Project
GSE53960: rat RNA-Seq transcriptomic Bodymap
Navigation
Downloads
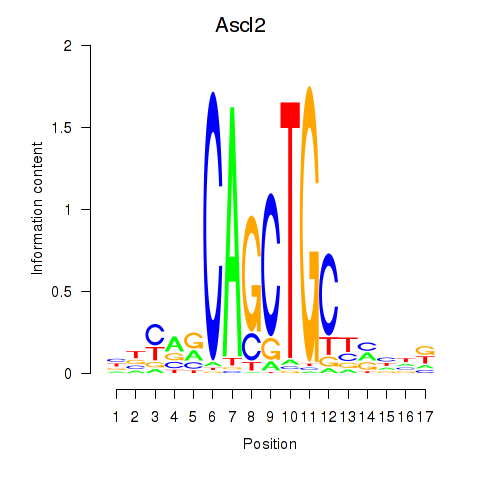
Results for Ascl2
Z-value: 1.17
Transcription factors associated with Ascl2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Ascl2
|
ENSRNOG00000020434 | achaete-scute family bHLH transcription factor 2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Ascl2 | rn6_v1_chr1_-_216156409_216156409 | -0.15 | 8.0e-03 | Click! |
Activity profile of Ascl2 motif
Sorted Z-values of Ascl2 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr10_+_87774552 | 71.68 |
ENSRNOT00000044342
|
Krtap9-1
|
keratin associated protein 9-1 |
| chr1_+_100199057 | 61.16 |
ENSRNOT00000025831
|
Klk1
|
kallikrein 1 |
| chr10_+_87759769 | 50.05 |
ENSRNOT00000017378
ENSRNOT00000046526 |
Krtap9-1
|
keratin associated protein 9-1 |
| chr10_+_87782376 | 42.04 |
ENSRNOT00000017415
|
LOC680396
|
hypothetical protein LOC680396 |
| chr12_+_24761210 | 40.28 |
ENSRNOT00000002003
|
Cldn4
|
claudin 4 |
| chr3_+_119015412 | 38.49 |
ENSRNOT00000013605
|
Slc27a2
|
solute carrier family 27 member 2 |
| chr19_-_43911057 | 32.32 |
ENSRNOT00000026017
|
Ctrb1
|
chymotrypsinogen B1 |
| chr5_-_151459037 | 32.23 |
ENSRNOT00000064472
ENSRNOT00000087836 |
Sytl1
|
synaptotagmin-like 1 |
| chr10_+_87788458 | 31.54 |
ENSRNOT00000042020
|
LOC100910814
|
keratin-associated protein 9-1-like |
| chr3_+_119014620 | 31.20 |
ENSRNOT00000067700
|
Slc27a2
|
solute carrier family 27 member 2 |
| chr4_-_10517832 | 31.11 |
ENSRNOT00000039953
ENSRNOT00000083964 |
Gsap
|
gamma-secretase activating protein |
| chr10_-_87564327 | 26.67 |
ENSRNOT00000064760
ENSRNOT00000068237 |
LOC680160
|
similar to keratin associated protein 4-7 |
| chr9_+_9721105 | 26.49 |
ENSRNOT00000073042
ENSRNOT00000075494 |
C3
|
complement C3 |
| chr6_-_138508753 | 26.06 |
ENSRNOT00000006888
|
Ighm
|
immunoglobulin heavy constant mu |
| chr17_-_43543172 | 25.47 |
ENSRNOT00000080684
ENSRNOT00000029626 ENSRNOT00000082719 |
Slc17a3
|
solute carrier family 17 member 3 |
| chr1_-_8751198 | 25.10 |
ENSRNOT00000030511
|
Adgrg6
|
adhesion G protein-coupled receptor G6 |
| chr1_-_260638816 | 24.87 |
ENSRNOT00000065632
ENSRNOT00000017938 ENSRNOT00000077962 |
Pik3ap1
|
phosphoinositide-3-kinase adaptor protein 1 |
| chr3_-_172537877 | 24.59 |
ENSRNOT00000072069
|
Ctsz
|
cathepsin Z |
| chr11_-_90406797 | 24.31 |
ENSRNOT00000073049
|
Snai2
|
snail family transcriptional repressor 2 |
| chr4_-_157252565 | 23.90 |
ENSRNOT00000079947
|
Ptpn6
|
protein tyrosine phosphatase, non-receptor type 6 |
| chr14_-_80973456 | 23.25 |
ENSRNOT00000013257
|
Hgfac
|
HGF activator |
| chr4_-_157252104 | 22.70 |
ENSRNOT00000082739
|
Ptpn6
|
protein tyrosine phosphatase, non-receptor type 6 |
| chr12_-_21832813 | 22.60 |
ENSRNOT00000075280
|
Cldn3
|
claudin 3 |
| chr1_+_101214593 | 22.46 |
ENSRNOT00000028086
|
Tead2
|
TEA domain transcription factor 2 |
| chr13_-_91427575 | 22.32 |
ENSRNOT00000012092
|
Apcs
|
amyloid P component, serum |
| chr14_+_3058993 | 21.90 |
ENSRNOT00000002807
|
Gfi1
|
growth factor independent 1 transcriptional repressor |
| chr18_+_56071478 | 21.80 |
ENSRNOT00000025344
ENSRNOT00000025354 |
Cd74
|
CD74 molecule |
| chr2_+_32820322 | 21.65 |
ENSRNOT00000013768
|
Cd180
|
CD180 molecule |
| chr8_-_50539331 | 20.73 |
ENSRNOT00000088997
|
AABR07073400.1
|
|
| chr7_+_94130852 | 20.67 |
ENSRNOT00000011485
|
Mal2
|
mal, T-cell differentiation protein 2 |
| chr1_-_53087474 | 20.33 |
ENSRNOT00000017302
|
Ccr6
|
C-C motif chemokine receptor 6 |
| chr1_+_86938138 | 19.94 |
ENSRNOT00000075601
|
Ccer2
|
coiled-coil glutamate-rich protein 2 |
| chr6_-_141291347 | 19.53 |
ENSRNOT00000008333
|
AABR07065789.1
|
|
| chr1_+_203160323 | 19.35 |
ENSRNOT00000027919
|
AABR07005844.1
|
|
| chr5_-_155772040 | 19.27 |
ENSRNOT00000036788
|
Cela3b
|
chymotrypsin-like elastase family, member 3B |
| chr3_-_60166013 | 19.20 |
ENSRNOT00000024922
|
Wipf1
|
WAS/WASL interacting protein family, member 1 |
| chr14_-_45797887 | 18.63 |
ENSRNOT00000029209
|
Tbc1d1
|
TBC1 domain family member 1 |
| chr16_-_19399851 | 18.48 |
ENSRNOT00000089056
ENSRNOT00000021073 |
Tpm4
|
tropomyosin 4 |
| chr12_+_2180150 | 18.41 |
ENSRNOT00000001322
|
Stxbp2
|
syntaxin binding protein 2 |
| chr3_+_19772056 | 18.41 |
ENSRNOT00000044455
|
AABR07051708.1
|
|
| chr11_-_38088753 | 18.13 |
ENSRNOT00000002713
|
Tmprss2
|
transmembrane protease, serine 2 |
| chr7_-_107634287 | 18.01 |
ENSRNOT00000093672
ENSRNOT00000087116 |
Sla
|
src-like adaptor |
| chr7_-_71226150 | 17.89 |
ENSRNOT00000005875
|
RGD1561812
|
similar to Retinol dehydrogenase type II (RODH II) (29 k-protein) |
| chr1_-_141188031 | 17.85 |
ENSRNOT00000044567
|
Polg
|
DNA polymerase gamma, catalytic subunit |
| chr19_-_19727081 | 17.74 |
ENSRNOT00000020700
|
Adcy7
|
adenylate cyclase 7 |
| chr7_-_143016040 | 17.56 |
ENSRNOT00000029697
|
Krt80
|
keratin 80 |
| chr9_-_9675110 | 17.07 |
ENSRNOT00000073294
|
Vav1
|
vav guanine nucleotide exchange factor 1 |
| chr2_+_54466280 | 16.99 |
ENSRNOT00000033112
|
C6
|
complement C6 |
| chr6_-_142060032 | 16.82 |
ENSRNOT00000064717
|
AABR07065811.1
|
|
| chr10_+_86399827 | 16.80 |
ENSRNOT00000009299
|
Grb7
|
growth factor receptor bound protein 7 |
| chr7_-_3386522 | 16.79 |
ENSRNOT00000010760
|
Mettl7b
|
methyltransferase like 7B |
| chr2_-_202816562 | 16.49 |
ENSRNOT00000020401
|
Fam46c
|
family with sequence similarity 46, member C |
| chr6_-_143065639 | 16.47 |
ENSRNOT00000070923
|
AABR07065827.1
|
|
| chr5_+_149047681 | 16.31 |
ENSRNOT00000015198
|
Laptm5
|
lysosomal protein transmembrane 5 |
| chr3_-_81282157 | 16.18 |
ENSRNOT00000051258
|
Large2
|
LARGE xylosyl- and glucuronyltransferase 2 |
| chr13_+_89805962 | 16.06 |
ENSRNOT00000074035
|
Tstd1
|
thiosulfate sulfurtransferase like domain containing 1 |
| chr9_-_52912293 | 16.04 |
ENSRNOT00000005228
|
Slc40a1
|
solute carrier family 40 member 1 |
| chr6_-_138550576 | 15.97 |
ENSRNOT00000075284
|
AABR07065645.1
|
|
| chr6_+_139158334 | 15.89 |
ENSRNOT00000089227
|
AABR07065673.1
|
|
| chr1_+_226687258 | 15.87 |
ENSRNOT00000079679
|
Vwce
|
von Willebrand factor C and EGF domains |
| chr12_-_22127021 | 15.83 |
ENSRNOT00000076261
|
Sap25
|
Sin3A-associated protein 25 |
| chr12_-_22126350 | 15.69 |
ENSRNOT00000076328
|
Sap25
|
Sin3A-associated protein 25 |
| chr7_-_51353068 | 15.33 |
ENSRNOT00000008222
|
Pawr
|
pro-apoptotic WT1 regulator |
| chr1_-_54748763 | 15.31 |
ENSRNOT00000074549
|
LOC100911027
|
protein MAL2-like |
| chr1_-_75154325 | 15.03 |
ENSRNOT00000074520
|
RGD1564801
|
similar to hepatic multiple inositol polyphosphate phosphatase |
| chr13_-_80745347 | 14.73 |
ENSRNOT00000041908
|
Fmo1
|
flavin containing monooxygenase 1 |
| chr1_+_251045440 | 14.35 |
ENSRNOT00000015017
|
Minpp1
|
multiple inositol-polyphosphate phosphatase 1 |
| chr12_+_25119355 | 14.19 |
ENSRNOT00000034629
|
Lat2
|
linker for activation of T cells family, member 2 |
| chr4_-_28437676 | 14.18 |
ENSRNOT00000012995
|
Hepacam2
|
HEPACAM family member 2 |
| chr12_-_22126858 | 14.15 |
ENSRNOT00000084480
|
Sap25
|
Sin3A-associated protein 25 |
| chr11_-_11585765 | 14.02 |
ENSRNOT00000066439
|
Robo2
|
roundabout guidance receptor 2 |
| chr9_-_14668297 | 13.99 |
ENSRNOT00000042404
|
Treml2
|
triggering receptor expressed on myeloid cells-like 2 |
| chr5_+_142845114 | 13.97 |
ENSRNOT00000039870
|
Yrdc
|
yrdC N(6)-threonylcarbamoyltransferase domain containing |
| chr14_-_82287706 | 13.91 |
ENSRNOT00000080695
|
Fgfr3
|
fibroblast growth factor receptor 3 |
| chr19_+_25391700 | 13.86 |
ENSRNOT00000011163
|
LOC100359539
|
ribonucleotide reductase M2 polypeptide |
| chr6_-_141488290 | 13.79 |
ENSRNOT00000067336
|
AABR07065792.1
|
|
| chr4_+_157126935 | 13.69 |
ENSRNOT00000056051
|
C1r
|
complement C1r |
| chr4_-_89695928 | 13.68 |
ENSRNOT00000039316
|
Gprin3
|
GPRIN family member 3 |
| chr4_-_170620703 | 13.59 |
ENSRNOT00000011930
|
Plbd1
|
phospholipase B domain containing 1 |
| chr10_+_103713045 | 13.32 |
ENSRNOT00000004351
|
Slc9a3r1
|
SLC9A3 regulator 1 |
| chr1_-_199439210 | 13.29 |
ENSRNOT00000026699
|
Pycard
|
PYD and CARD domain containing |
| chr5_-_134526089 | 12.83 |
ENSRNOT00000013321
|
Cyp4b1
|
cytochrome P450, family 4, subfamily b, polypeptide 1 |
| chr7_-_3342491 | 12.74 |
ENSRNOT00000081756
|
Rdh5
|
retinol dehydrogenase 5 |
| chr1_+_22332090 | 12.63 |
ENSRNOT00000091252
|
AABR07000663.1
|
trace amine-associated receptor 8c (Taar8c), mRNA |
| chrX_+_70461718 | 12.63 |
ENSRNOT00000078233
ENSRNOT00000003789 |
Kif4a
|
kinesin family member 4A |
| chr7_-_107616038 | 12.52 |
ENSRNOT00000088752
|
Sla
|
src-like adaptor |
| chr1_+_85003280 | 12.41 |
ENSRNOT00000057122
|
Fcgbp
|
Fc fragment of IgG binding protein |
| chr20_+_5446104 | 12.29 |
ENSRNOT00000000550
|
B3galt4
|
Beta-1,3-galactosyltransferase 4 |
| chr16_+_20426566 | 12.26 |
ENSRNOT00000026225
|
Ifi30
|
IFI30, lysosomal thiol reductase |
| chr16_+_20555395 | 12.26 |
ENSRNOT00000026652
|
Gdf15
|
growth differentiation factor 15 |
| chr16_+_20110148 | 12.25 |
ENSRNOT00000080146
ENSRNOT00000025312 |
Jak3
|
Janus kinase 3 |
| chr2_-_219262901 | 12.22 |
ENSRNOT00000037068
|
Gpr88
|
G-protein coupled receptor 88 |
| chr19_+_37476095 | 12.05 |
ENSRNOT00000092794
ENSRNOT00000023130 |
Hsd11b2
|
hydroxysteroid 11-beta dehydrogenase 2 |
| chr9_-_71445739 | 12.05 |
ENSRNOT00000019698
|
Fzd5
|
frizzled class receptor 5 |
| chr1_+_145715969 | 11.95 |
ENSRNOT00000037996
|
Tmc3
|
transmembrane channel-like 3 |
| chr17_+_76306585 | 11.88 |
ENSRNOT00000065978
|
Dhtkd1
|
dehydrogenase E1 and transketolase domain containing 1 |
