Project
PRJNA438478: RNAseq of wild type zebrafish germline
Navigation
Downloads
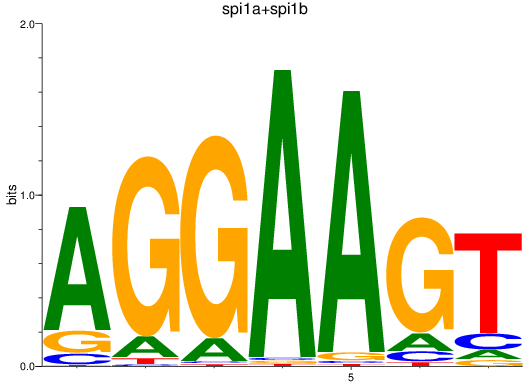
Results for spi1a+spi1b
Z-value: 3.26
Transcription factors associated with spi1a+spi1b
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
spi1b
|
ENSDARG00000000767 | Spi-1 proto-oncogene b |
|
spi1a
|
ENSDARG00000067797 | Spi-1 proto-oncogene a |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| spi1b | dr11_v1_chr7_-_32659048_32659048 | -0.84 | 1.3e-05 | Click! |
| spi1a | dr11_v1_chr25_+_35553542_35553542 | -0.77 | 1.8e-04 | Click! |
Activity profile of spi1a+spi1b motif
Sorted Z-values of spi1a+spi1b motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr15_+_12435975 | 11.38 |
ENSDART00000168011
|
tmprss4a
|
transmembrane protease, serine 4a |
| chr15_+_12436220 | 11.25 |
ENSDART00000169894
|
tmprss4a
|
transmembrane protease, serine 4a |
| chr18_+_15937610 | 8.31 |
ENSDART00000061134
ENSDART00000154505 |
itpr2
|
inositol 1,4,5-trisphosphate receptor, type 2 |
| chr7_-_26603743 | 8.19 |
ENSDART00000099003
|
plscr3b
|
phospholipid scramblase 3b |
| chr16_-_24832038 | 8.05 |
ENSDART00000153731
|
si:dkey-79d12.5
|
si:dkey-79d12.5 |
| chr12_-_17686404 | 7.47 |
ENSDART00000079065
|
ccz1
|
CCZ1 homolog, vacuolar protein trafficking and biogenesis associated |
| chr8_+_52442622 | 6.96 |
ENSDART00000012758
|
zgc:77112
|
zgc:77112 |
| chr24_-_31904924 | 6.93 |
ENSDART00000156060
ENSDART00000129741 ENSDART00000154276 |
si:ch73-78o10.1
|
si:ch73-78o10.1 |
| chr8_-_32385989 | 6.74 |
ENSDART00000143716
ENSDART00000098850 |
lipg
|
lipase, endothelial |
| chr13_-_17943135 | 6.52 |
ENSDART00000176027
|
march8
|
membrane-associated ring finger (C3HC4) 8 |
| chr14_-_16807206 | 6.44 |
ENSDART00000157957
|
tcirg1b
|
T cell immune regulator 1, ATPase H+ transporting V0 subunit a3b |
| chr1_+_27977297 | 6.43 |
ENSDART00000180692
ENSDART00000166819 |
sugt1
|
SGT1 homolog, MIS12 kinetochore complex assembly cochaperone |
| chr9_+_24088062 | 6.34 |
ENSDART00000126198
|
lrrfip1a
|
leucine rich repeat (in FLII) interacting protein 1a |
| chr8_+_52442785 | 6.31 |
ENSDART00000189958
|
zgc:77112
|
zgc:77112 |
| chr23_-_10177442 | 6.27 |
ENSDART00000144280
ENSDART00000129044 |
krt5
|
keratin 5 |
| chr21_-_4793686 | 6.21 |
ENSDART00000158232
|
notch1a
|
notch 1a |
| chr6_+_40563848 | 6.18 |
ENSDART00000154766
|
si:ch73-15b2.5
|
si:ch73-15b2.5 |
| chr19_-_43750659 | 6.14 |
ENSDART00000151309
|
ppt1
|
palmitoyl-protein thioesterase 1 (ceroid-lipofuscinosis, neuronal 1, infantile) |
| chr19_-_43750389 | 6.13 |
ENSDART00000147328
|
ppt1
|
palmitoyl-protein thioesterase 1 (ceroid-lipofuscinosis, neuronal 1, infantile) |
| chr18_-_25771553 | 6.05 |
ENSDART00000103046
|
zgc:162879
|
zgc:162879 |
| chr14_-_41467497 | 6.00 |
ENSDART00000181220
|
mid1ip1l
|
MID1 interacting protein 1, like |
| chr19_+_10661520 | 5.98 |
ENSDART00000091813
ENSDART00000165653 |
ago3b
|
argonaute RISC catalytic component 3b |
| chr6_-_39903393 | 5.91 |
ENSDART00000085945
|
tsfm
|
Ts translation elongation factor, mitochondrial |
| chr15_+_6459847 | 5.36 |
ENSDART00000157250
ENSDART00000065824 |
bace2
|
beta-site APP-cleaving enzyme 2 |
| chr16_-_42770064 | 5.26 |
ENSDART00000112879
|
slc4a7
|
solute carrier family 4, sodium bicarbonate cotransporter, member 7 |
| chr7_-_26601307 | 5.23 |
ENSDART00000188934
|
plscr3b
|
phospholipid scramblase 3b |
| chr16_+_29509133 | 5.20 |
ENSDART00000112116
|
ctss2.1
|
cathepsin S, ortholog2, tandem duplicate 1 |
| chr13_+_8840772 | 5.20 |
ENSDART00000059321
|
epcam
|
epithelial cell adhesion molecule |
| chr15_+_20239141 | 5.20 |
ENSDART00000101152
ENSDART00000152473 |
spint2
|
serine peptidase inhibitor, Kunitz type, 2 |
| chr9_-_443451 | 5.13 |
ENSDART00000165642
|
si:dkey-11f4.14
|
si:dkey-11f4.14 |
| chr5_-_54712159 | 5.07 |
ENSDART00000149207
|
ccnb1
|
cyclin B1 |
| chr8_-_22558773 | 4.96 |
ENSDART00000074309
|
porcnl
|
porcupine O-acyltransferase like |
| chr8_-_39884359 | 4.93 |
ENSDART00000131372
|
mlec
|
malectin |
| chr6_-_8360918 | 4.92 |
ENSDART00000004716
|
acp5a
|
acid phosphatase 5a, tartrate resistant |
| chr3_+_17939828 | 4.89 |
ENSDART00000185047
|
cnp
|
2',3'-cyclic nucleotide 3' phosphodiesterase |
| chr5_-_69482891 | 4.80 |
ENSDART00000109487
|
CABZ01032476.1
|
|
| chr3_-_34547000 | 4.70 |
ENSDART00000166623
|
sept9a
|
septin 9a |
| chr6_+_112579 | 4.65 |
ENSDART00000034505
|
ap1m2
|
adaptor-related protein complex 1, mu 2 subunit |
| chr5_-_47975440 | 4.63 |
ENSDART00000145665
ENSDART00000007057 |
ccnh
|
cyclin H |
| chr3_-_15119856 | 4.60 |
ENSDART00000138328
|
xpo6
|
exportin 6 |
| chr22_-_17652112 | 4.54 |
ENSDART00000189205
|
hmha1b
|
histocompatibility (minor) HA-1 b |
| chr17_+_22577472 | 4.54 |
ENSDART00000045099
|
yipf4
|
Yip1 domain family, member 4 |
| chr17_-_7733037 | 4.53 |
ENSDART00000064657
|
stx11a
|
syntaxin 11a |
| chr6_-_42949184 | 4.51 |
ENSDART00000147208
|
edem1
|
ER degradation enhancer, mannosidase alpha-like 1 |
| chr2_-_44777592 | 4.49 |
ENSDART00000113351
ENSDART00000169310 |
ncapd2
|
non-SMC condensin I complex, subunit D2 |
| chr4_-_5108844 | 4.46 |
ENSDART00000132666
ENSDART00000136096 |
tmem209
|
transmembrane protein 209 |
| chr20_+_23947004 | 4.46 |
ENSDART00000144195
|
casp8ap2
|
caspase 8 associated protein 2 |
| chr12_+_14149686 | 4.45 |
ENSDART00000123741
|
kbtbd2
|
kelch repeat and BTB (POZ) domain containing 2 |
| chr6_+_46309795 | 4.44 |
ENSDART00000154817
|
si:dkeyp-67f1.1
|
si:dkeyp-67f1.1 |
| chr25_+_3759553 | 4.42 |
ENSDART00000180601
ENSDART00000055845 ENSDART00000157050 ENSDART00000153905 |
thoc5
|
THO complex 5 |
| chr13_-_17860307 | 4.38 |
ENSDART00000135920
ENSDART00000054579 |
march8
|
membrane-associated ring finger (C3HC4) 8 |
| chr5_-_29195063 | 4.37 |
ENSDART00000109926
|
man1b1b
|
mannosidase, alpha, class 1B, member 1b |
| chr3_-_34561624 | 4.36 |
ENSDART00000129313
|
sept9a
|
septin 9a |
| chr10_+_15608326 | 4.29 |
ENSDART00000188770
|
zfand5b
|
zinc finger, AN1-type domain 5b |
| chr7_-_39378903 | 4.23 |
ENSDART00000173659
|
slc8b1
|
solute carrier family 8 (sodium/lithium/calcium exchanger), member B1 |
| chr3_+_18579806 | 4.18 |
ENSDART00000180967
ENSDART00000089765 |
arhgap17b
|
Rho GTPase activating protein 17b |
| chr4_+_4079418 | 4.17 |
ENSDART00000028016
|
waslb
|
Wiskott-Aldrich syndrome-like b |
| chr7_+_24153070 | 4.17 |
ENSDART00000076735
|
lrp10
|
low density lipoprotein receptor-related protein 10 |
| chr10_+_14982977 | 4.17 |
ENSDART00000140869
|
si:dkey-88l16.3
|
si:dkey-88l16.3 |
| chr7_+_19483277 | 4.16 |
ENSDART00000173750
|
si:ch211-212k18.7
|
si:ch211-212k18.7 |
| chr4_-_77561679 | 4.16 |
ENSDART00000180809
|
AL935186.9
|
|
| chr13_-_24717365 | 4.13 |
ENSDART00000137934
ENSDART00000003922 |
erlin1
|
ER lipid raft associated 1 |
| chr19_+_7559901 | 4.13 |
ENSDART00000141189
|
pbxip1a
|
pre-B-cell leukemia homeobox interacting protein 1a |
| chr12_-_35386910 | 4.11 |
ENSDART00000153453
|
camk2g1
|
calcium/calmodulin-dependent protein kinase (CaM kinase) II gamma 1 |
| chr21_+_28747069 | 4.11 |
ENSDART00000014058
|
zgc:100829
|
zgc:100829 |
| chr10_+_15025006 | 4.09 |
ENSDART00000145192
ENSDART00000140084 |
si:dkey-88l16.5
|
si:dkey-88l16.5 |
| chr12_-_18577983 | 4.08 |
ENSDART00000193262
|
zdhhc4
|
zinc finger, DHHC-type containing 4 |
| chr17_+_32623931 | 4.06 |
ENSDART00000144217
|
ctsba
|
cathepsin Ba |
| chr21_+_5192016 | 4.05 |
ENSDART00000139288
|
si:dkey-121h17.7
|
si:dkey-121h17.7 |
| chr20_-_28698172 | 4.03 |
ENSDART00000190635
|
sipa1l1
|
signal-induced proliferation-associated 1 like 1 |
| chr1_+_59321629 | 4.02 |
ENSDART00000161981
|
parn
|
poly(A)-specific ribonuclease (deadenylation nuclease) |
| chr23_-_22523303 | 4.01 |
ENSDART00000079019
|
spsb1
|
splA/ryanodine receptor domain and SOCS box containing 1 |
| chr7_+_39418869 | 4.01 |
ENSDART00000169195
|
CT030188.1
|
|
| chr8_-_7232413 | 4.01 |
ENSDART00000092426
|
grip2a
|
glutamate receptor interacting protein 2a |
| chr18_+_15271993 | 4.01 |
ENSDART00000099777
|
si:dkey-103i16.6
|
si:dkey-103i16.6 |
| chr2_+_24762567 | 3.93 |
ENSDART00000078866
|
ifi30
|
interferon, gamma-inducible protein 30 |
| chr5_-_29152457 | 3.92 |
ENSDART00000078469
|
noxa1
|
NADPH oxidase activator 1 |
| chr17_-_43031763 | 3.92 |
ENSDART00000132754
ENSDART00000050399 |
npc2
|
Niemann-Pick disease, type C2 |
| chr14_+_35428152 | 3.91 |
ENSDART00000172597
|
sytl4
|
synaptotagmin-like 4 |
| chr13_-_35760969 | 3.88 |
ENSDART00000127476
|
erlec1
|
endoplasmic reticulum lectin 1 |
| chr19_+_42086862 | 3.85 |
ENSDART00000151605
|
nfyc
|
nuclear transcription factor Y, gamma |
| chr23_+_40133136 | 3.84 |
ENSDART00000157616
|
gpsm2l
|
G protein signaling modulator 2, like |
| chr18_+_27598755 | 3.81 |
ENSDART00000193808
|
cd82b
|
CD82 molecule b |
| chr1_+_12301913 | 3.80 |
ENSDART00000165733
|
gne
|
glucosamine (UDP-N-acetyl)-2-epimerase/N-acetylmannosamine kinase |
| chr14_+_30285613 | 3.79 |
ENSDART00000173090
|
mtus1a
|
microtubule associated tumor suppressor 1a |
| chr8_+_7875110 | 3.79 |
ENSDART00000167423
ENSDART00000160267 ENSDART00000180490 |
mbd1a
|
methyl-CpG binding domain protein 1a |
| chr12_-_18577774 | 3.77 |
ENSDART00000078166
|
zdhhc4
|
zinc finger, DHHC-type containing 4 |
| chr15_+_38299385 | 3.77 |
ENSDART00000142403
|
si:dkey-24p1.6
|
si:dkey-24p1.6 |
| chr3_+_19665319 | 3.77 |
ENSDART00000007857
ENSDART00000193509 |
mettl2a
|
methyltransferase like 2A |
| chr23_-_31648026 | 3.77 |
ENSDART00000133569
|
sgk1
|
serum/glucocorticoid regulated kinase 1 |
| chr6_-_33916756 | 3.77 |
ENSDART00000137447
ENSDART00000138488 |
nasp
|
nuclear autoantigenic sperm protein (histone-binding) |
| chr20_-_14718801 | 3.76 |
ENSDART00000137605
|
suco
|
SUN domain containing ossification factor |
| chr6_-_17849786 | 3.76 |
ENSDART00000172709
|
rptor
|
regulatory associated protein of MTOR, complex 1 |
| chr15_+_21711671 | 3.76 |
ENSDART00000136151
|
NKAPD1
|
zgc:162339 |
| chr3_-_34136778 | 3.72 |
ENSDART00000131951
|
clpp
|
caseinolytic mitochondrial matrix peptidase proteolytic subunit |
| chr3_+_27664864 | 3.72 |
ENSDART00000126533
ENSDART00000180848 |
clcn7
|
chloride channel 7 |
| chr22_+_26600834 | 3.69 |
ENSDART00000157411
|
adcy9
|
adenylate cyclase 9 |
| chr3_-_40232615 | 3.69 |
ENSDART00000155969
|
flii
|
flightless I actin binding protein |
| chr8_+_20415824 | 3.68 |
ENSDART00000009081
ENSDART00000145444 |
mob3a
|
MOB kinase activator 3A |
| chr17_-_29271359 | 3.68 |
ENSDART00000104219
|
rcor1
|
REST corepressor 1 |
| chr14_-_46198373 | 3.68 |
ENSDART00000031640
ENSDART00000132966 |
zgc:113425
|
zgc:113425 |
| chr2_+_31942390 | 3.67 |
ENSDART00000138684
ENSDART00000146758 ENSDART00000137921 |
otulinb
|
OTU deubiquitinase with linear linkage specificity b |
| chr5_-_29512538 | 3.67 |
ENSDART00000098364
|
ehmt1a
|
euchromatic histone-lysine N-methyltransferase 1a |
| chr2_-_19234329 | 3.67 |
ENSDART00000161106
ENSDART00000160060 ENSDART00000174552 |
cdc20
|
cell division cycle 20 homolog |
| chr8_-_16650595 | 3.66 |
ENSDART00000135319
|
osbpl9
|
oxysterol binding protein-like 9 |
| chr12_-_10508952 | 3.66 |
ENSDART00000152806
|
zgc:152977
|
zgc:152977 |
| chr4_+_26056548 | 3.65 |
ENSDART00000171204
|
SCYL2
|
si:ch211-244b2.1 |
| chr7_+_34794829 | 3.65 |
ENSDART00000009698
ENSDART00000075089 ENSDART00000173456 |
esrp2
|
epithelial splicing regulatory protein 2 |
| chr4_-_4250317 | 3.64 |
ENSDART00000103316
|
cd9b
|
CD9 molecule b |
| chr5_-_29514689 | 3.64 |
ENSDART00000126018
ENSDART00000125175 |
ehmt1a
|
euchromatic histone-lysine N-methyltransferase 1a |
| chr19_-_8812891 | 3.62 |
ENSDART00000151134
ENSDART00000025385 ENSDART00000180291 |
cers2a
|
ceramide synthase 2a |
| chr7_+_24023653 | 3.61 |
ENSDART00000141165
|
tinf2
|
TERF1 (TRF1)-interacting nuclear factor 2 |
| chr4_-_5795309 | 3.61 |
ENSDART00000039987
|
pgm3
|
phosphoglucomutase 3 |
| chr3_-_26191960 | 3.61 |
ENSDART00000113843
|
ypel3
|
yippee-like 3 |
| chr6_-_49547680 | 3.59 |
ENSDART00000169678
|
ppp4r1l
|
protein phosphatase 4, regulatory subunit 1-like |
| chr19_+_43523690 | 3.59 |
ENSDART00000113031
|
wasf2
|
WAS protein family, member 2 |
| chr13_+_28785814 | 3.59 |
ENSDART00000039028
|
nsmce4a
|
NSE4 homolog A, SMC5-SMC6 complex component |
| chr21_-_43665537 | 3.58 |
ENSDART00000157610
|
si:dkey-229d11.3
|
si:dkey-229d11.3 |
| chr6_+_40554551 | 3.57 |
ENSDART00000017859
ENSDART00000155928 |
ddi2
|
DNA-damage inducible protein 2 |
| chr5_+_20453874 | 3.56 |
ENSDART00000124545
ENSDART00000008402 |
sart3
|
squamous cell carcinoma antigen recognized by T cells 3 |
| chr3_-_40051425 | 3.55 |
ENSDART00000146700
|
llgl1
|
lethal giant larvae homolog 1 (Drosophila) |
| chr10_+_44956660 | 3.53 |
ENSDART00000169225
ENSDART00000189298 |
il1b
|
interleukin 1, beta |
| chr13_+_22719789 | 3.53 |
ENSDART00000057672
|
nfkb2
|
nuclear factor of kappa light polypeptide gene enhancer in B-cells 2 (p49/p100) |
| chr3_-_50954607 | 3.52 |
ENSDART00000163810
|
cdc42ep4a
|
CDC42 effector protein (Rho GTPase binding) 4a |
| chr9_-_11655031 | 3.52 |
ENSDART00000044314
|
itgav
|
integrin, alpha V |
| chr1_-_49521407 | 3.51 |
ENSDART00000189845
ENSDART00000143474 |
zp3c
|
zona pellucida glycoprotein 3c |
| chr5_+_39099172 | 3.50 |
ENSDART00000006079
|
bmp2k
|
BMP2 inducible kinase |
| chr7_-_24112484 | 3.49 |
ENSDART00000111923
|
ajuba
|
ajuba LIM protein |
| chr6_+_29860776 | 3.48 |
ENSDART00000028406
|
dlg1
|
discs, large homolog 1 (Drosophila) |
| chr8_-_30742233 | 3.47 |
ENSDART00000098986
|
gucd1
|
guanylyl cyclase domain containing 1 |
| chr4_-_10835620 | 3.46 |
ENSDART00000150739
|
ppfibp1a
|
PTPRF interacting protein, binding protein 1a (liprin beta 1) |
| chr19_+_30867845 | 3.46 |
ENSDART00000047461
|
mfsd2ab
|
major facilitator superfamily domain containing 2ab |
| chr24_+_14801844 | 3.45 |
ENSDART00000141620
|
pi15a
|
peptidase inhibitor 15a |
| chr19_-_34063567 | 3.45 |
ENSDART00000157815
ENSDART00000183907 |
elmo1
|
engulfment and cell motility 1 (ced-12 homolog, C. elegans) |
| chr5_-_69948099 | 3.45 |
ENSDART00000034639
ENSDART00000191111 |
ugt2a4
|
UDP glucuronosyltransferase 2 family, polypeptide A4 |
| chr10_+_2842923 | 3.44 |
ENSDART00000181895
|
ykt6
|
YKT6 v-SNARE homolog (S. cerevisiae) |
| chr24_+_25210015 | 3.43 |
ENSDART00000081043
|
cip2a
|
cell proliferation regulating inhibitor of protein phosphatase 2A |
| chr14_+_24845941 | 3.42 |
ENSDART00000187513
|
arhgef37
|
Rho guanine nucleotide exchange factor (GEF) 37 |
| chr25_-_12410198 | 3.42 |
ENSDART00000182281
|
det1
|
DET1, COP1 ubiquitin ligase partner |
| chr5_+_40835601 | 3.42 |
ENSDART00000147767
|
si:dkey-3h3.3
|
si:dkey-3h3.3 |
| chr22_-_5822147 | 3.41 |
ENSDART00000011076
|
cers5
|
ceramide synthase 5 |
| chr23_+_28378543 | 3.40 |
ENSDART00000145327
|
zgc:153867
|
zgc:153867 |
| chr7_+_36472789 | 3.40 |
ENSDART00000136545
|
aktip
|
akt interacting protein |
| chr10_+_26944978 | 3.39 |
