Project
GSE49485: Hypoxia transcriptome sequencing of rat brain.
Navigation
Downloads
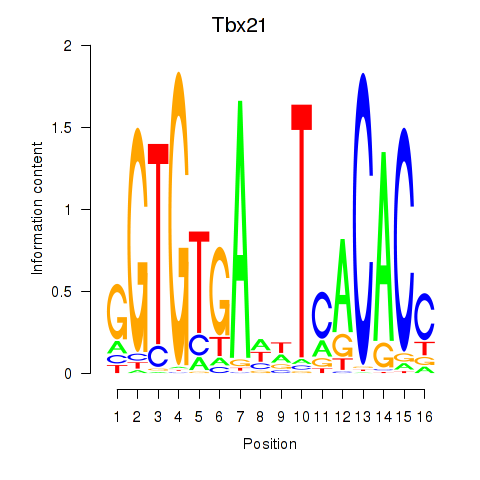
Results for Tbx21
Z-value: 0.30
Transcription factors associated with Tbx21
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Tbx21
|
ENSRNOG00000009427 | T-box 21 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Tbx21 | rn6_v1_chr10_-_85049331_85049331 | -0.24 | 7.0e-01 | Click! |
Activity profile of Tbx21 motif
Sorted Z-values of Tbx21 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr4_-_123494742 | 0.19 |
ENSRNOT00000073268
|
Slc41a3
|
solute carrier family 41, member 3 |
| chr14_-_82055290 | 0.18 |
ENSRNOT00000058062
|
RGD1560394
|
RGD1560394 |
| chr1_+_89162639 | 0.13 |
ENSRNOT00000028508
|
Atp4a
|
ATPase H+/K+ transporting alpha subunit |
| chr1_+_207508414 | 0.12 |
ENSRNOT00000054896
|
Nps
|
neuropeptide S |
| chr19_+_16415636 | 0.12 |
ENSRNOT00000089975
|
Irx5
|
iroquois homeobox 5 |
| chr9_+_14951047 | 0.11 |
ENSRNOT00000074267
|
LOC100911489
|
uncharacterized LOC100911489 |
| chr7_+_130542202 | 0.11 |
ENSRNOT00000079501
ENSRNOT00000045647 |
Acr
|
acrosin |
| chr1_-_216663720 | 0.11 |
ENSRNOT00000078944
ENSRNOT00000077409 |
Cdkn1c
|
cyclin-dependent kinase inhibitor 1C |
| chr5_-_151459037 | 0.10 |
ENSRNOT00000064472
ENSRNOT00000087836 |
Sytl1
|
synaptotagmin-like 1 |
| chr12_-_22478752 | 0.09 |
ENSRNOT00000089392
ENSRNOT00000086915 |
Ache
|
acetylcholinesterase |
| chr3_-_2938883 | 0.09 |
ENSRNOT00000084196
|
LOC108348105
|
allergen Fel d 4-like |
| chr9_+_16139101 | 0.09 |
ENSRNOT00000070803
|
LOC100910668
|
uncharacterized LOC100910668 |
| chr10_+_109786222 | 0.08 |
ENSRNOT00000054953
|
Npb
|
neuropeptide B |
| chr2_+_58724855 | 0.08 |
ENSRNOT00000089609
|
Capsl
|
calcyphosine-like |
| chr10_+_72486152 | 0.08 |
ENSRNOT00000004193
|
RGD1310166
|
similar to Chromodomain-helicase-DNA-binding protein 1 (CHD-1) |
| chr4_-_179700130 | 0.08 |
ENSRNOT00000021306
|
Lmntd1
|
lamin tail domain containing 1 |
| chr13_+_96195836 | 0.07 |
ENSRNOT00000042547
|
RGD1563812
|
similar to basic transcription factor 3 |
| chr1_-_222250980 | 0.07 |
ENSRNOT00000028734
|
Nudt22
|
nudix hydrolase 22 |
| chr1_-_24982606 | 0.07 |
ENSRNOT00000074143
|
RGD1562660
|
RGD1562660 |
| chr1_-_24891461 | 0.07 |
ENSRNOT00000049208
|
LOC501467
|
similar to spermatogenesis associated glutamate (E)-rich protein 4d |
| chr6_+_75429729 | 0.06 |
ENSRNOT00000043250
|
Rps10l1
|
ribosomal protein S10-like 1 |
| chr19_-_28751584 | 0.06 |
ENSRNOT00000079270
