Project
PRJNA438478: RNAseq of wild type zebrafish germline
Navigation
Downloads
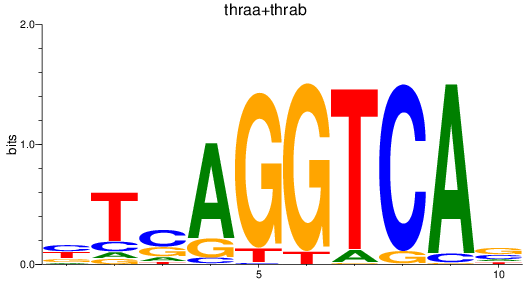
Results for thraa+thrab
Z-value: 0.90
Transcription factors associated with thraa+thrab
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
thraa
|
ENSDARG00000000151 | thyroid hormone receptor alpha a |
|
thrab
|
ENSDARG00000052654 | thyroid hormone receptor alpha b |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| thraa | dr11_v1_chr3_-_34724879_34724879 | -0.91 | 2.2e-07 | Click! |
| thrab | dr11_v1_chr12_-_22039350_22039350 | -0.82 | 3.6e-05 | Click! |
Activity profile of thraa+thrab motif
Sorted Z-values of thraa+thrab motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_+_57311901 | 2.90 |
ENSDART00000149397
|
mycbpap
|
mycbp associated protein |
| chr17_-_24838927 | 2.42 |
ENSDART00000123259
|
CR388164.1
|
|
| chr12_+_49125510 | 2.15 |
ENSDART00000185804
|
FO704607.1
|
|
| chr16_-_42969598 | 1.98 |
ENSDART00000156011
|
si:ch211-135n15.3
|
si:ch211-135n15.3 |
| chr4_+_18489207 | 1.98 |
ENSDART00000135276
|
si:dkey-202b22.5
|
si:dkey-202b22.5 |
| chr20_-_43462190 | 1.85 |
ENSDART00000100697
|
pimr133
|
Pim proto-oncogene, serine/threonine kinase, related 133 |
| chr25_+_4954220 | 1.80 |
ENSDART00000156034
|
si:ch73-265h17.2
|
si:ch73-265h17.2 |
| chr5_+_13353866 | 1.72 |
ENSDART00000099611
|
ccl19a.1
|
chemokine (C-C motif) ligand 19a, tandem duplicate 1 |
| chr6_+_475264 | 1.67 |
ENSDART00000193615
|
LO017974.1
|
|
| chr7_-_40738774 | 1.65 |
ENSDART00000084179
|
rnf32
|
ring finger protein 32 |
| chr20_+_42761881 | 1.64 |
ENSDART00000113625
|
pimr113
|
Pim proto-oncogene, serine/threonine kinase, related 113 |
| chr22_-_20761759 | 1.63 |
ENSDART00000013803
|
amh
|
anti-Mullerian hormone |
| chr13_-_36703164 | 1.57 |
ENSDART00000044357
|
cdkl1
|
cyclin-dependent kinase-like 1 (CDC2-related kinase) |
| chr20_+_39457598 | 1.56 |
ENSDART00000140931
ENSDART00000156176 |
pimr128
|
Pim proto-oncogene, serine/threonine kinase, related 128 |
| chr1_-_30039331 | 1.50 |
ENSDART00000086935
ENSDART00000143800 |
MARCH4 (1 of many)
|
zgc:153256 |
| chr20_+_54738210 | 1.45 |
ENSDART00000151399
|
pak7
|
p21 protein (Cdc42/Rac)-activated kinase 7 |
| chr16_-_41867833 | 1.44 |
ENSDART00000133066
|
zgc:194418
|
zgc:194418 |
| chr14_-_24397718 | 1.43 |
ENSDART00000110164
|
si:ch211-163m17.4
|
si:ch211-163m17.4 |
| chr10_-_44027391 | 1.39 |
ENSDART00000145404
|
crybb1
|
crystallin, beta B1 |
| chr20_+_42780162 | 1.37 |
ENSDART00000127069
|
pimr213
|
Pim proto-oncogene, serine/threonine kinase, related 213 |
| chr16_-_41840668 | 1.33 |
ENSDART00000146150
|
si:dkey-199f5.4
|
si:dkey-199f5.4 |
| chr11_-_6001845 | 1.32 |
ENSDART00000108628
|
ano8b
|
anoctamin 8b |
| chr20_-_43572763 | 1.25 |
ENSDART00000153251
|
pimr125
|
Pim proto-oncogene, serine/threonine kinase, related 125 |
| chr24_-_39610585 | 1.25 |
ENSDART00000066506
|
cox6b1
|
cytochrome c oxidase subunit VIb polypeptide 1 |
| chr14_+_39255437 | 1.23 |
ENSDART00000147443
|
diaph2
|
diaphanous-related formin 2 |
| chr13_+_8677166 | 1.22 |
ENSDART00000181016
ENSDART00000135028 |
prop1
|
PROP paired-like homeobox 1 |
| chr9_+_52613820 | 1.19 |
ENSDART00000168753
ENSDART00000165580 ENSDART00000164769 |
si:ch211-241j8.2
|
si:ch211-241j8.2 |
| chr2_+_6181383 | 1.18 |
ENSDART00000153307
|
si:ch73-344o19.1
|
si:ch73-344o19.1 |
| chr12_-_47782623 | 1.17 |
ENSDART00000115742
|
selenou1b
|
selenoprotein U1b |
| chr14_-_8724290 | 1.15 |
ENSDART00000161171
|
pimr56
|
Pim proto-oncogene, serine/threonine kinase, related 56 |
| chr19_+_3653976 | 1.12 |
ENSDART00000125673
|
nedd9
|
neural precursor cell expressed, developmentally down-regulated 9 |
| chr15_-_28095532 | 1.11 |
ENSDART00000191490
|
cryba1a
|
crystallin, beta A1a |
| chr12_+_18533198 | 1.10 |
ENSDART00000189729
|
meiob
|
meiosis specific with OB domains |
| chr23_+_42810055 | 1.09 |
ENSDART00000186647
|
myl9a
|
myosin, light chain 9a, regulatory |
| chr8_+_25902170 | 1.07 |