ENSDART00000192148
|
frmd8
|
FERM domain containing 8 |
| chr10_+_14963898 | 3.36 |
ENSDART00000187363
ENSDART00000175732 |
si:dkey-88l16.3
|
si:dkey-88l16.3 |
| chr1_+_12394205 | 3.35 |
ENSDART00000138622
ENSDART00000136421 ENSDART00000139440 ENSDART00000184296 ENSDART00000008127 |
zgc:77739
|
zgc:77739 |
| chr22_-_3299100 | 3.34 |
ENSDART00000160305
|
si:zfos-943e10.1
|
si:zfos-943e10.1 |
| chr3_-_1263047 | 3.34 |
ENSDART00000184388
|
tcf20
|
transcription factor 20 |
| chr10_+_5234327 | 3.34 |
ENSDART00000133927
ENSDART00000063120 |
sptlc1
|
serine palmitoyltransferase, long chain base subunit 1 |
| chr2_+_30249977 | 3.33 |
ENSDART00000109160
ENSDART00000135171 |
tmem70
|
transmembrane protein 70 |
| chr2_-_21167652 | 3.33 |
ENSDART00000185792
|
bmi1b
|
bmi1 polycomb ring finger oncogene 1b |
| chr19_-_31765615 | 3.33 |
ENSDART00000103636
|
si:dkeyp-120h9.1
|
si:dkeyp-120h9.1 |
| chr23_+_40139765 | 3.32 |
ENSDART00000185376
|
gpsm2l
|
G protein signaling modulator 2, like |
| chr13_-_25408387 | 3.32 |
ENSDART00000002741
|
itprip
|
inositol 1,4,5-trisphosphate receptor interacting protein |
| chr21_+_25120546 | 3.31 |
ENSDART00000149507
|
ddx10
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 10 |
| chr15_+_24737599 | 3.28 |
ENSDART00000078024
|
crk
|
v-crk avian sarcoma virus CT10 oncogene homolog |
| chr8_+_15269423 | 3.27 |
ENSDART00000020386
|
gclm
|
glutamate-cysteine ligase, modifier subunit |
| chr9_+_2762270 | 3.26 |
ENSDART00000123342
ENSDART00000001795 ENSDART00000177563 |
sp3a
|
sp3a transcription factor |
| chr15_+_20281305 | 3.26 |
ENSDART00000155065
|
plekhg2
|
pleckstrin homology domain containing, family G (with RhoGef domain) member 2 |
| chr13_-_8785610 | 3.26 |
ENSDART00000021083
|
calm2b
|
calmodulin 2b, (phosphorylase kinase, delta) |
| chr3_-_40254634 | 3.26 |
ENSDART00000154562
|
top3a
|
DNA topoisomerase III alpha |
| chr20_+_9223514 | 3.25 |
ENSDART00000023293
|
kcnk5b
|
potassium channel, subfamily K, member 5b |
| chr3_+_27665160 | 3.24 |
ENSDART00000103660
|
clcn7
|
chloride channel 7 |
| chr14_+_32918484 | 3.23 |
ENSDART00000105721
|
lnx2b
|
ligand of numb-protein X 2b |
| chr18_-_11595567 | 3.23 |
ENSDART00000098565
|
CRACR2A
|
calcium release activated channel regulator 2A |
| chr7_-_60351876 | 3.23 |
ENSDART00000098563
|
plcb3
|
phospholipase C, beta 3 (phosphatidylinositol-specific) |
| chr8_+_3431671 | 3.22 |
ENSDART00000017850
|
ctu1
|
cytosolic thiouridylase subunit 1 homolog (S. pombe) |
| chr18_+_3634652 | 3.21 |
ENSDART00000159913
|
lrch3
|
leucine-rich repeats and calponin homology (CH) domain containing 3 |
| chr20_-_34028967 | 3.19 |
ENSDART00000153408
ENSDART00000033817 |
scyl3
|
SCY1-like, kinase-like 3 |
| chr1_-_31105376 | 3.19 |
ENSDART00000132466
|
ppp1r9alb
|
protein phosphatase 1 regulatory subunit 9A-like B |
| chr10_-_28380919 | 3.18 |
ENSDART00000183409
ENSDART00000183105 ENSDART00000100207 ENSDART00000185392 ENSDART00000131220 |
btg3
|
B-cell translocation gene 3 |
| chr20_-_54198130 | 3.18 |
ENSDART00000160409
|
arf6a
|
ADP-ribosylation factor 6a |
| chr18_+_45666489 | 3.17 |
ENSDART00000180147
ENSDART00000151351 |
prrg4
|
proline rich Gla (G-carboxyglutamic acid) 4 (transmembrane) |
| chr10_-_24765988 | 3.17 |
ENSDART00000064463
|
timm10b
|
translocase of inner mitochondrial membrane 10 homolog B (yeast) |
| chr20_-_35470891 | 3.17 |
ENSDART00000152993
ENSDART00000016090 |
pla2g7
|
phospholipase A2, group VII (platelet-activating factor acetylhydrolase, plasma) |
| chr15_+_11693624 | 3.16 |
ENSDART00000193630
ENSDART00000161930 |
strn4
|
striatin, calmodulin binding protein 4 |
| chr17_+_21126495 | 3.15 |
ENSDART00000022830
|
pygb
|
phosphorylase, glycogen; brain |
| chr18_-_20444296 | 3.13 |
ENSDART00000132993
|
kif23
|
kinesin family member 23 |
| chr21_-_13856689 | 3.12 |
ENSDART00000102197
|
fam129ba
|
family with sequence similarity 129, member Ba |
| chr21_+_25765734 | 3.12 |
ENSDART00000021664
|
cldnb
|
claudin b |
| chr21_-_44771213 | 3.12 |
ENSDART00000188534
|
kif4
|
kinesin family member 4 |
| chr6_-_3998199 | 3.11 |
ENSDART00000059212
|
unc50
|
unc-50 homolog (C. elegans) |
| chr25_+_2361721 | 3.11 |
ENSDART00000172905
|
zmp:0000000932
|
zmp:0000000932 |
| chr7_-_60351537 | 3.11 |
ENSDART00000159875
|
plcb3
|
phospholipase C, beta 3 (phosphatidylinositol-specific) |
| chr15_+_12429206 | 3.09 |
ENSDART00000168997
|
tmprss4a
|
transmembrane protease, serine 4a |
| chr21_-_25295087 | 3.08 |
ENSDART00000087910
ENSDART00000147860 |
st14b
|
suppression of tumorigenicity 14 (colon carcinoma) b |
| chr21_+_28747236 | 3.08 |
ENSDART00000137874
|
zgc:100829
|
zgc:100829 |
| chr5_+_28260158 | 3.07 |
ENSDART00000181434
|
ncaph
|
non-SMC condensin I complex, subunit H |
| chr17_-_30652738 | 3.07 |
ENSDART00000154960
|
sh3yl1
|
SH3 and SYLF domain containing 1 |
| chr7_+_34487833 | 3.07 |
ENSDART00000173854
|
cln6a
|
CLN6, transmembrane ER protein a |
| chr15_-_28904371 | 3.05 |
ENSDART00000155154
|
eml2
|
echinoderm microtubule associated protein like 2 |
| chr12_-_8958353 | 3.04 |
ENSDART00000041728
|
cyp26a1
|
cytochrome P450, family 26, subfamily A, polypeptide 1 |
| chr14_+_32918172 | 3.02 |
ENSDART00000182867
|
lnx2b
|
ligand of numb-protein X 2b |
| chr7_-_59165640 | 3.02 |
ENSDART00000170853
|
haus6
|
HAUS augmin-like complex, subunit 6 |
| chr25_+_19238175 | 3.02 |
ENSDART00000110730
ENSDART00000193619 ENSDART00000154420 |
ppip5k1b
|
diphosphoinositol pentakisphosphate kinase 1b |
| chr12_+_27231607 | 3.02 |
ENSDART00000066270
|
tmem106a
|
transmembrane protein 106A |
| chr8_-_22531817 | 3.01 |
ENSDART00000140606
|
csde1
|
cold shock domain containing E1, RNA-binding |
| chr20_+_2642855 | 3.01 |
ENSDART00000058775
|
zgc:101562
|
zgc:101562 |
| chr9_+_33154841 | 3.00 |
ENSDART00000132465
|
dopey2
|
dopey family member 2 |
| chr9_-_12269847 | 2.99 |
ENSDART00000136558
ENSDART00000144734 ENSDART00000131766 ENSDART00000032344 |
nup35
|
nucleoporin 35 |
Network of associatons between targets according to the STRING database.
First level regulatory network of spi1a+spi1b
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.7 | 10.9 | GO:0002495 | antigen processing and presentation of peptide antigen via MHC class II(GO:0002495) |
| 2.2 | 6.5 | GO:0070125 | mitochondrial translational elongation(GO:0070125) |
| 2.1 | 6.2 | GO:0021531 | spinal cord radial glial cell differentiation(GO:0021531) |
| 1.7 | 5.0 | GO:0061355 | Wnt protein secretion(GO:0061355) |
| 1.3 | 5.3 | GO:2000659 | regulation of interleukin-1-mediated signaling pathway(GO:2000659) negative regulation of interleukin-1-mediated signaling pathway(GO:2000660) |
| 1.2 | 16.2 | GO:0002084 | protein depalmitoylation(GO:0002084) macromolecule depalmitoylation(GO:0098734) |
| 1.2 | 4.8 | GO:0071871 | response to monoamine(GO:0071867) response to catecholamine(GO:0071869) response to epinephrine(GO:0071871) |
| 1.2 | 3.6 | GO:0090085 | regulation of protein deubiquitination(GO:0090085) positive regulation of protein deubiquitination(GO:1903003) |
| 1.2 | 5.9 | GO:0016103 | diterpenoid catabolic process(GO:0016103) terpenoid catabolic process(GO:0016115) retinoic acid catabolic process(GO:0034653) |
| 1.2 | 3.5 | GO:0060148 | negative regulation of hippo signaling(GO:0035331) positive regulation of posttranscriptional gene silencing(GO:0060148) positive regulation of gene silencing by miRNA(GO:2000637) |
| 1.1 | 4.5 | GO:0010032 | meiotic chromosome condensation(GO:0010032) |
| 1.1 | 3.3 | GO:0033615 | mitochondrial proton-transporting ATP synthase complex assembly(GO:0033615) |
| 1.1 | 3.3 | GO:0006480 | N-terminal protein amino acid methylation(GO:0006480) |
| 1.1 | 4.4 | GO:0071788 | endoplasmic reticulum tubular network maintenance(GO:0071788) |
| 1.0 | 3.0 | GO:0060031 | Wnt signaling pathway involved in digestive tract morphogenesis(GO:0044333) mediolateral intercalation(GO:0060031) |
| 0.9 | 3.8 | GO:0034723 | DNA replication-dependent nucleosome assembly(GO:0006335) DNA replication-dependent nucleosome organization(GO:0034723) |
| 0.9 | 7.1 | GO:0003365 | establishment of cell polarity involved in ameboidal cell migration(GO:0003365) |
| 0.9 | 4.4 | GO:0051792 | medium-chain fatty acid biosynthetic process(GO:0051792) |
| 0.9 | 3.5 | GO:0006046 | N-acetylglucosamine catabolic process(GO:0006046) |
| 0.9 | 2.6 | GO:0003218 | cardiac left ventricle morphogenesis(GO:0003214) cardiac left ventricle formation(GO:0003218) pericardium development(GO:0060039) |
| 0.8 | 3.4 | GO:0061158 | 3'-UTR-mediated mRNA destabilization(GO:0061158) |
| 0.8 | 2.5 | GO:0021512 | spinal cord anterior/posterior patterning(GO:0021512) cell proliferation involved in pronephros development(GO:0039015) cell proliferation involved in kidney development(GO:0072111) |
| 0.8 | 3.3 | GO:0006750 | glutathione biosynthetic process(GO:0006750) |
| 0.8 | 2.4 | GO:0035046 | pronuclear migration(GO:0035046) |
| 0.8 | 2.4 | GO:1901004 | ubiquinone-6 metabolic process(GO:1901004) ubiquinone-6 biosynthetic process(GO:1901006) |
| 0.8 | 3.9 | GO:0006521 | regulation of cellular amino acid metabolic process(GO:0006521) regulation of cellular amine metabolic process(GO:0033238) |
| 0.8 | 2.3 | GO:1901223 | negative regulation of NIK/NF-kappaB signaling(GO:1901223) |
| 0.8 | 2.3 | GO:0071236 | cellular response to antibiotic(GO:0071236) cellular response to toxic substance(GO:0097237) |
| 0.7 | 2.2 | GO:0097095 | frontonasal suture morphogenesis(GO:0097095) |
| 0.7 | 2.2 | GO:0010657 | negative regulation of muscle cell apoptotic process(GO:0010656) muscle cell apoptotic process(GO:0010657) striated muscle cell apoptotic process(GO:0010658) regulation of muscle cell apoptotic process(GO:0010660) regulation of striated muscle cell apoptotic process(GO:0010662) negative regulation of striated muscle cell apoptotic process(GO:0010664) |
| 0.7 | 16.3 | GO:0032885 | regulation of polysaccharide biosynthetic process(GO:0032885) |
| 0.7 | 6.4 | GO:0006465 | signal peptide processing(GO:0006465) |
| 0.7 | 3.4 | GO:0000738 | DNA catabolic process, exonucleolytic(GO:0000738) |
| 0.7 | 2.7 | GO:0033540 | fatty acid beta-oxidation using acyl-CoA oxidase(GO:0033540) |
| 0.7 | 1.4 | GO:0007220 | Notch receptor processing(GO:0007220) |
| 0.7 | 5.3 | GO:0033314 | mitotic DNA replication checkpoint(GO:0033314) |
| 0.7 | 2.0 | GO:1901836 | regulation of transcription of nuclear large rRNA transcript from RNA polymerase I promoter(GO:1901836) |
| 0.7 | 3.9 | GO:0032367 | intracellular cholesterol transport(GO:0032367) |
| 0.6 | 3.2 | GO:0045056 | transcytosis(GO:0045056) |
| 0.6 | 3.7 | GO:1990108 | protein linear deubiquitination(GO:1990108) |
| 0.6 | 3.7 | GO:1904668 | positive regulation of ubiquitin protein ligase activity(GO:1904668) |
| 0.6 | 6.0 | GO:0072091 | regulation of stem cell proliferation(GO:0072091) |
| 0.6 | 2.4 | GO:0034729 | histone H3-K79 methylation(GO:0034729) regulation of transcription regulatory region DNA binding(GO:2000677) |
| 0.6 | 3.5 | GO:0090245 | axis elongation involved in somitogenesis(GO:0090245) |
| 0.6 | 1.7 | GO:0019852 | L-ascorbic acid transport(GO:0015882) L-ascorbic acid metabolic process(GO:0019852) |
| 0.6 | 3.9 | GO:2001045 | negative regulation of integrin-mediated signaling pathway(GO:2001045) |
| 0.6 | 5.1 | GO:0007080 | mitotic metaphase plate congression(GO:0007080) |
| 0.6 | 2.8 | GO:0030327 | prenylated protein catabolic process(GO:0030327) prenylcysteine catabolic process(GO:0030328) prenylcysteine metabolic process(GO:0030329) |
| 0.6 | 1.7 | GO:0043903 | regulation of symbiosis, encompassing mutualism through parasitism(GO:0043903) |
| 0.5 | 2.7 | GO:0042264 | peptidyl-aspartic acid modification(GO:0018197) peptidyl-aspartic acid hydroxylation(GO:0042264) negative regulation of transcription from RNA polymerase II promoter in response to stress(GO:0097201) |
| 0.5 | 2.7 | GO:0051988 | regulation of attachment of spindle microtubules to kinetochore(GO:0051988) |
| 0.5 | 6.0 | GO:0007076 | mitotic chromosome condensation(GO:0007076) |
| 0.5 | 9.2 | GO:0035803 | binding of sperm to zona pellucida(GO:0007339) egg coat formation(GO:0035803) regulation of acrosome reaction(GO:0060046) positive regulation of acrosome reaction(GO:2000344) |
| 0.5 | 1.1 | GO:0045730 | respiratory burst(GO:0045730) |
| 0.5 | 5.4 | GO:0031274 | pseudopodium organization(GO:0031268) pseudopodium assembly(GO:0031269) regulation of pseudopodium assembly(GO:0031272) positive regulation of pseudopodium assembly(GO:0031274) |
| 0.5 | 3.2 | GO:0032447 | tRNA wobble position uridine thiolation(GO:0002143) protein urmylation(GO:0032447) |
| 0.5 | 2.1 | GO:2000815 | regulation of mRNA stability involved in response to stress(GO:0010610) regulation of mRNA stability involved in response to oxidative stress(GO:2000815) |
| 0.5 | 4.3 | GO:0046477 | glycosylceramide catabolic process(GO:0046477) |
| 0.5 | 2.6 | GO:0001961 | positive regulation of cytokine-mediated signaling pathway(GO:0001961) positive regulation of response to cytokine stimulus(GO:0060760) |
| 0.5 | 3.7 | GO:0030511 | positive regulation of transforming growth factor beta receptor signaling pathway(GO:0030511) positive regulation of cellular response to transforming growth factor beta stimulus(GO:1903846) |
| 0.5 | 14.1 | GO:0006515 | misfolded or incompletely synthesized protein catabolic process(GO:0006515) |
| 0.5 | 5.7 | GO:0046512 | diol biosynthetic process(GO:0034312) sphingosine biosynthetic process(GO:0046512) |
| 0.5 | 3.1 | GO:0048714 | positive regulation of glial cell differentiation(GO:0045687) positive regulation of oligodendrocyte differentiation(GO:0048714) |
| 0.5 | 6.6 | GO:0006363 | termination of RNA polymerase I transcription(GO:0006363) |
| 0.5 | 2.0 | GO:0099563 | modification of synaptic structure(GO:0099563) |
| 0.5 | 4.0 | GO:1902902 | negative regulation of autophagosome assembly(GO:1902902) |
| 0.5 | 2.5 | GO:0007191 | adenylate cyclase-activating dopamine receptor signaling pathway(GO:0007191) |
| 0.5 | 1.5 | GO:0005997 | xylulose metabolic process(GO:0005997) |
| 0.5 | 2.5 | GO:0033499 | galactose catabolic process(GO:0019388) galactose catabolic process via UDP-galactose(GO:0033499) |
| 0.5 | 3.9 | GO:2000650 | negative regulation of sodium ion transmembrane transporter activity(GO:2000650) |
| 0.5 | 1.0 | GO:0031062 | positive regulation of histone methylation(GO:0031062) |
| 0.5 | 2.8 | GO:0051570 | regulation of histone H3-K9 methylation(GO:0051570) |
| 0.5 | 1.4 | GO:0045648 | positive regulation of erythrocyte differentiation(GO:0045648) |
| 0.5 | 2.3 | GO:0070498 | interleukin-1-mediated signaling pathway(GO:0070498) |
| 0.5 | 1.9 | GO:0016332 | establishment or maintenance of polarity of embryonic epithelium(GO:0016332) cell-cell junction maintenance(GO:0045217) |
