Project
PRJNA438478: RNAseq of wild type zebrafish germline
Navigation
Downloads
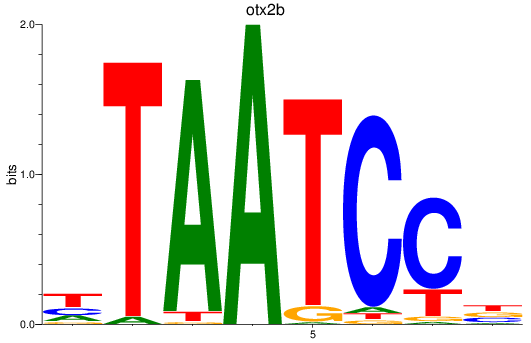
Results for otx2b
Z-value: 0.46
Transcription factors associated with otx2b
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
otx2b
|
ENSDARG00000011235 | orthodenticle homeobox 2b |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| otx2b | dr11_v1_chr17_-_44249538_44249538 | 0.20 | 4.3e-01 | Click! |
Activity profile of otx2b motif
Sorted Z-values of otx2b motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr21_-_40782393 | 0.59 |
ENSDART00000075808
|
apbb3
|
amyloid beta (A4) precursor protein-binding, family B, member 3 |
| chr3_-_61593274 | 0.53 |
ENSDART00000154132
ENSDART00000055071 |
nptx2a
|
neuronal pentraxin 2a |
| chr10_-_7988396 | 0.53 |
ENSDART00000141445
ENSDART00000024282 |
ewsr1a
|
EWS RNA-binding protein 1a |
| chr17_+_10593398 | 0.51 |
ENSDART00000168897
ENSDART00000193989 ENSDART00000191664 ENSDART00000167188 |
mapkbp1
|
mitogen-activated protein kinase binding protein 1 |
| chr18_+_7639401 | 0.50 |
ENSDART00000092416
|
rabl2
|
RAB, member of RAS oncogene family-like 2 |
| chr16_+_42829735 | 0.46 |
ENSDART00000014956
|
polr3glb
|
polymerase (RNA) III (DNA directed) polypeptide G like b |
| chr2_-_1569250 | 0.46 |
ENSDART00000167202
|
dab1b
|
Dab, reelin signal transducer, homolog 1b (Drosophila) |
| chrM_+_15308 | 0.46 |
ENSDART00000093625
|
mt-cyb
|
cytochrome b, mitochondrial |
| chr19_-_8880688 | 0.43 |
ENSDART00000039629
|
celf3a
|
cugbp, Elav-like family member 3a |
| chr22_+_30184039 | 0.42 |
ENSDART00000049075
|
add3a
|
adducin 3 (gamma) a |
| chr3_+_26144765 | 0.38 |
ENSDART00000146267
ENSDART00000043932 |
atp2a1
|
ATPase sarcoplasmic/endoplasmic reticulum Ca2+ transporting 1 |
| chr16_+_42830152 | 0.38 |
ENSDART00000159730
|
polr3glb
|
polymerase (RNA) III (DNA directed) polypeptide G like b |
| chr24_+_3478871 | 0.37 |
ENSDART00000111491
ENSDART00000134598 ENSDART00000142407 |
wdr37
|
WD repeat domain 37 |
| chr9_+_26103814 | 0.36 |
ENSDART00000026011
|
efnb2a
|
ephrin-B2a |
| chr21_-_41369370 | 0.35 |
ENSDART00000159290
|
cpeb4b
|
cytoplasmic polyadenylation element binding protein 4b |
| chr22_-_910926 | 0.33 |
ENSDART00000180075
|
FP016205.1
|
|
| chr20_-_16156419 | 0.33 |
ENSDART00000037420
|
ralgps2
|
Ral GEF with PH domain and SH3 binding motif 2 |
| chr24_+_24923166 | 0.33 |
ENSDART00000065288
|
pcyt1ba
|
phosphate cytidylyltransferase 1, choline, beta a |
| chr7_+_38395197 | 0.33 |
ENSDART00000138669
|
cep89
|
centrosomal protein 89 |
| chr5_-_39171302 | 0.31 |
ENSDART00000020808
|
paqr3a
|
progestin and adipoQ receptor family member IIIa |
| chr18_+_10784730 | 0.31 |
ENSDART00000028938
|
mical3a
|
microtubule associated monooxygenase, calponin and LIM domain containing 3a |
| chr25_+_19041329 | 0.29 |
ENSDART00000153467
|
lrtm2b
|
leucine-rich repeats and transmembrane domains 2b |
| chr7_-_53117131 | 0.29 |
ENSDART00000169211
ENSDART00000168890 ENSDART00000172179 ENSDART00000167882 |
cdh1
|
cadherin 1, type 1, E-cadherin (epithelial) |
| chr17_+_50701748 | 0.29 |
ENSDART00000191938
ENSDART00000183220 ENSDART00000049464 |
fermt2
|
fermitin family member 2 |
| chr25_+_19753890 | 0.29 |
ENSDART00000190268
ENSDART00000128852 |
celsr1b
|
cadherin EGF LAG seven-pass G-type receptor 1b |
