Project
PRJNA438478: RNAseq of wild type zebrafish germline
Navigation
Downloads
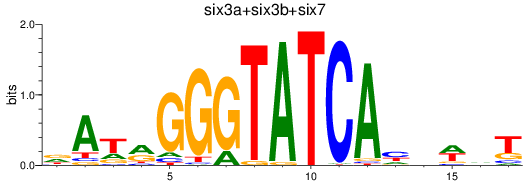
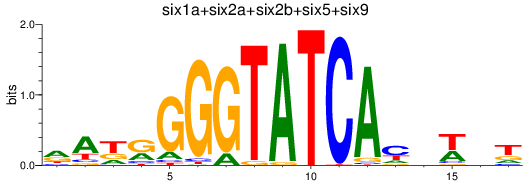
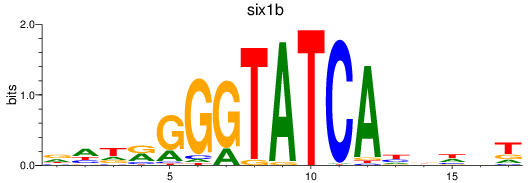
Results for six3a+six3b+six7_six1a+six2a+six2b+six5+six9_six1b
Z-value: 0.75
Transcription factors associated with six3a+six3b+six7_six1a+six2a+six2b+six5+six9_six1b
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
six3b
|
ENSDARG00000054879 | SIX homeobox 3b |
|
six3a
|
ENSDARG00000058008 | SIX homeobox 3a |
|
six7
|
ENSDARG00000070107 | SIX homeobox 7 |
|
six3a
|
ENSDARG00000114971 | SIX homeobox 3a |
|
six1a
|
ENSDARG00000039304 | SIX homeobox 1a |
|
six2b
|
ENSDARG00000054878 | SIX homeobox 2b |
|
six2a
|
ENSDARG00000058004 | SIX homeobox 2a |
|
six5
|
ENSDARG00000068406 | SIX homeobox 5 |
|
six9
|
ENSDARG00000068407 | SIX homeobox 9 |
|
six2a
|
ENSDARG00000115214 | SIX homeobox 2a |
|
six1b
|
ENSDARG00000026473 | SIX homeobox 1b |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| six2b | dr11_v1_chr12_-_25612170_25612170 | -0.94 | 4.0e-09 | Click! |
| six3b | dr11_v1_chr12_+_25600685_25600685 | -0.94 | 1.3e-08 | Click! |
| six1b | dr11_v1_chr20_+_20499869_20499869 | -0.91 | 1.6e-07 | Click! |
| six7 | dr11_v1_chr7_-_7420301_7420301 | -0.90 | 3.7e-07 | Click! |
| six9 | dr11_v1_chr18_-_37241080_37241115 | -0.86 | 4.4e-06 | Click! |
| six5 | dr11_v1_chr18_-_37252036_37252036 | 0.81 | 3.9e-05 | Click! |
| six3a | dr11_v1_chr13_-_10261383_10261383 | -0.77 | 1.8e-04 | Click! |
| six1a | dr11_v1_chr13_-_31622195_31622195 | -0.54 | 2.2e-02 | Click! |
| six2a | dr11_v1_chr13_+_10232695_10232695 | 0.09 | 7.3e-01 | Click! |
Activity profile of six3a+six3b+six7_six1a+six2a+six2b+six5+six9_six1b motif
Sorted Z-values of six3a+six3b+six7_six1a+six2a+six2b+six5+six9_six1b motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr10_-_42898220 | 3.87 |
ENSDART00000099270
|
CU326366.2
|
|
| chr18_-_49283058 | 2.44 |
ENSDART00000076554
|
si:zfos-464b6.2
|
si:zfos-464b6.2 |
| chr11_-_44194132 | 2.44 |
ENSDART00000182954
ENSDART00000111271 |
CABZ01080074.1
|
|
| chr17_+_16429826 | 2.28 |
ENSDART00000136078
|
efcab11
|
EF-hand calcium binding domain 11 |
| chr21_-_7940043 | 2.14 |
ENSDART00000099733
ENSDART00000136671 |
f2rl1.1
|
coagulation factor II (thrombin) receptor-like 1, tandem duplicate 1 |
| chr10_+_42898103 | 1.74 |
ENSDART00000015872
|
zcchc9
|
zinc finger, CCHC domain containing 9 |
| chr25_-_21763750 | 1.58 |
ENSDART00000089596
|
tmem168b
|
transmembrane protein 168b |
