Project
PRJNA438478: RNAseq of wild type zebrafish germline
Navigation
Downloads
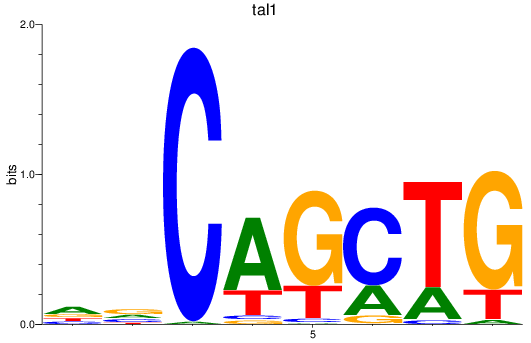
Results for tal1
Z-value: 1.74
Transcription factors associated with tal1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
tal1
|
ENSDARG00000019930 | T-cell acute lymphocytic leukemia 1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| tal1 | dr11_v1_chr22_+_16535575_16535575 | 0.83 | 1.7e-05 | Click! |
Activity profile of tal1 motif
Sorted Z-values of tal1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr5_+_42467867 | 6.61 |
ENSDART00000172028
|
pimr58
|
Pim proto-oncogene, serine/threonine kinase, related 58 |
| chr23_-_45319592 | 6.07 |
ENSDART00000189861
|
ccdc171
|
coiled-coil domain containing 171 |
| chr5_+_338154 | 4.84 |
ENSDART00000191743
|
rnf170
|
ring finger protein 170 |
| chr15_-_551177 | 4.82 |
ENSDART00000066774
ENSDART00000154617 |
tagln
|
transgelin |
| chr17_-_39786222 | 4.65 |
ENSDART00000154515
|
pimr62
|
Pim proto-oncogene, serine/threonine kinase, related 62 |
| chr17_-_24838927 | 4.63 |
ENSDART00000123259
|
CR388164.1
|
|
| chr3_-_61592417 | 4.54 |
ENSDART00000155082
|
nptx2a
|
neuronal pentraxin 2a |
| chr25_-_18470695 | 4.17 |
ENSDART00000034377
|
cpa5
|
carboxypeptidase A5 |
| chr7_-_6470431 | 4.02 |
ENSDART00000081359
|
zgc:110425
|
zgc:110425 |
| chr12_-_656540 | 3.93 |
ENSDART00000172651
|
sult2st2
|
sulfotransferase family 2, cytosolic sulfotransferase 2 |
| chr17_+_53156530 | 3.47 |
ENSDART00000126277
ENSDART00000156774 |
dph6
|
diphthamine biosynthesis 6 |
| chr24_+_17269849 | 3.37 |
ENSDART00000017605
|
spag6
|
sperm associated antigen 6 |
| chr3_+_58833306 | 3.37 |
ENSDART00000113223
|
igl1c3
|
immunoglobulin light 1 constant 3 |
| chr7_+_44715224 | 3.36 |
ENSDART00000184630
|
si:dkey-56m19.5
|
si:dkey-56m19.5 |
| chr2_+_49864219 | 3.29 |
ENSDART00000187744
|
rpl37
|
ribosomal protein L37 |
| chr11_+_25259058 | 3.26 |
ENSDART00000154109
|
tp53inp2
|
tumor protein p53 inducible nuclear protein 2 |
| chr7_+_24049776 | 3.23 |
ENSDART00000166559
|
efs
|
embryonal Fyn-associated substrate |
| chr24_-_26310854 | 3.23 |
ENSDART00000080113
|
apodb
|
apolipoprotein Db |
| chr8_-_48704050 | 3.23 |
ENSDART00000163916
|
pimr182
|
Pim proto-oncogene, serine/threonine kinase, related 182 |
| chr6_+_9241121 | 3.22 |
ENSDART00000064989
|
pimr70
|
Pim proto-oncogene, serine/threonine kinase, related 70 |
| chr3_-_1190132 | 3.19 |
ENSDART00000149709
|
smdt1a
|
single-pass membrane protein with aspartate-rich tail 1a |
| chr5_-_15692030 | 3.14 |
ENSDART00000099545
|
CR384085.1
|
|
| chr5_-_10007897 | 3.10 |
ENSDART00000109052
|
CR936408.1
|
Danio rerio uncharacterized LOC799523 (LOC799523), mRNA. |
| chr19_+_27589201 | 3.06 |
ENSDART00000182060
|
si:dkeyp-46h3.1
|
si:dkeyp-46h3.1 |
| chr20_+_25350435 | 2.99 |
ENSDART00000063034
|
fam228a
|
family with sequence similarity 228, member A |
| chr17_-_681142 | 2.95 |
ENSDART00000165583
|
soul3
|
heme-binding protein soul3 |
| chr14_-_36862745 | 2.94 |
ENSDART00000109293
|
rnf130
|
ring finger protein 130 |
| chr20_-_31075972 | 2.87 |
ENSDART00000122927
|
si:ch211-198b3.4
|
si:ch211-198b3.4 |
| chr18_+_5547185 | 2.85 |
ENSDART00000193977
|
nnt2
|
nicotinamide nucleotide transhydrogenase 2 |
| chr8_+_47817454 | 2.83 |
ENSDART00000144825
|
pimr185
|
Pim proto-oncogene, serine/threonine kinase, related 185 |
| chr13_-_9525527 | 2.80 |
ENSDART00000190618
|
CR848040.5
|
|
| chr18_-_85294 | 2.78 |
ENSDART00000044387
|
gabpb1
|
GA binding protein transcription factor, beta subunit 1 |
| chr10_+_4987766 | 2.73 |
ENSDART00000121959
|
si:ch73-234b20.5
|
si:ch73-234b20.5 |
| chr18_+_54354 | 2.63 |
ENSDART00000097163
|
zgc:158482
|
zgc:158482 |
| chr7_-_50917255 | 2.63 |
ENSDART00000022918
|
ankrd46b
|
ankyrin repeat domain 46b |
| chr3_-_32180796 | 2.60 |
ENSDART00000133191
ENSDART00000055308 |
pih1d1
|
PIH1 domain containing 1 |
| chr24_+_35827766 | 2.55 |
ENSDART00000144700
|
si:dkeyp-7a3.1
|
si:dkeyp-7a3.1 |
| chr23_+_578218 | 2.55 |
ENSDART00000055134
|
ogfr
|
opioid growth factor receptor |
| chr12_+_18533198 | 2.55 |
ENSDART00000189729
|
meiob
|
meiosis specific with OB domains |
| chr19_+_43978814 | 2.52 |
ENSDART00000102314
|
nkd3
|
naked cuticle homolog 3 |
| chr11_-_306533 | 2.51 |
ENSDART00000173437
|
poc1a
|
POC1 centriolar protein A |
| chr16_+_19014886 | 2.51 |
ENSDART00000079298
|
si:ch211-254p10.2
|
si:ch211-254p10.2 |
| chr16_+_9495583 | 2.48 |
ENSDART00000150750
ENSDART00000150457 |
pimr208
|
Pim proto-oncogene, serine/threonine kinase, related 208 |
| chr24_-_26399623 | 2.46 |
ENSDART00000112317
|
zgc:194621
|
zgc:194621 |
| chr13_-_438705 | 2.44 |
ENSDART00000082142
|
CU570800.1
|
|
| chr3_-_32590164 | 2.40 |
ENSDART00000151151
|
tspan4b
|
tetraspanin 4b |
| chr22_-_164944 | 2.40 |
ENSDART00000145379
|
fblim1
|
filamin binding LIM protein 1 |
| chr22_+_20720808 | 2.32 |
ENSDART00000171321
|
si:dkey-211f22.5
|
si:dkey-211f22.5 |
| chr1_+_22003173 | 2.28 |
ENSDART00000054388
|
dnah6
|
dynein, axonemal, heavy chain 6 |
| chr4_-_64703 | 2.27 |
ENSDART00000167851
|
CU856344.1
|
|
| chr14_+_9421510 | 2.27 |
ENSDART00000123652
|
hmgn6
|
high mobility group nucleosome binding domain 6 |
| chr25_+_1732838 | 2.26 |
ENSDART00000159555
ENSDART00000168161 |
FBLN1
|
fibulin 1 |
| chr3_-_1232384 | 2.22 |
ENSDART00000053892
|
si:ch1073-314i13.4
|
si:ch1073-314i13.4 |
| chr9_-_23747264 | 2.17 |
ENSDART00000141461
ENSDART00000010311 |
ndufa10
|
NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 10 |
| chr5_-_34185115 | 2.16 |
ENSDART00000192771
|
fibcd1
|
fibrinogen C domain containing 1 |
| chr1_+_22002649 | 2.16 |
ENSDART00000141317
|
dnah6
|
dynein, axonemal, heavy chain 6 |
| chr25_-_4482449 | 2.14 |
ENSDART00000056278
ENSDART00000149425 |
slc25a22a
|
solute carrier family 25 member 22a |
| chr19_+_31904836 | 2.03 |