| chr5_+_43072951 | 0.28 |
ENSDART00000193905
ENSDART00000026020 |
wdr54
|
WD repeat domain 54 |
| chr19_-_9503473 | 0.28 |
ENSDART00000091615
|
iffo1a
|
intermediate filament family orphan 1a |
| chr20_-_21672970 | 0.28 |
ENSDART00000133286
|
si:ch211-207i1.2
|
si:ch211-207i1.2 |
| chr5_-_27994679 | 0.27 |
ENSDART00000132740
|
ppp3cca
|
protein phosphatase 3, catalytic subunit, gamma isozyme, a |
| chr18_+_6155258 | 0.27 |
ENSDART00000092804
|
otogl
|
otogelin-like |
| chr22_+_16446052 | 0.26 |
ENSDART00000142454
|
si:dkey-121a11.3
|
si:dkey-121a11.3 |
| chr11_+_2506516 | 0.26 |
ENSDART00000130886
ENSDART00000189767 |
NABP2
|
si:ch73-190f16.2 |
| chr1_-_17650223 | 0.25 |
ENSDART00000043484
|
si:dkey-256e7.5
|
si:dkey-256e7.5 |
| chr14_+_34966598 | 0.24 |
ENSDART00000004550
|
rnf145a
|
ring finger protein 145a |
| chr7_+_66884570 | 0.24 |
ENSDART00000082664
|
sbf2
|
SET binding factor 2 |
| chr7_+_20471315 | 0.24 |
ENSDART00000173714
|
si:dkey-19b23.13
|
si:dkey-19b23.13 |
| chr15_-_18574716 | 0.24 |
ENSDART00000142010
ENSDART00000019006 |
ncam1b
|
neural cell adhesion molecule 1b |
| chr21_-_41369539 | 0.23 |
ENSDART00000187546
|
cpeb4b
|
cytoplasmic polyadenylation element binding protein 4b |
| chr24_-_31194847 | 0.23 |
ENSDART00000158808
|
cnn3a
|
calponin 3, acidic a |
| chr11_-_27702778 | 0.23 |
ENSDART00000045942
ENSDART00000125352 |
phf2
|
PHD finger protein 2 |
| chr14_-_25034142 | 0.23 |
ENSDART00000000392
|
pde6a
|
phosphodiesterase 6A, cGMP-specific, rod, alpha |
| chr18_+_6638726 | 0.23 |
ENSDART00000142755
ENSDART00000167781 |
c2cd5
|
C2 calcium-dependent domain containing 5 |
| chr7_+_19903924 | 0.23 |
ENSDART00000159112
|
si:ch211-285j22.3
|
si:ch211-285j22.3 |
| chr12_-_35582521 | 0.22 |
ENSDART00000162175
ENSDART00000168958 |
sec24c
|
SEC24 homolog C, COPII coat complex component |
| chr15_-_15469079 | 0.22 |
ENSDART00000132637
ENSDART00000004220 |
rab34a
|
RAB34, member RAS oncogene family a |
| chr18_+_6638974 | 0.21 |
ENSDART00000162398
|
c2cd5
|
C2 calcium-dependent domain containing 5 |
| chr8_-_25566347 | 0.21 |
ENSDART00000138289
ENSDART00000078022 |
prex1
|
phosphatidylinositol-3,4,5-trisphosphate-dependent Rac exchange factor 1 |
| chr7_+_66884291 | 0.20 |
ENSDART00000187499
|
sbf2
|
SET binding factor 2 |
| chr7_+_19904136 | 0.20 |
ENSDART00000173452
|
si:ch211-285j22.3
|
si:ch211-285j22.3 |
| chr7_-_18656069 | 0.20 |
ENSDART00000021559
|
coro1b
|
coronin, actin binding protein, 1B |
| chr16_-_33059246 | 0.20 |
ENSDART00000171718
ENSDART00000168305 ENSDART00000166401 |
snap91
|
synaptosomal-associated protein 91 |
| chr12_-_35582683 | 0.20 |
ENSDART00000167933
|
sec24c
|
SEC24 homolog C, COPII coat complex component |
| chr25_-_7925269 | 0.20 |
ENSDART00000014274
|
glcea
|
glucuronic acid epimerase a |
| chr15_+_37559570 | 0.20 |
ENSDART00000085522
|
hspb6
|
heat shock protein, alpha-crystallin-related, b6 |
| chr25_-_17552299 | 0.20 |
ENSDART00000154327
|
si:dkey-44k1.5
|
si:dkey-44k1.5 |
| chr14_+_46313135 | 0.20 |
ENSDART00000172902
|
cryba1l1
|
crystallin, beta A1, like 1 |
| chr15_-_15468326 | 0.19 |
ENSDART00000161192
|
rab34a
|
RAB34, member RAS oncogene family a |
| chr12_+_15622621 | 0.19 |
ENSDART00000079784
|
plcd3b
|
phospholipase C, delta 3b |
| chr25_-_7925019 | 0.19 |
ENSDART00000183309
