Project
PRJNA438478: RNAseq of wild type zebrafish germline
Navigation
Downloads
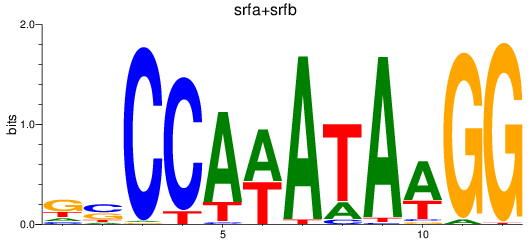
Results for srfa+srfb
Z-value: 0.31
Transcription factors associated with srfa+srfb
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
srfa
|
ENSDARG00000053918 | serum response factor a |
|
srfb
|
ENSDARG00000102867 | serum response factor b |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| srfb | dr11_v1_chr13_-_51922290_51922290 | -0.67 | 2.2e-03 | Click! |
| srfa | dr11_v1_chr22_+_35089031_35089031 | 0.59 | 1.0e-02 | Click! |
Activity profile of srfa+srfb motif
Sorted Z-values of srfa+srfb motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr14_-_21336385 | 2.34 |
ENSDART00000054460
|
egr1
|
early growth response 1 |
| chr15_-_551177 | 1.93 |
ENSDART00000066774
ENSDART00000154617 |
tagln
|
transgelin |
| chr23_+_42813415 | 1.32 |
ENSDART00000055577
|
myl9a
|
myosin, light chain 9a, regulatory |
| chr8_+_40628926 | 1.06 |
ENSDART00000163598
|
dusp2
|
dual specificity phosphatase 2 |
| chr5_+_37087583 | 1.02 |
ENSDART00000049900
|
tagln2
|
transgelin 2 |
| chr10_-_690072 | 0.95 |
ENSDART00000164871
ENSDART00000142833 |
glis3
|
GLIS family zinc finger 3 |
| chr23_-_45439903 | 0.92 |
ENSDART00000170729
|
NPNT (1 of many)
|
nephronectin |
| chr12_+_17100021 | 0.83 |
ENSDART00000177923
|
acta2
|
actin, alpha 2, smooth muscle, aorta |
| chr1_+_9994811 | 0.81 |
ENSDART00000143719
ENSDART00000110749 |
si:dkeyp-75b4.10
|
si:dkeyp-75b4.10 |
| chr18_+_36769758 | 0.72 |
ENSDART00000180375
ENSDART00000136463 ENSDART00000133487 ENSDART00000130206 |
fosb
|
FBJ murine osteosarcoma viral oncogene homolog B |
| chr10_-_29816467 | 0.62 |
ENSDART00000055913
|
hist2h2l
|
histone 2, H2, like |
| chr1_+_53945934 | 0.61 |
ENSDART00000052838
|
acta1a
|
actin, alpha 1a, skeletal muscle |
| chr11_-_27867024 | 0.60 |
ENSDART00000182136
ENSDART00000187587 |
eif4g3a
|
eukaryotic translation initiation factor 4 gamma, 3a |
| chr8_+_25902170 | 0.56 |
ENSDART00000193130
|
rhoab
|
ras homolog gene family, member Ab |
| chr11_-_472547 | 0.54 |
ENSDART00000005923
|
zgc:77375
|
zgc:77375 |
| chr2_-_24554416 | 0.54 |
ENSDART00000052061
|
cnn2
|
calponin 2 |
| chr20_-_14875308 | 0.54 |
ENSDART00000141290
|
dnm3a
|
dynamin 3a |
| chr17_-_20979077 | 0.49 |
ENSDART00000006676
|
phyhipla
|
phytanoyl-CoA 2-hydroxylase interacting protein-like a |
| chr17_-_43666166 | 0.49 |
ENSDART00000077990
|
egr2a
|
early growth response 2a |
| chr14_+_49251331 | 0.48 |
ENSDART00000148882
|
anxa6
|
annexin A6 |
| chr3_+_58167288 | 0.48 |
ENSDART00000155874
ENSDART00000010395 |
uqcrc2a
|
ubiquinol-cytochrome c reductase core protein 2a |
| chr1_+_50976975 | 0.46 |
ENSDART00000022290
ENSDART00000140982 |
mdh1aa
|
malate dehydrogenase 1Aa, NAD (soluble) |
| chr5_+_49744713 | 0.46 |
ENSDART00000133384
|
nr2f1a
|
nuclear receptor subfamily 2, group F, member 1a |
| chr14_+_17137023 | 0.46 |
ENSDART00000080712
|
slc43a3b
|
solute carrier family 43, member 3b |
| chr6_+_33897624 | 0.45 |
ENSDART00000166706
|
ccdc17
|
coiled-coil domain containing 17 |
| chr6_+_1787160 | 0.45 |
ENSDART00000113505
|
myl9b
|
myosin, light chain 9b, regulatory |
| chr18_+_36770166 | 0.44 |
ENSDART00000078151
|
fosb
|
FBJ murine osteosarcoma viral oncogene homolog B |