ENSDART00000162297
ENSDART00000088340 ENSDART00000151280 ENSDART00000151218 |
tpd52
|
tumor protein D52 |
| chr7_+_71683853 | 2.02 |
ENSDART00000163002
|
emilin2b
|
elastin microfibril interfacer 2b |
| chr13_+_31402067 | 2.02 |
ENSDART00000019202
|
tdrd9
|
tudor domain containing 9 |
| chr1_-_59141715 | 2.01 |
ENSDART00000164941
ENSDART00000138870 |
si:ch1073-110a20.1
|
si:ch1073-110a20.1 |
| chr1_+_57311901 | 2.01 |
ENSDART00000149397
|
mycbpap
|
mycbp associated protein |
| chr11_-_1392468 | 2.00 |
ENSDART00000004423
|
iars
|
isoleucyl-tRNA synthetase |
| chr25_+_258883 | 1.97 |
ENSDART00000155256
|
zgc:92481
|
zgc:92481 |
| chr17_+_53311618 | 1.97 |
ENSDART00000166517
|
asb2b
|
ankyrin repeat and SOCS box containing 2b |
| chr13_+_51981764 | 1.96 |
ENSDART00000160698
|
klhdc3
|
kelch domain containing 3 |
| chr6_+_9893554 | 1.96 |
ENSDART00000064979
|
pimr74
|
Pim proto-oncogene, serine/threonine kinase, related 74 |
| chr2_-_16565690 | 1.94 |
ENSDART00000022549
|
atp1b3a
|
ATPase Na+/K+ transporting subunit beta 3a |
| chr15_+_28368823 | 1.92 |
ENSDART00000142298
|
slc43a2a
|
solute carrier family 43 (amino acid system L transporter), member 2a |
| chr12_+_18532866 | 1.91 |
ENSDART00000152126
ENSDART00000152443 ENSDART00000089589 |
meiob
|
meiosis specific with OB domains |
| chr3_+_54581987 | 1.89 |
ENSDART00000018071
|
eif3g
|
eukaryotic translation initiation factor 3, subunit G |
| chr11_-_22372072 | 1.89 |
ENSDART00000065996
|
tmem183a
|
transmembrane protein 183A |
| chr21_-_551014 | 1.88 |
ENSDART00000099252
|
vimr1
|
vimentin-related 1 |
| chr1_+_55002583 | 1.88 |
ENSDART00000037250
|
si:ch211-196h16.12
|
si:ch211-196h16.12 |
| chr2_-_42360329 | 1.88 |
ENSDART00000144274
ENSDART00000140863 |
si:dkey-7l6.3
|
si:dkey-7l6.3 |
| chr16_+_30575901 | 1.87 |
ENSDART00000077231
|
mc2r
|
melanocortin 2 receptor |
| chr23_+_26079467 | 1.86 |
ENSDART00000129617
|
atp6ap1b
|
ATPase H+ transporting accessory protein 1b |
| chr15_-_34051457 | 1.84 |
ENSDART00000189764
|
VWDE
|
si:dkey-30e9.7 |
| chr14_-_32258759 | 1.82 |
ENSDART00000052949
|
fgf13a
|
fibroblast growth factor 13a |
| chr11_-_1509773 | 1.82 |
ENSDART00000050762
|
phactr3b
|
phosphatase and actin regulator 3b |
| chr8_-_48770163 | 1.81 |
ENSDART00000160959
|
pimr184
|
Pim proto-oncogene, serine/threonine kinase, related 184 |
| chr7_-_6346859 | 1.78 |
ENSDART00000172913
|
si:ch73-368j24.11
|
si:ch73-368j24.11 |
| chr17_+_51940768 | 1.75 |
ENSDART00000053422
|
ttll5
|
tubulin tyrosine ligase-like family, member 5 |
| chr17_+_53439606 | 1.74 |
ENSDART00000154946
|
si:zfos-1714f5.3
|
si:zfos-1714f5.3 |
| chr5_+_28297254 | 1.74 |
ENSDART00000139862
ENSDART00000098635 |
vamp5
|
vesicle-associated membrane protein 5 |
| chr2_-_26620803 | 1.74 |
ENSDART00000003946
ENSDART00000126826 |
tmem59
|
transmembrane protein 59 |
| chr24_+_42132962 | 1.73 |
ENSDART00000187739
|
wwp1
|
WW domain containing E3 ubiquitin protein ligase 1 |
| chr2_+_19633493 | 1.70 |
ENSDART00000147989
|
pimr54
|
Pim proto-oncogene, serine/threonine kinase, related 54 |
| chr23_-_44848961 | 1.69 |
ENSDART00000136839
|
wu:fb72h05
|
wu:fb72h05 |
| chr17_+_25397070 | 1.68 |
ENSDART00000164254
|
zgc:154055
|
zgc:154055 |
| chr1_+_52560549 | 1.68 |
ENSDART00000167514
|
abca1a
|
ATP-binding cassette, sub-family A (ABC1), member 1A |
| chr19_+_233143 | 1.67 |
ENSDART00000175273
|
syngap1a
|
synaptic Ras GTPase activating protein 1a |
| chr21_+_2742823 | 1.63 |
ENSDART00000189004
|
CABZ01073424.2
|
|
| chr1_-_52494122 | 1.63 |
ENSDART00000131407
|
acy3.2
|
aspartoacylase (aminocyclase) 3, tandem duplicate 2 |
| chr15_-_4485828 | 1.63 |
ENSDART00000062868
|
tfdp2
|
transcription factor Dp-2 |
| chr8_-_27687095 | 1.62 |
ENSDART00000086946
|
mov10b.1
|
Moloney leukemia virus 10b, tandem duplicate 1 |
| chr3_-_62380146 | 1.61 |
ENSDART00000155853
|
gprc5ba
|
G protein-coupled receptor, class C, group 5, member Ba |
| chr8_+_7144066 | 1.61 |
ENSDART00000146306
|
slc6a6a
|
solute carrier family 6 (neurotransmitter transporter), member 6a |
| chr9_-_105135 | 1.61 |
ENSDART00000180126
|
FQ377903.3
|
|
| chr16_-_41503925 | 1.59 |
ENSDART00000029492
|
cmtm7
|
CKLF-like MARVEL transmembrane domain containing 7 |
| chr22_+_38310957 | 1.59 |
ENSDART00000040550
|
traf5
|
Tnf receptor-associated factor 5 |
| chr20_+_33981946 | 1.59 |
ENSDART00000131775
|
si:dkey-51e6.1
|
si:dkey-51e6.1 |
| chr24_-_7632187 | 1.58 |
ENSDART00000041714
|
atp6v0a1b
|
ATPase H+ transporting V0 subunit a1b |
| chr14_-_28432699 | 1.58 |
ENSDART00000054062
|
nek12
|
NIMA-related kinase 12 |
| chr5_+_41322783 | 1.57 |
ENSDART00000097546
|
arid3c
|
AT rich interactive domain 3C (BRIGHT-like) |
| chr3_-_1388936 | 1.54 |
ENSDART00000171278
|
ddx47
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 47 |
| chr25_-_204019 | 1.54 |
ENSDART00000188440
ENSDART00000191735 |
FP236318.2
|
|
| chr14_+_17125428 | 1.54 |
ENSDART00000161489
|
slc43a3b
|
solute carrier family 43, member 3b |
| chr3_-_53486169 | 1.51 |
ENSDART00000115243
|
soul5
|
heme-binding protein soul5 |
| chr18_-_92046 | 1.50 |
ENSDART00000157709
|
gabpb1
|
GA binding protein transcription factor, beta subunit 1 |
| chr20_-_284865 | 1.49 |
ENSDART00000104806
|
wisp3
|
WNT1 inducible signaling pathway protein 3 |
| chr20_+_54280549 | 1.48 |
ENSDART00000151048
|
actr10
|
ARP10 actin related protein 10 homolog |
| chr4_-_38033800 | 1.46 |
ENSDART00000159662
|
si:dkeyp-82b4.4
|
si:dkeyp-82b4.4 |
| chr2_+_16160906 | 1.45 |
ENSDART00000135783
|
selenoj
|
selenoprotein J |
| chr19_-_24555935 | 1.42 |
ENSDART00000132660
ENSDART00000162801 |
polr3gla
|
polymerase (RNA) III (DNA directed) polypeptide G like a |
| chr8_-_45759137 | 1.42 |
ENSDART00000123878
|
ppiaa
|
peptidylprolyl isomerase Aa (cyclophilin A) |
| chr20_-_49729446 | 1.42 |
ENSDART00000111089
|
filip1b
|