|
glcea
|
glucuronic acid epimerase a |
| chr17_-_20979077 | 0.19 |
ENSDART00000006676
|
phyhipla
|
phytanoyl-CoA 2-hydroxylase interacting protein-like a |
| chr14_-_32089117 | 0.19 |
ENSDART00000158014
|
si:ch211-69b22.5
|
si:ch211-69b22.5 |
| chr10_+_9222476 | 0.19 |
ENSDART00000064973
ENSDART00000145879 |
paqr3b
|
progestin and adipoQ receptor family member IIIb |
| chr16_+_32059785 | 0.18 |
ENSDART00000134459
|
si:dkey-40m6.8
|
si:dkey-40m6.8 |
| chr16_+_16265850 | 0.18 |
ENSDART00000181265
|
setd2
|
SET domain containing 2 |
| chr11_-_3860534 | 0.17 |
ENSDART00000082425
|
gata2a
|
GATA binding protein 2a |
| chr21_+_32339158 | 0.17 |
ENSDART00000161723
|
si:ch211-247j9.1
|
si:ch211-247j9.1 |
| chr15_+_32727848 | 0.17 |
ENSDART00000161361
|
postnb
|
periostin, osteoblast specific factor b |
| chr25_+_29688671 | 0.17 |
ENSDART00000073478
|
brd1b
|
bromodomain containing 1b |
| chr5_+_59494079 | 0.16 |
ENSDART00000148727
|
gtf2ird1
|
GTF2I repeat domain containing 1 |
| chr25_+_7299488 | 0.16 |
ENSDART00000184836
|
hmg20a
|
high mobility group 20A |
| chr7_-_13882988 | 0.16 |
ENSDART00000169828
|
rlbp1a
|
retinaldehyde binding protein 1a |
| chr15_-_17074393 | 0.16 |
ENSDART00000155526
|
si:ch211-24o10.6
|
si:ch211-24o10.6 |
| chr11_-_3860199 | 0.16 |
ENSDART00000082420
|
gata2a
|
GATA binding protein 2a |
| chr1_+_44523516 | 0.16 |
ENSDART00000147702
|
zdhhc5a
|
zinc finger, DHHC-type containing 5a |
| chr8_+_53928472 | 0.15 |
ENSDART00000063242
|
MMP23B
|
matrix metallopeptidase 23B |
| chr3_-_30885250 | 0.14 |
ENSDART00000109104
|
kmt5c
|
lysine methyltransferase 5C |
| chr1_-_11075403 | 0.14 |
ENSDART00000102903
ENSDART00000170290 |
dmd
|
dystrophin |
| chr17_-_2685026 | 0.14 |
ENSDART00000191014
ENSDART00000179309 |
ptpn21
|
protein tyrosine phosphatase, non-receptor type 21 |
| chr7_+_39011355 | 0.13 |
ENSDART00000173855
|
dgkza
|
diacylglycerol kinase, zeta a |
| chr15_+_32711172 | 0.13 |
ENSDART00000163936
ENSDART00000168135 |
postnb
|
periostin, osteoblast specific factor b |
| chr15_+_42599501 | 0.12 |
ENSDART00000177646
|
grik1b
|
glutamate receptor, ionotropic, kainate 1b |
| chr2_+_1989941 | 0.12 |
ENSDART00000190814
|
ssx2ipa
|
synovial sarcoma, X breakpoint 2 interacting protein a |
| chr16_+_11151699 | 0.12 |
ENSDART00000140674
|
cicb
|
capicua transcriptional repressor b |
| chr19_-_2115040 | 0.12 |
ENSDART00000020497
|
snx13
|
sorting nexin 13 |
| chr2_-_37103622 | 0.12 |
ENSDART00000137849
|
zgc:101744
|
zgc:101744 |
| chr14_-_46395408 | 0.11 |
ENSDART00000147537
|
bbs7
|
Bardet-Biedl syndrome 7 |
| chr13_+_233482 | 0.11 |
ENSDART00000102511
|
cfap36
|
cilia and flagella associated protein 36 |
| chr10_-_27199135 | 0.11 |
ENSDART00000189511
ENSDART00000180314 |
auts2a
|
autism susceptibility candidate 2a |
| chr14_-_30587814 | 0.11 |
ENSDART00000144912
ENSDART00000149714 |
tmem265
|
transmembrane protein 265 |
| chr14_+_21754521 | 0.11 |
ENSDART00000111839
|
kdm2ab
|
lysine (K)-specific demethylase 2Ab |
| chr22_+_20427170 | 0.11 |
ENSDART00000136744
|
foxq2
|
forkhead box Q2 |
| chr25_-_16818195 | 0.10 |
ENSDART00000185215
|
dyrk4
|
dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 4 |
| chr4_+_1722899 | 0.10 |
ENSDART00000021139
|
fgfr1op2
|
FGFR1 oncogene partner 2 |
| chr2_-_48171441 | 0.10 |
ENSDART00000123040
