1.7 | 1.7 | GO:2000562 | negative regulation of CD4-positive, alpha-beta T cell proliferation(GO:2000562) |
| 1.7 | 5.0 | GO:0043091 | L-arginine import(GO:0043091) arginine import(GO:0090467) L-arginine transport(GO:1902023) |
| 1.6 | 102.0 | GO:0007004 | RNA-dependent DNA biosynthetic process(GO:0006278) telomere maintenance via telomerase(GO:0007004) |
| 1.6 | 24.3 | GO:0042448 | progesterone metabolic process(GO:0042448) |
| 1.6 | 19.2 | GO:0006958 | complement activation, classical pathway(GO:0006958) |
| 1.5 | 19.9 | GO:0002227 | innate immune response in mucosa(GO:0002227) |
| 1.5 | 7.6 | GO:0010032 | meiotic chromosome condensation(GO:0010032) |
| 1.5 | 31.8 | GO:0016578 | histone deubiquitination(GO:0016578) |
| 1.5 | 6.0 | GO:1903576 | response to L-arginine(GO:1903576) |
| 1.5 | 10.2 | GO:0015722 | canalicular bile acid transport(GO:0015722) |
| 1.4 | 2.8 | GO:2000857 | positive regulation of mineralocorticoid secretion(GO:2000857) positive regulation of aldosterone secretion(GO:2000860) |
| 1.4 | 27.2 | GO:0071108 | protein K48-linked deubiquitination(GO:0071108) |
| 1.3 | 5.4 | GO:0021785 | branchiomotor neuron axon guidance(GO:0021785) |
| 1.3 | 9.2 | GO:0097428 | protein maturation by iron-sulfur cluster transfer(GO:0097428) |
| 1.3 | 9.2 | GO:0030579 | ubiquitin-dependent SMAD protein catabolic process(GO:0030579) |
| 1.3 | 5.2 | GO:0043328 | protein targeting to vacuole involved in ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway(GO:0043328) |
| 1.3 | 11.7 | GO:0046501 | protoporphyrinogen IX metabolic process(GO:0046501) |
| 1.3 | 10.3 | GO:0019388 | galactose catabolic process(GO:0019388) |
| 1.3 | 13.9 | GO:0032728 | positive regulation of interferon-beta production(GO:0032728) |
| 1.2 | 11.1 | GO:0043402 | glucocorticoid mediated signaling pathway(GO:0043402) |
| 1.2 | 3.6 | GO:2000853 | negative regulation of corticosterone secretion(GO:2000853) |
| 1.2 | 22.5 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 1.2 | 3.5 | GO:0043605 | cellular amide catabolic process(GO:0043605) |
| 1.2 | 14.0 | GO:0035814 | negative regulation of renal sodium excretion(GO:0035814) |
| 1.2 | 141.2 | GO:0042157 | lipoprotein metabolic process(GO:0042157) |
| 1.2 | 9.3 | GO:0019372 | lipoxygenase pathway(GO:0019372) |
| 1.2 | 2.3 | GO:0035789 | cell migration involved in metanephros development(GO:0035788) metanephric mesenchymal cell migration(GO:0035789) positive regulation of metanephric mesenchymal cell migration by platelet-derived growth factor receptor-beta signaling pathway(GO:0035793) metanephric glomerular mesangial cell proliferation involved in metanephros development(GO:0072262) regulation of metanephric mesenchymal cell migration by platelet-derived growth factor receptor-beta signaling pathway(GO:1900238) regulation of metanephric mesenchymal cell migration(GO:2000589) positive regulation of metanephric mesenchymal cell migration(GO:2000591) |
| 1.1 | 3.4 | GO:0070914 | UV-damage excision repair(GO:0070914) |
| 1.1 | 10.9 | GO:0051151 | negative regulation of smooth muscle cell differentiation(GO:0051151) |
| 1.1 | 3.3 | GO:0090521 | glomerular visceral epithelial cell migration(GO:0090521) |
| 1.1 | 9.6 | GO:0032000 | positive regulation of fatty acid beta-oxidation(GO:0032000) |
| 1.1 | 15.9 | GO:0043312 | neutrophil degranulation(GO:0043312) |
| 1.0 | 5.2 | GO:1902951 | negative regulation of dendritic spine maintenance(GO:1902951) |
| 1.0 | 55.7 | GO:0070206 | protein trimerization(GO:0070206) |
| 1.0 | 6.2 | GO:0006449 | regulation of translational termination(GO:0006449) |
| 1.0 | 6.2 | GO:0031087 | deadenylation-independent decapping of nuclear-transcribed mRNA(GO:0031087) |
| 1.0 | 6.1 | GO:0019919 | peptidyl-arginine methylation, to asymmetrical-dimethyl arginine(GO:0019919) |
| 1.0 | 3.1 | GO:0030208 | dermatan sulfate biosynthetic process(GO:0030208) |
| 1.0 | 3.0 | GO:0019343 | cysteine biosynthetic process via cystathionine(GO:0019343) |
| 1.0 | 9.9 | GO:0033539 | fatty acid beta-oxidation using acyl-CoA dehydrogenase(GO:0033539) |
| 1.0 | 22.6 | GO:0046825 | regulation of protein export from nucleus(GO:0046825) |
| 1.0 | 7.8 | GO:0015712 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 1.0 | 70.2 | GO:0034440 | fatty acid oxidation(GO:0019395) lipid oxidation(GO:0034440) |
| 1.0 | 31.0 | GO:0010718 | positive regulation of epithelial to mesenchymal transition(GO:0010718) |
| 1.0 | 5.8 | GO:0006398 | mRNA 3'-end processing by stem-loop binding and cleavage(GO:0006398) |
| 0.9 | 3.7 | GO:0006369 | termination of RNA polymerase II transcription(GO:0006369) |
| 0.9 | 3.7 | GO:1902683 | regulation of receptor localization to synapse(GO:1902683) |
| 0.9 | 0.9 | GO:0009730 | detection of carbohydrate stimulus(GO:0009730) detection of hexose stimulus(GO:0009732) detection of monosaccharide stimulus(GO:0034287) detection of glucose(GO:0051594) |
| 0.9 | 4.6 | GO:0003383 | apical constriction(GO:0003383) |
| 0.9 | 17.7 | GO:0045671 | negative regulation of osteoclast differentiation(GO:0045671) |
| 0.9 | 13.1 | GO:0042407 | cristae formation(GO:0042407) |
| 0.9 | 6.8 | GO:0035021 | negative regulation of Rac protein signal transduction(GO:0035021) |
| 0.8 | 31.1 | GO:0002224 | toll-like receptor signaling pathway(GO:0002224) |
| 0.8 | 14.5 | GO:0046415 | urate metabolic process(GO:0046415) |
| 0.8 | 5.5 | GO:0035988 | chondrocyte proliferation(GO:0035988) |
| 0.8 | 6.3 | GO:1900113 | negative regulation of histone H3-K9 trimethylation(GO:1900113) |
| 0.8 | 12.4 | GO:0045653 | negative regulation of megakaryocyte differentiation(GO:0045653) |
| 0.8 | 17.8 | GO:0071353 | cellular response to interleukin-4(GO:0071353) |
| 0.8 | 13.1 | GO:0010971 | positive regulation of G2/M transition of mitotic cell cycle(GO:0010971) |
| 0.8 | 6.1 | GO:0042795 | snRNA transcription from RNA polymerase II promoter(GO:0042795) |
| 0.8 | 4.6 | GO:2000400 | positive regulation of T cell differentiation in thymus(GO:0033089) positive regulation of thymocyte aggregation(GO:2000400) |
| 0.8 | 29.7 | GO:0002260 | lymphocyte homeostasis(GO:0002260) |
| 0.8 | 15.2 | GO:0007095 | mitotic G2 DNA damage checkpoint(GO:0007095) G2 DNA damage checkpoint(GO:0031572) |
| 0.7 | 36.5 | GO:0050715 | positive regulation of cytokine secretion(GO:0050715) |
| 0.7 | 8.2 | GO:0006337 | nucleosome disassembly(GO:0006337) |
| 0.7 | 10.3 | GO:0017121 | phospholipid scrambling(GO:0017121) |
| 0.7 | 13.1 | GO:0072520 | seminiferous tubule development(GO:0072520) |
| 0.7 | 26.3 | GO:0071526 | semaphorin-plexin signaling pathway(GO:0071526) |
| 0.7 | 5.7 | GO:0051315 | attachment of mitotic spindle microtubules to kinetochore(GO:0051315) |
| 0.7 | 5.0 | GO:0071373 | cellular response to luteinizing hormone stimulus(GO:0071373) |
| 0.7 | 39.6 | GO:0007157 | heterophilic cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0007157) |
| 0.7 | 22.5 | GO:0030101 | natural killer cell activation(GO:0030101) |
| 0.7 | 18.2 | GO:0006270 | DNA replication initiation(GO:0006270) |
| 0.7 | 10.3 | GO:0006622 | protein targeting to lysosome(GO:0006622) |
| 0.7 | 18.6 | GO:0000027 | ribosomal large subunit assembly(GO:0000027) |
| 0.7 | 4.7 | GO:0006307 | DNA dealkylation involved in DNA repair(GO:0006307) |
| 0.7 | 2.7 | GO:0090234 | regulation of kinetochore assembly(GO:0090234) |
| 0.7 | 10.6 | GO:0060992 | response to fungicide(GO:0060992) |
| 0.7 | 2.0 | GO:0032825 | positive regulation of natural killer cell differentiation(GO:0032825) |
| 0.6 | 8.3 | GO:0009263 | deoxyribonucleotide biosynthetic process(GO:0009263) |
| 0.6 | 25.1 | GO:0019915 | lipid storage(GO:0019915) |
| 0.6 | 55.8 | GO:0042098 | T cell proliferation(GO:0042098) |
| 0.6 | 26.7 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.6 | 13.5 | GO:0060216 | definitive hemopoiesis(GO:0060216) |
| 0.6 | 7.3 | GO:0060218 | hematopoietic stem cell differentiation(GO:0060218) |
| 0.6 | 4.7 | GO:0006729 | tetrahydrobiopterin biosynthetic process(GO:0006729) |
| 0.6 | 3.5 | GO:0030950 | establishment or maintenance of actin cytoskeleton polarity(GO:0030950) |