filamin A interacting protein 1b |
| chr14_+_48964628 | 1.41 |
ENSDART00000105427
|
mif
|
macrophage migration inhibitory factor |
| chr25_-_36369057 | 1.40 |
ENSDART00000064400
|
si:ch211-113a14.24
|
si:ch211-113a14.24 |
| chr8_+_40210398 | 1.38 |
ENSDART00000167612
|
rnf34a
|
ring finger protein 34a |
| chr4_+_61171310 | 1.34 |
ENSDART00000141738
|
si:dkey-9p20.18
|
si:dkey-9p20.18 |
| chr20_+_2460864 | 1.32 |
ENSDART00000131642
|
akap7
|
A kinase (PRKA) anchor protein 7 |
| chr1_+_52563298 | 1.32 |
ENSDART00000142465
|
abca1a
|
ATP-binding cassette, sub-family A (ABC1), member 1A |
| chr25_-_35153985 | 1.32 |
ENSDART00000154851
|
zgc:153405
|
zgc:153405 |
| chr4_-_1015896 | 1.31 |
ENSDART00000170292
|
FAM180A
|
family with sequence similarity 180 member A |
| chr14_-_17563773 | 1.31 |
ENSDART00000082667
|
fgfrl1a
|
fibroblast growth factor receptor like 1a |
| chr17_+_135590 | 1.30 |
ENSDART00000166339
|
klhdc2
|
kelch domain containing 2 |
| chr25_+_36045072 | 1.29 |
ENSDART00000126326
|
rpgrip1l
|
RPGRIP1-like |
| chr9_+_51225345 | 1.29 |
ENSDART00000132896
|
fap
|
fibroblast activation protein, alpha |
| chr23_-_40790196 | 1.28 |
ENSDART00000141216
|
si:dkey-194e6.2
|
si:dkey-194e6.2 |
| chr2_+_17524278 | 1.28 |
ENSDART00000165633
|
pimr196
|
Pim proto-oncogene, serine/threonine kinase, related 196 |
| chr7_+_26029672 | 1.27 |
ENSDART00000101126
|
alox12
|
arachidonate 12-lipoxygenase |
| chr4_-_67589158 | 1.26 |
ENSDART00000184361
|
BX088712.2
|
|
| chr5_-_26181863 | 1.26 |
ENSDART00000098500
|
ccdc125
|
coiled-coil domain containing 125 |
| chr5_-_5243079 | 1.25 |
ENSDART00000130576
ENSDART00000164377 |
mvb12ba
|
multivesicular body subunit 12Ba |
| chr19_-_24555623 | 1.22 |
ENSDART00000176022
|
polr3gla
|
polymerase (RNA) III (DNA directed) polypeptide G like a |
| chr13_+_31479814 | 1.20 |
ENSDART00000148112
|
lrrc9
|
leucine rich repeat containing 9 |
| chr7_-_27033080 | 1.19 |
ENSDART00000173516
|
nucb2a
|
nucleobindin 2a |
| chr5_-_417495 | 1.19 |
ENSDART00000180586
ENSDART00000189408 |
HOOK3
|
hook microtubule tethering protein 3 |
| chr14_-_38809561 | 1.19 |
ENSDART00000159159
|
sil1
|
SIL1 nucleotide exchange factor |
| chr12_-_48960308 | 1.19 |
ENSDART00000176247
|
CABZ01112647.1
|
|
| chr25_-_1333676 | 1.18 |
ENSDART00000183182
ENSDART00000188907 |
cln6b
|
CLN6, transmembrane ER protein b |
| chr7_+_5964296 | 1.16 |
ENSDART00000173380
|
si:dkey-23a13.17
|
si:dkey-23a13.17 |
| chr2_-_36350082 | 1.15 |
ENSDART00000193104
|
CU571384.1
|
|
| chr23_+_22656477 | 1.15 |
ENSDART00000009337
ENSDART00000133322 |
eno1a
|
enolase 1a, (alpha) |
| chr1_-_53468160 | 1.14 |
ENSDART00000143349
|
zgc:66455
|
zgc:66455 |
| chr6_-_51573975 | 1.14 |
ENSDART00000073865
|
rbl1
|
retinoblastoma-like 1 (p107) |
| chr10_-_16470648 | 1.14 |
ENSDART00000149104
|
fbn2a
|
fibrillin 2a |
| chr4_-_63848305 | 1.13 |
ENSDART00000097308
|
BX901974.1
|
|
| chr7_-_6415991 | 1.13 |
ENSDART00000173349
|
CU457819.3
|
Histone H3.2 |
| chr21_-_43040780 | 1.13 |
ENSDART00000187037
|
jakmip2
|
janus kinase and microtubule interacting protein 2 |
| chr7_+_25053331 | 1.12 |
ENSDART00000173998
|
si:dkey-23i12.7
|
si:dkey-23i12.7 |
| chr7_+_69470142 | 1.12 |
ENSDART00000073861
|
gabarapb
|
GABA(A) receptor-associated protein b |
| chr2_+_20793982 | 1.12 |
ENSDART00000014785
|
prg4a
|
proteoglycan 4a |
| chr10_-_25069155 | 1.10 |
ENSDART00000078226
ENSDART00000181941 |
mtnr1bb
|
melatonin receptor 1Bb |
| chr18_-_8398972 | 1.09 |
ENSDART00000136512
|
BEND7
|
si:ch211-220f12.4 |
| chr24_+_18714212 | 1.09 |
ENSDART00000171181
|
cspp1a
|
centrosome and spindle pole associated protein 1a |
| chr5_-_72376973 | 1.08 |
ENSDART00000164969
|
CABZ01100198.1
|
|
| chr10_-_8295294 | 1.08 |
ENSDART00000075412
ENSDART00000163803 |
plpp1a
|
phospholipid phosphatase 1a |
| chr9_-_1499721 | 1.07 |
ENSDART00000062940
|
si:ch211-222f23.6
|
si:ch211-222f23.6 |
| chr17_-_51938663 | 1.07 |
ENSDART00000179784
|
ERG28
|
ergosterol biosynthesis 28 homolog |
| chr8_+_49149907 | 1.07 |
ENSDART00000138746
|
snpha
|
syntaphilin a |
| chr3_+_17537352 | 1.06 |
ENSDART00000104549
|
hcrt
|
hypocretin (orexin) neuropeptide precursor |
| chr20_-_438646 | 1.05 |
ENSDART00000009196
|
ufl1
|
UFM1-specific ligase 1 |
| chr4_+_77943184 | 1.04 |
ENSDART00000159094
|
pacsin2
|
protein kinase C and casein kinase substrate in neurons 2 |
| chr10_-_8294965 | 1.04 |
ENSDART00000167380
|
plpp1a
|
phospholipid phosphatase 1a |
| chr4_-_5455506 | 1.04 |
ENSDART00000156593
ENSDART00000154676 |
si:dkey-14d8.22
|
si:dkey-14d8.22 |
| chr17_+_53311243 | 1.04 |
ENSDART00000160241
ENSDART00000160009 ENSDART00000162239 |
asb2b
|
ankyrin repeat and SOCS box containing 2b |
| chr4_-_64709908 | 1.01 |
ENSDART00000161032
|
si:dkey-9i5.2
|
si:dkey-9i5.2 |
| chr5_+_58520266 | 1.01 |
ENSDART00000108889
|
oafb
|
OAF homolog b (Drosophila) |
| chr4_+_2267641 | 1.01 |
ENSDART00000165503
|
si:ch73-89b15.3
|
si:ch73-89b15.3 |
| chr22_-_16494406 | 1.01 |
ENSDART00000062727
|
stx6
|
syntaxin 6 |
| chr1_-_38816685 | 1.01 |
ENSDART00000075230
|
asb5b
|
ankyrin repeat and SOCS box containing 5b |
| chr3_+_30500968 | 1.01 |
ENSDART00000103447
|
si:dkey-13n23.3
|
si:dkey-13n23.3 |
| chr23_-_16682186 | 1.00 |
ENSDART00000020810
|
sdcbp2
|
syndecan binding protein (syntenin) 2 |
| chr21_+_22840246 | 1.00 |
ENSDART00000151621
|
birc2
|
baculoviral IAP repeat containing 2 |
| chr4_+_332709 | 1.00 |
ENSDART00000067486
|
tulp4a
|
tubby like protein 4a |
| chr5_-_30984010 | 0.99 |
ENSDART00000182367
|
spns3
|
spinster homolog 3 (Drosophila) |
| chr25_+_36046287 | 0.98 |
ENSDART00000185324
|
rpgrip1l
|
RPGRIP1-like |
| chr1_+_41849152 | 0.98 |
ENSDART00000053685
|
smox
|
spermine oxidase |
| chr9_-_356501 | 0.97 |
ENSDART00000168654
|
zgc:158320
|
zgc:158320 |
| chr25_-_18454016 | 0.96 |
ENSDART00000005877
|
cpa1
|