| chr1_+_137014272 | 11.84 |
ENSRNOT00000014802
|
Akap13
|
A-kinase anchoring protein 13 |
| chr14_-_33150509 | 11.76 |
ENSRNOT00000002837
|
Rest
|
RE1-silencing transcription factor |
| chr8_+_5993941 | 11.44 |
ENSRNOT00000014065
|
Tmem123
|
transmembrane protein 123 |
| chr3_+_164822111 | 11.43 |
ENSRNOT00000014568
|
Pard6b
|
par-6 family cell polarity regulator beta |
| chr17_-_43776460 | 11.41 |
ENSRNOT00000089055
|
Hist2h3c2
|
histone cluster 2, H3c2 |
| chr1_-_98570949 | 11.40 |
ENSRNOT00000033648
|
Siglec5
|
sialic acid binding Ig-like lectin 5 |
| chr6_-_138536321 | 11.35 |
ENSRNOT00000077743
|
AABR07065643.1
|
|
| chr6_+_10912383 | 11.23 |
ENSRNOT00000061747
ENSRNOT00000086247 |
Ttc7a
|
tetratricopeptide repeat domain 7A |
| chr6_-_138744480 | 11.12 |
ENSRNOT00000089387
|
AABR07065651.5
|
|
| chr20_-_22004209 | 11.08 |
ENSRNOT00000086250
ENSRNOT00000068778 |
Rtkn2
|
rhotekin 2 |
| chr1_+_107344904 | 10.99 |
ENSRNOT00000082582
|
Gas2
|
growth arrest-specific 2 |
| chr1_-_213987053 | 10.98 |
ENSRNOT00000072774
|
LOC100911519
|
p53-induced protein with a death domain-like |
| chr9_+_100285804 | 10.93 |
ENSRNOT00000079305
|
Agxt
|
alanine-glyoxylate aminotransferase |
| chr4_+_14001761 | 10.81 |
ENSRNOT00000076519
|
Cd36
|
CD36 molecule |
| chr1_+_192233910 | 10.81 |
ENSRNOT00000016418
ENSRNOT00000016442 |
Prkcb
|
protein kinase C, beta |
| chr10_-_31419235 | 10.76 |
ENSRNOT00000059496
|
Cyfip2
|
cytoplasmic FMR1 interacting protein 2 |
| chr9_+_94425252 | 10.56 |
ENSRNOT00000064965
ENSRNOT00000076099 |
Gigyf2
|
GRB10 interacting GYF protein 2 |
| chr4_-_100099517 | 10.43 |
ENSRNOT00000014277
|
Atoh8
|
atonal bHLH transcription factor 8 |
| chr20_+_8165307 | 10.31 |
ENSRNOT00000000637
|
Pim1
|
Pim-1 proto-oncogene, serine/threonine kinase |
| chr4_-_180505916 | 10.29 |
ENSRNOT00000086465
|
AABR07062512.1
|
|
| chr3_+_161298962 | 10.21 |
ENSRNOT00000066028
|
Ctsa
|
cathepsin A |
| chr1_-_169463760 | 10.19 |
ENSRNOT00000023100
|
Trim30c
|
tripartite motif-containing 30C |
| chr4_-_62438958 | 10.16 |
ENSRNOT00000014010
|
Wdr91
|
WD repeat domain 91 |
| chr20_+_3677474 | 10.03 |
ENSRNOT00000047663
|
Mt1m
|
metallothionein 1M |
| chr5_-_155258392 | 9.99 |
ENSRNOT00000017065
|
C1qc
|
complement C1q C chain |
| chr11_+_17538063 | 9.98 |
ENSRNOT00000031889
ENSRNOT00000090878 |
Chodl
|
chondrolectin |
| chr1_-_42971208 | 9.95 |
ENSRNOT00000088535
|
Gm5414
|
predicted gene 5414 |
| chr5_+_157222636 | 9.92 |
ENSRNOT00000022579
|
Pla2g2d
|
phospholipase A2, group IID |
| chr3_-_46621873 | 9.92 |
ENSRNOT00000011003
|
Pla2r1
|
phospholipase A2 receptor 1 |
| chr2_-_29121104 | 9.86 |
ENSRNOT00000020543
|
Tnpo1
|
transportin 1 |
| chr20_-_3397039 | 9.82 |
ENSRNOT00000001084
ENSRNOT00000085259 |
Ppp1r18
|
protein phosphatase 1, regulatory subunit 18 |
| chr8_-_62616828 | 9.76 |
ENSRNOT00000068340
|
Arid3b
|
AT-rich interaction domain 3B |
| chr1_-_64446818 | 9.73 |
ENSRNOT00000081980
|
Myadm
|
myeloid-associated differentiation marker |
| chr10_+_14122878 | 9.72 |
ENSRNOT00000052008
|
Hs3st6
|
heparan sulfate-glucosamine 3-sulfotransferase 6 |
| chr14_+_13192347 | 9.70 |
ENSRNOT00000000092
|
Antxr2
|
anthrax toxin receptor 2 |
| chr1_-_88176610 | 9.62 |
ENSRNOT00000032394
|
LOC100909725
|
potassium channel subfamily K member 6-like |
| chr8_-_61917125 | 9.61 |
ENSRNOT00000085049
|
RGD1305464
|
similar to human chromosome 15 open reading frame 39 |
| chr17_+_43734461 | 9.59 |
ENSRNOT00000072564
|
Hist1h1d
|
histone cluster 1, H1d |
| chr12_+_9728486 | 9.59 |
ENSRNOT00000001263
|
Lnx2
|
ligand of numb-protein X 2 |
| chr8_+_132828091 | 9.51 |
ENSRNOT00000008269
|
Ccr9
|
C-C motif chemokine receptor 9 |
| chr4_+_88834066 | 9.32 |
ENSRNOT00000009546
|
Abcg2
|
ATP-binding cassette, subfamily G (WHITE), member 2 |
| chr2_-_88113029 | 9.31 |
ENSRNOT00000013354
|
Car2
|
carbonic anhydrase 2 |
| chr12_+_22165486 | 9.24 |
ENSRNOT00000001890
|
Mospd3
|
motile sperm domain containing 3 |
| chr19_-_11341863 | 9.24 |
ENSRNOT00000025694
|
Mt4
|
metallothionein 4 |
| chr8_+_40078269 | 9.10 |
ENSRNOT00000044293
|
Siae
|
sialic acid acetylesterase |
| chr10_+_7041510 | 9.03 |
ENSRNOT00000003514
|
Carhsp1
|
calcium regulated heat stable protein 1 |
| chr7_+_12146642 | 9.03 |
ENSRNOT00000085899
|
Tcf3
|
transcription factor 3 |
| chr7_-_116920507 | 9.02 |
ENSRNOT00000048363
|
Mroh6
|
maestro heat-like repeat family member 6 |
| chr8_+_22368745 | 9.02 |
ENSRNOT00000049973
|
Slc44a2
|
solute carrier family 44 member 2 |
| chr14_-_84189266 | 8.98 |
ENSRNOT00000005934
|
Tcn2
|
transcobalamin 2 |
| chr16_+_74531564 | 8.94 |
ENSRNOT00000078971
|
Slc25a15
|
solute carrier family 25 member 15 |
| chr7_-_67116980 | 8.93 |
ENSRNOT00000005798
|
Ppm1h
|
protein phosphatase, Mg2+/Mn2+ dependent, 1H |
| chr6_-_139654508 | 8.91 |
ENSRNOT00000082576
|
AABR07065705.5
|
|
| chr3_+_7422820 | 8.86 |
ENSRNOT00000064323
|
Ddx31
|
DEAD-box helicase 31 |
| chr18_+_60392376 | 8.85 |
ENSRNOT00000023890
|
Nedd4l
|
neural precursor cell expressed, developmentally down-regulated 4-like, E3 ubiquitin protein ligase |
| chr6_-_138772894 | 8.82 |
ENSRNOT00000080779
|
AABR07065651.1
|
|
| chr7_-_119996824 | 8.81 |
ENSRNOT00000011079
|
Mfng
|
MFNG O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase |
| chr3_-_51297852 | 8.75 |
ENSRNOT00000001607
|
Cobll1
|
cordon-bleu WH2 repeat protein-like 1 |
| chr7_-_2941122 | 8.73 |
ENSRNOT00000082107
|
Esyt1
|
extended synaptotagmin 1 |
| chr10_-_39373437 | 8.65 |
ENSRNOT00000058907
|
Slc22a5
|
solute carrier family 22 member 5 |
| chr13_+_113373578 | 8.61 |
ENSRNOT00000009900
|
Plxna2
|
plexin A2 |
| chr6_-_24563246 | 8.51 |
ENSRNOT00000074294
|
LOC685881
|
hypothetical protein LOC685881 |
| chr7_-_3102142 | 8.49 |
ENSRNOT00000008018
|
Suox
|
sulfite oxidase |
| chr10_-_4910305 | 8.47 |
ENSRNOT00000033122
|
Rmi2
|
RecQ mediated genome instability 2 |
| chr1_+_85213652 | 8.41 |
ENSRNOT00000092044
|
Nccrp1
|
non-specific cytotoxic cell receptor protein 1 homolog (zebrafish) |
| chr4_-_155923079 | 8.38 |
ENSRNOT00000013308
|
Clec4a3
|
C-type lectin domain family 4, member A3 |
| chr8_+_48716939 | 8.32 |
ENSRNOT00000015810
|
Slc37a4
|
solute carrier family 37 member 4 |
| chr9_+_20048121 | 8.32 |
ENSRNOT00000014791
|
Mep1a
|
meprin 1 subunit alpha |
| chr6_+_43884678 | 8.26 |
ENSRNOT00000091551
|
Rrm2
|
ribonucleotide reductase regulatory subunit M2 |
| chr19_+_30936703 | 8.24 |
ENSRNOT00000024568
|
Smarca5
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 5 |
| chrX_+_123404518 | 8.15 |
ENSRNOT00000015085
|
Slc25a5
|
solute carrier family 25 member 5 |
| chr6_-_80334522 | 8.10 |
ENSRNOT00000059316
|
Fbxo33
|
F-box protein 33 |
| chr6_-_138536162 | 8.07 |
ENSRNOT00000083031
|
AABR07065643.1
|
|
| chr18_+_24708115 | 8.05 |
ENSRNOT00000061054
|
Lims2
|
LIM zinc finger domain containing 2 |
| chr17_+_76079720 | 8.04 |
ENSRNOT00000073933
|
Proser2
|
proline and serine rich 2 |
| chr1_-_214423881 | 8.01 |
ENSRNOT00000025290
|
Pidd1
|
p53-induced death domain protein 1 |
| chr6_-_141008427 | 7.94 |
ENSRNOT00000074472
|
AABR07065778.2
|
|
| chr1_-_252808380 | 7.94 |
ENSRNOT00000025856
|
Ch25h
|
cholesterol 25-hydroxylase |
| chrX_+_80213332 | 7.93 |
ENSRNOT00000042827
|
Sh3bgrl
|
SH3 domain binding glutamate-rich protein like |
| chr11_-_53575060 | 7.89 |
ENSRNOT00000050142
ENSRNOT00000078221 ENSRNOT00000078434 |
Cd47
|
Cd47 molecule |
| chr2_-_112831476 | 7.88 |
ENSRNOT00000018055
|
Ect2
|
epithelial cell transforming 2 |
| chr16_+_69048730 | 7.86 |
ENSRNOT00000086082
ENSRNOT00000078128 |
Rab11fip1
|
RAB11 family interacting protein 1 |
| chr6_-_140418831 | 7.86 |
ENSRNOT00000086301
|
AABR07065768.2
|
|
| chr14_+_84306466 | 7.80 |
ENSRNOT00000006116
|
Sec14l4
|
SEC14-like lipid binding 4 |
| chr7_-_75597087 | 7.78 |
ENSRNOT00000080676
|
Ywhaz
|
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, zeta |
| chr6_-_140715174 | 7.71 |
ENSRNOT00000085345
|
AABR07065773.1
|
|
| chr4_-_115157263 | 7.67 |
ENSRNOT00000015296
|
Tet3
|
tet methylcytosine dioxygenase 3 |
| chr15_+_30612409 | 7.60 |
ENSRNOT00000072977
|
AABR07017748.1
|
|
| chr20_+_5441876 | 7.56 |
ENSRNOT00000092476
|
Rps18
|
ribosomal protein S18 |
| chr16_-_41234095 | 7.56 |
ENSRNOT00000000120
|
Aga
|
aspartylglucosaminidase |
| chr8_-_122841477 | 7.56 |
ENSRNOT00000014861
|
Cmtm7
|
CKLF-like MARVEL transmembrane domain containing 7 |
| chr13_-_52752997 | 7.52 |
ENSRNOT00000013531
|
Pkp1
|
plakophilin 1 |
| chr17_+_18029124 | 7.46 |
ENSRNOT00000022085
|
Tpmt
|
thiopurine S-methyltransferase |
| chr2_-_34185682 | 7.42 |
ENSRNOT00000066925
ENSRNOT00000082755 |
Nln
|
neurolysin |
| chr14_-_46054022 | 7.38 |
ENSRNOT00000002982
|
LOC498368
|
similar to RIKEN cDNA 0610040J01 |
| chr3_+_20163337 | 7.37 |
ENSRNOT00000075136
|
AABR07051726.1
|
|
| chr1_-_101741441 | 7.37 |
ENSRNOT00000028570
|
Sult2b1
|
sulfotransferase family 2B member 1 |
| chr2_+_251863069 | 7.33 |
ENSRNOT00000036282
|
Syde2
|
synapse defective Rho GTPase homolog 2 |
| chr1_-_32272476 | 7.31 |
ENSRNOT00000022683
|
Tert
|
telomerase reverse transcriptase |
| chr17_+_76079907 | 7.30 |
ENSRNOT00000092710
|
Proser2
|
proline and serine rich 2 |
| chr14_-_8510138 | 7.25 |
ENSRNOT00000080758
|
Arhgap24
|
Rho GTPase activating protein 24 |
| chr5_-_137372524 | 7.22 |
ENSRNOT00000009061
|
Tmem125
|
transmembrane protein 125 |
| chr18_+_70427007 | 7.22 |
ENSRNOT00000087959
ENSRNOT00000019512 |
Myo5b
|
myosin Vb |
| chr20_-_2210033 | 7.20 |
ENSRNOT00000084596
|
Trim26
|
tripartite motif-containing 26 |
Network of associatons between targets according to the STRING database.