| 0.6 | 8.1 | GO:0060124 | positive regulation of growth hormone secretion(GO:0060124) |
| 0.6 | 18.3 | GO:0006695 | cholesterol biosynthetic process(GO:0006695) |
| 0.5 | 2.7 | GO:0010990 | regulation of SMAD protein complex assembly(GO:0010990) negative regulation of SMAD protein complex assembly(GO:0010991) |
| 0.5 | 8.5 | GO:0097286 | iron ion import(GO:0097286) |
| 0.5 | 3.2 | GO:0060965 | negative regulation of gene silencing by miRNA(GO:0060965) |
| 0.5 | 3.2 | GO:0090435 | protein localization to nuclear envelope(GO:0090435) |
| 0.5 | 2.1 | GO:1904158 | axonemal central apparatus assembly(GO:1904158) |
| 0.5 | 6.9 | GO:0010839 | negative regulation of keratinocyte proliferation(GO:0010839) |
| 0.5 | 11.6 | GO:0044783 | mitotic G1 DNA damage checkpoint(GO:0031571) G1 DNA damage checkpoint(GO:0044783) |
| 0.5 | 12.7 | GO:1900026 | positive regulation of substrate adhesion-dependent cell spreading(GO:1900026) |
| 0.5 | 4.5 | GO:0048820 | hair follicle maturation(GO:0048820) |
| 0.5 | 84.9 | GO:0045087 | innate immune response(GO:0045087) |
| 0.5 | 12.5 | GO:0000188 | inactivation of MAPK activity(GO:0000188) |
| 0.5 | 5.5 | GO:0006388 | tRNA splicing, via endonucleolytic cleavage and ligation(GO:0006388) |
| 0.5 | 1.5 | GO:0002329 | pre-B cell differentiation(GO:0002329) |
| 0.5 | 4.5 | GO:0021707 | cerebellar granular layer formation(GO:0021684) cerebellar granule cell differentiation(GO:0021707) |
| 0.5 | 26.3 | GO:0019319 | hexose biosynthetic process(GO:0019319) |
| 0.5 | 3.8 | GO:0007000 | nucleolus organization(GO:0007000) |
| 0.5 | 4.8 | GO:2000234 | positive regulation of ribosome biogenesis(GO:0090070) positive regulation of rRNA processing(GO:2000234) |
| 0.5 | 8.6 | GO:0006336 | DNA replication-independent nucleosome assembly(GO:0006336) |
| 0.5 | 1.9 | GO:0061484 | hematopoietic stem cell homeostasis(GO:0061484) |
| 0.5 | 4.7 | GO:0006308 | DNA catabolic process(GO:0006308) |
| 0.5 | 1.8 | GO:0070966 | nuclear-transcribed mRNA catabolic process, no-go decay(GO:0070966) |
| 0.4 | 2.6 | GO:0045196 | establishment or maintenance of neuroblast polarity(GO:0045196) establishment of neuroblast polarity(GO:0045200) |
| 0.4 | 3.7 | GO:2000641 | regulation of early endosome to late endosome transport(GO:2000641) |
| 0.4 | 6.4 | GO:0050830 | defense response to Gram-positive bacterium(GO:0050830) |
| 0.4 | 11.3 | GO:0070098 | chemokine-mediated signaling pathway(GO:0070098) |
| 0.4 | 6.8 | GO:0009404 | toxin metabolic process(GO:0009404) |
| 0.4 | 13.2 | GO:0050873 | brown fat cell differentiation(GO:0050873) |
| 0.4 | 3.6 | GO:0048280 | vesicle fusion with Golgi apparatus(GO:0048280) |
| 0.4 | 5.1 | GO:0031163 | iron-sulfur cluster assembly(GO:0016226) metallo-sulfur cluster assembly(GO:0031163) |
| 0.4 | 4.1 | GO:0033137 | negative regulation of peptidyl-serine phosphorylation(GO:0033137) |
| 0.4 | 5.2 | GO:0032801 | receptor catabolic process(GO:0032801) |
| 0.4 | 6.3 | GO:0006509 | membrane protein ectodomain proteolysis(GO:0006509) |
| 0.4 | 6.3 | GO:0034067 | protein localization to Golgi apparatus(GO:0034067) |
| 0.3 | 20.3 | GO:0002250 | adaptive immune response(GO:0002250) |
| 0.3 | 75.8 | GO:0010951 | negative regulation of endopeptidase activity(GO:0010951) |
| 0.3 | 12.4 | GO:0042149 | cellular response to glucose starvation(GO:0042149) |
| 0.3 | 7.1 | GO:0045292 | mRNA cis splicing, via spliceosome(GO:0045292) |
| 0.3 | 12.9 | GO:0000462 | maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000462) |
| 0.3 | 10.5 | GO:1903317 | regulation of protein processing(GO:0070613) regulation of protein maturation(GO:1903317) |
| 0.3 | 10.4 | GO:0006767 | water-soluble vitamin metabolic process(GO:0006767) |
| 0.3 | 0.3 | GO:0007493 | endodermal cell fate determination(GO:0007493) |
| 0.3 | 20.4 | GO:0006006 | glucose metabolic process(GO:0006006) |
| 0.3 | 2.8 | GO:0005981 | regulation of glycogen catabolic process(GO:0005981) |
| 0.3 | 4.3 | GO:0060746 | parental behavior(GO:0060746) |
| 0.3 | 10.1 | GO:0007274 | neuromuscular synaptic transmission(GO:0007274) |
| 0.3 | 3.0 | GO:0033148 | positive regulation of intracellular estrogen receptor signaling pathway(GO:0033148) |
| 0.3 | 1.8 | GO:0051255 | spindle midzone assembly(GO:0051255) |
| 0.3 | 4.5 | GO:0014877 | response to muscle inactivity involved in regulation of muscle adaptation(GO:0014877) response to denervation involved in regulation of muscle adaptation(GO:0014894) |
| 0.3 | 15.4 | GO:0019228 | neuronal action potential(GO:0019228) |
| 0.3 | 2.6 | GO:0030575 | nuclear body organization(GO:0030575) |
| 0.3 | 2.3 | GO:2000001 | regulation of DNA damage checkpoint(GO:2000001) |
| 0.3 | 53.5 | GO:0006898 | receptor-mediated endocytosis(GO:0006898) |
| 0.3 | 3.6 | GO:0007256 | activation of JNKK activity(GO:0007256) |
| 0.3 | 8.9 | GO:0000387 | spliceosomal snRNP assembly(GO:0000387) |
| 0.3 | 1.1 | GO:0035992 | tendon cell differentiation(GO:0035990) tendon formation(GO:0035992) |
| 0.3 | 1.9 | GO:0045648 | positive regulation of erythrocyte differentiation(GO:0045648) |
| 0.3 | 14.2 | GO:0006334 | nucleosome assembly(GO:0006334) |
| 0.3 | 6.4 | GO:0045638 | negative regulation of myeloid cell differentiation(GO:0045638) |
| 0.3 | 19.9 | GO:0034968 | histone lysine methylation(GO:0034968) |
| 0.3 | 1.3 | GO:0032262 | pyrimidine ribonucleotide salvage(GO:0010138) pyrimidine nucleotide salvage(GO:0032262) UMP salvage(GO:0044206) |
| 0.2 | 5.0 | GO:0009250 | glycogen biosynthetic process(GO:0005978) glucan biosynthetic process(GO:0009250) |
| 0.2 | 16.8 | GO:0090630 | activation of GTPase activity(GO:0090630) |
| 0.2 | 6.1 | GO:0033119 | negative regulation of RNA splicing(GO:0033119) |
| 0.2 | 13.9 | GO:2000177 | regulation of neural precursor cell proliferation(GO:2000177) |
| 0.2 | 2.9 | GO:0042574 | retinal metabolic process(GO:0042574) |
| 0.2 | 6.5 | GO:0060976 | coronary vasculature development(GO:0060976) |
| 0.2 | 14.6 | GO:0051209 | release of sequestered calcium ion into cytosol(GO:0051209) negative regulation of sequestering of calcium ion(GO:0051283) |
| 0.2 | 1.9 | GO:0048026 | positive regulation of mRNA splicing, via spliceosome(GO:0048026) |
| 0.2 | 1.2 | GO:0006689 | ganglioside catabolic process(GO:0006689) |
| 0.2 | 1.2 | GO:0001927 | exocyst assembly(GO:0001927) entry of bacterium into host cell(GO:0035635) regulation of entry of bacterium into host cell(GO:2000535) |
| 0.2 | 5.4 | GO:0048662 | negative regulation of smooth muscle cell proliferation(GO:0048662) |
| 0.2 | 0.5 | GO:2000398 | regulation of T cell differentiation in thymus(GO:0033081) regulation of thymocyte aggregation(GO:2000398) |
| 0.2 | 7.0 | GO:0034446 | substrate adhesion-dependent cell spreading(GO:0034446) |
| 0.2 | 7.4 | GO:0006998 | nuclear envelope organization(GO:0006998) |
| 0.2 | 4.1 | GO:0006879 | cellular iron ion homeostasis(GO:0006879) |
| 0.2 | 0.3 | GO:0034374 | low-density lipoprotein particle remodeling(GO:0034374) |
| 0.2 | 2.9 | GO:0031648 | protein destabilization(GO:0031648) |
| 0.2 | 2.4 | GO:0046519 | sphingoid metabolic process(GO:0046519) |
| 0.2 | 6.0 | GO:0009615 | response to virus(GO:0009615) |
| 0.2 | 7.9 | GO:0031016 | pancreas development(GO:0031016) |
| 0.1 | 53.3 | GO:0043547 | positive regulation of GTPase activity(GO:0043547) |
| 0.1 | 3.6 | GO:0030520 | intracellular estrogen receptor signaling pathway(GO:0030520) |
| 0.1 | 2.5 | GO:0050918 | positive chemotaxis(GO:0050918) |
| 0.1 | 15.4 | GO:0008202 | steroid metabolic process(GO:0008202) |
| 0.1 | 0.3 | GO:0016598 | protein arginylation(GO:0016598) |
| 0.1 | 4.2 | GO:0071427 | mRNA export from nucleus(GO:0006406) mRNA-containing ribonucleoprotein complex export from nucleus(GO:0071427) |
| 0.1 | 0.8 | GO:0071732 | cellular response to nitric oxide(GO:0071732) |
| 0.1 | 11.6 | GO:0045471 | response to ethanol(GO:0045471) |
| 0.1 | 0.8 | GO:0018022 | peptidyl-lysine methylation(GO:0018022) |
| 0.1 | 2.3 | GO:0097502 | mannosylation(GO:0097502) |
| 0.1 | 0.4 | GO:0006353 | DNA-templated transcription, termination(GO:0006353) |
