Project
PRJNA438478: RNAseq of wild type zebrafish germline
Navigation
Downloads
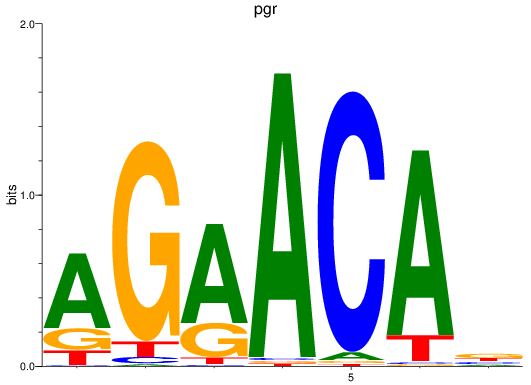
Results for pgr
Z-value: 0.52
Transcription factors associated with pgr
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
pgr
|
ENSDARG00000035966 | progesterone receptor |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| pgr | dr11_v1_chr18_+_41820039_41820039 | 0.72 | 8.4e-04 | Click! |
Activity profile of pgr motif
Sorted Z-values of pgr motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_+_57311901 | 1.50 |
ENSDART00000149397
|
mycbpap
|
mycbp associated protein |
| chr16_-_41807397 | 1.42 |
ENSDART00000138577
|
si:dkey-199f5.7
|
si:dkey-199f5.7 |
| chr17_-_24838927 | 1.35 |
ENSDART00000123259
|
CR388164.1
|
|
| chr15_-_19677511 | 1.23 |
ENSDART00000043743
|
si:dkey-4p15.3
|
si:dkey-4p15.3 |
| chr6_+_515181 | 0.93 |
ENSDART00000171374
|
CACNA1I (1 of many)
|
si:ch73-379f7.5 |
| chr7_+_23997518 | 0.92 |
ENSDART00000173372
|
ccdc166
|
coiled-coil domain containing 166 |
| chr16_-_41840668 | 0.91 |
ENSDART00000146150
|
si:dkey-199f5.4
|
si:dkey-199f5.4 |
| chr2_-_14272083 | 0.88 |
ENSDART00000169986
|
tctex1d1
|
Tctex1 domain containing 1 |
| chr19_+_43978814 | 0.88 |
ENSDART00000102314
|
nkd3
|
naked cuticle homolog 3 |
| chr16_-_41059329 | 0.82 |
ENSDART00000145556
|
si:dkey-201i6.8
|
si:dkey-201i6.8 |
| chr8_+_25902170 | 0.80 |
ENSDART00000193130
|
rhoab
|
ras homolog gene family, member Ab |
| chr6_+_475264 | 0.79 |
ENSDART00000193615
|
LO017974.1
|
|
| chr24_+_33589667 | 0.76 |
ENSDART00000152097
|
dnah5l
|
dynein, axonemal, heavy chain 5 like |
| chr18_+_1145571 | 0.76 |
ENSDART00000135055
|
rec114
|
REC114 meiotic recombination protein |
| chr14_-_36862745 | 0.75 |
ENSDART00000109293
|
rnf130
|
ring finger protein 130 |
| chr10_+_26669177 | 0.72 |
ENSDART00000143402
|
si:ch73-52f15.5
|
si:ch73-52f15.5 |
| chr17_-_12407106 | 0.71 |
ENSDART00000183540
|
ankef1b
|
ankyrin repeat and EF-hand domain containing 1b |
| chr13_-_37619159 | 0.66 |
ENSDART00000186348
|
zgc:152791
|
zgc:152791 |
| chr7_+_49681040 | 0.65 |
ENSDART00000176372
ENSDART00000192172 |
rassf7b
|
Ras association (RalGDS/AF-6) domain family (N-terminal) member 7b |
| chr22_-_24285432 | 0.64 |
ENSDART00000164083
|
si:ch211-117l17.4
|
si:ch211-117l17.4 |
| chr4_-_16406737 | 0.64 |
ENSDART00000013085
|
dcn
|
decorin |
| chr2_+_26288301 | 0.64 |
ENSDART00000017668
|
ptbp1a
|
polypyrimidine tract binding protein 1a |
| chr20_-_43462190 | 0.61 |
ENSDART00000100697
|
pimr133
|
Pim proto-oncogene, serine/threonine kinase, related 133 |
| chr2_-_29485408 | 0.60 |
ENSDART00000013411
|
cahz
|
carbonic anhydrase |
| chr18_+_5547185 | 0.59 |
ENSDART00000193977
|
nnt2
|
nicotinamide nucleotide transhydrogenase 2 |
| chr6_-_49078263 | 0.58 |
ENSDART00000032982
|
slc5a8l
|
solute carrier family 5 (iodide transporter), member 8-like |
| chr11_+_35364445 | 0.57 |
ENSDART00000125766
|
camkvb
|
CaM kinase-like vesicle-associated b |
| chr12_-_5188413 | 0.57 |
ENSDART00000161988
|
fra10ac1
|
FRA10A associated CGG repeat 1 |
| chr8_+_53311965 | 0.52 |
ENSDART00000130104
|
gnb1a
|
guanine nucleotide binding protein (G protein), beta polypeptide 1a |
| chr20_+_38837238 | 0.50 |
ENSDART00000061334
|
ift172
|
intraflagellar transport 172 |
| chr13_-_36535128 | 0.49 |
ENSDART00000043312
|
srsf5a
|
serine/arginine-rich splicing factor 5a |
| chr15_-_17870090 | 0.49 |
ENSDART00000155066
|
atf5b
|
activating transcription factor 5b |
| chr9_-_42696408 | 0.49 |
ENSDART00000144744
|
col5a2a
|
collagen, type V, alpha 2a |
| chr6_+_7250824 | 0.47 |
ENSDART00000177226
|
dzip1
|
DAZ interacting zinc finger protein 1 |
| chr20_-_26420939 | 0.47 |
ENSDART00000110883
|
akap12b
|
A kinase (PRKA) anchor protein 12b |
| chr5_+_69981453 | 0.47 |
ENSDART00000143860
|
si:ch211-154e10.1
|
si:ch211-154e10.1 |
| chr16_-_35579200 | 0.46 |
ENSDART00000162172
|
scmh1
|
Scm polycomb group protein homolog 1 |