ENSDART00000193130
|
rhoab
|
ras homolog gene family, member Ab |
| chr12_-_3133483 | 1.06 |
ENSDART00000015092
|
col1a1b
|
collagen, type I, alpha 1b |
| chr11_-_31172276 | 1.05 |
ENSDART00000171520
|
si:dkey-238i5.3
|
si:dkey-238i5.3 |
| chr13_+_51869025 | 1.04 |
ENSDART00000187066
|
LT631684.1
|
|
| chr21_-_45073764 | 1.00 |
ENSDART00000181390
ENSDART00000063714 |
rapgef6
|
Rap guanine nucleotide exchange factor (GEF) 6 |
| chr1_+_54626491 | 0.96 |
ENSDART00000136063
|
si:ch211-202h22.9
|
si:ch211-202h22.9 |
| chr12_+_7491690 | 0.95 |
ENSDART00000152564
|
phyhiplb
|
phytanoyl-CoA 2-hydroxylase interacting protein-like b |
| chr10_+_26813710 | 0.94 |
ENSDART00000136122
|
si:ch211-263p13.7
|
si:ch211-263p13.7 |
| chr18_-_15932704 | 0.93 |
ENSDART00000127769
|
plekhg7
|
pleckstrin homology domain containing, family G (with RhoGef domain) member 7 |
| chr23_+_4373360 | 0.93 |
ENSDART00000144061
|
ptpdc1b
|
protein tyrosine phosphatase domain containing 1b |
| chr5_+_7989210 | 0.92 |
ENSDART00000168071
|
gdnfb
|
glial cell derived neurotrophic factor b |
| chr2_+_49864219 | 0.92 |
ENSDART00000187744
|
rpl37
|
ribosomal protein L37 |
| chr5_-_26118855 | 0.92 |
ENSDART00000009028
|
ela3l
|
elastase 3 like |
| chr4_+_17280868 | 0.91 |
ENSDART00000145349
|
bcat1
|
branched chain amino-acid transaminase 1, cytosolic |
| chr13_-_31435137 | 0.89 |
ENSDART00000057441
|
rtn1a
|
reticulon 1a |
| chr16_-_45327616 | 0.89 |
ENSDART00000158733
|
si:dkey-33i11.1
|
si:dkey-33i11.1 |
| chr11_-_11471857 | 0.87 |
ENSDART00000030103
|
krt94
|
keratin 94 |
| chr6_+_30689239 | 0.86 |
ENSDART00000065206
|
wdr78
|
WD repeat domain 78 |
| chr2_+_24786765 | 0.85 |
ENSDART00000141030
|
pde4ca
|
phosphodiesterase 4C, cAMP-specific a |
| chr2_+_33368414 | 0.84 |
ENSDART00000077462
|
slc6a9
|
solute carrier family 6 (neurotransmitter transporter, glycine), member 9 |
| chr23_+_44634187 | 0.84 |
ENSDART00000143688
|
si:ch73-265d7.2
|
si:ch73-265d7.2 |
| chr1_-_59176949 | 0.82 |
ENSDART00000128742
|
CABZ01118678.1
|
|
| chr7_+_26716321 | 0.81 |
ENSDART00000189750
|
cd82a
|
CD82 molecule a |
| chr8_-_6918721 | 0.80 |
ENSDART00000014915
|
asb6
|
ankyrin repeat and SOCS box containing 6 |
| chr4_-_9609634 | 0.79 |
ENSDART00000067188
|
dclre1c
|
DNA cross-link repair 1C, PSO2 homolog (S. cerevisiae) |
| chr8_-_6918290 | 0.77 |
ENSDART00000138259
ENSDART00000142496 |
asb6
|
ankyrin repeat and SOCS box containing 6 |
| chr6_+_41463786 | 0.76 |
ENSDART00000065006
|
twf2a
|
twinfilin actin-binding protein 2a |
| chr8_-_48675411 | 0.75 |
ENSDART00000165081
|
pimr183
|
Pim proto-oncogene, serine/threonine kinase, related 183 |
| chr5_-_32882162 | 0.74 |
ENSDART00000085769
|
lrsam1
|
leucine rich repeat and sterile alpha motif containing 1 |
| chr12_+_18532866 | 0.74 |
ENSDART00000152126
ENSDART00000152443 ENSDART00000089589 |
meiob
|
meiosis specific with OB domains |
| chr4_+_38344 | 0.73 |
ENSDART00000170197
ENSDART00000175348 |
phtf2
|
putative homeodomain transcription factor 2 |
| chr3_-_38915998 | 0.70 |
ENSDART00000141886
|
si:dkey-106c17.2
|
si:dkey-106c17.2 |
| chr12_-_30841679 | 0.69 |
ENSDART00000105594
|
crygmx
|
crystallin, gamma MX |
| chr10_+_22782522 | 0.69 |
ENSDART00000079498
ENSDART00000145558 |
si:ch211-237l4.6
|
si:ch211-237l4.6 |
| chr10_+_8629275 | 0.68 |
ENSDART00000129643
|
aplnrb
|
apelin receptor b |
| chr21_-_8420830 | 0.67 |
ENSDART00000138040
|
lhx2a
|
LIM homeobox 2a |
| chr2_+_36015049 | 0.66 |
ENSDART00000158276
|
lamc2
|
laminin, gamma 2 |
| chr20_-_40717900 | 0.64 |
ENSDART00000181663
|
cx43
|
connexin 43 |
| chr11_+_6010177 | 0.64 |
ENSDART00000170047
ENSDART00000022526 ENSDART00000161001 ENSDART00000188999 |
gtpbp3
|
GTP binding protein 3, mitochondrial |
| chr16_+_38338721 | 0.63 |
ENSDART00000076528
ENSDART00000142885 |
gabpb2b
|
GA binding protein transcription factor, beta subunit 2b |
| chr4_-_16853464 | 0.61 |
ENSDART00000125743
ENSDART00000164570 |
slc25a3a
|