| 0.1 | 0.8 | GO:0045943 | positive regulation of transcription from RNA polymerase I promoter(GO:0045943) |
| 0.1 | 3.1 | GO:0018149 | peptide cross-linking(GO:0018149) |
| 0.1 | 0.7 | GO:1900264 | regulation of DNA-directed DNA polymerase activity(GO:1900262) positive regulation of DNA-directed DNA polymerase activity(GO:1900264) |
| 0.1 | 2.6 | GO:0042274 | ribosomal small subunit biogenesis(GO:0042274) |
| 0.1 | 1.2 | GO:0031297 | replication fork processing(GO:0031297) |
| 0.1 | 1.1 | GO:0030866 | cortical actin cytoskeleton organization(GO:0030866) |
| 0.0 | 1.2 | GO:0030488 | tRNA methylation(GO:0030488) |
| 0.0 | 5.5 | GO:0006260 | DNA replication(GO:0006260) |
| 0.0 | 1.4 | GO:0043966 | histone H3 acetylation(GO:0043966) |
| 0.0 | 1.2 | GO:0007052 | mitotic spindle organization(GO:0007052) |
| 0.0 | 0.1 | GO:0033005 | positive regulation of mast cell activation(GO:0033005) |
| 0.0 | 0.4 | GO:0016180 | snRNA processing(GO:0016180) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 99.8 | 598.7 | GO:1990111 | spermatoproteasome complex(GO:1990111) |
| 60.1 | 420.9 | GO:0042825 | TAP complex(GO:0042825) |
| 57.3 | 229.2 | GO:0020005 | symbiont-containing vacuole membrane(GO:0020005) |
| 56.7 | 56.7 | GO:0032398 | MHC class Ib protein complex(GO:0032398) |
| 37.2 | 111.6 | GO:0008537 | proteasome activator complex(GO:0008537) |
| 33.5 | 234.6 | GO:0072559 | NLRP3 inflammasome complex(GO:0072559) |
| 31.4 | 501.9 | GO:0042612 | MHC class I protein complex(GO:0042612) |
| 22.1 | 22.1 | GO:0097169 | AIM2 inflammasome complex(GO:0097169) |
| 20.3 | 61.0 | GO:0005896 | interleukin-6 receptor complex(GO:0005896) ciliary neurotrophic factor receptor complex(GO:0070110) |
| 20.3 | 101.4 | GO:0031265 | CD95 death-inducing signaling complex(GO:0031265) |
| 19.7 | 59.0 | GO:0035692 | macrophage migration inhibitory factor receptor complex(GO:0035692) |
| 14.7 | 14.7 | GO:0020003 | symbiont-containing vacuole(GO:0020003) |
| 14.6 | 58.3 | GO:0070022 | transforming growth factor beta receptor homodimeric complex(GO:0070022) |
| 10.6 | 232.4 | GO:0031233 | intrinsic component of external side of plasma membrane(GO:0031233) |
| 10.6 | 31.7 | GO:1990682 | CSF1-CSF1R complex(GO:1990682) |
| 9.9 | 198.4 | GO:0043196 | varicosity(GO:0043196) |
| 9.6 | 38.2 | GO:0036398 | TCR signalosome(GO:0036398) microspike(GO:0044393) |
| 9.0 | 89.7 | GO:0043020 | NADPH oxidase complex(GO:0043020) |
| 8.5 | 59.8 | GO:0071458 | integral component of cytoplasmic side of endoplasmic reticulum membrane(GO:0071458) integral component of lumenal side of endoplasmic reticulum membrane(GO:0071556) lumenal side of endoplasmic reticulum membrane(GO:0098553) |
| 7.0 | 48.9 | GO:0042406 | extrinsic component of endoplasmic reticulum membrane(GO:0042406) |
| 5.5 | 16.6 | GO:1990826 | nucleoplasmic periphery of the nuclear pore complex(GO:1990826) |
| 5.4 | 16.2 | GO:0098554 | cytoplasmic side of endoplasmic reticulum membrane(GO:0098554) |
| 5.4 | 262.9 | GO:0016235 | aggresome(GO:0016235) |
| 5.1 | 25.7 | GO:0031262 | Ndc80 complex(GO:0031262) |
| 5.0 | 140.5 | GO:0001891 | phagocytic cup(GO:0001891) |
| 5.0 | 189.5 | GO:0001772 | immunological synapse(GO:0001772) |
| 4.6 | 18.3 | GO:0031904 | endosome lumen(GO:0031904) |
| 4.5 | 13.6 | GO:1990622 | CHOP-ATF3 complex(GO:1990622) |
| 4.4 | 30.8 | GO:0097136 | Bcl-2 family protein complex(GO:0097136) |
| 4.0 | 15.9 | GO:0044194 | cytolytic granule(GO:0044194) |
| 4.0 | 71.2 | GO:0031258 | lamellipodium membrane(GO:0031258) |
| 3.9 | 23.5 | GO:0042105 | alpha-beta T cell receptor complex(GO:0042105) |
| 3.5 | 10.4 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 3.4 | 10.2 | GO:0046691 | intracellular canaliculus(GO:0046691) |
| 3.4 | 131.9 | GO:0005771 | multivesicular body(GO:0005771) |
| 3.4 | 84.6 | GO:0035327 | transcriptionally active chromatin(GO:0035327) |
| 3.3 | 6.7 | GO:0044279 | other organism cell membrane(GO:0044218) other organism membrane(GO:0044279) |
| 3.3 | 10.0 | GO:0070826 | paraferritin complex(GO:0070826) |
| 3.1 | 9.4 | GO:0000110 | nucleotide-excision repair factor 1 complex(GO:0000110) |
| 2.7 | 5.4 | GO:0031166 | integral component of vacuolar membrane(GO:0031166) intrinsic component of vacuolar membrane(GO:0031310) |
| 2.6 | 192.6 | GO:0005811 | lipid particle(GO:0005811) |
| 2.5 | 5.0 | GO:0044322 | endoplasmic reticulum quality control compartment(GO:0044322) |
| 2.4 | 259.1 | GO:0016605 | PML body(GO:0016605) |
| 2.4 | 19.0 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 2.3 | 11.4 | GO:1990604 | IRE1-TRAF2-ASK1 complex(GO:1990604) |
| 2.1 | 4.3 | GO:0032059 | bleb(GO:0032059) |
| 2.1 | 612.1 | GO:0009897 | external side of plasma membrane(GO:0009897) |
| 2.1 | 8.3 | GO:0005971 | ribonucleoside-diphosphate reductase complex(GO:0005971) |
| 2.1 | 6.2 | GO:0044614 | nuclear pore cytoplasmic filaments(GO:0044614) |
| 1.9 | 13.1 | GO:0061617 | MICOS complex(GO:0061617) |
| 1.9 | 11.2 | GO:0032444 | activin responsive factor complex(GO:0032444) |
| 1.7 | 24.4 | GO:0005832 | chaperonin-containing T-complex(GO:0005832) |
| 1.7 | 10.4 | GO:0014731 | spectrin-associated cytoskeleton(GO:0014731) |
| 1.7 | 5.2 | GO:0000814 | ESCRT II complex(GO:0000814) |
| 1.7 | 42.8 | GO:0042629 | mast cell granule(GO:0042629) |
| 1.7 | 111.5 | GO:0005876 | spindle microtubule(GO:0005876) |
| 1.7 | 18.2 | GO:0042555 | MCM complex(GO:0042555) |
| 1.6 | 23.6 | GO:0042582 | primary lysosome(GO:0005766) azurophil granule(GO:0042582) |
| 1.6 | 146.9 | GO:0000786 | nucleosome(GO:0000786) |
| 1.5 | 7.3 | GO:0044530 | supraspliceosomal complex(GO:0044530) |
| 1.4 | 11.2 | GO:0005658 | alpha DNA polymerase:primase complex(GO:0005658) |
| 1.4 | 10.9 | GO:0000940 | condensed chromosome outer kinetochore(GO:0000940) |
| 1.3 | 18.4 | GO:0016580 | Sin3 complex(GO:0016580) |
| 1.3 | 12.6 | GO:0043202 | lysosomal lumen(GO:0043202) |
| 1.3 | 135.3 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 1.2 | 22.5 | GO:0030056 | hemidesmosome(GO:0030056) |
| 1.2 | 6.2 | GO:0035032 | phosphatidylinositol 3-kinase complex, class III(GO:0035032) |
| 1.2 | 123.2 | GO:0072562 | blood microparticle(GO:0072562) |
| 1.2 | 891.4 | GO:0005789 | endoplasmic reticulum membrane(GO:0005789) |
| 1.1 | 85.0 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 1.1 | 8.8 | GO:0031080 | nuclear pore outer ring(GO:0031080) |
| 1.1 | 26.3 | GO:0000145 | exocyst(GO:0000145) |
| 1.1 | 74.2 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 1.1 | 3.3 | GO:0044611 | nuclear pore inner ring(GO:0044611) |
| 1.1 | 440.9 | GO:0005764 | lytic vacuole(GO:0000323) lysosome(GO:0005764) |
| 1.1 | 14.9 | GO:0070938 | contractile ring(GO:0070938) |
| 1.0 | 3.1 | GO:1990589 | ATF4-CREB1 transcription factor complex(GO:1990589) |
| 1.0 | 5.2 | GO:0033588 | Elongator holoenzyme complex(GO:0033588) |
| 1.0 | 25.6 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 1.0 | 29.4 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 1.0 | 7.6 | GO:0044815 | DNA packaging complex(GO:0044815) |
| 1.0 | 4.8 | GO:0034455 | t-UTP complex(GO:0034455) |
| 0.9 | 13.1 | GO:0008278 | cohesin complex(GO:0008278) |
| 0.9 | 65.6 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.9 | 19.0 | GO:0035861 | site of double-strand break(GO:0035861) |
| 0.9 | 30.7 | GO:0002102 | podosome(GO:0002102) |
| 0.9 | 39.3 | GO:0031519 | PcG protein complex(GO:0031519) |
| 0.9 | 5.2 | GO:0030688 | preribosome, small subunit precursor(GO:0030688) |
| 0.8 | 3.4 | GO:0070545 | PeBoW complex(GO:0070545) |
| 0.8 | 23.7 | GO:0005689 | U12-type spliceosomal complex(GO:0005689) |
| 0.8 | 13.4 | GO:0071011 | precatalytic spliceosome(GO:0071011) |
| 0.8 | 15.4 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
| 0.8 | 16.9 | GO:0005732 | small nucleolar ribonucleoprotein complex(GO:0005732) |
| 0.7 | 9.6 | GO:0000801 | central element(GO:0000801) |
| 0.7 | 16.2 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.7 | 34.7 | GO:0005793 | endoplasmic reticulum-Golgi intermediate compartment(GO:0005793) |