| chr25_-_12982193 | 0.45 |
ENSDART00000159617
|
ccl39.5
|
chemokine (C-C motif) ligand 39, duplicate 5 |
| chr14_-_25949713 | 0.44 |
ENSDART00000181455
|
sparc
|
secreted protein, acidic, cysteine-rich (osteonectin) |
| chr18_+_16744307 | 0.43 |
ENSDART00000179872
ENSDART00000133490 |
lyve1b
|
lymphatic vessel endothelial hyaluronic receptor 1b |
| chr20_-_26421112 | 0.42 |
ENSDART00000183767
ENSDART00000182330 |
akap12b
|
A kinase (PRKA) anchor protein 12b |
| chr17_+_26815021 | 0.42 |
ENSDART00000086885
|
asmt2
|
acetylserotonin O-methyltransferase 2 |
| chr25_-_6420800 | 0.42 |
ENSDART00000153768
|
ptpn9a
|
protein tyrosine phosphatase, non-receptor type 9, a |
| chr19_+_48251273 | 0.42 |
ENSDART00000157424
|
CU693379.1
|
|
| chr9_-_22831836 | 0.42 |
ENSDART00000142585
|
neb
|
nebulin |
| chr1_-_10914523 | 0.41 |
ENSDART00000007013
|
dmd
|
dystrophin |
| chr2_-_50053006 | 0.40 |
ENSDART00000083654
|
si:ch211-106n13.3
|
si:ch211-106n13.3 |
| chr5_+_20693724 | 0.40 |
ENSDART00000141368
|
si:ch211-240b21.2
|
si:ch211-240b21.2 |
| chr5_-_51756210 | 0.40 |
ENSDART00000163464
|
lhfpl2b
|
LHFPL tetraspan subfamily member 2b |
| chr18_+_9171778 | 0.40 |
ENSDART00000101192
|
sema3d
|
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3D |
| chr9_-_46842179 | 0.39 |
ENSDART00000054137
|
igfbp5b
|
insulin-like growth factor binding protein 5b |
| chr8_-_12468744 | 0.38 |
ENSDART00000135019
|
FIBCD1 (1 of many)
|
si:dkeyp-51b7.3 |
| chr10_+_8968203 | 0.38 |
ENSDART00000110443
ENSDART00000080772 |
fstb
|
follistatin b |
| chr20_+_16750177 | 0.38 |
ENSDART00000185357
|
calm1b
|
calmodulin 1b |
| chr23_-_26077038 | 0.37 |
ENSDART00000126299
|
gdi1
|
GDP dissociation inhibitor 1 |
| chr5_-_25174420 | 0.37 |
ENSDART00000141554
|
abca2
|
ATP-binding cassette, sub-family A (ABC1), member 2 |
| chr21_-_14803554 | 0.37 |
ENSDART00000121853
|
si:dkey-11o18.5
|
si:dkey-11o18.5 |
| chr1_-_51615437 | 0.36 |
ENSDART00000152185
ENSDART00000152237 ENSDART00000129052 ENSDART00000152595 |
FBXO48
|
si:dkey-202b17.4 |
| chr5_-_28029558 | 0.35 |
ENSDART00000078649
|
abcb9
|
ATP-binding cassette, sub-family B (MDR/TAP), member 9 |
| chr12_-_1951233 | 0.34 |
ENSDART00000005676
ENSDART00000127937 |
sox9a
|
SRY (sex determining region Y)-box 9a |
| chr4_+_12358822 | 0.34 |
ENSDART00000172557
ENSDART00000150627 |
pimr168
|
Pim proto-oncogene, serine/threonine kinase, related 168 |
| chr5_-_67911111 | 0.34 |
ENSDART00000051833
|
gsx1
|
GS homeobox 1 |
| chr20_+_35445462 | 0.33 |
ENSDART00000124497
|
tdrd6
|
tudor domain containing 6 |
| chr8_-_36475328 | 0.33 |
ENSDART00000048448
|
si:busm1-266f07.2
|
si:busm1-266f07.2 |
| chr14_-_2602445 | 0.33 |
ENSDART00000166910
|
etf1a
|
eukaryotic translation termination factor 1a |
| chr21_+_18353703 | 0.32 |
ENSDART00000181396
ENSDART00000166359 |
si:ch73-287m6.1
|
si:ch73-287m6.1 |
| chr24_+_23742690 | 0.32 |
ENSDART00000130162
|
TCF24
|
transcription factor 24 |
| chr20_+_9785952 | 0.32 |
ENSDART00000152403
|
ddhd1b
|
DDHD domain containing 1b |
| chr20_+_539852 | 0.31 |
ENSDART00000185994
|
dse
|
dermatan sulfate epimerase |
| chr18_+_21061216 | 0.31 |
ENSDART00000141739
|
fam169b
|
family with sequence similarity 169, member B |
| chr2_+_47581997 | 0.31 |
ENSDART00000112579
|
scg2b
|
secretogranin II (chromogranin C), b |
| chr16_+_16968682 | 0.31 |
ENSDART00000111074
|
si:ch211-120k19.1
|
si:ch211-120k19.1 |
| chr21_+_25187210 | 0.30 |
ENSDART00000101147
ENSDART00000167528 |
si:dkey-183i3.5
|
si:dkey-183i3.5 |
| chr17_+_26783828 | 0.30 |
ENSDART00000154921
|
larp1b
|
La ribonucleoprotein domain family, member 1B |
| chr8_+_20951590 | 0.30 |
ENSDART00000138728
|
si:dkeyp-82a1.1
|
si:dkeyp-82a1.1 |
| chr6_-_39764995 | 0.29 |
ENSDART00000085277
|
pfkmb
|
phosphofructokinase, muscle b |
| chr19_-_27542433 | 0.29 |
ENSDART00000136414
|
si:ch211-152p11.4
|
si:ch211-152p11.4 |
| chr2_+_44571200 | 0.29 |
ENSDART00000098132