|
pfkpb
|
phosphofructokinase, platelet b |
| chr3_+_49521106 | 0.10 |
ENSDART00000162799
|
crb3a
|
crumbs homolog 3a |
| chr4_-_18436899 | 0.10 |
ENSDART00000141671
|
socs2
|
suppressor of cytokine signaling 2 |
| chr14_-_2933185 | 0.10 |
ENSDART00000161677
ENSDART00000162446 ENSDART00000109378 |
si:dkey-201i24.6
|
si:dkey-201i24.6 |
| chr10_+_4095917 | 0.09 |
ENSDART00000114533
|
mn1a
|
meningioma 1a |
| chr6_-_17999776 | 0.09 |
ENSDART00000183048
ENSDART00000181577 ENSDART00000170597 |
rptor
|
regulatory associated protein of MTOR, complex 1 |
| chr6_-_36795111 | 0.09 |
ENSDART00000160669
ENSDART00000104256 ENSDART00000187751 ENSDART00000161928 ENSDART00000183264 |
opa1
|
optic atrophy 1 (autosomal dominant) |
| chr18_+_7594012 | 0.09 |
ENSDART00000062150
|
zgc:77752
|
zgc:77752 |
| chr2_-_48171112 | 0.08 |
ENSDART00000156258
|
pfkpb
|
phosphofructokinase, platelet b |
| chr23_-_3759692 | 0.08 |
ENSDART00000028885
|
hmga1a
|
high mobility group AT-hook 1a |
| chr14_+_14847745 | 0.08 |
ENSDART00000189302
ENSDART00000188336 |
ccdc142
|
coiled-coil domain containing 142 |
| chr25_-_35996141 | 0.08 |
ENSDART00000149074
|
sall1b
|
spalt-like transcription factor 1b |
| chr25_-_19090479 | 0.08 |
ENSDART00000027465
ENSDART00000177670 |
cacna2d4b
|
calcium channel, voltage-dependent, alpha 2/delta subunit 4b |
| chr23_-_35195908 | 0.07 |
ENSDART00000122429
|
klf15
|
Kruppel-like factor 15 |
| chr22_+_5103349 | 0.07 |
ENSDART00000083474
|
atcaya
|
ataxia, cerebellar, Cayman type a |
| chr2_+_45159855 | 0.07 |
ENSDART00000056333
|
CU929150.1
|
|
| chr3_+_34220194 | 0.07 |
ENSDART00000145859
|
slc25a23b
|
solute carrier family 25 (mitochondrial carrier; phosphate carrier), member 23b |
| chr7_-_28058442 | 0.07 |
ENSDART00000173842
|
si:ch211-235p24.2
|
si:ch211-235p24.2 |
| chr9_-_8454060 | 0.07 |
ENSDART00000110158
|
irs2b
|
insulin receptor substrate 2b |
| chr3_-_28665291 | 0.07 |
ENSDART00000151670
|
fbxl16
|
F-box and leucine-rich repeat protein 16 |
| chr19_+_2631565 | 0.07 |
ENSDART00000171487
|
fam126a
|
family with sequence similarity 126, member A |
| chr3_+_1492174 | 0.07 |
ENSDART00000112979
|
sox10
|
SRY (sex determining region Y)-box 10 |
| chr7_-_64770456 | 0.07 |
ENSDART00000192618
|
zdhhc21
|
zinc finger, DHHC-type containing 21 |
| chr5_+_63767376 | 0.07 |
ENSDART00000138898
|
rgs3b
|
regulator of G protein signaling 3b |
| chr14_+_21699129 | 0.07 |
ENSDART00000073707
|
stx3a
|
syntaxin 3A |
| chr3_-_49504023 | 0.06 |
ENSDART00000168108
|
prkacaa
|
protein kinase, cAMP-dependent, catalytic, alpha, genome duplicate a |
| chr3_+_23703704 | 0.06 |
ENSDART00000024256
|
hoxb6a
|
homeobox B6a |
| chr14_+_14847304 | 0.06 |
ENSDART00000169932
|
ccdc142
|
coiled-coil domain containing 142 |
| chr1_-_54107321 | 0.06 |
ENSDART00000148382
ENSDART00000150357 |
rfx1b
|
regulatory factor X, 1b (influences HLA class II expression) |
| chr18_+_16053455 | 0.06 |
ENSDART00000189163
ENSDART00000188269 |
FO834850.1
|
|
| chr4_-_9836342 | 0.06 |
ENSDART00000150519
|
ttc41
|
tetratricopeptide repeat domain 41 |
| chr12_-_10275320 | 0.06 |
ENSDART00000170078
|
mpp2b
|
membrane protein, palmitoylated 2b (MAGUK p55 subfamily member 2) |
| chr14_-_32503363 | 0.06 |
ENSDART00000034883
|
mcf2a
|
MCF.2 cell line derived transforming sequence a |
