Project
PRJNA438478: RNAseq of wild type zebrafish germline
Navigation
Downloads
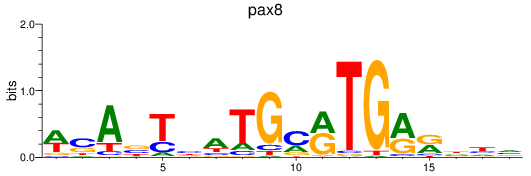
Results for pax8
Z-value: 0.34
Transcription factors associated with pax8
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
pax8
|
ENSDARG00000015879 | paired box 8 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| pax8 | dr11_v1_chr5_-_72415578_72415578 | 0.94 | 4.9e-09 | Click! |
Activity profile of pax8 motif
Sorted Z-values of pax8 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr25_-_7520937 | 1.55 |
ENSDART00000170050
|
cdkn1cb
|
cyclin-dependent kinase inhibitor 1Cb |
| chr10_+_2715548 | 0.78 |
ENSDART00000130793
|
grk5
|
G protein-coupled receptor kinase 5 |
| chr9_+_17348745 | 0.67 |
ENSDART00000147488
|
slain1a
|
SLAIN motif family, member 1a |
| chr1_+_55239710 | 0.66 |
ENSDART00000174846
|
si:ch211-286b5.2
|
si:ch211-286b5.2 |
| chr9_+_38712474 | 0.63 |
ENSDART00000182874
|
kansl1l
|
KAT8 regulatory NSL complex subunit 1-like |
| chr22_-_31752937 | 0.62 |
ENSDART00000169611
|
CABZ01028298.1
|
|
| chr23_+_42813415 | 0.59 |
ENSDART00000055577
|
myl9a
|
myosin, light chain 9a, regulatory |
| chr7_+_19374683 | 0.59 |
ENSDART00000162700
|
snrpf
|
small nuclear ribonucleoprotein polypeptide F |
| chr6_-_47840465 | 0.55 |
ENSDART00000141986
|
lrig2
|
leucine-rich repeats and immunoglobulin-like domains 2 |
| chr15_+_16387088 | 0.51 |
ENSDART00000101789
|
flot2b
|
flotillin 2b |
| chr22_-_25033105 | 0.50 |
ENSDART00000124220
|
nptxrb
|
neuronal pentraxin receptor b |
| chr7_-_52388090 | 0.46 |
ENSDART00000174104
|
wdr93
|
WD repeat domain 93 |
| chr23_+_26733232 | 0.37 |
ENSDART00000035080
|
zgc:158263
|
zgc:158263 |
| chr4_+_25630555 | 0.37 |
ENSDART00000133425
|
acot15
|
acyl-CoA thioesterase 15 |
| chr7_+_19552381 | 0.35 |
ENSDART00000169060
|
si:ch211-212k18.5
|
si:ch211-212k18.5 |
| chr14_+_48062180 | 0.34 |
ENSDART00000056713
|
ppid
|
peptidylprolyl isomerase D |
| chr6_+_168962 | 0.32 |
ENSDART00000182631
|
CABZ01066694.1
|
|
| chr19_+_46372115 | 0.32 |
ENSDART00000163935
|
med30
|
mediator complex subunit 30 |
| chr21_-_22928214 | 0.32 |
ENSDART00000182760
|
dub
|
duboraya |
| chr16_+_43347966 | 0.30 |
ENSDART00000171308
|
zmp:0000000930
|
zmp:0000000930 |
| chr11_-_165288 | 0.29 |
ENSDART00000108703
ENSDART00000173151 |
tegt
|
testis enhanced gene transcript (BAX inhibitor 1) |
| chr7_-_28549361 | 0.29 |
ENSDART00000173918
ENSDART00000054368 ENSDART00000113313 |
st5
|
suppression of tumorigenicity 5 |
| chr11_+_355305 | 0.28 |
ENSDART00000147426
|
pdrg1
|
p53 and DNA-damage regulated 1 |
| chr7_+_10610791 | 0.27 |
ENSDART00000166064
|
fah
|
fumarylacetoacetate hydrolase (fumarylacetoacetase) |
| chr6_-_436658 | 0.27 |
ENSDART00000191515
|
grap2b
|
GRB2-related adaptor protein 2b |
| chr12_-_13556536 | 0.26 |