| chr10_+_10972795 | 1.27 |
ENSDART00000127331
|
cdc37l1
|
cell division cycle 37-like 1 |
| chr17_-_2573021 | 1.22 |
ENSDART00000074181
|
zp3.2
|
zona pellucida glycoprotein 3, tandem duplicate 2 |
| chr19_+_34230108 | 1.10 |
ENSDART00000141950
|
galnt12
|
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 12 |
| chr20_+_34502606 | 1.08 |
ENSDART00000139739
|
gorab
|
golgin, rab6-interacting |
| chr5_+_37903790 | 1.07 |
ENSDART00000162470
|
tmprss4b
|
transmembrane protease, serine 4b |
| chr5_+_41477526 | 1.05 |
ENSDART00000153567
|
pias2
|
protein inhibitor of activated STAT, 2 |
| chr11_-_41220794 | 1.01 |
ENSDART00000192895
|
mrps16
|
mitochondrial ribosomal protein S16 |
| chr5_+_36899691 | 1.00 |
ENSDART00000132322
|
hnrnpl
|
heterogeneous nuclear ribonucleoprotein L |
| chr15_+_41815703 | 0.92 |
ENSDART00000059508
|
pxylp1
|
2-phosphoxylose phosphatase 1 |
| chr16_+_50755133 | 0.91 |
ENSDART00000029283
|
IGLON5
|
zgc:110372 |
| chr8_-_26961779 | 0.90 |
ENSDART00000099214
|
slc16a1b
|
solute carrier family 16 (monocarboxylate transporter), member 1b |
| chr16_+_26735894 | 0.82 |
ENSDART00000191605
ENSDART00000185175 |
rad54b
|
RAD54 homolog B (S. cerevisiae) |
| chr1_-_24255149 | 0.78 |
ENSDART00000146960
|
lrba
|
LPS-responsive vesicle trafficking, beach and anchor containing |
| chr21_-_26028205 | 0.72 |
ENSDART00000034875
|
sdf2
|
stromal cell-derived factor 2 |
| chr7_-_72423666 | 0.71 |
ENSDART00000191214
|
rph3ab
|
rabphilin 3A homolog (mouse), b |
| chr7_+_56517402 | 0.70 |
ENSDART00000073597
|
dhodh
|
dihydroorotate dehydrogenase |
| chr5_+_41477954 | 0.67 |
ENSDART00000185871
|
pias2
|
protein inhibitor of activated STAT, 2 |
| chr20_+_37866861 | 0.65 |
ENSDART00000153220
|
vash2
|
vasohibin 2 |
| chr5_+_36900157 | 0.64 |
ENSDART00000183533
ENSDART00000051184 |
hnrnpl
|
heterogeneous nuclear ribonucleoprotein L |
| chr4_-_27398385 | 0.60 |
ENSDART00000142117
ENSDART00000150553 ENSDART00000182746 |
fam19a5a
|
family with sequence similarity 19 (chemokine (C-C motif)-like), member A5a |
| chr22_-_5171362 | 0.57 |
ENSDART00000124889
|
tnfaip8l1
|
tumor necrosis factor, alpha-induced protein 8-like 1 |
| chr7_-_29356084 | 0.52 |
ENSDART00000075757
|
gtf2a2
|
general transcription factor IIA, 2 |
| chr5_-_26247973 | 0.51 |
ENSDART00000098527
|
erap1b
|
endoplasmic reticulum aminopeptidase 1b |
| chr16_-_46567344 | 0.51 |
ENSDART00000127721
|
si:dkey-152b24.7
|
si:dkey-152b24.7 |
| chr17_+_13099476 | 0.51 |
ENSDART00000012670
|
pnn
|