|
klhl24a
|
kelch-like family member 24a |
| chr15_-_21678634 | 0.29 |
ENSDART00000061139
|
bco2b
|
beta-carotene oxygenase 2b |
| chr9_+_19489514 | 0.29 |
ENSDART00000152032
ENSDART00000114256 ENSDART00000190572 ENSDART00000147571 ENSDART00000151918 ENSDART00000152034 |
si:ch211-140m22.7
|
si:ch211-140m22.7 |
| chr2_-_37896965 | 0.29 |
ENSDART00000129852
|
hbl1
|
hexose-binding lectin 1 |
| chr16_+_16969060 | 0.29 |
ENSDART00000182819
ENSDART00000191876 |
si:ch211-120k19.1
rpl18
|
si:ch211-120k19.1 ribosomal protein L18 |
| chr10_+_44146287 | 0.28 |
ENSDART00000162176
|
grk3
|
G protein-coupled receptor kinase 3 |
| chr20_+_29634653 | 0.28 |
ENSDART00000101556
|
asap2b
|
ArfGAP with SH3 domain, ankyrin repeat and PH domain 2b |
| chr20_-_34711528 | 0.28 |
ENSDART00000061555
|
si:ch211-63o20.7
|
si:ch211-63o20.7 |
| chr1_-_35924495 | 0.28 |
ENSDART00000184424
|
smad1
|
SMAD family member 1 |
| chr7_+_52761841 | 0.28 |
ENSDART00000111444
|
ppip5k1a
|
diphosphoinositol pentakisphosphate kinase 1a |
| chr22_+_20427170 | 0.28 |
ENSDART00000136744
|
foxq2
|
forkhead box Q2 |
| chr20_+_18580176 | 0.28 |
ENSDART00000185310
|
si:dkeyp-72h1.1
|
si:dkeyp-72h1.1 |
| chr1_-_45614481 | 0.27 |
ENSDART00000148413
|
atf7ip
|
activating transcription factor 7 interacting protein |
| chr12_-_19007834 | 0.27 |
ENSDART00000153248
|
chadlb
|
chondroadherin-like b |
| chr2_+_10134345 | 0.27 |
ENSDART00000100725
|
ahsg2
|
alpha-2-HS-glycoprotein 2 |
| chr6_-_8489810 | 0.27 |
ENSDART00000124643
|
rasal3
|
RAS protein activator like 3 |
| chr25_+_4541211 | 0.27 |
ENSDART00000129978
|
pnpla2
|
patatin-like phospholipase domain containing 2 |
| chr16_-_51271962 | 0.27 |
ENSDART00000164021
ENSDART00000046420 |
serpinb1l1
|
serpin peptidase inhibitor, clade B (ovalbumin), member 1, like 1 |
| chr15_-_32383529 | 0.27 |
ENSDART00000028349
|
c4
|
complement component 4 |
| chr23_+_25210953 | 0.26 |
ENSDART00000132799
|
si:dkey-151g10.3
|
si:dkey-151g10.3 |
| chr9_+_12948511 | 0.26 |
ENSDART00000135797
|
si:dkey-230p4.1
|
si:dkey-230p4.1 |
| chr2_+_29976419 | 0.26 |
ENSDART00000056748
|
en2b
|
engrailed homeobox 2b |
| chr7_-_27033080 | 0.26 |
ENSDART00000173516
|
nucb2a
|
nucleobindin 2a |
| chr24_-_22702017 | 0.26 |
ENSDART00000179403
|
ctnnd2a
|
catenin (cadherin-associated protein), delta 2a |
| chr1_+_144284 | 0.26 |
ENSDART00000064061
|
prozb
|
protein Z, vitamin K-dependent plasma glycoprotein b |
| chr18_+_19120984 | 0.26 |
ENSDART00000141501
|
si:dkey-242h9.3
|
si:dkey-242h9.3 |
| chr15_+_1796313 | 0.25 |
ENSDART00000126253
|
fam124b
|
family with sequence similarity 124B |
| chr21_-_25601648 | 0.25 |
ENSDART00000042578
|
efemp2b
|
EGF containing fibulin extracellular matrix protein 2b |
| chr15_+_44472356 | 0.25 |
ENSDART00000155763
ENSDART00000028011 |
msantd4
|
Myb/SANT-like DNA-binding domain containing 4 with coiled-coils |
| chr13_-_36391496 | 0.25 |
ENSDART00000100217
ENSDART00000140243 |
actn1
|
actinin, alpha 1 |
| chr17_-_28097760 | 0.24 |
ENSDART00000149861
|
kdm1a
|
lysine (K)-specific demethylase 1a |
| chr17_-_28100741 | 0.24 |
ENSDART00000180532
|
kdm1a
|
lysine (K)-specific demethylase 1a |
| chr3_-_2589704 | 0.24 |
ENSDART00000187239
|
si:dkey-217f16.5
|
si:dkey-217f16.5 |
| chr24_-_33756003 | 0.24 |
ENSDART00000079283
|
tmeff1b
|
transmembrane protein with EGF-like and two follistatin-like domains 1b |
| chr20_+_6630540 | 0.24 |
ENSDART00000138361
|
tns3.2
|
tensin 3, tandem duplicate 2 |
| chr5_+_30680804 | 0.24 |
ENSDART00000078049
|
pdzd3b
|
PDZ domain containing 3b |
| chr13_+_35745572 | 0.24 |
ENSDART00000159690
|
gpr75
|
G protein-coupled receptor 75 |
| chr1_-_23274038 | 0.24 |
ENSDART00000181658
|
rfc1
|
replication factor C (activator 1) 1 |
| chr15_+_38046435 | 0.24 |
ENSDART00000156481
|
si:ch73-380l3.4
|
si:ch73-380l3.4 |
| chr4_-_11577253 | 0.23 |
ENSDART00000144452
|
net1
|
neuroepithelial cell transforming 1 |
| chr6_+_11250033 | 0.23 |