| 0.7 | 10.3 | GO:0016010 | dystrophin-associated glycoprotein complex(GO:0016010) glycoprotein complex(GO:0090665) |
| 0.7 | 22.0 | GO:0032154 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.7 | 8.2 | GO:0071564 | npBAF complex(GO:0071564) |
| 0.7 | 45.4 | GO:0030863 | cortical cytoskeleton(GO:0030863) |
| 0.7 | 584.8 | GO:0005730 | nucleolus(GO:0005730) |
| 0.7 | 21.5 | GO:0005801 | cis-Golgi network(GO:0005801) |
| 0.6 | 5.8 | GO:0005847 | mRNA cleavage and polyadenylation specificity factor complex(GO:0005847) |
| 0.6 | 6.3 | GO:0005721 | pericentric heterochromatin(GO:0005721) |
| 0.6 | 12.5 | GO:0030140 | trans-Golgi network transport vesicle(GO:0030140) |
| 0.6 | 11.8 | GO:0005782 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.6 | 727.1 | GO:0005615 | extracellular space(GO:0005615) |
| 0.6 | 42.4 | GO:0016363 | nuclear matrix(GO:0016363) |
| 0.6 | 65.5 | GO:0005741 | mitochondrial outer membrane(GO:0005741) |
| 0.6 | 44.4 | GO:0005777 | peroxisome(GO:0005777) microbody(GO:0042579) |
| 0.6 | 36.5 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.6 | 3.5 | GO:0031932 | TORC2 complex(GO:0031932) |
| 0.6 | 13.8 | GO:0031304 | intrinsic component of mitochondrial inner membrane(GO:0031304) integral component of mitochondrial inner membrane(GO:0031305) |
| 0.6 | 4.5 | GO:0070187 | telosome(GO:0070187) |
| 0.6 | 3.4 | GO:0033202 | Ino80 complex(GO:0031011) DNA helicase complex(GO:0033202) |
| 0.6 | 1378.9 | GO:0005829 | cytosol(GO:0005829) |
| 0.5 | 8.6 | GO:0032588 | trans-Golgi network membrane(GO:0032588) |
| 0.4 | 1.3 | GO:0045098 | type III intermediate filament(GO:0045098) |
| 0.4 | 58.8 | GO:0005925 | focal adhesion(GO:0005925) |
| 0.4 | 3.0 | GO:0032045 | guanyl-nucleotide exchange factor complex(GO:0032045) |
| 0.3 | 152.0 | GO:0005783 | endoplasmic reticulum(GO:0005783) |
| 0.3 | 15.3 | GO:0005913 | cell-cell adherens junction(GO:0005913) |
| 0.3 | 2.3 | GO:0000242 | pericentriolar material(GO:0000242) |
| 0.3 | 6.1 | GO:0008023 | transcription elongation factor complex(GO:0008023) |
| 0.3 | 2.3 | GO:0030014 | CCR4-NOT complex(GO:0030014) |
| 0.2 | 10.0 | GO:0001750 | photoreceptor outer segment(GO:0001750) |
| 0.2 | 3.1 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.2 | 1.8 | GO:0080008 | Cul4-RING E3 ubiquitin ligase complex(GO:0080008) |
| 0.1 | 0.7 | GO:0005663 | DNA replication factor C complex(GO:0005663) |
| 0.1 | 1.2 | GO:0071821 | FANCM-MHF complex(GO:0071821) |
| 0.1 | 7.1 | GO:0036064 | ciliary basal body(GO:0036064) |
| 0.1 | 5.7 | GO:0005923 | bicellular tight junction(GO:0005923) |
| 0.1 | 2.6 | GO:0005758 | mitochondrial intermembrane space(GO:0005758) |
| 0.1 | 1.1 | GO:0031201 | SNARE complex(GO:0031201) |
| 0.1 | 1.3 | GO:0005884 | actin filament(GO:0005884) |
| 0.0 | 0.4 | GO:0032039 | integrator complex(GO:0032039) |
| 0.0 | 0.5 | GO:0030131 | clathrin adaptor complex(GO:0030131) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 81.6 | 244.8 | GO:0015433 | peptide antigen-transporting ATPase activity(GO:0015433) tapasin binding(GO:0046980) |
| 53.6 | 214.6 | GO:0003726 | double-stranded RNA adenosine deaminase activity(GO:0003726) |
| 45.2 | 135.5 | GO:0046977 | TAP binding(GO:0046977) TAP1 binding(GO:0046978) TAP2 binding(GO:0046979) |
| 42.4 | 84.8 | GO:0015440 | peptide-transporting ATPase activity(GO:0015440) |
| 36.3 | 108.8 | GO:0015322 | proton-dependent oligopeptide secondary active transmembrane transporter activity(GO:0005427) secondary active oligopeptide transmembrane transporter activity(GO:0015322) |
| 32.9 | 65.9 | GO:0031735 | CCR10 chemokine receptor binding(GO:0031735) |
| 25.7 | 592.1 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 23.3 | 116.4 | GO:0048248 | CXCR3 chemokine receptor binding(GO:0048248) |
| 22.0 | 110.2 | GO:0008310 | single-stranded DNA 3'-5' exodeoxyribonuclease activity(GO:0008310) |
| 21.7 | 195.7 | GO:0016174 | NAD(P)H oxidase activity(GO:0016174) |
| 21.6 | 86.5 | GO:0005143 | interleukin-12 receptor binding(GO:0005143) |
| 21.1 | 570.4 | GO:0070003 | threonine-type endopeptidase activity(GO:0004298) threonine-type peptidase activity(GO:0070003) |
| 20.1 | 260.9 | GO:0097200 | cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:0097200) |
| 18.7 | 112.1 | GO:0003956 | NAD(P)+-protein-arginine ADP-ribosyltransferase activity(GO:0003956) |
| 18.6 | 130.0 | GO:1901612 | cardiolipin binding(GO:1901612) |
| 16.4 | 65.6 | GO:0004771 | sterol esterase activity(GO:0004771) |
| 15.9 | 111.6 | GO:0061133 | endopeptidase activator activity(GO:0061133) |
| 15.5 | 46.4 | GO:0070976 | TIR domain binding(GO:0070976) |
| 15.5 | 185.6 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 15.4 | 123.3 | GO:0031386 | protein tag(GO:0031386) |
| 15.2 | 61.0 | GO:0004897 | ciliary neurotrophic factor receptor activity(GO:0004897) |
| 14.2 | 170.7 | GO:0031730 | CCR5 chemokine receptor binding(GO:0031730) |
| 14.1 | 84.7 | GO:0048030 | disaccharide binding(GO:0048030) |
| 13.9 | 527.0 | GO:0042605 | peptide antigen binding(GO:0042605) |
| 13.8 | 124.5 | GO:0001730 | 2'-5'-oligoadenylate synthetase activity(GO:0001730) |
| 12.6 | 62.8 | GO:0008503 | benzodiazepine receptor activity(GO:0008503) |
| 11.6 | 243.7 | GO:0005035 | tumor necrosis factor-activated receptor activity(GO:0005031) death receptor activity(GO:0005035) |
| 11.2 | 33.6 | GO:0004478 | methionine adenosyltransferase activity(GO:0004478) |
| 11.1 | 55.5 | GO:0034714 | type III transforming growth factor beta receptor binding(GO:0034714) |
| 10.6 | 296.8 | GO:0008009 | chemokine activity(GO:0008009) |
| 10.6 | 31.7 | GO:0005157 | macrophage colony-stimulating factor receptor binding(GO:0005157) |
| 10.1 | 40.3 | GO:0008265 | Mo-molybdopterin cofactor sulfurase activity(GO:0008265) |
| 10.0 | 59.8 | GO:0042500 | aspartic endopeptidase activity, intramembrane cleaving(GO:0042500) |
| 9.8 | 59.0 | GO:0042289 | MHC class II protein binding(GO:0042289) |
| 8.9 | 26.8 | GO:0050656 | 3'-phosphoadenosine 5'-phosphosulfate binding(GO:0050656) |
| 8.9 | 26.7 | GO:0004769 | steroid delta-isomerase activity(GO:0004769) |
| 8.7 | 43.3 | GO:0005138 | interleukin-6 receptor binding(GO:0005138) |
| 8.5 | 17.1 | GO:0050785 | advanced glycation end-product receptor activity(GO:0050785) |
| 8.4 | 92.3 | GO:0008641 | small protein activating enzyme activity(GO:0008641) |
| 8.1 | 40.5 | GO:0070892 | lipoteichoic acid receptor activity(GO:0070892) |
| 7.6 | 45.9 | GO:0048019 | receptor antagonist activity(GO:0048019) |
| 7.0 | 112.6 | GO:0070628 | proteasome binding(GO:0070628) |
| 6.9 | 68.9 | GO:0005384 | manganese ion transmembrane transporter activity(GO:0005384) |
| 6.9 | 20.6 | GO:0016822 | hydrolase activity, acting on acid carbon-carbon bonds(GO:0016822) hydrolase activity, acting on acid carbon-carbon bonds, in ketonic substances(GO:0016823) |
| 6.7 | 60.2 | GO:0022833 | mechanically-gated ion channel activity(GO:0008381) mechanically gated channel activity(GO:0022833) |
| 6.3 | 18.8 | GO:0004556 | alpha-amylase activity(GO:0004556) |
| 6.1 | 18.3 | GO:0035651 | AP-3 adaptor complex binding(GO:0035651) |
| 6.0 | 78.1 | GO:0015924 | mannosyl-oligosaccharide mannosidase activity(GO:0015924) |
| 5.8 | 35.0 | GO:0004694 | eukaryotic translation initiation factor 2alpha kinase activity(GO:0004694) |
| 5.7 | 79.3 | GO:0051400 | BH domain binding(GO:0051400) |
| 5.7 | 50.9 | GO:1904264 | ubiquitin protein ligase activity involved in ERAD pathway(GO:1904264) |
| 5.4 | 16.2 | GO:0030626 | U12 snRNA binding(GO:0030626) |
| 5.3 | 26.3 | GO:0031798 | type 1 metabotropic glutamate receptor binding(GO:0031798) |
| 5.0 | 40.2 | GO:0035184 | histone threonine kinase activity(GO:0035184) |
| 5.0 | 1501.0 | GO:0003924 | GTPase activity(GO:0003924) |
| 4.6 | 18.6 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 4.6 | 23.1 | GO:0072345 | NAADP-sensitive calcium-release channel activity(GO:0072345) |
| 4.5 | 13.5 | GO:0031852 | mu-type opioid receptor binding(GO:0031852) |
| 4.4 | 17.5 | GO:0003854 | 3-beta-hydroxy-delta5-steroid dehydrogenase activity(GO:0003854) |