ENSDART00000166396
|
CR354395.1
|
|
| chr25_-_3139805 | 0.25 |
ENSDART00000166625
|
epx
|
eosinophil peroxidase |
| chr5_-_68826177 | 0.24 |
ENSDART00000136605
|
si:ch211-283h6.4
|
si:ch211-283h6.4 |
| chr25_-_17579701 | 0.20 |
ENSDART00000073684
|
mmp15a
|
matrix metallopeptidase 15a |
| chr23_+_32039386 | 0.20 |
ENSDART00000133801
|
mylk2
|
myosin light chain kinase 2 |
| chr20_+_32224405 | 0.19 |
ENSDART00000062993
ENSDART00000147448 |
sesn1
|
sestrin 1 |
| chr7_+_20019125 | 0.18 |
ENSDART00000186391
|
bcl6b
|
B-cell CLL/lymphoma 6, member B |
| chr11_-_37693019 | 0.17 |
ENSDART00000102898
|
zgc:158258
|
zgc:158258 |
| chr1_-_30457062 | 0.17 |
ENSDART00000185318
ENSDART00000157924 ENSDART00000161380 |
igf2bp2b
|
insulin-like growth factor 2 mRNA binding protein 2b |
| chr20_+_35445462 | 0.17 |
ENSDART00000124497
|
tdrd6
|
tudor domain containing 6 |
| chr6_-_130849 | 0.16 |
ENSDART00000108710
|
LRRC8E
|
si:zfos-323e3.4 |
| chr2_+_45479841 | 0.16 |
ENSDART00000151856
|
si:ch211-66k16.28
|
si:ch211-66k16.28 |
| chr24_-_35699595 | 0.15 |
ENSDART00000167990
|
mapre2
|
microtubule-associated protein, RP/EB family, member 2 |
| chr8_+_4809156 | 0.14 |
ENSDART00000143720
|
tmem127
|
transmembrane protein 127 |
| chr14_-_29906209 | 0.14 |
ENSDART00000192952
|
sorbs2b
|
sorbin and SH3 domain containing 2b |
| chr7_-_8981507 | 0.13 |
ENSDART00000161422
|
si:ch211-183d5.2
|
si:ch211-183d5.2 |
| chr10_+_40598791 | 0.13 |
ENSDART00000131895
|
taar17a
|
trace amine associated receptor 17a |
| chr17_-_33405301 | 0.13 |
ENSDART00000157089
|
BX323819.1
|
|
| chr23_+_44049509 | 0.13 |
ENSDART00000102003
|
txk
|
TXK tyrosine kinase |
| chr12_+_4900981 | 0.12 |
ENSDART00000171507
|
cd79b
|
CD79b molecule, immunoglobulin-associated beta |
| chr6_+_25215944 | 0.12 |
ENSDART00000165456
|
si:ch73-97h19.2
|
si:ch73-97h19.2 |
| chr23_+_7505943 | 0.12 |
ENSDART00000135787
|
hck
|
HCK proto-oncogene, Src family tyrosine kinase |
| chr20_-_14680897 | 0.11 |
ENSDART00000063857
ENSDART00000161314 |
scrn2
|
secernin 2 |
| chr3_+_46444999 | 0.11 |
ENSDART00000186530
|
si:ch211-66e2.3
|
si:ch211-66e2.3 |
| chr10_+_24660017 | 0.11 |
ENSDART00000079597
|
vps36
|
vacuolar protein sorting 36 homolog (S. cerevisiae) |
| chr6_-_10927766 | 0.11 |
ENSDART00000134327
|
ccr7
|
chemokine (C-C motif) receptor 7 |
| chr15_+_15314189 | 0.09 |
ENSDART00000156830
|
camkk1b
|
calcium/calmodulin-dependent protein kinase kinase 1, alpha b |
| chr21_-_45891262 | 0.09 |
ENSDART00000169816
|
galnt10
|
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 10 (GalNAc-T10) |
| chr10_+_24660225 | 0.09 |
ENSDART00000190695
|
vps36
|
vacuolar protein sorting 36 homolog (S. cerevisiae) |
| chr9_+_23665777 | 0.08 |
ENSDART00000060905
|
gypc
|
glycophorin C (Gerbich blood group) |
| chr22_+_2598998 | 0.07 |