ENSDART00000065411
ENSDART00000132677 |
atg9a
|
ATG9 autophagy related 9 homolog A (S. cerevisiae) |
| chr9_+_44832032 | 0.23 |
ENSDART00000002633
|
frzb
|
frizzled related protein |
| chr7_+_46368520 | 0.23 |
ENSDART00000192821
|
znf536
|
zinc finger protein 536 |
| chr4_-_52165969 | 0.23 |
ENSDART00000171130
|
si:dkeyp-44b5.4
|
si:dkeyp-44b5.4 |
| chr7_-_74090168 | 0.23 |
ENSDART00000050528
|
tyrp1a
|
tyrosinase-related protein 1a |
| chr25_+_3564220 | 0.23 |
ENSDART00000170005
|
zgc:194202
|
zgc:194202 |
| chr1_-_26398361 | 0.22 |
ENSDART00000160183
|
ppa2
|
pyrophosphatase (inorganic) 2 |
| chr8_+_68864 | 0.22 |
ENSDART00000164574
|
prr16
|
proline rich 16 |
| chr19_-_10656667 | 0.22 |
ENSDART00000081379
ENSDART00000151456 ENSDART00000143271 ENSDART00000182126 |
olah
|
oleoyl-ACP hydrolase |
| chr6_+_11249706 | 0.22 |
ENSDART00000186547
ENSDART00000193287 |
atg9a
|
ATG9 autophagy related 9 homolog A (S. cerevisiae) |
| chr6_-_54290227 | 0.22 |
ENSDART00000050483
|
SPDEF
|
SAM pointed domain containing ETS transcription factor |
| chr17_-_7501916 | 0.22 |
ENSDART00000141987
|
si:ch211-278p9.3
|
si:ch211-278p9.3 |
| chr19_-_29437108 | 0.22 |
ENSDART00000052108
ENSDART00000074497 |
fndc5b
|
fibronectin type III domain containing 5b |
| chr5_+_22459087 | 0.21 |
ENSDART00000134781
|
BX546499.1
|
|
| chr23_+_30048849 | 0.21 |
ENSDART00000126027
|
uts2a
|
urotensin 2, alpha |
| chr20_+_33904258 | 0.21 |
ENSDART00000170930
|
rxrgb
|
retinoid X receptor, gamma b |
| chr3_+_42497814 | 0.21 |
ENSDART00000184653
|
BX537305.1
|
|
| chr3_-_29977495 | 0.21 |
ENSDART00000077111
|
hsd17b14
|
hydroxysteroid (17-beta) dehydrogenase 14 |
| chr6_+_41096058 | 0.20 |
ENSDART00000028373
|
fkbp5
|
FK506 binding protein 5 |
| chr13_+_28495419 | 0.20 |
ENSDART00000025583
|
fgf8a
|
fibroblast growth factor 8a |
| chr3_-_42125655 | 0.20 |
ENSDART00000040753
|
nudt1
|
nudix (nucleoside diphosphate linked moiety X)-type motif 1 |
| chr4_-_20181964 | 0.20 |
ENSDART00000022539
|
fgl2a
|
fibrinogen-like 2a |
| chr22_-_15602593 | 0.20 |
ENSDART00000036075
|
tpm4a
|
tropomyosin 4a |
| chr2_-_32366287 | 0.20 |
ENSDART00000144758
|
ubtfl
|
upstream binding transcription factor, like |
| chr9_-_8454060 | 0.20 |
ENSDART00000110158
|
irs2b
|
insulin receptor substrate 2b |
| chr7_-_51710015 | 0.20 |
ENSDART00000009184
|
hdac8
|
histone deacetylase 8 |
| chr10_+_34394454 | 0.19 |
ENSDART00000110121
|
stard13a
|
StAR-related lipid transfer (START) domain containing 13a |
| chr10_+_6121558 | 0.19 |
ENSDART00000166799
ENSDART00000157947 |
tln1
|
talin 1 |
| chr20_-_39735952 | 0.19 |
ENSDART00000101049
ENSDART00000137485 ENSDART00000062402 |
tpd52l1
|
tumor protein D52-like 1 |
| chr3_+_32698424 | 0.19 |
ENSDART00000055340
|
fus
|
FUS RNA binding protein |
| chr23_+_7379728 | 0.19 |
ENSDART00000012194
|
gata5
|
GATA binding protein 5 |
| chr14_-_7141550 | 0.19 |
ENSDART00000172020
|
trpt1
|
tRNA phosphotransferase 1 |
| chr9_+_31752628 | 0.19 |
ENSDART00000060054
|
itgbl1
|
integrin, beta-like 1 |
| chr9_-_14992730 | 0.19 |
ENSDART00000137117
|
pard3bb
|
par-3 family cell polarity regulator beta b |
| chr21_+_10866421 | 0.19 |
ENSDART00000137858
|
alpk2
|
alpha-kinase 2 |
| chr13_-_32577386 | 0.19 |
ENSDART00000016535
|
kcns3a
|
potassium voltage-gated channel, delayed-rectifier, subfamily S, member 3a |
| chr5_-_52215926 | 0.19 |
ENSDART00000163973
ENSDART00000193602 |
lnpep
|
leucyl/cystinyl aminopeptidase |
| chr2_+_45159855 | 0.18 |
ENSDART00000056333
|
CU929150.1
|
|
| chr6_-_11136786 | 0.18 |
ENSDART00000186522
|
myo10l1
|
myosin X-like 1 |
| chr8_-_11229523 | 0.18 |
ENSDART00000002164
|
unc45b
|
unc-45 myosin chaperone B |
| chr23_+_21663631 | 0.18 |
ENSDART00000066125
|
dhrs3a
|
dehydrogenase/reductase (SDR family) member 3a |
| chr6_-_18400548 | 0.18 |
ENSDART00000179797
ENSDART00000164891 |
trim25
|
tripartite motif containing 25 |
| chr21_+_25221940 | 0.18 |