First level regulatory network of Ascl2
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 23.2 | 69.7 | GO:0097089 | methyl-branched fatty acid metabolic process(GO:0097089) |
| 14.5 | 43.5 | GO:0001905 | activation of membrane attack complex(GO:0001905) regulation of activation of membrane attack complex(GO:0001969) |
| 11.6 | 46.6 | GO:0033277 | abortive mitotic cell cycle(GO:0033277) |
| 8.5 | 50.7 | GO:0010957 | negative regulation of vitamin D biosynthetic process(GO:0010957) |
| 5.6 | 22.3 | GO:0052203 | modulation of catalytic activity in other organism involved in symbiotic interaction(GO:0052203) modulation by host of symbiont catalytic activity(GO:0052422) regulation of exo-alpha-sialidase activity(GO:1903015) |
| 5.4 | 10.8 | GO:0035408 | histone H3-T6 phosphorylation(GO:0035408) |
| 5.1 | 15.3 | GO:1904457 | hindgut contraction(GO:0043133) regulation of hindgut contraction(GO:0043134) positive regulation of neuronal action potential(GO:1904457) |
| 5.1 | 20.3 | GO:0060474 | positive regulation of sperm motility involved in capacitation(GO:0060474) |
| 4.7 | 14.0 | GO:0002949 | tRNA threonylcarbamoyladenosine modification(GO:0002949) |
| 4.5 | 36.0 | GO:0031580 | membrane raft polarization(GO:0001766) membrane raft distribution(GO:0031580) |
| 4.4 | 13.3 | GO:0034635 | glutathione transport(GO:0034635) tripeptide transport(GO:0042939) |
| 4.4 | 13.3 | GO:0002277 | myeloid dendritic cell activation involved in immune response(GO:0002277) positive regulation of antigen processing and presentation of peptide antigen(GO:0002585) |
| 4.4 | 21.8 | GO:0002906 | mature B cell apoptotic process(GO:0002901) regulation of mature B cell apoptotic process(GO:0002905) negative regulation of mature B cell apoptotic process(GO:0002906) |
| 4.1 | 12.3 | GO:0002731 | negative regulation of dendritic cell cytokine production(GO:0002731) FasL biosynthetic process(GO:0045210) |
| 4.0 | 12.0 | GO:0060061 | Spemann organizer formation(GO:0060061) |
| 4.0 | 16.0 | GO:0070839 | regulation of transcription from RNA polymerase II promoter in response to iron(GO:0034395) divalent metal ion export(GO:0070839) |
| 3.9 | 11.8 | GO:2000705 | regulation of dense core granule biogenesis(GO:2000705) |
| 3.6 | 10.9 | GO:0019265 | glycine biosynthetic process, by transamination of glyoxylate(GO:0019265) oxalic acid secretion(GO:0046724) |
| 3.6 | 21.6 | GO:0031666 | positive regulation of lipopolysaccharide-mediated signaling pathway(GO:0031666) |
| 3.6 | 24.9 | GO:0034154 | toll-like receptor 7 signaling pathway(GO:0034154) |
| 3.5 | 24.6 | GO:0010757 | negative regulation of plasminogen activation(GO:0010757) |
| 3.2 | 9.7 | GO:0090038 | negative regulation of protein kinase C signaling(GO:0090038) |
| 3.2 | 34.8 | GO:0015747 | urate transport(GO:0015747) |
| 3.1 | 9.3 | GO:0042938 | dipeptide transport(GO:0042938) |
| 3.1 | 18.5 | GO:0006287 | base-excision repair, gap-filling(GO:0006287) |
| 3.0 | 11.8 | GO:0071883 | activation of MAPK activity by adrenergic receptor signaling pathway(GO:0071883) |
| 2.9 | 8.6 | GO:0042891 | antibiotic transport(GO:0042891) |
| 2.8 | 14.0 | GO:0050925 | negative regulation of negative chemotaxis(GO:0050925) |
| 2.8 | 13.9 | GO:0061144 | alveolar secondary septum development(GO:0061144) |
| 2.6 | 31.1 | GO:1902993 | positive regulation of beta-amyloid formation(GO:1902004) positive regulation of amyloid precursor protein catabolic process(GO:1902993) |
| 2.5 | 10.0 | GO:0070650 | actin filament bundle distribution(GO:0070650) |
| 2.5 | 7.4 | GO:1902809 | regulation of skeletal muscle fiber differentiation(GO:1902809) |
| 2.4 | 12.2 | GO:0061743 | motor learning(GO:0061743) |
| 2.4 | 7.3 | GO:1900368 | regulation of RNA interference(GO:1900368) |
| 2.4 | 9.7 | GO:0014835 | myoblast differentiation involved in skeletal muscle regeneration(GO:0014835) |
| 2.4 | 7.2 | GO:0006296 | nucleotide-excision repair, DNA incision, 5'-to lesion(GO:0006296) |
| 2.4 | 7.2 | GO:0050968 | detection of chemical stimulus involved in sensory perception of pain(GO:0050968) regulation of establishment of blood-brain barrier(GO:0090210) |
| 2.4 | 4.7 | GO:0072021 | ascending thin limb development(GO:0072021) metanephric ascending thin limb development(GO:0072218) |
| 2.2 | 17.8 | GO:0016584 | nucleosome positioning(GO:0016584) |
| 2.2 | 15.2 | GO:0006610 | ribosomal protein import into nucleus(GO:0006610) |
| 2.1 | 8.5 | GO:0042126 | nitrate metabolic process(GO:0042126) |
| 2.1 | 25.1 | GO:0010578 | regulation of adenylate cyclase activity involved in G-protein coupled receptor signaling pathway(GO:0010578) positive regulation of adenylate cyclase activity involved in G-protein coupled receptor signaling pathway(GO:0010579) |
| 2.0 | 12.3 | GO:0042590 | antigen processing and presentation of exogenous peptide antigen via MHC class I(GO:0042590) |
| 2.0 | 6.1 | GO:0060931 | sinoatrial node cell development(GO:0060931) His-Purkinje system cell differentiation(GO:0060932) |
| 2.0 | 6.0 | GO:0021577 | hindbrain structural organization(GO:0021577) cerebellum structural organization(GO:0021589) |
| 2.0 | 49.0 | GO:0061436 | establishment of skin barrier(GO:0061436) |
| 1.9 | 5.8 | GO:1901423 | response to benzene(GO:1901423) |
| 1.9 | 7.7 | GO:0044725 | chromatin reprogramming in the zygote(GO:0044725) |
| 1.9 | 7.6 | GO:0002337 | B-1a B cell differentiation(GO:0002337) |
| 1.9 | 9.4 | GO:0015755 | fructose transport(GO:0015755) cellular response to fructose stimulus(GO:0071332) |
| 1.8 | 12.8 | GO:0018879 | biphenyl metabolic process(GO:0018879) |
| 1.8 | 14.6 | GO:1990034 | calcium ion export from cell(GO:1990034) |
| 1.8 | 5.4 | GO:1904742 | regulation of telomeric DNA binding(GO:1904742) |
| 1.8 | 12.6 | GO:0051256 | mitotic spindle elongation(GO:0000022) mitotic spindle midzone assembly(GO:0051256) |
| 1.8 | 10.8 | GO:2000334 | blood microparticle formation(GO:0072564) regulation of blood microparticle formation(GO:2000332) positive regulation of blood microparticle formation(GO:2000334) |
| 1.8 | 9.0 | GO:0015889 | cobalamin transport(GO:0015889) |
| 1.8 | 62.9 | GO:0002675 | positive regulation of acute inflammatory response(GO:0002675) |
| 1.8 | 7.2 | GO:0032304 | negative regulation of icosanoid secretion(GO:0032304) |
| 1.8 | 8.9 | GO:0000066 | mitochondrial ornithine transport(GO:0000066) |
| 1.8 | 5.4 | GO:0046985 | positive regulation of hemoglobin biosynthetic process(GO:0046985) |
| 1.7 | 22.6 | GO:0016338 | calcium-independent cell-cell adhesion via plasma membrane cell-adhesion molecules(GO:0016338) |
| 1.7 | 5.1 | GO:0015015 | heparan sulfate proteoglycan biosynthetic process, enzymatic modification(GO:0015015) |
| 1.7 | 10.2 | GO:1904714 | regulation of chaperone-mediated autophagy(GO:1904714) |
| 1.7 | 5.1 | GO:0014707 | branchiomeric skeletal muscle development(GO:0014707) |
| 1.6 | 18.1 | GO:0046598 | positive regulation of viral entry into host cell(GO:0046598) |
| 1.6 | 4.9 | GO:0010424 | DNA methylation on cytosine within a CG sequence(GO:0010424) |
| 1.6 | 25.8 | GO:0009263 | deoxyribonucleotide biosynthetic process(GO:0009263) |
| 1.6 | 7.9 | GO:0070164 | negative regulation of adiponectin secretion(GO:0070164) |
| 1.6 | 4.7 | GO:0018094 | protein polyglycylation(GO:0018094) |
| 1.6 | 7.8 | GO:0090168 | Golgi reassembly(GO:0090168) |
| 1.5 | 6.1 | GO:0060447 | bud outgrowth involved in lung branching(GO:0060447) |
| 1.5 | 6.1 | GO:1905231 | carbohydrate export(GO:0033231) daunorubicin transport(GO:0043215) response to borneol(GO:1905230) cellular response to borneol(GO:1905231) response to codeine(GO:1905233) response to quercetin(GO:1905235) drug transport across blood-brain barrier(GO:1990962) establishment of blood-retinal barrier(GO:1990963) |
| 1.5 | 9.0 | GO:0002326 | B cell lineage commitment(GO:0002326) |
| 1.5 | 10.3 | GO:0031659 | positive regulation of cyclin-dependent protein serine/threonine kinase activity involved in G1/S transition of mitotic cell cycle(GO:0031659) |
| 1.5 | 14.7 | GO:0070995 | NADPH oxidation(GO:0070995) |
| 1.5 | 10.3 | GO:0010216 | maintenance of DNA methylation(GO:0010216) |
| 1.4 | 4.3 | GO:0009826 | unidimensional cell growth(GO:0009826) regulation of establishment or maintenance of cell polarity regulating cell shape(GO:2000769) positive regulation of establishment or maintenance of cell polarity regulating cell shape(GO:2000771) |
| 1.4 | 10.0 | GO:0045650 | negative regulation of macrophage differentiation(GO:0045650) |
| 1.4 | 7.1 | GO:0032020 | ISG15-protein conjugation(GO:0032020) |
| 1.4 | 9.9 | GO:0090403 | positive regulation of arachidonic acid secretion(GO:0090238) oxidative stress-induced premature senescence(GO:0090403) |
| 1.4 | 5.6 | GO:0008078 | mesodermal cell migration(GO:0008078) |
| 1.4 | 8.3 | GO:0035166 | post-embryonic hemopoiesis(GO:0035166) |
| 1.4 | 9.7 | GO:0043686 | co-translational protein modification(GO:0043686) |
| 1.3 | 5.4 | GO:1903225 | negative regulation of endodermal cell differentiation(GO:1903225) |
| 1.3 | 8.0 | GO:2000346 | negative regulation of hepatocyte proliferation(GO:2000346) |
| 1.3 | 2.6 | GO:0090271 | positive regulation of fibroblast growth factor production(GO:0090271) |
| 1.3 | 6.5 | GO:0001582 | detection of chemical stimulus involved in sensory perception of sweet taste(GO:0001582) |
| 1.3 | 16.8 | GO:0034063 | stress granule assembly(GO:0034063) |
| 1.3 | 5.2 | GO:1902202 | regulation of hepatocyte growth factor receptor signaling pathway(GO:1902202) |
| 1.3 | 5.1 | GO:1903691 | positive regulation of wound healing, spreading of epidermal cells(GO:1903691) |
| 1.3 | 8.8 | GO:0002315 | marginal zone B cell differentiation(GO:0002315) |
| 1.3 | 7.5 | GO:0045110 | intermediate filament bundle assembly(GO:0045110) |
| 1.2 | 3.7 | GO:0006436 | tryptophanyl-tRNA aminoacylation(GO:0006436) |
| 1.2 | 9.9 | GO:0051388 | positive regulation of neurotrophin TRK receptor signaling pathway(GO:0051388) |
| 1.2 | 7.2 | GO:0032439 | endosome localization(GO:0032439) |
| 1.2 | 22.5 | GO:0048368 | lateral mesoderm development(GO:0048368) |
| 1.2 | 5.9 | GO:0010360 | negative regulation of anion channel activity(GO:0010360) |
| 1.2 | 7.0 | GO:0019372 | lipoxygenase pathway(GO:0019372) |
| 1.1 | 18.0 | GO:0043312 | neutrophil degranulation(GO:0043312) |