ENSDART00000176665
|
CU570894.1
|
|
| chr14_-_32491127 | 0.06 |
ENSDART00000186724
|
mcf2a
|
MCF.2 cell line derived transforming sequence a |
| chr2_-_5942355 | 0.06 |
ENSDART00000110469
|
tmem125b
|
transmembrane protein 125b |
| chr6_-_9565526 | 0.06 |
ENSDART00000151470
|
map3k2
|
mitogen-activated protein kinase kinase kinase 2 |
| chr16_-_31435020 | 0.06 |
ENSDART00000138508
|
zgc:194210
|
zgc:194210 |
| chr15_+_638457 | 0.06 |
ENSDART00000157162
|
znf1013
|
zinc finger protein 1013 |
| chr25_+_10834903 | 0.05 |
ENSDART00000155247
|
si:ch211-147g22.5
|
si:ch211-147g22.5 |
| chr24_+_684924 | 0.05 |
ENSDART00000186103
|
si:ch211-188f17.1
|
si:ch211-188f17.1 |
| chr22_+_1751640 | 0.05 |
ENSDART00000162093
|
znf1169
|
zinc finger protein 1169 |
| chr22_-_16412496 | 0.04 |
ENSDART00000137497
|
cts12
|
cathepsin 12 |
| chr23_+_43255328 | 0.04 |
ENSDART00000102712
|
tgm2a
|
transglutaminase 2, C polypeptide A |
| chr1_+_51721851 | 0.04 |
ENSDART00000040397
|
prdx2
|
peroxiredoxin 2 |
| chr4_+_36489448 | 0.03 |
ENSDART00000143181
|
znf1149
|
zinc finger protein 1149 |
| chr2_+_45552901 | 0.03 |
ENSDART00000131701
|
fndc7a
|
fibronectin type III domain containing 7a |
| chr5_-_27438812 | 0.03 |
ENSDART00000078755
|
drd7
|
dopamine receptor D7 |
| chr7_+_2797795 | 0.02 |
ENSDART00000161208
|
si:ch211-217i17.1
|
si:ch211-217i17.1 |
| chr6_-_131401 | 0.02 |
ENSDART00000151251
|
LRRC8E
|
si:zfos-323e3.4 |
| chr16_+_41135018 | 0.02 |
ENSDART00000139435
|
si:ch211-53m15.2
|
si:ch211-53m15.2 |
| chr5_-_29112956 | 0.02 |
ENSDART00000132726
|
whrnb
|
whirlin b |
| chr22_+_5959984 | 0.02 |
ENSDART00000179378
ENSDART00000178985 ENSDART00000174635 |
BX322587.2
|
|
| chr22_-_4813345 | 0.02 |
ENSDART00000190650
|
CU570881.2
|
|
| chr4_-_17803784 | 0.01 |
ENSDART00000187195
|
spi2
|
Spi-2 proto-oncogene |
| chr15_-_34892664 | 0.01 |
ENSDART00000153787
ENSDART00000099721 |
rnf183
|
ring finger protein 183 |
| chr2_-_985417 | 0.01 |
ENSDART00000140540
|
si:ch211-241e1.3
|
si:ch211-241e1.3 |
| chr14_-_1313480 | 0.00 |
ENSDART00000097748
|
il21
|
interleukin 21 |
| chr16_+_49601838 | 0.00 |
ENSDART00000168570
ENSDART00000159236 |
si:dkey-82o10.4
|
si:dkey-82o10.4 |
| chr5_+_20437124 | 0.00 |
ENSDART00000159001
|
tmem119a
|
transmembrane protein 119a |
| chr25_+_37285737 | 0.00 |
ENSDART00000126879
|
tmed6
|
transmembrane p24 trafficking protein 6 |
| chr5_+_4298636 | 0.00 |
ENSDART00000100061
|
prdx4
|
peroxiredoxin 4 |
| chr10_+_36037977 | 0.00 |
ENSDART00000164678
|
katnal1
|
katanin p60 subunit A-like 1 |
| chr10_+_25726694 | 0.00 |
ENSDART00000140308
|
ugt5d1
|
UDP glucuronosyltransferase 5 family, polypeptide D1 |
| chr3_-_16110100 | 0.00 |
ENSDART00000051807
|
lasp1
|
LIM and SH3 protein 1 |
Network of associatons between targets according to the STRING database.