|
AABR07043456.1
|
|
| chr16_+_81616604 | 0.06 |
ENSRNOT00000026392
ENSRNOT00000057740 |
Adprhl1
Grtp1
|
ADP-ribosylhydrolase like 1 growth hormone regulated TBC protein 1 |
| chr10_-_104054805 | 0.06 |
ENSRNOT00000004853
|
Nt5c
|
5', 3'-nucleotidase, cytosolic |
| chr11_+_67451549 | 0.06 |
ENSRNOT00000042777
|
Stfa2l2
|
stefin A2-like 2 |
| chr4_-_122337389 | 0.06 |
ENSRNOT00000024222
|
Cfap100
|
cilia and flagella associated protein 100 |
| chr3_-_7278758 | 0.06 |
ENSRNOT00000017004
|
Spaca9
|
sperm acrosome associated 9 |
| chr10_-_57618527 | 0.06 |
ENSRNOT00000037517
|
C1qbp
|
complement C1q binding protein |
| chr16_-_79793619 | 0.06 |
ENSRNOT00000073971
|
Arhgef10
|
Rho guanine nucleotide exchange factor 10 |
| chr2_-_200513564 | 0.06 |
ENSRNOT00000056173
|
Phgdh
|
phosphoglycerate dehydrogenase |
| chr1_+_201429771 | 0.05 |
ENSRNOT00000027836
|
Plekha1
|
pleckstrin homology domain containing A1 |
| chr1_-_20417637 | 0.05 |
ENSRNOT00000036049
|
RGD1559962
|
similar to High mobility group protein 2 (HMG-2) |
| chr13_+_52553775 | 0.05 |
ENSRNOT00000011991
|
Csrp1
|
cysteine and glycine-rich protein 1 |
| chr11_-_36479868 | 0.05 |
ENSRNOT00000075762
|
LOC100911295
|
non-histone chromosomal protein HMG-14-like |
| chr7_-_119158173 | 0.05 |
ENSRNOT00000067483
ENSRNOT00000078528 |
Txn2
|
thioredoxin 2 |
| chr7_+_120273117 | 0.05 |
ENSRNOT00000014349
|
Galr3
|
galanin receptor 3 |
| chr1_+_248132090 | 0.05 |
ENSRNOT00000022056
|
Il33
|
interleukin 33 |
| chr19_-_42180981 | 0.04 |
ENSRNOT00000019577
|
Pmfbp1
|
polyamine modulated factor 1 binding protein 1 |
| chr7_-_73270308 | 0.04 |
ENSRNOT00000007430
|
Rida
|
reactive intermediate imine deaminase A homolog |
| chr5_+_171242897 | 0.04 |
ENSRNOT00000032859
|
RGD1304567
|
similar to RIKEN cDNA A430005L14 |
| chr1_+_80383050 | 0.04 |
ENSRNOT00000023451
|
Exoc3l2
|
exocyst complex component 3-like 2 |
| chr5_+_172648950 | 0.04 |
ENSRNOT00000055361
|
Faap20
|
Fanconi anemia core complex associated protein 20 |
| chr17_-_44520240 | 0.04 |
ENSRNOT00000086538
|
Hist1h2bh
|
histone cluster 1 H2B family member h |
| chr7_+_11769400 | 0.04 |
ENSRNOT00000044417
|
Jsrp1
|
junctional sarcoplasmic reticulum protein 1 |
| chr18_-_6782996 | 0.04 |
ENSRNOT00000090320
|
Aqp4
|
aquaporin 4 |
| chr14_-_7054548 | 0.04 |
ENSRNOT00000002973
|
Nudt9
|
nudix hydrolase 9 |
| chr9_-_11114843 | 0.04 |
ENSRNOT00000072031
|
Ebi3
|
Epstein-Barr virus induced 3 |
| chr13_+_52854031 | 0.04 |
ENSRNOT00000013738
ENSRNOT00000086004 |