| 1.1 | 3.3 | GO:0071899 | odontoblast differentiation(GO:0071895) regulation of estrogen receptor binding(GO:0071898) negative regulation of estrogen receptor binding(GO:0071899) |
| 1.1 | 6.7 | GO:0070827 | chromatin maintenance(GO:0070827) |
| 1.1 | 6.6 | GO:0045329 | carnitine biosynthetic process(GO:0045329) |
| 1.1 | 23.8 | GO:0046827 | positive regulation of protein export from nucleus(GO:0046827) |
| 1.1 | 7.4 | GO:0000103 | sulfate assimilation(GO:0000103) |
| 1.0 | 6.2 | GO:0090367 | negative regulation of mRNA modification(GO:0090367) |
| 1.0 | 5.1 | GO:0050917 | sensory perception of umami taste(GO:0050917) |
| 1.0 | 3.0 | GO:0060697 | glucosylceramide catabolic process(GO:0006680) positive regulation of phospholipid catabolic process(GO:0060697) |
| 1.0 | 7.9 | GO:0035754 | B cell chemotaxis(GO:0035754) |
| 1.0 | 5.0 | GO:0033625 | positive regulation of integrin activation(GO:0033625) |
| 1.0 | 7.9 | GO:0008228 | opsonization(GO:0008228) |
| 1.0 | 5.8 | GO:2000741 | positive regulation of mesenchymal stem cell differentiation(GO:2000741) |
| 0.9 | 4.7 | GO:0060174 | limb bud formation(GO:0060174) |
| 0.9 | 12.1 | GO:0001977 | renal system process involved in regulation of blood volume(GO:0001977) |
| 0.9 | 8.1 | GO:0015867 | ADP transport(GO:0015866) ATP transport(GO:0015867) |
| 0.9 | 2.6 | GO:0098749 | cerebellar neuron development(GO:0098749) |
| 0.8 | 13.2 | GO:1902187 | negative regulation of viral release from host cell(GO:1902187) |
| 0.8 | 18.5 | GO:0044819 | mitotic G1 DNA damage checkpoint(GO:0031571) mitotic G1/S transition checkpoint(GO:0044819) |
| 0.8 | 4.8 | GO:0035426 | extracellular matrix-cell signaling(GO:0035426) progesterone secretion(GO:0042701) |
| 0.8 | 3.1 | GO:0036481 | intrinsic apoptotic signaling pathway in response to hydrogen peroxide(GO:0036481) regulation of intrinsic apoptotic signaling pathway in response to hydrogen peroxide(GO:1903750) negative regulation of intrinsic apoptotic signaling pathway in response to hydrogen peroxide(GO:1903751) |
| 0.8 | 3.9 | GO:0044691 | tooth eruption(GO:0044691) |
| 0.8 | 2.3 | GO:0001545 | primary ovarian follicle growth(GO:0001545) |
| 0.8 | 3.8 | GO:0035405 | histone-threonine phosphorylation(GO:0035405) |
| 0.8 | 9.9 | GO:0002361 | CD4-positive, CD25-positive, alpha-beta regulatory T cell differentiation(GO:0002361) |
| 0.8 | 5.3 | GO:0006268 | DNA unwinding involved in DNA replication(GO:0006268) |
| 0.7 | 2.2 | GO:0032497 | lipopolysaccharide transport(GO:0015920) detection of lipopolysaccharide(GO:0032497) |
| 0.7 | 3.7 | GO:2000504 | positive regulation of blood vessel remodeling(GO:2000504) |
| 0.7 | 7.2 | GO:0035021 | negative regulation of Rac protein signal transduction(GO:0035021) |
| 0.7 | 3.6 | GO:0009133 | nucleoside diphosphate biosynthetic process(GO:0009133) |
| 0.7 | 2.2 | GO:0018199 | peptidyl-glutamine modification(GO:0018199) |
| 0.7 | 2.1 | GO:0071529 | cementum mineralization(GO:0071529) |
| 0.7 | 5.0 | GO:0034773 | histone H4-K20 trimethylation(GO:0034773) |
| 0.7 | 10.6 | GO:0035269 | protein O-linked mannosylation(GO:0035269) |
| 0.7 | 6.3 | GO:0003373 | dynamin polymerization involved in membrane fission(GO:0003373) dynamin polymerization involved in mitochondrial fission(GO:0003374) |
| 0.7 | 2.1 | GO:0090158 | endoplasmic reticulum membrane organization(GO:0090158) |
| 0.7 | 12.3 | GO:1901741 | positive regulation of myoblast fusion(GO:1901741) |
| 0.7 | 2.0 | GO:0032380 | regulation of intracellular lipid transport(GO:0032377) regulation of intracellular sterol transport(GO:0032380) regulation of intracellular cholesterol transport(GO:0032383) |
| 0.6 | 3.2 | GO:1904717 | regulation of AMPA glutamate receptor clustering(GO:1904717) |
| 0.6 | 3.9 | GO:2001271 | negative regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:2001271) |
| 0.6 | 1.9 | GO:0002780 | antimicrobial peptide biosynthetic process(GO:0002777) antibacterial peptide biosynthetic process(GO:0002780) |
| 0.6 | 1.9 | GO:2000853 | positive regulation of Schwann cell proliferation(GO:0010625) negative regulation of corticosterone secretion(GO:2000853) |
| 0.6 | 17.1 | GO:0045954 | positive regulation of natural killer cell mediated cytotoxicity(GO:0045954) |
| 0.6 | 8.9 | GO:2000810 | regulation of bicellular tight junction assembly(GO:2000810) |
| 0.6 | 10.4 | GO:0045603 | positive regulation of endothelial cell differentiation(GO:0045603) |
| 0.6 | 5.3 | GO:0051152 | positive regulation of smooth muscle cell differentiation(GO:0051152) |
| 0.6 | 2.8 | GO:0035063 | nuclear speck organization(GO:0035063) |
| 0.6 | 7.9 | GO:0051988 | regulation of attachment of spindle microtubules to kinetochore(GO:0051988) |
| 0.6 | 2.2 | GO:0071802 | negative regulation of podosome assembly(GO:0071802) |
| 0.6 | 5.0 | GO:0010457 | centriole-centriole cohesion(GO:0010457) |
| 0.5 | 2.7 | GO:2000574 | regulation of microtubule motor activity(GO:2000574) |
| 0.5 | 5.4 | GO:0001886 | endothelial cell morphogenesis(GO:0001886) |
| 0.5 | 6.9 | GO:0009312 | oligosaccharide biosynthetic process(GO:0009312) |
| 0.5 | 18.3 | GO:0043303 | mast cell activation involved in immune response(GO:0002279) mast cell degranulation(GO:0043303) |
| 0.5 | 1.9 | GO:0034760 | negative regulation of iron ion transmembrane transport(GO:0034760) |
| 0.5 | 17.2 | GO:0071361 | cellular response to ethanol(GO:0071361) |
| 0.5 | 2.4 | GO:1903899 | positive regulation of PERK-mediated unfolded protein response(GO:1903899) |
| 0.5 | 1.4 | GO:0044240 | multicellular organism lipid catabolic process(GO:0044240) |
| 0.4 | 27.4 | GO:0032094 | response to food(GO:0032094) |
| 0.4 | 11.4 | GO:0034723 | DNA replication-dependent nucleosome assembly(GO:0006335) DNA replication-dependent nucleosome organization(GO:0034723) |
| 0.4 | 0.9 | GO:0044800 | fusion of virus membrane with host plasma membrane(GO:0019064) membrane fusion involved in viral entry into host cell(GO:0039663) multi-organism membrane fusion(GO:0044800) |
| 0.4 | 4.2 | GO:1990001 | inhibition of cysteine-type endopeptidase activity involved in apoptotic process(GO:1990001) |
| 0.4 | 5.0 | GO:0032525 | somite rostral/caudal axis specification(GO:0032525) |
| 0.4 | 0.8 | GO:1903969 | regulation of macrophage colony-stimulating factor signaling pathway(GO:1902226) regulation of response to macrophage colony-stimulating factor(GO:1903969) regulation of cellular response to macrophage colony-stimulating factor stimulus(GO:1903972) |
| 0.4 | 4.8 | GO:0042492 | gamma-delta T cell differentiation(GO:0042492) |
| 0.4 | 5.6 | GO:0051044 | positive regulation of membrane protein ectodomain proteolysis(GO:0051044) |
| 0.4 | 1.5 | GO:0007182 | common-partner SMAD protein phosphorylation(GO:0007182) |
| 0.4 | 1.5 | GO:0090204 | protein localization to nuclear pore(GO:0090204) |
| 0.4 | 6.1 | GO:0016339 | calcium-dependent cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0016339) |
| 0.4 | 10.8 | GO:0006099 | tricarboxylic acid cycle(GO:0006099) |
| 0.4 | 1.1 | GO:0044339 | canonical Wnt signaling pathway involved in osteoblast differentiation(GO:0044339) |
| 0.3 | 2.4 | GO:0031848 | protection from non-homologous end joining at telomere(GO:0031848) |
| 0.3 | 5.1 | GO:0006517 | protein deglycosylation(GO:0006517) |
| 0.3 | 0.7 | GO:0097187 | dentinogenesis(GO:0097187) |
| 0.3 | 3.6 | GO:1900017 | positive regulation of cytokine production involved in inflammatory response(GO:1900017) |
| 0.3 | 1.9 | GO:0071409 | cellular response to cycloheximide(GO:0071409) |
| 0.3 | 2.9 | GO:1903715 | regulation of aerobic respiration(GO:1903715) |
| 0.3 | 2.4 | GO:0046543 | development of secondary female sexual characteristics(GO:0046543) |
| 0.3 | 1.2 | GO:0000379 | tRNA-type intron splice site recognition and cleavage(GO:0000379) |
| 0.3 | 13.7 | GO:0006956 | complement activation(GO:0006956) |
| 0.3 | 13.7 | GO:0045773 | positive regulation of axon extension(GO:0045773) |
| 0.3 | 11.4 | GO:0070265 | necrotic cell death(GO:0070265) |
| 0.3 | 7.4 | GO:0018279 | protein N-linked glycosylation via asparagine(GO:0018279) |
| 0.3 | 9.7 | GO:1901998 | toxin transport(GO:1901998) |
| 0.3 | 2.8 | GO:0000722 | telomere maintenance via recombination(GO:0000722) |
| 0.3 | 11.2 | GO:0006879 | cellular iron ion homeostasis(GO:0006879) |
| 0.3 | 0.8 | GO:0046013 | T cell homeostatic proliferation(GO:0001777) regulation of T cell homeostatic proliferation(GO:0046013) |
| 0.3 | 3.8 | GO:0060670 | branching involved in labyrinthine layer morphogenesis(GO:0060670) |
| 0.3 | 3.5 | GO:0070072 | vacuolar proton-transporting V-type ATPase complex assembly(GO:0070072) |
| 0.3 | 2.7 | GO:0042074 | cell migration involved in gastrulation(GO:0042074) |
| 0.3 | 2.9 | GO:0042487 | regulation of odontogenesis of dentin-containing tooth(GO:0042487) |
| 0.3 | 3.4 | GO:0007379 | segment specification(GO:0007379) |
| 0.3 | 3.9 | GO:0035855 | megakaryocyte development(GO:0035855) |
| 0.2 | 2.5 | GO:0046341 | CDP-diacylglycerol metabolic process(GO:0046341) |
| 0.2 | 13.2 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
| 0.2 | 6.5 | GO:0006829 | zinc II ion transport(GO:0006829) |
| 0.2 | 21.0 | GO:0008203 | cholesterol metabolic process(GO:0008203) |
| 0.2 | 2.3 | GO:0010839 | negative regulation of keratinocyte proliferation(GO:0010839) |
| 0.2 | 6.7 | GO:0006506 | GPI anchor biosynthetic process(GO:0006506) |
| 0.2 | 5.4 | GO:0019373 | epoxygenase P450 pathway(GO:0019373) |
| 0.2 | 11.2 | GO:0038083 | peptidyl-tyrosine autophosphorylation(GO:0038083) |
| 0.2 | 1.2 | GO:0033387 | putrescine biosynthetic process from ornithine(GO:0033387) |
| 0.2 | 8.9 | GO:0010501 | RNA secondary structure unwinding(GO:0010501) |
| 0.2 | 2.1 | GO:2000811 | negative regulation of anoikis(GO:2000811) |
| 0.2 | 4.8 | GO:0007064 | mitotic sister chromatid cohesion(GO:0007064) |
| 0.2 | 1.6 | GO:0060056 | mammary gland involution(GO:0060056) |
| 0.2 | 5.7 | GO:0018345 | protein palmitoylation(GO:0018345) |
| 0.2 | 0.5 | GO:0015942 | formate metabolic process(GO:0015942) |
| 0.2 | 2.5 | GO:0032060 | bleb assembly(GO:0032060) |
| 0.2 | 2.1 | GO:2000696 | regulation of epithelial cell differentiation involved in kidney development(GO:2000696) |
| 0.2 | 0.8 | GO:0071955 | recycling endosome to Golgi transport(GO:0071955) |