| 4.3 | 30.1 | GO:0046976 | histone methyltransferase activity (H3-K27 specific)(GO:0046976) |
| 3.9 | 274.2 | GO:0003725 | double-stranded RNA binding(GO:0003725) |
| 3.9 | 31.3 | GO:0047134 | protein-disulfide reductase activity(GO:0047134) |
| 3.7 | 25.8 | GO:0050700 | CARD domain binding(GO:0050700) |
| 3.7 | 36.5 | GO:0019957 | C-C chemokine binding(GO:0019957) |
| 3.6 | 97.1 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) |
| 3.6 | 10.7 | GO:0008475 | procollagen-lysine 5-dioxygenase activity(GO:0008475) procollagen glucosyltransferase activity(GO:0033823) |
| 3.5 | 10.4 | GO:0050265 | RNA uridylyltransferase activity(GO:0050265) |
| 3.2 | 19.5 | GO:0004126 | cytidine deaminase activity(GO:0004126) |
| 3.2 | 15.9 | GO:0030348 | syntaxin-3 binding(GO:0030348) |
| 3.1 | 68.8 | GO:0032183 | SUMO binding(GO:0032183) |
| 3.1 | 30.8 | GO:0016176 | superoxide-generating NADPH oxidase activator activity(GO:0016176) |
| 3.1 | 207.9 | GO:0030971 | receptor tyrosine kinase binding(GO:0030971) |
| 3.0 | 54.9 | GO:0042288 | MHC class I protein binding(GO:0042288) |
| 3.0 | 33.5 | GO:0030346 | protein phosphatase 2B binding(GO:0030346) |
| 3.0 | 27.2 | GO:0046703 | natural killer cell lectin-like receptor binding(GO:0046703) |
| 3.0 | 12.0 | GO:0047023 | androsterone dehydrogenase activity(GO:0047023) |
| 3.0 | 12.0 | GO:0070644 | vitamin D response element binding(GO:0070644) |
| 3.0 | 11.8 | GO:0004165 | dodecenoyl-CoA delta-isomerase activity(GO:0004165) |
| 3.0 | 20.7 | GO:0035251 | UDP-glucosyltransferase activity(GO:0035251) |
| 2.9 | 14.6 | GO:0002151 | G-quadruplex RNA binding(GO:0002151) |
| 2.9 | 8.6 | GO:0019809 | spermidine binding(GO:0019809) |
| 2.9 | 8.6 | GO:0031731 | CCR6 chemokine receptor binding(GO:0031731) |
| 2.8 | 14.0 | GO:1990829 | C-rich single-stranded DNA binding(GO:1990829) |
| 2.8 | 139.3 | GO:0001786 | phosphatidylserine binding(GO:0001786) |
| 2.7 | 16.0 | GO:0004982 | N-formyl peptide receptor activity(GO:0004982) |
| 2.7 | 16.0 | GO:0048039 | ubiquinone binding(GO:0048039) |
| 2.5 | 200.9 | GO:0000979 | RNA polymerase II core promoter sequence-specific DNA binding(GO:0000979) |
| 2.5 | 7.4 | GO:0004483 | mRNA (nucleoside-2'-O-)-methyltransferase activity(GO:0004483) |
| 2.5 | 7.4 | GO:0016508 | long-chain-enoyl-CoA hydratase activity(GO:0016508) testosterone dehydrogenase [NAD(P)] activity(GO:0030283) |
| 2.4 | 9.6 | GO:0016416 | carnitine O-palmitoyltransferase activity(GO:0004095) O-palmitoyltransferase activity(GO:0016416) |
| 2.4 | 38.2 | GO:0044548 | S100 protein binding(GO:0044548) |
| 2.4 | 229.8 | GO:0036459 | thiol-dependent ubiquitinyl hydrolase activity(GO:0036459) |
| 2.4 | 105.8 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 2.3 | 7.0 | GO:0009384 | N-acylmannosamine kinase activity(GO:0009384) |
| 2.3 | 7.0 | GO:0005171 | hepatocyte growth factor receptor binding(GO:0005171) |
| 2.3 | 9.3 | GO:0004052 | arachidonate 12-lipoxygenase activity(GO:0004052) |
| 2.3 | 27.2 | GO:0042609 | CD4 receptor binding(GO:0042609) |
| 2.3 | 6.8 | GO:0003865 | 3-oxo-5-alpha-steroid 4-dehydrogenase activity(GO:0003865) cholestenone 5-alpha-reductase activity(GO:0047751) |
| 2.2 | 31.2 | GO:0030550 | acetylcholine receptor inhibitor activity(GO:0030550) |
| 2.1 | 8.5 | GO:0047184 | 1-acylglycerophosphocholine O-acyltransferase activity(GO:0047184) |
| 2.1 | 10.4 | GO:0045340 | mercury ion binding(GO:0045340) |
| 2.1 | 29.1 | GO:0023026 | MHC class II protein complex binding(GO:0023026) |
| 2.1 | 10.3 | GO:0004614 | phosphoglucomutase activity(GO:0004614) |
| 2.0 | 8.0 | GO:0016647 | oxidoreductase activity, acting on the CH-NH group of donors, oxygen as acceptor(GO:0016647) |
| 2.0 | 15.8 | GO:1990247 | N6-methyladenosine-containing RNA binding(GO:1990247) |
| 2.0 | 9.8 | GO:0008970 | phosphatidylcholine 1-acylhydrolase activity(GO:0008970) |
| 1.9 | 23.0 | GO:0070008 | serine-type exopeptidase activity(GO:0070008) |
| 1.9 | 53.7 | GO:0008395 | steroid hydroxylase activity(GO:0008395) |
| 1.9 | 17.2 | GO:0005072 | transforming growth factor beta receptor, cytoplasmic mediator activity(GO:0005072) |
| 1.9 | 34.1 | GO:0004190 | aspartic-type endopeptidase activity(GO:0004190) |
| 1.9 | 5.6 | GO:0071208 | histone pre-mRNA DCP binding(GO:0071208) |
| 1.8 | 7.3 | GO:0032558 | adenyl deoxyribonucleotide binding(GO:0032558) |
| 1.8 | 65.2 | GO:0051539 | 4 iron, 4 sulfur cluster binding(GO:0051539) |
| 1.8 | 25.0 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 1.7 | 20.9 | GO:0001608 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 1.7 | 34.8 | GO:0008199 | ferric iron binding(GO:0008199) |
| 1.7 | 12.1 | GO:0005078 | MAP-kinase scaffold activity(GO:0005078) |
| 1.7 | 36.0 | GO:0005070 | SH3/SH2 adaptor activity(GO:0005070) |
| 1.7 | 116.4 | GO:0038024 | cargo receptor activity(GO:0038024) |
| 1.7 | 8.6 | GO:0045322 | unmethylated CpG binding(GO:0045322) |
| 1.7 | 40.6 | GO:0043395 | heparan sulfate proteoglycan binding(GO:0043395) |
| 1.7 | 11.7 | GO:0016634 | oxidoreductase activity, acting on the CH-CH group of donors, oxygen as acceptor(GO:0016634) |
| 1.7 | 8.3 | GO:0016728 | ribonucleoside-diphosphate reductase activity, thioredoxin disulfide as acceptor(GO:0004748) oxidoreductase activity, acting on CH or CH2 groups, disulfide as acceptor(GO:0016728) ribonucleoside-diphosphate reductase activity(GO:0061731) |
| 1.6 | 91.8 | GO:0000049 | tRNA binding(GO:0000049) |
| 1.6 | 32.3 | GO:0043325 | phosphatidylinositol-3,4-bisphosphate binding(GO:0043325) |
| 1.6 | 11.1 | GO:0043423 | 3-phosphoinositide-dependent protein kinase binding(GO:0043423) |
| 1.6 | 3.1 | GO:0001225 | RNA polymerase II transcription coactivator binding(GO:0001225) |
| 1.5 | 4.5 | GO:1990955 | G-rich single-stranded DNA binding(GO:1990955) |
| 1.5 | 301.7 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 1.5 | 433.9 | GO:0030246 | carbohydrate binding(GO:0030246) |
| 1.5 | 10.2 | GO:0003720 | telomerase activity(GO:0003720) RNA-directed DNA polymerase activity(GO:0003964) |
| 1.5 | 40.9 | GO:0001784 | phosphotyrosine binding(GO:0001784) |
| 1.4 | 14.5 | GO:0015143 | urate transmembrane transporter activity(GO:0015143) salt transmembrane transporter activity(GO:1901702) |
| 1.4 | 21.6 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 1.4 | 24.2 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 1.4 | 7.0 | GO:0046974 | histone methyltransferase activity (H3-K9 specific)(GO:0046974) |
| 1.4 | 88.8 | GO:0019843 | rRNA binding(GO:0019843) |
| 1.4 | 8.3 | GO:0004028 | 3-chloroallyl aldehyde dehydrogenase activity(GO:0004028) |
| 1.4 | 76.5 | GO:0004520 | endodeoxyribonuclease activity(GO:0004520) |
| 1.3 | 5.2 | GO:0089720 | caspase binding(GO:0089720) |
| 1.3 | 8.9 | GO:0046975 | histone methyltransferase activity (H3-K36 specific)(GO:0046975) |
| 1.3 | 3.8 | GO:0001716 | L-amino-acid oxidase activity(GO:0001716) |
| 1.3 | 6.3 | GO:0032454 | histone demethylase activity (H3-K9 specific)(GO:0032454) |
| 1.2 | 30.8 | GO:0070840 | dynein complex binding(GO:0070840) |
| 1.2 | 6.1 | GO:0035241 | protein-arginine omega-N monomethyltransferase activity(GO:0035241) histone methyltransferase activity (H4-R3 specific)(GO:0044020) |
| 1.2 | 10.9 | GO:0043515 | kinetochore binding(GO:0043515) |
| 1.2 | 40.8 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 1.2 | 3.6 | GO:0004995 | tachykinin receptor activity(GO:0004995) |
| 1.1 | 7.8 | GO:0015526 | hexose phosphate transmembrane transporter activity(GO:0015119) organophosphate:inorganic phosphate antiporter activity(GO:0015315) hexose-phosphate:inorganic phosphate antiporter activity(GO:0015526) glucose 6-phosphate:inorganic phosphate antiporter activity(GO:0061513) |
| 1.1 | 5.3 | GO:0004337 | dimethylallyltranstransferase activity(GO:0004161) geranyltranstransferase activity(GO:0004337) |
| 1.0 | 3.1 | GO:0009019 | tRNA (guanine-N1-)-methyltransferase activity(GO:0009019) |
| 1.0 | 23.3 | GO:0070064 | proline-rich region binding(GO:0070064) |
| 1.0 | 45.5 | GO:0030374 | ligand-dependent nuclear receptor transcription coactivator activity(GO:0030374) |