ENSDART00000108972
|
sycn.1
|
syncollin, tandem duplicate 1 |
| chr2_+_258698 | 0.18 |
ENSDART00000181330
ENSDART00000181645 |
phlpp1
|
PH domain and leucine rich repeat protein phosphatase 1 |
| chr6_-_39765546 | 0.18 |
ENSDART00000185767
|
pfkmb
|
phosphofructokinase, muscle b |
| chr18_+_30567945 | 0.18 |
ENSDART00000078894
|
irf8
|
interferon regulatory factor 8 |
| chr19_-_18626515 | 0.17 |
ENSDART00000160624
|
rps18
|
ribosomal protein S18 |
| chr5_+_61863996 | 0.17 |
ENSDART00000082879
|
sympk
|
symplekin |
| chr18_-_39200557 | 0.17 |
ENSDART00000132367
ENSDART00000183672 |
si:ch211-235f12.2
mapk6
|
si:ch211-235f12.2 mitogen-activated protein kinase 6 |
| chr10_+_25204626 | 0.17 |
ENSDART00000024514
|
chordc1b
|
cysteine and histidine-rich domain (CHORD) containing 1b |
| chr19_-_18626952 | 0.17 |
ENSDART00000168004
ENSDART00000162034 ENSDART00000165486 ENSDART00000167971 |
rps18
|
ribosomal protein S18 |
| chr13_+_39188737 | 0.16 |
ENSDART00000083641
|
fam135a
|
family with sequence similarity 135, member A |
| chr22_-_38508112 | 0.16 |
ENSDART00000192004
|
klc4
|
kinesin light chain 4 |
| chr12_-_33646010 | 0.16 |
ENSDART00000111259
|
tmem94
|
transmembrane protein 94 |
| chr23_-_45504991 | 0.16 |
ENSDART00000148761
|
col24a1
|
collagen type XXIV alpha 1 |
| chr23_+_2760573 | 0.16 |
ENSDART00000129719
|
top1
|
DNA topoisomerase I |
| chr6_-_40429411 | 0.16 |
ENSDART00000156005
ENSDART00000156357 |
si:dkey-28n18.9
|
si:dkey-28n18.9 |
| chr6_-_60031693 | 0.16 |
ENSDART00000160275
|
CABZ01079262.1
|
|
| chr5_-_21422390 | 0.16 |
ENSDART00000144198
|
tenm1
|
teneurin transmembrane protein 1 |
| chr11_-_4787083 | 0.15 |
ENSDART00000159425
|
FHIT
|
si:ch211-63i20.3 |
| chr25_-_210730 | 0.15 |
ENSDART00000187580
|
FP236318.1
|
|
| chr7_-_44672095 | 0.15 |
ENSDART00000073745
|
cmtm4
|
CKLF-like MARVEL transmembrane domain containing 4 |
| chr14_-_25985698 | 0.15 |
ENSDART00000172909
ENSDART00000123053 |
atox1
|
antioxidant 1 copper chaperone |
| chr17_+_25677458 | 0.15 |
ENSDART00000029703
|
kcnh1a
|
potassium voltage-gated channel, subfamily H (eag-related), member 1a |
| chr18_-_2639351 | 0.15 |
ENSDART00000168106
|
relt
|
RELT, TNF receptor |
| chr3_-_58582663 | 0.15 |
ENSDART00000180055
|
CABZ01038521.1
|
|
| chr20_-_20611063 | 0.14 |
ENSDART00000063492
|
ppm1ab
|
protein phosphatase, Mg2+/Mn2+ dependent, 1Ab |
| chr20_-_30377221 | 0.14 |
ENSDART00000126229
|
rps7
|
ribosomal protein S7 |
| chr15_-_32383340 | 0.14 |
ENSDART00000185632
|
c4
|
complement component 4 |
| chr21_-_26089964 | 0.14 |
ENSDART00000027848
|
tlcd1
|
TLC domain containing 1 |
| chr8_+_23093155 | 0.14 |
ENSDART00000063075
|
zgc:100920
|
zgc:100920 |
| chr5_+_62356304 | 0.14 |
ENSDART00000148381
|
aspa
|
aspartoacylase |
| chr4_+_77021784 | 0.14 |
ENSDART00000135345
ENSDART00000133855 |
trpm2
|
transient receptor potential cation channel, subfamily M, member 2 |
| chr1_-_59348118 | 0.14 |
ENSDART00000170901
|
cyp3a65
|
cytochrome P450, family 3, subfamily A, polypeptide 65 |
| chr17_-_28100501 | 0.14 |
ENSDART00000149543
|
kdm1a
|
lysine (K)-specific demethylase 1a |
| chr5_-_41307550 | 0.14 |
ENSDART00000143446
|
npr3
|
natriuretic peptide receptor 3 |
| chr22_-_4778018 | 0.14 |
ENSDART00000143968
|
si:ch73-256j6.5
|
si:ch73-256j6.5 |
| chr8_+_44613135 | 0.14 |
ENSDART00000063392
|
lsm1
|
LSM1, U6 small nuclear RNA associated |
| chr20_-_26846028 | 0.14 |
ENSDART00000136687
|
mylk4b
|
myosin light chain kinase family, member 4b |
| chr20_-_47704973 | 0.13 |
ENSDART00000174808
|
tfap2b
|
transcription factor AP-2 beta |
| chr22_+_1384104 | 0.13 |
ENSDART00000187191
|
BX322657.3
|
|
| chr4_-_28958601 | 0.13 |
ENSDART00000111294
|
zgc:174315
|
zgc:174315 |
| chr12_+_2995887 | 0.13 |
ENSDART00000189499
ENSDART00000182597 ENSDART00000184918 ENSDART00000182292 |
lrrc45
|
leucine rich repeat containing 45 |
Network of associatons between targets according to the STRING database.