| 0.2 | 0.5 | GO:0019255 | glucose 1-phosphate metabolic process(GO:0019255) |
| 0.1 | 0.7 | GO:0051596 | methylglyoxal catabolic process to D-lactate via S-lactoyl-glutathione(GO:0019243) methylglyoxal catabolic process(GO:0051596) methylglyoxal catabolic process to lactate(GO:0061727) |
| 0.1 | 1.4 | GO:0006283 | transcription-coupled nucleotide-excision repair(GO:0006283) |
| 0.1 | 9.0 | GO:0043488 | regulation of mRNA stability(GO:0043488) |
| 0.1 | 2.1 | GO:0060445 | branching involved in salivary gland morphogenesis(GO:0060445) |
| 0.1 | 7.3 | GO:0033045 | regulation of sister chromatid segregation(GO:0033045) |
| 0.1 | 0.7 | GO:0008295 | spermidine biosynthetic process(GO:0008295) |
| 0.1 | 7.1 | GO:0031338 | regulation of vesicle fusion(GO:0031338) |
| 0.1 | 0.5 | GO:0071494 | cellular response to UV-C(GO:0071494) |
| 0.1 | 2.8 | GO:0019432 | triglyceride biosynthetic process(GO:0019432) |
| 0.1 | 2.3 | GO:0070884 | regulation of calcineurin-NFAT signaling cascade(GO:0070884) |
| 0.1 | 5.2 | GO:0006893 | Golgi to plasma membrane transport(GO:0006893) |
| 0.1 | 6.5 | GO:0045454 | cell redox homeostasis(GO:0045454) |
| 0.1 | 2.2 | GO:0032757 | positive regulation of interleukin-8 production(GO:0032757) |
| 0.1 | 2.6 | GO:0033137 | negative regulation of peptidyl-serine phosphorylation(GO:0033137) |
| 0.1 | 0.7 | GO:0032287 | peripheral nervous system myelin maintenance(GO:0032287) |
| 0.1 | 1.5 | GO:0045109 | intermediate filament organization(GO:0045109) |
| 0.1 | 0.6 | GO:0035735 | intraciliary transport involved in cilium morphogenesis(GO:0035735) |
| 0.1 | 2.9 | GO:0050829 | defense response to Gram-negative bacterium(GO:0050829) |
| 0.1 | 3.2 | GO:0034605 | cellular response to heat(GO:0034605) |
| 0.1 | 4.8 | GO:0007093 | mitotic cell cycle checkpoint(GO:0007093) |
| 0.1 | 12.6 | GO:0043413 | protein glycosylation(GO:0006486) macromolecule glycosylation(GO:0043413) |
| 0.1 | 1.6 | GO:0015985 | energy coupled proton transport, down electrochemical gradient(GO:0015985) ATP synthesis coupled proton transport(GO:0015986) |
| 0.1 | 2.0 | GO:0042073 | intraciliary transport(GO:0042073) |
| 0.1 | 1.7 | GO:0097352 | autophagosome maturation(GO:0097352) |
| 0.1 | 3.6 | GO:0032024 | positive regulation of insulin secretion(GO:0032024) |
| 0.1 | 2.7 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.1 | 0.3 | GO:0036414 | protein citrullination(GO:0018101) histone citrullination(GO:0036414) |
| 0.1 | 0.5 | GO:0071218 | cellular response to misfolded protein(GO:0071218) |
| 0.1 | 0.2 | GO:1901187 | regulation of ephrin receptor signaling pathway(GO:1901187) |
| 0.1 | 2.8 | GO:0070373 | negative regulation of ERK1 and ERK2 cascade(GO:0070373) |
| 0.1 | 0.9 | GO:0051597 | response to methylmercury(GO:0051597) |
| 0.0 | 1.0 | GO:0051642 | centrosome localization(GO:0051642) |
| 0.0 | 2.1 | GO:0030148 | sphingolipid biosynthetic process(GO:0030148) |
| 0.0 | 2.0 | GO:0071427 | mRNA export from nucleus(GO:0006406) mRNA-containing ribonucleoprotein complex export from nucleus(GO:0071427) |
| 0.0 | 2.3 | GO:0030032 | lamellipodium assembly(GO:0030032) |
| 0.0 | 4.4 | GO:0042787 | protein ubiquitination involved in ubiquitin-dependent protein catabolic process(GO:0042787) |
| 0.0 | 1.1 | GO:0006953 | acute-phase response(GO:0006953) |
| 0.0 | 2.0 | GO:0007218 | neuropeptide signaling pathway(GO:0007218) |
| 0.0 | 1.7 | GO:0030183 | B cell differentiation(GO:0030183) |
| 0.0 | 0.8 | GO:0042462 | eye photoreceptor cell development(GO:0042462) |
| 0.0 | 5.1 | GO:0032259 | methylation(GO:0032259) |
| 0.0 | 0.5 | GO:0051491 | positive regulation of filopodium assembly(GO:0051491) |
| 0.0 | 1.4 | GO:1902476 | chloride transmembrane transport(GO:1902476) |
| 0.0 | 1.8 | GO:0006401 | RNA catabolic process(GO:0006401) |
| 0.0 | 1.0 | GO:0009791 | post-embryonic development(GO:0009791) |
| 0.0 | 2.3 | GO:0006869 | lipid transport(GO:0006869) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 7.5 | 22.5 | GO:0071149 | TEAD-2-YAP complex(GO:0071149) |
| 7.3 | 21.8 | GO:0035692 | macrophage migration inhibitory factor receptor complex(GO:0035692) |
| 6.0 | 17.9 | GO:0005760 | gamma DNA polymerase complex(GO:0005760) |
| 5.8 | 46.6 | GO:0042105 | alpha-beta T cell receptor complex(GO:0042105) |
| 3.7 | 69.7 | GO:0005779 | integral component of peroxisomal membrane(GO:0005779) intrinsic component of peroxisomal membrane(GO:0031231) |
| 2.7 | 13.3 | GO:0097169 | NLRP1 inflammasome complex(GO:0072558) AIM2 inflammasome complex(GO:0097169) |
| 2.6 | 62.9 | GO:0016327 | apicolateral plasma membrane(GO:0016327) |
| 2.5 | 17.6 | GO:0097524 | sperm plasma membrane(GO:0097524) |
| 2.4 | 17.0 | GO:0005579 | membrane attack complex(GO:0005579) |
| 2.4 | 11.9 | GO:0045252 | oxoglutarate dehydrogenase complex(GO:0045252) |
| 2.3 | 18.4 | GO:0070820 | cytolytic granule(GO:0044194) tertiary granule(GO:0070820) |
| 2.1 | 8.3 | GO:0005971 | ribonucleoside-diphosphate reductase complex(GO:0005971) |
| 2.0 | 21.7 | GO:1990635 | proximal dendrite(GO:1990635) |
| 2.0 | 7.9 | GO:0097149 | centralspindlin complex(GO:0097149) |
| 1.5 | 4.6 | GO:1990423 | RZZ complex(GO:1990423) |
| 1.5 | 21.1 | GO:0044232 | organelle membrane contact site(GO:0044232) |
| 1.4 | 103.2 | GO:0045095 | keratin filament(GO:0045095) |
| 1.4 | 8.1 | GO:0071817 | MMXD complex(GO:0071817) |
| 1.3 | 6.5 | GO:1903767 | sweet taste receptor complex(GO:1903767) taste receptor complex(GO:1903768) |
| 1.2 | 15.2 | GO:0032591 | dendritic spine membrane(GO:0032591) |
| 1.1 | 3.3 | GO:1990907 | beta-catenin-TCF complex(GO:1990907) |
| 1.1 | 30.0 | GO:0000788 | nuclear nucleosome(GO:0000788) |
| 1.0 | 8.2 | GO:0016589 | NURF complex(GO:0016589) |
| 1.0 | 7.1 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 1.0 | 9.1 | GO:0045179 | apical cortex(GO:0045179) |
| 1.0 | 3.9 | GO:0002947 | tumor necrosis factor receptor superfamily complex(GO:0002947) |
| 1.0 | 3.8 | GO:0070436 | Grb2-EGFR complex(GO:0070436) |
| 1.0 | 13.3 | GO:0032426 | stereocilium tip(GO:0032426) |
| 0.9 | 26.1 | GO:0042629 | mast cell granule(GO:0042629) |
| 0.9 | 14.7 | GO:0008250 | oligosaccharyltransferase complex(GO:0008250) |
| 0.9 | 7.7 | GO:0001940 | male pronucleus(GO:0001940) |
| 0.8 | 7.6 | GO:0032437 | cuticular plate(GO:0032437) |
| 0.8 | 9.0 | GO:0000177 | cytoplasmic exosome (RNase complex)(GO:0000177) |
| 0.7 | 3.3 | GO:0045098 | type III intermediate filament(GO:0045098) |
| 0.6 | 19.1 | GO:0099738 | cell cortex region(GO:0099738) |
| 0.6 | 11.7 | GO:0030057 | desmosome(GO:0030057) |
| 0.6 | 4.3 | GO:0000127 | transcription factor TFIIIC complex(GO:0000127) |
| 0.6 | 72.5 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.6 | 6.1 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.6 | 2.2 | GO:0031528 | microvillus membrane(GO:0031528) |
| 0.5 | 5.4 | GO:0070187 | telosome(GO:0070187) |
| 0.5 | 6.0 | GO:0034663 | endoplasmic reticulum chaperone complex(GO:0034663) |
| 0.5 | 6.5 | GO:0008385 | IkappaB kinase complex(GO:0008385) |
| 0.5 | 6.8 | GO:0042581 | specific granule(GO:0042581) |
| 0.5 | 44.4 | GO:0030863 | cortical cytoskeleton(GO:0030863) |
| 0.5 | 9.1 | GO:0031907 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.5 | 5.3 | GO:0042555 | MCM complex(GO:0042555) |
| 0.5 | 36.5 | GO:0031526 | brush border membrane(GO:0031526) |
| 0.4 | 3.9 | GO:0031209 | SCAR complex(GO:0031209) |
| 0.4 | 2.1 | GO:0071256 | Sec61 translocon complex(GO:0005784) translocon complex(GO:0071256) |
| 0.4 | 32.2 | GO:0042470 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.4 | 24.1 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.4 | 35.2 | GO:0017053 | transcriptional repressor complex(GO:0017053) |
| 0.4 | 7.9 | GO:0030670 | phagocytic vesicle membrane(GO:0030670) |
| 0.4 | 16.8 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
| 0.4 | 2.1 | GO:0065010 | extracellular membrane-bounded organelle(GO:0065010) |
| 0.3 | 6.7 | GO:0035327 | transcriptionally active chromatin(GO:0035327) |
| 0.3 | 6.3 | GO:0032588 | trans-Golgi network membrane(GO:0032588) |
| 0.3 | 2.8 | GO:0005610 | laminin-5 complex(GO:0005610) |
| 0.3 | 6.1 | GO:0005614 | interstitial matrix(GO:0005614) |
| 0.3 | 0.9 | GO:0032173 | septin collar(GO:0032173) |
| 0.3 | 12.6 | GO:0045171 | intercellular bridge(GO:0045171) |
| 0.3 | 0.8 | GO:1990682 | CSF1-CSF1R complex(GO:1990682) |
| 0.3 | 6.6 | GO:0008074 | guanylate cyclase complex, soluble(GO:0008074) |
| 0.3 | 107.0 | GO:0045177 | apical part of cell(GO:0045177) |
| 0.3 | 2.4 | GO:0044292 | dendrite terminus(GO:0044292) |
| 0.3 | 6.8 | GO:0071782 | endoplasmic reticulum tubular network(GO:0071782) |
| 0.3 | 10.3 | GO:0000791 | euchromatin(GO:0000791) |
| 0.2 | 1.2 | GO:0000214 | tRNA-intron endonuclease complex(GO:0000214) |
| 0.2 | 6.9 | GO:0005697 | telomerase holoenzyme complex(GO:0005697) |
| 0.2 | 3.7 | GO:0031235 | intrinsic component of the cytoplasmic side of the plasma membrane(GO:0031235) |
| 0.2 | 26.9 | GO:0031234 | extrinsic component of cytoplasmic side of plasma membrane(GO:0031234) |
| 0.2 | 3.0 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.2 | 14.5 | GO:0005811 | lipid particle(GO:0005811) |
| 0.2 | 19.2 | GO:0005923 | bicellular tight junction(GO:0005923) |
| 0.2 | 1.6 | GO:0000275 | mitochondrial proton-transporting ATP synthase complex, catalytic core F(1)(GO:0000275) proton-transporting ATP synthase complex, catalytic core F(1)(GO:0045261) |
| 0.2 | 2.0 | GO:0044613 | nuclear pore central transport channel(GO:0044613) nuclear pore nuclear basket(GO:0044615) |
| 0.2 | 1.8 | GO:0000178 | exosome (RNase complex)(GO:0000178) |
| 0.2 | 4.4 | GO:1902711 | GABA-A receptor complex(GO:1902711) |
| 0.2 | 16.8 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.2 | 14.5 | GO:0034399 | nuclear periphery(GO:0034399) |
| 0.2 | 2.3 | GO:0031233 | intrinsic component of external side of plasma membrane(GO:0031233) |
| 0.2 | 3.4 | GO:0000421 | autophagosome membrane(GO:0000421) |
| 0.2 | 8.4 | GO:0005791 | rough endoplasmic reticulum(GO:0005791) |
| 0.1 | 3.8 | GO:0032154 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.1 | 3.8 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.1 | 1.0 | GO:0005641 | nuclear envelope lumen(GO:0005641) |