carboxypeptidase A1 (pancreatic) |
| chr25_-_37501371 | 0.95 |
ENSDART00000160498
|
CABZ01088346.1
|
|
| chr1_-_8980665 | 0.95 |
ENSDART00000148182
|
si:ch73-191k20.3
|
si:ch73-191k20.3 |
| chr8_-_1051438 | 0.95 |
ENSDART00000067093
ENSDART00000170737 |
smyd1b
|
SET and MYND domain containing 1b |
| chr24_+_42074143 | 0.95 |
ENSDART00000170514
|
top1mt
|
DNA topoisomerase I mitochondrial |
| chr11_-_8782871 | 0.94 |
ENSDART00000158546
|
si:ch211-51h4.2
|
si:ch211-51h4.2 |
| chr25_-_3549321 | 0.93 |
ENSDART00000181214
ENSDART00000160600 |
hdhd5
|
haloacid dehalogenase like hydrolase domain containing 5 |
| chr25_+_37340722 | 0.93 |
ENSDART00000137025
|
pamr1
|
peptidase domain containing associated with muscle regeneration 1 |
| chr19_+_31404686 | 0.91 |
ENSDART00000078459
|
pip4p2
|
phosphatidylinositol-4,5-bisphosphate 4-phosphatase 2 |
| chr7_-_18508815 | 0.91 |
ENSDART00000173539
|
rgs12a
|
regulator of G protein signaling 12a |
| chr4_-_52165969 | 0.90 |
ENSDART00000171130
|
si:dkeyp-44b5.4
|
si:dkeyp-44b5.4 |
| chr8_-_54223316 | 0.89 |
ENSDART00000018054
|
trh
|
thyrotropin-releasing hormone |
| chr25_-_173165 | 0.87 |
ENSDART00000193594
|
CABZ01114053.1
|
|
| chr1_+_22851261 | 0.87 |
ENSDART00000193925
|
gtf2e2
|
general transcription factor IIE, polypeptide 2, beta |
| chr21_-_45073764 | 0.87 |
ENSDART00000181390
ENSDART00000063714 |
rapgef6
|
Rap guanine nucleotide exchange factor (GEF) 6 |
| chr3_+_1209006 | 0.86 |
ENSDART00000158633
|
poldip3
|
polymerase (DNA-directed), delta interacting protein 3 |
| chr20_-_54377933 | 0.85 |
ENSDART00000182664
|
entpd5b
|
ectonucleoside triphosphate diphosphohydrolase 5b |
| chr4_+_16787040 | 0.85 |
ENSDART00000039027
|
golt1ba
|
golgi transport 1Ba |
| chr4_-_20208166 | 0.85 |
ENSDART00000066891
|
gsap
|
gamma-secretase activating protein |
Network of associatons between targets according to the STRING database.
First level regulatory network of tal1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.8 | 3.0 | GO:0010874 | regulation of cholesterol efflux(GO:0010874) |
| 0.7 | 2.9 | GO:0006740 | NADPH regeneration(GO:0006740) |
| 0.6 | 3.2 | GO:0090527 | actin filament reorganization(GO:0090527) |
| 0.6 | 3.5 | GO:0017182 | peptidyl-diphthamide metabolic process(GO:0017182) peptidyl-diphthamide biosynthetic process from peptidyl-histidine(GO:0017183) |
| 0.5 | 2.7 | GO:0070254 | mucus secretion(GO:0070254) |
| 0.5 | 1.6 | GO:0035971 | peptidyl-histidine dephosphorylation(GO:0035971) |
| 0.5 | 1.9 | GO:0042755 | eating behavior(GO:0042755) |
| 0.4 | 1.9 | GO:1901207 | regulation of heart looping(GO:1901207) |
| 0.4 | 2.6 | GO:0000492 | box C/D snoRNP assembly(GO:0000492) |
| 0.3 | 6.2 | GO:0016584 | nucleosome positioning(GO:0016584) |
| 0.3 | 1.6 | GO:0098795 | mRNA cleavage involved in gene silencing by miRNA(GO:0035279) mRNA cleavage involved in gene silencing(GO:0098795) |
| 0.3 | 2.5 | GO:0034755 | iron ion transmembrane transport(GO:0034755) |
| 0.3 | 1.3 | GO:1902592 | virion assembly(GO:0019068) virus maturation(GO:0019075) multi-organism membrane organization(GO:0044803) viral budding(GO:0046755) multi-organism organelle organization(GO:1902590) multi-organism membrane budding(GO:1902592) |
| 0.3 | 1.5 | GO:0032289 | central nervous system myelin formation(GO:0032289) |
| 0.3 | 2.0 | GO:0007141 | male meiosis I(GO:0007141) |
| 0.3 | 1.0 | GO:0060547 | negative regulation of necroptotic process(GO:0060546) negative regulation of necrotic cell death(GO:0060547) |
| 0.2 | 0.7 | GO:0006432 | phenylalanyl-tRNA aminoacylation(GO:0006432) |
| 0.2 | 0.5 | GO:0065001 | specification of axis polarity(GO:0065001) |
| 0.2 | 1.1 | GO:1990592 | IRE1-mediated unfolded protein response(GO:0036498) negative regulation of response to endoplasmic reticulum stress(GO:1903573) protein polyufmylation(GO:1990564) protein K69-linked ufmylation(GO:1990592) |
| 0.2 | 2.1 | GO:0015813 | acidic amino acid transport(GO:0015800) aspartate transport(GO:0015810) L-glutamate transport(GO:0015813) malate-aspartate shuttle(GO:0043490) |
| 0.2 | 1.1 | GO:0051659 | maintenance of mitochondrion location(GO:0051659) |
| 0.2 | 4.4 | GO:0060287 | epithelial cilium movement involved in determination of left/right asymmetry(GO:0060287) |
| 0.2 | 2.3 | GO:0046548 | retinal rod cell development(GO:0046548) |
| 0.2 | 0.8 | GO:0002532 | production of molecular mediator involved in inflammatory response(GO:0002532) |
| 0.2 | 1.4 | GO:2001270 | negative regulation of execution phase of apoptosis(GO:1900118) regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:2001270) negative regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:2001271) |
| 0.2 | 5.0 | GO:0010508 | positive regulation of autophagy(GO:0010508) |
| 0.2 | 0.8 | GO:0014743 | regulation of muscle hypertrophy(GO:0014743) |
| 0.2 | 3.3 | GO:0097034 | mitochondrial respiratory chain complex IV assembly(GO:0033617) mitochondrial respiratory chain complex IV biogenesis(GO:0097034) |
| 0.2 | 4.5 | GO:0000712 | resolution of meiotic recombination intermediates(GO:0000712) |
| 0.2 | 2.1 | GO:0032098 | regulation of appetite(GO:0032098) |
| 0.2 | 1.3 | GO:0032889 | regulation of vacuole fusion, non-autophagic(GO:0032889) |
| 0.2 | 0.5 | GO:0080120 | CAAX-box protein processing(GO:0071586) CAAX-box protein maturation(GO:0080120) |
| 0.2 | 2.1 | GO:1905150 | regulation of voltage-gated sodium channel activity(GO:1905150) |
| 0.2 | 3.2 | GO:0048251 | elastic fiber assembly(GO:0048251) |
| 0.1 | 3.9 | GO:0051923 | sulfation(GO:0051923) |
| 0.1 | 0.3 | GO:1904861 | excitatory synapse assembly(GO:1904861) |
| 0.1 | 1.9 | GO:0030325 | adrenal gland development(GO:0030325) |
| 0.1 | 1.7 | GO:0035493 | SNARE complex assembly(GO:0035493) |
| 0.1 | 4.0 | GO:0055003 | cardiac myofibril assembly(GO:0055003) |