| 0.5 | 1.8 | GO:0010668 | ectodermal cell fate commitment(GO:0001712) ectodermal cell fate specification(GO:0001715) ectodermal cell differentiation(GO:0010668) regulation of ectodermal cell fate specification(GO:0042665) regulation of ectoderm development(GO:2000383) |
| 0.5 | 5.5 | GO:0060236 | regulation of mitotic spindle organization(GO:0060236) |
| 0.5 | 3.2 | GO:1905097 | regulation of guanyl-nucleotide exchange factor activity(GO:1905097) regulation of Rho guanyl-nucleotide exchange factor activity(GO:2001106) |
| 0.5 | 6.4 | GO:0051452 | vacuolar acidification(GO:0007035) pH reduction(GO:0045851) intracellular pH reduction(GO:0051452) |
| 0.5 | 6.8 | GO:0070816 | phosphorylation of RNA polymerase II C-terminal domain(GO:0070816) |
| 0.5 | 3.2 | GO:0036010 | protein localization to endosome(GO:0036010) |
| 0.4 | 3.1 | GO:0040016 | embryonic cleavage(GO:0040016) |
| 0.4 | 4.9 | GO:0006611 | protein export from nucleus(GO:0006611) |
| 0.4 | 1.3 | GO:0045453 | bone resorption(GO:0045453) |
| 0.4 | 2.7 | GO:0098957 | anterograde axonal transport of mitochondrion(GO:0098957) |
| 0.4 | 2.2 | GO:0015744 | thiosulfate transport(GO:0015709) oxaloacetate transport(GO:0015729) malate transport(GO:0015743) succinate transport(GO:0015744) succinate transmembrane transport(GO:0071422) malate transmembrane transport(GO:0071423) |
| 0.4 | 3.9 | GO:0016246 | RNA interference(GO:0016246) |
| 0.4 | 1.3 | GO:0006505 | GPI anchor metabolic process(GO:0006505) |
| 0.4 | 6.5 | GO:0051127 | positive regulation of actin nucleation(GO:0051127) positive regulation of Arp2/3 complex-mediated actin nucleation(GO:2000601) |
| 0.4 | 4.3 | GO:0006685 | sphingomyelin catabolic process(GO:0006685) |
| 0.4 | 1.3 | GO:0000390 | spliceosomal complex disassembly(GO:0000390) |
| 0.4 | 8.4 | GO:0030149 | sphingolipid catabolic process(GO:0030149) membrane lipid catabolic process(GO:0046466) |
| 0.4 | 2.1 | GO:0010586 | miRNA metabolic process(GO:0010586) |
| 0.4 | 5.8 | GO:0035020 | regulation of Rac protein signal transduction(GO:0035020) |
| 0.4 | 12.7 | GO:0017121 | phospholipid scrambling(GO:0017121) |
| 0.4 | 2.0 | GO:0016191 | synaptic vesicle uncoating(GO:0016191) |
| 0.4 | 2.0 | GO:0006627 | protein processing involved in protein targeting to mitochondrion(GO:0006627) |
| 0.4 | 0.8 | GO:0035973 | aggrephagy(GO:0035973) |
| 0.4 | 3.5 | GO:0000491 | small nucleolar ribonucleoprotein complex assembly(GO:0000491) |
| 0.4 | 6.6 | GO:0006047 | UDP-N-acetylglucosamine metabolic process(GO:0006047) |
| 0.4 | 4.2 | GO:0050435 | beta-amyloid metabolic process(GO:0050435) |
| 0.4 | 2.3 | GO:0045721 | negative regulation of gluconeogenesis(GO:0045721) |
| 0.4 | 3.8 | GO:0046850 | regulation of bone remodeling(GO:0046850) |
| 0.4 | 2.6 | GO:1990253 | response to leucine(GO:0043201) cellular response to leucine(GO:0071233) regulation of response to reactive oxygen species(GO:1901031) cellular response to leucine starvation(GO:1990253) |
| 0.4 | 6.6 | GO:0043114 | regulation of vascular permeability(GO:0043114) |
| 0.4 | 1.1 | GO:0071470 | cellular response to osmotic stress(GO:0071470) |
| 0.4 | 1.1 | GO:0042787 | protein ubiquitination involved in ubiquitin-dependent protein catabolic process(GO:0042787) regulation of protein ubiquitination involved in ubiquitin-dependent protein catabolic process(GO:2000058) |
| 0.4 | 1.1 | GO:0043416 | regulation of skeletal muscle tissue regeneration(GO:0043416) |
| 0.4 | 7.0 | GO:0007032 | endosome organization(GO:0007032) |
| 0.4 | 6.7 | GO:0008354 | germ cell migration(GO:0008354) |
| 0.4 | 1.1 | GO:0048205 | COPI-coated vesicle budding(GO:0035964) Golgi transport vesicle coating(GO:0048200) COPI coating of Golgi vesicle(GO:0048205) |
| 0.3 | 1.4 | GO:0070131 | regulation of mitochondrial translation(GO:0070129) positive regulation of mitochondrial translation(GO:0070131) |
| 0.3 | 1.4 | GO:0034219 | carbohydrate transmembrane transport(GO:0034219) |
| 0.3 | 1.0 | GO:0034421 | post-translational protein acetylation(GO:0034421) |
| 0.3 | 5.8 | GO:2000369 | regulation of clathrin-mediated endocytosis(GO:2000369) |
| 0.3 | 1.0 | GO:0046443 | FAD biosynthetic process(GO:0006747) FAD metabolic process(GO:0046443) flavin adenine dinucleotide metabolic process(GO:0072387) flavin adenine dinucleotide biosynthetic process(GO:0072388) |
| 0.3 | 4.1 | GO:0021979 | hypothalamus cell differentiation(GO:0021979) |
| 0.3 | 2.4 | GO:1901341 | activation of store-operated calcium channel activity(GO:0032237) positive regulation of store-operated calcium channel activity(GO:1901341) |
| 0.3 | 1.7 | GO:0010828 | positive regulation of glucose transport(GO:0010828) |
| 0.3 | 2.0 | GO:0016255 | attachment of GPI anchor to protein(GO:0016255) |
| 0.3 | 1.3 | GO:0036149 | phosphatidylinositol acyl-chain remodeling(GO:0036149) |
| 0.3 | 4.3 | GO:0032933 | response to sterol depletion(GO:0006991) SREBP signaling pathway(GO:0032933) cellular response to sterol depletion(GO:0071501) |
| 0.3 | 5.6 | GO:0019433 | triglyceride catabolic process(GO:0019433) |
| 0.3 | 2.0 | GO:0019427 | acetate metabolic process(GO:0006083) acetyl-CoA biosynthetic process from acetate(GO:0019427) |
| 0.3 | 1.0 | GO:1902767 | farnesyl diphosphate biosynthetic process, mevalonate pathway(GO:0010142) isoprenoid biosynthetic process via mevalonate(GO:1902767) |
| 0.3 | 4.5 | GO:0008625 | extrinsic apoptotic signaling pathway via death domain receptors(GO:0008625) |
| 0.3 | 1.9 | GO:2000042 | negative regulation of double-strand break repair via homologous recombination(GO:2000042) |
| 0.3 | 2.8 | GO:0043328 | protein targeting to vacuole involved in ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway(GO:0043328) |
| 0.3 | 2.2 | GO:0018027 | peptidyl-lysine dimethylation(GO:0018027) |
| 0.3 | 3.7 | GO:0019747 | regulation of isoprenoid metabolic process(GO:0019747) regulation of vitamin metabolic process(GO:0030656) regulation of retinoic acid biosynthetic process(GO:1900052) |
| 0.3 | 2.4 | GO:0034315 | regulation of Arp2/3 complex-mediated actin nucleation(GO:0034315) |
| 0.3 | 1.8 | GO:0046716 | muscle cell cellular homeostasis(GO:0046716) |
| 0.3 | 0.9 | GO:0032197 | regulation of transposition, RNA-mediated(GO:0010525) negative regulation of transposition, RNA-mediated(GO:0010526) transposition, RNA-mediated(GO:0032197) positive regulation of neuron apoptotic process(GO:0043525) positive regulation of neuron death(GO:1901216) |
| 0.3 | 0.9 | GO:0048194 | Golgi vesicle budding(GO:0048194) |
| 0.3 | 3.6 | GO:1904356 | regulation of telomere maintenance via telomere lengthening(GO:1904356) |
| 0.3 | 0.9 | GO:0000973 | posttranscriptional tethering of RNA polymerase II gene DNA at nuclear periphery(GO:0000973) |
| 0.3 | 1.5 | GO:0090024 | negative regulation of leukocyte chemotaxis(GO:0002689) negative regulation of granulocyte chemotaxis(GO:0071623) negative regulation of neutrophil chemotaxis(GO:0090024) negative regulation of neutrophil migration(GO:1902623) |
| 0.3 | 6.7 | GO:0006491 | N-glycan processing(GO:0006491) |
| 0.3 | 1.4 | GO:0045053 | protein retention in Golgi apparatus(GO:0045053) |
| 0.3 | 1.1 | GO:0036306 | embryonic heart tube elongation(GO:0036306) |
| 0.3 | 2.3 | GO:0044528 | regulation of mitochondrial mRNA stability(GO:0044528) |
| 0.3 | 6.5 | GO:0007205 | protein kinase C-activating G-protein coupled receptor signaling pathway(GO:0007205) |
| 0.3 | 1.4 | GO:0008635 | activation of cysteine-type endopeptidase activity involved in apoptotic process by cytochrome c(GO:0008635) |
| 0.3 | 8.9 | GO:0048920 | posterior lateral line neuromast primordium migration(GO:0048920) |
| 0.3 | 1.9 | GO:0060394 | negative regulation of pathway-restricted SMAD protein phosphorylation(GO:0060394) |
| 0.3 | 1.9 | GO:0008063 | Toll signaling pathway(GO:0008063) |
| 0.3 | 3.0 | GO:0006999 | nuclear pore organization(GO:0006999) |
| 0.3 | 8.3 | GO:0070936 | protein K48-linked ubiquitination(GO:0070936) |
| 0.3 | 1.1 | GO:0000722 | telomere maintenance via recombination(GO:0000722) |
| 0.3 | 1.1 | GO:0018008 | N-terminal protein myristoylation(GO:0006499) N-terminal peptidyl-glycine N-myristoylation(GO:0018008) peptidyl-glycine modification(GO:0018201) protein myristoylation(GO:0018377) |
| 0.3 | 2.4 | GO:0016188 | synaptic vesicle maturation(GO:0016188) |
| 0.3 | 2.3 | GO:0006857 | oligopeptide transport(GO:0006857) |
| 0.3 | 2.8 | GO:0038203 | TORC2 signaling(GO:0038203) |
| 0.3 | 13.3 | GO:0071166 | ribonucleoprotein complex localization(GO:0071166) |
| 0.3 | 1.8 | GO:0045050 | protein insertion into ER membrane by stop-transfer membrane-anchor sequence(GO:0045050) |
| 0.3 | 0.8 | GO:0050427 | sulfate assimilation(GO:0000103) purine ribonucleoside bisphosphate metabolic process(GO:0034035) purine ribonucleoside bisphosphate biosynthetic process(GO:0034036) 3'-phosphoadenosine 5'-phosphosulfate metabolic process(GO:0050427) 3'-phosphoadenosine 5'-phosphosulfate biosynthetic process(GO:0050428) |
| 0.2 | 2.2 | GO:1900077 | negative regulation of insulin receptor signaling pathway(GO:0046627) negative regulation of cellular response to insulin stimulus(GO:1900077) |
| 0.2 | 2.5 | GO:0032878 | regulation of establishment or maintenance of cell polarity(GO:0032878) |
| 0.2 | 5.4 | GO:2000134 | negative regulation of cell cycle G1/S phase transition(GO:1902807) negative regulation of G1/S transition of mitotic cell cycle(GO:2000134) |
| 0.2 | 1.7 | GO:0032475 | otolith formation(GO:0032475) |
| 0.2 | 0.7 | GO:2000055 | positive regulation of Wnt signaling pathway involved in dorsal/ventral axis specification(GO:2000055) |
| 0.2 | 3.2 | GO:0006020 | inositol metabolic process(GO:0006020) |
| 0.2 | 2.2 | GO:0006622 | protein targeting to lysosome(GO:0006622) |
| 0.2 | 2.1 | GO:0030104 | water homeostasis(GO:0030104) |
| 0.2 | 1.9 | GO:0042554 | superoxide anion generation(GO:0042554) |
| 0.2 | 3.2 | GO:0036353 | histone H2A-K119 monoubiquitination(GO:0036353) |
| 0.2 | 2.0 | GO:0001682 | tRNA 5'-leader removal(GO:0001682) |
| 0.2 | 1.1 | GO:1904036 | negative regulation of epithelial cell apoptotic process(GO:1904036) |
| 0.2 | 8.7 | GO:0031647 | regulation of protein stability(GO:0031647) |
| 0.2 | 0.4 | GO:1901097 | negative regulation of autophagosome maturation(GO:1901097) |
| 0.2 | 7.5 | GO:0051209 | release of sequestered calcium ion into cytosol(GO:0051209) |
| 0.2 | 0.9 | GO:0046900 | tetrahydrofolylpolyglutamate metabolic process(GO:0046900) |
| 0.2 | 1.1 | GO:0003173 | ventriculo bulbo valve development(GO:0003173) |
| 0.2 | 0.9 | GO:0042754 | negative regulation of circadian rhythm(GO:0042754) |
| 0.2 | 1.1 | GO:0070650 | actin filament bundle distribution(GO:0070650) |
| 0.2 | 1.9 | GO:0046247 | carotene metabolic process(GO:0016119) carotene catabolic process(GO:0016121) terpene metabolic process(GO:0042214) terpene catabolic process(GO:0046247) |
| 0.2 | 1.3 | GO:0030033 | microvillus assembly(GO:0030033) microvillus organization(GO:0032528) |
| 0.2 | 1.7 | GO:0019430 | response to superoxide(GO:0000303) response to oxygen radical(GO:0000305) removal of superoxide radicals(GO:0019430) cellular response to oxygen radical(GO:0071450) cellular response to superoxide(GO:0071451) cellular oxidant detoxification(GO:0098869) cellular detoxification(GO:1990748) |
| 0.2 | 1.1 | GO:0044205 | 'de novo' UMP biosynthetic process(GO:0044205) |
| 0.2 | 1.5 | GO:0048823 | nucleate erythrocyte development(GO:0048823) |
| 0.2 | 1.7 | GO:0006686 | sphingomyelin biosynthetic process(GO:0006686) |
| 0.2 | 3.5 | GO:0038202 | TORC1 signaling(GO:0038202) |
| 0.2 | 0.8 | GO:0036498 | IRE1-mediated unfolded protein response(GO:0036498) |
| 0.2 | 0.4 | GO:0042256 | mature ribosome assembly(GO:0042256) |
| 0.2 | 2.6 | GO:2000054 | negative regulation of Wnt signaling pathway involved in dorsal/ventral axis specification(GO:2000054) |
| 0.2 | 1.4 | GO:2001256 | regulation of store-operated calcium entry(GO:2001256) |
| 0.2 | 1.6 | GO:0001672 | regulation of chromatin assembly or disassembly(GO:0001672) |
| 0.2 | 2.9 | GO:2000223 | regulation of BMP signaling pathway involved in heart jogging(GO:2000223) |
| 0.2 | 1.0 | GO:0060368 | regulation of Fc receptor mediated stimulatory signaling pathway(GO:0060368) |
| 0.2 | 1.6 | GO:0032515 | negative regulation of phosphoprotein phosphatase activity(GO:0032515) |
| 0.2 | 6.4 | GO:0018230 | peptidyl-L-cysteine S-palmitoylation(GO:0018230) peptidyl-S-diacylglycerol-L-cysteine biosynthetic process from peptidyl-cysteine(GO:0018231) |
| 0.2 | 4.2 | GO:0015701 | bicarbonate transport(GO:0015701) |
| 0.2 | 1.5 | GO:0034063 | stress granule assembly(GO:0034063) |
| 0.2 | 1.3 | GO:0050820 | positive regulation of blood coagulation(GO:0030194) positive regulation of coagulation(GO:0050820) positive regulation of hemostasis(GO:1900048) |
| 0.2 | 1.7 | GO:0007584 | response to nutrient(GO:0007584) |
| 0.2 | 2.8 | GO:0045292 | mRNA cis splicing, via spliceosome(GO:0045292) |
| 0.2 | 1.1 | GO:0031284 | regulation of cGMP metabolic process(GO:0030823) positive regulation of cGMP metabolic process(GO:0030825) regulation of cGMP biosynthetic process(GO:0030826) positive regulation of cGMP biosynthetic process(GO:0030828) regulation of guanylate cyclase activity(GO:0031282) positive regulation of guanylate cyclase activity(GO:0031284) |
| 0.2 | 0.4 | GO:0060823 | canonical Wnt signaling pathway involved in neural plate anterior/posterior pattern formation(GO:0060823) |
| 0.2 | 2.0 | GO:0035587 | adenosine receptor signaling pathway(GO:0001973) purinergic receptor signaling pathway(GO:0035587) G-protein coupled purinergic receptor signaling pathway(GO:0035588) |
| 0.2 | 2.8 | GO:0061055 | myotome development(GO:0061055) |
| 0.2 | 2.0 | GO:0051597 | response to methylmercury(GO:0051597) |
| 0.2 | 1.7 | GO:0015015 | heparan sulfate proteoglycan biosynthetic process, enzymatic modification(GO:0015015) |
| 0.2 | 1.8 | GO:0032785 | negative regulation of DNA-templated transcription, elongation(GO:0032785) negative regulation of transcription elongation from RNA polymerase II promoter(GO:0034244) |
| 0.2 | 0.9 | GO:0071939 | vitamin A transport(GO:0071938) vitamin A import(GO:0071939) |
| 0.2 | 4.8 | GO:0035469 | determination of pancreatic left/right asymmetry(GO:0035469) |
| 0.2 | 3.9 | GO:0048814 | regulation of dendrite morphogenesis(GO:0048814) |
| 0.2 | 2.3 | GO:0051123 | RNA polymerase II transcriptional preinitiation complex assembly(GO:0051123) |
| 0.2 | 5.3 | GO:0051014 | actin filament severing(GO:0051014) |
| 0.2 | 1.6 | GO:0035479 | angioblast cell migration from lateral mesoderm to midline(GO:0035479) |
| 0.2 | 1.7 | GO:0043931 | ossification involved in bone maturation(GO:0043931) bone maturation(GO:0070977) |
| 0.2 | 2.7 | GO:0006003 | fructose 2,6-bisphosphate metabolic process(GO:0006003) |
| 0.2 | 6.2 | GO:0097205 | renal filtration(GO:0097205) |
| 0.2 | 1.7 | GO:0038007 | netrin-activated signaling pathway(GO:0038007) |
| 0.2 | 0.7 | GO:0006297 | nucleotide-excision repair, DNA gap filling(GO:0006297) |
| 0.2 | 1.2 | GO:0070572 | positive regulation of axon regeneration(GO:0048680) positive regulation of neuron projection regeneration(GO:0070572) |
| 0.2 | 0.5 | GO:0009193 | pyrimidine ribonucleoside diphosphate metabolic process(GO:0009193) UDP metabolic process(GO:0046048) |
| 0.2 | 0.7 | GO:1903292 | protein localization to Golgi membrane(GO:1903292) |
| 0.2 | 1.1 | GO:0006283 | transcription-coupled nucleotide-excision repair(GO:0006283) |
| 0.2 | 5.6 | GO:0030488 | tRNA methylation(GO:0030488) |
| 0.2 | 1.4 | GO:0043162 | ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway(GO:0043162) |
| 0.2 | 2.2 | GO:0015781 | pyrimidine nucleotide-sugar transport(GO:0015781) pyrimidine nucleotide-sugar transmembrane transport(GO:0090481) |
| 0.2 | 1.7 | GO:0044818 | mitotic G2/M transition checkpoint(GO:0044818) |
| 0.2 | 1.3 | GO:0030948 | negative regulation of vascular endothelial growth factor receptor signaling pathway(GO:0030948) reelin-mediated signaling pathway(GO:0038026) |
| 0.2 | 5.8 | GO:0071897 | DNA biosynthetic process(GO:0071897) |
| 0.2 | 6.8 | GO:0046513 | ceramide biosynthetic process(GO:0046513) |
| 0.2 | 1.4 | GO:0003428 | growth plate cartilage morphogenesis(GO:0003422) chondrocyte intercalation involved in growth plate cartilage morphogenesis(GO:0003428) |
| 0.2 | 6.4 | GO:0006890 | retrograde vesicle-mediated transport, Golgi to ER(GO:0006890) |
| 0.2 | 1.5 | GO:0002154 | thyroid hormone mediated signaling pathway(GO:0002154) |
| 0.2 | 1.8 | GO:0033198 | response to ATP(GO:0033198) |
| 0.1 | 7.9 | GO:0006892 | post-Golgi vesicle-mediated transport(GO:0006892) |
| 0.1 | 3.3 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
| 0.1 | 0.4 | GO:0009397 | 10-formyltetrahydrofolate metabolic process(GO:0009256) 10-formyltetrahydrofolate catabolic process(GO:0009258) folic acid-containing compound catabolic process(GO:0009397) pteridine-containing compound catabolic process(GO:0042560) |
| 0.1 | 0.4 | GO:0060967 | negative regulation of posttranscriptional gene silencing(GO:0060149) negative regulation of gene silencing by miRNA(GO:0060965) negative regulation of gene silencing by RNA(GO:0060967) |
| 0.1 | 0.4 | GO:0006656 | phosphatidylcholine biosynthetic process(GO:0006656) |
| 0.1 | 4.1 | GO:0007338 | single fertilization(GO:0007338) |
| 0.1 | 1.5 | GO:0046146 | tetrahydrobiopterin biosynthetic process(GO:0006729) tetrahydrobiopterin metabolic process(GO:0046146) |
| 0.1 | 5.2 | GO:0000289 | nuclear-transcribed mRNA poly(A) tail shortening(GO:0000289) |
| 0.1 | 4.9 | GO:0031103 | axon regeneration(GO:0031103) |
| 0.1 | 2.9 | GO:0009250 | glycogen biosynthetic process(GO:0005978) glucan biosynthetic process(GO:0009250) |
| 0.1 | 0.6 | GO:0050926 | regulation of positive chemotaxis(GO:0050926) positive regulation of positive chemotaxis(GO:0050927) induction of positive chemotaxis(GO:0050930) |
| 0.1 | 0.8 | GO:0033169 | histone H3-K9 demethylation(GO:0033169) |
| 0.1 | 0.8 | GO:1900108 | negative regulation of nodal signaling pathway(GO:1900108) |
| 0.1 | 2.4 | GO:0015693 | magnesium ion transport(GO:0015693) |
| 0.1 | 1.5 | GO:0044247 | polysaccharide catabolic process(GO:0000272) glycogen catabolic process(GO:0005980) glucan catabolic process(GO:0009251) cellular polysaccharide catabolic process(GO:0044247) |
| 0.1 | 0.7 | GO:0043508 | negative regulation of JUN kinase activity(GO:0043508) |
| 0.1 | 2.4 | GO:0043486 | histone exchange(GO:0043486) |
| 0.1 | 10.3 | GO:0045930 | negative regulation of mitotic cell cycle(GO:0045930) |
| 0.1 | 4.5 | GO:0099500 | synaptic vesicle fusion to presynaptic active zone membrane(GO:0031629) vesicle fusion to plasma membrane(GO:0099500) |
| 0.1 | 5.4 | GO:0046856 | phosphatidylinositol dephosphorylation(GO:0046856) |
| 0.1 | 1.5 | GO:0060325 | head morphogenesis(GO:0060323) face morphogenesis(GO:0060325) |
| 0.1 | 6.8 | GO:0006661 | phosphatidylinositol biosynthetic process(GO:0006661) |
| 0.1 | 11.0 | GO:0061640 | cytoskeleton-dependent cytokinesis(GO:0061640) |
| 0.1 | 0.8 | GO:0044341 | sodium-dependent phosphate transport(GO:0044341) |
| 0.1 | 2.1 | GO:0043249 | erythrocyte maturation(GO:0043249) |
| 0.1 | 2.9 | GO:0007040 | lysosome organization(GO:0007040) lytic vacuole organization(GO:0080171) |
| 0.1 | 1.3 | GO:0006098 | pentose-phosphate shunt(GO:0006098) |
| 0.1 | 2.2 | GO:0034472 | snRNA 3'-end processing(GO:0034472) |
| 0.1 | 0.9 | GO:0018344 | protein geranylgeranylation(GO:0018344) |
| 0.1 | 0.4 | GO:1904871 | positive regulation of protein localization to nucleus(GO:1900182) regulation of protein localization to Cajal body(GO:1904869) positive regulation of protein localization to Cajal body(GO:1904871) |
| 0.1 | 0.6 | GO:2000001 | regulation of cell cycle checkpoint(GO:1901976) regulation of DNA damage checkpoint(GO:2000001) |
| 0.1 | 1.3 | GO:0046037 | GMP biosynthetic process(GO:0006177) GMP metabolic process(GO:0046037) |
| 0.1 | 0.7 | GO:0016048 | detection of temperature stimulus(GO:0016048) sensory perception of temperature stimulus(GO:0050951) |
| 0.1 | 0.4 | GO:0015712 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 0.1 | 1.6 | GO:0032008 | positive regulation of TOR signaling(GO:0032008) |
| 0.1 | 2.3 | GO:0006613 | cotranslational protein targeting to membrane(GO:0006613) |
| 0.1 | 7.8 | GO:0016571 | histone methylation(GO:0016571) |
| 0.1 | 2.5 | GO:0031163 | iron-sulfur cluster assembly(GO:0016226) metallo-sulfur cluster assembly(GO:0031163) |
| 0.1 | 2.8 | GO:0006884 | cell volume homeostasis(GO:0006884) |
| 0.1 | 2.3 | GO:0071456 | cellular response to hypoxia(GO:0071456) |
| 0.1 | 3.0 | GO:0008286 | insulin receptor signaling pathway(GO:0008286) |
| 0.1 | 22.2 | GO:0018105 | peptidyl-serine phosphorylation(GO:0018105) |
| 0.1 | 0.8 | GO:0090497 | mesenchymal cell migration(GO:0090497) |
| 0.1 | 0.7 | GO:0060397 | JAK-STAT cascade involved in growth hormone signaling pathway(GO:0060397) |
| 0.1 | 0.8 | GO:0006646 | phosphatidylethanolamine biosynthetic process(GO:0006646) |
| 0.1 | 0.9 | GO:0043153 | entrainment of circadian clock by photoperiod(GO:0043153) |
| 0.1 | 6.9 | GO:0007528 | neuromuscular junction development(GO:0007528) |
| 0.1 | 1.6 | GO:0050910 | detection of mechanical stimulus involved in sensory perception of sound(GO:0050910) |
| 0.1 | 1.0 | GO:0061098 | positive regulation of epidermal growth factor-activated receptor activity(GO:0045741) positive regulation of protein tyrosine kinase activity(GO:0061098) positive regulation of receptor activity(GO:2000273) |
| 0.1 | 0.8 | GO:0035264 | multicellular organism growth(GO:0035264) |
| 0.1 | 5.8 | GO:0045766 | positive regulation of angiogenesis(GO:0045766) |
| 0.1 | 0.3 | GO:0071586 | CAAX-box protein processing(GO:0071586) CAAX-box protein maturation(GO:0080120) |
| 0.1 | 4.6 | GO:0032922 | circadian regulation of gene expression(GO:0032922) |
| 0.1 | 2.6 | GO:0001878 | response to yeast(GO:0001878) |
| 0.1 | 2.9 | GO:0009072 | aromatic amino acid family metabolic process(GO:0009072) |
| 0.1 | 8.8 | GO:0007051 | spindle organization(GO:0007051) |
| 0.1 | 2.8 | GO:0006828 | manganese ion transport(GO:0006828) |
| 0.1 | 1.2 | GO:0002098 | tRNA wobble uridine modification(GO:0002098) |
| 0.1 | 2.2 | GO:0034112 | positive regulation of homotypic cell-cell adhesion(GO:0034112) positive regulation of T cell activation(GO:0050870) positive regulation of leukocyte cell-cell adhesion(GO:1903039) |
| 0.1 | 2.5 | GO:0030322 | stabilization of membrane potential(GO:0030322) |
| 0.1 | 0.6 | GO:0070584 | mitochondrion morphogenesis(GO:0070584) |
| 0.1 | 1.4 | GO:0048263 | determination of dorsal identity(GO:0048263) |
| 0.1 | 0.4 | GO:0006953 | acute-phase response(GO:0006953) |
| 0.1 | 1.0 | GO:0070309 | lens fiber cell development(GO:0070307) lens fiber cell morphogenesis(GO:0070309) |
| 0.1 | 2.2 | GO:0006418 | tRNA aminoacylation for protein translation(GO:0006418) |
| 0.1 | 0.8 | GO:0031063 | regulation of histone deacetylation(GO:0031063) |
| 0.1 | 0.4 | GO:0015682 | ferric iron transport(GO:0015682) transferrin transport(GO:0033572) trivalent inorganic cation transport(GO:0072512) |
| 0.1 | 2.3 | GO:0072114 | pronephros morphogenesis(GO:0072114) |
| 0.1 | 1.5 | GO:0016926 | protein desumoylation(GO:0016926) |
| 0.1 | 2.7 | GO:0006998 | nuclear envelope organization(GO:0006998) |
| 0.1 | 1.4 | GO:0032958 | inositol phosphate biosynthetic process(GO:0032958) |
| 0.1 | 0.1 | GO:0002228 | natural killer cell mediated immunity(GO:0002228) positive regulation of lymphocyte mediated immunity(GO:0002708) regulation of natural killer cell mediated immunity(GO:0002715) positive regulation of natural killer cell mediated immunity(GO:0002717) natural killer cell mediated cytotoxicity(GO:0042267) regulation of natural killer cell mediated cytotoxicity(GO:0042269) positive regulation of natural killer cell mediated cytotoxicity(GO:0045954) |
| 0.1 | 1.8 | GO:0051016 | barbed-end actin filament capping(GO:0051016) |
| 0.1 | 0.4 | GO:0031048 | chromatin silencing by small RNA(GO:0031048) |
| 0.1 | 2.9 | GO:0032924 | activin receptor signaling pathway(GO:0032924) |
| 0.1 | 1.8 | GO:0032233 | positive regulation of actin filament bundle assembly(GO:0032233) |
| 0.1 | 1.5 | GO:0098962 | regulation of postsynaptic neurotransmitter receptor activity(GO:0098962) |
| 0.1 | 0.4 | GO:0070933 | histone H4 deacetylation(GO:0070933) |
| 0.1 | 1.3 | GO:0010738 | regulation of protein kinase A signaling(GO:0010738) |
| 0.1 | 0.9 | GO:0019883 | antigen processing and presentation of endogenous peptide antigen(GO:0002483) antigen processing and presentation of endogenous antigen(GO:0019883) antigen processing and presentation of endogenous peptide antigen via MHC class I(GO:0019885) |
| 0.1 | 2.0 | GO:0000245 | spliceosomal complex assembly(GO:0000245) |
| 0.1 | 0.7 | GO:1900026 | regulation of substrate adhesion-dependent cell spreading(GO:1900024) positive regulation of substrate adhesion-dependent cell spreading(GO:1900026) |
| 0.1 | 1.4 | GO:0097034 | mitochondrial respiratory chain complex IV assembly(GO:0033617) mitochondrial respiratory chain complex IV biogenesis(GO:0097034) |
| 0.1 | 1.2 | GO:0000186 | activation of MAPKK activity(GO:0000186) |
| 0.1 | 2.1 | GO:0072348 | sulfur compound transport(GO:0072348) |
| 0.1 | 1.3 | GO:0034629 | cellular protein complex localization(GO:0034629) |
| 0.1 | 0.7 | GO:2001240 | histone dephosphorylation(GO:0016576) negative regulation of signal transduction in absence of ligand(GO:1901099) regulation of extrinsic apoptotic signaling pathway in absence of ligand(GO:2001239) negative regulation of extrinsic apoptotic signaling pathway in absence of ligand(GO:2001240) |
| 0.1 | 0.6 | GO:0018196 | peptidyl-asparagine modification(GO:0018196) |
| 0.1 | 0.5 | GO:0006301 | postreplication repair(GO:0006301) |
| 0.1 | 1.2 | GO:1903288 | positive regulation of sodium ion transport(GO:0010765) positive regulation of sodium ion transmembrane transport(GO:1902307) regulation of potassium ion import(GO:1903286) positive regulation of potassium ion import(GO:1903288) positive regulation of sodium ion transmembrane transporter activity(GO:2000651) |
| 0.1 | 1.1 | GO:0034453 | microtubule anchoring(GO:0034453) |
| 0.1 | 6.8 | GO:0006821 | chloride transport(GO:0006821) |
| 0.1 | 0.2 | GO:0016056 | rhodopsin mediated signaling pathway(GO:0016056) |
| 0.1 | 0.5 | GO:0051897 | positive regulation of protein kinase B signaling(GO:0051897) |
| 0.1 | 1.3 | GO:1990126 | retrograde transport, endosome to plasma membrane(GO:1990126) |
| 0.1 | 3.7 | GO:0044744 | protein import into nucleus(GO:0006606) protein targeting to nucleus(GO:0044744) single-organism nuclear import(GO:1902593) |
| 0.1 | 2.1 | GO:0009395 | phospholipid catabolic process(GO:0009395) |
| 0.1 | 1.3 | GO:0061462 | protein localization to lysosome(GO:0061462) |
| 0.1 | 0.7 | GO:0043951 | regulation of cAMP-mediated signaling(GO:0043949) negative regulation of cAMP-mediated signaling(GO:0043951) |
| 0.1 | 5.5 | GO:0031101 | fin regeneration(GO:0031101) |
| 0.1 | 1.1 | GO:0016441 | posttranscriptional gene silencing(GO:0016441) posttranscriptional gene silencing by RNA(GO:0035194) |
| 0.1 | 3.0 | GO:0070830 | apical junction assembly(GO:0043297) bicellular tight junction assembly(GO:0070830) |
| 0.1 | 1.8 | GO:0070593 | dendrite self-avoidance(GO:0070593) |
| 0.1 | 1.8 | GO:0006368 | transcription elongation from RNA polymerase II promoter(GO:0006368) |
| 0.1 | 0.5 | GO:0034033 | coenzyme A biosynthetic process(GO:0015937) nucleoside bisphosphate biosynthetic process(GO:0033866) ribonucleoside bisphosphate biosynthetic process(GO:0034030) purine nucleoside bisphosphate biosynthetic process(GO:0034033) |
| 0.1 | 2.0 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.1 | 3.2 | GO:0048015 | phosphatidylinositol-mediated signaling(GO:0048015) |
| 0.1 | 0.3 | GO:0050938 | regulation of xanthophore differentiation(GO:0050938) |
| 0.1 | 2.0 | GO:0030866 | cortical actin cytoskeleton organization(GO:0030866) |
| 0.1 | 0.5 | GO:1902742 | apoptotic process involved in development(GO:1902742) |
| 0.1 | 0.2 | GO:0015074 | DNA integration(GO:0015074) |
| 0.1 | 2.4 | GO:0035336 | long-chain fatty-acyl-CoA metabolic process(GO:0035336) |
| 0.1 | 1.6 | GO:0007492 | endoderm development(GO:0007492) |
| 0.1 | 0.7 | GO:0035269 | protein O-linked mannosylation(GO:0035269) |
| 0.1 | 1.2 | GO:0006446 | regulation of translational initiation(GO:0006446) |
| 0.1 | 1.6 | GO:0000184 | nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:0000184) |
| 0.1 | 3.2 | GO:0006637 | acyl-CoA metabolic process(GO:0006637) thioester metabolic process(GO:0035383) |
| 0.1 | 0.3 | GO:0045176 | asymmetric protein localization(GO:0008105) apical protein localization(GO:0045176) |
| 0.1 | 3.6 | GO:0035249 | synaptic transmission, glutamatergic(GO:0035249) |
| 0.1 | 0.5 | GO:0071684 | hatching(GO:0035188) organism emergence from protective structure(GO:0071684) |
| 0.1 | 0.4 | GO:0042249 | establishment of planar polarity of embryonic epithelium(GO:0042249) establishment of planar polarity involved in neural tube closure(GO:0090177) regulation of establishment of planar polarity involved in neural tube closure(GO:0090178) planar cell polarity pathway involved in neural tube closure(GO:0090179) |
| 0.1 | 0.5 | GO:0046320 | regulation of fatty acid beta-oxidation(GO:0031998) regulation of fatty acid oxidation(GO:0046320) |
| 0.1 | 0.5 | GO:0046632 | alpha-beta T cell differentiation(GO:0046632) |
| 0.0 | 0.1 | GO:0006012 | galactose metabolic process(GO:0006012) |
| 0.0 | 0.6 | GO:0021702 | cerebellar Purkinje cell layer formation(GO:0021694) cerebellar Purkinje cell differentiation(GO:0021702) |
| 0.0 | 1.1 | GO:0022011 | myelination in peripheral nervous system(GO:0022011) |
| 0.0 | 7.5 | GO:0043484 | regulation of RNA splicing(GO:0043484) |
| 0.0 | 2.4 | GO:0045765 | regulation of angiogenesis(GO:0045765) |