| 0.1 | 11.0 | GO:0030496 | midbody(GO:0030496) |
| 0.1 | 44.4 | GO:0000323 | lytic vacuole(GO:0000323) lysosome(GO:0005764) |
| 0.1 | 38.0 | GO:0016607 | nuclear speck(GO:0016607) |
| 0.1 | 7.6 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.1 | 0.9 | GO:0043596 | nuclear replication fork(GO:0043596) |
| 0.1 | 5.6 | GO:0005798 | Golgi-associated vesicle(GO:0005798) |
| 0.1 | 25.5 | GO:0000790 | nuclear chromatin(GO:0000790) |
| 0.1 | 8.6 | GO:0005795 | Golgi stack(GO:0005795) |
| 0.1 | 2.0 | GO:0030992 | intraciliary transport particle B(GO:0030992) |
| 0.1 | 1.7 | GO:0032420 | stereocilium(GO:0032420) |
| 0.1 | 5.2 | GO:0016605 | PML body(GO:0016605) |
| 0.1 | 7.3 | GO:0030176 | integral component of endoplasmic reticulum membrane(GO:0030176) |
| 0.1 | 1.1 | GO:0034364 | high-density lipoprotein particle(GO:0034364) |
| 0.1 | 4.7 | GO:0005930 | axoneme(GO:0005930) ciliary plasm(GO:0097014) |
| 0.1 | 5.8 | GO:0044798 | nuclear transcription factor complex(GO:0044798) |
| 0.1 | 14.2 | GO:0005925 | focal adhesion(GO:0005925) |
| 0.0 | 0.8 | GO:0097546 | ciliary base(GO:0097546) |
| 0.0 | 0.7 | GO:0055038 | recycling endosome membrane(GO:0055038) |
| 0.0 | 0.2 | GO:0030896 | checkpoint clamp complex(GO:0030896) |
| 0.0 | 0.2 | GO:0034388 | Pwp2p-containing subcomplex of 90S preribosome(GO:0034388) |
| 0.0 | 4.3 | GO:0001650 | fibrillar center(GO:0001650) |
| 0.0 | 0.8 | GO:0000118 | histone deacetylase complex(GO:0000118) |
| 0.0 | 3.3 | GO:0032993 | protein-DNA complex(GO:0032993) |
| 0.0 | 0.1 | GO:0005927 | muscle tendon junction(GO:0005927) |
| 0.0 | 2.5 | GO:0005814 | centriole(GO:0005814) |
| 0.0 | 1.4 | GO:0071013 | catalytic step 2 spliceosome(GO:0071013) |
| 0.0 | 1.1 | GO:0035770 | ribonucleoprotein granule(GO:0035770) |
| 0.0 | 1.3 | GO:0005758 | mitochondrial intermembrane space(GO:0005758) |
| 0.0 | 1.5 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.0 | 0.6 | GO:0036064 | ciliary basal body(GO:0036064) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 8.8 | 26.5 | GO:0031714 | C5a anaphylatoxin chemotactic receptor binding(GO:0031714) |
| 7.7 | 76.9 | GO:0031957 | very long-chain fatty acid-CoA ligase activity(GO:0031957) |
| 6.1 | 18.4 | GO:0030348 | syntaxin-3 binding(GO:0030348) |
| 5.2 | 46.6 | GO:0005001 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 4.8 | 14.3 | GO:0052743 | inositol tetrakisphosphate phosphatase activity(GO:0052743) |
| 4.0 | 12.1 | GO:0003845 | 11-beta-hydroxysteroid dehydrogenase [NAD(P)] activity(GO:0003845) |
| 4.0 | 11.9 | GO:0004591 | oxoglutarate dehydrogenase (succinyl-transferring) activity(GO:0004591) |
| 3.8 | 15.2 | GO:0004691 | cAMP-dependent protein kinase activity(GO:0004691) |
| 3.7 | 22.1 | GO:0061731 | ribonucleoside-diphosphate reductase activity, thioredoxin disulfide as acceptor(GO:0004748) oxidoreductase activity, acting on CH or CH2 groups, disulfide as acceptor(GO:0016728) ribonucleoside-diphosphate reductase activity(GO:0061731) |
| 3.6 | 10.8 | GO:0035403 | histone kinase activity (H3-T6 specific)(GO:0035403) |
| 3.3 | 13.3 | GO:0045159 | myosin II binding(GO:0045159) |
| 3.0 | 36.0 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 3.0 | 17.9 | GO:0042289 | MHC class II protein binding(GO:0042289) |
| 2.9 | 8.6 | GO:0042895 | amino-acid betaine transmembrane transporter activity(GO:0015199) carnitine transmembrane transporter activity(GO:0015226) antibiotic transporter activity(GO:0042895) |
| 2.8 | 25.5 | GO:0019534 | toxin transporter activity(GO:0019534) |
| 2.6 | 10.3 | GO:0044729 | hemi-methylated DNA-binding(GO:0044729) |
| 2.6 | 10.2 | GO:0004185 | serine-type carboxypeptidase activity(GO:0004185) |
| 2.5 | 22.3 | GO:0046790 | virion binding(GO:0046790) |
| 2.5 | 7.4 | GO:0070012 | oligopeptidase activity(GO:0070012) |
| 2.5 | 7.4 | GO:0050294 | steroid sulfotransferase activity(GO:0050294) |
| 2.4 | 7.2 | GO:0000703 | oxidized pyrimidine nucleobase lesion DNA N-glycosylase activity(GO:0000703) |
| 2.4 | 7.2 | GO:0097603 | temperature-gated ion channel activity(GO:0097603) |
| 2.3 | 18.7 | GO:0070053 | thrombospondin receptor activity(GO:0070053) |
| 2.3 | 32.2 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 2.3 | 9.0 | GO:0070644 | vitamin D response element binding(GO:0070644) |
| 2.2 | 6.7 | GO:0000991 | transcription factor activity, core RNA polymerase II binding(GO:0000991) |
| 2.2 | 10.9 | GO:0008453 | alanine-glyoxylate transaminase activity(GO:0008453) |
| 2.0 | 20.3 | GO:0019957 | C-C chemokine binding(GO:0019957) |
| 1.9 | 5.8 | GO:0016748 | succinyltransferase activity(GO:0016748) |
| 1.9 | 7.7 | GO:0070579 | methylcytosine dioxygenase activity(GO:0070579) |
| 1.9 | 13.3 | GO:0005138 | interleukin-6 receptor binding(GO:0005138) |
| 1.8 | 16.2 | GO:0042285 | xylosyltransferase activity(GO:0042285) |
| 1.8 | 16.0 | GO:0015093 | ferrous iron transmembrane transporter activity(GO:0015093) |
| 1.8 | 7.0 | GO:0004052 | arachidonate 12-lipoxygenase activity(GO:0004052) |
| 1.7 | 5.2 | GO:0001716 | L-amino-acid oxidase activity(GO:0001716) |
| 1.7 | 13.9 | GO:0005007 | fibroblast growth factor-activated receptor activity(GO:0005007) |
| 1.7 | 5.2 | GO:0005171 | hepatocyte growth factor receptor binding(GO:0005171) |
| 1.7 | 15.4 | GO:0008559 | xenobiotic-transporting ATPase activity(GO:0008559) |
| 1.6 | 6.5 | GO:0003842 | 1-pyrroline-5-carboxylate dehydrogenase activity(GO:0003842) |
| 1.6 | 9.7 | GO:0008467 | [heparan sulfate]-glucosamine 3-sulfotransferase 1 activity(GO:0008467) |
| 1.6 | 9.5 | GO:0016493 | C-C chemokine receptor activity(GO:0016493) |
| 1.6 | 4.7 | GO:0070735 | protein-glycine ligase activity(GO:0070735) |
| 1.4 | 5.6 | GO:0051377 | mannose-ethanolamine phosphotransferase activity(GO:0051377) |
| 1.4 | 6.9 | GO:0032564 | dATP binding(GO:0032564) |
| 1.4 | 6.8 | GO:0019834 | phospholipase A2 inhibitor activity(GO:0019834) |
| 1.3 | 5.3 | GO:0061665 | SUMO ligase activity(GO:0061665) |
| 1.3 | 9.3 | GO:0004064 | arylesterase activity(GO:0004064) |
| 1.3 | 10.1 | GO:0003964 | telomerase activity(GO:0003720) RNA-directed DNA polymerase activity(GO:0003964) |
| 1.2 | 3.7 | GO:0004830 | tryptophan-tRNA ligase activity(GO:0004830) |
| 1.2 | 7.5 | GO:0008172 | S-methyltransferase activity(GO:0008172) |
| 1.2 | 24.9 | GO:0036312 | phosphatidylinositol 3-kinase regulatory subunit binding(GO:0036312) |
| 1.2 | 14.7 | GO:0004499 | N,N-dimethylaniline monooxygenase activity(GO:0004499) |
| 1.2 | 9.7 | GO:0048273 | mitogen-activated protein kinase p38 binding(GO:0048273) NFAT protein binding(GO:0051525) |
| 1.2 | 8.5 | GO:0043546 | molybdopterin cofactor binding(GO:0043546) |
| 1.2 | 4.8 | GO:0019002 | GMP binding(GO:0019002) |
| 1.2 | 8.3 | GO:0015119 | hexose phosphate transmembrane transporter activity(GO:0015119) organophosphate:inorganic phosphate antiporter activity(GO:0015315) hexose-phosphate:inorganic phosphate antiporter activity(GO:0015526) glucose 6-phosphate:inorganic phosphate antiporter activity(GO:0061513) |
| 1.2 | 14.0 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 1.2 | 2.3 | GO:0004963 | follicle-stimulating hormone receptor activity(GO:0004963) |
| 1.1 | 22.5 | GO:0001134 | transcription factor activity, transcription factor recruiting(GO:0001134) |
| 1.1 | 9.0 | GO:0031419 | cobalamin binding(GO:0031419) |
| 1.1 | 8.9 | GO:0000064 | L-ornithine transmembrane transporter activity(GO:0000064) |
| 1.0 | 9.4 | GO:0070061 | fructose binding(GO:0070061) |
| 1.0 | 6.1 | GO:0045545 | syndecan binding(GO:0045545) |
| 1.0 | 14.3 | GO:0043522 | leucine zipper domain binding(GO:0043522) |
| 1.0 | 3.0 | GO:0070976 | TIR domain binding(GO:0070976) |
| 1.0 | 5.9 | GO:0047696 | beta-adrenergic receptor kinase activity(GO:0047696) |
| 1.0 | 2.9 | GO:0004167 | dopachrome isomerase activity(GO:0004167) |
| 0.9 | 8.1 | GO:0005347 | ATP transmembrane transporter activity(GO:0005347) ADP transmembrane transporter activity(GO:0015217) |
| 0.9 | 2.7 | GO:0038100 | nodal binding(GO:0038100) |
| 0.9 | 19.3 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
| 0.9 | 2.6 | GO:0008948 | oxaloacetate decarboxylase activity(GO:0008948) hydrolase activity, acting on acid carbon-carbon bonds(GO:0016822) hydrolase activity, acting on acid carbon-carbon bonds, in ketonic substances(GO:0016823) |
| 0.8 | 15.0 | GO:0003993 | acid phosphatase activity(GO:0003993) |
| 0.8 | 12.6 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.8 | 3.8 | GO:0035184 | histone threonine kinase activity(GO:0035184) |
| 0.7 | 166.8 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.7 | 2.2 | GO:0070506 | high-density lipoprotein particle receptor activity(GO:0070506) |
| 0.7 | 9.6 | GO:0043024 | ribosomal small subunit binding(GO:0043024) |
| 0.7 | 3.7 | GO:0004586 | ornithine decarboxylase activity(GO:0004586) |
| 0.7 | 13.6 | GO:0015643 | toxic substance binding(GO:0015643) |
| 0.7 | 4.9 | GO:0046976 | histone methyltransferase activity (H3-K27 specific)(GO:0046976) |
| 0.7 | 9.7 | GO:0004576 | oligosaccharyl transferase activity(GO:0004576) dolichyl-diphosphooligosaccharide-protein glycotransferase activity(GO:0004579) |
| 0.7 | 5.4 | GO:0030274 | LIM domain binding(GO:0030274) |
| 0.7 | 2.0 | GO:0005049 | nuclear export signal receptor activity(GO:0005049) |
| 0.7 | 41.3 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.7 | 25.1 | GO:0043236 | laminin binding(GO:0043236) |
| 0.7 | 6.6 | GO:1904264 | ubiquitin protein ligase activity involved in ERAD pathway(GO:1904264) |
| 0.6 | 5.2 | GO:0008429 | phosphatidylethanolamine binding(GO:0008429) |
| 0.6 | 7.1 | GO:0008641 | small protein activating enzyme activity(GO:0008641) |
| 0.6 | 19.0 | GO:0034061 | DNA-directed DNA polymerase activity(GO:0003887) DNA polymerase activity(GO:0034061) |
| 0.6 | 16.2 | GO:0070064 | proline-rich region binding(GO:0070064) |
| 0.6 | 2.3 | GO:0008597 | calcium-dependent protein serine/threonine phosphatase regulator activity(GO:0008597) |
| 0.6 | 12.7 | GO:0004745 | retinol dehydrogenase activity(GO:0004745) |
| 0.6 | 5.1 | GO:0034483 | heparan sulfate sulfotransferase activity(GO:0034483) |
| 0.6 | 7.9 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.6 | 12.3 | GO:0035250 | UDP-galactosyltransferase activity(GO:0035250) |