solute carrier family 25 (mitochondrial carrier; phosphate carrier), member 3a |
| chr7_-_37812176 | 0.60 |
ENSDART00000164485
|
adcy7
|
adenylate cyclase 7 |
| chr7_-_41627956 | 0.58 |
ENSDART00000083929
|
plxdc2
|
plexin domain containing 2 |
| chr3_-_18575868 | 0.58 |
ENSDART00000122968
|
aqp8b
|
aquaporin 8b |
| chr9_-_202805 | 0.57 |
ENSDART00000182260
|
CABZ01067557.1
|
|
| chr18_+_3530769 | 0.57 |
ENSDART00000163469
|
si:ch73-338a16.3
|
si:ch73-338a16.3 |
| chr22_-_38258053 | 0.56 |
ENSDART00000132516
|
elavl2
|
ELAV like neuron-specific RNA binding protein 2 |
| chr6_+_1787160 | 0.55 |
ENSDART00000113505
|
myl9b
|
myosin, light chain 9b, regulatory |
| chr4_-_16412084 | 0.54 |
ENSDART00000188460
|
dcn
|
decorin |
| chr7_-_11638126 | 0.54 |
ENSDART00000125827
|
si:dkey-15b23.3
|
si:dkey-15b23.3 |
| chr5_-_67661102 | 0.54 |
ENSDART00000013605
|
zbtb20
|
zinc finger and BTB domain containing 20 |
| chr2_-_43123949 | 0.52 |
ENSDART00000075347
ENSDART00000132346 |
lrrc6
|
leucine rich repeat containing 6 |
| chr17_+_6217704 | 0.51 |
ENSDART00000129100
|
BX000363.1
|
|
| chr8_+_53269657 | 0.50 |
ENSDART00000184212
|
gnb1a
|
guanine nucleotide binding protein (G protein), beta polypeptide 1a |
| chr2_-_53481912 | 0.49 |
ENSDART00000189610
|
hsd11b1lb
|
hydroxysteroid (11-beta) dehydrogenase 1-like b |
| chr3_-_29977495 | 0.49 |
ENSDART00000077111
|
hsd17b14
|
hydroxysteroid (17-beta) dehydrogenase 14 |
| chr4_-_1360495 | 0.49 |
ENSDART00000164623
|
ptn
|
pleiotrophin |
| chr18_-_26675699 | 0.48 |
ENSDART00000113280
|
FRMD5
|
si:ch211-69m14.1 |
| chr7_-_26308098 | 0.47 |
ENSDART00000146440
ENSDART00000146935 ENSDART00000164627 |
zgc:77439
|
zgc:77439 |
| chr17_-_15149192 | 0.47 |
ENSDART00000180511
ENSDART00000103405 |
gch1
|
GTP cyclohydrolase 1 |
| chr19_+_4912817 | 0.47 |
ENSDART00000101658
ENSDART00000165082 |
ppp1r1b
|
protein phosphatase 1, regulatory (inhibitor) subunit 1B |
| chr16_+_33655890 | 0.46 |
ENSDART00000143757
|
fhl3a
|
four and a half LIM domains 3a |
| chr20_-_7992496 | 0.46 |
ENSDART00000145521
|
plpp3
|
phospholipid phosphatase 3 |
| chr21_-_41870029 | 0.45 |
ENSDART00000182035
|
endou2
|
endonuclease, polyU-specific 2 |
| chr23_+_44633858 | 0.45 |
ENSDART00000180728
|
si:ch73-265d7.2
|
si:ch73-265d7.2 |
| chr3_-_37681824 | 0.45 |
ENSDART00000185858
|
gpatch8
|
G patch domain containing 8 |
| chr23_+_30048849 | 0.44 |
ENSDART00000126027
|
uts2a
|
urotensin 2, alpha |
| chr12_-_37653685 | 0.44 |
ENSDART00000085121
|
sdk2b
|
sidekick cell adhesion molecule 2b |
| chr11_+_12897004 | 0.44 |
ENSDART00000152007
|
si:dkey-11m19.5
|
si:dkey-11m19.5 |
| chr22_-_15578402 | 0.43 |
ENSDART00000062986
|
hsh2d
|
hematopoietic SH2 domain containing |
| chr15_+_24644016 | 0.43 |
ENSDART00000043292
|
smtnl
|
smoothelin, like |
| chr16_-_13612650 | 0.43 |
ENSDART00000080372
|
dbpb
|
D site albumin promoter binding protein b |
| chr11_-_22997506 | 0.43 |
ENSDART00000167817
|
atp2b2
|
ATPase plasma membrane Ca2+ transporting 2 |
| chr3_+_12484008 | 0.43 |
ENSDART00000182229
|
vasnb
|
vasorin b |
| chr8_+_20776654 | 0.43 |
ENSDART00000135850
|
nfic
|
nuclear factor I/C |
| chr6_-_49873020 | 0.42 |
ENSDART00000148511
|
gnas
|
GNAS complex locus |
| chr11_+_45421761 | 0.42 |
ENSDART00000167347
|
hrasls
|
HRAS-like suppressor |
| chr19_-_7043355 | 0.42 |
ENSDART00000104845
|
tapbp.1
|
TAP binding protein (tapasin), tandem duplicate 1 |
| chr7_+_29133321 | 0.41 |
ENSDART00000052346
|
gnao1b
|
guanine nucleotide binding protein (G protein), alpha activating activity polypeptide O, b |
| chr23_-_42810664 | 0.41 |
ENSDART00000102328
|
pfkfb2a
|
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 2a |
| chr13_+_18663600 | 0.41 |
ENSDART00000141647
|
selenou1a
|
selenoprotein U 1a |
| chr14_+_28438947 | 0.40 |
ENSDART00000006489
|
acsl4a
|
acyl-CoA synthetase long chain family member 4a |
| chr3_+_13440900 | 0.39 |
ENSDART00000143715
|
GNG14
|
si:dkey-117i10.1 |
| chr1_-_46343999 | 0.39 |
ENSDART00000145117
ENSDART00000193233 |
atp11a
|