| 0.0 | 3.3 | GO:0008033 | tRNA processing(GO:0008033) |
| 0.0 | 0.2 | GO:1903723 | negative regulation of centriole elongation(GO:1903723) |
| 0.0 | 1.5 | GO:0030336 | negative regulation of cell migration(GO:0030336) |
| 0.0 | 0.6 | GO:0032483 | regulation of Rab protein signal transduction(GO:0032483) |
| 0.0 | 3.4 | GO:0032147 | activation of protein kinase activity(GO:0032147) |
| 0.0 | 0.9 | GO:0035987 | endodermal cell differentiation(GO:0035987) |
| 0.0 | 1.6 | GO:0007031 | peroxisome organization(GO:0007031) |
| 0.0 | 0.5 | GO:1903311 | regulation of mRNA processing(GO:0050684) regulation of mRNA metabolic process(GO:1903311) |
| 0.0 | 0.2 | GO:0006296 | nucleotide-excision repair, DNA incision, 5'-to lesion(GO:0006296) |
| 0.0 | 3.1 | GO:0043065 | positive regulation of cell death(GO:0010942) positive regulation of apoptotic process(GO:0043065) positive regulation of programmed cell death(GO:0043068) |
| 0.0 | 2.1 | GO:0055088 | lipid homeostasis(GO:0055088) |
| 0.0 | 2.6 | GO:0045055 | regulated exocytosis(GO:0045055) |
| 0.0 | 0.2 | GO:0006370 | 7-methylguanosine mRNA capping(GO:0006370) |
| 0.0 | 0.2 | GO:0034975 | protein folding in endoplasmic reticulum(GO:0034975) |
| 0.0 | 3.6 | GO:0051604 | protein maturation(GO:0051604) |
| 0.0 | 1.1 | GO:0030199 | collagen fibril organization(GO:0030199) |
| 0.0 | 0.9 | GO:0000387 | spliceosomal snRNP assembly(GO:0000387) |
| 0.0 | 1.5 | GO:0032011 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.0 | 1.6 | GO:0006487 | protein N-linked glycosylation(GO:0006487) |
| 0.0 | 2.6 | GO:0030178 | negative regulation of Wnt signaling pathway(GO:0030178) |
| 0.0 | 1.0 | GO:0032465 | regulation of cytokinesis(GO:0032465) |
| 0.0 | 0.2 | GO:0045444 | fat cell differentiation(GO:0045444) |
| 0.0 | 0.4 | GO:0046928 | regulation of neurotransmitter secretion(GO:0046928) |
| 0.0 | 0.6 | GO:0030513 | positive regulation of BMP signaling pathway(GO:0030513) |
| 0.0 | 0.1 | GO:0055014 | atrial cardiac muscle cell development(GO:0055014) |
| 0.0 | 0.3 | GO:0044042 | glycogen metabolic process(GO:0005977) cellular glucan metabolic process(GO:0006073) glucan metabolic process(GO:0044042) |
| 0.0 | 0.6 | GO:0007520 | myoblast fusion(GO:0007520) |
| 0.0 | 0.1 | GO:1903232 | melanosome assembly(GO:1903232) |
| 0.0 | 0.6 | GO:0045454 | cell redox homeostasis(GO:0045454) |
| 0.0 | 0.9 | GO:0030968 | endoplasmic reticulum unfolded protein response(GO:0030968) |
| 0.0 | 1.8 | GO:0009190 | cyclic nucleotide biosynthetic process(GO:0009190) |
| 0.0 | 0.1 | GO:0009972 | cytidine catabolic process(GO:0006216) cytidine deamination(GO:0009972) cytidine metabolic process(GO:0046087) pyrimidine ribonucleoside catabolic process(GO:0046133) |
| 0.0 | 0.1 | GO:0002283 | neutrophil activation involved in immune response(GO:0002283) |
| 0.0 | 0.2 | GO:0000380 | alternative mRNA splicing, via spliceosome(GO:0000380) |
| 0.0 | 0.2 | GO:0043981 | histone H4-K5 acetylation(GO:0043981) histone H4-K8 acetylation(GO:0043982) |
| 0.0 | 0.4 | GO:0045879 | negative regulation of smoothened signaling pathway(GO:0045879) |
| 0.0 | 0.1 | GO:1990168 | protein K29-linked deubiquitination(GO:0035523) protein K33-linked deubiquitination(GO:1990168) |
| 0.0 | 1.3 | GO:0006401 | RNA catabolic process(GO:0006401) |
| 0.0 | 0.2 | GO:0007197 | adenylate cyclase-inhibiting G-protein coupled acetylcholine receptor signaling pathway(GO:0007197) |
| 0.0 | 0.1 | GO:0043649 | dicarboxylic acid catabolic process(GO:0043649) |
| 0.0 | 0.3 | GO:0006513 | protein monoubiquitination(GO:0006513) |
| 0.0 | 0.8 | GO:0007019 | microtubule depolymerization(GO:0007019) |
| 0.0 | 0.3 | GO:0050728 | negative regulation of inflammatory response(GO:0050728) |
| 0.0 | 0.1 | GO:0019336 | phenol-containing compound catabolic process(GO:0019336) catechol-containing compound catabolic process(GO:0019614) dopamine metabolic process(GO:0042417) catecholamine catabolic process(GO:0042424) |
| 0.0 | 0.1 | GO:0015800 | acidic amino acid transport(GO:0015800) aspartate transport(GO:0015810) L-glutamate transport(GO:0015813) malate-aspartate shuttle(GO:0043490) |
| 0.0 | 0.6 | GO:0032436 | positive regulation of proteasomal ubiquitin-dependent protein catabolic process(GO:0032436) positive regulation of proteasomal protein catabolic process(GO:1901800) |
| 0.0 | 1.0 | GO:0048846 | axon extension involved in axon guidance(GO:0048846) neuron projection extension involved in neuron projection guidance(GO:1902284) |
| 0.0 | 0.1 | GO:0031102 | neuron projection regeneration(GO:0031102) |
| 0.0 | 1.1 | GO:0006887 | exocytosis(GO:0006887) |
| 0.0 | 0.4 | GO:0034204 | lipid translocation(GO:0034204) |
| 0.0 | 0.6 | GO:0031146 | SCF-dependent proteasomal ubiquitin-dependent protein catabolic process(GO:0031146) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.7 | 5.1 | GO:0097125 | cyclin B1-CDK1 complex(GO:0097125) |
| 1.2 | 4.8 | GO:0097524 | sperm plasma membrane(GO:0097524) |
| 1.1 | 3.2 | GO:0042719 | mitochondrial intermembrane space protein transporter complex(GO:0042719) |
| 1.1 | 4.2 | GO:1990498 | mitotic spindle microtubule(GO:1990498) |
| 1.1 | 4.2 | GO:0044609 | DBIRD complex(GO:0044609) |
| 1.0 | 4.1 | GO:0005787 | signal peptidase complex(GO:0005787) |
| 1.0 | 8.7 | GO:0032045 | guanyl-nucleotide exchange factor complex(GO:0032045) |
| 0.9 | 6.9 | GO:0031332 | RISC complex(GO:0016442) RNAi effector complex(GO:0031332) |
| 0.8 | 2.5 | GO:0097189 | apoptotic body(GO:0097189) |
| 0.8 | 2.4 | GO:0098826 | endoplasmic reticulum tubular network membrane(GO:0098826) |
| 0.8 | 2.3 | GO:0034663 | endoplasmic reticulum chaperone complex(GO:0034663) |
| 0.8 | 12.8 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.7 | 6.0 | GO:0000796 | condensin complex(GO:0000796) |
| 0.7 | 5.2 | GO:0030121 | AP-1 adaptor complex(GO:0030121) |
| 0.7 | 5.2 | GO:0000445 | THO complex(GO:0000347) THO complex part of transcription export complex(GO:0000445) |
| 0.7 | 2.7 | GO:0043540 | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase complex(GO:0043540) |
| 0.7 | 7.3 | GO:0016602 | CCAAT-binding factor complex(GO:0016602) |
| 0.6 | 6.4 | GO:0000220 | vacuolar proton-transporting V-type ATPase, V0 domain(GO:0000220) |
| 0.6 | 6.8 | GO:0005675 | holo TFIIH complex(GO:0005675) |
| 0.6 | 3.7 | GO:0031428 | box C/D snoRNP complex(GO:0031428) |
| 0.6 | 4.8 | GO:0030670 | phagocytic vesicle membrane(GO:0030670) |
| 0.6 | 3.0 | GO:0031515 | tRNA (m1A) methyltransferase complex(GO:0031515) |
| 0.6 | 1.8 | GO:0008275 | gamma-tubulin small complex(GO:0008275) |
| 0.6 | 2.9 | GO:0034657 | GID complex(GO:0034657) |
| 0.6 | 2.3 | GO:0071556 | integral component of cytoplasmic side of endoplasmic reticulum membrane(GO:0071458) integral component of lumenal side of endoplasmic reticulum membrane(GO:0071556) lumenal side of endoplasmic reticulum membrane(GO:0098553) |
| 0.6 | 2.3 | GO:0005672 | transcription factor TFIIA complex(GO:0005672) |
| 0.6 | 3.4 | GO:0070695 | FHF complex(GO:0070695) |
| 0.6 | 1.7 | GO:0005947 | mitochondrial alpha-ketoglutarate dehydrogenase complex(GO:0005947) mitochondrial tricarboxylic acid cycle enzyme complex(GO:0030062) |
| 0.5 | 8.5 | GO:0032588 | trans-Golgi network membrane(GO:0032588) |
| 0.5 | 5.2 | GO:0045095 | keratin filament(GO:0045095) |
| 0.5 | 3.8 | GO:0031931 | TORC1 complex(GO:0031931) |
| 0.4 | 1.8 | GO:0098890 | extrinsic component of postsynaptic membrane(GO:0098890) |
| 0.4 | 10.1 | GO:0030667 | secretory granule membrane(GO:0030667) |
| 0.4 | 2.6 | GO:0005913 | cell-cell adherens junction(GO:0005913) zonula adherens(GO:0005915) |
| 0.4 | 3.7 | GO:0071797 | LUBAC complex(GO:0071797) |
| 0.4 | 2.0 | GO:1990513 | CLOCK-BMAL transcription complex(GO:1990513) |
| 0.4 | 3.2 | GO:0000306 | extrinsic component of vacuolar membrane(GO:0000306) |
| 0.4 | 3.9 | GO:0000812 | Swr1 complex(GO:0000812) |
| 0.4 | 6.6 | GO:0015030 | Cajal body(GO:0015030) |
| 0.4 | 2.3 | GO:0070876 | SOSS complex(GO:0070876) |
| 0.4 | 13.8 | GO:0032156 | septin ring(GO:0005940) septin complex(GO:0031105) septin cytoskeleton(GO:0032156) |
| 0.4 | 5.1 | GO:0048179 | activin receptor complex(GO:0048179) |
| 0.4 | 1.1 | GO:0033065 | Rad51C-XRCC3 complex(GO:0033065) |
| 0.3 | 2.0 | GO:0098837 | postsynaptic recycling endosome(GO:0098837) |
| 0.3 | 2.0 | GO:0042765 | GPI-anchor transamidase complex(GO:0042765) |
| 0.3 | 3.0 | GO:0044613 | nuclear pore central transport channel(GO:0044613) |
| 0.3 | 1.3 | GO:0044233 | ER-mitochondrion membrane contact site(GO:0044233) |
| 0.3 | 3.9 | GO:0016580 | Sin3 complex(GO:0016580) |
| 0.3 | 2.6 | GO:0061700 | GATOR2 complex(GO:0061700) |
| 0.3 | 1.9 | GO:0000127 | transcription factor TFIIIC complex(GO:0000127) |
| 0.3 | 1.3 | GO:0046695 | SLIK (SAGA-like) complex(GO:0046695) |
| 0.3 | 1.6 | GO:0034448 | EGO complex(GO:0034448) Gtr1-Gtr2 GTPase complex(GO:1990131) |
| 0.3 | 1.9 | GO:0031907 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.3 | 1.4 | GO:0043625 | delta DNA polymerase complex(GO:0043625) |
| 0.3 | 1.1 | GO:0005655 | nucleolar ribonuclease P complex(GO:0005655) |
| 0.3 | 5.5 | GO:0044453 | integral component of nuclear inner membrane(GO:0005639) intrinsic component of nuclear inner membrane(GO:0031229) nuclear membrane part(GO:0044453) |
| 0.3 | 0.8 | GO:1990604 | IRE1-TRAF2-ASK1 complex(GO:1990604) |
| 0.3 | 1.8 | GO:0032797 | SMN complex(GO:0032797) |
| 0.3 | 2.8 | GO:0031932 | TORC2 complex(GO:0031932) TOR complex(GO:0038201) |
| 0.3 | 14.7 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.2 | 4.0 | GO:0031209 | SCAR complex(GO:0031209) |
| 0.2 | 3.6 | GO:0070187 | telosome(GO:0070187) |
| 0.2 | 2.3 | GO:0017059 | serine C-palmitoyltransferase complex(GO:0017059) endoplasmic reticulum palmitoyltransferase complex(GO:0031211) |
| 0.2 | 1.9 | GO:0043020 | NADPH oxidase complex(GO:0043020) |
| 0.2 | 0.9 | GO:0005968 | Rab-protein geranylgeranyltransferase complex(GO:0005968) |
| 0.2 | 3.4 | GO:0071014 | post-mRNA release spliceosomal complex(GO:0071014) |
| 0.2 | 1.5 | GO:0072669 | tRNA-splicing ligase complex(GO:0072669) |
| 0.2 | 5.4 | GO:0055038 | recycling endosome membrane(GO:0055038) |
| 0.2 | 1.3 | GO:0005784 | Sec61 translocon complex(GO:0005784) |
| 0.2 | 3.6 | GO:0000813 | ESCRT I complex(GO:0000813) |
| 0.2 | 2.9 | GO:0034751 | aryl hydrocarbon receptor complex(GO:0034751) |
| 0.2 | 1.6 | GO:0071782 | endoplasmic reticulum tubular network(GO:0071782) endoplasmic reticulum subcompartment(GO:0098827) |
| 0.2 | 2.4 | GO:0097431 | mitotic spindle pole(GO:0097431) |
| 0.2 | 1.8 | GO:0032021 | NELF complex(GO:0032021) |
| 0.2 | 0.6 | GO:0005958 | DNA-dependent protein kinase-DNA ligase 4 complex(GO:0005958) |
| 0.2 | 1.8 | GO:0072546 | ER membrane protein complex(GO:0072546) |
| 0.2 | 2.0 | GO:0071007 | U2-type catalytic step 2 spliceosome(GO:0071007) |
| 0.2 | 4.5 | GO:0045180 | basal cortex(GO:0045180) |
| 0.2 | 1.3 | GO:0045335 | phagocytic vesicle(GO:0045335) |
| 0.2 | 1.7 | GO:0030904 | retromer complex(GO:0030904) |
| 0.2 | 4.5 | GO:0048787 | presynaptic active zone membrane(GO:0048787) |
| 0.2 | 14.0 | GO:0005769 | early endosome(GO:0005769) |
| 0.2 | 0.9 | GO:0070390 | transcription export complex 2(GO:0070390) |
| 0.2 | 19.1 | GO:0016323 | basolateral plasma membrane(GO:0016323) |
| 0.2 | 6.9 | GO:0016605 | PML body(GO:0016605) |
| 0.2 | 1.4 | GO:0070765 | gamma-secretase complex(GO:0070765) |
| 0.2 | 4.0 | GO:0035102 | PRC1 complex(GO:0035102) |
| 0.2 | 3.2 | GO:0005903 | brush border(GO:0005903) |
| 0.2 | 40.1 | GO:0000323 | lytic vacuole(GO:0000323) |
| 0.2 | 1.2 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.2 | 0.5 | GO:0010369 | chromocenter(GO:0010369) |
| 0.2 | 1.1 | GO:0070652 | HAUS complex(GO:0070652) |
| 0.2 | 1.1 | GO:0034045 | pre-autophagosomal structure membrane(GO:0034045) |
| 0.2 | 3.4 | GO:0005637 | nuclear inner membrane(GO:0005637) |
| 0.1 | 0.4 | GO:0097361 | CIA complex(GO:0097361) |
| 0.1 | 1.8 | GO:0030126 | COPI vesicle coat(GO:0030126) |
| 0.1 | 2.9 | GO:0071144 | SMAD2-SMAD3 protein complex(GO:0071144) |
| 0.1 | 1.5 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.1 | 17.4 | GO:0005923 | bicellular tight junction(GO:0005923) occluding junction(GO:0070160) |
| 0.1 | 0.7 | GO:0000814 | ESCRT II complex(GO:0000814) |
| 0.1 | 0.5 | GO:0061673 | mitotic spindle astral microtubule(GO:0061673) |
| 0.1 | 1.2 | GO:0033588 | Elongator holoenzyme complex(GO:0033588) |
| 0.1 | 0.6 | GO:0000172 | ribonuclease MRP complex(GO:0000172) |
| 0.1 | 3.2 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.1 | 8.5 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.1 | 1.1 | GO:0035145 | exon-exon junction complex(GO:0035145) |
| 0.1 | 1.8 | GO:0071564 | npBAF complex(GO:0071564) |
| 0.1 | 2.2 | GO:0032039 | integrator complex(GO:0032039) |
| 0.1 | 1.0 | GO:0070822 | Sin3-type complex(GO:0070822) |
| 0.1 | 2.6 | GO:0033116 | endoplasmic reticulum-Golgi intermediate compartment membrane(GO:0033116) |
| 0.1 | 1.3 | GO:0005956 | protein kinase CK2 complex(GO:0005956) |
| 0.1 | 1.4 | GO:0043189 | NuA4 histone acetyltransferase complex(GO:0035267) H4/H2A histone acetyltransferase complex(GO:0043189) H4 histone acetyltransferase complex(GO:1902562) |
| 0.1 | 1.2 | GO:0043296 | apical junction complex(GO:0043296) |
| 0.1 | 2.4 | GO:0005921 | gap junction(GO:0005921) |
| 0.1 | 6.2 | GO:0001726 | ruffle(GO:0001726) |
| 0.1 | 1.2 | GO:0005682 | U5 snRNP(GO:0005682) |
| 0.1 | 2.4 | GO:0000145 | exocyst(GO:0000145) |
| 0.1 | 6.2 | GO:0030027 | lamellipodium(GO:0030027) |
| 0.1 | 3.4 | GO:0072686 | mitotic spindle(GO:0072686) |
| 0.1 | 4.4 | GO:0008287 | protein serine/threonine phosphatase complex(GO:0008287) phosphatase complex(GO:1903293) |
| 0.1 | 4.4 | GO:0000779 | condensed chromosome, centromeric region(GO:0000779) |
| 0.1 | 1.8 | GO:0097610 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.1 | 1.6 | GO:0034399 | nuclear periphery(GO:0034399) |
| 0.1 | 4.9 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.1 | 0.6 | GO:0016586 | RSC complex(GO:0016586) |
| 0.1 | 4.5 | GO:0036464 | cytoplasmic ribonucleoprotein granule(GO:0036464) |
| 0.1 | 1.0 | GO:0005876 | spindle microtubule(GO:0005876) |
| 0.1 | 10.2 | GO:0099572 | postsynaptic specialization(GO:0099572) |
| 0.1 | 1.8 | GO:0071004 | U2-type prespliceosome(GO:0071004) prespliceosome(GO:0071010) |
| 0.1 | 2.0 | GO:0030496 | midbody(GO:0030496) |
| 0.1 | 3.7 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.1 | 1.4 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.1 | 4.7 | GO:0016324 | apical plasma membrane(GO:0016324) |
| 0.1 | 0.9 | GO:0043209 | myelin sheath(GO:0043209) |
| 0.1 | 66.8 | GO:0005783 | endoplasmic reticulum(GO:0005783) |
| 0.1 | 0.4 | GO:0030134 | ER to Golgi transport vesicle(GO:0030134) |
| 0.1 | 6.5 | GO:0042579 | peroxisome(GO:0005777) microbody(GO:0042579) |
| 0.1 | 1.2 | GO:0090545 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) |