| 0.5 | 3.2 | GO:0030284 | estrogen receptor activity(GO:0030284) |
| 0.5 | 16.6 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.5 | 18.5 | GO:0004180 | carboxypeptidase activity(GO:0004180) |
| 0.5 | 16.4 | GO:0042169 | SH2 domain binding(GO:0042169) |
| 0.5 | 6.0 | GO:0051787 | misfolded protein binding(GO:0051787) |
| 0.4 | 6.3 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.4 | 6.2 | GO:0008143 | poly(A) binding(GO:0008143) |
| 0.4 | 2.1 | GO:0001729 | ceramide kinase activity(GO:0001729) |
| 0.4 | 14.1 | GO:0008536 | Ran GTPase binding(GO:0008536) |
| 0.4 | 57.6 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 0.4 | 10.8 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.4 | 1.9 | GO:0005176 | ErbB-2 class receptor binding(GO:0005176) |
| 0.4 | 3.7 | GO:1990825 | sequence-specific mRNA binding(GO:1990825) |
| 0.4 | 8.1 | GO:0005388 | calcium-transporting ATPase activity(GO:0005388) |
| 0.4 | 1.1 | GO:0000026 | alpha-1,2-mannosyltransferase activity(GO:0000026) |
| 0.4 | 1.4 | GO:0004452 | isopentenyl-diphosphate delta-isomerase activity(GO:0004452) |
| 0.4 | 7.5 | GO:0005521 | lamin binding(GO:0005521) |
| 0.3 | 3.1 | GO:0000774 | adenyl-nucleotide exchange factor activity(GO:0000774) |
| 0.3 | 5.4 | GO:0070182 | DNA polymerase binding(GO:0070182) |
| 0.3 | 2.4 | GO:0035312 | 5'-3' exodeoxyribonuclease activity(GO:0035312) |
| 0.3 | 1.7 | GO:0072345 | NAADP-sensitive calcium-release channel activity(GO:0072345) |
| 0.3 | 2.5 | GO:0017169 | CDP-alcohol phosphatidyltransferase activity(GO:0017169) |
| 0.3 | 9.9 | GO:0004623 | phospholipase A2 activity(GO:0004623) |
| 0.3 | 9.8 | GO:0001205 | transcriptional activator activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001205) |
| 0.3 | 7.9 | GO:0008395 | steroid hydroxylase activity(GO:0008395) |
| 0.3 | 6.6 | GO:0004383 | guanylate cyclase activity(GO:0004383) |
| 0.3 | 0.8 | GO:0005134 | interleukin-2 receptor binding(GO:0005134) |
| 0.3 | 0.8 | GO:0005157 | macrophage colony-stimulating factor receptor binding(GO:0005157) |
| 0.3 | 2.1 | GO:0004035 | alkaline phosphatase activity(GO:0004035) |
| 0.2 | 3.6 | GO:0017110 | nucleoside-diphosphatase activity(GO:0017110) |
| 0.2 | 1.2 | GO:0000213 | tRNA-intron endonuclease activity(GO:0000213) |
| 0.2 | 5.4 | GO:0016709 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, NAD(P)H as one donor, and incorporation of one atom of oxygen(GO:0016709) |
| 0.2 | 5.4 | GO:0008392 | arachidonic acid epoxygenase activity(GO:0008392) |
| 0.2 | 6.3 | GO:0005328 | neurotransmitter:sodium symporter activity(GO:0005328) |
| 0.2 | 3.2 | GO:0099604 | calcium-release channel activity(GO:0015278) ligand-gated calcium channel activity(GO:0099604) |
| 0.2 | 2.9 | GO:0003688 | DNA replication origin binding(GO:0003688) |
| 0.2 | 4.7 | GO:0071837 | HMG box domain binding(GO:0071837) |
| 0.2 | 1.8 | GO:0070700 | BMP receptor binding(GO:0070700) |
| 0.2 | 14.0 | GO:0003725 | double-stranded RNA binding(GO:0003725) |
| 0.2 | 3.1 | GO:0046965 | retinoid X receptor binding(GO:0046965) |
| 0.2 | 10.6 | GO:0004004 | RNA helicase activity(GO:0003724) ATP-dependent RNA helicase activity(GO:0004004) RNA-dependent ATPase activity(GO:0008186) |
| 0.2 | 6.7 | GO:0035255 | ionotropic glutamate receptor binding(GO:0035255) |
| 0.2 | 20.0 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.2 | 5.7 | GO:0019706 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.2 | 20.9 | GO:0017124 | SH3 domain binding(GO:0017124) |
| 0.2 | 4.4 | GO:0004890 | GABA-A receptor activity(GO:0004890) |
| 0.2 | 6.6 | GO:0016706 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, 2-oxoglutarate as one donor, and incorporation of one atom each of oxygen into both donors(GO:0016706) |
| 0.2 | 10.4 | GO:0033613 | activating transcription factor binding(GO:0033613) |
| 0.2 | 2.8 | GO:0008195 | phosphatidate phosphatase activity(GO:0008195) |
| 0.2 | 9.0 | GO:0004386 | helicase activity(GO:0004386) |
| 0.1 | 0.7 | GO:0004416 | hydroxyacylglutathione hydrolase activity(GO:0004416) |
| 0.1 | 2.4 | GO:0002162 | dystroglycan binding(GO:0002162) |
| 0.1 | 9.4 | GO:0003730 | mRNA 3'-UTR binding(GO:0003730) |
| 0.1 | 4.4 | GO:0030332 | cyclin binding(GO:0030332) |
| 0.1 | 3.2 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.1 | 4.8 | GO:0070063 | RNA polymerase binding(GO:0070063) |
| 0.1 | 40.6 | GO:0045296 | cadherin binding(GO:0045296) |
| 0.1 | 3.4 | GO:0001105 | RNA polymerase II transcription coactivator activity(GO:0001105) |
| 0.1 | 0.9 | GO:0004065 | arylsulfatase activity(GO:0004065) |
| 0.1 | 2.1 | GO:0001106 | RNA polymerase II transcription corepressor activity(GO:0001106) |
| 0.1 | 6.4 | GO:0019843 | rRNA binding(GO:0019843) |
| 0.1 | 0.5 | GO:0004614 | phosphoglucomutase activity(GO:0004614) |
| 0.1 | 5.4 | GO:0046332 | SMAD binding(GO:0046332) |
| 0.1 | 0.5 | GO:0004477 | methenyltetrahydrofolate cyclohydrolase activity(GO:0004477) methylenetetrahydrofolate dehydrogenase (NADP+) activity(GO:0004488) |
| 0.1 | 2.0 | GO:0016755 | transferase activity, transferring amino-acyl groups(GO:0016755) |
| 0.1 | 1.6 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 0.1 | 7.9 | GO:0030165 | PDZ domain binding(GO:0030165) |
| 0.1 | 4.9 | GO:0016667 | oxidoreductase activity, acting on a sulfur group of donors(GO:0016667) |
| 0.1 | 6.3 | GO:0008565 | protein transporter activity(GO:0008565) |
| 0.1 | 2.4 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.1 | 1.5 | GO:0070273 | phosphatidylinositol-4-phosphate binding(GO:0070273) |
| 0.1 | 3.7 | GO:0002039 | p53 binding(GO:0002039) |
| 0.1 | 1.0 | GO:0001972 | retinoic acid binding(GO:0001972) |
| 0.1 | 2.7 | GO:0003755 | peptidyl-prolyl cis-trans isomerase activity(GO:0003755) |
| 0.1 | 0.3 | GO:0004668 | protein-arginine deiminase activity(GO:0004668) |
| 0.1 | 4.6 | GO:0017048 | Rho GTPase binding(GO:0017048) |
| 0.0 | 4.5 | GO:0008080 | N-acetyltransferase activity(GO:0008080) |
| 0.0 | 5.5 | GO:0005506 | iron ion binding(GO:0005506) |
| 0.0 | 3.3 | GO:0005254 | chloride channel activity(GO:0005254) |
| 0.0 | 3.9 | GO:0031490 | chromatin DNA binding(GO:0031490) |
| 0.0 | 2.0 | GO:0008375 | acetylglucosaminyltransferase activity(GO:0008375) |
| 0.0 | 6.8 | GO:0035091 | phosphatidylinositol binding(GO:0035091) |
| 0.0 | 0.5 | GO:0045125 | bioactive lipid receptor activity(GO:0045125) |
| 0.0 | 4.5 | GO:0008168 | methyltransferase activity(GO:0008168) |
| 0.0 | 0.4 | GO:0005251 | delayed rectifier potassium channel activity(GO:0005251) |
| 0.0 | 3.2 | GO:0031625 | ubiquitin protein ligase binding(GO:0031625) |
| 0.0 | 3.5 | GO:0008233 | peptidase activity(GO:0008233) |
| 0.0 | 0.2 | GO:0046875 | ephrin receptor binding(GO:0046875) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.7 | 29.4 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 1.6 | 17.6 | ST JAK STAT PATHWAY | Jak-STAT Pathway |
| 1.5 | 10.3 | PID IL5 PATHWAY | IL5-mediated signaling events |
| 1.3 | 91.8 | PID KIT PATHWAY | Signaling events mediated by Stem cell factor receptor (c-Kit) |
| 1.1 | 10.8 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
| 0.8 | 15.2 | PID BETA CATENIN DEG PATHWAY | Degradation of beta catenin |
| 0.7 | 9.7 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
| 0.6 | 9.7 | PID LYMPH ANGIOGENESIS PATHWAY | VEGFR3 signaling in lymphatic endothelium |
| 0.6 | 28.1 | PID FCER1 PATHWAY | Fc-epsilon receptor I signaling in mast cells |
| 0.6 | 7.8 | PID GMCSF PATHWAY | GMCSF-mediated signaling events |
| 0.6 | 2.9 | SA B CELL RECEPTOR COMPLEXES | Antigen binding to B cell receptors activates protein tyrosine kinases, such as the Src family, which ultimate activate MAP kinases. |
| 0.5 | 13.5 | PID RHODOPSIN PATHWAY | Visual signal transduction: Rods |
| 0.5 | 9.3 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 0.5 | 16.8 | PID EPHB FWD PATHWAY | EPHB forward signaling |
| 0.5 | 4.3 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.5 | 13.3 | PID TXA2PATHWAY | Thromboxane A2 receptor signaling |
| 0.5 | 14.1 | PID ECADHERIN NASCENT AJ PATHWAY | E-cadherin signaling in the nascent adherens junction |
| 0.5 | 18.7 | PID THROMBIN PAR1 PATHWAY | PAR1-mediated thrombin signaling events |
| 0.4 | 6.1 | PID INTEGRIN A9B1 PATHWAY | Alpha9 beta1 integrin signaling events |
| 0.4 | 15.5 | PID BCR 5PATHWAY | BCR signaling pathway |
| 0.4 | 25.1 | PID AR PATHWAY | Coregulation of Androgen receptor activity |
| 0.4 | 7.2 | PID MAPK TRK PATHWAY | Trk receptor signaling mediated by the MAPK pathway |
| 0.4 | 7.3 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.4 | 17.2 | PID FGF PATHWAY | FGF signaling pathway |
| 0.4 | 81.4 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.3 | 12.6 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.3 | 9.3 | PID HIF2PATHWAY | HIF-2-alpha transcription factor network |
| 0.3 | 8.1 | PID IL2 PI3K PATHWAY | IL2 signaling events mediated by PI3K |
| 0.3 | 15.8 | PID TAP63 PATHWAY | Validated transcriptional targets of TAp63 isoforms |
| 0.3 | 11.2 | PID HES HEY PATHWAY | Notch-mediated HES/HEY network |
| 0.3 | 15.6 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
| 0.3 | 4.1 | PID ERBB2 ERBB3 PATHWAY | ErbB2/ErbB3 signaling events |
| 0.2 | 5.9 | PID CXCR4 PATHWAY | CXCR4-mediated signaling events |
| 0.2 | 13.2 | PID PDGFRB PATHWAY | PDGFR-beta signaling pathway |
| 0.2 | 12.0 | PID E2F PATHWAY | E2F transcription factor network |
| 0.2 | 6.5 | ST JNK MAPK PATHWAY | JNK MAPK Pathway |
| 0.2 | 11.0 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.2 | 11.4 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.2 | 5.2 | PID ATR PATHWAY | ATR signaling pathway |
| 0.2 | 31.3 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.2 | 6.3 | PID ARF6 TRAFFICKING PATHWAY | Arf6 trafficking events |
| 0.2 | 7.9 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.2 | 0.8 | PID TCPTP PATHWAY | Signaling events mediated by TCPTP |
| 0.2 | 2.9 | PID ERBB4 PATHWAY | ErbB4 signaling events |
| 0.2 | 13.3 | PID P53 DOWNSTREAM PATHWAY | Direct p53 effectors |
| 0.1 | 3.9 | PID TNF PATHWAY | TNF receptor signaling pathway |
| 0.1 | 2.2 | PID PS1 PATHWAY | Presenilin action in Notch and Wnt signaling |
| 0.1 | 5.4 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.1 | 3.4 | PID P73PATHWAY | p73 transcription factor network |