| chr1_+_52560549 | 0.43 |
ENSDART00000167514
|
abca1a
|
ATP-binding cassette, sub-family A (ABC1), member 1A |
| chr7_-_18168493 | 0.43 |
ENSDART00000127428
|
peli3
|
pellino E3 ubiquitin protein ligase family member 3 |
| chr20_-_46554440 | 0.42 |
ENSDART00000043298
ENSDART00000060680 |
fosab
|
v-fos FBJ murine osteosarcoma viral oncogene homolog Ab |
| chr5_+_9428876 | 0.42 |
ENSDART00000081791
|
ugt2a7
|
UDP glucuronosyltransferase 2 family, polypeptide A7 |
| chr4_-_17642168 | 0.41 |
ENSDART00000007030
|
klhl42
|
kelch-like family, member 42 |
| chr1_-_8651718 | 0.41 |
ENSDART00000133319
|
actb1
|
actin, beta 1 |
| chr22_-_15602593 | 0.40 |
ENSDART00000036075
|
tpm4a
|
tropomyosin 4a |
| chr19_-_42424599 | 0.39 |
ENSDART00000077042
|
zgc:153441
|
zgc:153441 |
| chr14_-_30747686 | 0.39 |
ENSDART00000008373
|
fosl1a
|
FOS-like antigen 1a |
| chr21_+_8533533 | 0.38 |
ENSDART00000077924
|
FO834888.1
|
|
| chr16_+_23816933 | 0.38 |
ENSDART00000183428
|
si:dkey-7f3.9
|
si:dkey-7f3.9 |
| chr14_+_48964628 | 0.38 |
ENSDART00000105427
|
mif
|
macrophage migration inhibitory factor |
| chr11_+_25459697 | 0.37 |
ENSDART00000161481
|
opn1sw2
|
opsin 1 (cone pigments), short-wave-sensitive 2 |
| chr11_+_16152316 | 0.36 |
ENSDART00000081054
|
tada3l
|
transcriptional adaptor 3 (NGG1 homolog, yeast)-like |
| chr15_-_39945036 | 0.35 |
ENSDART00000192481
|
msh5
|
mutS homolog 5 |
| chr15_-_15910226 | 0.35 |
ENSDART00000154219
|
synrg
|
synergin, gamma |
| chr20_+_6590220 | 0.34 |
ENSDART00000136567
|
tns3.2
|
tensin 3, tandem duplicate 2 |
| chr7_-_28549361 | 0.34 |
ENSDART00000173918
ENSDART00000054368 ENSDART00000113313 |
st5
|
suppression of tumorigenicity 5 |
| chr10_+_29816681 | 0.34 |
ENSDART00000100022
|
h2afx1
|
H2A histone family member X1 |
| chr7_+_5494565 | 0.32 |
ENSDART00000135271
|
MYADM
|
si:dkeyp-67a8.2 |
| chr22_+_35089031 | 0.32 |
ENSDART00000076040
|
srfa
|
serum response factor a |
| chr1_-_8652648 | 0.32 |
ENSDART00000138324
ENSDART00000141407 ENSDART00000054987 |
actb1
|
actin, beta 1 |
| chr12_-_20373058 | 0.32 |
ENSDART00000066382
|
aqp8a.1
|
aquaporin 8a, tandem duplicate 1 |
| chr7_+_29951997 | 0.30 |
ENSDART00000173453
|
tpma
|
alpha-tropomyosin |
| chr21_-_17603182 | 0.30 |
ENSDART00000020048
ENSDART00000177270 |
gsna
|
gelsolin a |
| chr9_-_202805 | 0.29 |
ENSDART00000182260
|
CABZ01067557.1
|
|
| chr11_+_6281647 | 0.28 |
ENSDART00000002459
|
ctns
|
cystinosin, lysosomal cystine transporter |
| chr3_-_40529782 | 0.27 |
ENSDART00000055194
ENSDART00000141737 |
actb2
|
actin, beta 2 |
| chr13_-_3155243 | 0.27 |
ENSDART00000139183
ENSDART00000050934 |
pkdcca
|
protein kinase domain containing, cytoplasmic a |
| chr22_-_15602760 | 0.26 |
ENSDART00000009054
|
tpm4a
|
tropomyosin 4a |
| chr18_-_17074155 | 0.26 |
ENSDART00000143304
|
tango6
|
transport and golgi organization 6 homolog (Drosophila) |
| chr16_-_30421275 | 0.25 |
ENSDART00000003752
|
cct3
|
chaperonin containing TCP1, subunit 3 (gamma) |
| chr23_-_44207349 | 0.25 |
ENSDART00000186276
|
zgc:158659
|
zgc:158659 |
| chr1_+_57331813 | 0.25 |
ENSDART00000152440
ENSDART00000062841 |
epn3b
|
epsin 3b |
| chr3_-_40945710 | 0.24 |
ENSDART00000138719
ENSDART00000102416 |
cyp3c4
|
cytochrome P450, family 3, subfamily C, polypeptide 4 |
| chr12_+_35654749 | 0.24 |
ENSDART00000169889
