Project
PRJNA438478: RNAseq of wild type zebrafish germline
Navigation
Downloads
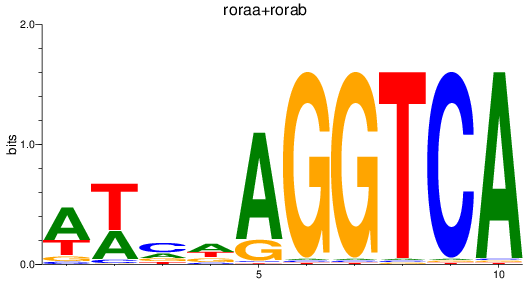
Results for roraa+rorab
Z-value: 0.28
Transcription factors associated with roraa+rorab
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
rorab
|
ENSDARG00000001910 | RAR-related orphan receptor A, paralog b |
|
roraa
|
ENSDARG00000031768 | RAR-related orphan receptor A, paralog a |
|
rorab
|
ENSDARG00000109324 | RAR-related orphan receptor A, paralog b |
|
rorab
|
ENSDARG00000113275 | RAR-related orphan receptor A, paralog b |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| roraa | dr11_v1_chr25_+_33849647_33849647 | -0.67 | 2.4e-03 | Click! |
| rorab | dr11_v1_chr7_-_29571615_29571615 | 0.57 | 1.4e-02 | Click! |
Activity profile of roraa+rorab motif
Sorted Z-values of roraa+rorab motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr13_+_50375800 | 1.12 |
ENSDART00000099537
|
cox5b2
|
cytochrome c oxidase subunit Vb 2 |
| chr5_-_29780752 | 0.68 |
ENSDART00000137400
ENSDART00000145021 |
cfap77
|
cilia and flagella associated protein 77 |
| chr1_-_44704261 | 0.58 |
ENSDART00000133210
|
si:dkey-28b4.8
|
si:dkey-28b4.8 |
| chr18_-_29962234 | 0.52 |
ENSDART00000144996
|
si:ch73-103b9.2
|
si:ch73-103b9.2 |
| chr25_+_3326885 | 0.50 |
ENSDART00000104866
|
ldhbb
|
lactate dehydrogenase Bb |
| chr21_+_11885404 | 0.50 |
ENSDART00000092015
|
dcaf12
|
DDB1 and CUL4 associated factor 12 |
| chr5_-_69523816 | 0.49 |
ENSDART00000112692
|
si:ch211-157p22.10
|
si:ch211-157p22.10 |
| chr3_+_1182315 | 0.38 |
ENSDART00000055430
|
ndufa6
|
NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 6 |
| chr11_-_37995501 | 0.36 |
ENSDART00000192096
|
slc41a1
|
solute carrier family 41 (magnesium transporter), member 1 |
| chr11_+_5842632 | 0.36 |
ENSDART00000111374
ENSDART00000158599 |
ndufs7
|
NADH dehydrogenase (ubiquinone) Fe-S protein 7, (NADH-coenzyme Q reductase) |
| chr6_-_49873020 | 0.35 |
ENSDART00000148511
|
gnas
|
GNAS complex locus |
| chr6_+_30689239 | 0.35 |
ENSDART00000065206
|
wdr78
|
WD repeat domain 78 |
| chr25_+_3327071 | 0.35 |
ENSDART00000136131
ENSDART00000133243 |
ldhbb
|
lactate dehydrogenase Bb |
| chr16_+_41517188 | 0.34 |
ENSDART00000049976
|
si:dkey-11p23.7
|
si:dkey-11p23.7 |
| chr1_+_54115839 | 0.34 |
ENSDART00000180214
|
LO017722.2
|
|
| chr5_+_20693724 | 0.33 |
ENSDART00000141368
|
si:ch211-240b21.2
|
si:ch211-240b21.2 |
| chr23_+_42813415 | 0.33 |
ENSDART00000055577
|
myl9a
|
myosin, light chain 9a, regulatory |
| chr8_+_25902170 | 0.33 |
ENSDART00000193130
|
rhoab
|
ras homolog gene family, member Ab |
| chr6_-_10835849 | 0.31 |
ENSDART00000005903
ENSDART00000135065 |
atp5mc3b
|
ATP synthase membrane subunit c locus 3b |
| chr4_+_7677318 | 0.29 |
ENSDART00000149218
|
elk3
|
ELK3, ETS-domain protein |
| chr6_-_9922266 | 0.29 |
ENSDART00000151549
|
pimr73
|
Pim proto-oncogene, serine/threonine kinase, related 73 |
| chr16_-_28856112 | 0.28 |
ENSDART00000078543
|
syt11b
|
synaptotagmin XIb |
| chr13_-_29406534 | 0.28 |
ENSDART00000100877
|
zgc:153142
|
zgc:153142 |
| chr3_+_58167288 | 0.28 |
ENSDART00000155874
ENSDART00000010395 |
uqcrc2a
|
ubiquinol-cytochrome c reductase core protein 2a |
| chr14_+_23518110 | 0.25 |
ENSDART00000112930
|
si:ch211-221f10.2
|
si:ch211-221f10.2 |
| chr21_+_20771082 | 0.25 |
ENSDART00000079732
|
oxct1b
|
3-oxoacid CoA transferase 1b |
| chr22_-_24285432 | 0.24 |