First level regulatory network of pgr
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.9 | GO:1904969 | bleb assembly(GO:0032060) slow muscle cell migration(GO:1904969) |
| 0.2 | 0.7 | GO:0060467 | negative regulation of fertilization(GO:0060467) prevention of polyspermy(GO:0060468) cortical granule exocytosis(GO:0060471) |
| 0.2 | 0.6 | GO:0015670 | carbon dioxide transport(GO:0015670) |
| 0.1 | 0.6 | GO:0006740 | NADPH regeneration(GO:0006740) |
| 0.1 | 0.4 | GO:0030186 | melatonin metabolic process(GO:0030186) |
| 0.1 | 0.8 | GO:1902766 | skeletal muscle satellite cell migration(GO:1902766) |
| 0.1 | 0.6 | GO:0034720 | histone H3-K4 demethylation(GO:0034720) |
| 0.1 | 0.4 | GO:0007508 | larval development(GO:0002164) larval heart development(GO:0007508) |
| 0.1 | 0.4 | GO:0071691 | cardiac muscle thin filament assembly(GO:0071691) |
| 0.1 | 0.3 | GO:0010898 | regulation of triglyceride catabolic process(GO:0010896) positive regulation of triglyceride catabolic process(GO:0010898) |
| 0.1 | 0.3 | GO:0002184 | cytoplasmic translational termination(GO:0002184) |
| 0.1 | 0.3 | GO:0030205 | dermatan sulfate metabolic process(GO:0030205) |
| 0.1 | 0.2 | GO:0039529 | RIG-I signaling pathway(GO:0039529) regulation of viral-induced cytoplasmic pattern recognition receptor signaling pathway(GO:0039531) regulation of RIG-I signaling pathway(GO:0039535) |
| 0.1 | 0.2 | GO:0045649 | regulation of macrophage differentiation(GO:0045649) |
| 0.1 | 0.8 | GO:0036158 | outer dynein arm assembly(GO:0036158) |
| 0.1 | 0.3 | GO:0001773 | myeloid dendritic cell activation(GO:0001773) myeloid dendritic cell differentiation(GO:0043011) |
| 0.1 | 0.5 | GO:1902624 | positive regulation of neutrophil migration(GO:1902624) |
| 0.0 | 0.2 | GO:0051660 | establishment of centrosome localization(GO:0051660) |
| 0.0 | 0.2 | GO:0035988 | chondrocyte proliferation(GO:0035988) |
| 0.0 | 0.4 | GO:0044805 | late nucleophagy(GO:0044805) |
| 0.0 | 0.3 | GO:0038065 | collagen-activated tyrosine kinase receptor signaling pathway(GO:0038063) collagen-activated signaling pathway(GO:0038065) |
| 0.0 | 0.5 | GO:0006007 | glucose catabolic process(GO:0006007) NADH regeneration(GO:0006735) glycolytic process through fructose-6-phosphate(GO:0061615) glycolytic process through glucose-6-phosphate(GO:0061620) canonical glycolysis(GO:0061621) glucose catabolic process to pyruvate(GO:0061718) |
| 0.0 | 0.1 | GO:0045218 | zonula adherens maintenance(GO:0045218) |
| 0.0 | 0.3 | GO:0043587 | tongue development(GO:0043586) tongue morphogenesis(GO:0043587) taste bud development(GO:0061193) taste bud morphogenesis(GO:0061194) taste bud formation(GO:0061195) |
| 0.0 | 0.2 | GO:0030890 | B cell homeostasis(GO:0001782) positive regulation of B cell proliferation(GO:0030890) |
| 0.0 | 0.3 | GO:2000312 | regulation of kainate selective glutamate receptor activity(GO:2000312) |
| 0.0 | 0.3 | GO:0070933 | histone H4 deacetylation(GO:0070933) |
| 0.0 | 0.5 | GO:0006607 | NLS-bearing protein import into nucleus(GO:0006607) |
| 0.0 | 0.1 | GO:0033210 | leptin-mediated signaling pathway(GO:0033210) cellular response to leptin stimulus(GO:0044320) response to leptin(GO:0044321) |
| 0.0 | 0.4 | GO:0043567 | regulation of insulin-like growth factor receptor signaling pathway(GO:0043567) |
| 0.0 | 0.1 | GO:0055091 | phospholipid homeostasis(GO:0055091) |
| 0.0 | 0.4 | GO:0007525 | somatic muscle development(GO:0007525) |
| 0.0 | 0.4 | GO:0032926 | negative regulation of activin receptor signaling pathway(GO:0032926) |
| 0.0 | 0.1 | GO:0003261 | cardiac muscle progenitor cell migration to the midline involved in heart field formation(GO:0003261) |
| 0.0 | 0.3 | GO:0046247 | carotene metabolic process(GO:0016119) carotene catabolic process(GO:0016121) terpene metabolic process(GO:0042214) terpene catabolic process(GO:0046247) |
| 0.0 | 0.3 | GO:0019885 | antigen processing and presentation of endogenous peptide antigen(GO:0002483) antigen processing and presentation of endogenous antigen(GO:0019883) antigen processing and presentation of endogenous peptide antigen via MHC class I(GO:0019885) |
| 0.0 | 0.1 | GO:0097623 | potassium ion export(GO:0071435) potassium ion export across plasma membrane(GO:0097623) |
| 0.0 | 0.3 | GO:0030719 | P granule organization(GO:0030719) |
| 0.0 | 0.4 | GO:0090308 | methylation-dependent chromatin silencing(GO:0006346) positive regulation of chromatin silencing(GO:0031937) regulation of methylation-dependent chromatin silencing(GO:0090308) positive regulation of methylation-dependent chromatin silencing(GO:0090309) |
| 0.0 | 0.1 | GO:1903385 | dendrite guidance(GO:0070983) regulation of homophilic cell adhesion(GO:1903385) |
| 0.0 | 0.3 | GO:0032096 | negative regulation of response to food(GO:0032096) negative regulation of appetite(GO:0032099) |
| 0.0 | 0.3 | GO:0003315 | heart rudiment formation(GO:0003315) |
| 0.0 | 0.1 | GO:1901492 | positive regulation of lymphangiogenesis(GO:1901492) |
| 0.0 | 0.2 | GO:0042438 | melanin biosynthetic process(GO:0042438) |
| 0.0 | 0.1 | GO:0030237 | female sex determination(GO:0030237) male sex determination(GO:0030238) |