| 0.1 | 0.4 | GO:0090278 | negative regulation of peptide secretion(GO:0002792) negative regulation of peptide hormone secretion(GO:0090278) |
| 0.1 | 2.2 | GO:0006120 | mitochondrial electron transport, NADH to ubiquinone(GO:0006120) |
| 0.1 | 0.8 | GO:0060855 | venous endothelial cell migration involved in lymph vessel development(GO:0060855) |
| 0.1 | 1.6 | GO:0051452 | vacuolar acidification(GO:0007035) pH reduction(GO:0045851) intracellular pH reduction(GO:0051452) |
| 0.1 | 0.5 | GO:0048245 | eosinophil chemotaxis(GO:0048245) eosinophil migration(GO:0072677) |
| 0.1 | 1.3 | GO:0021592 | fourth ventricle development(GO:0021592) |
| 0.1 | 1.9 | GO:0036376 | cellular potassium ion homeostasis(GO:0030007) sodium ion export from cell(GO:0036376) sodium ion export(GO:0071436) |
| 0.1 | 3.2 | GO:0007568 | aging(GO:0007568) |
| 0.1 | 0.4 | GO:0051876 | pigment granule dispersal(GO:0051876) |
| 0.1 | 0.4 | GO:1903146 | regulation of mitophagy(GO:1903146) |
| 0.1 | 1.0 | GO:0001765 | membrane raft assembly(GO:0001765) membrane raft organization(GO:0031579) plasma membrane raft assembly(GO:0044854) plasma membrane raft organization(GO:0044857) caveola assembly(GO:0070836) |
| 0.1 | 1.9 | GO:0043562 | cellular response to nitrogen starvation(GO:0006995) cellular response to nitrogen levels(GO:0043562) |
| 0.1 | 0.4 | GO:0038003 | opioid receptor signaling pathway(GO:0038003) |
| 0.1 | 1.7 | GO:0036444 | calcium ion transmembrane import into mitochondrion(GO:0036444) |
| 0.1 | 1.8 | GO:0018095 | protein polyglutamylation(GO:0018095) |
| 0.1 | 3.2 | GO:0006910 | phagocytosis, recognition(GO:0006910) |
| 0.1 | 1.3 | GO:0035024 | negative regulation of Rho protein signal transduction(GO:0035024) |
| 0.1 | 0.8 | GO:0034205 | beta-amyloid formation(GO:0034205) amyloid precursor protein catabolic process(GO:0042987) |
| 0.1 | 0.8 | GO:0000056 | ribosomal small subunit export from nucleus(GO:0000056) |
| 0.1 | 2.0 | GO:0030183 | B cell differentiation(GO:0030183) |
| 0.1 | 0.5 | GO:0003210 | cardiac atrium formation(GO:0003210) |
| 0.1 | 2.6 | GO:0006383 | transcription from RNA polymerase III promoter(GO:0006383) |
| 0.1 | 0.3 | GO:0010847 | regulation of chromatin assembly(GO:0010847) |
| 0.1 | 0.5 | GO:1900028 | wound healing, spreading of epidermal cells(GO:0035313) negative regulation of ruffle assembly(GO:1900028) |
| 0.1 | 30.0 | GO:0046777 | protein autophosphorylation(GO:0046777) |
| 0.1 | 0.5 | GO:0032218 | riboflavin transport(GO:0032218) |
| 0.1 | 0.9 | GO:0051122 | lipoxygenase pathway(GO:0019372) linoleic acid metabolic process(GO:0043651) hepoxilin metabolic process(GO:0051121) hepoxilin biosynthetic process(GO:0051122) |
| 0.1 | 1.6 | GO:0033209 | tumor necrosis factor-mediated signaling pathway(GO:0033209) |
| 0.1 | 0.4 | GO:0098535 | de novo centriole assembly(GO:0098535) |
| 0.1 | 0.8 | GO:0032481 | positive regulation of type I interferon production(GO:0032481) |
| 0.1 | 0.6 | GO:0035721 | intraciliary retrograde transport(GO:0035721) |
| 0.1 | 0.5 | GO:0048680 | positive regulation of axon regeneration(GO:0048680) positive regulation of neuron projection regeneration(GO:0070572) |
| 0.1 | 1.9 | GO:0001732 | formation of cytoplasmic translation initiation complex(GO:0001732) |
| 0.1 | 5.0 | GO:0051966 | regulation of synaptic transmission, glutamatergic(GO:0051966) |
| 0.1 | 2.0 | GO:0015807 | L-amino acid transport(GO:0015807) |
| 0.1 | 0.9 | GO:0006265 | DNA topological change(GO:0006265) |
| 0.1 | 0.4 | GO:0051754 | meiotic sister chromatid cohesion, centromeric(GO:0051754) |
| 0.1 | 1.3 | GO:0043124 | negative regulation of I-kappaB kinase/NF-kappaB signaling(GO:0043124) |
| 0.1 | 0.3 | GO:0098529 | neuromuscular junction development, skeletal muscle fiber(GO:0098529) |
| 0.1 | 1.2 | GO:0010507 | negative regulation of autophagy(GO:0010507) |
| 0.1 | 0.3 | GO:0060282 | positive regulation of oocyte development(GO:0060282) |
| 0.1 | 0.9 | GO:0009134 | nucleoside diphosphate catabolic process(GO:0009134) |
| 0.0 | 1.4 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.0 | 1.4 | GO:0060117 | auditory receptor cell development(GO:0060117) |
| 0.0 | 1.6 | GO:0072348 | sulfur compound transport(GO:0072348) |
| 0.0 | 0.1 | GO:0060945 | activation of transmembrane receptor protein tyrosine kinase activity(GO:0007171) Schwann cell migration(GO:0036135) cardiac neuron differentiation(GO:0060945) cardiac neuron development(GO:0060959) |
| 0.0 | 0.4 | GO:0050732 | negative regulation of peptidyl-tyrosine phosphorylation(GO:0050732) negative regulation of protein tyrosine kinase activity(GO:0061099) |
| 0.0 | 0.7 | GO:0035435 | phosphate ion transmembrane transport(GO:0035435) |
| 0.0 | 0.1 | GO:0010265 | SCF complex assembly(GO:0010265) |
| 0.0 | 0.2 | GO:0000183 | chromatin silencing at rDNA(GO:0000183) |
| 0.0 | 1.2 | GO:0007040 | lysosome organization(GO:0007040) lytic vacuole organization(GO:0080171) |
| 0.0 | 0.2 | GO:0030242 | pexophagy(GO:0030242) |
| 0.0 | 1.1 | GO:0007157 | heterophilic cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0007157) |
| 0.0 | 1.7 | GO:0046928 | regulation of neurotransmitter secretion(GO:0046928) |
| 0.0 | 0.7 | GO:0007274 | neuromuscular synaptic transmission(GO:0007274) |
| 0.0 | 0.6 | GO:0036092 | phosphatidylinositol-3-phosphate biosynthetic process(GO:0036092) |
| 0.0 | 0.3 | GO:0071376 | response to corticotropin-releasing hormone(GO:0043435) cellular response to corticotropin-releasing hormone stimulus(GO:0071376) |
| 0.0 | 2.3 | GO:0046839 | phospholipid dephosphorylation(GO:0046839) |
| 0.0 | 0.4 | GO:0019934 | cGMP-mediated signaling(GO:0019934) |
| 0.0 | 0.7 | GO:0030223 | neutrophil differentiation(GO:0030223) |
| 0.0 | 0.1 | GO:0051725 | protein de-ADP-ribosylation(GO:0051725) |
| 0.0 | 0.5 | GO:0034625 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.0 | 0.9 | GO:0016339 | calcium-dependent cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0016339) |