| chr14_+_7892383 | 0.05 |
ENSDART00000063837
|
ube2d2
|
ubiquitin-conjugating enzyme E2D 2 (UBC4/5 homolog, yeast) |
| chr11_+_25010491 | 0.05 |
ENSDART00000167285
|
zgc:92107
|
zgc:92107 |
| chr7_+_39011852 | 0.05 |
ENSDART00000093009
|
dgkza
|
diacylglycerol kinase, zeta a |
| chr12_+_17042754 | 0.05 |
ENSDART00000066439
|
ch25h
|
cholesterol 25-hydroxylase |
| chr6_+_36942966 | 0.05 |
ENSDART00000028895
|
negr1
|
neuronal growth regulator 1 |
| chr23_+_7505943 | 0.04 |
ENSDART00000135787
|
hck
|
HCK proto-oncogene, Src family tyrosine kinase |
| chr20_+_6543674 | 0.04 |
ENSDART00000134204
|
tns3.1
|
tensin 3, tandem duplicate 1 |
| chr5_-_14344647 | 0.04 |
ENSDART00000188456
|
tet3
|
tet methylcytosine dioxygenase 3 |
| chr3_-_32170850 | 0.04 |
ENSDART00000055307
ENSDART00000157366 |
tnnt1
|
troponin T type 1 (skeletal, slow) |
| chr16_+_14710436 | 0.04 |
ENSDART00000027982
|
col14a1a
|
collagen, type XIV, alpha 1a |
| chr13_-_47800342 | 0.04 |
ENSDART00000121817
|
fbln7
|
fibulin 7 |
| chr8_+_14381272 | 0.04 |
ENSDART00000057642
|
acbd6
|
acyl-CoA binding domain containing 6 |
| chr11_+_31680513 | 0.04 |
ENSDART00000139900
ENSDART00000040305 |
diaph3
|
diaphanous-related formin 3 |
| chr5_-_38107741 | 0.04 |
ENSDART00000156853
|
si:ch211-284e13.14
|
si:ch211-284e13.14 |
| chr15_+_5973909 | 0.04 |
ENSDART00000126886
ENSDART00000189618 |
igsf5b
|
immunoglobulin superfamily, member 5b |
| chr25_-_18913336 | 0.04 |
ENSDART00000171010
|
parietopsin
|
parietopsin |
| chr17_+_18117029 | 0.04 |
ENSDART00000154646
ENSDART00000179739 |
bcl11ba
|
B cell CLL/lymphoma 11Ba |
| chr1_-_45146834 | 0.04 |
ENSDART00000144997
|
si:ch211-239f4.6
|
si:ch211-239f4.6 |
| chr13_-_50614639 | 0.03 |
ENSDART00000170527
|
vent
|
ventral expressed homeobox |
| chr14_-_29799993 | 0.03 |
ENSDART00000133775
ENSDART00000005568 |
pdlim3b
|
PDZ and LIM domain 3b |
| chr19_+_19729506 | 0.03 |
ENSDART00000171610
|
hoxa13a
|
homeobox A13a |
| chr14_-_36378494 | 0.03 |
ENSDART00000058503
|
gpm6aa
|
glycoprotein M6Aa |
| chr16_-_17188294 | 0.03 |
ENSDART00000165883
|
opn9
|
opsin 9 |
| chr14_-_4273396 | 0.03 |
ENSDART00000127318
|
frmpd1b
|
FERM and PDZ domain containing 1b |
| chr10_+_375042 | 0.03 |
ENSDART00000171854
|
si:ch1073-303d10.1
|
si:ch1073-303d10.1 |
| chr11_+_29671661 | 0.03 |
ENSDART00000024318
ENSDART00000165024 |
rnf207a
|
ring finger protein 207a |
| chr14_+_21699414 | 0.03 |
ENSDART00000169942
|
stx3a
|
syntaxin 3A |
| chr20_-_34768301 | 0.03 |
ENSDART00000142482
|
paqr8
|
progestin and adipoQ receptor family member VIII |
| chr17_+_16564921 | 0.03 |
ENSDART00000151904
|
foxn3
|
forkhead box N3 |
| chr8_+_23142946 | 0.02 |
ENSDART00000152933
|
si:ch211-196c10.13
|
si:ch211-196c10.13 |
| chr24_-_1303935 | 0.02 |
ENSDART00000159212
ENSDART00000159267 ENSDART00000164904 |
nrp1a
|
neuropilin 1a |
| chr2_-_50272077 | 0.02 |
ENSDART00000127623
|
cul1a
|
cullin 1a |
| chr16_+_20926673 | 0.02 |
ENSDART00000009827
|
hoxa2b
|
homeobox A2b |
| chr23_-_24394719 | 0.02 |
ENSDART00000044918
|
epha2b
|
eph receptor A2 b |
| chr18_+_5172848 | 0.02 |
ENSDART00000190642
|
CABZ01080599.1
|
|
| chr7_-_52842605 | 0.02 |
ENSDART00000083002
|
map1aa
|
microtubule-associated protein 1Aa |
| chr2_+_37110504 | 0.02 |
ENSDART00000132794
ENSDART00000042974 |
slc1a8b
|