First level regulatory network of pax8
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.6 | GO:0045736 | negative regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0045736) negative regulation of cyclin-dependent protein kinase activity(GO:1904030) |
| 0.1 | 0.3 | GO:0061341 | non-canonical Wnt signaling pathway involved in heart development(GO:0061341) |
| 0.1 | 0.3 | GO:0035994 | response to muscle stretch(GO:0035994) |
| 0.0 | 0.3 | GO:0006572 | tyrosine catabolic process(GO:0006572) |
| 0.0 | 0.5 | GO:0045661 | regulation of myoblast differentiation(GO:0045661) |
| 0.0 | 0.2 | GO:1901031 | response to leucine(GO:0043201) cellular response to leucine(GO:0071233) regulation of response to reactive oxygen species(GO:1901031) |
| 0.0 | 0.1 | GO:0038083 | peptidyl-tyrosine autophosphorylation(GO:0038083) |
| 0.0 | 0.2 | GO:0043328 | protein targeting to vacuole involved in ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway(GO:0043328) |
| 0.0 | 0.6 | GO:0000387 | spliceosomal snRNP assembly(GO:0000387) |
| 0.0 | 0.1 | GO:1904825 | protein localization to microtubule(GO:0035372) protein localization to microtubule plus-end(GO:1904825) |
| 0.0 | 0.2 | GO:0030719 | P granule organization(GO:0030719) |
| 0.0 | 0.3 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0016600 | flotillin complex(GO:0016600) |
| 0.1 | 0.2 | GO:0033391 | chromatoid body(GO:0033391) |
| 0.0 | 0.6 | GO:0044545 | NSL complex(GO:0044545) |
| 0.0 | 0.2 | GO:0000814 | ESCRT II complex(GO:0000814) |
| 0.0 | 0.3 | GO:0016272 | prefoldin complex(GO:0016272) |
| 0.0 | 0.1 | GO:0019815 | B cell receptor complex(GO:0019815) |
| 0.0 | 0.6 | GO:0005732 | small nucleolar ribonucleoprotein complex(GO:0005732) |
| 0.0 | 0.2 | GO:0061700 | GATOR2 complex(GO:0061700) |
| 0.0 | 0.5 | GO:0045271 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.6 | GO:0004861 | cyclin-dependent protein serine/threonine kinase inhibitor activity(GO:0004861) |
| 0.1 | 0.4 | GO:0001729 | ceramide kinase activity(GO:0001729) |
| 0.1 | 0.6 | GO:0035035 | histone acetyltransferase binding(GO:0035035) |
| 0.1 | 0.3 | GO:0016823 | hydrolase activity, acting on acid carbon-carbon bonds(GO:0016822) hydrolase activity, acting on acid carbon-carbon bonds, in ketonic substances(GO:0016823) |
| 0.0 | 0.8 | GO:0004703 | G-protein coupled receptor kinase activity(GO:0004703) |
| 0.0 | 0.2 | GO:0070728 | leucine binding(GO:0070728) |
| 0.0 | 0.1 | GO:0070004 | cysteine-type exopeptidase activity(GO:0070004) |
| 0.0 | 0.3 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 0.0 | 0.2 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 0.0 | 0.3 | GO:0008307 | structural constituent of muscle(GO:0008307) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.9 | PID ALPHA SYNUCLEIN PATHWAY | Alpha-synuclein signaling |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | REACTOME SLBP DEPENDENT PROCESSING OF REPLICATION DEPENDENT HISTONE PRE MRNAS | Genes involved in SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs |
| 0.0 | 0.1 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |