Project
GSE49485: Hypoxia transcriptome sequencing of rat brain.
Navigation
Downloads
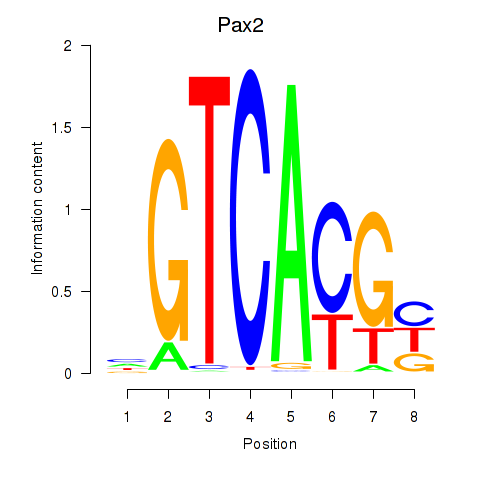
Results for Pax2
Z-value: 0.09
Transcription factors associated with Pax2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Pax2
|
ENSRNOG00000014253 | paired box 2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Pax2 | rn6_v1_chr1_+_264504591_264504591 | -0.35 | 5.6e-01 | Click! |
Activity profile of Pax2 motif
Sorted Z-values of Pax2 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr4_+_33638709 | 0.03 |
ENSRNOT00000009888
ENSRNOT00000034719 ENSRNOT00000052333 |
Tac1
|
tachykinin, precursor 1 |
| chr3_-_161299024 | 0.01 |
ENSRNOT00000021216
|
Neurl2
|
neuralized E3 ubiquitin protein ligase 2 |
| chr2_-_197991198 | 0.01 |
ENSRNOT00000056322
|
Ciart
|
circadian associated repressor of transcription |
| chr2_-_197991574 | 0.01 |
ENSRNOT00000085632
|
Ciart
|
circadian associated repressor of transcription |
| chr10_-_56276764 | 0.01 |
ENSRNOT00000049048
|
Eif4a1
|
eukaryotic translation initiation factor 4A1 |
| chr19_+_37229120 | 0.01 |
ENSRNOT00000076335
|
Hsf4
|
heat shock transcription factor 4 |
| chr1_+_154377247 | 0.01 |
ENSRNOT00000092945
|
Picalm
|
phosphatidylinositol binding clathrin assembly protein |
| chrX_-_54303729 | 0.01 |
ENSRNOT00000087919
ENSRNOT00000064340 ENSRNOT00000051249 ENSRNOT00000087547 |
Gk
|
glycerol kinase |
| chr13_-_42263024 | 0.01 |
ENSRNOT00000004741
|
Lypd1
|
Ly6/Plaur domain containing 1 |
| chr6_+_108167716 | 0.01 |
ENSRNOT00000064426
|
Lin52
|
lin-52 DREAM MuvB core complex component |
| chr5_+_64326733 | 0.01 |
ENSRNOT00000065775
|
Tmeff1
|
transmembrane protein with EGF-like and two follistatin-like domains 1 |
| chr2_+_189400696 | 0.01 |
ENSRNOT00000046919
ENSRNOT00000089801 |
LOC361985
|
similar to NICE-3 |
| chr7_-_92882068 | 0.01 |
ENSRNOT00000037809
|
Ext1
|
exostosin glycosyltransferase 1 |
| chrX_+_77263359 | 0.01 |
ENSRNOT00000077604
|
Pgk1
|
phosphoglycerate kinase 1 |
| chr11_-_60613718 | 0.00 |
ENSRNOT00000002906
|
Atg3
|
autophagy related 3 |
| chr6_+_28663602 | 0.00 |
ENSRNOT00000005402
|
Ptrhd1
|
peptidyl-tRNA hydrolase domain containing 1 |
| chr10_+_55940533 | 0.00 |
ENSRNOT00000012061
|
RGD1563441
|
similar to RIKEN cDNA A030009H04 |
| chr10_-_56839437 | 0.00 |
ENSRNOT00000090338
|
Bcl6b
|
B-cell CLL/lymphoma 6B |
| chr9_-_113331319 | 0.00 |
ENSRNOT00000020681
|
Vapa
|
VAMP associated protein A |
| chr12_+_19303387 | 0.00 |
ENSRNOT00000001820
|
Cops6
|
COP9 signalosome subunit 6 |
| chr8_-_58253688 | 0.00 |
ENSRNOT00000010956
|
Cul5
|
cullin 5 |
| chr5_+_128629942 | 0.00 |
ENSRNOT00000010462
ENSRNOT00000010440 |
Nrdc
|
nardilysin convertase |