solute carrier family 1 (glutamate transporter), member 8b |
| chr3_+_34919810 | 0.02 |
ENSDART00000055264
|
ca10b
|
carbonic anhydrase Xb |
| chr6_+_32415132 | 0.02 |
ENSDART00000155790
|
kank4
|
KN motif and ankyrin repeat domains 4 |
| chr9_-_27738110 | 0.02 |
ENSDART00000060347
|
crygs2
|
crystallin, gamma S2 |
| chr10_+_29260096 | 0.02 |
ENSDART00000088973
|
sytl2a
|
synaptotagmin-like 2a |
| chr1_-_16507812 | 0.01 |
ENSDART00000169081
|
mtmr7b
|
myotubularin related protein 7b |
| chr3_-_10970502 | 0.01 |
ENSDART00000127500
|
CR382337.1
|
|
| chr9_-_48184823 | 0.01 |
ENSDART00000180264
|
klhl23
|
kelch-like family member 23 |
| chr1_-_58009216 | 0.01 |
ENSDART00000143829
|
nxnl1
|
nucleoredoxin like 1 |
| chr1_+_55080797 | 0.01 |
ENSDART00000077390
|
lgalsla
|
lectin, galactoside-binding-like a |
| chr24_-_6260333 | 0.01 |
ENSDART00000136041
|
myo3a
|
myosin IIIA |
| chr1_-_53988017 | 0.01 |
ENSDART00000003097
|
RHOU (1 of many)
|
si:ch211-133l11.10 |
| chr20_-_1141722 | 0.01 |
ENSDART00000152675
|
gabrr1
|
gamma-aminobutyric acid (GABA) A receptor, rho 1 |
| chr7_+_41303572 | 0.01 |
ENSDART00000012692
|
rgs9bp
|
regulator of G protein signaling 9 binding protein |
| chr7_+_4384863 | 0.01 |
ENSDART00000042955
ENSDART00000134653 |
slc12a10.3
|
slc12a10.3 solute carrier family 12 (sodium/potassium/chloride transporters), member 10, tandem duplicate 3 |
| chr8_+_27743550 | 0.01 |
ENSDART00000046004
|
wnt2bb
|
wingless-type MMTV integration site family, member 2Bb |
| chr14_-_8080416 | 0.01 |
ENSDART00000045109
|
zgc:92242
|
zgc:92242 |
| chr16_-_37372613 | 0.01 |
ENSDART00000124090
|
si:ch211-208k15.1
|
si:ch211-208k15.1 |
| chr2_-_30784198 | 0.01 |
ENSDART00000182523
ENSDART00000147355 |
rgs20
|
regulator of G protein signaling 20 |
| chr9_+_33357011 | 0.01 |
ENSDART00000088569
|
nyx
|
nyctalopin |
| chr9_+_24095677 | 0.01 |
ENSDART00000150443
|
lrrfip1a
|
leucine rich repeat (in FLII) interacting protein 1a |
| chr16_+_24684107 | 0.00 |
ENSDART00000183920
|
ywhabl
|
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, beta polypeptide like |
| chr2_+_11923824 | 0.00 |
ENSDART00000153873
|
trove2
|
TROVE domain family, member 2 |
| chr8_-_22965214 | 0.00 |
ENSDART00000148178
|
emilin3a
|
elastin microfibril interfacer 3a |
| chr25_+_33002963 | 0.00 |
ENSDART00000187366
|
CABZ01046980.1
|
|
| chr13_-_25199260 | 0.00 |
ENSDART00000057605
|
adka
|
adenosine kinase a |
| chr2_+_31476065 | 0.00 |
ENSDART00000049219
|
cacnb2b
|
calcium channel, voltage-dependent, beta 2b |
| chr4_-_5077158 | 0.00 |
ENSDART00000155915
|
ahcyl2
|
adenosylhomocysteinase-like 2 |
| chr1_-_45177373 | 0.00 |
ENSDART00000143142
ENSDART00000034549 |
zgc:111983
|
zgc:111983 |
| chr16_+_37876779 | 0.00 |
ENSDART00000140148
|
si:ch211-198c19.1
|
si:ch211-198c19.1 |
| chr7_-_12821277 | 0.00 |
ENSDART00000091584
|
zgc:158785
|
zgc:158785 |
| chr17_-_12389259 | 0.00 |
ENSDART00000185724
|
snap25b
|
synaptosomal-associated protein, 25b |
| chr2_+_29795833 | 0.00 |
ENSDART00000056750
|
si:ch211-207d6.2
|
si:ch211-207d6.2 |
| chr18_+_9323211 | 0.00 |
ENSDART00000166114
|
sema3ab
|
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3Ab |
| chr18_+_49969568 | 0.00 |
ENSDART00000126916
|
mob2b
|
MOB kinase activator 2b |
Network of associatons between targets according to the STRING database.