| 0.1 | 5.0 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.1 | 0.1 | GO:0031085 | BLOC-3 complex(GO:0031085) |
| 0.1 | 3.5 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.1 | 2.2 | GO:0031941 | filamentous actin(GO:0031941) |
| 0.0 | 0.4 | GO:0005952 | cAMP-dependent protein kinase complex(GO:0005952) |
| 0.0 | 1.2 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.0 | 0.6 | GO:0017119 | Golgi transport complex(GO:0017119) |
| 0.0 | 31.6 | GO:0005794 | Golgi apparatus(GO:0005794) |
| 0.0 | 4.3 | GO:0005681 | spliceosomal complex(GO:0005681) |
| 0.0 | 0.3 | GO:0000221 | vacuolar proton-transporting V-type ATPase, V1 domain(GO:0000221) |
| 0.0 | 4.5 | GO:0048471 | perinuclear region of cytoplasm(GO:0048471) |
| 0.0 | 0.3 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.0 | 0.3 | GO:0005964 | phosphorylase kinase complex(GO:0005964) |
| 0.0 | 4.0 | GO:0070382 | exocytic vesicle(GO:0070382) |
| 0.0 | 0.2 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.0 | 1.0 | GO:0008305 | integrin complex(GO:0008305) protein complex involved in cell adhesion(GO:0098636) |
| 0.0 | 1.5 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.0 | 0.4 | GO:0042470 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.0 | 0.1 | GO:0008622 | epsilon DNA polymerase complex(GO:0008622) |
| 0.0 | 0.1 | GO:0018444 | translation release factor complex(GO:0018444) |
| 0.0 | 0.4 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.0 | 3.5 | GO:0030425 | dendrite(GO:0030425) |
| 0.0 | 1.2 | GO:0035097 | histone methyltransferase complex(GO:0035097) |
| 0.0 | 0.2 | GO:0031902 | late endosome membrane(GO:0031902) |
| 0.0 | 0.2 | GO:0043204 | perikaryon(GO:0043204) |
| 0.0 | 0.8 | GO:0008023 | transcription elongation factor complex(GO:0008023) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.1 | 8.4 | GO:0008117 | sphinganine-1-phosphate aldolase activity(GO:0008117) |
| 2.0 | 5.9 | GO:0008401 | retinoic acid 4-hydroxylase activity(GO:0008401) |
| 1.6 | 4.9 | GO:0004113 | 2',3'-cyclic-nucleotide 3'-phosphodiesterase activity(GO:0004113) |
| 1.3 | 3.8 | GO:0009384 | UDP-N-acetylglucosamine 2-epimerase activity(GO:0008761) N-acylmannosamine kinase activity(GO:0009384) |
| 1.2 | 13.3 | GO:0017128 | phospholipid scramblase activity(GO:0017128) |
| 1.1 | 3.3 | GO:0071885 | N-terminal protein N-methyltransferase activity(GO:0071885) |
| 1.0 | 6.2 | GO:0008126 | acetylesterase activity(GO:0008126) |
| 1.0 | 20.9 | GO:0098599 | palmitoyl-(protein) hydrolase activity(GO:0008474) palmitoyl hydrolase activity(GO:0098599) |
| 1.0 | 3.9 | GO:0047134 | protein-disulfide reductase activity(GO:0047134) |
| 1.0 | 2.9 | GO:0043812 | phosphatidylinositol-4-phosphate phosphatase activity(GO:0043812) inositol monophosphate 4-phosphatase activity(GO:0052833) inositol monophosphate phosphatase activity(GO:0052834) |
| 0.9 | 2.8 | GO:0003868 | 4-hydroxyphenylpyruvate dioxygenase activity(GO:0003868) |
| 0.9 | 3.8 | GO:0016427 | tRNA (cytosine) methyltransferase activity(GO:0016427) |
| 0.9 | 2.7 | GO:0071532 | ankyrin repeat binding(GO:0071532) |
| 0.9 | 9.1 | GO:0015924 | mannosyl-oligosaccharide 1,2-alpha-mannosidase activity(GO:0004571) mannosyl-oligosaccharide mannosidase activity(GO:0015924) |
| 0.9 | 12.9 | GO:0051117 | ATPase binding(GO:0051117) |
| 0.9 | 12.8 | GO:2001069 | glycogen binding(GO:2001069) |
| 0.8 | 5.1 | GO:0016361 | activin receptor activity, type I(GO:0016361) |
| 0.8 | 4.1 | GO:0003980 | UDP-glucose:glycoprotein glucosyltransferase activity(GO:0003980) |
| 0.8 | 5.7 | GO:0016454 | serine C-palmitoyltransferase activity(GO:0004758) C-palmitoyltransferase activity(GO:0016454) |
| 0.8 | 8.1 | GO:0035198 | miRNA binding(GO:0035198) |
| 0.8 | 4.0 | GO:0004032 | alditol:NADP+ 1-oxidoreductase activity(GO:0004032) |
| 0.8 | 3.0 | GO:0033857 | inositol heptakisphosphate kinase activity(GO:0000829) diphosphoinositol-pentakisphosphate kinase activity(GO:0033857) |
| 0.7 | 2.2 | GO:0004557 | alpha-galactosidase activity(GO:0004557) |
| 0.7 | 3.7 | GO:1990757 | anaphase-promoting complex binding(GO:0010997) ubiquitin ligase activator activity(GO:1990757) |
| 0.7 | 5.8 | GO:0016530 | metallochaperone activity(GO:0016530) |
| 0.7 | 2.9 | GO:0004373 | glycogen (starch) synthase activity(GO:0004373) |
| 0.7 | 2.1 | GO:0008440 | inositol-1,4,5-trisphosphate 3-kinase activity(GO:0008440) |
| 0.7 | 8.3 | GO:0005220 | inositol 1,4,5-trisphosphate-sensitive calcium-release channel activity(GO:0005220) |
| 0.7 | 3.4 | GO:1990756 | protein binding, bridging involved in substrate recognition for ubiquitination(GO:1990756) |
| 0.7 | 2.0 | GO:0004342 | glucosamine-6-phosphate deaminase activity(GO:0004342) |
| 0.7 | 2.7 | GO:0051377 | mannose-ethanolamine phosphotransferase activity(GO:0051377) |
| 0.6 | 1.9 | GO:0003978 | UDP-glucose 4-epimerase activity(GO:0003978) |
| 0.6 | 2.5 | GO:0003827 | alpha-1,3-mannosylglycoprotein 2-beta-N-acetylglucosaminyltransferase activity(GO:0003827) |
| 0.6 | 6.7 | GO:0004465 | lipoprotein lipase activity(GO:0004465) |
| 0.6 | 3.0 | GO:0016429 | tRNA (adenine-N1-)-methyltransferase activity(GO:0016429) |
| 0.6 | 6.5 | GO:0052658 | inositol-1,4,5-trisphosphate 5-phosphatase activity(GO:0052658) |
| 0.6 | 2.4 | GO:0031151 | histone methyltransferase activity (H3-K79 specific)(GO:0031151) |
| 0.6 | 2.3 | GO:0004534 | 5'-3' exoribonuclease activity(GO:0004534) |
| 0.6 | 4.0 | GO:0004430 | 1-phosphatidylinositol 4-kinase activity(GO:0004430) |
| 0.6 | 10.1 | GO:0002039 | p53 binding(GO:0002039) |
| 0.6 | 2.8 | GO:0001735 | prenylcysteine oxidase activity(GO:0001735) |
| 0.6 | 2.2 | GO:0016308 | 1-phosphatidylinositol-4-phosphate 5-kinase activity(GO:0016308) |
| 0.6 | 1.7 | GO:0004394 | heparan sulfate 2-O-sulfotransferase activity(GO:0004394) |
| 0.5 | 2.7 | GO:0003997 | acyl-CoA oxidase activity(GO:0003997) |
| 0.5 | 9.2 | GO:0032190 | acrosin binding(GO:0032190) |
| 0.5 | 1.6 | GO:0016165 | linoleate 13S-lipoxygenase activity(GO:0016165) |
| 0.5 | 1.6 | GO:0046403 | polynucleotide 3'-phosphatase activity(GO:0046403) |
| 0.5 | 3.2 | GO:0070016 | armadillo repeat domain binding(GO:0070016) |
| 0.5 | 5.1 | GO:0005049 | nuclear export signal receptor activity(GO:0005049) |
| 0.5 | 6.0 | GO:0035612 | AP-2 adaptor complex binding(GO:0035612) |
| 0.5 | 1.5 | GO:0005128 | erythropoietin receptor binding(GO:0005128) |
| 0.5 | 3.9 | GO:0019871 | sodium channel inhibitor activity(GO:0019871) |
| 0.5 | 1.4 | GO:0032184 | SUMO polymer binding(GO:0032184) |
| 0.5 | 6.9 | GO:0050291 | sphingosine N-acyltransferase activity(GO:0050291) |
| 0.5 | 8.2 | GO:0005247 | voltage-gated chloride channel activity(GO:0005247) |
| 0.4 | 4.0 | GO:0016176 | superoxide-generating NADPH oxidase activator activity(GO:0016176) |
| 0.4 | 3.6 | GO:0017070 | U6 snRNA binding(GO:0017070) |
| 0.4 | 8.4 | GO:0004190 | aspartic-type endopeptidase activity(GO:0004190) aspartic-type peptidase activity(GO:0070001) |
| 0.4 | 3.5 | GO:0048019 | receptor antagonist activity(GO:0048019) |
| 0.4 | 1.8 | GO:0004525 | ribonuclease III activity(GO:0004525) double-stranded RNA-specific ribonuclease activity(GO:0032296) |
| 0.4 | 2.2 | GO:0015131 | thiosulfate transmembrane transporter activity(GO:0015117) oxaloacetate transmembrane transporter activity(GO:0015131) malate transmembrane transporter activity(GO:0015140) |
| 0.4 | 3.4 | GO:0046912 | transferase activity, transferring acyl groups, acyl groups converted into alkyl on transfer(GO:0046912) |
| 0.4 | 2.6 | GO:0005068 | transmembrane receptor protein tyrosine kinase adaptor activity(GO:0005068) |
| 0.4 | 3.4 | GO:0008515 | sugar:proton symporter activity(GO:0005351) cation:sugar symporter activity(GO:0005402) sucrose:proton symporter activity(GO:0008506) sucrose transmembrane transporter activity(GO:0008515) disaccharide transmembrane transporter activity(GO:0015154) oligosaccharide transmembrane transporter activity(GO:0015157) |
| 0.4 | 5.0 | GO:0016868 | intramolecular transferase activity, phosphotransferases(GO:0016868) |
| 0.4 | 3.6 | GO:0071933 | Arp2/3 complex binding(GO:0071933) |
| 0.4 | 1.9 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.4 | 8.5 | GO:0050321 | tau-protein kinase activity(GO:0050321) |
| 0.4 | 1.5 | GO:0070182 | DNA polymerase binding(GO:0070182) |
| 0.4 | 5.3 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 0.4 | 2.6 | GO:0070728 | leucine binding(GO:0070728) |
| 0.4 | 0.4 | GO:0030547 | receptor inhibitor activity(GO:0030547) |
| 0.4 | 3.0 | GO:0072518 | Rho-dependent protein serine/threonine kinase activity(GO:0072518) |
| 0.4 | 4.7 | GO:0016864 | protein disulfide isomerase activity(GO:0003756) intramolecular oxidoreductase activity, transposing S-S bonds(GO:0016864) |
| 0.4 | 3.2 | GO:0072320 | volume-sensitive chloride channel activity(GO:0072320) |
| 0.4 | 11.0 | GO:0042287 | MHC protein binding(GO:0042287) |
| 0.3 | 1.0 | GO:0003919 | FMN adenylyltransferase activity(GO:0003919) |
| 0.3 | 2.7 | GO:0003836 | beta-galactoside (CMP) alpha-2,3-sialyltransferase activity(GO:0003836) |
| 0.3 | 1.0 | GO:0030267 | hydroxypyruvate reductase activity(GO:0016618) glyoxylate reductase (NADP) activity(GO:0030267) |
| 0.3 | 7.5 | GO:0000993 | RNA polymerase II core binding(GO:0000993) |
| 0.3 | 2.0 | GO:0003987 | acetate-CoA ligase activity(GO:0003987) |
| 0.3 | 4.2 | GO:0005432 | calcium:sodium antiporter activity(GO:0005432) |
| 0.3 | 2.9 | GO:0051920 | peroxiredoxin activity(GO:0051920) |
| 0.3 | 1.3 | GO:0016842 | amidine-lyase activity(GO:0016842) |
| 0.3 | 4.3 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.3 | 3.7 | GO:0043495 | protein anchor(GO:0043495) |
| 0.3 | 1.5 | GO:0008184 | phosphorylase activity(GO:0004645) glycogen phosphorylase activity(GO:0008184) |
| 0.3 | 1.5 | GO:0043139 | 5'-3' DNA helicase activity(GO:0043139) |
| 0.3 | 6.0 | GO:0001671 | ATPase activator activity(GO:0001671) |
| 0.3 | 19.5 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.3 | 2.0 | GO:0070888 | E-box binding(GO:0070888) |
| 0.3 | 5.5 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.3 | 0.9 | GO:0016436 | rRNA (uridine) methyltransferase activity(GO:0016436) |
| 0.3 | 4.3 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.3 | 1.1 | GO:0061656 | SUMO conjugating enzyme activity(GO:0061656) |
| 0.3 | 1.7 | GO:0033188 | sphingomyelin synthase activity(GO:0033188) ceramide cholinephosphotransferase activity(GO:0047493) |
| 0.3 | 10.1 | GO:0004435 | phosphatidylinositol phospholipase C activity(GO:0004435) phospholipase C activity(GO:0004629) |
| 0.3 | 1.4 | GO:0035256 | G-protein coupled glutamate receptor binding(GO:0035256) |
| 0.3 | 3.3 | GO:0004711 | ribosomal protein S6 kinase activity(GO:0004711) |
| 0.3 | 1.4 | GO:0001223 | transcription coactivator binding(GO:0001223) |
| 0.3 | 0.5 | GO:0034594 | phosphatidylinositol trisphosphate phosphatase activity(GO:0034594) |
| 0.3 | 6.5 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.3 | 1.1 | GO:0016635 | oxidoreductase activity, acting on the CH-CH group of donors, quinone or related compound as acceptor(GO:0016635) |
| 0.3 | 2.1 | GO:0003993 | acid phosphatase activity(GO:0003993) |
| 0.3 | 1.1 | GO:0004379 | glycylpeptide N-tetradecanoyltransferase activity(GO:0004379) |
| 0.3 | 2.6 | GO:0052629 | phosphatidylinositol-3,5-bisphosphate 3-phosphatase activity(GO:0052629) |
| 0.3 | 6.5 | GO:0005112 | Notch binding(GO:0005112) |
| 0.3 | 5.8 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.3 | 0.8 | GO:0004781 | adenylylsulfate kinase activity(GO:0004020) sulfate adenylyltransferase activity(GO:0004779) sulfate adenylyltransferase (ATP) activity(GO:0004781) |
| 0.2 | 4.2 | GO:0005487 | nucleocytoplasmic transporter activity(GO:0005487) nuclear import signal receptor activity(GO:0061608) |
| 0.2 | 5.6 | GO:0042800 | histone methyltransferase activity (H3-K4 specific)(GO:0042800) |
| 0.2 | 1.9 | GO:0017169 | CDP-alcohol phosphatidyltransferase activity(GO:0017169) |
| 0.2 | 1.2 | GO:0034046 | poly(G) binding(GO:0034046) |
| 0.2 | 0.7 | GO:0070051 | fibrinogen binding(GO:0070051) |
| 0.2 | 0.9 | GO:0004663 | Rab geranylgeranyltransferase activity(GO:0004663) |
| 0.2 | 2.2 | GO:0002161 | aminoacyl-tRNA editing activity(GO:0002161) |
| 0.2 | 3.1 | GO:0070403 | NAD+ binding(GO:0070403) |
| 0.2 | 2.2 | GO:0005459 | UDP-galactose transmembrane transporter activity(GO:0005459) |
| 0.2 | 2.6 | GO:0008422 | beta-glucosidase activity(GO:0008422) |
| 0.2 | 1.9 | GO:0010436 | beta-carotene 15,15'-monooxygenase activity(GO:0003834) carotenoid dioxygenase activity(GO:0010436) |
| 0.2 | 1.1 | GO:0008821 | crossover junction endodeoxyribonuclease activity(GO:0008821) |
| 0.2 | 2.7 | GO:0005332 | gamma-aminobutyric acid:sodium symporter activity(GO:0005332) |
| 0.2 | 1.3 | GO:0003938 | IMP dehydrogenase activity(GO:0003938) |
| 0.2 | 5.0 | GO:0047617 | acyl-CoA hydrolase activity(GO:0047617) |
| 0.2 | 4.2 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.2 | 3.6 | GO:0045159 | myosin II binding(GO:0045159) |
| 0.2 | 3.3 | GO:0015174 | basic amino acid transmembrane transporter activity(GO:0015174) |
| 0.2 | 2.7 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.2 | 3.2 | GO:0052744 | phosphatidylinositol-3-phosphatase activity(GO:0004438) phosphatidylinositol monophosphate phosphatase activity(GO:0052744) |
| 0.2 | 0.6 | GO:0003910 | DNA ligase activity(GO:0003909) DNA ligase (ATP) activity(GO:0003910) |
| 0.2 | 9.2 | GO:0019707 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.2 | 4.3 | GO:0097153 | cysteine-type endopeptidase activity involved in apoptotic process(GO:0097153) |
| 0.2 | 0.6 | GO:1901612 | cardiolipin binding(GO:1901612) |
| 0.2 | 2.4 | GO:0005158 | insulin receptor binding(GO:0005158) |
| 0.2 | 2.9 | GO:0015271 | outward rectifier potassium channel activity(GO:0015271) |
| 0.2 | 2.7 | GO:0003873 | 6-phosphofructo-2-kinase activity(GO:0003873) |
| 0.2 | 1.5 | GO:0070139 | ubiquitin-like protein-specific endopeptidase activity(GO:0070137) SUMO-specific endopeptidase activity(GO:0070139) |
| 0.2 | 1.7 | GO:0008494 | translation activator activity(GO:0008494) |
| 0.2 | 1.6 | GO:0004791 | thioredoxin-disulfide reductase activity(GO:0004791) |
| 0.2 | 2.3 | GO:0005537 | mannose binding(GO:0005537) |
| 0.2 | 4.7 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.2 | 2.3 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.2 | 4.2 | GO:0031267 | small GTPase binding(GO:0031267) |
| 0.2 | 3.1 | GO:0004532 | exoribonuclease activity(GO:0004532) exoribonuclease activity, producing 5'-phosphomonoesters(GO:0016896) |
| 0.2 | 2.6 | GO:0031682 | G-protein gamma-subunit binding(GO:0031682) |
| 0.2 | 3.6 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.2 | 1.3 | GO:0031419 | cobalamin binding(GO:0031419) |
| 0.2 | 3.4 | GO:0016251 | obsolete general RNA polymerase II transcription factor activity(GO:0016251) |