ATPase phospholipid transporting 11A |
| chr5_-_69425224 | 0.39 |
ENSDART00000158096
|
FQ311924.1
|
|
| chr25_-_36492779 | 0.39 |
ENSDART00000042271
|
irx3b
|
iroquois homeobox 3b |
| chr5_-_56412262 | 0.38 |
ENSDART00000083079
|
acaca
|
acetyl-CoA carboxylase alpha |
| chr14_-_46675850 | 0.37 |
ENSDART00000113285
|
tapt1a
|
transmembrane anterior posterior transformation 1a |
| chr1_-_58975098 | 0.37 |
ENSDART00000189899
|
CABZ01114581.1
|
|
| chr25_-_29134654 | 0.36 |
ENSDART00000067066
|
parp6b
|
poly (ADP-ribose) polymerase family, member 6b |
| chr2_-_127945 | 0.36 |
ENSDART00000056453
|
igfbp1b
|
insulin-like growth factor binding protein 1b |
| chr25_+_37443194 | 0.35 |
ENSDART00000163178
ENSDART00000190262 |
slc10a3
|
solute carrier family 10, member 3 |
| chr5_-_23473312 | 0.35 |
ENSDART00000140310
|
si:dkeyp-20g2.3
|
si:dkeyp-20g2.3 |
| chr25_-_15214161 | 0.34 |
ENSDART00000031499
|
wt1a
|
wilms tumor 1a |
| chr23_+_41912151 | 0.34 |
ENSDART00000191115
|
podn
|
podocan |
| chr5_-_51747278 | 0.34 |
ENSDART00000192232
|
lhfpl2b
|
LHFPL tetraspan subfamily member 2b |
| chr17_+_24613255 | 0.34 |
ENSDART00000064738
|
atp5if1b
|
ATP synthase inhibitory factor subunit 1b |
| chr15_+_28096152 | 0.34 |
ENSDART00000100293
ENSDART00000140092 |
crybb1l3
|
crystallin, beta B1, like 3 |
| chr3_-_36612877 | 0.33 |
ENSDART00000167164
|
si:dkeyp-72e1.7
|
si:dkeyp-72e1.7 |
| chr7_-_30082931 | 0.33 |
ENSDART00000075600
|
tspan3b
|
tetraspanin 3b |
| chr7_+_31871830 | 0.33 |
ENSDART00000139899
|
mybpc3
|
myosin binding protein C, cardiac |
| chr6_+_33885828 | 0.32 |
ENSDART00000179994
|
gpbp1l1
|
GC-rich promoter binding protein 1-like 1 |
| chr17_+_53424415 | 0.31 |
ENSDART00000157022
|
slc9a1b
|
solute carrier family 9 member A1b |
| chr8_+_28259347 | 0.31 |
ENSDART00000110857
|
fam212b
|
family with sequence similarity 212, member B |
| chr19_-_18855717 | 0.31 |
ENSDART00000158192
|
ppt2
|
palmitoyl-protein thioesterase 2 |
| chr23_+_24272421 | 0.30 |
ENSDART00000029974
|
clcnk
|
chloride channel K |
| chr9_-_9225980 | 0.30 |
ENSDART00000180301
|
cbsb
|
cystathionine-beta-synthase b |
| chr8_+_23711842 | 0.30 |
ENSDART00000128783
|
ppardb
|
peroxisome proliferator-activated receptor delta b |
| chr16_+_5774977 | 0.29 |
ENSDART00000134202
|
ccka
|
cholecystokinin a |
| chr9_-_374693 | 0.29 |
ENSDART00000166571
|
si:dkey-11f4.7
|
si:dkey-11f4.7 |
| chr12_-_42397002 | 0.28 |
ENSDART00000160497
|
glrx3
|
glutaredoxin 3 |
| chr11_+_6819050 | 0.27 |
ENSDART00000104289
|
rab3ab
|
RAB3A, member RAS oncogene family, b |
| chr10_+_44146287 | 0.26 |
ENSDART00000162176
|
grk3
|
G protein-coupled receptor kinase 3 |
| chr5_-_67629263 | 0.26 |
ENSDART00000133753
|
zbtb20
|
zinc finger and BTB domain containing 20 |
| chr6_-_60031693 | 0.26 |
ENSDART00000160275
|
CABZ01079262.1
|
|
| chr25_-_5740334 | 0.26 |
ENSDART00000169622
ENSDART00000168720 |
LO017739.1
|
|
| chr2_-_6182098 | 0.26 |
ENSDART00000156167
|
si:ch73-182a11.2
|
si:ch73-182a11.2 |
| chr3_+_1942219 | 0.26 |
ENSDART00000114520
|
zgc:165583
|
zgc:165583 |
| chr9_-_22892838 | 0.26 |
ENSDART00000143888
|
neb
|
nebulin |
| chr18_+_17959681 | 0.25 |
ENSDART00000142700
|
znf423
|
zinc finger protein 423 |
| chr4_-_77432218 | 0.25 |
ENSDART00000158683
|
slco1d1
|
solute carrier organic anion transporter family, member 1D1 |
| chr14_+_36738069 | 0.25 |
ENSDART00000105590
|
tdo2a
|
tryptophan 2,3-dioxygenase a |
| chr17_+_22587356 | 0.25 |
ENSDART00000157328
|
birc6
|
baculoviral IAP repeat containing 6 |
| chr10_-_39153959 | 0.24 |
ENSDART00000150193
ENSDART00000111362 |
slc37a4b
|
solute carrier family 37 (glucose-6-phosphate transporter), member 4b |
| chr16_-_46605160 | 0.24 |
ENSDART00000189070
ENSDART00000181194 |
tmem176l.3a
|
transmembrane protein 176l.3a |
| chr2_-_44199722 | 0.24 |
ENSDART00000140633
ENSDART00000145728 |
sdhc
|
succinate dehydrogenase complex, subunit C, integral membrane protein |
| chr3_+_12710350 | 0.24 |
ENSDART00000157959
|
cyp2k18
|