ENSDART00000167873 |
baiap2b
|
BAI1-associated protein 2b |
| chr6_+_58915889 | 0.24 |
ENSDART00000083628
|
ddit3
|
DNA-damage-inducible transcript 3 |
| chr3_-_35602233 | 0.23 |
ENSDART00000055269
|
gng13b
|
guanine nucleotide binding protein (G protein), gamma 13b |
| chr22_-_36856405 | 0.23 |
ENSDART00000029588
|
kng1
|
kininogen 1 |
| chr5_+_37785152 | 0.22 |
ENSDART00000053511
ENSDART00000189812 |
myo1ca
|
myosin Ic, paralog a |
| chr3_-_32898626 | 0.21 |
ENSDART00000103201
|
kat7a
|
K(lysine) acetyltransferase 7a |
| chr1_+_55293424 | 0.20 |
ENSDART00000152464
|
zgc:172106
|
zgc:172106 |
| chr23_-_35064785 | 0.20 |
ENSDART00000172240
|
BX294434.1
|
|
| chr16_-_12496632 | 0.19 |
ENSDART00000019941
|
slc2a3b
|
solute carrier family 2 (facilitated glucose transporter), member 3b |
| chr12_+_27129659 | 0.19 |
ENSDART00000076161
|
hoxb5b
|
homeobox B5b |
| chr6_+_41191482 | 0.19 |
ENSDART00000000877
|
opn1mw3
|
opsin 1 (cone pigments), medium-wave-sensitive, 3 |
| chr3_-_2592350 | 0.19 |
ENSDART00000192325
|
si:dkey-217f16.5
|
si:dkey-217f16.5 |
| chr11_-_16152400 | 0.18 |
ENSDART00000123665
|
arpc4l
|
actin related protein 2/3 complex, subunit 4, like |
| chr20_-_42702832 | 0.18 |
ENSDART00000134689
ENSDART00000045816 |
plg
|
plasminogen |
| chr9_+_34232503 | 0.18 |
ENSDART00000132836
|
nxpe3
|
neurexophilin and PC-esterase domain family, member 3 |
| chr20_-_29420713 | 0.17 |
ENSDART00000147464
|
ryr3
|
ryanodine receptor 3 |
| chr2_+_38264964 | 0.17 |
ENSDART00000182068
|
dhrs1
|
dehydrogenase/reductase (SDR family) member 1 |
| chr14_-_25309058 | 0.17 |
ENSDART00000159569
|
htr4
|
5-hydroxytryptamine receptor 4 |
| chr3_+_23707691 | 0.17 |
ENSDART00000025449
|
hoxb5a
|
homeobox B5a |
| chr17_+_45454943 | 0.16 |
ENSDART00000074838
|
kcnk3b
|
potassium channel, subfamily K, member 3b |
| chr6_-_45905746 | 0.16 |
ENSDART00000025428
|
epha2a
|
eph receptor A2 a |
| chr18_-_46258612 | 0.16 |
ENSDART00000153930
|
si:dkey-244a7.1
|
si:dkey-244a7.1 |
| chr4_-_22310956 | 0.16 |
ENSDART00000162585
|
hcls1
|
hematopoietic cell-specific Lyn substrate 1 |
| chr17_-_25326296 | 0.16 |
ENSDART00000168822
|
pabpc4
|
poly(A) binding protein, cytoplasmic 4 (inducible form) |
| chr24_+_31277360 | 0.15 |
ENSDART00000165993
|
f3a
|
coagulation factor IIIa |
| chr12_-_15584479 | 0.15 |
ENSDART00000150134
|
acbd4
|
acyl-CoA binding domain containing 4 |
| chr15_-_42760110 | 0.14 |
ENSDART00000152490
|
si:ch211-181d7.3
|
si:ch211-181d7.3 |
| chr8_+_10561922 | 0.14 |
ENSDART00000133348
|
fam19a5l
|
family with sequence similarity 19 (chemokine (C-C motif)-like), member A5-like |
| chr11_-_16152105 | 0.14 |
ENSDART00000081062
|
arpc4l
|
actin related protein 2/3 complex, subunit 4, like |
| chr1_-_14332283 | 0.14 |
ENSDART00000090025
|
wfs1a
|
Wolfram syndrome 1a (wolframin) |
| chr18_+_45504362 | 0.14 |
ENSDART00000140089
|
cngb1a
|
cyclic nucleotide gated channel beta 1a |
| chr12_+_19362335 | 0.13 |
ENSDART00000041711
|
gspt1
|
G1 to S phase transition 1 |
| chr21_-_25555355 | 0.13 |
ENSDART00000144228
|
si:dkey-17e16.9
|
si:dkey-17e16.9 |
| chr1_-_37383741 | 0.13 |
ENSDART00000193155
ENSDART00000191887 ENSDART00000189077 |
scpp1
|
secretory calcium-binding phosphoprotein 1 |
| chr1_+_45754868 | 0.12 |
ENSDART00000084512
|
pkn1a
|
protein kinase N1a |
| chr15_+_2534740 | 0.12 |
ENSDART00000138469
|
cux1b
|
cut-like homeobox 1b |
| chr12_-_29624638 | 0.12 |
ENSDART00000126744
|
nrg3b
|
neuregulin 3b |
| chr1_-_6494384 | 0.12 |
ENSDART00000109356
|
klf7a
|
Kruppel-like factor 7a |
| chr25_+_1591964 | 0.12 |
ENSDART00000093277
|
ppm1h
|
protein phosphatase, Mg2+/Mn2+ dependent, 1H |
| chr13_-_40398556 | 0.11 |
ENSDART00000191921
|
nkx3.3
|
NK3 homeobox 3 |
| chr11_+_40657612 | 0.11 |
ENSDART00000183271
|
slc45a1
|
solute carrier family 45, member 1 |
| chr4_+_5180650 | 0.11 |
ENSDART00000067390
|
fgf6b
|
fibroblast growth factor 6b |
| chr23_-_44466257 | 0.11 |
ENSDART00000150126
|
si:ch1073-228j22.2
|
si:ch1073-228j22.2 |
| chr21_+_32332616 | 0.11 |
ENSDART00000154105
|
si:ch211-247j9.1
|
si:ch211-247j9.1 |
| chr3_-_56896702 | 0.11 |
ENSDART00000023265
|
ush1ga
|
Usher syndrome 1Ga (autosomal recessive) |
| chr16_-_31754102 | 0.11 |
ENSDART00000185043
|
ptpn6
|
protein tyrosine phosphatase, non-receptor type 6 |
| chr20_-_47732703 | 0.11 |
ENSDART00000193975
|
tfap2d
|
transcription factor AP-2 delta (activating enhancer binding protein 2 delta) |
| chr15_-_23761580 | 0.11 |
ENSDART00000137918
|
bbc3
|
BCL2 binding component 3 |
| chr21_+_6780340 | 0.11 |
ENSDART00000139493
ENSDART00000140478 |
olfm1b
|
olfactomedin 1b |
| chr20_-_48485354 | 0.11 |
ENSDART00000124040
ENSDART00000148437 |
insm1a
|
insulinoma-associated 1a |
| chr3_+_1211242 | 0.11 |
ENSDART00000171287
ENSDART00000165769 |
poldip3
|
polymerase (DNA-directed), delta interacting protein 3 |
| chr11_+_37178271 | 0.10 |
ENSDART00000161771
|
itih3b
|
inter-alpha-trypsin inhibitor heavy chain 3b |
| chr20_-_31018569 | 0.10 |
ENSDART00000136039
|
fndc1
|
fibronectin type III domain containing 1 |
| chr23_+_24272421 | 0.10 |
ENSDART00000029974
|
clcnk
|
chloride channel K |
| chr7_+_29952169 | 0.09 |
ENSDART00000173540
ENSDART00000173940 ENSDART00000173906 ENSDART00000173772 ENSDART00000173506 ENSDART00000039657 |
tpma
|
alpha-tropomyosin |
| chr1_+_8492379 | 0.09 |
ENSDART00000184518
|
myo15ab
|
myosin XVAb |
| chr10_-_22057001 | 0.09 |
ENSDART00000016575
|
tlx3b
|
T cell leukemia homeobox 3b |
| chr22_+_33362552 | 0.09 |
ENSDART00000101580
|
nicn1
|
nicolin 1 |
| chr13_+_29238850 | 0.09 |
ENSDART00000026000
|
myofl
|
myoferlin like |
| chr4_-_6623645 | 0.09 |
ENSDART00000060861
|
foxp2
|
forkhead box P2 |
| chr13_+_24279021 | 0.08 |
ENSDART00000058629
|
acta1b
|
actin, alpha 1b, skeletal muscle |
| chr5_-_32292965 | 0.08 |
ENSDART00000183522
ENSDART00000131983 |
myhz1.2
|
myosin, heavy polypeptide 1.2, skeletal muscle |
| chr18_-_41650648 | 0.08 |
ENSDART00000024087
|
fzd9b
|
frizzled class receptor 9b |
| chr9_-_44948488 | 0.08 |
ENSDART00000059228
|
vil1
|
villin 1 |
| chr23_+_45579497 | 0.08 |
ENSDART00000110381
|
egr4
|
early growth response 4 |
| chr18_-_26675699 | 0.07 |
ENSDART00000113280
|
FRMD5
|
si:ch211-69m14.1 |
| chr10_+_8101729 | 0.07 |
ENSDART00000138875
ENSDART00000123447 ENSDART00000185333 |
pstpip2
|
proline-serine-threonine phosphatase interacting protein 2 |
| chr22_-_29336268 | 0.07 |
ENSDART00000132776
ENSDART00000186351 ENSDART00000121599 |
pdgfba
|
platelet-derived growth factor beta polypeptide a |
| chr21_+_20950161 | 0.07 |
ENSDART00000079698
|
htr1ab
|
5-hydroxytryptamine (serotonin) receptor 1A b |
| chr3_+_54761569 | 0.07 |
ENSDART00000135913
ENSDART00000180983 |
si:ch211-74m13.1
|
si:ch211-74m13.1 |
| chr20_+_9211237 | 0.07 |
ENSDART00000139527
|
si:ch211-59d15.4
|
si:ch211-59d15.4 |
| chr17_+_18031899 | 0.07 |
ENSDART00000022758
|
setd3
|
SET domain containing 3 |
| chr12_+_19362547 | 0.07 |
ENSDART00000180148
|
gspt1
|
G1 to S phase transition 1 |
| chr23_-_37514493 | 0.06 |
ENSDART00000083267
|
dnajc16
|
DnaJ (Hsp40) homolog, subfamily C, member 16 |
| chr19_+_19759577 | 0.06 |
ENSDART00000169480
|
hoxa5a
|
homeobox A5a |
| chr10_-_7892192 | 0.06 |
ENSDART00000145480
|
smtna
|
smoothelin a |
| chr2_+_10771787 | 0.06 |
ENSDART00000187782
|
gfi1aa
|
growth factor independent 1A transcription repressor a |
| chr21_+_20949976 | 0.06 |
ENSDART00000135342
|
htr1ab
|
5-hydroxytryptamine (serotonin) receptor 1A b |
| chr20_-_2949028 | 0.06 |
ENSDART00000104667
ENSDART00000193151 ENSDART00000131946 |
cdk19
|
cyclin-dependent kinase 19 |
| chr8_+_50727220 | 0.06 |
ENSDART00000127062
|
egr3
|
early growth response 3 |
| chr16_+_28383758 | 0.06 |
ENSDART00000059038
ENSDART00000141061 |
itga8
|
integrin, alpha 8 |
| chr9_+_6587364 | 0.06 |
ENSDART00000122279
|
fhl2a
|
four and a half LIM domains 2a |
| chr9_+_6587056 | 0.06 |
ENSDART00000193421
|
fhl2a
|
four and a half LIM domains 2a |
| chr2_-_24270062 | 0.06 |
ENSDART00000192445
|
myh7
|
myosin heavy chain 7 |
| chr22_-_17458070 | 0.06 |
ENSDART00000139658
|
si:ch211-197g15.10
|
si:ch211-197g15.10 |
| chr20_+_20186706 | 0.05 |
ENSDART00000002507
|
rhoj
|
ras homolog family member J |
| chr17_-_42213285 | 0.05 |
ENSDART00000140549
|
nkx2.2a
|
NK2 homeobox 2a |
| chr4_+_33461796 | 0.05 |
ENSDART00000150445
|
si:dkey-247i3.1
|
si:dkey-247i3.1 |
| chr2_-_24269911 | 0.05 |
ENSDART00000099532
|
myh7
|
myosin heavy chain 7 |
| chr1_-_50791280 | 0.05 |
ENSDART00000181224
|
CABZ01031870.1
|
|
| chr4_-_49133107 | 0.05 |
ENSDART00000150806
|
znf1146
|
zinc finger protein 1146 |
| chr2_+_30916188 | 0.05 |
ENSDART00000137012
|
myom1a
|
myomesin 1a (skelemin) |
| chr13_-_9598320 | 0.05 |
ENSDART00000184613
|
cpxm1a
|
carboxypeptidase X (M14 family), member 1a |
| chr8_+_8937723 | 0.05 |
ENSDART00000145970
|
si:dkey-83k24.5
|
si:dkey-83k24.5 |
| chr21_+_10756154 | 0.05 |
ENSDART00000074833
|
rx3
|
retinal homeobox gene 3 |
| chr3_-_18410968 | 0.05 |
ENSDART00000041842
|
ndufb10
|
NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 10 |
| chr13_+_29773153 | 0.05 |
ENSDART00000144443
ENSDART00000133796 ENSDART00000141310 ENSDART00000139782 |
pax2a
|
paired box 2a |
| chr21_-_19906786 | 0.05 |
ENSDART00000136084
|
mfhas1
|
malignant fibrous histiocytoma amplified sequence 1 |
| chr2_+_31804582 | 0.05 |
ENSDART00000086646
|
rnf182
|
ring finger protein 182 |
| chr3_+_32553714 | 0.05 |
ENSDART00000165638
|
pax10
|
paired box 10 |
| chr15_+_7064819 | 0.04 |
ENSDART00000155268
|
pik3cb
|
phosphatidylinositol-4,5-bisphosphate 3-kinase, catalytic subunit beta |
| chr11_-_20987378 | 0.04 |
ENSDART00000110140
|
taf4a
|
TAF4A RNA polymerase II, TATA box binding protein (TBP)-associated factor |
| chr4_-_49582758 | 0.04 |
ENSDART00000180834
ENSDART00000187608 |
si:dkey-159n16.2
|
si:dkey-159n16.2 |
| chr7_+_56651759 | 0.04 |
ENSDART00000073600
|
kcng4b
|
potassium voltage-gated channel, subfamily G, member 4b |
| chr22_-_382955 | 0.04 |
ENSDART00000082406
|
ora3
|
olfactory receptor class A related 3 |
| chr6_+_41200500 | 0.04 |
ENSDART00000000678
|
opn1mw4
|
opsin 1 (cone pigments), medium-wave-sensitive, 4 |
| chr13_-_28688104 | 0.03 |
ENSDART00000133827
|
pcgf6
|
polycomb group ring finger 6 |
| chr10_-_7857494 | 0.03 |
ENSDART00000143215
|
inpp5ja
|
inositol polyphosphate-5-phosphatase Ja |
| chr11_-_34097418 | 0.03 |
ENSDART00000028301
ENSDART00000138579 |
tdo2b
|
tryptophan 2,3-dioxygenase b |
| chr20_-_40750953 | 0.03 |
ENSDART00000061256
|
cx28.9
|
connexin 28.9 |
| chr19_-_20403507 | 0.03 |
ENSDART00000052603
ENSDART00000137590 |
dazl
|
deleted in azoospermia-like |
| chr21_+_30043054 | 0.03 |
ENSDART00000065448
|
fabp6
|
fatty acid binding protein 6, ileal (gastrotropin) |
| chr12_-_26064105 | 0.03 |
ENSDART00000168825
|
ldb3b
|
LIM domain binding 3b |
| chr16_-_31675669 | 0.03 |
ENSDART00000168848
ENSDART00000158331 |
c1r
|
complement component 1, r subcomponent |
| chr10_-_373575 | 0.03 |
ENSDART00000114487
|
DMPK
|
DM1 protein kinase |
| chr12_-_26064480 | 0.02 |
ENSDART00000158215
ENSDART00000171206 ENSDART00000171212 ENSDART00000182956 ENSDART00000186779 |
ldb3b
|
LIM domain binding 3b |
| chr20_+_35279968 | 0.02 |
ENSDART00000168216
ENSDART00000153332 |
fam49a
|
family with sequence similarity 49, member A |
| chr12_-_35054354 | 0.02 |
ENSDART00000075351
|
zgc:112285
|
zgc:112285 |
| chr3_+_3046909 | 0.02 |
ENSDART00000122435
|
BX004816.2
|
|
| chr5_+_38462121 | 0.02 |
ENSDART00000144425
|
gltpd2
|
glycolipid transfer protein domain containing 2 |
| chr19_+_43715911 | 0.02 |
ENSDART00000006344
|
cap1
|
CAP, adenylate cyclase-associated protein 1 (yeast) |
| chr13_-_25303303 | 0.02 |
ENSDART00000133153
|
plaua
|
plasminogen activator, urokinase a |
| chr11_-_6068375 | 0.02 |
ENSDART00000167672
ENSDART00000122262 |
babam1
|
BRISC and BRCA1 A complex member 1 |
| chr8_-_11229523 | 0.02 |
ENSDART00000002164
|
unc45b
|
unc-45 myosin chaperone B |
| chr21_+_42930558 | 0.02 |
ENSDART00000135234
|
stk32a
|
serine/threonine kinase 32A |
| chr14_-_10492072 | 0.02 |
ENSDART00000182525
|
lpar4
|
lysophosphatidic acid receptor 4 |
| chr10_-_15910974 | 0.01 |
ENSDART00000148169
|
pip5k1ba
|
phosphatidylinositol-4-phosphate 5-kinase, type I, beta a |
| chr1_+_53714734 | 0.01 |
ENSDART00000114689
|
pus10
|
pseudouridylate synthase 10 |
| chr23_+_36118738 | 0.01 |
ENSDART00000139319
|
hoxc5a
|
homeobox C5a |
| chr20_+_1996202 | 0.01 |
ENSDART00000184143
|
CABZ01092781.1
|
|
| chr16_+_12632428 | 0.01 |
ENSDART00000184600
ENSDART00000180537 |
nat14
|
N-acetyltransferase 14 (GCN5-related, putative) |
| chr6_-_13498745 | 0.01 |
ENSDART00000027684
ENSDART00000189438 |
mylkb
|
myosin light chain kinase b |
| chr3_+_45401472 | 0.00 |
ENSDART00000156693
|
baiap3
|
BAI1-associated protein 3 |
| chr12_+_42436328 | 0.00 |
ENSDART00000167324
|
ebf3a
|
early B cell factor 3a |
| chr8_+_36942262 | 0.00 |
ENSDART00000188173
|
iqsec2b
|
IQ motif and Sec7 domain 2b |
| chr5_-_32309129 | 0.00 |
ENSDART00000123003
|
myhz1.1
|
myosin, heavy polypeptide 1.1, skeletal muscle |
| chr5_-_26330313 | 0.00 |
ENSDART00000148656
|
arvcfb
|
ARVCF, delta catenin family member b |
Network of associatons between targets according to the STRING database.
First level regulatory network of srfa+srfb
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.4 | GO:0090131 | mesenchyme migration(GO:0090131) |
| 0.1 | 0.5 | GO:0003413 | plasma membrane repair(GO:0001778) chondrocyte differentiation involved in endochondral bone morphogenesis(GO:0003413) growth plate cartilage chondrocyte differentiation(GO:0003418) |
| 0.1 | 0.4 | GO:0010874 | regulation of cholesterol efflux(GO:0010874) |
| 0.1 | 0.5 | GO:0006107 | oxaloacetate metabolic process(GO:0006107) |
| 0.1 | 0.4 | GO:0008592 | regulation of Toll signaling pathway(GO:0008592) |
| 0.1 | 0.6 | GO:1902766 | skeletal muscle satellite cell migration(GO:1902766) |
| 0.1 | 0.4 | GO:0055011 | atrial cardiac muscle cell differentiation(GO:0055011) |
| 0.1 | 0.8 | GO:0035970 | peptidyl-threonine dephosphorylation(GO:0035970) |
| 0.1 | 0.2 | GO:0042730 | fibrinolysis(GO:0042730) |
| 0.1 | 0.2 | GO:2001244 | positive regulation of intrinsic apoptotic signaling pathway(GO:2001244) |
| 0.1 | 0.2 | GO:0030195 | negative regulation of blood coagulation(GO:0030195) negative regulation of wound healing(GO:0061045) negative regulation of hemostasis(GO:1900047) |
| 0.1 | 1.4 | GO:1990798 | pancreas regeneration(GO:1990798) |
| 0.1 | 2.3 | GO:0060059 | embryonic retina morphogenesis in camera-type eye(GO:0060059) |
| 0.0 | 0.2 | GO:0002184 | cytoplasmic translational termination(GO:0002184) |
| 0.0 | 0.1 | GO:0090199 | regulation of release of cytochrome c from mitochondria(GO:0090199) positive regulation of release of cytochrome c from mitochondria(GO:0090200) |
| 0.0 | 0.4 | GO:0000710 | meiotic mismatch repair(GO:0000710) |
| 0.0 | 0.4 | GO:0061026 | cardiac muscle tissue regeneration(GO:0061026) |
| 0.0 | 0.3 | GO:0070293 | renal absorption(GO:0070293) |
| 0.0 | 0.2 | GO:2000251 | positive regulation of actin cytoskeleton reorganization(GO:2000251) |
| 0.0 | 0.5 | GO:0098884 | postsynaptic neurotransmitter receptor internalization(GO:0098884) |
| 0.0 | 0.8 | GO:0055003 | cardiac myofibril assembly(GO:0055003) |
| 0.0 | 0.3 | GO:0050909 | sensory perception of taste(GO:0050909) |
| 0.0 | 0.2 | GO:0010002 | cardioblast differentiation(GO:0010002) |
| 0.0 | 0.5 | GO:0043535 | regulation of blood vessel endothelial cell migration(GO:0043535) |
| 0.0 | 0.1 | GO:0072673 | lamellipodium morphogenesis(GO:0072673) regulation of lamellipodium morphogenesis(GO:2000392) |
| 0.0 | 0.1 | GO:0044319 | wound healing, spreading of cells(GO:0044319) epiboly involved in wound healing(GO:0090505) |
| 0.0 | 0.1 | GO:0021530 | spinal cord oligodendrocyte cell fate specification(GO:0021530) |
| 0.0 | 0.0 | GO:0060898 | eye field cell fate commitment involved in camera-type eye formation(GO:0060898) |
| 0.0 | 0.3 | GO:0036092 | phosphatidylinositol-3-phosphate biosynthetic process(GO:0036092) |
| 0.0 | 0.1 | GO:0060221 | retinal rod cell differentiation(GO:0060221) |
| 0.0 | 0.5 | GO:0060117 | auditory receptor cell development(GO:0060117) |
| 0.0 | 0.3 | GO:0007198 | adenylate cyclase-inhibiting serotonin receptor signaling pathway(GO:0007198) |
| 0.0 | 0.0 | GO:0055057 | neuroblast division(GO:0055057) |
| 0.0 | 0.6 | GO:0018298 | protein-chromophore linkage(GO:0018298) |
| 0.0 | 0.4 | GO:0043967 | histone H4 acetylation(GO:0043967) |
| 0.0 | 0.1 | GO:0051122 | lipoxygenase pathway(GO:0019372) linoleic acid metabolic process(GO:0043651) hepoxilin metabolic process(GO:0051121) hepoxilin biosynthetic process(GO:0051122) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.0 | GO:0097433 | dense body(GO:0097433) |
| 0.1 | 1.5 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.1 | 0.2 | GO:0060171 | stereocilium membrane(GO:0060171) |
| 0.0 | 0.4 | GO:0005671 | Ada2/Gcn5/Ada3 transcription activator complex(GO:0005671) |
| 0.0 | 0.2 | GO:0018444 | translation release factor complex(GO:0018444) |
| 0.0 | 0.6 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.0 | 0.5 | GO:0098844 | postsynaptic endocytic zone membrane(GO:0098844) |
| 0.0 | 0.3 | GO:0005943 | phosphatidylinositol 3-kinase complex, class IA(GO:0005943) |
| 0.0 | 0.3 | GO:0005832 | chaperonin-containing T-complex(GO:0005832) |
| 0.0 | 0.3 | GO:0030130 | trans-Golgi network transport vesicle membrane(GO:0012510) clathrin coat of trans-Golgi network vesicle(GO:0030130) |
| 0.0 | 0.1 | GO:0032433 | filopodium tip(GO:0032433) |
| 0.0 | 0.6 | GO:0032154 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.0 | 0.6 | GO:0001750 | photoreceptor outer segment(GO:0001750) |
| 0.0 | 0.5 | GO:0048770 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.0 | 1.2 | GO:0005884 | actin filament(GO:0005884) |
| 0.0 | 0.2 | GO:0030125 | clathrin vesicle coat(GO:0030125) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.0 | GO:0098973 | structural constituent of postsynaptic actin cytoskeleton(GO:0098973) |
| 0.1 | 2.3 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.1 | 0.5 | GO:0030060 | L-malate dehydrogenase activity(GO:0030060) |
| 0.1 | 0.3 | GO:0000099 | sulfur amino acid transmembrane transporter activity(GO:0000099) |
| 0.1 | 0.4 | GO:0090556 | phospholipid-translocating ATPase activity(GO:0004012) phosphatidylcholine-translocating ATPase activity(GO:0090554) phosphatidylserine-translocating ATPase activity(GO:0090556) |
| 0.1 | 0.8 | GO:0008330 | protein tyrosine/threonine phosphatase activity(GO:0008330) |
| 0.1 | 0.2 | GO:0005252 | open rectifier potassium channel activity(GO:0005252) |
| 0.0 | 0.3 | GO:0015250 | water channel activity(GO:0015250) |
| 0.0 | 0.3 | GO:0052812 | phosphatidylinositol-4,5-bisphosphate 3-kinase activity(GO:0046934) phosphatidylinositol-3,4-bisphosphate 5-kinase activity(GO:0052812) |
| 0.0 | 0.6 | GO:0000339 | RNA cap binding(GO:0000339) |
| 0.0 | 0.4 | GO:0030983 | mismatched DNA binding(GO:0030983) |
| 0.0 | 0.2 | GO:0008079 | translation release factor activity(GO:0003747) translation termination factor activity(GO:0008079) |
| 0.0 | 0.1 | GO:0005351 | sugar:proton symporter activity(GO:0005351) cation:sugar symporter activity(GO:0005402) sucrose:proton symporter activity(GO:0008506) sucrose transmembrane transporter activity(GO:0008515) disaccharide transmembrane transporter activity(GO:0015154) oligosaccharide transmembrane transporter activity(GO:0015157) |
| 0.0 | 0.1 | GO:0005219 | ryanodine-sensitive calcium-release channel activity(GO:0005219) calcium-induced calcium release activity(GO:0048763) |
| 0.0 | 0.3 | GO:0043176 | amine binding(GO:0043176) serotonin binding(GO:0051378) |
| 0.0 | 0.6 | GO:0008020 | G-protein coupled photoreceptor activity(GO:0008020) |
| 0.0 | 0.0 | GO:0004833 | tryptophan 2,3-dioxygenase activity(GO:0004833) |
| 0.0 | 0.2 | GO:0008266 | poly(U) RNA binding(GO:0008266) |
| 0.0 | 0.2 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.0 | 0.1 | GO:0043855 | intracellular cyclic nucleotide activated cation channel activity(GO:0005221) intracellular cAMP activated cation channel activity(GO:0005222) intracellular cGMP activated cation channel activity(GO:0005223) cyclic nucleotide-gated ion channel activity(GO:0043855) |
| 0.0 | 0.5 | GO:0001786 | phosphatidylserine binding(GO:0001786) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.5 | PID NFAT TFPATHWAY | Calcineurin-regulated NFAT-dependent transcription in lymphocytes |
| 0.0 | 2.8 | PID PDGFRB PATHWAY | PDGFR-beta signaling pathway |
| 0.0 | 1.2 | PID AP1 PATHWAY | AP-1 transcription factor network |
| 0.0 | 0.2 | PID SYNDECAN 2 PATHWAY | Syndecan-2-mediated signaling events |
| 0.0 | 0.2 | PID SYNDECAN 4 PATHWAY | Syndecan-4-mediated signaling events |
| 0.0 | 0.1 | PID ERBB1 RECEPTOR PROXIMAL PATHWAY | EGF receptor (ErbB1) signaling pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.4 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 0.0 | 0.2 | REACTOME SEROTONIN RECEPTORS | Genes involved in Serotonin receptors |
| 0.0 | 0.6 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.0 | 0.2 | REACTOME ACTIVATION OF CHAPERONE GENES BY ATF6 ALPHA | Genes involved in Activation of Chaperone Genes by ATF6-alpha |
| 0.0 | 0.2 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.0 | 0.2 | REACTOME INTRINSIC PATHWAY | Genes involved in Intrinsic Pathway |
| 0.0 | 0.3 | REACTOME FORMATION OF TUBULIN FOLDING INTERMEDIATES BY CCT TRIC | Genes involved in Formation of tubulin folding intermediates by CCT/TriC |
| 0.0 | 0.1 | REACTOME ACTIVATION OF BH3 ONLY PROTEINS | Genes involved in Activation of BH3-only proteins |