| chr5_+_147375350 | 0.00 |
ENSRNOT00000010674
|
Yars
|
tyrosyl-tRNA synthetase |
| chr1_+_102849889 | 0.00 |
ENSRNOT00000066791
|
Gtf2h1
|
general transcription factor IIH subunit 1 |
| chr1_-_198232344 | 0.00 |
ENSRNOT00000080988
|
Aldoa
|
aldolase, fructose-bisphosphate A |
| chr4_-_60358562 | 0.00 |
ENSRNOT00000018001
|
Chchd3
|
coiled-coil-helix-coiled-coil-helix domain containing 3 |
| chr3_+_113415774 | 0.00 |
ENSRNOT00000056151
|
Serf2
|
small EDRK-rich factor 2 |
| chr12_+_37594185 | 0.00 |
ENSRNOT00000088787
|
Sbno1
|
strawberry notch homolog 1 |
| chr1_-_219450451 | 0.00 |
ENSRNOT00000025317
|
Rad9a
|
RAD9 checkpoint clamp component A |
| chr20_-_5533600 | 0.00 |
ENSRNOT00000072319
|
Cuta
|
cutA divalent cation tolerance homolog |
| chr6_-_109205004 | 0.00 |
ENSRNOT00000010512
|
Tmed10
|
transmembrane p24 trafficking protein 10 |
| chr20_-_5533448 | 0.00 |
ENSRNOT00000000568
|
Cuta
|
cutA divalent cation tolerance homolog |
| chr17_+_85356042 | 0.00 |
ENSRNOT00000022201
|
Commd3
|
COMM domain containing 3 |
| chr10_+_38692211 | 0.00 |
ENSRNOT00000009440
|
Aff4
|
AF4/FMR2 family, member 4 |
| chr7_-_70498992 | 0.00 |
ENSRNOT00000067774
ENSRNOT00000079327 |
Pip4k2c
|
phosphatidylinositol-5-phosphate 4-kinase type 2 gamma |
| chr5_+_78222504 | 0.00 |
ENSRNOT00000019544
|
Slc31a1
|
solute carrier family 31 member 1 |
| chr2_+_207108552 | 0.00 |
ENSRNOT00000027234
|
Slc16a1
|
solute carrier family 16 member 1 |
| chr19_-_37525762 | 0.00 |
ENSRNOT00000023606
|
Atp6v0d1
|
ATPase H+ transporting V0 subunit D1 |
| chr8_-_48674748 | 0.00 |
ENSRNOT00000014127
|
Hmbs
|
hydroxymethylbilane synthase |
| chr9_-_16647458 | 0.00 |
ENSRNOT00000024380
|
Mrpl2
|
mitochondrial ribosomal protein L2 |
| chr20_+_6288267 | 0.00 |
ENSRNOT00000000627
|
Srsf3
|
serine and arginine rich splicing factor 3 |
| chr3_-_23510959 | 0.00 |
ENSRNOT00000021176
|
Ppp6c
|
protein phosphatase 6, catalytic subunit |
| chr16_-_10941414 | 0.00 |
ENSRNOT00000086627
ENSRNOT00000085414 ENSRNOT00000081631 ENSRNOT00000087521 ENSRNOT00000083623 |
Ldb3
|
LIM domain binding 3 |
| chr3_+_161298962 | 0.00 |
ENSRNOT00000066028
|
Ctsa
|
cathepsin A |
| chr7_-_12741296 | 0.00 |
ENSRNOT00000060648
|
Arhgap45
|
Rho GTPase activating protein 45 |
| chr13_+_26172243 | 0.00 |
ENSRNOT00000003840
|
Phlpp1
|
PH domain and leucine rich repeat protein phosphatase 1 |
| chr7_+_53878610 | 0.00 |
ENSRNOT00000091910
|
Osbpl8
|
oxysterol binding protein-like 8 |
| chr12_+_37593874 | 0.00 |
ENSRNOT00000057902
|
Sbno1
|
strawberry notch homolog 1 |
| chr10_+_84135116 | 0.00 |
ENSRNOT00000031035
|
Hoxb7
|
homeo box B7 |
| chr13_-_101697684 | 0.00 |
ENSRNOT00000078834
|
Brox
|
BRO1 domain and CAAX motif containing |