| 0.1 | 5.2 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.1 | 2.3 | PID ARF6 PATHWAY | Arf6 signaling events |
| 0.1 | 1.9 | ST ERK1 ERK2 MAPK PATHWAY | ERK1/ERK2 MAPK Pathway |
| 0.1 | 4.8 | WNT SIGNALING | Genes related to Wnt-mediated signal transduction |
| 0.1 | 0.6 | PID DNA PK PATHWAY | DNA-PK pathway in nonhomologous end joining |
| 0.1 | 3.7 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.0 | 0.5 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.8 | 69.7 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 24 HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 24-hydroxycholesterol |
| 5.1 | 20.3 | REACTOME BETA DEFENSINS | Genes involved in Beta defensins |
| 3.6 | 58.3 | REACTOME SIGNAL REGULATORY PROTEIN SIRP FAMILY INTERACTIONS | Genes involved in Signal regulatory protein (SIRP) family interactions |
| 2.2 | 73.2 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |
| 1.9 | 26.5 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 1.6 | 9.9 | REACTOME DESTABILIZATION OF MRNA BY TRISTETRAPROLIN TTP | Genes involved in Destabilization of mRNA by Tristetraprolin (TTP) |
| 1.4 | 23.5 | REACTOME ACYL CHAIN REMODELLING OF PI | Genes involved in Acyl chain remodelling of PI |
| 1.4 | 8.1 | REACTOME INTERACTIONS OF VPR WITH HOST CELLULAR PROTEINS | Genes involved in Interactions of Vpr with host cellular proteins |
| 1.4 | 14.9 | REACTOME APOPTOSIS INDUCED DNA FRAGMENTATION | Genes involved in Apoptosis induced DNA fragmentation |
| 1.3 | 27.0 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 1.2 | 34.2 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 1.1 | 8.6 | REACTOME ORGANIC CATION ANION ZWITTERION TRANSPORT | Genes involved in Organic cation/anion/zwitterion transport |
| 1.1 | 13.9 | REACTOME SIGNALING BY FGFR3 MUTANTS | Genes involved in Signaling by FGFR3 mutants |
| 1.0 | 9.3 | REACTOME ABACAVIR TRANSPORT AND METABOLISM | Genes involved in Abacavir transport and metabolism |
| 1.0 | 24.6 | REACTOME LYSOSOME VESICLE BIOGENESIS | Genes involved in Lysosome Vesicle Biogenesis |
| 1.0 | 12.3 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.9 | 16.8 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 0.9 | 10.6 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 0.9 | 7.9 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS | Genes involved in Synthesis of bile acids and bile salts |
| 0.8 | 11.8 | REACTOME REGULATION OF SIGNALING BY CBL | Genes involved in Regulation of signaling by CBL |
| 0.8 | 22.5 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.8 | 17.5 | REACTOME DESTABILIZATION OF MRNA BY KSRP | Genes involved in Destabilization of mRNA by KSRP |
| 0.7 | 7.2 | REACTOME REGULATION OF WATER BALANCE BY RENAL AQUAPORINS | Genes involved in Regulation of Water Balance by Renal Aquaporins |
| 0.7 | 12.9 | REACTOME SEMA3A PAK DEPENDENT AXON REPULSION | Genes involved in Sema3A PAK dependent Axon repulsion |
| 0.7 | 5.4 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |
| 0.7 | 12.6 | REACTOME RECYCLING PATHWAY OF L1 | Genes involved in Recycling pathway of L1 |
| 0.6 | 23.4 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.6 | 27.6 | REACTOME PHASE1 FUNCTIONALIZATION OF COMPOUNDS | Genes involved in Phase 1 - Functionalization of compounds |
| 0.6 | 9.3 | REACTOME ADENYLATE CYCLASE ACTIVATING PATHWAY | Genes involved in Adenylate cyclase activating pathway |
| 0.6 | 8.3 | REACTOME GLUCOSE TRANSPORT | Genes involved in Glucose transport |
| 0.6 | 7.5 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.6 | 10.8 | REACTOME TRAFFICKING OF GLUR2 CONTAINING AMPA RECEPTORS | Genes involved in Trafficking of GluR2-containing AMPA receptors |
| 0.6 | 7.4 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.6 | 1.7 | REACTOME TRANSFERRIN ENDOCYTOSIS AND RECYCLING | Genes involved in Transferrin endocytosis and recycling |
| 0.6 | 9.4 | REACTOME FACILITATIVE NA INDEPENDENT GLUCOSE TRANSPORTERS | Genes involved in Facilitative Na+-independent glucose transporters |
| 0.6 | 16.0 | REACTOME METAL ION SLC TRANSPORTERS | Genes involved in Metal ion SLC transporters |
| 0.6 | 7.2 | REACTOME BASE FREE SUGAR PHOSPHATE REMOVAL VIA THE SINGLE NUCLEOTIDE REPLACEMENT PATHWAY | Genes involved in Base-free sugar-phosphate removal via the single-nucleotide replacement pathway |
| 0.5 | 6.0 | REACTOME NFKB ACTIVATION THROUGH FADD RIP1 PATHWAY MEDIATED BY CASPASE 8 AND10 | Genes involved in NF-kB activation through FADD/RIP-1 pathway mediated by caspase-8 and -10 |
| 0.5 | 22.3 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.5 | 6.0 | REACTOME ACTIVATION OF CHAPERONE GENES BY ATF6 ALPHA | Genes involved in Activation of Chaperone Genes by ATF6-alpha |
| 0.5 | 5.3 | REACTOME UNWINDING OF DNA | Genes involved in Unwinding of DNA |
| 0.5 | 11.5 | REACTOME CLASS C 3 METABOTROPIC GLUTAMATE PHEROMONE RECEPTORS | Genes involved in Class C/3 (Metabotropic glutamate/pheromone receptors) |
| 0.5 | 12.0 | REACTOME G1 S SPECIFIC TRANSCRIPTION | Genes involved in G1/S-Specific Transcription |
| 0.5 | 14.8 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
| 0.5 | 3.9 | REACTOME SIGNALLING BY NGF | Genes involved in Signalling by NGF |
| 0.5 | 8.8 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.5 | 11.8 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |
| 0.5 | 11.0 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.4 | 32.0 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.4 | 3.3 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |
| 0.4 | 5.3 | REACTOME REGULATION OF IFNG SIGNALING | Genes involved in Regulation of IFNG signaling |
| 0.4 | 5.6 | REACTOME SYNTHESIS OF SUBSTRATES IN N GLYCAN BIOSYTHESIS | Genes involved in Synthesis of substrates in N-glycan biosythesis |
| 0.3 | 8.2 | REACTOME DEPOSITION OF NEW CENPA CONTAINING NUCLEOSOMES AT THE CENTROMERE | Genes involved in Deposition of New CENPA-containing Nucleosomes at the Centromere |
| 0.3 | 5.8 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.3 | 22.5 | REACTOME ANTIVIRAL MECHANISM BY IFN STIMULATED GENES | Genes involved in Antiviral mechanism by IFN-stimulated genes |
| 0.3 | 5.6 | REACTOME SYNTHESIS OF GLYCOSYLPHOSPHATIDYLINOSITOL GPI | Genes involved in Synthesis of glycosylphosphatidylinositol (GPI) |
| 0.3 | 8.5 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 0.3 | 7.2 | REACTOME EXTENSION OF TELOMERES | Genes involved in Extension of Telomeres |
| 0.3 | 0.8 | REACTOME OLFACTORY SIGNALING PATHWAY | Genes involved in Olfactory Signaling Pathway |
| 0.2 | 5.4 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.2 | 5.8 | REACTOME CELL CELL JUNCTION ORGANIZATION | Genes involved in Cell-cell junction organization |
| 0.2 | 2.3 | REACTOME HORMONE LIGAND BINDING RECEPTORS | Genes involved in Hormone ligand-binding receptors |
| 0.2 | 2.9 | REACTOME SIGNALING BY FGFR1 FUSION MUTANTS | Genes involved in Signaling by FGFR1 fusion mutants |
| 0.2 | 9.5 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.2 | 4.3 | REACTOME RNA POL III TRANSCRIPTION INITIATION FROM TYPE 2 PROMOTER | Genes involved in RNA Polymerase III Transcription Initiation From Type 2 Promoter |
| 0.2 | 3.1 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.2 | 3.5 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.2 | 12.7 | REACTOME ASPARAGINE N LINKED GLYCOSYLATION | Genes involved in Asparagine N-linked glycosylation |
| 0.2 | 3.0 | REACTOME CA DEPENDENT EVENTS | Genes involved in Ca-dependent events |
| 0.2 | 4.1 | REACTOME ANTIGEN ACTIVATES B CELL RECEPTOR LEADING TO GENERATION OF SECOND MESSENGERS | Genes involved in Antigen Activates B Cell Receptor Leading to Generation of Second Messengers |
| 0.2 | 4.9 | REACTOME RNA POL I TRANSCRIPTION INITIATION | Genes involved in RNA Polymerase I Transcription Initiation |
| 0.2 | 2.3 | REACTOME GROWTH HORMONE RECEPTOR SIGNALING | Genes involved in Growth hormone receptor signaling |
| 0.2 | 6.3 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.2 | 10.8 | REACTOME ANTIGEN PROCESSING CROSS PRESENTATION | Genes involved in Antigen processing-Cross presentation |
| 0.1 | 1.9 | REACTOME SHC1 EVENTS IN ERBB4 SIGNALING | Genes involved in SHC1 events in ERBB4 signaling |
| 0.1 | 12.5 | REACTOME TOLL RECEPTOR CASCADES | Genes involved in Toll Receptor Cascades |
| 0.1 | 7.4 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.1 | 2.2 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.1 | 18.7 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.1 | 2.0 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.1 | 1.4 | REACTOME TRANSCRIPTION COUPLED NER TC NER | Genes involved in Transcription-coupled NER (TC-NER) |
| 0.1 | 1.5 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.1 | 1.6 | REACTOME FORMATION OF ATP BY CHEMIOSMOTIC COUPLING | Genes involved in Formation of ATP by chemiosmotic coupling |
| 0.1 | 0.9 | REACTOME THE ACTIVATION OF ARYLSULFATASES | Genes involved in The activation of arylsulfatases |
| 0.1 | 4.3 | REACTOME BIOLOGICAL OXIDATIONS | Genes involved in Biological oxidations |
| 0.1 | 0.9 | REACTOME HYALURONAN UPTAKE AND DEGRADATION | Genes involved in Hyaluronan uptake and degradation |
| 0.1 | 1.3 | REACTOME MITOCHONDRIAL TRNA AMINOACYLATION | Genes involved in Mitochondrial tRNA aminoacylation |
| 0.1 | 0.8 | REACTOME IL RECEPTOR SHC SIGNALING | Genes involved in Interleukin receptor SHC signaling |
| 0.1 | 1.8 | REACTOME CELL SURFACE INTERACTIONS AT THE VASCULAR WALL | Genes involved in Cell surface interactions at the vascular wall |
| 0.0 | 0.5 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GIP | Genes involved in Synthesis, Secretion, and Inactivation of Glucose-dependent Insulinotropic Polypeptide (GIP) |
| 0.0 | 1.9 | REACTOME GLYCEROPHOSPHOLIPID BIOSYNTHESIS | Genes involved in Glycerophospholipid biosynthesis |
| 0.0 | 0.3 | REACTOME TERMINATION OF O GLYCAN BIOSYNTHESIS | Genes involved in Termination of O-glycan biosynthesis |
| 0.0 | 2.0 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.0 | 0.6 | REACTOME SIGNALING BY NODAL | Genes involved in Signaling by NODAL |
| 0.0 | 1.1 | REACTOME INFLUENZA VIRAL RNA TRANSCRIPTION AND REPLICATION | Genes involved in Influenza Viral RNA Transcription and Replication |