Project
PRJNA438478: RNAseq of wild type zebrafish germline
Navigation
Downloads
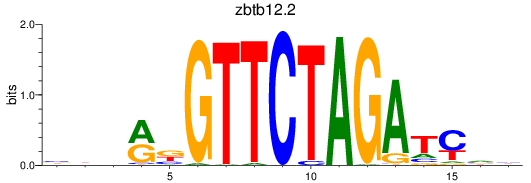
Results for zbtb12.2
Z-value: 0.93
Transcription factors associated with zbtb12.2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
zbtb12.2
|
ENSDARG00000070658 | zinc finger and BTB domain containing 12, tandem duplicate 2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| zbtb12.2 | dr11_v1_chr19_-_26964348_26964348 | -0.14 | 5.9e-01 | Click! |
Activity profile of zbtb12.2 motif
Sorted Z-values of zbtb12.2 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr2_-_17114852 | 3.82 |
ENSDART00000006549
|
pif1
|
PIF1 5'-to-3' DNA helicase homolog (S. cerevisiae) |
| chr2_-_17115256 | 3.59 |
ENSDART00000190488
|
pif1
|
PIF1 5'-to-3' DNA helicase homolog (S. cerevisiae) |
| chr19_-_27579805 | 2.97 |
ENSDART00000132980
ENSDART00000142263 |
si:dkeyp-46h3.3
|
si:dkeyp-46h3.3 |
| chr5_+_25311309 | 1.81 |
ENSDART00000169638
|
wu:fa19b12
|
wu:fa19b12 |
| chr15_+_38299563 | 1.73 |
ENSDART00000099375
|
si:dkey-24p1.6
|
si:dkey-24p1.6 |
| chr14_-_6527010 | 1.73 |
ENSDART00000125058
|
nipsnap3a
|
nipsnap homolog 3A (C. elegans) |
| chr16_-_42175617 | 1.52 |
ENSDART00000084715
|
alkbh8
|
alkB homolog 8, tRNA methyltransferase |
| chr20_-_29532939 | 1.50 |
ENSDART00000049224
ENSDART00000062377 |
taf1b
|
TATA box binding protein (Tbp)-associated factor, RNA polymerase I, B |
| chr13_+_46941930 | 1.43 |
ENSDART00000056962
|
fbxo5
|
F-box protein 5 |
| chr19_+_26640096 | 1.29 |
ENSDART00000067793
|
ints3
|
integrator complex subunit 3 |
| chr3_+_43373867 | 1.25 |
ENSDART00000159455
ENSDART00000172425 |
zfand2a
|
zinc finger, AN1-type domain 2A |
| chr17_-_12249990 | 1.21 |
ENSDART00000177889
ENSDART00000155545 |
ahctf1
|
AT hook containing transcription factor 1 |
| chr16_+_53526135 | 1.15 |
ENSDART00000083558
|
smpd5
|
sphingomyelin phosphodiesterase 5 |
| chr19_-_24136233 | 1.15 |
ENSDART00000143365
|
thap7
|
THAP domain containing 7 |
| chr24_-_7957581 | 1.06 |
ENSDART00000145815
|
txndc5
|
thioredoxin domain containing 5 |
| chr24_-_28259127 | 1.04 |
ENSDART00000149589
|
prkag2a
|
protein kinase, AMP-activated, gamma 2 non-catalytic subunit a |
| chr20_+_38458084 | 1.02 |
ENSDART00000020153
ENSDART00000135912 |
coq8a
|
coenzyme Q8A |
| chr9_-_180334 | 0.99 |
ENSDART00000180339
|
zgc:158619
|
zgc:158619 |
| chr3_+_43374571 | 0.96 |
ENSDART00000182497
|
zfand2a
|
zinc finger, AN1-type domain 2A |
| chr5_-_66823750 | 0.91 |
ENSDART00000041441
ENSDART00000112488 |
stip1
|
stress-induced phosphoprotein 1 |
| chr21_+_1587722 | 0.86 |
ENSDART00000013581
|
wdr91
|
WD repeat domain 91 |