| 1.0 | 92.6 | GO:0031490 | chromatin DNA binding(GO:0031490) |
| 1.0 | 68.4 | GO:0033613 | activating transcription factor binding(GO:0033613) |
| 1.0 | 4.0 | GO:0000822 | inositol hexakisphosphate binding(GO:0000822) |
| 1.0 | 171.3 | GO:0061135 | endopeptidase regulator activity(GO:0061135) |
| 1.0 | 3.0 | GO:0002114 | interleukin-33 receptor activity(GO:0002114) |
| 1.0 | 2.9 | GO:0000253 | 3-keto sterol reductase activity(GO:0000253) |
| 1.0 | 69.2 | GO:0001948 | glycoprotein binding(GO:0001948) |
| 1.0 | 4.8 | GO:0003691 | double-stranded telomeric DNA binding(GO:0003691) |
| 0.9 | 164.0 | GO:0042393 | histone binding(GO:0042393) |
| 0.9 | 32.9 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.9 | 14.6 | GO:0001671 | ATPase activator activity(GO:0001671) |
| 0.9 | 3.6 | GO:0004706 | JUN kinase kinase kinase activity(GO:0004706) |
| 0.9 | 22.9 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.9 | 9.7 | GO:0005172 | vascular endothelial growth factor receptor binding(GO:0005172) |
| 0.9 | 13.1 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.9 | 24.4 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.9 | 27.3 | GO:0001205 | transcriptional activator activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001205) |
| 0.8 | 17.5 | GO:0005112 | Notch binding(GO:0005112) |
| 0.8 | 9.9 | GO:0017166 | vinculin binding(GO:0017166) |
| 0.8 | 5.5 | GO:0038062 | protein tyrosine kinase collagen receptor activity(GO:0038062) |
| 0.8 | 60.4 | GO:0031072 | heat shock protein binding(GO:0031072) |
| 0.8 | 2.3 | GO:0005017 | platelet-derived growth factor-activated receptor activity(GO:0005017) |
| 0.8 | 3.1 | GO:0047756 | chondroitin 4-sulfotransferase activity(GO:0047756) |
| 0.8 | 15.1 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.7 | 5.1 | GO:0003958 | NADPH-hemoprotein reductase activity(GO:0003958) |
| 0.7 | 2.9 | GO:0004563 | beta-N-acetylhexosaminidase activity(GO:0004563) |
| 0.7 | 22.9 | GO:0003678 | DNA helicase activity(GO:0003678) |
| 0.7 | 5.5 | GO:0030368 | interleukin-17 receptor activity(GO:0030368) |
| 0.7 | 26.7 | GO:0003755 | peptidyl-prolyl cis-trans isomerase activity(GO:0003755) |
| 0.6 | 1.8 | GO:0070052 | collagen V binding(GO:0070052) |
| 0.6 | 2.3 | GO:0002046 | opsin binding(GO:0002046) |
| 0.5 | 8.2 | GO:0050431 | transforming growth factor beta binding(GO:0050431) |
| 0.5 | 2.6 | GO:1903763 | gap junction channel activity involved in cell communication by electrical coupling(GO:1903763) |
| 0.5 | 21.4 | GO:0004601 | peroxidase activity(GO:0004601) |
| 0.5 | 15.4 | GO:0005248 | voltage-gated sodium channel activity(GO:0005248) |
| 0.5 | 6.1 | GO:0031491 | nucleosome binding(GO:0031491) |
| 0.5 | 5.5 | GO:0004180 | carboxypeptidase activity(GO:0004180) |
| 0.5 | 38.1 | GO:0005178 | integrin binding(GO:0005178) |
| 0.5 | 5.9 | GO:0017160 | Ral GTPase binding(GO:0017160) |
| 0.4 | 13.4 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.4 | 1.8 | GO:0060698 | endoribonuclease inhibitor activity(GO:0060698) |
| 0.4 | 8.2 | GO:0001099 | basal transcription machinery binding(GO:0001098) basal RNA polymerase II transcription machinery binding(GO:0001099) |
| 0.4 | 1.7 | GO:0003989 | acetyl-CoA carboxylase activity(GO:0003989) |
| 0.4 | 19.3 | GO:0016835 | carbon-oxygen lyase activity(GO:0016835) |
| 0.4 | 4.1 | GO:0070742 | C2H2 zinc finger domain binding(GO:0070742) |
| 0.4 | 5.5 | GO:0008198 | ferrous iron binding(GO:0008198) |
| 0.4 | 346.4 | GO:0008270 | zinc ion binding(GO:0008270) |
| 0.4 | 8.1 | GO:0070006 | metalloaminopeptidase activity(GO:0070006) |
| 0.4 | 5.0 | GO:0016813 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amidines(GO:0016813) |
| 0.4 | 3.0 | GO:0016846 | carbon-sulfur lyase activity(GO:0016846) |
| 0.4 | 102.3 | GO:0003712 | transcription cofactor activity(GO:0003712) |
| 0.4 | 10.4 | GO:0008519 | ammonium transmembrane transporter activity(GO:0008519) |
| 0.3 | 3.3 | GO:0005149 | interleukin-1 receptor binding(GO:0005149) |
| 0.3 | 13.9 | GO:0016811 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amides(GO:0016811) |
| 0.3 | 7.2 | GO:0005487 | nucleocytoplasmic transporter activity(GO:0005487) |
| 0.3 | 8.1 | GO:0001076 | transcription factor activity, RNA polymerase II transcription factor binding(GO:0001076) |
| 0.3 | 3.4 | GO:0003995 | acyl-CoA dehydrogenase activity(GO:0003995) |
| 0.3 | 9.4 | GO:0003684 | damaged DNA binding(GO:0003684) |
| 0.3 | 1.5 | GO:0004875 | complement receptor activity(GO:0004875) |
| 0.3 | 1.8 | GO:0005007 | fibroblast growth factor-activated receptor activity(GO:0005007) |
| 0.3 | 1.2 | GO:0052796 | exo-alpha-(2->3)-sialidase activity(GO:0052794) exo-alpha-(2->6)-sialidase activity(GO:0052795) exo-alpha-(2->8)-sialidase activity(GO:0052796) |
| 0.3 | 29.0 | GO:0004497 | monooxygenase activity(GO:0004497) |
| 0.3 | 2.8 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.3 | 4.4 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) |
| 0.3 | 2.8 | GO:0098641 | cadherin binding involved in cell-cell adhesion(GO:0098641) |
| 0.2 | 1.9 | GO:0004439 | phosphatidylinositol-4,5-bisphosphate 5-phosphatase activity(GO:0004439) |
| 0.2 | 4.3 | GO:0017049 | GTP-Rho binding(GO:0017049) |
| 0.2 | 6.0 | GO:0030515 | snoRNA binding(GO:0030515) |
| 0.2 | 1.1 | GO:0042731 | PH domain binding(GO:0042731) |
| 0.2 | 9.4 | GO:0008375 | acetylglucosaminyltransferase activity(GO:0008375) |
| 0.2 | 0.8 | GO:0019871 | sodium channel inhibitor activity(GO:0019871) |
| 0.2 | 0.7 | GO:0042974 | retinoic acid receptor binding(GO:0042974) |
| 0.1 | 3.1 | GO:0071889 | 14-3-3 protein binding(GO:0071889) |
| 0.1 | 34.6 | GO:0005096 | GTPase activator activity(GO:0005096) |
| 0.1 | 0.4 | GO:0001042 | RNA polymerase I core binding(GO:0001042) |
| 0.1 | 1.3 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.1 | 2.4 | GO:0004402 | histone acetyltransferase activity(GO:0004402) |
| 0.1 | 1.4 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.1 | 2.3 | GO:0000030 | mannosyltransferase activity(GO:0000030) |
| 0.1 | 1.2 | GO:0017134 | fibroblast growth factor binding(GO:0017134) |
| 0.1 | 8.1 | GO:0004725 | protein tyrosine phosphatase activity(GO:0004725) |
| 0.1 | 0.7 | GO:0043142 | single-stranded DNA-dependent ATPase activity(GO:0043142) |
| 0.1 | 1.7 | GO:0005242 | inward rectifier potassium channel activity(GO:0005242) |
| 0.1 | 1.3 | GO:0019200 | carbohydrate kinase activity(GO:0019200) |
| 0.1 | 4.6 | GO:0008757 | S-adenosylmethionine-dependent methyltransferase activity(GO:0008757) |
| 0.0 | 1.3 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.0 | 1.3 | GO:0016763 | transferase activity, transferring pentosyl groups(GO:0016763) |
| 0.0 | 0.5 | GO:0030306 | ADP-ribosylation factor binding(GO:0030306) |
| 0.0 | 0.3 | GO:0047498 | calcium-dependent phospholipase A2 activity(GO:0047498) |
| 0.0 | 4.9 | GO:0005085 | guanyl-nucleotide exchange factor activity(GO:0005085) |
| 0.0 | 0.3 | GO:0005547 | phosphatidylinositol-3,4,5-trisphosphate binding(GO:0005547) |
| 0.0 | 0.3 | GO:0016755 | transferase activity, transferring amino-acyl groups(GO:0016755) |
| 0.0 | 0.8 | GO:0019842 | vitamin binding(GO:0019842) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 12.2 | 218.8 | SA MMP CYTOKINE CONNECTION | Cytokines can induce activation of matrix metalloproteinases, which degrade extracellular matrix. |
| 9.6 | 250.7 | PID IL27 PATHWAY | IL27-mediated signaling events |
| 9.2 | 101.4 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 6.6 | 99.0 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
| 6.0 | 107.3 | SA CASPASE CASCADE | Apoptosis is mediated by caspases, cysteine proteases arranged in a proteolytic cascade. |
| 4.1 | 77.1 | ST G ALPHA S PATHWAY | G alpha s Pathway |
| 4.0 | 214.5 | PID IL12 2PATHWAY | IL12-mediated signaling events |
| 3.8 | 30.8 | SA PROGRAMMED CELL DEATH | Programmed cell death, or apoptosis, eliminates damaged or unneeded cells. |
| 3.6 | 85.7 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 3.5 | 38.9 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 3.4 | 119.8 | PID CXCR3 PATHWAY | CXCR3-mediated signaling events |