| 0.0 | 0.1 | GO:0015803 | branched-chain amino acid transport(GO:0015803) leucine transport(GO:0015820) |
| 0.0 | 0.1 | GO:0003272 | endocardial cushion formation(GO:0003272) |
| 0.0 | 0.6 | GO:0010508 | positive regulation of autophagy(GO:0010508) |
| 0.0 | 0.3 | GO:0032332 | positive regulation of chondrocyte differentiation(GO:0032332) |
| 0.0 | 0.3 | GO:0006020 | inositol metabolic process(GO:0006020) |
| 0.0 | 0.1 | GO:2000042 | negative regulation of double-strand break repair via homologous recombination(GO:2000042) |
| 0.0 | 0.1 | GO:0008584 | male gonad development(GO:0008584) |
| 0.0 | 0.2 | GO:0032925 | regulation of activin receptor signaling pathway(GO:0032925) |
| 0.0 | 0.3 | GO:0042572 | retinol metabolic process(GO:0042572) |
| 0.0 | 0.1 | GO:0000290 | deadenylation-dependent decapping of nuclear-transcribed mRNA(GO:0000290) |
| 0.0 | 0.2 | GO:0006388 | tRNA splicing, via endonucleolytic cleavage and ligation(GO:0006388) |
| 0.0 | 0.1 | GO:0006878 | cellular copper ion homeostasis(GO:0006878) |
| 0.0 | 0.5 | GO:0045494 | photoreceptor cell maintenance(GO:0045494) |
| 0.0 | 0.5 | GO:1903670 | regulation of sprouting angiogenesis(GO:1903670) |
| 0.0 | 0.1 | GO:0060333 | interferon-gamma-mediated signaling pathway(GO:0060333) |
| 0.0 | 0.1 | GO:0043091 | L-arginine import(GO:0043091) amino acid import across plasma membrane(GO:0089718) arginine import(GO:0090467) L-arginine import across plasma membrane(GO:0097638) L-arginine transport(GO:1902023) L-arginine import into cell(GO:1902765) amino acid import into cell(GO:1902837) L-arginine transmembrane transport(GO:1903400) arginine transmembrane transport(GO:1903826) |
| 0.0 | 0.4 | GO:0048752 | otolith morphogenesis(GO:0032474) semicircular canal morphogenesis(GO:0048752) |
| 0.0 | 0.2 | GO:2000651 | positive regulation of sodium ion transport(GO:0010765) positive regulation of sodium ion transmembrane transport(GO:1902307) regulation of potassium ion import(GO:1903286) positive regulation of potassium ion import(GO:1903288) positive regulation of sodium ion transmembrane transporter activity(GO:2000651) |
| 0.0 | 0.4 | GO:0043249 | erythrocyte maturation(GO:0043249) |
| 0.0 | 0.2 | GO:0051764 | actin crosslink formation(GO:0051764) |
| 0.0 | 0.3 | GO:0007274 | neuromuscular synaptic transmission(GO:0007274) |
| 0.0 | 0.1 | GO:0070293 | renal absorption(GO:0070293) |
| 0.0 | 0.4 | GO:0060997 | dendritic spine morphogenesis(GO:0060997) |
| 0.0 | 0.1 | GO:0034389 | lipid particle organization(GO:0034389) |
| 0.0 | 0.3 | GO:0048251 | elastic fiber assembly(GO:0048251) |
| 0.0 | 0.3 | GO:0043171 | peptide catabolic process(GO:0043171) |
| 0.0 | 0.1 | GO:0042554 | superoxide anion generation(GO:0042554) |
| 0.0 | 0.5 | GO:0034341 | response to interferon-gamma(GO:0034341) cellular response to interferon-gamma(GO:0071346) |
| 0.0 | 0.2 | GO:0055003 | cardiac myofibril assembly(GO:0055003) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 0.1 | 0.3 | GO:0033391 | chromatoid body(GO:0033391) |
| 0.1 | 0.3 | GO:0005592 | collagen type XI trimer(GO:0005592) |
| 0.1 | 0.3 | GO:0042824 | MHC class I peptide loading complex(GO:0042824) |
| 0.1 | 0.3 | GO:0018444 | translation release factor complex(GO:0018444) |
| 0.1 | 0.8 | GO:0036157 | outer dynein arm(GO:0036157) |
| 0.0 | 0.5 | GO:0005945 | 6-phosphofructokinase complex(GO:0005945) |
| 0.0 | 0.5 | GO:0030992 | intraciliary transport particle B(GO:0030992) |
| 0.0 | 0.1 | GO:0005913 | cell-cell adherens junction(GO:0005913) zonula adherens(GO:0005915) |
| 0.0 | 0.8 | GO:0097610 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.0 | 0.1 | GO:1990726 | Lsm1-7-Pat1 complex(GO:1990726) |
| 0.0 | 0.2 | GO:0033162 | melanosome membrane(GO:0033162) chitosome(GO:0045009) |
| 0.0 | 0.4 | GO:0090665 | dystrophin-associated glycoprotein complex(GO:0016010) glycoprotein complex(GO:0090665) |
| 0.0 | 0.4 | GO:0000407 | pre-autophagosomal structure(GO:0000407) |
| 0.0 | 0.2 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.0 | 0.1 | GO:0043020 | NADPH oxidase complex(GO:0043020) |
| 0.0 | 0.2 | GO:0005847 | mRNA cleavage and polyadenylation specificity factor complex(GO:0005847) |
| 0.0 | 0.3 | GO:0005791 | rough endoplasmic reticulum(GO:0005791) |
| 0.0 | 0.1 | GO:0005892 | acetylcholine-gated channel complex(GO:0005892) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | GO:0008746 | NAD(P)+ transhydrogenase activity(GO:0008746) oxidoreductase activity, acting on NAD(P)H, NAD(P) as acceptor(GO:0016652) |
| 0.1 | 0.6 | GO:0015355 | secondary active monocarboxylate transmembrane transporter activity(GO:0015355) |
| 0.1 | 0.4 | GO:0005093 | Rab GDP-dissociation inhibitor activity(GO:0005093) |
| 0.1 | 0.3 | GO:0046978 | TAP1 binding(GO:0046978) |
| 0.1 | 0.3 | GO:1990825 | sequence-specific mRNA binding(GO:1990825) |
| 0.1 | 0.6 | GO:0034648 | histone demethylase activity (H3-dimethyl-K4 specific)(GO:0034648) |
| 0.1 | 0.2 | GO:0010852 | cyclase inhibitor activity(GO:0010852) guanylate cyclase inhibitor activity(GO:0030251) |
| 0.1 | 0.3 | GO:0005528 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.1 | 0.3 | GO:0033857 | inositol heptakisphosphate kinase activity(GO:0000829) diphosphoinositol-pentakisphosphate kinase activity(GO:0033857) |