ENSDART00000164083
|
si:ch211-117l17.4
|
si:ch211-117l17.4 |
| chr8_+_14158021 | 0.24 |
ENSDART00000080832
|
si:dkey-6n6.2
|
si:dkey-6n6.2 |
| chr18_+_7594012 | 0.24 |
ENSDART00000062150
|
zgc:77752
|
zgc:77752 |
| chr9_-_21067971 | 0.23 |
ENSDART00000004333
|
tbx15
|
T-box 15 |
| chr12_-_47648538 | 0.23 |
ENSDART00000108477
|
fh
|
fumarate hydratase |
| chr16_+_21242491 | 0.22 |
ENSDART00000145886
|
osbpl3b
|
oxysterol binding protein-like 3b |
| chr5_+_41322783 | 0.22 |
ENSDART00000097546
|
arid3c
|
AT rich interactive domain 3C (BRIGHT-like) |
| chr14_+_33882973 | 0.21 |
ENSDART00000019396
|
clic2
|
chloride intracellular channel 2 |
| chr7_-_24046999 | 0.19 |
ENSDART00000144616
ENSDART00000124653 ENSDART00000127813 |
dhrs4
|
dehydrogenase/reductase (SDR family) member 4 |
| chr16_-_45327616 | 0.19 |
ENSDART00000158733
|
si:dkey-33i11.1
|
si:dkey-33i11.1 |
| chr6_-_39764995 | 0.19 |
ENSDART00000085277
|
pfkmb
|
phosphofructokinase, muscle b |
| chr18_+_14529005 | 0.19 |
ENSDART00000186379
|
kcng4a
|
potassium voltage-gated channel, subfamily G, member 4a |
| chr20_+_16750177 | 0.18 |
ENSDART00000185357
|
calm1b
|
calmodulin 1b |
| chr11_+_24900123 | 0.18 |
ENSDART00000044987
ENSDART00000148023 |
timm17a
|
translocase of inner mitochondrial membrane 17 homolog A (yeast) |
| chr23_+_4324625 | 0.18 |
ENSDART00000146302
ENSDART00000136792 ENSDART00000135027 ENSDART00000179819 |
sgk2a
|
serum/glucocorticoid regulated kinase 2a |
| chr21_-_22724980 | 0.18 |
ENSDART00000035469
|
c1qa
|
complement component 1, q subcomponent, A chain |
| chr9_-_53666031 | 0.18 |
ENSDART00000126314
|
pcdh8
|
protocadherin 8 |
| chr22_+_18816662 | 0.17 |
ENSDART00000132476
|
cbarpb
|
calcium channel, voltage-dependent, beta subunit associated regulatory protein b |
| chr9_+_22359919 | 0.17 |
ENSDART00000009591
|
crygs4
|
crystallin, gamma S4 |
| chr13_-_31452516 | 0.17 |
ENSDART00000193268
|
rtn1a
|
reticulon 1a |
| chr7_+_20475788 | 0.17 |
ENSDART00000171155
|
si:dkey-19b23.13
|
si:dkey-19b23.13 |
| chr7_+_48805534 | 0.17 |
ENSDART00000145375
ENSDART00000148744 |
cpt1aa
|
carnitine palmitoyltransferase 1Aa (liver) |
| chr7_+_17255705 | 0.17 |
ENSDART00000053357
ENSDART00000167849 ENSDART00000165833 |
nitr9
si:ch73-46n24.1
|
novel immune-type receptor 9 si:ch73-46n24.1 |
| chr17_-_53353653 | 0.17 |
ENSDART00000180744
ENSDART00000026879 |
unm_sa911
|
un-named sa911 |
| chr22_-_17574511 | 0.16 |
ENSDART00000181496
|
pip5k1ca
|
phosphatidylinositol-4-phosphate 5-kinase, type I, gamma a |
| chr17_-_26610814 | 0.16 |
ENSDART00000133402
ENSDART00000016608 |
mrpl57
|
mitochondrial ribosomal protein L57 |
| chr10_-_17103651 | 0.16 |
ENSDART00000108959
|
RNF208
|
ring finger protein 208 |
| chr5_+_31860043 | 0.16 |
ENSDART00000036235
ENSDART00000140541 |
iscub
|
iron-sulfur cluster assembly enzyme b |
| chr22_-_5252005 | 0.15 |
ENSDART00000132942
ENSDART00000081801 |
ncln
|
nicalin |
| chr10_-_7988396 | 0.15 |
ENSDART00000141445
ENSDART00000024282 |
ewsr1a
|
EWS RNA-binding protein 1a |
| chr8_+_30664077 | 0.15 |
ENSDART00000138750
|
adora2aa
|
adenosine A2a receptor a |
| chr1_+_580642 | 0.15 |
ENSDART00000147633
|
mrpl39
|
mitochondrial ribosomal protein L39 |
| chr20_-_14680897 | 0.14 |
ENSDART00000063857
ENSDART00000161314 |
scrn2
|
secernin 2 |
| chr10_+_17592273 | 0.14 |
ENSDART00000141221
|
KCNN2
|
si:dkey-76p7.5 |
| chr8_+_18464235 | 0.14 |
ENSDART00000110571