| 0.2 | 2.8 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.2 | 4.2 | GO:0030159 | receptor signaling complex scaffold activity(GO:0030159) |
| 0.2 | 1.7 | GO:0005042 | netrin receptor activity(GO:0005042) |
| 0.2 | 0.5 | GO:0009041 | uridylate kinase activity(GO:0009041) |
| 0.2 | 1.3 | GO:0015168 | glycerol transmembrane transporter activity(GO:0015168) |
| 0.2 | 1.1 | GO:0004176 | ATP-dependent peptidase activity(GO:0004176) |
| 0.2 | 1.1 | GO:0004517 | nitric-oxide synthase activity(GO:0004517) |
| 0.2 | 1.0 | GO:0008427 | calcium-dependent protein kinase inhibitor activity(GO:0008427) calcium-dependent protein kinase regulator activity(GO:0010858) |
| 0.2 | 3.9 | GO:0015377 | cation:chloride symporter activity(GO:0015377) potassium:chloride symporter activity(GO:0015379) |
| 0.2 | 0.6 | GO:0004579 | oligosaccharyl transferase activity(GO:0004576) dolichyl-diphosphooligosaccharide-protein glycotransferase activity(GO:0004579) |
| 0.2 | 0.9 | GO:0004997 | thyrotropin-releasing hormone receptor activity(GO:0004997) |
| 0.2 | 4.1 | GO:0004016 | adenylate cyclase activity(GO:0004016) |
| 0.2 | 4.7 | GO:0031683 | G-protein beta/gamma-subunit complex binding(GO:0031683) |
| 0.2 | 7.3 | GO:0004683 | calmodulin-dependent protein kinase activity(GO:0004683) |
| 0.2 | 1.8 | GO:0004931 | extracellular ATP-gated cation channel activity(GO:0004931) ATP-gated ion channel activity(GO:0035381) |
| 0.1 | 4.9 | GO:0017147 | Wnt-protein binding(GO:0017147) |
| 0.1 | 0.4 | GO:0016155 | formyltetrahydrofolate dehydrogenase activity(GO:0016155) |
| 0.1 | 2.9 | GO:0070411 | I-SMAD binding(GO:0070411) |
| 0.1 | 0.7 | GO:0060182 | apelin receptor activity(GO:0060182) |
| 0.1 | 3.8 | GO:0001103 | RNA polymerase II repressing transcription factor binding(GO:0001103) |
| 0.1 | 2.6 | GO:0016423 | tRNA (guanine) methyltransferase activity(GO:0016423) |
| 0.1 | 2.0 | GO:0004169 | dolichyl-phosphate-mannose-protein mannosyltransferase activity(GO:0004169) |
| 0.1 | 6.7 | GO:0005546 | phosphatidylinositol-4,5-bisphosphate binding(GO:0005546) |
| 0.1 | 0.6 | GO:0042609 | CD4 receptor binding(GO:0042609) |
| 0.1 | 5.8 | GO:0019902 | phosphatase binding(GO:0019902) |
| 0.1 | 3.4 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.1 | 1.4 | GO:0030628 | pre-mRNA 3'-splice site binding(GO:0030628) |
| 0.1 | 21.9 | GO:0060090 | binding, bridging(GO:0060090) |
| 0.1 | 1.4 | GO:0005025 | transforming growth factor beta receptor activity, type I(GO:0005025) |
| 0.1 | 1.0 | GO:0016212 | kynurenine-oxoglutarate transaminase activity(GO:0016212) kynurenine aminotransferase activity(GO:0036137) |
| 0.1 | 6.7 | GO:0030276 | clathrin binding(GO:0030276) |
| 0.1 | 0.8 | GO:0017040 | ceramidase activity(GO:0017040) |
| 0.1 | 0.5 | GO:0070324 | thyroid hormone binding(GO:0070324) |
| 0.1 | 7.3 | GO:0019208 | phosphatase regulator activity(GO:0019208) |
| 0.1 | 0.8 | GO:0005436 | sodium:phosphate symporter activity(GO:0005436) |
| 0.1 | 6.0 | GO:0001786 | phosphatidylserine binding(GO:0001786) |
| 0.1 | 0.7 | GO:0032217 | riboflavin transporter activity(GO:0032217) |
| 0.1 | 0.9 | GO:0034632 | retinol transporter activity(GO:0034632) |
| 0.1 | 0.4 | GO:0061513 | hexose phosphate transmembrane transporter activity(GO:0015119) organophosphate:inorganic phosphate antiporter activity(GO:0015315) hexose-phosphate:inorganic phosphate antiporter activity(GO:0015526) glucose 6-phosphate:inorganic phosphate antiporter activity(GO:0061513) |
| 0.1 | 2.2 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.1 | 6.7 | GO:0017124 | SH3 domain binding(GO:0017124) |
| 0.1 | 2.6 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 0.1 | 0.6 | GO:0004122 | cystathionine beta-synthase activity(GO:0004122) |
| 0.1 | 0.3 | GO:0030882 | lipid antigen binding(GO:0030882) |
| 0.1 | 2.6 | GO:0004709 | MAP kinase kinase kinase activity(GO:0004709) |
| 0.1 | 3.7 | GO:0099529 | neurotransmitter receptor activity involved in regulation of postsynaptic membrane potential(GO:0099529) transmitter-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1904315) |
| 0.1 | 3.5 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.1 | 4.2 | GO:0015459 | potassium channel regulator activity(GO:0015459) |
| 0.1 | 4.0 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.1 | 3.0 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.1 | 1.6 | GO:0008301 | DNA binding, bending(GO:0008301) |
| 0.1 | 1.4 | GO:0004551 | nucleotide diphosphatase activity(GO:0004551) |
| 0.1 | 1.5 | GO:0005123 | death receptor binding(GO:0005123) |
| 0.1 | 3.2 | GO:0000049 | tRNA binding(GO:0000049) |
| 0.1 | 5.8 | GO:0004521 | endoribonuclease activity(GO:0004521) |
| 0.1 | 3.1 | GO:0070006 | metalloaminopeptidase activity(GO:0070006) |
| 0.1 | 0.4 | GO:0004998 | transferrin receptor activity(GO:0004998) |
| 0.1 | 4.6 | GO:0003755 | peptidyl-prolyl cis-trans isomerase activity(GO:0003755) |
| 0.1 | 0.4 | GO:0004558 | alpha-1,4-glucosidase activity(GO:0004558) |
| 0.1 | 0.7 | GO:0004740 | pyruvate dehydrogenase (acetyl-transferring) kinase activity(GO:0004740) |
| 0.1 | 12.3 | GO:0003714 | transcription corepressor activity(GO:0003714) |
| 0.1 | 0.7 | GO:0043035 | chromatin insulator sequence binding(GO:0043035) |
| 0.1 | 1.7 | GO:0043539 | protein serine/threonine kinase activator activity(GO:0043539) |
| 0.1 | 2.5 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.1 | 0.2 | GO:0098518 | polynucleotide phosphatase activity(GO:0098518) |
| 0.1 | 1.8 | GO:1901476 | carbohydrate transmembrane transporter activity(GO:0015144) carbohydrate transporter activity(GO:1901476) |
| 0.1 | 0.8 | GO:0031013 | troponin C binding(GO:0030172) troponin I binding(GO:0031013) |
| 0.1 | 0.4 | GO:0047184 | 1-acylglycerophosphocholine O-acyltransferase activity(GO:0047184) |
| 0.1 | 2.4 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.1 | 0.5 | GO:0004594 | pantothenate kinase activity(GO:0004594) |
| 0.1 | 1.5 | GO:0003688 | DNA replication origin binding(GO:0003688) |
| 0.1 | 5.4 | GO:0032934 | sterol binding(GO:0032934) |
| 0.1 | 0.6 | GO:0016780 | phosphotransferase activity, for other substituted phosphate groups(GO:0016780) |
| 0.1 | 0.6 | GO:0080019 | fatty-acyl-CoA reductase (alcohol-forming) activity(GO:0080019) |
| 0.1 | 0.3 | GO:0001596 | angiotensin type I receptor activity(GO:0001596) |
| 0.1 | 4.8 | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity(GO:0016538) |
| 0.1 | 0.5 | GO:1900750 | oligopeptide binding(GO:1900750) |
| 0.1 | 0.8 | GO:0043142 | single-stranded DNA-dependent ATP-dependent DNA helicase activity(GO:0017116) single-stranded DNA-dependent ATPase activity(GO:0043142) |
| 0.1 | 1.2 | GO:0004703 | G-protein coupled receptor kinase activity(GO:0004703) |
| 0.1 | 0.2 | GO:0005502 | 11-cis retinal binding(GO:0005502) |
| 0.1 | 1.1 | GO:0004724 | magnesium-dependent protein serine/threonine phosphatase activity(GO:0004724) |
| 0.1 | 0.4 | GO:0004743 | pyruvate kinase activity(GO:0004743) |
| 0.1 | 2.9 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.1 | 0.3 | GO:0016717 | oxidoreductase activity, acting on paired donors, with oxidation of a pair of donors resulting in the reduction of molecular oxygen to two molecules of water(GO:0016717) |
| 0.1 | 3.9 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.1 | 0.3 | GO:0008559 | xenobiotic-transporting ATPase activity(GO:0008559) |
| 0.1 | 0.4 | GO:0008349 | MAP kinase kinase kinase kinase activity(GO:0008349) |
| 0.1 | 1.9 | GO:0030165 | PDZ domain binding(GO:0030165) |
| 0.1 | 0.8 | GO:0003917 | DNA topoisomerase type I activity(GO:0003917) |
| 0.1 | 0.4 | GO:0008603 | cAMP-dependent protein kinase regulator activity(GO:0008603) |
| 0.1 | 1.8 | GO:0004467 | long-chain fatty acid-CoA ligase activity(GO:0004467) |
| 0.1 | 1.8 | GO:0005496 | steroid binding(GO:0005496) |
| 0.1 | 1.7 | GO:0019213 | deacetylase activity(GO:0019213) |
| 0.1 | 0.4 | GO:0071558 | histone demethylase activity (H3-K27 specific)(GO:0071558) |
| 0.1 | 2.1 | GO:0098632 | protein binding involved in cell-cell adhesion(GO:0098632) |
| 0.1 | 0.9 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.1 | 1.9 | GO:0018024 | histone-lysine N-methyltransferase activity(GO:0018024) |
| 0.1 | 1.6 | GO:1901682 | sulfur compound transmembrane transporter activity(GO:1901682) |
| 0.1 | 1.7 | GO:0051537 | 2 iron, 2 sulfur cluster binding(GO:0051537) |
| 0.1 | 0.8 | GO:0032454 | histone demethylase activity (H3-K9 specific)(GO:0032454) |
| 0.1 | 0.2 | GO:0000703 | oxidized pyrimidine nucleobase lesion DNA N-glycosylase activity(GO:0000703) |
| 0.1 | 0.5 | GO:0045236 | CXCR chemokine receptor binding(GO:0045236) |
| 0.1 | 0.2 | GO:2001070 | starch binding(GO:2001070) |
| 0.0 | 0.7 | GO:0008330 | protein tyrosine/threonine phosphatase activity(GO:0008330) |
| 0.0 | 3.8 | GO:0003724 | RNA helicase activity(GO:0003724) |
| 0.0 | 1.6 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.0 | 0.2 | GO:0016521 | pituitary adenylate cyclase activating polypeptide activity(GO:0016521) |
| 0.0 | 13.3 | GO:0005085 | guanyl-nucleotide exchange factor activity(GO:0005085) |
| 0.0 | 2.3 | GO:0042562 | hormone binding(GO:0042562) |
| 0.0 | 0.7 | GO:0004298 | threonine-type endopeptidase activity(GO:0004298) threonine-type peptidase activity(GO:0070003) |
| 0.0 | 1.5 | GO:0030170 | pyridoxal phosphate binding(GO:0030170) |
| 0.0 | 2.9 | GO:0005319 | lipid transporter activity(GO:0005319) |
| 0.0 | 0.7 | GO:0030515 | snoRNA binding(GO:0030515) |
| 0.0 | 3.0 | GO:0019838 | growth factor binding(GO:0019838) |
| 0.0 | 1.0 | GO:0019003 | GDP binding(GO:0019003) |
| 0.0 | 0.4 | GO:0008353 | RNA polymerase II carboxy-terminal domain kinase activity(GO:0008353) |
| 0.0 | 0.1 | GO:1990825 | sequence-specific mRNA binding(GO:1990825) |
| 0.0 | 0.7 | GO:0051018 | protein kinase A binding(GO:0051018) |
| 0.0 | 0.6 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.0 | 1.8 | GO:0004197 | cysteine-type endopeptidase activity(GO:0004197) |
| 0.0 | 0.3 | GO:0046527 | glucosyltransferase activity(GO:0046527) |
| 0.0 | 0.4 | GO:0004415 | hyalurononglucosaminidase activity(GO:0004415) |
| 0.0 | 0.9 | GO:0051539 | 4 iron, 4 sulfur cluster binding(GO:0051539) |
| 0.0 | 1.0 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
| 0.0 | 0.6 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.0 | 1.3 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.0 | 0.8 | GO:0008331 | high voltage-gated calcium channel activity(GO:0008331) |
| 0.0 | 0.1 | GO:0016206 | catechol O-methyltransferase activity(GO:0016206) |
| 0.0 | 0.1 | GO:0001875 | lipopolysaccharide receptor activity(GO:0001875) |
| 0.0 | 2.0 | GO:0001227 | transcriptional repressor activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001227) |
| 0.0 | 1.1 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.0 | 2.6 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 1.2 | GO:0003727 | single-stranded RNA binding(GO:0003727) |
| 0.0 | 11.8 | GO:0004842 | ubiquitin-protein transferase activity(GO:0004842) |
| 0.0 | 2.9 | GO:0036459 | thiol-dependent ubiquitin-specific protease activity(GO:0004843) thiol-dependent ubiquitinyl hydrolase activity(GO:0036459) ubiquitinyl hydrolase activity(GO:0101005) |
| 0.0 | 0.2 | GO:0016907 | G-protein coupled acetylcholine receptor activity(GO:0016907) |
| 0.0 | 0.2 | GO:0004067 | asparaginase activity(GO:0004067) |
| 0.0 | 0.7 | GO:0035014 | phosphatidylinositol 3-kinase regulator activity(GO:0035014) |
| 0.0 | 0.5 | GO:0016881 | acid-amino acid ligase activity(GO:0016881) |
| 0.0 | 0.3 | GO:0097602 | cullin family protein binding(GO:0097602) |
| 0.0 | 0.2 | GO:0070273 | phosphatidylinositol-4-phosphate binding(GO:0070273) |
| 0.0 | 1.5 | GO:0045296 | cadherin binding(GO:0045296) |
| 0.0 | 0.7 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.0 | 0.3 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.0 | 0.1 | GO:1904680 | peptide transporter activity(GO:0015197) oligopeptide transporter activity(GO:0015198) oligopeptide transmembrane transporter activity(GO:0035673) dipeptide transporter activity(GO:0042936) dipeptide transmembrane transporter activity(GO:0071916) peptide transmembrane transporter activity(GO:1904680) |
| 0.0 | 0.1 | GO:0015183 | L-aspartate transmembrane transporter activity(GO:0015183) |
| 0.0 | 0.3 | GO:0051011 | microtubule minus-end binding(GO:0051011) |
| 0.0 | 0.1 | GO:0010859 | calcium-dependent cysteine-type endopeptidase inhibitor activity(GO:0010859) |
| 0.0 | 1.0 | GO:0008375 | acetylglucosaminyltransferase activity(GO:0008375) |
| 0.0 | 0.2 | GO:0005243 | gap junction channel activity(GO:0005243) |
| 0.0 | 0.3 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.0 | 0.4 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.0 | 0.1 | GO:0004126 | cytidine deaminase activity(GO:0004126) |
| 0.0 | 0.1 | GO:0004157 | dihydropyrimidinase activity(GO:0004157) |
| 0.0 | 0.5 | GO:0071949 | FAD binding(GO:0071949) |
| 0.0 | 0.0 | GO:0070643 | vitamin D 25-hydroxylase activity(GO:0070643) |
| 0.0 | 0.3 | GO:0016831 | carboxy-lyase activity(GO:0016831) |
| 0.0 | 1.8 | GO:0004725 | protein tyrosine phosphatase activity(GO:0004725) |
| 0.0 | 1.2 | GO:0003713 | transcription coactivator activity(GO:0003713) |
| 0.0 | 0.2 | GO:0004065 | arylsulfatase activity(GO:0004065) |
| 0.0 | 0.6 | GO:0004722 | protein serine/threonine phosphatase activity(GO:0004722) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 8.4 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
| 0.6 | 3.5 | PID NECTIN PATHWAY | Nectin adhesion pathway |
| 0.4 | 6.2 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 0.4 | 3.0 | ST B CELL ANTIGEN RECEPTOR | B Cell Antigen Receptor |
| 0.4 | 3.3 | SA MMP CYTOKINE CONNECTION | Cytokines can induce activation of matrix metalloproteinases, which degrade extracellular matrix. |
| 0.4 | 7.5 | ST WNT CA2 CYCLIC GMP PATHWAY | Wnt/Ca2+/cyclic GMP signaling. |
| 0.4 | 7.0 | PID DNA PK PATHWAY | DNA-PK pathway in nonhomologous end joining |
| 0.4 | 17.1 | PID ECADHERIN NASCENT AJ PATHWAY | E-cadherin signaling in the nascent adherens junction |
| 0.4 | 2.2 | SA G2 AND M PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
| 0.3 | 12.8 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.3 | 14.1 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.3 | 2.2 | ST G ALPHA S PATHWAY | G alpha s Pathway |
| 0.3 | 1.2 | ST INTERFERON GAMMA PATHWAY | Interferon gamma pathway. |
| 0.3 | 4.9 | PID BETA CATENIN DEG PATHWAY | Degradation of beta catenin |
| 0.3 | 10.3 | PID PI3KCI PATHWAY | Class I PI3K signaling events |
| 0.3 | 7.5 | ST TUMOR NECROSIS FACTOR PATHWAY | Tumor Necrosis Factor Pathway. |
| 0.3 | 1.6 | ST TYPE I INTERFERON PATHWAY | Type I Interferon (alpha/beta IFN) Pathway |
| 0.3 | 3.2 | ST GA12 PATHWAY | G alpha 12 Pathway |
| 0.3 | 4.5 | PID RETINOIC ACID PATHWAY | Retinoic acid receptors-mediated signaling |
| 0.3 | 6.8 | PID IL8 CXCR2 PATHWAY | IL8- and CXCR2-mediated signaling events |
| 0.2 | 1.9 | SA PROGRAMMED CELL DEATH | Programmed cell death, or apoptosis, eliminates damaged or unneeded cells. |
| 0.2 | 5.2 | ST ERK1 ERK2 MAPK PATHWAY | ERK1/ERK2 MAPK Pathway |
| 0.2 | 6.6 | PID ATM PATHWAY | ATM pathway |
| 0.2 | 0.4 | ST STAT3 PATHWAY | STAT3 Pathway |
| 0.2 | 5.9 | PID TRKR PATHWAY | Neurotrophic factor-mediated Trk receptor signaling |
| 0.2 | 1.7 | PID THROMBIN PAR1 PATHWAY | PAR1-mediated thrombin signaling events |
| 0.2 | 5.3 | PID ATR PATHWAY | ATR signaling pathway |
| 0.2 | 2.2 | PID S1P META PATHWAY | Sphingosine 1-phosphate (S1P) pathway |
| 0.1 | 2.2 | PID HIF1A PATHWAY | Hypoxic and oxygen homeostasis regulation of HIF-1-alpha |
| 0.1 | 2.3 | PID ERBB2 ERBB3 PATHWAY | ErbB2/ErbB3 signaling events |
| 0.1 | 2.9 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 0.1 | 2.8 | PID CERAMIDE PATHWAY | Ceramide signaling pathway |
| 0.1 | 4.3 | PID MTOR 4PATHWAY | mTOR signaling pathway |
| 0.1 | 1.7 | PID P38 ALPHA BETA PATHWAY | Regulation of p38-alpha and p38-beta |
| 0.1 | 5.7 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
| 0.1 | 6.0 | PID ERA GENOMIC PATHWAY | Validated nuclear estrogen receptor alpha network |
| 0.1 | 2.8 | PID MYC PATHWAY | C-MYC pathway |
| 0.1 | 3.9 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.1 | 2.7 | PID RAC1 REG PATHWAY | Regulation of RAC1 activity |
| 0.1 | 3.1 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
| 0.1 | 1.6 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
| 0.1 | 2.1 | PID HDAC CLASSII PATHWAY | Signaling events mediated by HDAC Class II |
| 0.1 | 2.6 | PID AR TF PATHWAY | Regulation of Androgen receptor activity |
| 0.1 | 2.0 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.1 | 1.2 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.1 | 2.8 | PID REG GR PATHWAY | Glucocorticoid receptor regulatory network |
| 0.1 | 0.8 | SIG PIP3 SIGNALING IN CARDIAC MYOCTES | Genes related to PIP3 signaling in cardiac myocytes |
| 0.1 | 0.2 | ST MYOCYTE AD PATHWAY | Myocyte Adrenergic Pathway is a specific case of the generalized Adrenergic Pathway. |
| 0.1 | 2.0 | ST FAS SIGNALING PATHWAY | Fas Signaling Pathway |
| 0.1 | 2.9 | PID P73PATHWAY | p73 transcription factor network |
| 0.1 | 0.8 | PID INTEGRIN A9B1 PATHWAY | Alpha9 beta1 integrin signaling events |
| 0.1 | 0.9 | PID PRL SIGNALING EVENTS PATHWAY | Signaling events mediated by PRL |
| 0.1 | 0.8 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.1 | 0.4 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.0 | 0.7 | PID AJDISS 2PATHWAY | Posttranslational regulation of adherens junction stability and dissassembly |
| 0.0 | 0.6 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.0 | 0.3 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
| 0.0 | 1.4 | PID AVB3 INTEGRIN PATHWAY | Integrins in angiogenesis |
| 0.0 | 0.1 | PID UPA UPAR PATHWAY | Urokinase-type plasminogen activator (uPA) and uPAR-mediated signaling |
| 0.0 | 0.6 | PID ARF6 TRAFFICKING PATHWAY | Arf6 trafficking events |
| 0.0 | 0.0 | PID NFKAPPAB ATYPICAL PATHWAY | Atypical NF-kappaB pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 8.3 | REACTOME ELEVATION OF CYTOSOLIC CA2 LEVELS | Genes involved in Elevation of cytosolic Ca2+ levels |
| 0.7 | 14.3 | REACTOME N GLYCAN TRIMMING IN THE ER AND CALNEXIN CALRETICULIN CYCLE | Genes involved in N-glycan trimming in the ER and Calnexin/Calreticulin cycle |
| 0.7 | 6.3 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.7 | 11.9 | REACTOME CYCLIN A B1 ASSOCIATED EVENTS DURING G2 M TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 0.6 | 2.5 | REACTOME OLFACTORY SIGNALING PATHWAY | Genes involved in Olfactory Signaling Pathway |
| 0.6 | 7.2 | REACTOME SYNTHESIS SECRETION AND DEACYLATION OF GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |
| 0.5 | 4.2 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.5 | 2.0 | REACTOME ACYL CHAIN REMODELLING OF PI | Genes involved in Acyl chain remodelling of PI |
| 0.5 | 13.8 | REACTOME LYSOSOME VESICLE BIOGENESIS | Genes involved in Lysosome Vesicle Biogenesis |
| 0.5 | 6.4 | REACTOME ACTIVATION OF NMDA RECEPTOR UPON GLUTAMATE BINDING AND POSTSYNAPTIC EVENTS | Genes involved in Activation of NMDA receptor upon glutamate binding and postsynaptic events |
| 0.4 | 1.3 | REACTOME PASSIVE TRANSPORT BY AQUAPORINS | Genes involved in Passive Transport by Aquaporins |
| 0.4 | 6.0 | REACTOME IONOTROPIC ACTIVITY OF KAINATE RECEPTORS | Genes involved in Ionotropic activity of Kainate Receptors |
| 0.4 | 3.7 | REACTOME ADENYLATE CYCLASE ACTIVATING PATHWAY | Genes involved in Adenylate cyclase activating pathway |
| 0.4 | 3.1 | REACTOME ARMS MEDIATED ACTIVATION | Genes involved in ARMS-mediated activation |
| 0.4 | 2.0 | REACTOME ETHANOL OXIDATION | Genes involved in Ethanol oxidation |
| 0.4 | 3.1 | REACTOME P75NTR SIGNALS VIA NFKB | Genes involved in p75NTR signals via NF-kB |
| 0.4 | 5.4 | REACTOME TRAF3 DEPENDENT IRF ACTIVATION PATHWAY | Genes involved in TRAF3-dependent IRF activation pathway |
| 0.4 | 3.8 | REACTOME MTORC1 MEDIATED SIGNALLING | Genes involved in mTORC1-mediated signalling |
| 0.3 | 4.6 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.3 | 5.9 | REACTOME TRAF6 MEDIATED NFKB ACTIVATION | Genes involved in TRAF6 mediated NF-kB activation |
| 0.3 | 6.5 | REACTOME SYNTHESIS OF PIPS AT THE GOLGI MEMBRANE | Genes involved in Synthesis of PIPs at the Golgi membrane |
| 0.3 | 5.2 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.3 | 4.8 | REACTOME ASSOCIATION OF LICENSING FACTORS WITH THE PRE REPLICATIVE COMPLEX | Genes involved in Association of licensing factors with the pre-replicative complex |
| 0.3 | 3.5 | REACTOME GLYCOGEN BREAKDOWN GLYCOGENOLYSIS | Genes involved in Glycogen breakdown (glycogenolysis) |
| 0.3 | 5.3 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.3 | 3.6 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |
| 0.3 | 1.8 | REACTOME RECRUITMENT OF NUMA TO MITOTIC CENTROSOMES | Genes involved in Recruitment of NuMA to mitotic centrosomes |
| 0.2 | 7.2 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.2 | 5.9 | REACTOME CYTOCHROME P450 ARRANGED BY SUBSTRATE TYPE | Genes involved in Cytochrome P450 - arranged by substrate type |
| 0.2 | 0.7 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.2 | 1.6 | REACTOME REGULATION OF IFNA SIGNALING | Genes involved in Regulation of IFNA signaling |
| 0.2 | 2.3 | REACTOME AMINO ACID SYNTHESIS AND INTERCONVERSION TRANSAMINATION | Genes involved in Amino acid synthesis and interconversion (transamination) |
| 0.2 | 2.9 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |
| 0.2 | 2.2 | REACTOME IRAK2 MEDIATED ACTIVATION OF TAK1 COMPLEX UPON TLR7 8 OR 9 STIMULATION | Genes involved in IRAK2 mediated activation of TAK1 complex upon TLR7/8 or 9 stimulation |
| 0.2 | 2.8 | REACTOME NEGATIVE REGULATORS OF RIG I MDA5 SIGNALING | Genes involved in Negative regulators of RIG-I/MDA5 signaling |
| 0.2 | 1.9 | REACTOME RAP1 SIGNALLING | Genes involved in Rap1 signalling |
| 0.2 | 2.8 | REACTOME SYNTHESIS OF SUBSTRATES IN N GLYCAN BIOSYTHESIS | Genes involved in Synthesis of substrates in N-glycan biosythesis |
| 0.2 | 3.2 | REACTOME REGULATION OF HYPOXIA INDUCIBLE FACTOR HIF BY OXYGEN | Genes involved in Regulation of Hypoxia-inducible Factor (HIF) by Oxygen |
| 0.2 | 1.5 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |
| 0.2 | 2.0 | REACTOME PECAM1 INTERACTIONS | Genes involved in PECAM1 interactions |
| 0.2 | 1.8 | REACTOME COPI MEDIATED TRANSPORT | Genes involved in COPI Mediated Transport |
| 0.2 | 5.0 | REACTOME APC CDC20 MEDIATED DEGRADATION OF NEK2A | Genes involved in APC-Cdc20 mediated degradation of Nek2A |
| 0.2 | 1.3 | REACTOME PHOSPHORYLATION OF CD3 AND TCR ZETA CHAINS | Genes involved in Phosphorylation of CD3 and TCR zeta chains |
| 0.2 | 5.5 | REACTOME POST TRANSLATIONAL MODIFICATION SYNTHESIS OF GPI ANCHORED PROTEINS | Genes involved in Post-translational modification: synthesis of GPI-anchored proteins |
| 0.2 | 0.8 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.2 | 4.1 | REACTOME PRE NOTCH TRANSCRIPTION AND TRANSLATION | Genes involved in Pre-NOTCH Transcription and Translation |
| 0.2 | 2.2 | REACTOME AKT PHOSPHORYLATES TARGETS IN THE CYTOSOL | Genes involved in AKT phosphorylates targets in the cytosol |
| 0.2 | 2.0 | REACTOME PLATELET CALCIUM HOMEOSTASIS | Genes involved in Platelet calcium homeostasis |
| 0.2 | 3.7 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.2 | 3.7 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.2 | 0.8 | REACTOME CD28 DEPENDENT PI3K AKT SIGNALING | Genes involved in CD28 dependent PI3K/Akt signaling |
| 0.2 | 1.6 | REACTOME MEMBRANE BINDING AND TARGETTING OF GAG PROTEINS | Genes involved in Membrane binding and targetting of GAG proteins |
| 0.2 | 1.9 | REACTOME VITAMIN B5 PANTOTHENATE METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |
| 0.2 | 1.9 | REACTOME INHIBITION OF REPLICATION INITIATION OF DAMAGED DNA BY RB1 E2F1 | Genes involved in Inhibition of replication initiation of damaged DNA by RB1/E2F1 |
| 0.2 | 6.3 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.1 | 1.5 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.1 | 3.3 | REACTOME PEROXISOMAL LIPID METABOLISM | Genes involved in Peroxisomal lipid metabolism |
| 0.1 | 0.4 | REACTOME ACYL CHAIN REMODELLING OF PS | Genes involved in Acyl chain remodelling of PS |
| 0.1 | 2.1 | REACTOME RNA POL I TRANSCRIPTION INITIATION | Genes involved in RNA Polymerase I Transcription Initiation |
| 0.1 | 3.7 | REACTOME KERATAN SULFATE BIOSYNTHESIS | Genes involved in Keratan sulfate biosynthesis |
| 0.1 | 0.8 | REACTOME INTEGRATION OF PROVIRUS | Genes involved in Integration of provirus |
| 0.1 | 1.5 | REACTOME TELOMERE MAINTENANCE | Genes involved in Telomere Maintenance |
| 0.1 | 5.2 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |
| 0.1 | 2.6 | REACTOME THE ROLE OF NEF IN HIV1 REPLICATION AND DISEASE PATHOGENESIS | Genes involved in The role of Nef in HIV-1 replication and disease pathogenesis |
| 0.1 | 1.8 | REACTOME RNA POL III TRANSCRIPTION INITIATION FROM TYPE 2 PROMOTER | Genes involved in RNA Polymerase III Transcription Initiation From Type 2 Promoter |
| 0.1 | 1.4 | REACTOME POL SWITCHING | Genes involved in Polymerase switching |
| 0.1 | 3.7 | REACTOME GLUCOSE METABOLISM | Genes involved in Glucose metabolism |
| 0.1 | 4.7 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.1 | 5.0 | REACTOME RNA POL II TRANSCRIPTION PRE INITIATION AND PROMOTER OPENING | Genes involved in RNA Polymerase II Transcription Pre-Initiation And Promoter Opening |
| 0.1 | 2.1 | REACTOME GPVI MEDIATED ACTIVATION CASCADE | Genes involved in GPVI-mediated activation cascade |
| 0.1 | 4.7 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
| 0.1 | 2.2 | REACTOME MITOCHONDRIAL TRNA AMINOACYLATION | Genes involved in Mitochondrial tRNA aminoacylation |
| 0.1 | 1.4 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.1 | 1.2 | REACTOME SYNTHESIS OF PIPS AT THE PLASMA MEMBRANE | Genes involved in Synthesis of PIPs at the plasma membrane |
| 0.1 | 1.8 | REACTOME FGFR LIGAND BINDING AND ACTIVATION | Genes involved in FGFR ligand binding and activation |
| 0.1 | 1.7 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.1 | 0.7 | REACTOME SHC RELATED EVENTS | Genes involved in SHC-related events |
| 0.1 | 1.0 | REACTOME SRP DEPENDENT COTRANSLATIONAL PROTEIN TARGETING TO MEMBRANE | Genes involved in SRP-dependent cotranslational protein targeting to membrane |
| 0.1 | 1.4 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.1 | 5.5 | REACTOME TRANSPORT OF INORGANIC CATIONS ANIONS AND AMINO ACIDS OLIGOPEPTIDES | Genes involved in Transport of inorganic cations/anions and amino acids/oligopeptides |
| 0.1 | 7.4 | REACTOME MRNA SPLICING | Genes involved in mRNA Splicing |
| 0.1 | 0.8 | REACTOME THE ACTIVATION OF ARYLSULFATASES | Genes involved in The activation of arylsulfatases |
| 0.1 | 0.8 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 0.1 | 0.2 | REACTOME REGULATION OF SIGNALING BY CBL | Genes involved in Regulation of signaling by CBL |
| 0.1 | 0.5 | REACTOME GENERATION OF SECOND MESSENGER MOLECULES | Genes involved in Generation of second messenger molecules |
| 0.1 | 2.1 | REACTOME TRANSPORT OF MATURE MRNA DERIVED FROM AN INTRONLESS TRANSCRIPT | Genes involved in Transport of Mature mRNA Derived from an Intronless Transcript |
| 0.1 | 1.6 | REACTOME SIGNALING BY ROBO RECEPTOR | Genes involved in Signaling by Robo receptor |
| 0.0 | 1.7 | REACTOME TRANSPORT TO THE GOLGI AND SUBSEQUENT MODIFICATION | Genes involved in Transport to the Golgi and subsequent modification |
| 0.0 | 0.2 | REACTOME MRNA CAPPING | Genes involved in mRNA Capping |
| 0.0 | 1.0 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.0 | 1.0 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.0 | 0.9 | REACTOME TRNA AMINOACYLATION | Genes involved in tRNA Aminoacylation |
| 0.0 | 2.4 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.0 | 0.2 | REACTOME OPSINS | Genes involved in Opsins |
| 0.0 | 0.6 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |
| 0.0 | 0.2 | REACTOME ERKS ARE INACTIVATED | Genes involved in ERKs are inactivated |
| 0.0 | 0.2 | REACTOME FACILITATIVE NA INDEPENDENT GLUCOSE TRANSPORTERS | Genes involved in Facilitative Na+-independent glucose transporters |
| 0.0 | 0.2 | REACTOME IL 6 SIGNALING | Genes involved in Interleukin-6 signaling |
| 0.0 | 3.5 | REACTOME ANTIGEN PROCESSING UBIQUITINATION PROTEASOME DEGRADATION | Genes involved in Antigen processing: Ubiquitination & Proteasome degradation |
| 0.0 | 0.7 | REACTOME L1CAM INTERACTIONS | Genes involved in L1CAM interactions |
| 0.0 | 0.3 | REACTOME BIOSYNTHESIS OF THE N GLYCAN PRECURSOR DOLICHOL LIPID LINKED OLIGOSACCHARIDE LLO AND TRANSFER TO A NASCENT PROTEIN | Genes involved in Biosynthesis of the N-glycan precursor (dolichol lipid-linked oligosaccharide, LLO) and transfer to a nascent protein |
| 0.0 | 0.1 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION IN TLR7 8 OR 9 SIGNALING | Genes involved in TRAF6 mediated IRF7 activation in TLR7/8 or 9 signaling |
| 0.0 | 0.3 | REACTOME FANCONI ANEMIA PATHWAY | Genes involved in Fanconi Anemia pathway |
| 0.0 | 0.4 | REACTOME NOTCH1 INTRACELLULAR DOMAIN REGULATES TRANSCRIPTION | Genes involved in NOTCH1 Intracellular Domain Regulates Transcription |
| 0.0 | 0.1 | REACTOME DIGESTION OF DIETARY CARBOHYDRATE | Genes involved in Digestion of dietary carbohydrate |
| 0.0 | 0.2 | REACTOME BASE FREE SUGAR PHOSPHATE REMOVAL VIA THE SINGLE NUCLEOTIDE REPLACEMENT PATHWAY | Genes involved in Base-free sugar-phosphate removal via the single-nucleotide replacement pathway |
| 0.0 | 0.3 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.0 | 0.3 | REACTOME METABOLISM OF PROTEINS | Genes involved in Metabolism of proteins |