| 0.0 | 0.4 | GO:0043433 | negative regulation of sequence-specific DNA binding transcription factor activity(GO:0043433) |
| 0.0 | 0.1 | GO:0046333 | octopamine biosynthetic process(GO:0006589) dopamine catabolic process(GO:0042420) norepinephrine biosynthetic process(GO:0042421) octopamine metabolic process(GO:0046333) |
| 0.0 | 0.3 | GO:0099560 | synaptic membrane adhesion(GO:0099560) |
| 0.0 | 0.4 | GO:0071577 | zinc II ion transmembrane transport(GO:0071577) |
| 0.0 | 0.1 | GO:1904105 | regulation of Wnt signaling pathway, calcium modulating pathway(GO:0008591) positive regulation of convergent extension involved in gastrulation(GO:1904105) |
| 0.0 | 0.5 | GO:0014020 | neural tube closure(GO:0001843) primary neural tube formation(GO:0014020) |
| 0.0 | 0.9 | GO:0006367 | transcription initiation from RNA polymerase II promoter(GO:0006367) |
| 0.0 | 0.8 | GO:0042752 | regulation of circadian rhythm(GO:0042752) |
| 0.0 | 0.1 | GO:0060832 | oocyte animal/vegetal axis specification(GO:0060832) |
| 0.0 | 0.4 | GO:0046849 | bone remodeling(GO:0046849) |
| 0.0 | 0.6 | GO:0006919 | activation of cysteine-type endopeptidase activity involved in apoptotic process(GO:0006919) |
| 0.0 | 0.6 | GO:0030574 | collagen catabolic process(GO:0030574) multicellular organism catabolic process(GO:0044243) |
| 0.0 | 0.2 | GO:0070212 | protein poly-ADP-ribosylation(GO:0070212) |
| 0.0 | 0.3 | GO:0046051 | UTP biosynthetic process(GO:0006228) UTP metabolic process(GO:0046051) |
| 0.0 | 1.2 | GO:0048278 | vesicle docking(GO:0048278) |
| 0.0 | 0.4 | GO:0030322 | stabilization of membrane potential(GO:0030322) |
| 0.0 | 1.0 | GO:0007416 | synapse assembly(GO:0007416) |
| 0.0 | 0.1 | GO:0097510 | base-excision repair, AP site formation via deaminated base removal(GO:0097510) |
| 0.0 | 0.4 | GO:0051085 | chaperone mediated protein folding requiring cofactor(GO:0051085) |
| 0.0 | 0.1 | GO:1902624 | positive regulation of neutrophil migration(GO:1902624) |
| 0.0 | 0.7 | GO:0048484 | enteric nervous system development(GO:0048484) |
| 0.0 | 0.2 | GO:0007198 | adenylate cyclase-inhibiting serotonin receptor signaling pathway(GO:0007198) |
| 0.0 | 0.2 | GO:0098962 | regulation of postsynaptic neurotransmitter receptor activity(GO:0098962) |
| 0.0 | 0.3 | GO:0030513 | positive regulation of BMP signaling pathway(GO:0030513) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 2.6 | GO:0097255 | R2TP complex(GO:0097255) |
| 0.3 | 1.9 | GO:0033181 | plasma membrane proton-transporting V-type ATPase complex(GO:0033181) |
| 0.3 | 1.6 | GO:0070062 | extracellular exosome(GO:0070062) |
| 0.2 | 1.3 | GO:0031258 | lamellipodium membrane(GO:0031258) |
| 0.2 | 1.9 | GO:0005890 | sodium:potassium-exchanging ATPase complex(GO:0005890) cation-transporting ATPase complex(GO:0090533) |
| 0.2 | 0.7 | GO:0060171 | stereocilium membrane(GO:0060171) |
| 0.2 | 0.9 | GO:0005673 | transcription factor TFIIE complex(GO:0005673) |
| 0.2 | 1.3 | GO:0000015 | phosphopyruvate hydratase complex(GO:0000015) |
| 0.2 | 1.6 | GO:0000220 | vacuolar proton-transporting V-type ATPase, V0 domain(GO:0000220) |
| 0.2 | 2.3 | GO:0032391 | photoreceptor connecting cilium(GO:0032391) |
| 0.1 | 0.6 | GO:0061702 | inflammasome complex(GO:0061702) |
| 0.1 | 1.7 | GO:1990246 | uniplex complex(GO:1990246) |
| 0.1 | 3.6 | GO:0043186 | P granule(GO:0043186) |
| 0.1 | 3.2 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
| 0.1 | 1.5 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.1 | 1.9 | GO:0016282 | eukaryotic 43S preinitiation complex(GO:0016282) |
| 0.1 | 3.3 | GO:0016605 | PML body(GO:0016605) |
| 0.1 | 1.2 | GO:0030285 | integral component of synaptic vesicle membrane(GO:0030285) intrinsic component of synaptic vesicle membrane(GO:0098563) |
| 0.1 | 0.6 | GO:0030991 | intraciliary transport particle A(GO:0030991) |
| 0.1 | 4.0 | GO:0031305 | integral component of mitochondrial inner membrane(GO:0031305) |
| 0.1 | 0.8 | GO:0030688 | preribosome, small subunit precursor(GO:0030688) |
| 0.1 | 4.4 | GO:0030286 | dynein complex(GO:0030286) |
| 0.1 | 1.3 | GO:0000813 | ESCRT I complex(GO:0000813) |
| 0.1 | 3.2 | GO:0005901 | caveola(GO:0005901) plasma membrane raft(GO:0044853) |
| 0.1 | 0.5 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.1 | 0.2 | GO:1990072 | TRAPPIII protein complex(GO:1990072) |
| 0.1 | 6.9 | GO:0000786 | nucleosome(GO:0000786) |
| 0.1 | 0.6 | GO:0005943 | phosphatidylinositol 3-kinase complex, class IA(GO:0005943) |
| 0.1 | 5.0 | GO:0031201 | SNARE complex(GO:0031201) |
| 0.1 | 1.9 | GO:0000421 | autophagosome membrane(GO:0000421) |
| 0.1 | 2.2 | GO:0045271 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.0 | 1.0 | GO:0031430 | M band(GO:0031430) |
| 0.0 | 0.3 | GO:0071818 | BAT3 complex(GO:0071818) |
| 0.0 | 5.5 | GO:0005924 | cell-substrate adherens junction(GO:0005924) focal adhesion(GO:0005925) |
| 0.0 | 0.2 | GO:0033553 | rDNA heterochromatin(GO:0033553) |
| 0.0 | 0.4 | GO:0042583 | chromaffin granule(GO:0042583) |
| 0.0 | 1.1 | GO:0035861 | site of double-strand break(GO:0035861) |
| 0.0 | 1.8 | GO:0030175 | filopodium(GO:0030175) |
| 0.0 | 0.8 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.0 | 0.1 | GO:0032019 | mitochondrial cloud(GO:0032019) |
| 0.0 | 0.2 | GO:0042645 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.0 | 0.2 | GO:0031464 | Cul4A-RING E3 ubiquitin ligase complex(GO:0031464) |
| 0.0 | 1.1 | GO:0031941 | filamentous actin(GO:0031941) |
| 0.0 | 0.4 | GO:0000780 | condensed nuclear chromosome kinetochore(GO:0000778) condensed nuclear chromosome, centromeric region(GO:0000780) |
| 0.0 | 2.4 | GO:0005770 | late endosome(GO:0005770) |
| 0.0 | 2.0 | GO:0005912 | adherens junction(GO:0005912) |
| 0.0 | 0.8 | GO:0032580 | Golgi cisterna membrane(GO:0032580) |
| 0.0 | 1.0 | GO:0030141 | secretory granule(GO:0030141) |
| 0.0 | 2.6 | GO:0000785 | chromatin(GO:0000785) |
| 0.0 | 1.3 | GO:0016324 | apical plasma membrane(GO:0016324) |
| 0.0 | 0.3 | GO:0055038 | recycling endosome membrane(GO:0055038) |
| 0.0 | 0.8 | GO:0031228 | integral component of Golgi membrane(GO:0030173) intrinsic component of Golgi membrane(GO:0031228) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 4.5 | GO:0008310 | single-stranded DNA 3'-5' exodeoxyribonuclease activity(GO:0008310) |
| 0.7 | 2.9 | GO:0016652 | NAD(P)+ transhydrogenase activity(GO:0008746) oxidoreductase activity, acting on NAD(P)H, NAD(P) as acceptor(GO:0016652) |
| 0.7 | 2.1 | GO:0000810 | diacylglycerol diphosphate phosphatase activity(GO:0000810) |
| 0.6 | 3.0 | GO:0090556 | phospholipid-translocating ATPase activity(GO:0004012) phosphatidylcholine-translocating ATPase activity(GO:0090554) phosphatidylserine-translocating ATPase activity(GO:0090556) |
| 0.6 | 2.4 | GO:0031005 | filamin binding(GO:0031005) |
| 0.5 | 1.6 | GO:0032574 | 5'-3' RNA helicase activity(GO:0032574) |
| 0.4 | 1.6 | GO:0101006 | protein histidine phosphatase activity(GO:0101006) |
| 0.4 | 8.4 | GO:0031492 | nucleosomal DNA binding(GO:0031492) |
| 0.4 | 3.3 | GO:0004985 | opioid receptor activity(GO:0004985) |
| 0.4 | 1.1 | GO:0071568 | UFM1 transferase activity(GO:0071568) |
| 0.3 | 1.0 | GO:0045545 | syndecan binding(GO:0045545) |
| 0.3 | 2.5 | GO:0005381 | iron ion transmembrane transporter activity(GO:0005381) |
| 0.2 | 0.9 | GO:0033149 | FFAT motif binding(GO:0033149) |
| 0.2 | 1.9 | GO:0004977 | melanocortin receptor activity(GO:0004977) |
| 0.2 | 1.0 | GO:0046592 | polyamine oxidase activity(GO:0046592) |
| 0.2 | 1.1 | GO:0008502 | melatonin receptor activity(GO:0008502) |
| 0.2 | 0.7 | GO:0004826 | phenylalanine-tRNA ligase activity(GO:0004826) |
| 0.2 | 2.6 | GO:0005324 | long-chain fatty acid transporter activity(GO:0005324) |
| 0.2 | 5.2 | GO:0016881 | acid-amino acid ligase activity(GO:0016881) |
| 0.2 | 0.5 | GO:0008397 | sterol 12-alpha-hydroxylase activity(GO:0008397) |
| 0.2 | 1.0 | GO:0004104 | cholinesterase activity(GO:0004104) |
| 0.2 | 1.3 | GO:0004634 | phosphopyruvate hydratase activity(GO:0004634) |
| 0.1 | 1.6 | GO:0004046 | aminoacylase activity(GO:0004046) |
| 0.1 | 0.7 | GO:0042030 | ATPase inhibitor activity(GO:0042030) |
| 0.1 | 3.7 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.1 | 4.4 | GO:0045505 | dynein intermediate chain binding(GO:0045505) |
| 0.1 | 0.5 | GO:0051916 | granulocyte colony-stimulating factor binding(GO:0051916) |
| 0.1 | 1.6 | GO:0051117 | ATPase binding(GO:0051117) |
| 0.1 | 0.7 | GO:1990380 | Lys48-specific deubiquitinase activity(GO:1990380) |
| 0.1 | 0.9 | GO:0034597 | phosphatidylinositol-4,5-bisphosphate 4-phosphatase activity(GO:0034597) |
| 0.1 | 0.5 | GO:0019863 | IgE binding(GO:0019863) immunoglobulin binding(GO:0019865) |
| 0.1 | 3.2 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.1 | 1.8 | GO:0003823 | antigen binding(GO:0003823) |
| 0.1 | 1.4 | GO:0001102 | RNA polymerase II activating transcription factor binding(GO:0001102) |
| 0.1 | 1.3 | GO:0005007 | fibroblast growth factor-activated receptor activity(GO:0005007) |
| 0.1 | 1.9 | GO:0001671 | ATPase activator activity(GO:0001671) |
| 0.1 | 1.9 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.1 | 1.1 | GO:0030247 | pattern binding(GO:0001871) polysaccharide binding(GO:0030247) |
| 0.1 | 1.4 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 0.1 | 3.5 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.1 | 2.1 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.1 | 0.5 | GO:0032217 | riboflavin transporter activity(GO:0032217) |
| 0.1 | 1.0 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) cysteine-type endopeptidase regulator activity involved in apoptotic process(GO:0043028) |
| 0.1 | 1.3 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.1 | 3.3 | GO:0019843 | rRNA binding(GO:0019843) |
| 0.1 | 1.7 | GO:0004438 | phosphatidylinositol-3-phosphatase activity(GO:0004438) |
| 0.1 | 0.6 | GO:0005522 | profilin binding(GO:0005522) |
| 0.1 | 3.2 | GO:0005484 | SNAP receptor activity(GO:0005484) |
| 0.1 | 1.8 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.1 | 0.6 | GO:0046934 | phosphatidylinositol-4,5-bisphosphate 3-kinase activity(GO:0046934) phosphatidylinositol-3,4-bisphosphate 5-kinase activity(GO:0052812) |
| 0.1 | 0.5 | GO:0050694 | galactosylceramide sulfotransferase activity(GO:0001733) galactose 3-O-sulfotransferase activity(GO:0050694) |
| 0.1 | 3.9 | GO:0008146 | sulfotransferase activity(GO:0008146) |
| 0.1 | 0.3 | GO:0001042 | RNA polymerase I core binding(GO:0001042) |
| 0.1 | 0.9 | GO:0045134 | guanosine-diphosphatase activity(GO:0004382) uridine-diphosphatase activity(GO:0045134) |
| 0.0 | 0.6 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.0 | 0.8 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.0 | 0.5 | GO:0005391 | sodium:potassium-exchanging ATPase activity(GO:0005391) potassium-transporting ATPase activity(GO:0008556) |
| 0.0 | 0.7 | GO:0005315 | inorganic phosphate transmembrane transporter activity(GO:0005315) |
| 0.0 | 1.1 | GO:0051378 | amine binding(GO:0043176) serotonin binding(GO:0051378) |
| 0.0 | 3.6 | GO:0003724 | RNA helicase activity(GO:0003724) |
| 0.0 | 1.6 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.0 | 0.7 | GO:0031682 | G-protein gamma-subunit binding(GO:0031682) |
| 0.0 | 0.5 | GO:0032036 | myosin heavy chain binding(GO:0032036) |
| 0.0 | 2.1 | GO:0005347 | ATP transmembrane transporter activity(GO:0005347) |
| 0.0 | 1.0 | GO:0004312 | fatty acid synthase activity(GO:0004312) |
| 0.0 | 0.6 | GO:0097199 | cysteine-type endopeptidase activity involved in apoptotic signaling pathway(GO:0097199) |