Tmem9
|
transmembrane protein 9 |
| chr5_+_164972480 | 0.04 |
ENSRNOT00000012670
|
Fbxo2
|
F-box protein 2 |
| chr4_-_162025090 | 0.04 |
ENSRNOT00000085887
ENSRNOT00000009904 |
Klrb1a
|
killer cell lectin-like receptor subfamily B, member 1A |
| chr11_+_70056624 | 0.04 |
ENSRNOT00000002447
|
AC133403.1
|
|
| chr7_+_2479311 | 0.04 |
ENSRNOT00000003787
|
Ptges3
|
prostaglandin E synthase 3 |
| chr10_-_63219309 | 0.03 |
ENSRNOT00000076357
|
Blmh
|
bleomycin hydrolase |
| chr10_-_7240812 | 0.03 |
ENSRNOT00000003700
|
Mettl22
|
methyltransferase like 22 |
| chr7_-_12810570 | 0.03 |
ENSRNOT00000012578
|
Fstl3
|
follistatin like 3 |
| chr3_-_113405829 | 0.03 |
ENSRNOT00000036823
|
Ell3
|
elongation factor for RNA polymerase II 3 |
| chr16_+_20704807 | 0.03 |
ENSRNOT00000027241
|
Klhl26
|
kelch-like family member 26 |
| chr6_-_127534247 | 0.03 |
ENSRNOT00000012500
|
Serpina6
|
serpin family A member 6 |
| chr8_-_103608913 | 0.03 |
ENSRNOT00000013209
|
Pls1
|
plastin 1 |
| chr8_-_68525911 | 0.03 |
ENSRNOT00000080588
ENSRNOT00000011109 |
Iqch
|
IQ motif containing H |
| chr1_-_101095594 | 0.03 |
ENSRNOT00000027944
|
Fcgrt
|
Fc fragment of IgG receptor and transporter |
| chr6_+_72461977 | 0.03 |
ENSRNOT00000007887
ENSRNOT00000090716 |
Ap4s1
|
adaptor-related protein complex 4, sigma 1 subunit |
| chr5_-_138239306 | 0.03 |
ENSRNOT00000039305
|
Ermap
|
erythroblast membrane-associated protein |
| chr12_-_18526250 | 0.03 |
ENSRNOT00000001791
|
Asmt
|
acetylserotonin O-methyltransferase |
| chr16_+_7758996 | 0.03 |
ENSRNOT00000061063
|
Btd
|
biotinidase |
| chr9_+_73418607 | 0.03 |
ENSRNOT00000092547
|
Map2
|
microtubule-associated protein 2 |
| chr2_-_224802997 | 0.03 |
ENSRNOT00000032324
|
Tmem56
|
transmembrane protein 56 |
| chr11_+_86421106 | 0.03 |
ENSRNOT00000002599
|
LOC100362453
|
serine/threonine-protein phosphatase 2A catalytic subunit alpha-like |
| chr7_+_2458264 | 0.03 |
ENSRNOT00000003550
|
Naca
|
nascent polypeptide-associated complex alpha subunit |
| chr2_+_30664217 | 0.03 |
ENSRNOT00000076786
|
Ak6
|
adenylate kinase 6 |
| chr11_-_71938165 | 0.03 |
ENSRNOT00000037795
|
Cep19
|
centrosomal protein 19 |
| chr5_+_122508388 | 0.03 |
ENSRNOT00000038410
|
Tctex1d1
|
Tctex1 domain containing 1 |
| chr17_+_57040023 | 0.03 |
ENSRNOT00000020204
|
Crem
|
cAMP responsive element modulator |
| chr2_-_198439454 | 0.03 |
ENSRNOT00000028780
|
Fcgr1a
|
Fc fragment of IgG receptor Ia |
| chr8_-_79119720 | 0.03 |
ENSRNOT00000086562
|
Tex9
|
testis expressed 9 |