First level regulatory network of otx2b
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0061181 | regulation of chondrocyte development(GO:0061181) |
| 0.1 | 0.3 | GO:1902895 | pri-miRNA transcription from RNA polymerase II promoter(GO:0061614) regulation of pri-miRNA transcription from RNA polymerase II promoter(GO:1902893) positive regulation of pri-miRNA transcription from RNA polymerase II promoter(GO:1902895) |
| 0.1 | 0.4 | GO:0031448 | regulation of twitch skeletal muscle contraction(GO:0014724) regulation of fast-twitch skeletal muscle fiber contraction(GO:0031446) positive regulation of fast-twitch skeletal muscle fiber contraction(GO:0031448) negative regulation of striated muscle contraction(GO:0045988) relaxation of skeletal muscle(GO:0090076) |
| 0.1 | 0.4 | GO:0010828 | positive regulation of glucose transport(GO:0010828) |
| 0.1 | 0.2 | GO:0061187 | regulation of chromatin silencing at rDNA(GO:0061187) negative regulation of chromatin silencing at rDNA(GO:0061188) |
| 0.1 | 0.4 | GO:0007412 | axon target recognition(GO:0007412) |
| 0.1 | 0.3 | GO:0046373 | arabinose metabolic process(GO:0019566) L-arabinose metabolic process(GO:0046373) |
| 0.1 | 0.5 | GO:0097476 | spinal cord motor neuron migration(GO:0097476) lateral motor column neuron migration(GO:0097477) |
| 0.1 | 0.4 | GO:0090385 | phagosome-lysosome fusion(GO:0090385) |
| 0.0 | 0.3 | GO:0090134 | mesendoderm migration(GO:0090133) cell migration involved in mesendoderm migration(GO:0090134) |
| 0.0 | 0.1 | GO:0034770 | histone H4-K20 methylation(GO:0034770) histone H4-K20 trimethylation(GO:0034773) |
| 0.0 | 0.5 | GO:0006122 | mitochondrial electron transport, ubiquinol to cytochrome c(GO:0006122) |
| 0.0 | 0.6 | GO:2000766 | negative regulation of cytoplasmic translation(GO:2000766) |
| 0.0 | 0.1 | GO:0010662 | negative regulation of muscle cell apoptotic process(GO:0010656) muscle cell apoptotic process(GO:0010657) striated muscle cell apoptotic process(GO:0010658) regulation of muscle cell apoptotic process(GO:0010660) regulation of striated muscle cell apoptotic process(GO:0010662) negative regulation of striated muscle cell apoptotic process(GO:0010664) |
| 0.0 | 0.4 | GO:0030202 | heparin metabolic process(GO:0030202) heparin biosynthetic process(GO:0030210) |
| 0.0 | 0.2 | GO:0097198 | histone H3-K36 trimethylation(GO:0097198) |
| 0.0 | 0.8 | GO:0006383 | transcription from RNA polymerase III promoter(GO:0006383) |
| 0.0 | 0.4 | GO:0090110 | cargo loading into COPII-coated vesicle(GO:0090110) |
| 0.0 | 0.4 | GO:0016203 | muscle attachment(GO:0016203) |
| 0.0 | 0.4 | GO:0048026 | positive regulation of mRNA splicing, via spliceosome(GO:0048026) |
| 0.0 | 0.1 | GO:1900028 | wound healing, spreading of epidermal cells(GO:0035313) negative regulation of ruffle assembly(GO:1900028) |
| 0.0 | 0.1 | GO:0098881 | exocytic insertion of neurotransmitter receptor to plasma membrane(GO:0098881) exocytic insertion of neurotransmitter receptor to postsynaptic membrane(GO:0098967) |
| 0.0 | 0.2 | GO:0006007 | glucose catabolic process(GO:0006007) NADH regeneration(GO:0006735) glycolytic process through fructose-6-phosphate(GO:0061615) glycolytic process through glucose-6-phosphate(GO:0061620) canonical glycolysis(GO:0061621) glucose catabolic process to pyruvate(GO:0061718) |
| 0.0 | 0.1 | GO:0046426 | negative regulation of JAK-STAT cascade(GO:0046426) negative regulation of STAT cascade(GO:1904893) |
| 0.0 | 0.1 | GO:0070584 | mitochondrion morphogenesis(GO:0070584) |
| 0.0 | 0.2 | GO:0044154 | histone H3-K14 acetylation(GO:0044154) |
| 0.0 | 0.1 | GO:0051877 | pigment granule aggregation in cell center(GO:0051877) |
| 0.0 | 0.3 | GO:0040023 | nuclear migration(GO:0007097) establishment of nucleus localization(GO:0040023) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0031673 | H zone(GO:0031673) |
| 0.0 | 0.3 | GO:0031258 | lamellipodium membrane(GO:0031258) |
| 0.0 | 0.5 | GO:0005750 | mitochondrial respiratory chain complex III(GO:0005750) respiratory chain complex III(GO:0045275) |
| 0.0 | 0.3 | GO:0005955 | calcineurin complex(GO:0005955) |
| 0.0 | 0.6 | GO:1990124 | messenger ribonucleoprotein complex(GO:1990124) |
| 0.0 | 0.8 | GO:0005666 | DNA-directed RNA polymerase III complex(GO:0005666) |
| 0.0 | 0.2 | GO:0042622 | photoreceptor outer segment membrane(GO:0042622) |
| 0.0 | 0.2 | GO:0098888 | presynaptic endocytic zone(GO:0098833) presynaptic endocytic zone membrane(GO:0098835) extrinsic component of presynaptic membrane(GO:0098888) extrinsic component of presynaptic endocytic zone(GO:0098894) |
| 0.0 | 0.4 | GO:0030127 | COPII vesicle coat(GO:0030127) |
| 0.0 | 0.2 | GO:0005945 | 6-phosphofructokinase complex(GO:0005945) |
| 0.0 | 0.3 | GO:0005640 | nuclear outer membrane(GO:0005640) |
| 0.0 | 0.4 | GO:0030687 | preribosome, large subunit precursor(GO:0030687) |
| 0.0 | 0.1 | GO:0034464 | BBSome(GO:0034464) |
| 0.0 | 0.1 | GO:0031931 | TORC1 complex(GO:0031931) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0047464 | heparosan-N-sulfate-glucuronate 5-epimerase activity(GO:0047464) |
| 0.1 | 0.5 | GO:0008121 | ubiquinol-cytochrome-c reductase activity(GO:0008121) oxidoreductase activity, acting on diphenols and related substances as donors, cytochrome as acceptor(GO:0016681) |
| 0.1 | 0.3 | GO:0046556 | alpha-L-arabinofuranosidase activity(GO:0046556) |
| 0.1 | 0.3 | GO:0004105 | choline-phosphate cytidylyltransferase activity(GO:0004105) |
| 0.0 | 0.1 | GO:0042799 | histone methyltransferase activity (H4-K20 specific)(GO:0042799) |
| 0.0 | 0.3 | GO:0033192 | calcium-dependent protein serine/threonine phosphatase activity(GO:0004723) calmodulin-dependent protein phosphatase activity(GO:0033192) |
| 0.0 | 0.6 | GO:0000900 | translation repressor activity, nucleic acid binding(GO:0000900) |
| 0.0 | 0.1 | GO:1901612 | cardiolipin binding(GO:1901612) |
| 0.0 | 0.6 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.0 | 0.1 | GO:0033130 | acetylcholine receptor binding(GO:0033130) |
| 0.0 | 0.3 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.0 | 0.4 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.0 | 0.3 | GO:0048495 | Roundabout binding(GO:0048495) |
| 0.0 | 0.2 | GO:0047555 | 3',5'-cyclic-GMP phosphodiesterase activity(GO:0047555) |
| 0.0 | 0.4 | GO:0046875 | ephrin receptor binding(GO:0046875) |
| 0.0 | 0.2 | GO:0003872 | 6-phosphofructokinase activity(GO:0003872) |
| 0.0 | 0.2 | GO:0005545 | 1-phosphatidylinositol binding(GO:0005545) |
| 0.0 | 0.2 | GO:0005221 | intracellular cyclic nucleotide activated cation channel activity(GO:0005221) intracellular cAMP activated cation channel activity(GO:0005222) intracellular cGMP activated cation channel activity(GO:0005223) cyclic nucleotide-gated ion channel activity(GO:0043855) |
| 0.0 | 0.2 | GO:0032454 | histone demethylase activity (H3-K9 specific)(GO:0032454) |
| 0.0 | 0.1 | GO:0004679 | AMP-activated protein kinase activity(GO:0004679) |
| 0.0 | 0.1 | GO:0005159 | insulin-like growth factor receptor binding(GO:0005159) |
| 0.0 | 0.2 | GO:0043994 | H3 histone acetyltransferase activity(GO:0010484) histone acetyltransferase activity (H3-K23 specific)(GO:0043994) |
| 0.0 | 0.2 | GO:0046975 | histone methyltransferase activity (H3-K36 specific)(GO:0046975) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | ST G ALPHA S PATHWAY | G alpha s Pathway |
| 0.0 | 0.2 | ST WNT CA2 CYCLIC GMP PATHWAY | Wnt/Ca2+/cyclic GMP signaling. |
| 0.0 | 0.3 | PID ECADHERIN KERATINOCYTE PATHWAY | E-cadherin signaling in keratinocytes |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.0 | 0.3 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.0 | 0.4 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.0 | 0.4 | REACTOME ANTIGEN PRESENTATION FOLDING ASSEMBLY AND PEPTIDE LOADING OF CLASS I MHC | Genes involved in Antigen Presentation: Folding, assembly and peptide loading of class I MHC |
| 0.0 | 0.2 | REACTOME CGMP EFFECTS | Genes involved in cGMP effects |