|
alkal1
|
ALK and LTK ligand 1 |
| chr18_+_36631923 | 0.14 |
ENSDART00000098980
|
znf296
|
zinc finger protein 296 |
| chr7_-_24005268 | 0.14 |
ENSDART00000173608
|
si:dkey-183c6.9
|
si:dkey-183c6.9 |
| chr19_-_5103141 | 0.14 |
ENSDART00000150952
|
tpi1a
|
triosephosphate isomerase 1a |
| chr3_-_39198113 | 0.14 |
ENSDART00000102690
|
RETSAT
|
zgc:154169 |
| chr1_+_31725154 | 0.14 |
ENSDART00000112333
ENSDART00000189801 |
cnnm2b
|
cyclin and CBS domain divalent metal cation transport mediator 2b |
| chr7_+_40228422 | 0.13 |
ENSDART00000052222
|
ptprn2
|
protein tyrosine phosphatase, receptor type, N polypeptide 2 |
| chr19_+_40856534 | 0.13 |
ENSDART00000051950
|
gngt1
|
guanine nucleotide binding protein (G protein), gamma transducing activity polypeptide 1 |
| chr1_+_12763920 | 0.13 |
ENSDART00000189465
|
pcdh10a
|
protocadherin 10a |
| chr1_-_50859053 | 0.13 |
ENSDART00000132779
ENSDART00000137648 |
si:dkeyp-123h10.2
|
si:dkeyp-123h10.2 |
| chr19_-_5103313 | 0.13 |
ENSDART00000037007
|
tpi1a
|
triosephosphate isomerase 1a |
| chr6_-_1514767 | 0.13 |
ENSDART00000067586
|
chchd6b
|
coiled-coil-helix-coiled-coil-helix domain containing 6b |
| chr25_-_1235457 | 0.13 |
ENSDART00000093093
|
coro2bb
|
coronin, actin binding protein, 2Bb |
| chr25_-_21156678 | 0.13 |
ENSDART00000156257
|
wnk1a
|
WNK lysine deficient protein kinase 1a |
| chr2_-_127945 | 0.13 |
ENSDART00000056453
|
igfbp1b
|
insulin-like growth factor binding protein 1b |
| chr14_-_33454595 | 0.13 |
ENSDART00000109615
ENSDART00000173267 ENSDART00000185737 ENSDART00000190989 |
tmem255a
|
transmembrane protein 255A |
| chr14_-_29859067 | 0.12 |
ENSDART00000136380
|
sorbs2b
|
sorbin and SH3 domain containing 2b |
| chr5_+_36870737 | 0.12 |
ENSDART00000145182
|
slc8a2a
|
solute carrier family 8 (sodium/calcium exchanger), member 2a |
| chr8_+_18463797 | 0.12 |
ENSDART00000148958
|
alkal1
|
ALK and LTK ligand 1 |
| chr18_-_16922905 | 0.12 |
ENSDART00000187165
|
wee1
|
WEE1 G2 checkpoint kinase |
| chr2_-_56655769 | 0.12 |
ENSDART00000113589
|
gpx4b
|
glutathione peroxidase 4b |
| chr13_-_15082024 | 0.12 |
ENSDART00000157482
|
sfxn5a
|
sideroflexin 5a |
| chr14_+_48045193 | 0.12 |
ENSDART00000124773
|
ppid
|
peptidylprolyl isomerase D |
| chr25_+_31264155 | 0.12 |
ENSDART00000012256
|
tnni2a.3
|
troponin I type 2a (skeletal, fast), tandem duplicate 3 |
| chr15_-_30450898 | 0.12 |
ENSDART00000156584
|
msi2b
|
musashi RNA-binding protein 2b |
| chr24_-_12770357 | 0.12 |
ENSDART00000060826
|
ipo4
|
importin 4 |
| chr3_-_15889508 | 0.11 |
ENSDART00000148363
|
cramp1
|
cramped chromatin regulator homolog 1 |
| chr7_+_29133321 | 0.11 |
ENSDART00000052346
|
gnao1b
|
guanine nucleotide binding protein (G protein), alpha activating activity polypeptide O, b |
| chr21_+_7582036 | 0.11 |
ENSDART00000135485
ENSDART00000027268 |
otpa
|
orthopedia homeobox a |
| chr8_+_8012570 | 0.11 |
ENSDART00000183429
|
si:ch211-169p10.1
|
si:ch211-169p10.1 |
| chr15_-_25304710 | 0.11 |
ENSDART00000190690
|
pafah1b1a
|
platelet-activating factor acetylhydrolase 1b, regulatory subunit 1a |
| chr21_-_20840714 | 0.11 |
ENSDART00000144861
ENSDART00000139430 |
c6
|
complement component 6 |
| chr10_+_40973524 | 0.11 |
ENSDART00000176660
ENSDART00000114966 |
antxr1b
|
anthrax toxin receptor 1b |
| chr7_+_40884012 | 0.11 |
ENSDART00000149395
|
shha
|
sonic hedgehog a |
| chr6_-_39765546 | 0.10 |
ENSDART00000185767