| chr14_+_38030189 | 0.03 |
ENSRNOT00000035623
|
Txk
|
TXK tyrosine kinase |
| chr19_+_52217984 | 0.03 |
ENSRNOT00000079580
|
Dnaaf1
|
dynein, axonemal, assembly factor 1 |
| chr3_-_46601409 | 0.03 |
ENSRNOT00000079261
|
Pla2r1
|
phospholipase A2 receptor 1 |
| chr3_+_11396070 | 0.03 |
ENSRNOT00000037399
|
Ciz1
|
CDKN1A interacting zinc finger protein 1 |
| chr17_+_18059382 | 0.03 |
ENSRNOT00000031656
|
Nhlrc1
|
NHL repeat containing E3 ubiquitin protein ligase 1 |
| chr4_-_84127331 | 0.03 |
ENSRNOT00000085812
|
Cpvl
|
carboxypeptidase, vitellogenic-like |
| chr6_-_92643847 | 0.03 |
ENSRNOT00000009183
|
Pygl
|
glycogen phosphorylase L |
| chr8_-_23292682 | 0.02 |
ENSRNOT00000082252
|
LOC103690166
|
zinc finger protein OZF-like |
| chr16_+_69048730 | 0.02 |
ENSRNOT00000086082
ENSRNOT00000078128 |
Rab11fip1
|
RAB11 family interacting protein 1 |
| chr8_-_36760742 | 0.02 |
ENSRNOT00000017307
|
Ddx25
|
DEAD-box helicase 25 |
| chr1_-_198476476 | 0.02 |
ENSRNOT00000027525
|
Kif22
|
kinesin family member 22 |
| chr18_+_86307646 | 0.02 |
ENSRNOT00000091778
|
Cd226
|
CD226 molecule |
| chr19_-_43215281 | 0.02 |
ENSRNOT00000025052
|
Aars
|
alanyl-tRNA synthetase |
| chr8_-_2045817 | 0.02 |
ENSRNOT00000009542
ENSRNOT00000081171 ENSRNOT00000078765 |
Gria4
|
glutamate ionotropic receptor AMPA type subunit 4 |
| chr16_+_49185560 | 0.02 |
ENSRNOT00000014360
|
Helt
|
helt bHLH transcription factor |
| chr3_-_114235933 | 0.02 |
ENSRNOT00000023939
|
Duox2
|
dual oxidase 2 |
| chr1_-_101514547 | 0.02 |
ENSRNOT00000079633
|
Ppp1r15a
|
protein phosphatase 1, regulatory subunit 15A |
| chr10_+_7041510 | 0.02 |
ENSRNOT00000003514
|
Carhsp1
|
calcium regulated heat stable protein 1 |
| chr1_+_63263722 | 0.02 |
ENSRNOT00000088203
|
LOC100911184
|
vomeronasal type-1 receptor 4-like |
| chr12_-_9727489 | 0.02 |
ENSRNOT00000001261
|
Polr1d
|
RNA polymerase I subunit D |
| chr19_-_28666993 | 0.02 |
ENSRNOT00000083821
|
AABR07043453.1
|
|
| chr16_+_80729959 | 0.02 |
ENSRNOT00000082049
|
Tdrp
|
testis development related protein |
| chr15_+_18941431 | 0.02 |
ENSRNOT00000092092
|
AABR07017236.1
|
|
| chr7_+_117643206 | 0.02 |
ENSRNOT00000077588
|
Adck5
|
aarF domain containing kinase 5 |
| chr15_+_45422010 | 0.02 |
ENSRNOT00000012231
|
Rnaseh2b
|
ribonuclease H2, subunit B |
| chr1_+_201672528 | 0.02 |
ENSRNOT00000093490
|
Dmbt1
|
deleted in malignant brain tumors 1 |
| chr19_-_28044531 | 0.02 |
ENSRNOT00000086160
|
AABR07043395.1
|
|
| chr3_-_161067358 | 0.02 |
ENSRNOT00000055160
|