| 2.9 | 176.6 | PID IL4 2PATHWAY | IL4-mediated signaling events |
| 2.8 | 104.6 | PID HDAC CLASSII PATHWAY | Signaling events mediated by HDAC Class II |
| 2.6 | 36.1 | PID IL23 PATHWAY | IL23-mediated signaling events |
| 2.5 | 42.7 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 2.5 | 84.7 | PID SYNDECAN 4 PATHWAY | Syndecan-4-mediated signaling events |
| 2.5 | 94.5 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 2.4 | 75.4 | PID ALK1 PATHWAY | ALK1 signaling events |
| 2.4 | 52.7 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
| 2.4 | 116.4 | PID BCR 5PATHWAY | BCR signaling pathway |
| 2.2 | 336.8 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 2.0 | 47.8 | PID MYC PATHWAY | C-MYC pathway |
| 2.0 | 25.8 | PID NFKAPPAB CANONICAL PATHWAY | Canonical NF-kappaB pathway |
| 1.9 | 9.7 | PID LYMPH ANGIOGENESIS PATHWAY | VEGFR3 signaling in lymphatic endothelium |
| 1.9 | 26.8 | PID SYNDECAN 2 PATHWAY | Syndecan-2-mediated signaling events |
| 1.9 | 97.1 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 1.9 | 153.1 | PID SMAD2 3NUCLEAR PATHWAY | Regulation of nuclear SMAD2/3 signaling |
| 1.5 | 36.5 | PID TCPTP PATHWAY | Signaling events mediated by TCPTP |
| 1.4 | 26.6 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
| 1.3 | 13.1 | PID NECTIN PATHWAY | Nectin adhesion pathway |
| 1.3 | 40.7 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 1.1 | 310.0 | NABA SECRETED FACTORS | Genes encoding secreted soluble factors |
| 1.1 | 243.1 | NABA MATRISOME ASSOCIATED | Ensemble of genes encoding ECM-associated proteins including ECM-affilaited proteins, ECM regulators and secreted factors |
| 1.0 | 120.9 | PID P53 DOWNSTREAM PATHWAY | Direct p53 effectors |
| 1.0 | 16.0 | PID UPA UPAR PATHWAY | Urokinase-type plasminogen activator (uPA) and uPAR-mediated signaling |
| 1.0 | 8.6 | PID INTEGRIN A9B1 PATHWAY | Alpha9 beta1 integrin signaling events |
| 0.9 | 22.5 | PID A6B1 A6B4 INTEGRIN PATHWAY | a6b1 and a6b4 Integrin signaling |
| 0.9 | 38.2 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.9 | 20.5 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 0.8 | 12.1 | PID CERAMIDE PATHWAY | Ceramide signaling pathway |
| 0.8 | 24.4 | PID HEDGEHOG GLI PATHWAY | Hedgehog signaling events mediated by Gli proteins |
| 0.7 | 4.4 | PID P38 ALPHA BETA PATHWAY | Regulation of p38-alpha and p38-beta |
| 0.7 | 6.1 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.7 | 13.9 | PID NFAT TFPATHWAY | Calcineurin-regulated NFAT-dependent transcription in lymphocytes |
| 0.6 | 21.4 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.6 | 8.2 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.6 | 4.4 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.6 | 25.4 | PID E2F PATHWAY | E2F transcription factor network |
| 0.6 | 2.3 | SA G2 AND M PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
| 0.5 | 19.8 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.5 | 22.4 | PID ERA GENOMIC PATHWAY | Validated nuclear estrogen receptor alpha network |
| 0.5 | 10.2 | PID FAK PATHWAY | Signaling events mediated by focal adhesion kinase |
| 0.4 | 11.1 | PID PI3KCI PATHWAY | Class I PI3K signaling events |
| 0.4 | 14.6 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.4 | 12.8 | PID TGFBR PATHWAY | TGF-beta receptor signaling |
| 0.4 | 10.0 | PID HIF2PATHWAY | HIF-2-alpha transcription factor network |
| 0.4 | 20.2 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
| 0.4 | 3.0 | PID IL1 PATHWAY | IL1-mediated signaling events |
| 0.4 | 10.0 | PID CONE PATHWAY | Visual signal transduction: Cones |
| 0.3 | 4.1 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.2 | 2.6 | PID ERBB2 ERBB3 PATHWAY | ErbB2/ErbB3 signaling events |
| 0.2 | 5.5 | PID HES HEY PATHWAY | Notch-mediated HES/HEY network |
| 0.2 | 6.6 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.2 | 2.5 | PID CD40 PATHWAY | CD40/CD40L signaling |
| 0.2 | 2.6 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.2 | 5.0 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.2 | 5.1 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.1 | 1.2 | PID EPHRINB REV PATHWAY | Ephrin B reverse signaling |
| 0.1 | 3.8 | PID ARF6 TRAFFICKING PATHWAY | Arf6 trafficking events |
| 0.1 | 1.8 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 0.0 | 2.2 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.0 | 7.2 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 1.1 | PID MTOR 4PATHWAY | mTOR signaling pathway |
| 0.0 | 0.3 | SIG INSULIN RECEPTOR PATHWAY IN CARDIAC MYOCYTES | Genes related to the insulin receptor pathway |
| 0.0 | 0.3 | PID RAS PATHWAY | Regulation of Ras family activation |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 31.8 | 1907.8 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 22.1 | 66.2 | REACTOME ENDOSOMAL VACUOLAR PATHWAY | Genes involved in Endosomal/Vacuolar pathway |
| 20.3 | 263.5 | REACTOME RIP MEDIATED NFKB ACTIVATION VIA DAI | Genes involved in RIP-mediated NFkB activation via DAI |
| 20.2 | 222.0 | REACTOME NFKB ACTIVATION THROUGH FADD RIP1 PATHWAY MEDIATED BY CASPASE 8 AND10 | Genes involved in NF-kB activation through FADD/RIP-1 pathway mediated by caspase-8 and -10 |
| 16.9 | 286.9 | REACTOME ANTIGEN PRESENTATION FOLDING ASSEMBLY AND PEPTIDE LOADING OF CLASS I MHC | Genes involved in Antigen Presentation: Folding, assembly and peptide loading of class I MHC |
| 13.7 | 164.4 | REACTOME PD1 SIGNALING | Genes involved in PD-1 signaling |
| 12.2 | 61.0 | REACTOME IL 6 SIGNALING | Genes involved in Interleukin-6 signaling |
| 11.5 | 577.4 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 11.2 | 358.2 | REACTOME INTERFERON GAMMA SIGNALING | Genes involved in Interferon gamma signaling |
| 11.0 | 22.1 | REACTOME INFLAMMASOMES | Genes involved in Inflammasomes |
| 10.7 | 512.4 | REACTOME CROSS PRESENTATION OF SOLUBLE EXOGENOUS ANTIGENS ENDOSOMES | Genes involved in Cross-presentation of soluble exogenous antigens (endosomes) |
| 9.5 | 104.6 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 8.4 | 135.1 | REACTOME ANTIGEN PROCESSING CROSS PRESENTATION | Genes involved in Antigen processing-Cross presentation |
| 6.5 | 39.2 | REACTOME DEFENSINS | Genes involved in Defensins |
| 6.3 | 88.2 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 6.3 | 69.1 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 6.0 | 54.2 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 5.8 | 58.3 | REACTOME APOPTOSIS INDUCED DNA FRAGMENTATION | Genes involved in Apoptosis induced DNA fragmentation |
| 5.8 | 46.4 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION IN TLR7 8 OR 9 SIGNALING | Genes involved in TRAF6 mediated IRF7 activation in TLR7/8 or 9 signaling |
| 4.9 | 126.5 | REACTOME RNA POL I PROMOTER OPENING | Genes involved in RNA Polymerase I Promoter Opening |
| 4.8 | 180.6 | REACTOME IMMUNOREGULATORY INTERACTIONS BETWEEN A LYMPHOID AND A NON LYMPHOID CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 4.3 | 143.0 | REACTOME RIG I MDA5 MEDIATED INDUCTION OF IFN ALPHA BETA PATHWAYS | Genes involved in RIG-I/MDA5 mediated induction of IFN-alpha/beta pathways |
| 4.3 | 34.2 | REACTOME ADVANCED GLYCOSYLATION ENDPRODUCT RECEPTOR SIGNALING | Genes involved in Advanced glycosylation endproduct receptor signaling |
| 4.1 | 20.5 | REACTOME P53 INDEPENDENT G1 S DNA DAMAGE CHECKPOINT | Genes involved in p53-Independent G1/S DNA damage checkpoint |
| 3.6 | 138.0 | REACTOME INTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Intrinsic Pathway for Apoptosis |
| 3.5 | 55.8 | REACTOME TGF BETA RECEPTOR SIGNALING IN EMT EPITHELIAL TO MESENCHYMAL TRANSITION | Genes involved in TGF-beta receptor signaling in EMT (epithelial to mesenchymal transition) |
| 3.1 | 18.8 | REACTOME DIGESTION OF DIETARY CARBOHYDRATE | Genes involved in Digestion of dietary carbohydrate |
| 3.1 | 49.5 | REACTOME ACYL CHAIN REMODELLING OF PG | Genes involved in Acyl chain remodelling of PG |
| 2.5 | 22.2 | REACTOME GAMMA CARBOXYLATION TRANSPORT AND AMINO TERMINAL CLEAVAGE OF PROTEINS | Genes involved in Gamma-carboxylation, transport, and amino-terminal cleavage of proteins |
| 2.3 | 25.8 | REACTOME TAK1 ACTIVATES NFKB BY PHOSPHORYLATION AND ACTIVATION OF IKKS COMPLEX | Genes involved in TAK1 activates NFkB by phosphorylation and activation of IKKs complex |
| 1.9 | 11.7 | REACTOME CALNEXIN CALRETICULIN CYCLE | Genes involved in Calnexin/calreticulin cycle |
| 1.9 | 108.7 | REACTOME AMINO ACID AND OLIGOPEPTIDE SLC TRANSPORTERS | Genes involved in Amino acid and oligopeptide SLC transporters |
| 1.8 | 62.3 | REACTOME TRANSPORT TO THE GOLGI AND SUBSEQUENT MODIFICATION | Genes involved in Transport to the Golgi and subsequent modification |
| 1.8 | 35.9 | REACTOME PRE NOTCH TRANSCRIPTION AND TRANSLATION | Genes involved in Pre-NOTCH Transcription and Translation |
| 1.7 | 19.2 | REACTOME CREATION OF C4 AND C2 ACTIVATORS | Genes involved in Creation of C4 and C2 activators |
| 1.7 | 20.9 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 1.7 | 18.2 | REACTOME UNWINDING OF DNA | Genes involved in Unwinding of DNA |
| 1.6 | 24.4 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 7ALPHA HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 7alpha-hydroxycholesterol |
| 1.6 | 25.0 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 1.5 | 266.7 | REACTOME CLASS I MHC MEDIATED ANTIGEN PROCESSING PRESENTATION | Genes involved in Class I MHC mediated antigen processing & presentation |
| 1.4 | 32.1 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 1.4 | 23.0 | REACTOME INTRINSIC PATHWAY | Genes involved in Intrinsic Pathway |
| 1.4 | 134.1 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 1.3 | 38.1 | REACTOME RECYCLING PATHWAY OF L1 | Genes involved in Recycling pathway of L1 |
| 1.3 | 34.0 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 1.2 | 29.9 | REACTOME CHONDROITIN SULFATE BIOSYNTHESIS | Genes involved in Chondroitin sulfate biosynthesis |
| 1.2 | 33.6 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 1.2 | 4.7 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |
| 1.0 | 24.2 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 1.0 | 60.6 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 1.0 | 14.6 | REACTOME MRNA DECAY BY 5 TO 3 EXORIBONUCLEASE | Genes involved in mRNA Decay by 5' to 3' Exoribonuclease |
| 0.9 | 9.0 | REACTOME PURINE CATABOLISM | Genes involved in Purine catabolism |
| 0.9 | 10.6 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.9 | 133.7 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.8 | 10.2 | REACTOME PASSIVE TRANSPORT BY AQUAPORINS | Genes involved in Passive Transport by Aquaporins |
| 0.8 | 9.7 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.8 | 10.4 | REACTOME GRB2 SOS PROVIDES LINKAGE TO MAPK SIGNALING FOR INTERGRINS | Genes involved in GRB2:SOS provides linkage to MAPK signaling for Intergrins |
| 0.8 | 8.6 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |
| 0.8 | 10.9 | REACTOME RAP1 SIGNALLING | Genes involved in Rap1 signalling |
| 0.8 | 15.3 | REACTOME LYSOSOME VESICLE BIOGENESIS | Genes involved in Lysosome Vesicle Biogenesis |
| 0.7 | 5.2 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 0.7 | 17.8 | REACTOME DEPOSITION OF NEW CENPA CONTAINING NUCLEOSOMES AT THE CENTROMERE | Genes involved in Deposition of New CENPA-containing Nucleosomes at the Centromere |
| 0.7 | 5.8 | REACTOME TRANSPORT OF MATURE MRNA DERIVED FROM AN INTRONLESS TRANSCRIPT | Genes involved in Transport of Mature mRNA Derived from an Intronless Transcript |
| 0.7 | 18.6 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.7 | 18.6 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.7 | 19.1 | REACTOME ANTIGEN ACTIVATES B CELL RECEPTOR LEADING TO GENERATION OF SECOND MESSENGERS | Genes involved in Antigen Activates B Cell Receptor Leading to Generation of Second Messengers |
| 0.7 | 5.6 | REACTOME SLBP DEPENDENT PROCESSING OF REPLICATION DEPENDENT HISTONE PRE MRNAS | Genes involved in SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs |
| 0.7 | 11.2 | REACTOME DOWNREGULATION OF SMAD2 3 SMAD4 TRANSCRIPTIONAL ACTIVITY | Genes involved in Downregulation of SMAD2/3:SMAD4 transcriptional activity |
| 0.7 | 11.2 | REACTOME G1 S SPECIFIC TRANSCRIPTION | Genes involved in G1/S-Specific Transcription |
| 0.7 | 11.7 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.7 | 46.9 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.7 | 18.8 | REACTOME RORA ACTIVATES CIRCADIAN EXPRESSION | Genes involved in RORA Activates Circadian Expression |
| 0.7 | 6.7 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 0.7 | 4.0 | REACTOME ACTIVATION OF CHAPERONES BY ATF6 ALPHA | Genes involved in Activation of Chaperones by ATF6-alpha |
| 0.6 | 30.5 | REACTOME ANTIVIRAL MECHANISM BY IFN STIMULATED GENES | Genes involved in Antiviral mechanism by IFN-stimulated genes |
| 0.6 | 7.2 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GIP | Genes involved in Synthesis, Secretion, and Inactivation of Glucose-dependent Insulinotropic Polypeptide (GIP) |
| 0.6 | 6.8 | REACTOME ANDROGEN BIOSYNTHESIS | Genes involved in Androgen biosynthesis |
| 0.6 | 6.2 | REACTOME SYNTHESIS OF PIPS AT THE LATE ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the late endosome membrane |
| 0.6 | 23.6 | REACTOME COSTIMULATION BY THE CD28 FAMILY | Genes involved in Costimulation by the CD28 family |
| 0.5 | 5.3 | REACTOME FACILITATIVE NA INDEPENDENT GLUCOSE TRANSPORTERS | Genes involved in Facilitative Na+-independent glucose transporters |
| 0.5 | 13.6 | REACTOME ACTIVATION OF GENES BY ATF4 | Genes involved in Activation of Genes by ATF4 |
| 0.5 | 2.9 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.5 | 13.1 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |
| 0.5 | 15.4 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.4 | 3.6 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.4 | 8.3 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.4 | 5.6 | REACTOME SEMA4D INDUCED CELL MIGRATION AND GROWTH CONE COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.4 | 9.4 | REACTOME CIRCADIAN CLOCK | Genes involved in Circadian Clock |
| 0.4 | 12.0 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.4 | 4.9 | REACTOME G0 AND EARLY G1 | Genes involved in G0 and Early G1 |
| 0.3 | 7.3 | REACTOME PYRIMIDINE METABOLISM | Genes involved in Pyrimidine metabolism |
| 0.3 | 2.3 | REACTOME DOWNSTREAM SIGNAL TRANSDUCTION | Genes involved in Downstream signal transduction |
| 0.3 | 15.6 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.3 | 3.2 | REACTOME REGULATION OF MRNA STABILITY BY PROTEINS THAT BIND AU RICH ELEMENTS | Genes involved in Regulation of mRNA Stability by Proteins that Bind AU-rich Elements |
| 0.3 | 29.2 | REACTOME MRNA SPLICING | Genes involved in mRNA Splicing |
| 0.3 | 17.3 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.3 | 2.8 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 0.2 | 1.4 | REACTOME BASIGIN INTERACTIONS | Genes involved in Basigin interactions |
| 0.2 | 4.1 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.2 | 11.7 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.1 | 3.0 | REACTOME IL1 SIGNALING | Genes involved in Interleukin-1 signaling |
| 0.1 | 2.6 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.1 | 1.7 | REACTOME ACTIVATED AMPK STIMULATES FATTY ACID OXIDATION IN MUSCLE | Genes involved in Activated AMPK stimulates fatty-acid oxidation in muscle |
| 0.1 | 1.5 | REACTOME INWARDLY RECTIFYING K CHANNELS | Genes involved in Inwardly rectifying K+ channels |
| 0.1 | 2.3 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |
| 0.1 | 1.4 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.1 | 0.8 | REACTOME AMINE DERIVED HORMONES | Genes involved in Amine-derived hormones |
| 0.1 | 1.2 | REACTOME FANCONI ANEMIA PATHWAY | Genes involved in Fanconi Anemia pathway |
| 0.0 | 0.7 | REACTOME POL SWITCHING | Genes involved in Polymerase switching |
| 0.0 | 0.4 | REACTOME RNA POL I TRANSCRIPTION INITIATION | Genes involved in RNA Polymerase I Transcription Initiation |