|
pfkmb
|
phosphofructokinase, muscle b |
| chr23_+_19564392 | 0.10 |
ENSDART00000144746
|
atp6ap1lb
|
ATPase H+ transporting accessory protein 1 like b |
| chr19_+_1465004 | 0.10 |
ENSDART00000159157
|
CABZ01073736.1
|
|
| chr8_+_26034623 | 0.10 |
ENSDART00000004521
ENSDART00000142555 |
arih2
|
ariadne homolog 2 (Drosophila) |
| chr3_+_12484008 | 0.10 |
ENSDART00000182229
|
vasnb
|
vasorin b |
| chr4_-_1360495 | 0.10 |
ENSDART00000164623
|
ptn
|
pleiotrophin |
| chr11_+_28476298 | 0.10 |
ENSDART00000122319
|
lrrc38b
|
leucine rich repeat containing 38b |
| chr12_-_25201576 | 0.10 |
ENSDART00000077188
|
cox7a3
|
cytochrome c oxidase subunit VIIa polypeptide 3 |
| chr6_+_3710865 | 0.10 |
ENSDART00000170781
|
phospho2
|
phosphatase, orphan 2 |
| chr12_+_28799988 | 0.09 |
ENSDART00000022724
|
pnpo
|
pyridoxamine 5'-phosphate oxidase |
| chr4_-_17629444 | 0.09 |
ENSDART00000108814
|
nrip2
|
nuclear receptor interacting protein 2 |
| chr1_+_23783349 | 0.09 |
ENSDART00000007531
|
slit2
|
slit homolog 2 (Drosophila) |
| chr16_+_27349585 | 0.09 |
ENSDART00000142573
|
nr4a3
|
nuclear receptor subfamily 4, group A, member 3 |
| chr19_+_37620342 | 0.09 |
ENSDART00000158960
|
thsd7aa
|
thrombospondin, type I, domain containing 7Aa |
| chr3_-_26109322 | 0.09 |
ENSDART00000113780
|
zgc:162612
|
zgc:162612 |
| chr6_-_40312091 | 0.09 |
ENSDART00000154878
|
col7a1
|
collagen, type VII, alpha 1 |
| chr22_-_834106 | 0.08 |
ENSDART00000105873
|
cry4
|
cryptochrome circadian clock 4 |
| chr10_+_4924388 | 0.08 |
ENSDART00000108595
|
slc46a2
|
solute carrier family 46 member 2 |
| chr7_-_48805181 | 0.08 |
ENSDART00000015884
|
mfge8a
|
milk fat globule-EGF factor 8 protein a |
| chr7_-_16237861 | 0.08 |
ENSDART00000173647
|
btr07
|
bloodthirsty-related gene family, member 7 |
| chr22_-_11124419 | 0.08 |
ENSDART00000149634
|
atp6ap2
|
ATPase H+ transporting accessory protein 2 |
| chr7_+_17303944 | 0.08 |
ENSDART00000163505
ENSDART00000162169 ENSDART00000161291 |
nitr9
|
novel immune-type receptor 9 |
| chr1_+_34496855 | 0.08 |
ENSDART00000012873
|
klf12a
|
Kruppel-like factor 12a |
| chr21_+_39432248 | 0.07 |
ENSDART00000179938
|
pafah1b1b
|
platelet-activating factor acetylhydrolase 1b, regulatory subunit 1b |
| chr23_+_44741500 | 0.07 |
ENSDART00000166421
|
atp1b2a
|
ATPase Na+/K+ transporting subunit beta 2a |
| chr19_+_30662529 | 0.07 |
ENSDART00000175662
|
fam49al
|
family with sequence similarity 49, member A-like |
| chr10_+_1638876 | 0.07 |
ENSDART00000184484
ENSDART00000060946 ENSDART00000181251 |
sgsm1b
|
small G protein signaling modulator 1b |
| chr17_-_36529016 | 0.07 |
ENSDART00000025019
|
colec11
|
collectin sub-family member 11 |
| chr15_+_29362714 | 0.07 |
ENSDART00000099916
ENSDART00000126196 |
gucy2f
|
guanylate cyclase 2F, retinal |
| chr11_+_24819324 | 0.07 |
ENSDART00000184549
|
kdm5ba
|
lysine (K)-specific demethylase 5Ba |
| chr11_-_29650930 | 0.07 |
ENSDART00000166969
|
chd5
|
chromodomain helicase DNA binding protein 5 |
| chr16_+_12236339 | 0.07 |
ENSDART00000132468
|
tpi1b
|
triosephosphate isomerase 1b |
| chr10_+_20128267 | 0.06 |
ENSDART00000064615
|
dmtn
|
dematin actin binding protein |
| chr2_+_58841181 | 0.06 |
ENSDART00000164102
|
cirbpa
|
cold inducible RNA binding protein a |
| chr19_+_40856807 | 0.06 |
ENSDART00000139083
|
gngt1
|
guanine nucleotide binding protein (G protein), gamma transducing activity polypeptide 1 |
| chr19_+_30990815 | 0.06 |
ENSDART00000134645
|
sync
|
syncoilin, intermediate filament protein |
| chr18_+_8340886 | 0.06 |
ENSDART00000081132
|
cpt1b
|
carnitine palmitoyltransferase 1B (muscle) |
| chr7_+_48805725 | 0.06 |
ENSDART00000166543
|
cpt1aa
|
carnitine palmitoyltransferase 1Aa (liver) |
| chr24_+_29449690 | 0.06 |
ENSDART00000105743
ENSDART00000193556 ENSDART00000145816 |
ntng1a
|
netrin g1a |
| chr1_+_9708801 | 0.06 |
ENSDART00000189621
|
elfn1b
|
extracellular leucine-rich repeat and fibronectin type III domain containing 1b |
| chr10_-_26273629 | 0.06 |
ENSDART00000147790
|
dchs1b
|
dachsous cadherin-related 1b |
| chr17_-_36818176 | 0.05 |
ENSDART00000061762
|
myo6b
|
myosin VIb |
| chr24_+_39186940 | 0.05 |
ENSDART00000155817
|
spsb3b
|
splA/ryanodine receptor domain and SOCS box containing 3b |
| chr19_+_2590182 | 0.05 |
ENSDART00000162293
|
si:ch73-345f18.3
|
si:ch73-345f18.3 |
| chr20_-_31018569 | 0.05 |
ENSDART00000136039
|
fndc1
|
fibronectin type III domain containing 1 |
| chr6_-_10320676 | 0.05 |
ENSDART00000151247
|
scn1lab
|
sodium channel, voltage-gated, type I like, alpha b |
| chr17_+_33340675 | 0.05 |
ENSDART00000184396
ENSDART00000077553 |
xdh
|
xanthine dehydrogenase |
| chr24_-_10919588 | 0.05 |
ENSDART00000131204
|
asap1b
|
ArfGAP with SH3 domain, ankyrin repeat and PH domain 1b |
| chr15_+_19797918 | 0.05 |
ENSDART00000113314
|
si:ch211-229d2.5
|
si:ch211-229d2.5 |
| chr7_-_26446726 | 0.05 |
ENSDART00000173607
|
zgc:172079
|
zgc:172079 |
| chr7_-_30227878 | 0.05 |
ENSDART00000173514
|
znf710b
|
zinc finger protein 710b |
| chr19_+_11214007 | 0.05 |
ENSDART00000127362
|
si:ch73-109i22.2
|
si:ch73-109i22.2 |
| chr25_-_19374710 | 0.05 |
ENSDART00000184483
ENSDART00000188706 |
map1ab
|
microtubule-associated protein 1Ab |
| chr5_-_8636168 | 0.05 |
ENSDART00000134877
|
fyba
|
FYN binding protein a |
| chr6_-_15603675 | 0.05 |
ENSDART00000143502
|
lrrfip1b
|
leucine rich repeat (in FLII) interacting protein 1b |
| chr9_+_23224761 | 0.05 |
ENSDART00000142008
|
map3k19
|
mitogen-activated protein kinase kinase kinase 19 |
| chr11_+_10975481 | 0.05 |
ENSDART00000160488
|
itgb6
|
integrin, beta 6 |
| chr15_+_31820536 | 0.05 |
ENSDART00000045921
|
frya
|
furry homolog a (Drosophila) |
| chr15_-_14486534 | 0.05 |
ENSDART00000179368
|
numbl
|
numb homolog (Drosophila)-like |
| chr17_-_36529449 | 0.04 |
ENSDART00000187252
|
colec11
|
collectin sub-family member 11 |
| chr9_+_41914378 | 0.04 |
ENSDART00000130434
ENSDART00000007058 |
col18a1a
|
collagen type XVIII alpha 1 chain a |
| chr12_-_45876387 | 0.04 |
ENSDART00000043210
ENSDART00000149044 |
pax2b
|
paired box 2b |
| chr10_+_19528321 | 0.04 |
ENSDART00000184816
|
vsig8a
|
V-set and immunoglobulin domain containing 8a |
| chr20_+_43623064 | 0.04 |
ENSDART00000186659
|
si:dkey-206f10.1
|
si:dkey-206f10.1 |
| chr11_+_7432533 | 0.04 |
ENSDART00000180977
|
adgrl2a
|
adhesion G protein-coupled receptor L2a |
| chr16_+_5678071 | 0.04 |
ENSDART00000011166
ENSDART00000134198 ENSDART00000131575 |
zgc:158689
|
zgc:158689 |
| chr13_+_33688474 | 0.04 |
ENSDART00000161465
|
CABZ01087953.1
|
|
| chr23_-_43486714 | 0.04 |
ENSDART00000169726
|
e2f1
|
E2F transcription factor 1 |
| chr19_+_30990129 | 0.03 |
ENSDART00000052169
ENSDART00000193376 |
sync
|
syncoilin, intermediate filament protein |
| chr3_-_21402279 | 0.03 |
ENSDART00000164513
|
CT573446.1
|
|
| chr18_+_50461981 | 0.03 |
ENSDART00000158761
|
CU896640.1
|
|
| chr12_+_27156943 | 0.03 |
ENSDART00000153030
ENSDART00000001737 |
skap1
|
src kinase associated phosphoprotein 1 |
| chr6_+_51773873 | 0.03 |
ENSDART00000156516
|
tmem74b
|
transmembrane protein 74B |
| chr3_-_19091024 | 0.03 |
ENSDART00000188485
ENSDART00000110554 |
grin2ca
|
glutamate receptor, ionotropic, N-methyl D-aspartate 2Ca |
| chr9_-_1970071 | 0.03 |
ENSDART00000080608
|
hoxd10a
|
homeobox D10a |
| chr2_-_44971551 | 0.03 |
ENSDART00000018818
|
mul1a
|
mitochondrial E3 ubiquitin protein ligase 1a |
| chr21_+_37411547 | 0.03 |
ENSDART00000076320
|
mrps17
|
mitochondrial ribosomal protein S17 |
| chr19_+_20724347 | 0.03 |
ENSDART00000090757
|
kat2b
|
K(lysine) acetyltransferase 2B |
| chr18_+_16330025 | 0.03 |
ENSDART00000142353
|
nts
|
neurotensin |
| chr5_+_1219187 | 0.03 |
ENSDART00000129490
|
gb:bc139872
|
expressed sequence BC139872 |
| chr18_-_1185772 | 0.03 |
ENSDART00000143245
|
nptnb
|
neuroplastin b |
| chr4_+_76353418 | 0.03 |
ENSDART00000171289
|
si:ch73-158p21.2
|
si:ch73-158p21.2 |
| chr7_-_4036184 | 0.03 |
ENSDART00000019949
|
ndrg2
|
NDRG family member 2 |
| chr25_-_31396479 | 0.03 |
ENSDART00000156828
|
prr33
|
proline rich 33 |
| chr3_+_24207243 | 0.03 |
ENSDART00000023454
ENSDART00000136400 |
adsl
|
adenylosuccinate lyase |
| chr22_+_20546612 | 0.02 |
ENSDART00000141852
|
si:dkey-172o19.2
|
si:dkey-172o19.2 |
| chr17_-_7371564 | 0.02 |
ENSDART00000060336
|
rab32b
|
RAB32b, member RAS oncogene family |
| chr3_+_13637383 | 0.02 |
ENSDART00000166000
|
si:ch211-194b1.1
|
si:ch211-194b1.1 |
| chr14_+_9583588 | 0.02 |
ENSDART00000164101
|
tmem129
|
transmembrane protein 129, E3 ubiquitin protein ligase |
| chr11_+_30729745 | 0.02 |
ENSDART00000103270
|
slc22a7a
|
solute carrier family 22 (organic anion transporter), member 7a |
| chr12_+_17439580 | 0.02 |
ENSDART00000189257
|
atad1b
|
ATPase family, AAA domain containing 1b |
| chr10_+_29840725 | 0.02 |
ENSDART00000127268
|
BX120005.1
|
|
| chr10_-_28395620 | 0.02 |
ENSDART00000168907
|
CR392341.4
|
|
| chr15_-_21239416 | 0.02 |
ENSDART00000155787
|
A2ML1 (1 of many)
|
si:dkey-105h12.2 |
| chr14_+_33525196 | 0.02 |
ENSDART00000085335
|
zdhhc9
|
zinc finger, DHHC-type containing 9 |
| chr14_-_29858883 | 0.02 |
ENSDART00000141034
|
sorbs2b
|
sorbin and SH3 domain containing 2b |
| chr23_-_24488696 | 0.02 |
ENSDART00000155593
|
tmem82
|
transmembrane protein 82 |
| chr4_+_2620751 | 0.02 |
ENSDART00000013924
|
gpr22a
|
G protein-coupled receptor 22a |
| chr16_+_42471455 | 0.02 |
ENSDART00000166640
|
si:ch211-215k15.5
|
si:ch211-215k15.5 |
| chr10_+_20113830 | 0.02 |
ENSDART00000139722
|
dmtn
|
dematin actin binding protein |
| chr8_+_51084993 | 0.02 |
ENSDART00000127709
ENSDART00000053768 |
DISP3
|
si:dkey-32e23.6 |
| chr11_-_12198765 | 0.02 |
ENSDART00000104203
ENSDART00000128364 ENSDART00000166887 ENSDART00000041533 |
krt95
|
kertain 95 |
| chr6_+_8339298 | 0.02 |
ENSDART00000151672
|
si:ch211-276a17.5
|
si:ch211-276a17.5 |
| chr7_-_29341233 | 0.02 |
ENSDART00000140938
ENSDART00000147251 |
trpm1a
|
transient receptor potential cation channel, subfamily M, member 1a |
| chr2_-_16380283 | 0.02 |
ENSDART00000149992
|
si:dkey-231j24.3
|
si:dkey-231j24.3 |
| chr5_-_55395964 | 0.02 |
ENSDART00000145791
|
prune2
|
prune homolog 2 (Drosophila) |
Network of associatons between targets according to the STRING database.