Wfdc8
|
WAP four-disulfide core domain 8 |
| chr16_+_37177443 | 0.02 |
ENSRNOT00000014083
|
Cep44
|
centrosomal protein 44 |
| chr16_-_31301880 | 0.02 |
ENSRNOT00000084847
ENSRNOT00000083943 |
AABR07025272.1
|
|
| chr2_+_209661244 | 0.02 |
ENSRNOT00000091973
|
AABR07012826.1
|
|
| chr3_-_2841331 | 0.02 |
ENSRNOT00000061874
|
Ccdc183
|
coiled-coil domain containing 183 |
| chr1_+_222250995 | 0.02 |
ENSRNOT00000031458
|
Trpt1
|
tRNA phosphotransferase 1 |
| chr19_-_43215077 | 0.02 |
ENSRNOT00000082151
|
Aars
|
alanyl-tRNA synthetase |
| chr4_+_146276862 | 0.02 |
ENSRNOT00000009705
|
Slc6a1
|
solute carrier family 6 member 1 |
| chr5_-_143120165 | 0.02 |
ENSRNOT00000012314
|
Zc3h12a
|
zinc finger CCCH type containing 12A |
| chr4_+_51614676 | 0.02 |
ENSRNOT00000060494
|
Asb15
|
ankyrin repeat and SOCS box containing 15 |
| chr15_+_28295368 | 0.02 |
ENSRNOT00000013786
|
Slc39a2
|
solute carrier family 39 member 2 |
| chr1_-_189182306 | 0.02 |
ENSRNOT00000021249
|
Gp2
|
glycoprotein 2 |
| chr4_+_154391647 | 0.02 |
ENSRNOT00000081488
ENSRNOT00000079192 |
Mug1
|
murinoglobulin 1 |
| chr19_+_248622 | 0.02 |
ENSRNOT00000078228
|
AABR07042607.1
|
|
| chr4_+_145330457 | 0.02 |
ENSRNOT00000011937
|
Arpc4
|
actin related protein 2/3 complex, subunit 4 |
| chr7_-_123287721 | 0.02 |
ENSRNOT00000009152
|
LOC100359574
|
NHP2 non-histone chromosome protein 2-like 1-like |
| chr4_-_164406146 | 0.02 |
ENSRNOT00000090110
|
Klra22
|
killer cell lectin-like receptor subfamily A, member 22 |
| chr5_+_69833225 | 0.01 |
ENSRNOT00000013759
|
Nipsnap3b
|
nipsnap homolog 3B |
| chr16_-_7007287 | 0.01 |
ENSRNOT00000041216
|
Itih3
|
inter-alpha trypsin inhibitor, heavy chain 3 |
| chr7_-_15706893 | 0.01 |
ENSRNOT00000042742
|
Olr1092
|
olfactory receptor 1092 |
| chr13_-_74005486 | 0.01 |
ENSRNOT00000090173
|
AABR07021465.1
|
|
| chr20_-_14001269 | 0.01 |
ENSRNOT00000093187
|
Ggt5
|
gamma-glutamyltransferase 5 |
| chr7_+_14441476 | 0.01 |
ENSRNOT00000093687
|
Cyp4f39
|
cytochrome P450, family 4, subfamily f, polypeptide 39 |
| chr7_+_93975451 | 0.01 |
ENSRNOT00000011379
|
Colec10
|
collectin subfamily member 10 |
| chr5_-_150704117 | 0.01 |
ENSRNOT00000067455
|
Sesn2
|
sestrin 2 |
| chr20_+_47596575 | 0.01 |
ENSRNOT00000087230
ENSRNOT00000064905 |
Scml4
|
sex comb on midleg-like 4 (Drosophila) |
| chr8_+_132135187 | 0.01 |
ENSRNOT00000091791
|
Tgm4
|
transglutaminase 4 |
| chr5_-_151029233 | 0.01 |
ENSRNOT00000089155
|
Ppp1r8
|
protein phosphatase 1, regulatory subunit 8 |