| 0.0 | 0.8 | GO:0070492 | oligosaccharide binding(GO:0070492) |
| 0.0 | 0.7 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) alpha-actinin binding(GO:0051393) |
| 0.0 | 0.5 | GO:0001591 | dopamine neurotransmitter receptor activity, coupled via Gi/Go(GO:0001591) |
| 0.0 | 0.6 | GO:0015923 | mannosidase activity(GO:0015923) |
| 0.0 | 1.3 | GO:0030145 | manganese ion binding(GO:0030145) |
| 0.0 | 1.7 | GO:0019209 | kinase activator activity(GO:0019209) protein kinase activator activity(GO:0030295) |
| 0.0 | 0.2 | GO:0070008 | serine-type exopeptidase activity(GO:0070008) |
| 0.0 | 0.8 | GO:0008378 | galactosyltransferase activity(GO:0008378) |
| 0.0 | 0.2 | GO:0050431 | transforming growth factor beta binding(GO:0050431) |
| 0.0 | 1.1 | GO:0071855 | neuropeptide receptor binding(GO:0071855) |
| 0.0 | 0.1 | GO:0003875 | ADP-ribosylarginine hydrolase activity(GO:0003875) |
| 0.0 | 0.3 | GO:0035259 | glucocorticoid receptor binding(GO:0035259) |
| 0.0 | 0.4 | GO:0004983 | neuropeptide Y receptor activity(GO:0004983) |
| 0.0 | 0.1 | GO:0004500 | dopamine beta-monooxygenase activity(GO:0004500) |
| 0.0 | 4.5 | GO:0020037 | heme binding(GO:0020037) |
| 0.0 | 0.8 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
| 0.0 | 0.5 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.0 | 17.8 | GO:0004674 | protein serine/threonine kinase activity(GO:0004674) |
| 0.0 | 0.1 | GO:0008048 | calcium sensitive guanylate cyclase activator activity(GO:0008048) |
| 0.0 | 0.2 | GO:1990404 | protein ADP-ribosylase activity(GO:1990404) |
| 0.0 | 0.5 | GO:0030374 | ligand-dependent nuclear receptor transcription coactivator activity(GO:0030374) |
| 0.0 | 0.8 | GO:0017124 | SH3 domain binding(GO:0017124) |
| 0.0 | 0.4 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.0 | 0.4 | GO:0019210 | protein kinase inhibitor activity(GO:0004860) kinase inhibitor activity(GO:0019210) |
| 0.0 | 0.5 | GO:0008381 | mechanically-gated ion channel activity(GO:0008381) mechanically gated channel activity(GO:0022833) |
| 0.0 | 0.4 | GO:0022842 | leak channel activity(GO:0022840) potassium ion leak channel activity(GO:0022841) narrow pore channel activity(GO:0022842) |
| 0.0 | 0.3 | GO:0031267 | small GTPase binding(GO:0031267) |
| 0.0 | 0.3 | GO:0004115 | 3',5'-cyclic-AMP phosphodiesterase activity(GO:0004115) |
| 0.0 | 1.3 | GO:0019955 | cytokine binding(GO:0019955) |
| 0.0 | 0.1 | GO:0052650 | NADP-retinol dehydrogenase activity(GO:0052650) |
| 0.0 | 0.4 | GO:0005242 | inward rectifier potassium channel activity(GO:0005242) |
| 0.0 | 0.6 | GO:0019905 | syntaxin binding(GO:0019905) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.4 | ST INTERLEUKIN 4 PATHWAY | Interleukin 4 (IL-4) Pathway |
| 0.1 | 1.0 | SA CASPASE CASCADE | Apoptosis is mediated by caspases, cysteine proteases arranged in a proteolytic cascade. |
| 0.0 | 1.9 | PID ERBB4 PATHWAY | ErbB4 signaling events |
| 0.0 | 1.6 | SIG CD40PATHWAYMAP | Genes related to CD40 signaling |
| 0.0 | 6.8 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 3.2 | PID PDGFRB PATHWAY | PDGFR-beta signaling pathway |
| 0.0 | 0.5 | PID IL27 PATHWAY | IL27-mediated signaling events |
| 0.0 | 0.1 | PID FAS PATHWAY | FAS (CD95) signaling pathway |
| 0.0 | 0.4 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.0 | 0.5 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.0 | 1.1 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
| 0.0 | 0.9 | PID MTOR 4PATHWAY | mTOR signaling pathway |
| 0.0 | 0.1 | PID LYMPH ANGIOGENESIS PATHWAY | VEGFR3 signaling in lymphatic endothelium |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 0.1 | 1.3 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.1 | 0.5 | REACTOME SEROTONIN RECEPTORS | Genes involved in Serotonin receptors |
| 0.1 | 1.0 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |
| 0.1 | 0.4 | REACTOME TANDEM PORE DOMAIN POTASSIUM CHANNELS | Genes involved in Tandem pore domain potassium channels |
| 0.1 | 1.0 | REACTOME PROTEOLYTIC CLEAVAGE OF SNARE COMPLEX PROTEINS | Genes involved in Proteolytic cleavage of SNARE complex proteins |
| 0.1 | 4.1 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.0 | 0.5 | REACTOME IL 6 SIGNALING | Genes involved in Interleukin-6 signaling |
| 0.0 | 1.4 | REACTOME G0 AND EARLY G1 | Genes involved in G0 and Early G1 |
| 0.0 | 0.7 | REACTOME CGMP EFFECTS | Genes involved in cGMP effects |
| 0.0 | 5.2 | REACTOME 3 UTR MEDIATED TRANSLATIONAL REGULATION | Genes involved in 3' -UTR-mediated translational regulation |
| 0.0 | 0.4 | REACTOME SOS MEDIATED SIGNALLING | Genes involved in SOS-mediated signalling |
| 0.0 | 2.0 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.0 | 2.2 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.0 | 1.7 | REACTOME TRNA AMINOACYLATION | Genes involved in tRNA Aminoacylation |
| 0.0 | 0.6 | REACTOME INHIBITION OF INSULIN SECRETION BY ADRENALINE NORADRENALINE | Genes involved in Inhibition of Insulin Secretion by Adrenaline/Noradrenaline |
| 0.0 | 0.4 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.0 | 1.0 | REACTOME NOD1 2 SIGNALING PATHWAY | Genes involved in NOD1/2 Signaling Pathway |
| 0.0 | 1.9 | REACTOME SIGNALING BY ERBB4 | Genes involved in Signaling by ERBB4 |
| 0.0 | 0.2 | REACTOME CREATION OF C4 AND C2 ACTIVATORS | Genes involved in Creation of C4 and C2 activators |
| 0.0 | 1.0 | REACTOME TRANSPORT TO THE GOLGI AND SUBSEQUENT MODIFICATION | Genes involved in Transport to the Golgi and subsequent modification |
| 0.0 | 0.8 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |
| 0.0 | 0.8 | REACTOME DESTABILIZATION OF MRNA BY AUF1 HNRNP D0 | Genes involved in Destabilization of mRNA by AUF1 (hnRNP D0) |
| 0.0 | 0.9 | REACTOME RNA POL II TRANSCRIPTION PRE INITIATION AND PROMOTER OPENING | Genes involved in RNA Polymerase II Transcription Pre-Initiation And Promoter Opening |