cytochrome P450, family 2, subfamily K, polypeptide 18 |
| chr23_-_20429741 | 0.23 |
ENSDART00000104334
|
taf13
|
TATA-box binding protein associated factor 13 |
| chr5_+_4564233 | 0.23 |
ENSDART00000193435
|
CABZ01058647.1
|
|
| chr11_-_25213651 | 0.23 |
ENSDART00000097316
ENSDART00000152186 |
myh7ba
|
myosin, heavy chain 7B, cardiac muscle, beta a |
| chr18_-_26785861 | 0.23 |
ENSDART00000098361
|
nmba
|
neuromedin Ba |
| chr8_+_7097740 | 0.22 |
ENSDART00000159670
|
abtb1
|
ankyrin repeat and BTB (POZ) domain containing 1 |
| chr11_+_25477643 | 0.22 |
ENSDART00000065941
|
opn1lw1
|
opsin 1 (cone pigments), long-wave-sensitive, 1 |
| chr13_-_31878043 | 0.22 |
ENSDART00000187720
|
syt14a
|
synaptotagmin XIVa |
| chr10_-_30785501 | 0.21 |
ENSDART00000020054
|
opcml
|
opioid binding protein/cell adhesion molecule-like |
| chr7_+_20017211 | 0.21 |
ENSDART00000100808
|
bcl6b
|
B-cell CLL/lymphoma 6, member B |
| chr2_-_28102264 | 0.21 |
ENSDART00000013638
|
cdh6
|
cadherin 6 |
| chr13_-_31878263 | 0.21 |
ENSDART00000180411
|
syt14a
|
synaptotagmin XIVa |
| chr19_-_24443867 | 0.21 |
ENSDART00000163763
ENSDART00000043133 |
thbs3b
|
thrombospondin 3b |
| chr1_+_2112726 | 0.21 |
ENSDART00000131714
ENSDART00000138396 |
mbnl2
|
muscleblind-like splicing regulator 2 |
| chr23_-_3703569 | 0.21 |
ENSDART00000143731
|
pacsin1a
|
protein kinase C and casein kinase substrate in neurons 1a |
| chr11_-_3574311 | 0.20 |
ENSDART00000167064
|
si:dkey-33m11.7
|
si:dkey-33m11.7 |
| chr1_-_49250490 | 0.20 |
ENSDART00000150386
|
si:ch73-6k14.2
|
si:ch73-6k14.2 |
| chr1_+_23783349 | 0.20 |
ENSDART00000007531
|
slit2
|
slit homolog 2 (Drosophila) |
| chr10_+_17592273 | 0.20 |
ENSDART00000141221
|
KCNN2
|
si:dkey-76p7.5 |
| chr10_+_5422575 | 0.20 |
ENSDART00000063093
|
auh
|
AU RNA binding protein/enoyl-CoA hydratase |
| chr17_-_51829310 | 0.20 |
ENSDART00000154544
|
numb
|
numb homolog (Drosophila) |
| chr20_-_25877793 | 0.20 |
ENSDART00000177333
ENSDART00000184619 |
exoc1
|
exocyst complex component 1 |
| chr14_-_34771371 | 0.20 |
ENSDART00000160598
ENSDART00000150413 ENSDART00000168910 |
ablim3
|
actin binding LIM protein family, member 3 |
| chr18_-_16924221 | 0.20 |
ENSDART00000122102
|
wee1
|
WEE1 G2 checkpoint kinase |
| chr9_+_38458193 | 0.20 |
ENSDART00000008053
|
mcm3ap
|
minichromosome maintenance complex component 3 associated protein |
| chr6_-_51556308 | 0.19 |
ENSDART00000149033
|
rbl1
|
retinoblastoma-like 1 (p107) |
| chr13_-_30645965 | 0.19 |
ENSDART00000109307
|
zcchc24
|
zinc finger, CCHC domain containing 24 |
| chr19_-_24443394 | 0.19 |
ENSDART00000090825
|
thbs3b
|
thrombospondin 3b |
| chr6_-_1566581 | 0.19 |
ENSDART00000192993
|
trim107
|
tripartite motif containing 107 |
| chr24_-_1356668 | 0.19 |
ENSDART00000188935
|
nrp1a
|
neuropilin 1a |
| chr10_-_42923385 | 0.19 |
ENSDART00000076731
|
ACOT12
|
acyl-CoA thioesterase 12 |
| chr21_+_22738939 | 0.19 |
ENSDART00000151342
ENSDART00000079145 |
arhgap42a
|
Rho GTPase activating protein 42a |
| chr7_-_50604626 | 0.19 |
ENSDART00000073903
ENSDART00000174031 |
crtc3
|
CREB regulated transcription coactivator 3 |
| chr3_+_17030665 | 0.18 |
ENSDART00000159849
ENSDART00000174491 ENSDART00000104519 ENSDART00000080854 |
stat3
|
signal transducer and activator of transcription 3 (acute-phase response factor) |
| chr5_-_30535327 | 0.18 |
ENSDART00000040328
|
h2afx
|
H2A histone family, member X |
| chr11_-_1400507 | 0.18 |
ENSDART00000173029
ENSDART00000172953 ENSDART00000111140 |
rpl29
|
ribosomal protein L29 |
| chr8_-_39754621 | 0.17 |
ENSDART00000149243
|
si:ch211-170d8.8
|
si:ch211-170d8.8 |
| chr24_-_7995748 | 0.17 |
ENSDART00000158566
|
bloc1s5
|
biogenesis of lysosomal organelles complex-1, subunit 5, muted |
| chr19_-_6988837 | 0.17 |
ENSDART00000145741
ENSDART00000167640 |
znf384l
|
zinc finger protein 384 like |
| chr4_+_75314247 | 0.17 |
ENSDART00000162365
|
CABZ01041913.1
|
|
Network of associatons between targets according to the STRING database.
First level regulatory network of thraa+thrab
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.6 | GO:0032369 | negative regulation of lipid transport(GO:0032369) |
| 0.3 | 0.9 | GO:0061217 | regulation of morphogenesis of a branching structure(GO:0060688) positive regulation of mesonephros development(GO:0061213) regulation of mesonephros development(GO:0061217) regulation of kidney development(GO:0090183) positive regulation of kidney development(GO:0090184) regulation of branching involved in ureteric bud morphogenesis(GO:0090189) positive regulation of branching involved in ureteric bud morphogenesis(GO:0090190) |
| 0.3 | 0.8 | GO:0060092 | regulation of synaptic transmission, glycinergic(GO:0060092) |
| 0.2 | 0.9 | GO:0009098 | leucine biosynthetic process(GO:0009098) |
| 0.2 | 1.1 | GO:0090527 | actin filament reorganization(GO:0090527) |
| 0.2 | 1.2 | GO:0000706 | meiotic DNA double-strand break processing(GO:0000706) |
| 0.2 | 0.7 | GO:1903589 | regulation of blood vessel endothelial cell proliferation involved in sprouting angiogenesis(GO:1903587) positive regulation of blood vessel endothelial cell proliferation involved in sprouting angiogenesis(GO:1903589) |
| 0.2 | 1.1 | GO:1902766 | skeletal muscle satellite cell migration(GO:1902766) |
| 0.1 | 0.4 | GO:0045724 | positive regulation of cilium assembly(GO:0045724) |
| 0.1 | 0.5 | GO:0046654 | tetrahydrofolate biosynthetic process(GO:0046654) |
| 0.1 | 0.3 | GO:0032780 | negative regulation of ATPase activity(GO:0032780) |
| 0.1 | 0.3 | GO:0031335 | regulation of sulfur amino acid metabolic process(GO:0031335) regulation of homocysteine metabolic process(GO:0050666) |
| 0.1 | 0.4 | GO:2001295 | malonyl-CoA biosynthetic process(GO:2001295) |
| 0.1 | 0.3 | GO:0031630 | regulation of synaptic vesicle fusion to presynaptic membrane(GO:0031630) |
| 0.1 | 1.8 | GO:0000712 | resolution of meiotic recombination intermediates(GO:0000712) |
| 0.1 | 0.3 | GO:0072149 | visceral serous pericardium development(GO:0061032) glomerular visceral epithelial cell fate commitment(GO:0072149) glomerular epithelial cell fate commitment(GO:0072314) |
| 0.1 | 0.4 | GO:0007191 | adenylate cyclase-activating dopamine receptor signaling pathway(GO:0007191) |
| 0.1 | 0.8 | GO:0031848 | protection from non-homologous end joining at telomere(GO:0031848) telomere maintenance in response to DNA damage(GO:0043247) |
| 0.1 | 0.2 | GO:0007414 | axonal defasciculation(GO:0007414) |
| 0.1 | 0.4 | GO:1900744 | regulation of p38MAPK cascade(GO:1900744) |
| 0.1 | 0.3 | GO:0003210 | cardiac atrium formation(GO:0003210) |
| 0.1 | 0.2 | GO:0044320 | leptin-mediated signaling pathway(GO:0033210) cellular response to leptin stimulus(GO:0044320) response to leptin(GO:0044321) |
| 0.1 | 0.2 | GO:0090008 | hypoblast development(GO:0090008) |
| 0.1 | 1.2 | GO:0021984 | adenohypophysis development(GO:0021984) |
| 0.1 | 0.2 | GO:1905208 | negative regulation of striated muscle cell differentiation(GO:0051154) negative regulation of cardiac muscle tissue development(GO:0055026) cardiac muscle cell fate commitment(GO:0060923) canonical Wnt signaling pathway involved in cardiac muscle cell fate commitment(GO:0061317) regulation of canonical Wnt signaling pathway involved in cardiac muscle cell fate commitment(GO:1901295) negative regulation of canonical Wnt signaling pathway involved in cardiac muscle cell fate commitment(GO:1901296) negative regulation of cardiocyte differentiation(GO:1905208) negative regulation of cardiac muscle cell differentiation(GO:2000726) |
| 0.1 | 1.3 | GO:0007168 | receptor guanylyl cyclase signaling pathway(GO:0007168) |
| 0.1 | 0.9 | GO:0097034 | mitochondrial respiratory chain complex IV assembly(GO:0033617) mitochondrial respiratory chain complex IV biogenesis(GO:0097034) |
| 0.0 | 0.5 | GO:1902624 | positive regulation of neutrophil migration(GO:1902624) |
| 0.0 | 0.4 | GO:0035777 | pronephric distal tubule development(GO:0035777) |
| 0.0 | 0.2 | GO:0006121 | mitochondrial electron transport, succinate to ubiquinone(GO:0006121) |
| 0.0 | 0.9 | GO:0006198 | cAMP catabolic process(GO:0006198) |
| 0.0 | 0.3 | GO:0071691 | cardiac muscle thin filament assembly(GO:0071691) |
| 0.0 | 1.7 | GO:0048247 | lymphocyte chemotaxis(GO:0048247) |
| 0.0 | 0.2 | GO:0002164 | larval development(GO:0002164) larval heart development(GO:0007508) angiogenesis involved in wound healing(GO:0060055) |
| 0.0 | 0.4 | GO:0002483 | antigen processing and presentation of endogenous peptide antigen(GO:0002483) antigen processing and presentation of endogenous antigen(GO:0019883) antigen processing and presentation of endogenous peptide antigen via MHC class I(GO:0019885) |
| 0.0 | 0.1 | GO:0045671 | negative regulation of osteoclast differentiation(GO:0045671) |
| 0.0 | 0.2 | GO:0019441 | tryptophan catabolic process to kynurenine(GO:0019441) |
| 0.0 | 0.3 | GO:0010887 | negative regulation of cholesterol storage(GO:0010887) |
| 0.0 | 0.3 | GO:0071260 | cellular response to mechanical stimulus(GO:0071260) |
| 0.0 | 0.4 | GO:0043567 | regulation of insulin-like growth factor receptor signaling pathway(GO:0043567) |
| 0.0 | 0.1 | GO:1903385 | dendrite guidance(GO:0070983) regulation of homophilic cell adhesion(GO:1903385) |
| 0.0 | 0.3 | GO:0006611 | protein export from nucleus(GO:0006611) |
| 0.0 | 0.3 | GO:0003315 | heart rudiment formation(GO:0003315) |
| 0.0 | 0.3 | GO:0097369 | sodium ion import(GO:0097369) sodium ion import across plasma membrane(GO:0098719) sodium ion import into cell(GO:1990118) |
| 0.0 | 0.6 | GO:0002098 | tRNA wobble uridine modification(GO:0002098) |
| 0.0 | 0.2 | GO:0032793 | positive regulation of CREB transcription factor activity(GO:0032793) |
| 0.0 | 0.6 | GO:0035435 | phosphate ion transmembrane transport(GO:0035435) |
| 0.0 | 0.1 | GO:0044333 | Wnt signaling pathway involved in digestive tract morphogenesis(GO:0044333) |
| 0.0 | 0.2 | GO:0006574 | valine catabolic process(GO:0006574) |
| 0.0 | 9.3 | GO:0046777 | protein autophosphorylation(GO:0046777) |
| 0.0 | 0.4 | GO:0015721 | bile acid and bile salt transport(GO:0015721) |
| 0.0 | 0.2 | GO:0006542 | glutamine biosynthetic process(GO:0006542) |
| 0.0 | 0.2 | GO:0032570 | response to progesterone(GO:0032570) |
| 0.0 | 0.1 | GO:0090161 | Golgi ribbon formation(GO:0090161) |
| 0.0 | 0.1 | GO:0050861 | positive regulation of B cell receptor signaling pathway(GO:0050861) |
| 0.0 | 0.4 | GO:1903845 | negative regulation of transforming growth factor beta receptor signaling pathway(GO:0030512) negative regulation of cellular response to transforming growth factor beta stimulus(GO:1903845) |
| 0.0 | 0.5 | GO:0050654 | chondroitin sulfate proteoglycan metabolic process(GO:0050654) |
| 0.0 | 1.9 | GO:0002088 | lens development in camera-type eye(GO:0002088) |
| 0.0 | 1.1 | GO:0055113 | epiboly involved in gastrulation with mouth forming second(GO:0055113) |
| 0.0 | 0.6 | GO:0046058 | cAMP metabolic process(GO:0046058) |
| 0.0 | 0.3 | GO:0043252 | sodium-independent organic anion transport(GO:0043252) |
| 0.0 | 0.1 | GO:0003322 | pancreatic A cell development(GO:0003322) |
| 0.0 | 0.7 | GO:0043113 | receptor clustering(GO:0043113) |
| 0.0 | 0.2 | GO:2000134 | negative regulation of cell cycle G1/S phase transition(GO:1902807) negative regulation of G1/S transition of mitotic cell cycle(GO:2000134) |
| 0.0 | 0.0 | GO:0007638 | mechanosensory behavior(GO:0007638) |
| 0.0 | 0.3 | GO:0020027 | hemoglobin metabolic process(GO:0020027) |
| 0.0 | 0.4 | GO:0045332 | phospholipid translocation(GO:0045332) |
| 0.0 | 0.1 | GO:0006953 | acute-phase response(GO:0006953) |
| 0.0 | 0.4 | GO:0007212 | dopamine receptor signaling pathway(GO:0007212) |
| 0.0 | 0.3 | GO:0055064 | chloride ion homeostasis(GO:0055064) |
| 0.0 | 0.4 | GO:0060319 | primitive erythrocyte differentiation(GO:0060319) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.2 | GO:0000801 | central element(GO:0000801) |
| 0.1 | 0.4 | GO:0036156 | inner dynein arm(GO:0036156) |
| 0.0 | 0.2 | GO:0045283 | mitochondrial respiratory chain complex II, succinate dehydrogenase complex (ubiquinone)(GO:0005749) succinate dehydrogenase complex (ubiquinone)(GO:0045257) respiratory chain complex II(GO:0045273) succinate dehydrogenase complex(GO:0045281) fumarate reductase complex(GO:0045283) |
| 0.0 | 0.2 | GO:0070390 | transcription export complex 2(GO:0070390) |
| 0.0 | 1.1 | GO:0097610 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.0 | 1.5 | GO:0031305 | integral component of mitochondrial inner membrane(GO:0031305) |
| 0.0 | 0.2 | GO:0070062 | extracellular exosome(GO:0070062) |
| 0.0 | 1.3 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.0 | 0.7 | GO:0001725 | stress fiber(GO:0001725) contractile actin filament bundle(GO:0097517) |
| 0.0 | 0.2 | GO:0005669 | transcription factor TFIID complex(GO:0005669) |
| 0.0 | 0.1 | GO:0005697 | telomerase holoenzyme complex(GO:0005697) |
| 0.0 | 0.2 | GO:0031083 | BLOC-1 complex(GO:0031083) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.8 | GO:0008310 | single-stranded DNA 3'-5' exodeoxyribonuclease activity(GO:0008310) |
| 0.2 | 0.9 | GO:0030116 | glial cell-derived neurotrophic factor receptor binding(GO:0030116) |
| 0.2 | 0.9 | GO:0052656 | branched-chain-amino-acid transaminase activity(GO:0004084) L-leucine transaminase activity(GO:0052654) L-valine transaminase activity(GO:0052655) L-isoleucine transaminase activity(GO:0052656) |
| 0.2 | 0.8 | GO:0031852 | mu-type opioid receptor binding(GO:0031852) corticotropin-releasing hormone receptor 1 binding(GO:0051430) |
| 0.1 | 0.7 | GO:0060182 | apelin receptor activity(GO:0060182) |
| 0.1 | 0.8 | GO:0035312 | 5'-3' exodeoxyribonuclease activity(GO:0035312) |
| 0.1 | 0.4 | GO:0003989 | acetyl-CoA carboxylase activity(GO:0003989) |
| 0.1 | 0.5 | GO:0050431 | transforming growth factor beta binding(GO:0050431) |
| 0.1 | 0.2 | GO:0004833 | tryptophan 2,3-dioxygenase activity(GO:0004833) |
| 0.1 | 0.2 | GO:0000104 | succinate dehydrogenase activity(GO:0000104) |
| 0.1 | 0.3 | GO:0042030 | ATPase inhibitor activity(GO:0042030) |
| 0.1 | 0.8 | GO:0015187 | glycine transmembrane transporter activity(GO:0015187) |
| 0.1 | 0.6 | GO:0015250 | water channel activity(GO:0015250) |
| 0.1 | 1.6 | GO:0004693 | cyclin-dependent protein serine/threonine kinase activity(GO:0004693) cyclin-dependent protein kinase activity(GO:0097472) |
| 0.1 | 0.3 | GO:0015349 | thyroid hormone transmembrane transporter activity(GO:0015349) |
| 0.1 | 0.3 | GO:0004122 | cystathionine beta-synthase activity(GO:0004122) |
| 0.1 | 0.4 | GO:0008508 | bile acid:sodium symporter activity(GO:0008508) |
| 0.1 | 0.5 | GO:0004499 | N,N-dimethylaniline monooxygenase activity(GO:0004499) |
| 0.0 | 0.6 | GO:0005315 | inorganic phosphate transmembrane transporter activity(GO:0005315) |
| 0.0 | 1.7 | GO:0048020 | CCR chemokine receptor binding(GO:0048020) |
| 0.0 | 0.2 | GO:0070891 | lipoteichoic acid binding(GO:0070891) |
| 0.0 | 0.5 | GO:0019238 | cyclohydrolase activity(GO:0019238) |
| 0.0 | 0.2 | GO:0008442 | 3-hydroxyisobutyrate dehydrogenase activity(GO:0008442) |
| 0.0 | 0.2 | GO:0031705 | bombesin receptor binding(GO:0031705) |
| 0.0 | 0.4 | GO:0047676 | arachidonate-CoA ligase activity(GO:0047676) |
| 0.0 | 0.9 | GO:0004115 | 3',5'-cyclic-AMP phosphodiesterase activity(GO:0004115) |
| 0.0 | 0.5 | GO:0031682 | G-protein gamma-subunit binding(GO:0031682) |
| 0.0 | 0.4 | GO:0031995 | insulin-like growth factor I binding(GO:0031994) insulin-like growth factor II binding(GO:0031995) |
| 0.0 | 2.6 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.0 | 0.1 | GO:0046964 | 3'-phosphoadenosine 5'-phosphosulfate transmembrane transporter activity(GO:0046964) |
| 0.0 | 0.3 | GO:0032036 | myosin heavy chain binding(GO:0032036) |
| 0.0 | 0.3 | GO:0015386 | monovalent cation:proton antiporter activity(GO:0005451) sodium:proton antiporter activity(GO:0015385) potassium:proton antiporter activity(GO:0015386) |
| 0.0 | 0.2 | GO:0008140 | cAMP response element binding protein binding(GO:0008140) |
| 0.0 | 0.2 | GO:0004300 | enoyl-CoA hydratase activity(GO:0004300) |
| 0.0 | 0.2 | GO:0004356 | glutamate-ammonia ligase activity(GO:0004356) ammonia ligase activity(GO:0016211) acid-ammonia (or amide) ligase activity(GO:0016880) |
| 0.0 | 0.6 | GO:0004016 | adenylate cyclase activity(GO:0004016) |
| 0.0 | 0.7 | GO:0022833 | mechanically-gated ion channel activity(GO:0008381) mechanically gated channel activity(GO:0022833) |
| 0.0 | 0.1 | GO:0036042 | long-chain fatty acyl-CoA binding(GO:0036042) |
| 0.0 | 0.6 | GO:0030165 | PDZ domain binding(GO:0030165) |
| 0.0 | 0.1 | GO:0070034 | telomerase RNA binding(GO:0070034) |
| 0.0 | 0.1 | GO:0004095 | carnitine O-palmitoyltransferase activity(GO:0004095) O-palmitoyltransferase activity(GO:0016416) |
| 0.0 | 0.9 | GO:0019843 | rRNA binding(GO:0019843) |
| 0.0 | 0.5 | GO:0042577 | lipid phosphatase activity(GO:0042577) |
| 0.0 | 0.2 | GO:0005021 | vascular endothelial growth factor-activated receptor activity(GO:0005021) |
| 0.0 | 1.6 | GO:0001227 | transcriptional repressor activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001227) |
| 0.0 | 1.3 | GO:0051427 | hormone receptor binding(GO:0051427) |
| 0.0 | 0.3 | GO:0005247 | voltage-gated chloride channel activity(GO:0005247) |
| 0.0 | 0.3 | GO:0004703 | G-protein coupled receptor kinase activity(GO:0004703) |
| 0.0 | 0.2 | GO:0070700 | BMP receptor binding(GO:0070700) |
| 0.0 | 0.2 | GO:0001102 | RNA polymerase II activating transcription factor binding(GO:0001102) |
| 0.0 | 0.5 | GO:0005518 | collagen binding(GO:0005518) |
| 0.0 | 2.2 | GO:0008083 | growth factor activity(GO:0008083) |
| 0.0 | 0.4 | GO:0045505 | dynein intermediate chain binding(GO:0045505) |
| 0.0 | 0.1 | GO:0008048 | calcium sensitive guanylate cyclase activator activity(GO:0008048) |
| 0.0 | 0.3 | GO:0015035 | protein disulfide oxidoreductase activity(GO:0015035) |
| 0.0 | 0.0 | GO:0047961 | glycine N-acyltransferase activity(GO:0047961) |
| 0.0 | 0.2 | GO:0048495 | Roundabout binding(GO:0048495) |
| 0.0 | 1.2 | GO:0016209 | antioxidant activity(GO:0016209) |
| 0.0 | 0.0 | GO:0016206 | catechol O-methyltransferase activity(GO:0016206) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.6 | PID ALK2 PATHWAY | ALK2 signaling events |
| 0.1 | 1.5 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.1 | 0.7 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.0 | 0.6 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 0.0 | 0.5 | PID SYNDECAN 3 PATHWAY | Syndecan-3-mediated signaling events |
| 0.0 | 1.0 | PID ATM PATHWAY | ATM pathway |
| 0.0 | 0.2 | ST STAT3 PATHWAY | STAT3 Pathway |
| 0.0 | 0.8 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 0.2 | SA G2 AND M PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
| 0.0 | 0.9 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.0 | 0.4 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.7 | REACTOME ACTIVATION OF RAC | Genes involved in Activation of Rac |
| 0.1 | 0.5 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |
| 0.1 | 0.8 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.1 | 0.6 | REACTOME ADENYLATE CYCLASE ACTIVATING PATHWAY | Genes involved in Adenylate cyclase activating pathway |
| 0.0 | 0.8 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.0 | 0.5 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.0 | 0.4 | REACTOME GLUCAGON SIGNALING IN METABOLIC REGULATION | Genes involved in Glucagon signaling in metabolic regulation |
| 0.0 | 0.5 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.0 | 0.4 | REACTOME FATTY ACYL COA BIOSYNTHESIS | Genes involved in Fatty Acyl-CoA Biosynthesis |
| 0.0 | 1.6 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.0 | 0.2 | REACTOME DCC MEDIATED ATTRACTIVE SIGNALING | Genes involved in DCC mediated attractive signaling |
| 0.0 | 0.2 | REACTOME IL 6 SIGNALING | Genes involved in Interleukin-6 signaling |
| 0.0 | 0.2 | REACTOME G2 M DNA DAMAGE CHECKPOINT | Genes involved in G2/M DNA damage checkpoint |
| 0.0 | 0.9 | REACTOME CELL JUNCTION ORGANIZATION | Genes involved in Cell junction organization |
| 0.0 | 0.8 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.0 | 0.2 | REACTOME ACTIVATED AMPK STIMULATES FATTY ACID OXIDATION IN MUSCLE | Genes involved in Activated AMPK stimulates fatty-acid oxidation in muscle |
| 0.0 | 0.2 | REACTOME RNA POL I PROMOTER OPENING | Genes involved in RNA Polymerase I Promoter Opening |
| 0.0 | 0.0 | REACTOME APC C CDH1 MEDIATED DEGRADATION OF CDC20 AND OTHER APC C CDH1 TARGETED PROTEINS IN LATE MITOSIS EARLY G1 | Genes involved in APC/C:Cdh1 mediated degradation of Cdc20 and other APC/C:Cdh1 targeted proteins in late mitosis/early G1 |