| chr1_-_141533908 | 0.01 |
ENSRNOT00000020117
|
Mesp1
|
mesoderm posterior bHLH transcription factor 1 |
| chr11_+_39482408 | 0.01 |
ENSRNOT00000075126
|
Hmgn1
|
high mobility group nucleosome binding domain 1 |
| chr1_-_198298076 | 0.01 |
ENSRNOT00000027033
|
Ino80e
|
INO80 complex subunit E |
| chrX_-_64702441 | 0.01 |
ENSRNOT00000051132
|
Amer1
|
APC membrane recruitment protein 1 |
| chr9_-_104700609 | 0.01 |
ENSRNOT00000081815
ENSRNOT00000018503 |
Slco6c1
|
solute carrier organic anion transporter family, member 6c1 |
| chr5_+_152681101 | 0.01 |
ENSRNOT00000076052
ENSRNOT00000022574 |
Stmn1
|
stathmin 1 |
| chr13_+_82369493 | 0.01 |
ENSRNOT00000003733
|
Sell
|
selectin L |
| chr3_+_71020534 | 0.01 |
ENSRNOT00000007276
|
Zc3h15
|
zinc finger CCCH-type containing 15 |
| chr11_+_36497509 | 0.01 |
ENSRNOT00000002222
|
Wrb
|
tryptophan rich basic protein |
| chr16_+_80729400 | 0.01 |
ENSRNOT00000036383
|
Tdrp
|
testis development related protein |
| chr1_+_56242346 | 0.01 |
ENSRNOT00000019317
|
Smoc2
|
SPARC related modular calcium binding 2 |
| chr9_+_4269160 | 0.01 |
ENSRNOT00000050420
|
Sult1c2
|
sulfotransferase family 1C member 2 |
| chrX_-_140303686 | 0.01 |
ENSRNOT00000033219
|
Gpr101
|
G protein-coupled receptor 101 |
| chr7_-_11982141 | 0.01 |
ENSRNOT00000024980
|
Scamp4
|
secretory carrier membrane protein 4 |
| chr7_+_120176530 | 0.01 |
ENSRNOT00000087548
|
Triobp
|
TRIO and F-actin binding protein |
| chrX_-_123092217 | 0.01 |
ENSRNOT00000039710
|
Gm14569
|
predicted gene 14569 |
| chr7_+_15621989 | 0.01 |
ENSRNOT00000051928
|
Olr1095
|
olfactory receptor 1095 |
| chr3_+_160945359 | 0.01 |
ENSRNOT00000019761
|
Pigt
|
phosphatidylinositol glycan anchor biosynthesis, class T |
| chr10_-_107424710 | 0.01 |
ENSRNOT00000004320
|
Lgals3bp
|
galectin 3 binding protein |
| chr1_+_212714179 | 0.01 |
ENSRNOT00000054880
ENSRNOT00000025680 |
Olr292
|
olfactory receptor 292 |
| chr17_-_44841382 | 0.01 |
ENSRNOT00000080119
|
Hist1h2ak
|
histone cluster 1, H2ak |
| chr1_-_67284864 | 0.01 |
ENSRNOT00000082908
|
LOC691661
|
similar to zinc finger and SCAN domain containing 4 |
| chr1_-_81152384 | 0.01 |
ENSRNOT00000026263
ENSRNOT00000077305 |
Zfp94
|
zinc finger protein 94 |
| chr12_+_46316236 | 0.01 |
ENSRNOT00000001508
|
Prkab1
|
protein kinase AMP-activated non-catalytic subunit beta 1 |
| chr11_+_82373870 | 0.01 |
ENSRNOT00000002429
|
Tra2b
|
transformer 2 beta homolog (Drosophila) |
| chr6_-_103187275 | 0.01 |
ENSRNOT00000083272
|
LOC100359616
|
ribosomal protein L36-like |
| chr6_+_43363792 | 0.01 |
ENSRNOT00000091061
|
Cpsf3
|
cleavage and polyadenylation specific factor 3 |
| chr2_-_206274079 | 0.01 |
ENSRNOT00000056079
|
Hipk1
|
homeodomain interacting protein kinase 1 |
| chr4_+_118243132 | 0.01 |
ENSRNOT00000023654
|
RGD1306746
|
similar to Hypothetical protein MGC25529 |
| chr3_+_149802621 | 0.01 |
ENSRNOT00000071772
|
LOC690507
|
similar to Vomeromodulin |
| chr7_+_38897278 | 0.01 |
ENSRNOT00000006450
|
Epyc
|
epiphycan |
| chr4_+_181103774 | 0.01 |
ENSRNOT00000084207
ENSRNOT00000055473 ENSRNOT00000077619 |
Arntl2
|
aryl hydrocarbon receptor nuclear translocator-like 2 |
| chr3_-_125533600 | 0.01 |
ENSRNOT00000040802
|
Lrrn4
|
leucine rich repeat neuronal 4 |
| chr1_+_196095214 | 0.01 |
ENSRNOT00000080741
|
LOC691716
|
similar to ribosomal protein S15a |
| chr5_-_68139199 | 0.01 |
ENSRNOT00000009987
|
Plppr1
|
phospholipid phosphatase related 1 |
| chr8_+_70712513 | 0.01 |
ENSRNOT00000079548
|
Parp16
|
poly (ADP-ribose) polymerase family, member 16 |
| chr1_-_66212418 | 0.01 |
ENSRNOT00000026074
|
LOC691722
|
hypothetical protein LOC691722 |
| chr10_+_105498728 | 0.01 |
ENSRNOT00000032163
|
Sphk1
|
sphingosine kinase 1 |
| chr7_+_140758615 | 0.01 |
ENSRNOT00000089448
|
Troap
|
trophinin associated protein |
| chr20_-_34929965 | 0.00 |
ENSRNOT00000004499
|
Mcm9
|
minichromosome maintenance 9 homologous recombination repair factor |
| chr6_+_132551394 | 0.00 |
ENSRNOT00000019583
|
Evl
|
Enah/Vasp-like |
| chr15_-_42751330 | 0.00 |
ENSRNOT00000066478
|
Adam2
|
ADAM metallopeptidase domain 2 |
| chr13_+_103300932 | 0.00 |
ENSRNOT00000085214
ENSRNOT00000038264 |
Eprs
|
glutamyl-prolyl-tRNA synthetase |
| chr3_-_46051096 | 0.00 |
ENSRNOT00000081302
|
Baz2b
|
bromodomain adjacent to zinc finger domain, 2B |
| chr8_-_62410338 | 0.00 |
ENSRNOT00000026358
|
Csk
|
c-src tyrosine kinase |
| chr2_-_57196568 | 0.00 |
ENSRNOT00000060146
|
Wdr70
|
WD repeat domain 70 |
| chr7_+_107695227 | 0.00 |
ENSRNOT00000009673
|
Wisp1
|
WNT1 inducible signaling pathway protein 1 |
| chrX_+_123205869 | 0.00 |
ENSRNOT00000017101
ENSRNOT00000079231 |
Pgrmc1
|
progesterone receptor membrane component 1 |
| chr1_+_81328183 | 0.00 |
ENSRNOT00000074985
ENSRNOT00000075167 |
Plaur
|
plasminogen activator, urokinase receptor |
| chr17_+_90696019 | 0.00 |
ENSRNOT00000003438
|
Gpr137b
|
G protein-coupled receptor 137B |
| chr2_+_140541619 | 0.00 |
ENSRNOT00000017396
|
Rab33b
|
RAB33B, member RAS oncogene family |
| chr5_+_134593517 | 0.00 |
ENSRNOT00000013471
|
Efcab14
|
EF-hand calcium binding domain 14 |
| chr20_+_2044200 | 0.00 |
ENSRNOT00000092626
|
RT1-M4
|
RT1 class Ib, locus M4 |
| chr5_-_58288125 | 0.00 |
ENSRNOT00000068752
|
Fam205a
|
family with sequence similarity 205, member A |
| chrX_+_156274800 | 0.00 |
ENSRNOT00000080887
|
G6pd
|
glucose-6-phosphate dehydrogenase |
| chr19_+_52258947 | 0.00 |
ENSRNOT00000021072
|
Adad2
|
adenosine deaminase domain containing 2 |
Network of associatons between targets according to the STRING database.
First level regulatory network of Tbx21
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0010841 | positive regulation of circadian sleep/wake cycle, wakefulness(GO:0010841) |
| 0.0 | 0.1 | GO:1902746 | regulation of lens fiber cell differentiation(GO:1902746) |
| 0.0 | 0.1 | GO:1901165 | positive regulation of trophoblast cell migration(GO:1901165) |
| 0.0 | 0.1 | GO:0007341 | penetration of zona pellucida(GO:0007341) |
| 0.0 | 0.1 | GO:0051725 | protein de-ADP-ribosylation(GO:0051725) |
| 0.0 | 0.1 | GO:0009223 | pyrimidine deoxyribonucleotide catabolic process(GO:0009223) |
| 0.0 | 0.0 | GO:0061518 | microglial cell activation involved in immune response(GO:0002282) negative regulation of immunoglobulin secretion(GO:0051025) macrophage proliferation(GO:0061517) microglial cell proliferation(GO:0061518) |
| 0.0 | 0.1 | GO:0006564 | L-serine biosynthetic process(GO:0006564) threonine metabolic process(GO:0006566) |
| 0.0 | 0.1 | GO:0060040 | retinal bipolar neuron differentiation(GO:0060040) |
| 0.0 | 0.1 | GO:1900619 | acetylcholine metabolic process(GO:0008291) acetate ester metabolic process(GO:1900619) |
| 0.0 | 0.0 | GO:0030186 | melatonin metabolic process(GO:0030186) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0043159 | acrosomal matrix(GO:0043159) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0004040 | amidase activity(GO:0004040) |
| 0.0 | 0.1 | GO:0008900 | hydrogen:potassium-exchanging ATPase activity(GO:0008900) |
| 0.0 | 0.1 | GO:0003990 | acetylcholinesterase activity(GO:0003990) |
| 0.0 | 0.1 | GO:0030984 | kininogen binding(GO:0030984) |
| 0.0 | 0.0 | GO:0044378 | non-sequence-specific DNA binding, bending(GO:0044378) |
| 0.0 | 0.0 | GO:0033743 | peptide-methionine (R)-S-oxide reductase activity(GO:0033743) |
| 0.0 | 0.1 | GO:0019770 | IgG receptor activity(GO:0019770) |
| 0.0 | 0.0 | GO:0050220 | prostaglandin-E synthase activity(GO:0050220) |
| 0.0 | 0.0 | GO:0047708 | biotinidase activity(GO:0047708) |