| chr6_+_11397269 | 0.86 |
ENSDART00000114260
|
senp2
|
SUMO1/sentrin/SMT3 specific peptidase 2 |
| chr6_+_12482599 | 0.83 |
ENSDART00000090316
|
stk24b
|
serine/threonine kinase 24b (STE20 homolog, yeast) |
| chr19_+_3140313 | 0.82 |
ENSDART00000125504
|
zgc:86598
|
zgc:86598 |
| chr1_+_58981985 | 0.81 |
ENSDART00000170646
ENSDART00000171055 |
thumpd1
|
THUMP domain containing 1 |
| chr22_+_2048566 | 0.78 |
ENSDART00000169234
|
si:dkey-1b17.9
|
si:dkey-1b17.9 |
| chr6_+_20954400 | 0.76 |
ENSDART00000143248
ENSDART00000165806 |
stk11ip
|
serine/threonine kinase 11 interacting protein |
| chr25_-_7649736 | 0.75 |
ENSDART00000149548
|
myo5ab
|
myosin VAb |
| chr12_-_48312647 | 0.73 |
ENSDART00000114415
|
ascc1
|
activating signal cointegrator 1 complex subunit 1 |
| chr2_+_30547018 | 0.71 |
ENSDART00000193747
|
ankrd33bb
|
ankyrin repeat domain 33Bb |
| chr22_-_9861531 | 0.71 |
ENSDART00000193197
|
si:dkey-253d23.2
|
si:dkey-253d23.2 |
| chr21_+_20386865 | 0.70 |
ENSDART00000144366
|
si:dkey-30k6.5
|
si:dkey-30k6.5 |
| chr14_+_46342882 | 0.69 |
ENSDART00000193707
ENSDART00000060577 |
tmem33
|
transmembrane protein 33 |
| chr1_+_218524 | 0.69 |
ENSDART00000109529
|
tmco3
|
transmembrane and coiled-coil domains 3 |
| chr7_+_21768452 | 0.68 |
ENSDART00000127719
|
kdm6ba
|
lysine (K)-specific demethylase 6B, a |
| chr3_+_33300522 | 0.67 |
ENSDART00000114023
|
hspb9
|
heat shock protein, alpha-crystallin-related, 9 |
| chr10_+_29849497 | 0.63 |
ENSDART00000099994
ENSDART00000132212 |
hspa8
|
heat shock protein 8 |
| chr1_-_40341306 | 0.62 |
ENSDART00000190649
|
maml3
|
mastermind-like transcriptional coactivator 3 |
| chr21_-_28737320 | 0.61 |
ENSDART00000098696
ENSDART00000124826 ENSDART00000125652 |
nrg2a
|
neuregulin 2a |
| chr6_-_46053300 | 0.59 |
ENSDART00000169722
|
ca16b
|
carbonic anhydrase XVI b |
| chr18_+_8402129 | 0.57 |
ENSDART00000081154
ENSDART00000171974 |
prpf18
|
PRP18 pre-mRNA processing factor 18 homolog (yeast) |
| chr3_-_36364903 | 0.57 |
ENSDART00000028883
|
gna13b
|
guanine nucleotide binding protein (G protein), alpha 13b |
| chr22_+_2512154 | 0.54 |
ENSDART00000097363
|
zgc:173726
|
zgc:173726 |
| chr12_+_9849476 | 0.53 |
ENSDART00000110458
|
fam117ab
|
family with sequence similarity 117, member Ab |
| chr11_+_1551603 | 0.51 |
ENSDART00000185383
ENSDART00000121489 ENSDART00000040577 |
mybl2b
|
v-myb avian myeloblastosis viral oncogene homolog-like 2b |
| chr14_-_14607855 | 0.49 |
ENSDART00000162322
|
rab9b
|
RAB9B, member RAS oncogene family |
| chr5_-_42950963 | 0.47 |
ENSDART00000149868
|
grsf1
|
G-rich RNA sequence binding factor 1 |
| chr19_-_42945965 | 0.46 |
ENSDART00000142858
|
dclk3
|
doublecortin-like kinase 3 |
| chr6_+_2195625 | 0.45 |
ENSDART00000155659
|
acvr1bb
|
activin A receptor type 1Bb |
| chr11_+_2855430 | 0.43 |
ENSDART00000172837
|
kif21b
|
kinesin family member 21B |
| chr2_+_22602301 | 0.41 |
ENSDART00000038514
|
sept2
|
septin 2 |
| chr21_+_43171013 | 0.38 |
ENSDART00000151362
|
zcchc10
|
zinc finger, CCHC domain containing 10 |
| chr19_+_20793388 | 0.36 |
ENSDART00000142463
|
txnl4a
|
thioredoxin-like 4A |
| chr19_+_20793237 | 0.35 |
ENSDART00000014774
|
txnl4a
|
thioredoxin-like 4A |
| chr22_+_2686673 | 0.34 |
ENSDART00000161580
|
si:dkey-20i20.15
|
si:dkey-20i20.15 |
| chr19_+_3215466 | 0.31 |
ENSDART00000181288
|
zgc:86598
|
zgc:86598 |
| chr10_-_15048781 | 0.31 |
ENSDART00000038401
ENSDART00000155674 |
si:ch211-95j8.2
|
si:ch211-95j8.2 |
| chr19_-_25519612 | 0.28 |
ENSDART00000133150
|
C1GALT1 (1 of many)
|
si:dkey-202e17.1 |
| chr21_-_43171249 | 0.27 |
ENSDART00000021037
ENSDART00000148630 |
hspa4a
|
heat shock protein 4a |
| chr1_+_54194801 | 0.27 |
ENSDART00000186802
|
CABZ01067150.1
|
|
| chr22_+_1837448 | 0.26 |
ENSDART00000160015
|
znf1183
|
zinc finger protein 1183 |
| chr11_-_18254 | 0.25 |
ENSDART00000167814
|
prr13
|
proline rich 13 |
| chr22_+_1734981 | 0.25 |
ENSDART00000158195
|
znf1159
|
zinc finger protein 1159 |
| chr6_-_1566186 | 0.25 |
ENSDART00000156305
|
trim107
|
tripartite motif containing 107 |
| chr8_+_24281512 | 0.24 |
ENSDART00000062845
|
mmp9
|
matrix metallopeptidase 9 |
| chr22_+_1300587 | 0.22 |
ENSDART00000124161
|
si:ch73-138e16.5
|
si:ch73-138e16.5 |
| chr8_+_22491947 | 0.21 |
ENSDART00000125805
|
si:ch211-261n11.8
|
si:ch211-261n11.8 |
| chr4_+_16323970 | 0.21 |
ENSDART00000190651
|
BX322608.1
|
|
| chr11_-_18449 | 0.21 |
ENSDART00000172050
|
prr13
|
proline rich 13 |
| chr12_-_20120702 | 0.20 |
ENSDART00000153387
ENSDART00000158412 ENSDART00000112768 |
ubald1a
|
UBA-like domain containing 1a |
| chr1_+_16127825 | 0.19 |
ENSDART00000122503
|
tusc3
|
tumor suppressor candidate 3 |
| chr2_-_20981907 | 0.18 |
ENSDART00000113384
|
lyrm4
|
LYR motif containing 4 |
| chr19_+_8985230 | 0.17 |
ENSDART00000018973
|
scamp3
|
secretory carrier membrane protein 3 |
| chr6_-_1566407 | 0.17 |
ENSDART00000112118
|
trim107
|
tripartite motif containing 107 |
| chr8_-_50132860 | 0.16 |
ENSDART00000149964
|
antxr1a
|
anthrax toxin receptor 1a |
| chr3_-_2033105 | 0.16 |
ENSDART00000135249
|
si:dkey-88j15.3
|
si:dkey-88j15.3 |
| chr8_-_50133064 | 0.15 |
ENSDART00000030064
|
antxr1a
|
anthrax toxin receptor 1a |
| chr22_+_9862466 | 0.15 |
ENSDART00000146864
|
si:dkey-253d23.3
|
si:dkey-253d23.3 |
| chr8_+_43852743 | 0.14 |
ENSDART00000186485
|
AL808129.2
|
|
| chr23_+_3553516 | 0.13 |
ENSDART00000181050
|
si:dkey-9l20.3
|
si:dkey-9l20.3 |
| chr16_+_38659475 | 0.12 |
ENSDART00000023238
|
emc2
|
ER membrane protein complex subunit 2 |
| chr1_-_45888608 | 0.12 |
ENSDART00000139219
|
pnpla6
|
patatin-like phospholipase domain containing 6 |
| chr19_-_25519310 | 0.12 |
ENSDART00000089882
|
C1GALT1 (1 of many)
|
si:dkey-202e17.1 |
| chr13_+_30055171 | 0.11 |
ENSDART00000143581
ENSDART00000132027 |
spock2
|
sparc/osteonectin, cwcv and kazal-like domains proteoglycan (testican) 2 |
| chr5_+_31944270 | 0.10 |
ENSDART00000147850
|
ungb
|
uracil DNA glycosylase b |
| chr19_+_7864767 | 0.10 |
ENSDART00000137540
ENSDART00000151642 |
si:dkeyp-85e10.3
|
si:dkeyp-85e10.3 |
| chr12_-_26430507 | 0.09 |
ENSDART00000153214
|
synpo2lb
|
synaptopodin 2-like b |
| chr23_+_34005792 | 0.08 |
ENSDART00000132668
|
si:ch211-207e14.4
|
si:ch211-207e14.4 |
| chr22_+_9287929 | 0.07 |
ENSDART00000193522
|
si:ch211-250k18.7
|
si:ch211-250k18.7 |
| chr9_-_22099536 | 0.07 |
ENSDART00000101923
|
CR391987.1
|
|
| chr10_+_9595575 | 0.07 |
ENSDART00000091780
ENSDART00000184287 |
rc3h2
|
ring finger and CCCH-type domains 2 |
| chr3_-_61345723 | 0.07 |
ENSDART00000156749
|
si:dkey-111k8.5
|
si:dkey-111k8.5 |
| chr9_-_10357638 | 0.07 |
ENSDART00000132123
|
thsd7ba
|
thrombospondin, type I, domain containing 7Ba |
| chr22_+_2144278 | 0.05 |
ENSDART00000162173
ENSDART00000159914 ENSDART00000160192 |
znf1164
|
zinc finger protein 1164 |
| chr7_-_58867188 | 0.05 |
ENSDART00000187006
|
CU681855.1
|
|
| chr18_+_28106139 | 0.04 |
ENSDART00000089615
|
kiaa1549lb
|
KIAA1549-like b |
| chr7_+_29024282 | 0.04 |
ENSDART00000076425
|
cog4
|
component of oligomeric golgi complex 4 |
| chr16_-_10837245 | 0.04 |
ENSDART00000036891
|
rabac1
|
Rab acceptor 1 (prenylated) |
| chr16_-_44709832 | 0.04 |
ENSDART00000156784
|
si:ch211-151m7.6
|
si:ch211-151m7.6 |
| chr5_-_29570141 | 0.03 |
ENSDART00000043259
|
entpd2a.2
|
ectonucleoside triphosphate diphosphohydrolase 2a, tandem duplicate 2 |
| chr22_+_18469004 | 0.03 |
ENSDART00000061430
|
cilp2
|
cartilage intermediate layer protein 2 |
| chr13_-_50446872 | 0.02 |
ENSDART00000083883
|
herc4
|
HECT and RLD domain containing E3 ubiquitin protein ligase 4 |
| chr3_-_7129057 | 0.01 |
ENSDART00000125947
|
BX005085.1
|
|
| chr3_-_136043 | 0.01 |
ENSDART00000160266
|
zgc:110249
|
zgc:110249 |
| chr15_+_31426768 | 0.01 |
ENSDART00000014270
|
or103-1
|
odorant receptor, family C, subfamily 103, member 1 |
Network of associatons between targets according to the STRING database.
First level regulatory network of zbtb12.2
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 7.4 | GO:0000002 | mitochondrial genome maintenance(GO:0000002) |
| 0.4 | 1.4 | GO:1904667 | negative regulation of ligase activity(GO:0051352) negative regulation of ubiquitin-protein transferase activity(GO:0051444) negative regulation of ubiquitin protein ligase activity(GO:1904667) |
| 0.3 | 1.5 | GO:0042790 | transcription of nuclear large rRNA transcript from RNA polymerase I promoter(GO:0042790) |
| 0.2 | 1.2 | GO:0051292 | nuclear pore complex assembly(GO:0051292) |
| 0.2 | 0.7 | GO:0061383 | trabecula morphogenesis(GO:0061383) heart trabecula morphogenesis(GO:0061384) |
| 0.2 | 0.6 | GO:0000350 | generation of catalytic spliceosome for second transesterification step(GO:0000350) |
| 0.1 | 0.6 | GO:0031584 | activation of phospholipase D activity(GO:0031584) |
| 0.1 | 0.6 | GO:0007221 | positive regulation of transcription of Notch receptor target(GO:0007221) |
| 0.1 | 1.2 | GO:0006685 | sphingomyelin catabolic process(GO:0006685) |
| 0.1 | 0.6 | GO:1902946 | positive regulation of receptor-mediated endocytosis(GO:0048260) protein localization to early endosome(GO:1902946) |
| 0.1 | 0.6 | GO:0060347 | ventricular trabecula myocardium morphogenesis(GO:0003222) trabecula formation(GO:0060343) heart trabecula formation(GO:0060347) skin epidermis development(GO:0098773) |
| 0.1 | 0.7 | GO:1901339 | regulation of store-operated calcium channel activity(GO:1901339) |
| 0.1 | 0.2 | GO:0002544 | granuloma formation(GO:0002432) chronic inflammatory response(GO:0002544) regulation of granuloma formation(GO:0002631) regulation of chronic inflammatory response(GO:0002676) |
| 0.1 | 1.3 | GO:0044818 | mitotic G2/M transition checkpoint(GO:0044818) |
| 0.1 | 0.9 | GO:0043551 | regulation of lipid kinase activity(GO:0043550) regulation of phosphatidylinositol 3-kinase activity(GO:0043551) |
| 0.1 | 1.5 | GO:0002098 | tRNA wobble uridine modification(GO:0002098) |
| 0.1 | 0.9 | GO:0016926 | protein desumoylation(GO:0016926) |
| 0.0 | 0.7 | GO:0061055 | myotome development(GO:0061055) |
| 0.0 | 0.3 | GO:1901998 | toxin transport(GO:1901998) |
| 0.0 | 1.0 | GO:0009395 | phospholipid catabolic process(GO:0009395) |
| 0.0 | 1.0 | GO:0042149 | cellular response to glucose starvation(GO:0042149) |
| 0.0 | 0.1 | GO:0097510 | base-excision repair, AP site formation via deaminated base removal(GO:0097510) |
| 0.0 | 0.2 | GO:2000406 | positive regulation of lymphocyte migration(GO:2000403) positive regulation of T cell migration(GO:2000406) |
| 0.0 | 0.9 | GO:0043277 | apoptotic cell clearance(GO:0043277) |
| 0.0 | 1.1 | GO:0018107 | peptidyl-threonine phosphorylation(GO:0018107) |
| 0.0 | 0.1 | GO:0045050 | protein insertion into ER membrane by stop-transfer membrane-anchor sequence(GO:0045050) |
| 0.0 | 0.5 | GO:0032482 | Rab protein signal transduction(GO:0032482) |
| 0.0 | 0.8 | GO:0099515 | vesicle transport along actin filament(GO:0030050) actin filament-based transport(GO:0099515) |
| 0.0 | 0.8 | GO:0006400 | tRNA modification(GO:0006400) |
| 0.0 | 0.2 | GO:0015693 | magnesium ion transport(GO:0015693) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.5 | GO:0000120 | RNA polymerase I transcription factor complex(GO:0000120) |
| 0.2 | 1.3 | GO:0070876 | SOSS complex(GO:0070876) |
| 0.1 | 0.9 | GO:0031313 | extrinsic component of endosome membrane(GO:0031313) |
| 0.1 | 7.4 | GO:0005657 | replication fork(GO:0005657) |
| 0.1 | 1.1 | GO:0005956 | protein kinase CK2 complex(GO:0005956) |
| 0.1 | 1.0 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.1 | 1.3 | GO:0005682 | U5 snRNP(GO:0005682) |
| 0.1 | 0.6 | GO:0031526 | brush border membrane(GO:0031526) |
| 0.1 | 0.5 | GO:0030670 | phagocytic vesicle membrane(GO:0030670) |
| 0.0 | 0.7 | GO:0044666 | MLL3/4 complex(GO:0044666) |
| 0.0 | 0.4 | GO:0048179 | activin receptor complex(GO:0048179) |
| 0.0 | 1.2 | GO:0005758 | mitochondrial intermembrane space(GO:0005758) organelle envelope lumen(GO:0031970) |
| 0.0 | 1.1 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.0 | 0.4 | GO:0060170 | ciliary membrane(GO:0060170) |
| 0.0 | 0.2 | GO:0008250 | oligosaccharyltransferase complex(GO:0008250) |
| 0.0 | 0.1 | GO:0072546 | ER membrane protein complex(GO:0072546) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.5 | 7.4 | GO:0043139 | 5'-3' DNA helicase activity(GO:0043139) |
| 0.5 | 1.5 | GO:0016300 | tRNA (uracil) methyltransferase activity(GO:0016300) |
| 0.5 | 1.5 | GO:0001164 | RNA polymerase I regulatory region DNA binding(GO:0001013) RNA polymerase I regulatory region sequence-specific DNA binding(GO:0001163) RNA polymerase I CORE element sequence-specific DNA binding(GO:0001164) |
| 0.2 | 0.6 | GO:0031752 | D5 dopamine receptor binding(GO:0031752) |
| 0.1 | 1.2 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.1 | 1.1 | GO:0003756 | protein disulfide isomerase activity(GO:0003756) intramolecular oxidoreductase activity, transposing S-S bonds(GO:0016864) |
| 0.1 | 0.4 | GO:0016361 | activin receptor activity, type I(GO:0016361) |
| 0.1 | 0.7 | GO:0071558 | histone demethylase activity (H3-K27 specific)(GO:0071558) |
| 0.1 | 0.9 | GO:0051879 | Hsp90 protein binding(GO:0051879) |
| 0.1 | 0.9 | GO:0016929 | SUMO-specific protease activity(GO:0016929) |
| 0.0 | 0.6 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.0 | 1.0 | GO:0016208 | AMP binding(GO:0016208) |
| 0.0 | 0.9 | GO:0035014 | phosphatidylinositol 3-kinase regulator activity(GO:0035014) |
| 0.0 | 0.8 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.0 | 0.1 | GO:0032977 | membrane insertase activity(GO:0032977) |
| 0.0 | 3.0 | GO:0030246 | carbohydrate binding(GO:0030246) |
| 0.0 | 1.1 | GO:0004620 | phospholipase activity(GO:0004620) |
| 0.0 | 0.1 | GO:0035613 | RNA stem-loop binding(GO:0035613) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 7.4 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.0 | 1.4 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.0 | 0.9 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.0 | 0.8 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.0 | 0.2 | PID SYNDECAN 4 PATHWAY | Syndecan-4-mediated signaling events |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.5 | REACTOME RNA POL I TRANSCRIPTION TERMINATION | Genes involved in RNA Polymerase I Transcription Termination |
| 0.1 | 0.6 | REACTOME NOTCH HLH TRANSCRIPTION PATHWAY | Genes involved in Notch-HLH transcription pathway |
| 0.1 | 1.7 | REACTOME LYSOSOME VESICLE BIOGENESIS | Genes involved in Lysosome Vesicle Biogenesis |
| 0.1 | 1.4 | REACTOME G1 S SPECIFIC TRANSCRIPTION | Genes involved in G1/S-Specific Transcription |
| 0.0 | 0.2 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.0 | 0.7 | REACTOME MRNA SPLICING MINOR PATHWAY | Genes involved in mRNA Splicing - Minor Pathway |
| 0.0 | 1.2 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.0 | 0.9 | REACTOME L1CAM INTERACTIONS | Genes involved in L1CAM interactions |