pinin, desmosome associated protein |
| chr16_-_46567136 | 0.50 |
ENSDART00000159180
|
si:dkey-152b24.7
|
si:dkey-152b24.7 |
| chr6_-_28222592 | 0.47 |
ENSDART00000131126
|
bcl6a
|
B-cell CLL/lymphoma 6a (zinc finger protein 51) |
| chr7_-_59515569 | 0.46 |
ENSDART00000163343
ENSDART00000165457 ENSDART00000163745 |
slx1b
|
SLX1 homolog B, structure-specific endonuclease subunit |
| chr24_+_30435164 | 0.45 |
ENSDART00000193877
|
dpyda.1
|
dihydropyrimidine dehydrogenase a, tandem duplicate 1 |
| chr10_-_32610776 | 0.45 |
ENSDART00000017436
|
mogat2
|
monoacylglycerol O-acyltransferase 2 |
| chr23_-_27045231 | 0.42 |
ENSDART00000187979
|
zgc:66440
|
zgc:66440 |
| chr16_+_41060161 | 0.42 |
ENSDART00000141130
|
scap
|
SREBF chaperone |
| chr23_-_4019699 | 0.42 |
ENSDART00000159780
|
slc9a8
|
solute carrier family 9, subfamily A (NHE8, cation proton antiporter 8), member 8 |
| chr23_-_4019928 | 0.41 |
ENSDART00000021062
|
slc9a8
|
solute carrier family 9, subfamily A (NHE8, cation proton antiporter 8), member 8 |
| chr2_+_26240631 | 0.41 |
ENSDART00000129895
|
palm1b
|
paralemmin 1b |
| chr5_-_54395488 | 0.40 |
ENSDART00000160781
|
zmynd19
|
zinc finger, MYND-type containing 19 |
| chr4_-_7811925 | 0.39 |
ENSDART00000103070
|
cdk17
|
cyclin-dependent kinase 17 |
| chr21_-_21465111 | 0.39 |
ENSDART00000141487
|
nectin3b
|
nectin cell adhesion molecule 3b |
| chr25_+_3788074 | 0.38 |
ENSDART00000154008
|
chid1
|
chitinase domain containing 1 |
| chr1_+_37752171 | 0.38 |
ENSDART00000183247
ENSDART00000189756 ENSDART00000139448 |
GALNTL6
|
si:ch211-15e22.3 |
| chr9_-_13871935 | 0.38 |
ENSDART00000146597
|
raph1a
|
Ras association (RalGDS/AF-6) and pleckstrin homology domains 1a |
| chr7_-_71486162 | 0.35 |
ENSDART00000045253
|
aco1
|
aconitase 1, soluble |
| chr21_+_18405585 | 0.34 |
ENSDART00000139318
|
si:dkey-1d7.3
|
si:dkey-1d7.3 |
| chr12_+_13742778 | 0.34 |
ENSDART00000111401
ENSDART00000190552 |
ppp1r16a
|
protein phosphatase 1, regulatory subunit 16A |
| chr12_+_13742953 | 0.32 |
ENSDART00000152257
|
ppp1r16a
|
protein phosphatase 1, regulatory subunit 16A |
| chr16_+_14201401 | 0.32 |
ENSDART00000113679
|
dap3
|
death associated protein 3 |
| chr6_-_7686594 | 0.29 |
ENSDART00000091836
ENSDART00000151697 |
ubn2a
|
ubinuclein 2a |
| chr9_-_28990649 | 0.29 |
ENSDART00000078823
|
ptpn4a
|
protein tyrosine phosphatase, non-receptor type 4a |
| chr20_-_45058041 | 0.27 |
ENSDART00000124298
|
klhl29
|
kelch-like family member 29 |
| chr2_-_38312757 | 0.25 |
ENSDART00000167274
|
ap1g2
|
adaptor-related protein complex 1, gamma 2 subunit |
| chr9_+_26086135 | 0.25 |
ENSDART00000011910
|
arglu1a
|
arginine and glutamate rich 1a |
| chr24_-_36175365 | 0.23 |
ENSDART00000065338
|
pak1ip1
|
PAK1 interacting protein 1 |
| chr13_+_33368140 | 0.23 |
ENSDART00000033848
|
brf1a
|
BRF1, RNA polymerase III transcription initiation factor a |
| chr8_+_32722842 | 0.22 |
ENSDART00000147594
|
hmcn2
|
hemicentin 2 |
| chr22_-_11520405 | 0.22 |
ENSDART00000063157
|
slc26a11
|
solute carrier family 26 (anion exchanger), member 11 |
| chr24_-_18809433 | 0.21 |
ENSDART00000152009
|
arfgef1
|
ADP-ribosylation factor guanine nucleotide-exchange factor 1 (brefeldin A-inhibited) |
| chr20_+_22067337 | 0.20 |
ENSDART00000152636
|
clocka
|
clock circadian regulator a |
| chr19_-_15434813 | 0.19 |
ENSDART00000019843
|
ftr55
|
finTRIM family, member 55 |
| chr13_-_40120252 | 0.18 |
ENSDART00000157852
|
crtac1b
|
cartilage acidic protein 1b |
| chr10_+_23520952 | 0.17 |
ENSDART00000125317
|
zmp:0000000937
|
zmp:0000000937 |
| chr15_-_38218320 | 0.15 |
ENSDART00000152363
|
rhoga
|
ras homolog family member Ga |
| chr10_+_35468978 | 0.13 |
ENSDART00000131984
|
phldb2a
|
pleckstrin homology-like domain, family B, member 2a |
| chr15_+_37546391 | 0.13 |
ENSDART00000186625
|
psenen
|
presenilin enhancer gamma secretase subunit |
| chr22_+_2315996 | 0.13 |
ENSDART00000132489
|
znf1175
|
zinc finger protein 1175 |
| chr17_+_132555 | 0.13 |
ENSDART00000158159
|
zgc:77287
|
zgc:77287 |
| chr15_-_43270889 | 0.12 |
ENSDART00000166805
|
serpine2
|
serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 2 |
| chr23_+_39481537 | 0.12 |
ENSDART00000187222
|
mical1
|
microtubule associated monooxygenase, calponin and LIM domain containing 1 |
| chr13_+_9678427 | 0.12 |
ENSDART00000153907
|
sema4ga
|
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4Ga |
| chr17_-_44776224 | 0.12 |
ENSDART00000156195
|
zdhhc22
|
zinc finger, DHHC-type containing 22 |
| chr10_+_14488625 | 0.12 |
ENSDART00000026383
|
sigmar1
|
sigma non-opioid intracellular receptor 1 |
| chr15_+_40792667 | 0.11 |
ENSDART00000193186
ENSDART00000184559 |
fat3a
|
FAT atypical cadherin 3a |
| chr7_-_36113098 | 0.11 |
ENSDART00000064913
|
fto
|
fat mass and obesity associated |
| chr21_-_2158298 | 0.11 |
ENSDART00000182199
|
gb:ai877918
|
expressed sequence AI877918 |
| chr5_-_12823876 | 0.10 |
ENSDART00000019461
|
p2rx2
|
purinergic receptor P2X, ligand-gated ion channel, 2 |
| chr5_+_68112194 | 0.10 |
ENSDART00000162468
|
CABZ01083937.1
|
|
| chr14_+_22397251 | 0.09 |
ENSDART00000185239
ENSDART00000124072 ENSDART00000054977 |
atp7a
|
ATPase copper transporting alpha |
| chr11_-_24016761 | 0.09 |
ENSDART00000153601
ENSDART00000067817 ENSDART00000170531 |
chia.3
|
chitinase, acidic.3 |
| chr18_+_33284202 | 0.09 |
ENSDART00000132758
|
v2ra17
|
vomeronasal 2 receptor, a17 |
| chr1_+_532766 | 0.09 |
ENSDART00000179731
ENSDART00000060944 |
mrpl39
|
mitochondrial ribosomal protein L39 |
| chr8_-_25566347 | 0.08 |
ENSDART00000138289
ENSDART00000078022 |
prex1
|
phosphatidylinositol-3,4,5-trisphosphate-dependent Rac exchange factor 1 |
| chr18_+_5308392 | 0.08 |
ENSDART00000179072
|
DUT
|
deoxyuridine triphosphatase |
| chr11_+_24002503 | 0.07 |
ENSDART00000164702
|
chia.2
|
chitinase, acidic.2 |
| chr20_+_13783040 | 0.07 |
ENSDART00000115329
ENSDART00000152497 |
lpgat1
|
lysophosphatidylglycerol acyltransferase 1 |
| chr7_+_2244859 | 0.07 |
ENSDART00000114242
ENSDART00000064292 |
si:ch211-285c6.4
|
si:ch211-285c6.4 |
| chr3_+_1724941 | 0.07 |
ENSDART00000193402
|
BX321875.2
|
|
| chr13_+_25549425 | 0.07 |
ENSDART00000087553
ENSDART00000169199 |
sec23ip
|
SEC23 interacting protein |
| chr8_+_7801060 | 0.07 |
ENSDART00000161618
|
tfe3a
|
transcription factor binding to IGHM enhancer 3a |
| chr25_+_19008497 | 0.06 |
ENSDART00000104420
|
samm50
|
SAMM50 sorting and assembly machinery component |
| chr3_-_49815223 | 0.06 |
ENSDART00000181208
|
gcgra
|
glucagon receptor a |
| chr5_+_26806141 | 0.06 |
ENSDART00000142753
|
rabl6b
|
RAB, member RAS oncogene family-like 6b |
| chr9_-_22076368 | 0.06 |
ENSDART00000128486
|
crygm2a
|
crystallin, gamma M2a |
| chr3_-_5067585 | 0.05 |
ENSDART00000169609
|
tefb
|
thyrotrophic embryonic factor b |
| chr4_+_14826299 | 0.05 |
ENSDART00000067035
|
shisal1a
|
shisa like 1a |
| chr12_-_35095414 | 0.05 |
ENSDART00000153229
|
si:dkey-21e13.3
|
si:dkey-21e13.3 |
| chr5_+_13326765 | 0.05 |
ENSDART00000090851
|
ydjc
|
YdjC chitooligosaccharide deacetylase homolog |
| chr13_-_3324764 | 0.05 |
ENSDART00000102748
ENSDART00000114040 |
ubr2
|
ubiquitin protein ligase E3 component n-recognin 2 |
| chr15_+_15173611 | 0.04 |
ENSDART00000155267
|
si:ch211-149e23.4
|
si:ch211-149e23.4 |
| chr14_+_26805388 | 0.04 |
ENSDART00000044389
ENSDART00000173126 |
klhl4
|
kelch-like family member 4 |
| chr5_-_41841892 | 0.04 |
ENSDART00000167089
|
si:dkey-65b12.6
|
si:dkey-65b12.6 |
| chr24_-_21588914 | 0.04 |
ENSDART00000152054
|
gpr12
|
G protein-coupled receptor 12 |
| chr21_+_40307476 | 0.04 |
ENSDART00000132774
|
si:ch211-218m3.19
|
si:ch211-218m3.19 |
| chr15_-_37834433 | 0.04 |
ENSDART00000189748
|
si:dkey-238d18.3
|
si:dkey-238d18.3 |
| chr1_+_11691722 | 0.04 |
ENSDART00000134531
|
eif3bb
|
eukaryotic translation initiation factor 3, subunit Bb |
| chr13_+_25412307 | 0.03 |
ENSDART00000175339
|
calhm1
|
calcium homeostasis modulator 1 |
| chr13_+_9678262 | 0.03 |
ENSDART00000111365
|
sema4ga
|
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4Ga |
| chr10_+_40633990 | 0.03 |
ENSDART00000190489
ENSDART00000139474 |
si:ch211-238p8.31
|
si:ch211-238p8.31 |
| chr11_+_1602916 | 0.03 |
ENSDART00000184434
ENSDART00000112597 ENSDART00000192165 |
si:dkey-40c23.2
si:dkey-40c23.3
|
si:dkey-40c23.2 si:dkey-40c23.3 |
| chr15_-_31406093 | 0.02 |
ENSDART00000123444
|
or111-8
|
odorant receptor, family D, subfamily 111, member 8 |
| chr5_+_40224938 | 0.02 |
ENSDART00000142897
|
si:dkey-193c22.2
|
si:dkey-193c22.2 |
| chr10_+_9410304 | 0.02 |
ENSDART00000080843
|
ndufa8
|
NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 8 |
| chr9_+_35017702 | 0.02 |
ENSDART00000193640
|
gabpa
|
GA binding protein transcription factor, alpha subunit |
| chr21_-_43094210 | 0.02 |
ENSDART00000144747
|
si:ch73-68b22.2
|
si:ch73-68b22.2 |
| chr7_-_40657831 | 0.01 |
ENSDART00000084153
|
nom1
|
nucleolar protein with MIF4G domain 1 |
| chr4_-_27301356 | 0.01 |
ENSDART00000100444
|
fam19a5a
|
family with sequence similarity 19 (chemokine (C-C motif)-like), member A5a |
| chr10_+_21718468 | 0.01 |
ENSDART00000126622
|
pcdh1gb9
|
protocadherin 1 gamma b 9 |
| chr2_+_49631850 | 0.01 |
ENSDART00000114274
ENSDART00000114516 |
si:dkey-53k12.18
|
si:dkey-53k12.18 |
| chr19_-_22664128 | 0.01 |
ENSDART00000136295
|
eif3eb
|
eukaryotic translation initiation factor 3, subunit E, b |
| chr16_+_29630965 | 0.01 |
ENSDART00000185820
ENSDART00000192105 ENSDART00000153683 ENSDART00000186713 |
VPS72
|
si:ch211-203d17.1 |
| chr18_+_33322515 | 0.00 |
ENSDART00000136603
|
v2ra18
|
vomeronasal 2 receptor, a18 |
| chr5_-_24112175 | 0.00 |
ENSDART00000131595
|
si:ch211-114c12.5
|
si:ch211-114c12.5 |
| chr4_+_74396786 | 0.00 |
ENSDART00000127501
ENSDART00000174347 |
zmp:0000001020
|
zmp:0000001020 |
| chr10_+_23441488 | 0.00 |
ENSDART00000100783
|
si:ch73-122g19.1
|
si:ch73-122g19.1 |
| chr6_+_41186320 | 0.00 |
ENSDART00000025241
|
opn1mw2
|
opsin 1 (cone pigments), medium-wave-sensitive, 2 |
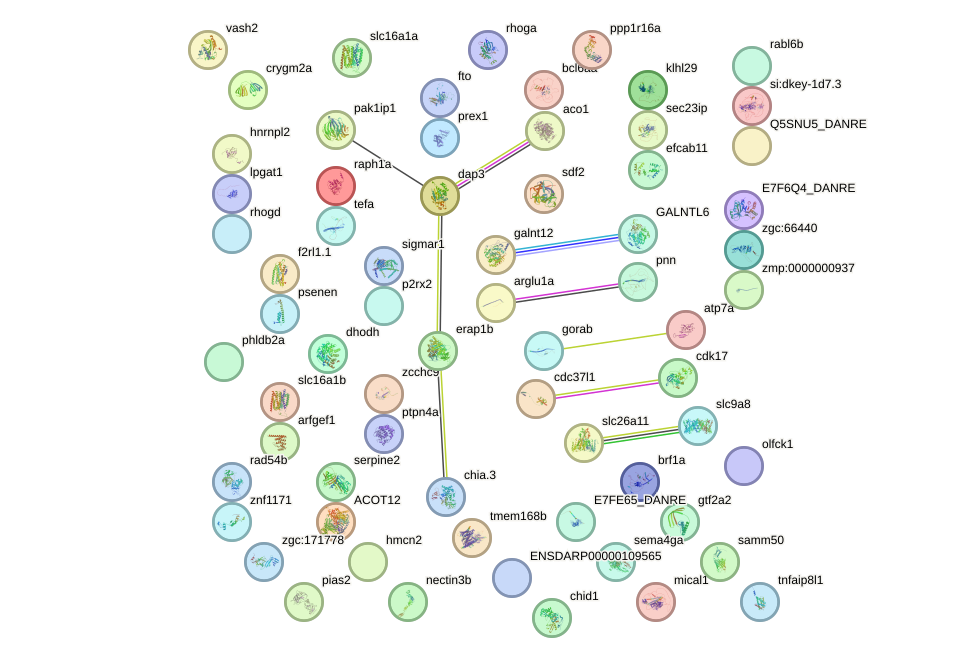
Network of associatons between targets according to the STRING database.
First level regulatory network of six3a+six3b+six7_six1a+six2a+six2b+six5+six9_six1b
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 2.1 | GO:0070493 | thrombin receptor signaling pathway(GO:0070493) |
| 0.2 | 0.9 | GO:0042762 | regulation of sulfur metabolic process(GO:0042762) |
| 0.1 | 0.7 | GO:0044205 | 'de novo' UMP biosynthetic process(GO:0044205) |
| 0.1 | 0.5 | GO:0006212 | uracil catabolic process(GO:0006212) uracil metabolic process(GO:0019860) |
| 0.1 | 0.4 | GO:0006651 | diacylglycerol biosynthetic process(GO:0006651) |
| 0.1 | 0.3 | GO:0010039 | response to iron ion(GO:0010039) |
| 0.1 | 1.7 | GO:0010923 | negative regulation of phosphatase activity(GO:0010923) |
| 0.1 | 1.2 | GO:0035803 | binding of sperm to zona pellucida(GO:0007339) egg coat formation(GO:0035803) regulation of acrosome reaction(GO:0060046) positive regulation of acrosome reaction(GO:2000344) |
| 0.1 | 1.7 | GO:0016925 | protein sumoylation(GO:0016925) |
| 0.0 | 0.5 | GO:0019883 | antigen processing and presentation of endogenous peptide antigen(GO:0002483) antigen processing and presentation of endogenous antigen(GO:0019883) antigen processing and presentation of endogenous peptide antigen via MHC class I(GO:0019885) |
| 0.0 | 0.7 | GO:0071712 | protein O-linked mannosylation(GO:0035269) ER-associated misfolded protein catabolic process(GO:0071712) |
| 0.0 | 0.4 | GO:0000083 | regulation of transcription involved in G1/S transition of mitotic cell cycle(GO:0000083) |
| 0.0 | 0.1 | GO:0019417 | sulfur oxidation(GO:0019417) |
| 0.0 | 0.5 | GO:0051123 | RNA polymerase II transcriptional preinitiation complex assembly(GO:0051123) |
| 0.0 | 1.3 | GO:0050821 | protein stabilization(GO:0050821) |
| 0.0 | 0.1 | GO:0042245 | RNA repair(GO:0042245) |
| 0.0 | 0.4 | GO:0006991 | response to sterol depletion(GO:0006991) SREBP signaling pathway(GO:0032933) cellular response to sterol depletion(GO:0071501) |
| 0.0 | 0.2 | GO:0090497 | mesenchymal cell migration(GO:0090497) |
| 0.0 | 0.2 | GO:0070897 | DNA-templated transcriptional preinitiation complex assembly(GO:0070897) |
| 0.0 | 0.1 | GO:0007008 | outer mitochondrial membrane organization(GO:0007008) protein import into mitochondrial outer membrane(GO:0045040) |
| 0.0 | 0.1 | GO:0035434 | copper ion transmembrane transport(GO:0035434) |
| 0.0 | 0.1 | GO:0036149 | phosphatidylinositol acyl-chain remodeling(GO:0036149) |
| 0.0 | 0.2 | GO:0006032 | chitin catabolic process(GO:0006032) |
| 0.0 | 0.1 | GO:0042987 | Notch receptor processing(GO:0007220) beta-amyloid formation(GO:0034205) amyloid precursor protein catabolic process(GO:0042987) |
| 0.0 | 0.8 | GO:0007131 | reciprocal meiotic recombination(GO:0007131) |
| 0.0 | 0.4 | GO:0007286 | spermatid development(GO:0007286) |
| 0.0 | 0.6 | GO:0032007 | negative regulation of TOR signaling(GO:0032007) |
| 0.0 | 0.3 | GO:0006896 | Golgi to vacuole transport(GO:0006896) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0005672 | transcription factor TFIIA complex(GO:0005672) |
| 0.1 | 0.5 | GO:0033557 | Slx1-Slx4 complex(GO:0033557) |
| 0.1 | 1.3 | GO:0000314 | organellar small ribosomal subunit(GO:0000314) mitochondrial small ribosomal subunit(GO:0005763) |
| 0.0 | 0.2 | GO:1990513 | CLOCK-BMAL transcription complex(GO:1990513) |
| 0.0 | 0.3 | GO:0030121 | AP-1 adaptor complex(GO:0030121) |
| 0.0 | 0.2 | GO:0000126 | transcription factor TFIIIB complex(GO:0000126) |
| 0.0 | 0.5 | GO:0030057 | desmosome(GO:0030057) |
| 0.0 | 0.1 | GO:0070765 | gamma-secretase complex(GO:0070765) |
| 0.0 | 0.4 | GO:0012507 | ER to Golgi transport vesicle membrane(GO:0012507) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 2.1 | GO:0015057 | thrombin receptor activity(GO:0015057) |
| 0.2 | 0.7 | GO:0016635 | oxidoreductase activity, acting on the CH-CH group of donors, quinone or related compound as acceptor(GO:0016635) |
| 0.2 | 0.7 | GO:0017020 | myosin phosphatase regulator activity(GO:0017020) |
| 0.1 | 0.8 | GO:0015616 | DNA translocase activity(GO:0015616) |
| 0.1 | 1.7 | GO:0061665 | SUMO ligase activity(GO:0061665) |
| 0.1 | 0.3 | GO:0003994 | aconitate hydratase activity(GO:0003994) |
| 0.1 | 0.5 | GO:0017113 | uracil binding(GO:0002058) pyrimidine nucleobase binding(GO:0002061) dihydropyrimidine dehydrogenase (NADP+) activity(GO:0017113) |
| 0.1 | 1.2 | GO:0035804 | structural constituent of egg coat(GO:0035804) |
| 0.1 | 0.5 | GO:0008821 | crossover junction endodeoxyribonuclease activity(GO:0008821) |
| 0.1 | 0.8 | GO:0005451 | monovalent cation:proton antiporter activity(GO:0005451) sodium:proton antiporter activity(GO:0015385) potassium:proton antiporter activity(GO:0015386) |
| 0.1 | 0.9 | GO:0015129 | lactate transmembrane transporter activity(GO:0015129) |
| 0.1 | 0.7 | GO:0004169 | dolichyl-phosphate-mannose-protein mannosyltransferase activity(GO:0004169) |
| 0.0 | 0.7 | GO:0017025 | TBP-class protein binding(GO:0017025) |
| 0.0 | 0.5 | GO:0008061 | chitin binding(GO:0008061) |
| 0.0 | 0.2 | GO:0070888 | E-box binding(GO:0070888) |
| 0.0 | 0.1 | GO:0035516 | oxidative DNA demethylase activity(GO:0035516) DNA-N1-methyladenine dioxygenase activity(GO:0043734) RNA N6-methyladenosine dioxygenase activity(GO:1990931) |
| 0.0 | 1.3 | GO:0031072 | heat shock protein binding(GO:0031072) |
| 0.0 | 0.4 | GO:0016411 | acylglycerol O-acyltransferase activity(GO:0016411) |
| 0.0 | 0.1 | GO:0005375 | copper ion transmembrane transporter activity(GO:0005375) |
| 0.0 | 0.5 | GO:0070006 | metalloaminopeptidase activity(GO:0070006) |
| 0.0 | 1.1 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.0 | 0.1 | GO:0016519 | gastric inhibitory peptide receptor activity(GO:0016519) |
| 0.0 | 0.3 | GO:0098748 | clathrin adaptor activity(GO:0035615) endocytic adaptor activity(GO:0098748) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.7 | PID IL12 STAT4 PATHWAY | IL12 signaling mediated by STAT4 |
| 0.0 | 0.4 | SA CASPASE CASCADE | Apoptosis is mediated by caspases, cysteine proteases arranged in a proteolytic cascade. |
| 0.0 | 0.3 | PID TRAIL PATHWAY | TRAIL signaling pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.5 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |
| 0.0 | 0.8 | REACTOME PYRIMIDINE METABOLISM | Genes involved in Pyrimidine metabolism |
| 0.0 | 0.5 | REACTOME RNA POL II TRANSCRIPTION PRE INITIATION AND PROMOTER OPENING | Genes involved in RNA Polymerase II Transcription Pre-Initiation And Promoter Opening |
| 0.0 | 0.1 | REACTOME SIGNALING BY NOTCH4 | Genes involved in Signaling by NOTCH4 |
| 0.0 | 1.5 | REACTOME MRNA SPLICING | Genes involved in mRNA Splicing |