First level regulatory network of roraa+rorab
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.1 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
| 0.1 | 0.4 | GO:0015990 | energy coupled proton transmembrane transport, against electrochemical gradient(GO:0015988) electron transport coupled proton transport(GO:0015990) |
| 0.1 | 0.3 | GO:0019242 | methylglyoxal biosynthetic process(GO:0019242) glyceraldehyde-3-phosphate biosynthetic process(GO:0046166) |
| 0.1 | 0.4 | GO:0007191 | adenylate cyclase-activating dopamine receptor signaling pathway(GO:0007191) |
| 0.1 | 0.3 | GO:0045124 | regulation of bone resorption(GO:0045124) |
| 0.1 | 0.3 | GO:0070376 | ERK5 cascade(GO:0070375) regulation of ERK5 cascade(GO:0070376) positive regulation of ERK5 cascade(GO:0070378) |
| 0.0 | 0.2 | GO:0046952 | ketone body catabolic process(GO:0046952) |
| 0.0 | 0.3 | GO:1902766 | skeletal muscle satellite cell migration(GO:1902766) |
| 0.0 | 0.2 | GO:0051012 | microtubule sliding(GO:0051012) |
| 0.0 | 0.5 | GO:0010960 | magnesium ion homeostasis(GO:0010960) |
| 0.0 | 0.2 | GO:0061181 | regulation of chondrocyte development(GO:0061181) |
| 0.0 | 0.1 | GO:0061469 | regulation of type B pancreatic cell proliferation(GO:0061469) |
| 0.0 | 0.1 | GO:2000171 | negative regulation of dendrite development(GO:2000171) |
| 0.0 | 0.1 | GO:0042823 | water-soluble vitamin biosynthetic process(GO:0042364) pyridoxal phosphate metabolic process(GO:0042822) pyridoxal phosphate biosynthetic process(GO:0042823) |
| 0.0 | 0.1 | GO:0007414 | axonal defasciculation(GO:0007414) |
| 0.0 | 0.2 | GO:0038107 | nodal signaling pathway involved in determination of left/right asymmetry(GO:0038107) regulation of nodal signaling pathway involved in determination of left/right asymmetry(GO:1900145) |
| 0.0 | 0.1 | GO:0048618 | post-embryonic foregut morphogenesis(GO:0048618) |
| 0.0 | 0.3 | GO:0006735 | glucose catabolic process(GO:0006007) NADH regeneration(GO:0006735) glycolytic process through fructose-6-phosphate(GO:0061615) glycolytic process through glucose-6-phosphate(GO:0061620) canonical glycolysis(GO:0061621) glucose catabolic process to pyruvate(GO:0061718) |
| 0.0 | 0.2 | GO:0045955 | negative regulation of calcium ion-dependent exocytosis(GO:0045955) negative regulation of regulated secretory pathway(GO:1903306) |
| 0.0 | 0.2 | GO:0006108 | malate metabolic process(GO:0006108) |
| 0.0 | 0.2 | GO:0031998 | regulation of fatty acid beta-oxidation(GO:0031998) |
| 0.0 | 0.1 | GO:0035176 | mammillary body development(GO:0021767) social behavior(GO:0035176) intraspecies interaction between organisms(GO:0051703) |
| 0.0 | 0.1 | GO:1903673 | mitotic cytokinetic process(GO:1902410) mitotic cleavage furrow formation(GO:1903673) |
| 0.0 | 0.1 | GO:1904059 | regulation of locomotor rhythm(GO:1904059) |
| 0.0 | 0.1 | GO:0006145 | purine nucleobase catabolic process(GO:0006145) |
| 0.0 | 0.1 | GO:0043567 | regulation of insulin-like growth factor receptor signaling pathway(GO:0043567) |
| 0.0 | 0.1 | GO:0071376 | response to corticotropin-releasing hormone(GO:0043435) cellular response to corticotropin-releasing hormone stimulus(GO:0071376) |
| 0.0 | 0.0 | GO:0090069 | regulation of ribosome biogenesis(GO:0090069) |
| 0.0 | 0.2 | GO:0030150 | protein import into mitochondrial matrix(GO:0030150) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | GO:0008247 | 1-alkyl-2-acetylglycerophosphocholine esterase complex(GO:0008247) |
| 0.0 | 0.3 | GO:0005945 | 6-phosphofructokinase complex(GO:0005945) |
| 0.0 | 0.6 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.0 | 0.3 | GO:0000276 | mitochondrial proton-transporting ATP synthase complex, coupling factor F(o)(GO:0000276) |
| 0.0 | 0.7 | GO:0005747 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.0 | 0.1 | GO:0033181 | plasma membrane proton-transporting V-type ATPase complex(GO:0033181) |
| 0.0 | 0.5 | GO:0080008 | Cul4-RING E3 ubiquitin ligase complex(GO:0080008) |
| 0.0 | 0.2 | GO:0005744 | mitochondrial inner membrane presequence translocase complex(GO:0005744) |
| 0.0 | 0.2 | GO:0097038 | perinuclear endoplasmic reticulum(GO:0097038) |
| 0.0 | 0.1 | GO:0061617 | MICOS complex(GO:0061617) |
| 0.0 | 0.2 | GO:0031680 | G-protein beta/gamma-subunit complex(GO:0031680) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.9 | GO:0004459 | L-lactate dehydrogenase activity(GO:0004459) |
| 0.1 | 0.5 | GO:0031852 | mu-type opioid receptor binding(GO:0031852) corticotropin-releasing hormone receptor 1 binding(GO:0051430) |
| 0.1 | 0.3 | GO:0008929 | triose-phosphate isomerase activity(GO:0004807) methylglyoxal synthase activity(GO:0008929) |
| 0.1 | 0.3 | GO:0030298 | receptor signaling protein tyrosine kinase activator activity(GO:0030298) |
| 0.0 | 0.2 | GO:0008260 | 3-oxoacid CoA-transferase activity(GO:0008260) |
| 0.0 | 1.1 | GO:0004129 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors(GO:0016675) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.0 | 0.3 | GO:0016416 | carnitine O-palmitoyltransferase activity(GO:0004095) O-palmitoyltransferase activity(GO:0016416) |
| 0.0 | 0.1 | GO:0042806 | fucose binding(GO:0042806) |
| 0.0 | 0.6 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.0 | 0.2 | GO:0015450 | P-P-bond-hydrolysis-driven protein transmembrane transporter activity(GO:0015450) |
| 0.0 | 0.1 | GO:0070004 | cysteine-type exopeptidase activity(GO:0070004) |
| 0.0 | 0.3 | GO:0003872 | 6-phosphofructokinase activity(GO:0003872) |
| 0.0 | 0.1 | GO:0047066 | phospholipid-hydroperoxide glutathione peroxidase activity(GO:0047066) |
| 0.0 | 0.1 | GO:0001609 | G-protein coupled adenosine receptor activity(GO:0001609) |
| 0.0 | 0.1 | GO:0015137 | citrate transmembrane transporter activity(GO:0015137) tricarboxylic acid transmembrane transporter activity(GO:0015142) |
| 0.0 | 0.4 | GO:0048038 | quinone binding(GO:0048038) |
| 0.0 | 0.1 | GO:0050431 | transforming growth factor beta binding(GO:0050431) |
| 0.0 | 0.2 | GO:0070840 | dynein complex binding(GO:0070840) |
| 0.0 | 0.1 | GO:0005113 | patched binding(GO:0005113) |
| 0.0 | 0.1 | GO:0031995 | insulin-like growth factor I binding(GO:0031994) insulin-like growth factor II binding(GO:0031995) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | PID RHODOPSIN PATHWAY | Visual signal transduction: Rods |
| 0.0 | 0.1 | SA G2 AND M PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | REACTOME OLFACTORY SIGNALING PATHWAY | Genes involved in Olfactory Signaling Pathway |
| 0.0 | 0.4 | REACTOME PROSTACYCLIN SIGNALLING THROUGH PROSTACYCLIN RECEPTOR | Genes involved in Prostacyclin signalling through prostacyclin receptor |
| 0.0 | 0.2 | REACTOME CREATION OF C4 AND C2 ACTIVATORS | Genes involved in Creation of C4 and C2 activators |
| 0.0 | 0.4 | REACTOME METAL ION SLC TRANSPORTERS | Genes involved in Metal ion SLC transporters |
| 0.0 | 0.7 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.0 | 0.2 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.0 | 0.1 | REACTOME PURINE CATABOLISM | Genes involved in Purine catabolism |