| 0.1 | 0.3 | GO:0047757 | chondroitin-glucuronate 5-epimerase activity(GO:0047757) |
| 0.0 | 0.2 | GO:0004427 | inorganic diphosphatase activity(GO:0004427) |
| 0.0 | 0.5 | GO:0003872 | 6-phosphofructokinase activity(GO:0003872) |
| 0.0 | 0.1 | GO:0070097 | delta-catenin binding(GO:0070097) |
| 0.0 | 0.4 | GO:0031994 | insulin-like growth factor I binding(GO:0031994) insulin-like growth factor II binding(GO:0031995) |
| 0.0 | 0.1 | GO:0072571 | ADP-D-ribose binding(GO:0072570) mono-ADP-D-ribose binding(GO:0072571) |
| 0.0 | 0.5 | GO:0031682 | G-protein gamma-subunit binding(GO:0031682) |
| 0.0 | 0.2 | GO:0047429 | nucleoside-triphosphate diphosphatase activity(GO:0047429) |
| 0.0 | 0.3 | GO:0003834 | beta-carotene 15,15'-monooxygenase activity(GO:0003834) carotenoid dioxygenase activity(GO:0010436) |
| 0.0 | 0.1 | GO:0005499 | vitamin D binding(GO:0005499) |
| 0.0 | 0.1 | GO:0016531 | copper chaperone activity(GO:0016531) |
| 0.0 | 0.4 | GO:0031078 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.0 | 1.1 | GO:0005518 | collagen binding(GO:0005518) |
| 0.0 | 0.5 | GO:0031726 | CCR1 chemokine receptor binding(GO:0031726) CCR4 chemokine receptor binding(GO:0031729) |
| 0.0 | 0.9 | GO:0051018 | protein kinase A binding(GO:0051018) |
| 0.0 | 0.2 | GO:0005105 | type 1 fibroblast growth factor receptor binding(GO:0005105) |
| 0.0 | 0.4 | GO:0038191 | neuropilin binding(GO:0038191) |
| 0.0 | 0.1 | GO:0005252 | open rectifier potassium channel activity(GO:0005252) |
| 0.0 | 0.1 | GO:0047173 | phosphatidylcholine-retinol O-acyltransferase activity(GO:0047173) |
| 0.0 | 0.8 | GO:0045505 | dynein intermediate chain binding(GO:0045505) |
| 0.0 | 0.2 | GO:0044323 | retinoic acid-responsive element binding(GO:0044323) |
| 0.0 | 0.2 | GO:0004745 | retinol dehydrogenase activity(GO:0004745) |
| 0.0 | 0.6 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.0 | 0.1 | GO:0005006 | epidermal growth factor-activated receptor activity(GO:0005006) |
| 0.0 | 0.1 | GO:0015193 | L-proline transmembrane transporter activity(GO:0015193) |
| 0.0 | 0.4 | GO:0048185 | activin binding(GO:0048185) |
| 0.0 | 0.3 | GO:0004703 | G-protein coupled receptor kinase activity(GO:0004703) |
| 0.0 | 0.1 | GO:0031544 | procollagen-proline 3-dioxygenase activity(GO:0019797) peptidyl-proline 3-dioxygenase activity(GO:0031544) |
| 0.0 | 0.2 | GO:0005158 | insulin receptor binding(GO:0005158) |
| 0.0 | 0.2 | GO:0048018 | receptor activator activity(GO:0030546) receptor agonist activity(GO:0048018) |
| 0.0 | 0.1 | GO:0015189 | L-lysine transmembrane transporter activity(GO:0015189) |
| 0.0 | 0.2 | GO:0003917 | DNA topoisomerase type I activity(GO:0003917) |
| 0.0 | 0.2 | GO:0019869 | chloride channel inhibitor activity(GO:0019869) potassium channel inhibitor activity(GO:0019870) |
| 0.0 | 0.4 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.0 | 0.1 | GO:0034714 | type III transforming growth factor beta receptor binding(GO:0034714) |
| 0.0 | 0.1 | GO:0005161 | platelet-derived growth factor receptor binding(GO:0005161) |
| 0.0 | 0.1 | GO:0050700 | CARD domain binding(GO:0050700) |
| 0.0 | 0.1 | GO:0016175 | superoxide-generating NADPH oxidase activity(GO:0016175) |
| 0.0 | 0.1 | GO:0005052 | peroxisome matrix targeting signal-1 binding(GO:0005052) |
| 0.0 | 0.3 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.0 | 0.1 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.7 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 0.3 | PID ALK2 PATHWAY | ALK2 signaling events |
| 0.0 | 0.8 | PID HES HEY PATHWAY | Notch-mediated HES/HEY network |
| 0.0 | 0.5 | PID HEDGEHOG GLI PATHWAY | Hedgehog signaling events mediated by Gli proteins |
| 0.0 | 0.3 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
| 0.0 | 0.2 | PID SYNDECAN 4 PATHWAY | Syndecan-4-mediated signaling events |
| 0.0 | 0.3 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.0 | 0.0 | PID THROMBIN PAR4 PATHWAY | PAR4-mediated thrombin signaling events |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | REACTOME ENDOSOMAL VACUOLAR PATHWAY | Genes involved in Endosomal/Vacuolar pathway |
| 0.0 | 0.6 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.0 | 0.4 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.0 | 0.2 | REACTOME NEGATIVE REGULATION OF THE PI3K AKT NETWORK | Genes involved in Negative regulation of the PI3K/AKT network |
| 0.0 | 0.8 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.0 | 0.1 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 0.0 | 0.3 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |
| 0.0 | 0.2 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.0 | 0.2 | REACTOME TRAF3 DEPENDENT IRF ACTIVATION PATHWAY | Genes involved in TRAF3-dependent IRF activation pathway |
| 0.0 | 0.6 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.0 | 0.2 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.0 | 0.1 | REACTOME IL 6 SIGNALING | Genes involved in Interleukin-6 signaling |
| 0.0 | 0.2 | REACTOME MITOCHONDRIAL TRNA AMINOACYLATION | Genes involved in Mitochondrial tRNA aminoacylation |
| 0.0 | 1.0 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |