Project
PRJNA438478: RNAseq of wild type zebrafish germline
Navigation
Downloads
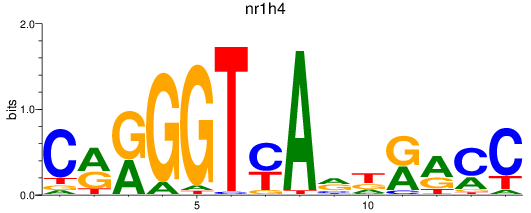
Results for nr1h4
Z-value: 0.40
Transcription factors associated with nr1h4
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
nr1h4
|
ENSDARG00000057741 | nuclear receptor subfamily 1, group H, member 4 |
|
nr1h4
|
ENSDARG00000113600 | nuclear receptor subfamily 1, group H, member 4 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| nr1h4 | dr11_v1_chr18_+_15644559_15644559 | -0.82 | 2.6e-05 | Click! |
Activity profile of nr1h4 motif
Sorted Z-values of nr1h4 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr5_+_20693724 | 1.39 |
ENSDART00000141368
|
si:ch211-240b21.2
|
si:ch211-240b21.2 |
| chr17_-_25395395 | 1.14 |
ENSDART00000170233
|
fam167b
|
family with sequence similarity 167, member B |
| chr10_-_27197044 | 0.84 |
ENSDART00000137928
|
auts2a
|
autism susceptibility candidate 2a |
| chr3_-_18030938 | 0.71 |
ENSDART00000013540
|
si:ch73-141c7.1
|
si:ch73-141c7.1 |
| chr16_+_54263921 | 0.71 |
ENSDART00000002856
|
drd2l
|
dopamine receptor D2 like |
| chr13_-_7766758 | 0.70 |
ENSDART00000171831
|
h2afy2
|
H2A histone family, member Y2 |
| chr2_+_49644803 | 0.70 |
ENSDART00000160342
|
si:ch211-209f23.7
|
si:ch211-209f23.7 |
| chr2_+_49799470 | 0.69 |
ENSDART00000146325
|
si:ch211-190k17.19
|
si:ch211-190k17.19 |
| chr11_-_22372072 | 0.66 |
ENSDART00000065996
|
tmem183a
|
transmembrane protein 183A |
| chr19_+_32166702 | 0.66 |
ENSDART00000021798
|
fabp11a
|
fatty acid binding protein 11a |
| chr5_+_25072894 | 0.66 |
ENSDART00000012268
|
mrpl41
|
mitochondrial ribosomal protein L41 |
| chr23_+_42810055 | 0.65 |
ENSDART00000186647
|
myl9a
|
myosin, light chain 9a, regulatory |
| chr1_-_17650223 | 0.65 |
ENSDART00000043484
|
si:dkey-256e7.5
|
si:dkey-256e7.5 |
| chr4_-_16545085 | 0.63 |
ENSDART00000033188
|
btg1
|
B-cell translocation gene 1, anti-proliferative |
| chr5_+_4366431 | 0.63 |
ENSDART00000168560
ENSDART00000149185 |
sat1a.2
|
spermidine/spermine N1-acetyltransferase 1a, duplicate 2 |
| chr20_+_54685143 | 0.62 |
ENSDART00000162535
|
pak7
|
p21 protein (Cdc42/Rac)-activated kinase 7 |
| chr8_-_48675775 | 0.62 |
ENSDART00000060785
|
pimr183
|
Pim proto-oncogene, serine/threonine kinase, related 183 |
| chr7_+_34688527 | 0.61 |
ENSDART00000108473
|
plekhg4
|
pleckstrin homology domain containing, family G (with RhoGef domain) member 4 |
| chr25_+_245018 | 0.60 |
ENSDART00000155344
|
zgc:92481
|
zgc:92481 |
| chr9_+_2499627 | 0.55 |
ENSDART00000160782
|
wipf1a
|
WAS/WASL interacting protein family, member 1a |
| chr12_-_30841679 | 0.53 |
ENSDART00000105594
|
crygmx
|
crystallin, gamma MX |
| chr15_-_15910226 | 0.53 |
ENSDART00000154219
|
synrg
|
synergin, gamma |
| chr11_+_5842632 | 0.53 |
ENSDART00000111374
ENSDART00000158599 |
ndufs7
|
NADH dehydrogenase (ubiquinone) Fe-S protein 7, (NADH-coenzyme Q reductase) |
| chr8_-_48675411 | 0.52 |
ENSDART00000165081
|
pimr183
|
Pim proto-oncogene, serine/threonine kinase, related 183 |
| chr25_+_5012791 | 0.51 |
ENSDART00000156970
|
si:ch73-265h17.5
|
si:ch73-265h17.5 |
| chr3_-_34027178 | 0.50 |
ENSDART00000170201
ENSDART00000151408 |
ighv1-4
ighv14-1
|
immunoglobulin heavy variable 1-4 immunoglobulin heavy variable 14-1 |
| chr23_-_22598002 | 0.47 |
ENSDART00000146966
|
si:ch211-28e16.5
|
si:ch211-28e16.5 |
| chr17_+_35431724 | 0.47 |
ENSDART00000190293
|
BX571665.1
|
|
| chr2_-_40191603 | 0.47 |
ENSDART00000180691
|
si:ch211-122l24.6
|
si:ch211-122l24.6 |
| chr15_+_6652396 | 0.47 |
ENSDART00000192813
ENSDART00000157678 |
nop53
|
NOP53 ribosome biogenesis factor |
| chr21_-_12036134 | 0.46 |
ENSDART00000031658
|
tpgs2
|
tubulin polyglutamylase complex subunit 2 |
| chr20_-_4738101 | 0.45 |
ENSDART00000050201
ENSDART00000152559 ENSDART00000053858 ENSDART00000125620 |
papola
|
poly(A) polymerase alpha |
| chr1_-_44704261 | 0.44 |
ENSDART00000133210
|
si:dkey-28b4.8
|
si:dkey-28b4.8 |
| chr6_+_9398074 | 0.43 |
ENSDART00000151772
|
kalrnb
|
kalirin RhoGEF kinase b |
| chr25_-_16969922 | 0.43 |
ENSDART00000111158
|
tigara
|
tp53-induced glycolysis and apoptosis regulator a |
| chr7_+_528593 | 0.43 |
ENSDART00000091955
|
nrxn2b
|
neurexin 2b |
| chr23_+_7471072 | 0.41 |
ENSDART00000135551
|
si:ch211-200e2.1
|
si:ch211-200e2.1 |
| chr11_-_14924138 | 0.39 |
ENSDART00000168348
|
si:dkey-6d5.1
|
si:dkey-6d5.1 |
| chr5_+_13385837 | 0.38 |
ENSDART00000191190
|
ccl19a.1
|
chemokine (C-C motif) ligand 19a, tandem duplicate 1 |
| chr4_-_18455480 | 0.38 |
ENSDART00000150221
|
sco2
|
SCO2 cytochrome c oxidase assembly protein |
| chr22_+_10645088 | 0.36 |
ENSDART00000142265
|
rassf1
|
Ras association (RalGDS/AF-6) domain family 1 |
| chr2_+_371759 | 0.35 |
ENSDART00000153788
|
si:dkey-33c14.7
|
si:dkey-33c14.7 |
| chr21_-_5879897 | 0.33 |
ENSDART00000184034
|
rpl35
|
ribosomal protein L35 |
| chr7_-_52531252 | 0.33 |
ENSDART00000174369
|
tcf12
|
transcription factor 12 |
| chr19_-_18855513 | 0.33 |
ENSDART00000162708
|
ppt2
|
palmitoyl-protein thioesterase 2 |
| chr1_-_21538329 | 0.33 |
ENSDART00000182798
ENSDART00000193518 ENSDART00000183873 ENSDART00000183376 |
wdr19
|
WD repeat domain 19 |
| chr21_-_14811058 | 0.32 |
ENSDART00000143100
|
phpt1
|
phosphohistidine phosphatase 1 |
| chr4_-_5597167 | 0.32 |
ENSDART00000132431
|
vegfab
|
vascular endothelial growth factor Ab |
| chr22_+_34663328 | 0.32 |
ENSDART00000183912
|
si:ch73-304f21.1
|
si:ch73-304f21.1 |
| chr3_-_52661242 | 0.31 |
ENSDART00000138018
|
zgc:113210
|
zgc:113210 |
| chr12_+_695619 | 0.31 |
ENSDART00000161691
|
abcc3
|
ATP-binding cassette, sub-family C (CFTR/MRP), member 3 |
| chr21_+_44112914 | 0.30 |
ENSDART00000062836
|
fgf1b
|
fibroblast growth factor 1b |
| chr7_+_39399747 | 0.30 |
ENSDART00000147037
|
tnni2b.1
|
troponin I type 2b (skeletal, fast), tandem duplicate 1 |
| chr17_+_996509 | 0.29 |
ENSDART00000158830
|
cyp1c2
|
cytochrome P450, family 1, subfamily C, polypeptide 2 |
| chr16_+_53203370 | 0.29 |
ENSDART00000154669
|
si:ch211-269k10.2
|
si:ch211-269k10.2 |
| chr18_+_22302635 | 0.29 |
ENSDART00000141051
|
carmil2
|
capping protein regulator and myosin 1 linker 2 |
| chr18_+_31056645 | 0.29 |
ENSDART00000159316
|
mvda
|
mevalonate (diphospho) decarboxylase a |
| chr2_-_6051836 | 0.28 |
ENSDART00000092479
|
si:ch211-284b7.3
|
si:ch211-284b7.3 |
| chr25_-_24046870 | 0.28 |
ENSDART00000047569
|
igf2b
|
insulin-like growth factor 2b |
| chr23_+_4253957 | 0.28 |
ENSDART00000008761
|
arl6ip5b
|
ADP-ribosylation factor-like 6 interacting protein 5b |
| chr5_-_38777852 | 0.28 |
ENSDART00000131603
|
si:dkey-58f10.4
|
si:dkey-58f10.4 |
| chr11_+_41540862 | 0.28 |
ENSDART00000173210
|
kcnab2a
|
potassium voltage-gated channel, shaker-related subfamily, beta member 2 a |
| chr5_+_54400971 | 0.28 |
ENSDART00000169695
|
bspry
|
B-box and SPRY domain containing |
| chr7_+_29153018 | 0.27 |
ENSDART00000138065
|
tmppe
|
transmembrane protein with metallophosphoesterase domain |
| chr16_-_4769877 | 0.27 |
ENSDART00000149421
ENSDART00000054078 |
rpa2
|
replication protein A2 |
| chr10_+_28428222 | 0.27 |
ENSDART00000135003
|
si:ch211-222e20.4
|
si:ch211-222e20.4 |
| chr1_-_9130136 | 0.27 |
ENSDART00000113013
|
palb2
|
partner and localizer of BRCA2 |
| chr9_-_28616436 | 0.27 |
ENSDART00000136985
|
si:ch73-7i4.2
|
si:ch73-7i4.2 |
| chr17_-_15149192 | 0.26 |
ENSDART00000180511
ENSDART00000103405 |
gch1
|
GTP cyclohydrolase 1 |
| chr8_-_36469117 | 0.26 |
ENSDART00000111240
|
mhc2dab
|
major histocompatibility complex class II DAB gene |
| chr19_-_18855717 | 0.26 |
ENSDART00000158192
|
ppt2
|
palmitoyl-protein thioesterase 2 |
| chr1_-_44940830 | 0.26 |
ENSDART00000097500
ENSDART00000134464 ENSDART00000137216 |
tmem176
|
transmembrane protein 176 |
| chr7_+_11197940 | 0.25 |
ENSDART00000081346
|
cemip
|
cell migration inducing protein, hyaluronan binding |
| chr23_+_28128453 | 0.25 |
ENSDART00000182618
|
c1galt1a
|
core 1 synthase, glycoprotein-N-acetylgalactosamine 3-beta-galactosyltransferase, 1a |
| chr1_+_58353661 | 0.25 |
ENSDART00000140074
|
si:dkey-222h21.2
|
si:dkey-222h21.2 |
| chr17_+_5768608 | 0.24 |
ENSDART00000157039
|
rp1l1a
|
retinitis pigmentosa 1-like 1a |
| chr8_-_29822527 | 0.23 |
ENSDART00000167487
|
slc20a2
|
solute carrier family 20 (phosphate transporter), member 2 |
| chr16_-_50157187 | 0.23 |
ENSDART00000155162
|
nek10
|
NIMA-related kinase 10 |
| chr24_-_25184553 | 0.23 |
ENSDART00000166917
|
plcxd2
|
phosphatidylinositol-specific phospholipase C, X domain containing 2 |
| chr8_+_45942431 | 0.22 |
ENSDART00000060955
|
nudt18
|
nudix (nucleoside diphosphate linked moiety X)-type motif 18 |
| chr22_+_28888781 | 0.22 |
ENSDART00000145629
|
pimr98
|
Pim proto-oncogene, serine/threonine kinase, related 98 |
| chr10_+_20070178 | 0.22 |
ENSDART00000027612
ENSDART00000145264 ENSDART00000172713 |
xpo7
|
exportin 7 |
| chr3_+_31953145 | 0.22 |
ENSDART00000148861
|
kcnc3a
|
potassium voltage-gated channel, Shaw-related subfamily, member 3a |
| chr5_+_69716458 | 0.22 |
ENSDART00000159594
|
mthfd2
|
methylenetetrahydrofolate dehydrogenase (NADP+ dependent) 2, methenyltetrahydrofolate cyclohydrolase |
| chr13_+_35746440 | 0.22 |
ENSDART00000187859
|
gpr75
|
G protein-coupled receptor 75 |
| chr10_-_20696518 | 0.21 |
ENSDART00000064581
|
kcnip3b
|
Kv channel interacting protein 3b, calsenilin |
| chr6_+_9677420 | 0.20 |
ENSDART00000064995
|
sumo1
|
small ubiquitin-like modifier 1 |
| chr21_-_20733615 | 0.20 |
ENSDART00000145544
|
si:ch211-22d5.2
|
si:ch211-22d5.2 |
| chr22_+_7486867 | 0.20 |
ENSDART00000034586
|
CELA1 (1 of many)
|
zgc:112302 |
| chr2_+_42072689 | 0.19 |
ENSDART00000134203
|
vcpip1
|
valosin containing protein (p97)/p47 complex interacting protein 1 |
| chr24_+_28448387 | 0.19 |
ENSDART00000154077
ENSDART00000018037 |
arhgap29a
|
Rho GTPase activating protein 29a |
| chr12_-_25201576 | 0.18 |
ENSDART00000077188
|
cox7a3
|
cytochrome c oxidase subunit VIIa polypeptide 3 |
| chr3_+_4346854 | 0.18 |
ENSDART00000004273
|
si:dkey-73p2.3
|
si:dkey-73p2.3 |
| chr13_-_23665580 | 0.18 |
ENSDART00000144282
|
map3k21
|
mitogen-activated protein kinase kinase kinase 21 |
| chr11_+_21910752 | 0.18 |
ENSDART00000114288
|
foxp4
|
forkhead box P4 |
| chr16_+_5184402 | 0.17 |
ENSDART00000156685
|
soga3a
|
SOGA family member 3a |
| chr6_-_40312091 | 0.17 |
ENSDART00000154878
|
col7a1
|
collagen, type VII, alpha 1 |
| chr11_-_6974022 | 0.17 |
ENSDART00000172851
|
COMP
|
si:ch211-43f4.1 |
| chr7_-_16237861 | 0.17 |
ENSDART00000173647
|
btr07
|
bloodthirsty-related gene family, member 7 |
| chr3_-_58582663 | 0.17 |
ENSDART00000180055
|
CABZ01038521.1
|
|
| chr5_+_65946222 | 0.17 |
ENSDART00000190969
ENSDART00000161578 |
mymk
|
myomaker, myoblast fusion factor |
| chr16_+_5901835 | 0.16 |
ENSDART00000060519
|
ulk4
|
unc-51 like kinase 4 |
| chr25_+_23591990 | 0.16 |
ENSDART00000151938
ENSDART00000089199 |
cpt1ab
|
carnitine palmitoyltransferase 1Ab (liver) |
| chr20_-_19864131 | 0.16 |
ENSDART00000057819
|
ptk2bb
|
protein tyrosine kinase 2 beta, b |
| chr18_+_3338228 | 0.16 |
ENSDART00000161520
|
gdpd4a
|
glycerophosphodiester phosphodiesterase domain containing 4a |
| chr18_-_41754020 | 0.16 |
ENSDART00000077898
|
grk7b
|
G protein-coupled receptor kinase 7b |
| chr19_+_30990815 | 0.15 |
ENSDART00000134645
|
sync
|
syncoilin, intermediate filament protein |
| chr18_+_19990412 | 0.15 |
ENSDART00000155054
ENSDART00000090310 |
pias1b
|
protein inhibitor of activated STAT, 1b |
| chr24_-_21689146 | 0.14 |
ENSDART00000105917
|
urad
|
ureidoimidazoline (2-oxo-4-hydroxy-4-carboxy-5-) decarboxylase |
| chr16_-_50181206 | 0.14 |
ENSDART00000148478
|
lim2.5
|
lens intrinsic membrane protein 2.5 |
| chr15_-_462783 | 0.14 |
ENSDART00000161049
|
CABZ01033205.1
|
Danio rerio uncharacterized LOC100002960 (LOC100002960), mRNA. |
| chr16_-_13281380 | 0.13 |
ENSDART00000103882
|
grin2db
|
glutamate receptor, ionotropic, N-methyl D-aspartate 2D, b |
| chr9_+_46644633 | 0.13 |
ENSDART00000160285
|
slc4a3
|
solute carrier family 4 (anion exchanger), member 3 |
| chr22_-_17474583 | 0.13 |
ENSDART00000148027
|
si:ch211-197g15.8
|
si:ch211-197g15.8 |
| chr8_-_13013123 | 0.13 |
ENSDART00000147802
|
dennd2da
|
DENN/MADD domain containing 2Da |
| chr15_+_44472356 | 0.13 |
ENSDART00000155763
ENSDART00000028011 |
msantd4
|
Myb/SANT-like DNA-binding domain containing 4 with coiled-coils |
| chr23_+_35684756 | 0.13 |
ENSDART00000132320
|
pmelb
|
premelanosome protein b |
| chr23_+_43954809 | 0.12 |
ENSDART00000164080
|
corin
|
corin, serine peptidase |
| chr7_-_41338923 | 0.12 |
ENSDART00000099138
|
ncf2
|
neutrophil cytosolic factor 2 |
| chr3_+_32142382 | 0.12 |
ENSDART00000133035
|
syt5a
|
synaptotagmin Va |
| chr6_+_27418541 | 0.12 |
ENSDART00000187410
|
crocc2
|
ciliary rootlet coiled-coil, rootletin family member 2 |
| chr12_+_34119439 | 0.11 |
ENSDART00000032821
|
cyth1b
|
cytohesin 1b |
| chr6_-_1514767 | 0.11 |
ENSDART00000067586
|
chchd6b
|
coiled-coil-helix-coiled-coil-helix domain containing 6b |
| chr11_+_12175162 | 0.11 |
ENSDART00000125446
|
si:ch211-156l18.7
|
si:ch211-156l18.7 |
| chr14_-_34771371 | 0.11 |
ENSDART00000160598
ENSDART00000150413 ENSDART00000168910 |
ablim3
|
actin binding LIM protein family, member 3 |
| chr5_+_1079423 | 0.11 |
ENSDART00000172231
|
si:zfos-128g4.2
|
si:zfos-128g4.2 |
| chr1_+_44941031 | 0.11 |
ENSDART00000141145
|
si:dkey-9i23.16
|
si:dkey-9i23.16 |
| chr11_+_11200550 | 0.11 |
ENSDART00000181339
ENSDART00000187116 |
myom2a
|
myomesin 2a |
| chr25_+_12494079 | 0.10 |
ENSDART00000163508
|
map1lc3b
|
microtubule-associated protein 1 light chain 3 beta |
| chr11_-_17713987 | 0.10 |
ENSDART00000090401
|
fam19a4b
|
family with sequence similarity 19 (chemokine (C-C motif)-like), member A4b |
| chr14_-_34276574 | 0.10 |
ENSDART00000021437
|
gria1a
|
glutamate receptor, ionotropic, AMPA 1a |
| chr13_+_29238850 | 0.10 |
ENSDART00000026000
|
myofl
|
myoferlin like |
| chr19_-_23369282 | 0.10 |
ENSDART00000147824
|
pimr90
|
Pim proto-oncogene, serine/threonine kinase, related 90 |
| chr20_-_37522569 | 0.10 |
ENSDART00000177011
ENSDART00000058502 |
hivep2a
|
human immunodeficiency virus type I enhancer binding protein 2a |
| chr21_+_29179887 | 0.10 |
ENSDART00000161941
|
si:ch211-57b15.1
|
si:ch211-57b15.1 |
| chr14_+_22297257 | 0.09 |
ENSDART00000113676
|
atp10b
|
ATPase phospholipid transporting 10B |
| chr21_-_22547496 | 0.09 |
ENSDART00000166835
ENSDART00000089030 |
myo5b
|
myosin VB |
| chr15_+_37331585 | 0.09 |
ENSDART00000170715
|
zgc:171592
|
zgc:171592 |
| chr8_+_10305400 | 0.09 |
ENSDART00000172400
|
pim1
|
Pim-1 proto-oncogene, serine/threonine kinase |
| chr22_-_17474781 | 0.09 |
ENSDART00000186817
|
si:ch211-197g15.8
|
si:ch211-197g15.8 |
| chr8_+_8735285 | 0.08 |
ENSDART00000165655
|
elk1
|
ELK1, member of ETS oncogene family |
| chr24_-_36337395 | 0.08 |
ENSDART00000154402
|
si:ch211-40k21.9
|
si:ch211-40k21.9 |
| chr15_-_31000275 | 0.08 |
ENSDART00000132493
ENSDART00000100183 |
lgals9l6
lgals9l5
|
lectin, galactoside-binding, soluble, 9 (galectin 9)-like 6 lectin, galactoside-binding, soluble, 9 (galectin 9)-like 5 |
| chr15_-_923981 | 0.08 |
ENSDART00000155285
|
si:dkey-77f5.14
|
si:dkey-77f5.14 |
| chr21_+_27370671 | 0.08 |
ENSDART00000009234
ENSDART00000142071 |
rbm14a
|
RNA binding motif protein 14a |
| chr17_+_20173882 | 0.08 |
ENSDART00000155379
|
si:ch211-248a14.8
|
si:ch211-248a14.8 |
| chr22_-_11136625 | 0.08 |
ENSDART00000016873
ENSDART00000125561 |
atp6ap2
|
ATPase H+ transporting accessory protein 2 |
| chr4_+_70013627 | 0.08 |
ENSDART00000180432
|
si:dkey-190j3.4
|
si:dkey-190j3.4 |
| chr14_-_3174570 | 0.08 |
ENSDART00000163585
|
csf1ra
|
colony stimulating factor 1 receptor, a |
| chr15_-_14552101 | 0.07 |
ENSDART00000171169
|
numbl
|
numb homolog (Drosophila)-like |
| chr6_-_609880 | 0.07 |
ENSDART00000149248
ENSDART00000148867 ENSDART00000149414 ENSDART00000148552 ENSDART00000148391 |
lgals2b
|
lectin, galactoside-binding, soluble, 2b |
| chr16_-_13730152 | 0.07 |
ENSDART00000138772
|
ttyh1
|
tweety family member 1 |
| chr5_+_1219187 | 0.07 |
ENSDART00000129490
|
gb:bc139872
|
expressed sequence BC139872 |
| chr15_-_19705707 | 0.07 |
ENSDART00000047643
|
sytl2b
|
synaptotagmin-like 2b |
| chr13_-_37200120 | 0.07 |
ENSDART00000135001
|
si:dkeyp-77c8.3
|
si:dkeyp-77c8.3 |
| chr23_-_45504991 | 0.07 |
ENSDART00000148761
|
col24a1
|
collagen type XXIV alpha 1 |
| chr4_+_20812900 | 0.06 |
ENSDART00000005847
|
nav3
|
neuron navigator 3 |
| chr20_+_7243879 | 0.06 |
ENSDART00000122298
|
bsnd
|
barttin CLCNK-type chloride channel accessory beta subunit |
| chr9_-_48184823 | 0.06 |
ENSDART00000180264
|
klhl23
|
kelch-like family member 23 |
| chr19_+_30990129 | 0.06 |
ENSDART00000052169
ENSDART00000193376 |
sync
|
syncoilin, intermediate filament protein |
| chr7_+_36539124 | 0.06 |
ENSDART00000173653
|
chd9
|
chromodomain helicase DNA binding protein 9 |
| chr21_-_32082130 | 0.06 |
ENSDART00000003978
ENSDART00000182050 |
mat2b
|
methionine adenosyltransferase II, beta |
| chr2_+_42112503 | 0.06 |
ENSDART00000136022
|
crha
|
corticotropin releasing hormone a |
| chr3_-_19899914 | 0.06 |
ENSDART00000134969
|
rnd2
|
Rho family GTPase 2 |
| chr11_+_24340166 | 0.06 |
ENSDART00000188719
|
rbm39a
|
RNA binding motif protein 39a |
| chr23_-_42810664 | 0.05 |
ENSDART00000102328
|
pfkfb2a
|
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 2a |
| chr20_+_9828769 | 0.05 |
ENSDART00000160480
ENSDART00000053844 ENSDART00000080691 |
stxbp6l
|
syntaxin binding protein 6 (amisyn), like |
| chr4_+_1530287 | 0.05 |
ENSDART00000067446
|
slc38a4
|
solute carrier family 38, member 4 |
| chr1_+_127250 | 0.05 |
ENSDART00000003463
|
f7i
|
coagulation factor VIIi |
| chr5_-_50748081 | 0.05 |
ENSDART00000113985
|
mctp1a
|
multiple C2 domains, transmembrane 1a |
| chr12_-_4370585 | 0.05 |
ENSDART00000129502
|
CA4 (1 of many)
|
si:ch211-173d10.4 |
| chr2_-_37862380 | 0.05 |
ENSDART00000186005
|
si:ch211-284o19.8
|
si:ch211-284o19.8 |
| chr5_+_28830388 | 0.05 |
ENSDART00000149150
|
serpina7
|
serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 7 |
| chr16_-_20312146 | 0.04 |
ENSDART00000134980
|
si:dkeyp-86h10.3
|
si:dkeyp-86h10.3 |
| chr8_+_53311965 | 0.04 |
ENSDART00000130104
|
gnb1a
|
guanine nucleotide binding protein (G protein), beta polypeptide 1a |
| chr22_-_11137268 | 0.04 |
ENSDART00000178882
|
atp6ap2
|
ATPase H+ transporting accessory protein 2 |
| chr17_-_465285 | 0.04 |
ENSDART00000168718
|
chrm5a
|
cholinergic receptor, muscarinic 5a |
| chr23_-_21515182 | 0.04 |
ENSDART00000142000
|
rnf207b
|
ring finger protein 207b |
| chr2_-_52020049 | 0.04 |
ENSDART00000128198
ENSDART00000135331 |
scn12aa
|
sodium channel, voltage gated, type XII, alpha a |
| chr13_-_37108310 | 0.04 |
ENSDART00000163845
|
syne2b
|
spectrin repeat containing, nuclear envelope 2b |
| chr14_-_2318590 | 0.04 |
ENSDART00000192735
|
pcdh2ab8
|
protocadherin 2 alpha b 8 |
| chr21_-_45186431 | 0.03 |
ENSDART00000193449
|
si:ch73-269m14.2
|
si:ch73-269m14.2 |
| chr10_+_22012389 | 0.03 |
ENSDART00000035188
|
kcnip1b
|
Kv channel interacting protein 1 b |
| chr19_-_1929310 | 0.03 |
ENSDART00000144113
|
si:ch211-149a19.3
|
si:ch211-149a19.3 |
| chr18_-_16791331 | 0.03 |
ENSDART00000148222
|
ampd3b
|
adenosine monophosphate deaminase 3b |
| chr17_+_14711765 | 0.02 |
ENSDART00000012889
|
cx28.6
|
connexin 28.6 |
| chr1_-_22512063 | 0.02 |
ENSDART00000031546
ENSDART00000190987 |
chrna6
|
cholinergic receptor, nicotinic, alpha 6 |
| chr5_-_69571572 | 0.02 |
ENSDART00000145966
|
mapkapk5
|
mitogen-activated protein kinase-activated protein kinase 5 |
| chr14_+_904850 | 0.02 |
ENSDART00000161847
|
si:ch73-208h1.1
|
si:ch73-208h1.1 |
| chr16_-_17175731 | 0.02 |
ENSDART00000183057
|
opn9
|
opsin 9 |
| chr7_-_1918971 | 0.01 |
ENSDART00000181759
|
CABZ01040626.1
|
|
| chr15_-_34322915 | 0.01 |
ENSDART00000190543
ENSDART00000019651 ENSDART00000193601 |
dgkb
|
diacylglycerol kinase, beta |
| chr7_+_53755054 | 0.01 |
ENSDART00000181629
|
neo1a
|
neogenin 1a |
| chr12_+_13405445 | 0.01 |
ENSDART00000089042
|
kcnh4b
|
potassium voltage-gated channel, subfamily H (eag-related), member 4b |
| chr5_+_16496273 | 0.00 |
ENSDART00000168969
ENSDART00000147442 |
htr7c
|
5-hydroxytryptamine (serotonin) receptor 7c |
| chr21_+_40092301 | 0.00 |
ENSDART00000145150
|
serpinf2a
|
serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), member 2a |
| chr7_-_8014186 | 0.00 |
ENSDART00000190012
|
si:cabz01030277.1
|
si:cabz01030277.1 |
| chr3_+_59117136 | 0.00 |
ENSDART00000182745
|
MGAT5B
|
mannosyl (alpha-1,6-)-glycoprotein beta-1,6-N-acetyl-glucosaminyltransferase, isozyme B |
Network of associatons between targets according to the STRING database.
First level regulatory network of nr1h4
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.6 | GO:0009447 | polyamine catabolic process(GO:0006598) putrescine catabolic process(GO:0009447) |
| 0.1 | 0.4 | GO:0043455 | regulation of secondary metabolic process(GO:0043455) |
| 0.1 | 0.5 | GO:0015990 | energy coupled proton transmembrane transport, against electrochemical gradient(GO:0015988) electron transport coupled proton transport(GO:0015990) |
| 0.1 | 0.3 | GO:0035971 | peptidyl-histidine dephosphorylation(GO:0035971) |
| 0.1 | 0.2 | GO:0009191 | ribonucleoside diphosphate catabolic process(GO:0009191) |
| 0.1 | 0.3 | GO:0019287 | isopentenyl diphosphate biosynthetic process, mevalonate pathway(GO:0019287) |
| 0.1 | 0.3 | GO:0046654 | tetrahydrofolate biosynthetic process(GO:0046654) |
| 0.1 | 0.3 | GO:0016267 | O-glycan processing, core 1(GO:0016267) |
| 0.1 | 0.2 | GO:0071276 | cellular response to cadmium ion(GO:0071276) |
| 0.1 | 0.7 | GO:0051481 | negative regulation of cytosolic calcium ion concentration(GO:0051481) negative regulation of synaptic transmission, glutamatergic(GO:0051967) |
| 0.0 | 0.2 | GO:0048308 | organelle inheritance(GO:0048308) Golgi inheritance(GO:0048313) |
| 0.0 | 0.4 | GO:0006878 | cellular copper ion homeostasis(GO:0006878) |
| 0.0 | 0.1 | GO:0019628 | urate catabolic process(GO:0019628) urate metabolic process(GO:0046415) |
| 0.0 | 0.2 | GO:0014743 | regulation of muscle hypertrophy(GO:0014743) |
| 0.0 | 0.7 | GO:0006744 | ubiquinone metabolic process(GO:0006743) ubiquinone biosynthetic process(GO:0006744) quinone biosynthetic process(GO:1901663) |
| 0.0 | 0.8 | GO:0021884 | forebrain neuron development(GO:0021884) |
| 0.0 | 0.3 | GO:0060754 | mast cell chemotaxis(GO:0002551) regulation of mast cell chemotaxis(GO:0060753) positive regulation of mast cell chemotaxis(GO:0060754) mast cell migration(GO:0097531) |
| 0.0 | 0.1 | GO:0045730 | respiratory burst(GO:0045730) |
| 0.0 | 0.3 | GO:0035721 | intraciliary retrograde transport(GO:0035721) |
| 0.0 | 0.3 | GO:0098900 | regulation of action potential(GO:0098900) |
| 0.0 | 0.5 | GO:0018095 | protein polyglutamylation(GO:0018095) |
| 0.0 | 0.1 | GO:0003245 | growth involved in heart morphogenesis(GO:0003241) cardiac chamber ballooning(GO:0003242) cardiac muscle tissue growth involved in heart morphogenesis(GO:0003245) |
| 0.0 | 0.2 | GO:0006611 | protein export from nucleus(GO:0006611) |
| 0.0 | 0.5 | GO:0000027 | ribosomal large subunit assembly(GO:0000027) |
| 0.0 | 0.3 | GO:0030011 | maintenance of cell polarity(GO:0030011) |
| 0.0 | 0.1 | GO:0010758 | regulation of macrophage chemotaxis(GO:0010758) positive regulation of macrophage chemotaxis(GO:0010759) regulation of odontogenesis(GO:0042481) mononuclear cell migration(GO:0071674) regulation of mononuclear cell migration(GO:0071675) |
| 0.0 | 0.3 | GO:0030323 | respiratory tube development(GO:0030323) lung development(GO:0030324) |
| 0.0 | 0.3 | GO:0046626 | regulation of insulin receptor signaling pathway(GO:0046626) |
| 0.0 | 0.1 | GO:2000562 | negative regulation of interferon-gamma production(GO:0032689) CD4-positive, alpha-beta T cell proliferation(GO:0035739) regulation of CD4-positive, alpha-beta T cell proliferation(GO:2000561) negative regulation of CD4-positive, alpha-beta T cell proliferation(GO:2000562) |
| 0.0 | 0.5 | GO:0006378 | mRNA polyadenylation(GO:0006378) RNA polyadenylation(GO:0043631) |
| 0.0 | 0.2 | GO:0031998 | regulation of fatty acid beta-oxidation(GO:0031998) |
| 0.0 | 0.5 | GO:0006910 | phagocytosis, recognition(GO:0006910) |
| 0.0 | 0.2 | GO:0035999 | tetrahydrofolate interconversion(GO:0035999) |
| 0.0 | 0.2 | GO:0007172 | signal complex assembly(GO:0007172) |
| 0.0 | 0.1 | GO:0038063 | collagen-activated tyrosine kinase receptor signaling pathway(GO:0038063) collagen-activated signaling pathway(GO:0038065) |
| 0.0 | 0.2 | GO:0000463 | maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000463) |
| 0.0 | 0.2 | GO:0035435 | phosphate ion transmembrane transport(GO:0035435) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | GO:0030991 | intraciliary transport particle A(GO:0030991) |
| 0.0 | 0.3 | GO:0005662 | DNA replication factor A complex(GO:0005662) |
| 0.0 | 0.1 | GO:0005790 | smooth endoplasmic reticulum(GO:0005790) |
| 0.0 | 0.3 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 0.0 | 0.5 | GO:0012510 | trans-Golgi network transport vesicle membrane(GO:0012510) clathrin coat of trans-Golgi network vesicle(GO:0030130) |
| 0.0 | 0.1 | GO:0005592 | collagen type XI trimer(GO:0005592) |
| 0.0 | 0.3 | GO:0044224 | juxtaparanode region of axon(GO:0044224) |
| 0.0 | 0.4 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.0 | 0.7 | GO:0098978 | glutamatergic synapse(GO:0098978) |
| 0.0 | 0.1 | GO:0035253 | ciliary rootlet(GO:0035253) |
| 0.0 | 0.5 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
| 0.0 | 0.1 | GO:0048269 | methionine adenosyltransferase complex(GO:0048269) |
| 0.0 | 0.1 | GO:0043020 | NADPH oxidase complex(GO:0043020) |
| 0.0 | 0.5 | GO:0030964 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.0 | 0.2 | GO:0032809 | neuronal cell body membrane(GO:0032809) cell body membrane(GO:0044298) |
| 0.0 | 0.5 | GO:0005762 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.0 | 0.1 | GO:0061617 | MICOS complex(GO:0061617) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0004652 | polynucleotide adenylyltransferase activity(GO:0004652) |
| 0.1 | 0.4 | GO:0034416 | bisphosphoglycerate phosphatase activity(GO:0034416) |
| 0.1 | 0.3 | GO:0050649 | testosterone 6-beta-hydroxylase activity(GO:0050649) |
| 0.1 | 0.5 | GO:0008097 | 5S rRNA binding(GO:0008097) |
| 0.1 | 0.6 | GO:0019808 | polyamine binding(GO:0019808) spermidine binding(GO:0019809) |
| 0.1 | 0.3 | GO:0101006 | protein histidine phosphatase activity(GO:0101006) |
| 0.1 | 0.3 | GO:0043183 | vascular endothelial growth factor receptor 1 binding(GO:0043183) |
| 0.1 | 0.7 | GO:0048039 | ubiquinone binding(GO:0048039) |
| 0.1 | 0.3 | GO:0016263 | glycoprotein-N-acetylgalactosamine 3-beta-galactosyltransferase activity(GO:0016263) beta-1,3-galactosyltransferase activity(GO:0048531) |
| 0.1 | 0.7 | GO:0001591 | dopamine neurotransmitter receptor activity, coupled via Gi/Go(GO:0001591) |
| 0.0 | 0.5 | GO:0048038 | quinone binding(GO:0048038) |
| 0.0 | 0.5 | GO:0019238 | cyclohydrolase activity(GO:0019238) |
| 0.0 | 0.3 | GO:0005159 | insulin-like growth factor receptor binding(GO:0005159) |
| 0.0 | 0.2 | GO:0050254 | rhodopsin kinase activity(GO:0050254) |
| 0.0 | 0.4 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.0 | 0.2 | GO:0004095 | carnitine O-palmitoyltransferase activity(GO:0004095) O-palmitoyltransferase activity(GO:0016416) |
| 0.0 | 0.2 | GO:0005049 | nuclear export signal receptor activity(GO:0005049) |
| 0.0 | 0.2 | GO:0004706 | JUN kinase kinase kinase activity(GO:0004706) |
| 0.0 | 0.5 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.0 | 0.2 | GO:0005315 | inorganic phosphate transmembrane transporter activity(GO:0005315) |
| 0.0 | 0.2 | GO:0016936 | galactoside binding(GO:0016936) |
| 0.0 | 0.1 | GO:0016176 | superoxide-generating NADPH oxidase activator activity(GO:0016176) |
| 0.0 | 0.3 | GO:0004033 | aldo-keto reductase (NADP) activity(GO:0004033) |
| 0.0 | 0.4 | GO:0015035 | protein disulfide oxidoreductase activity(GO:0015035) |
| 0.0 | 0.2 | GO:0061665 | SUMO ligase activity(GO:0061665) |
| 0.0 | 0.3 | GO:0005104 | fibroblast growth factor receptor binding(GO:0005104) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.0 | 0.2 | PID RANBP2 PATHWAY | Sumoylation by RanBP2 regulates transcriptional repression |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | REACTOME RECYCLING OF BILE ACIDS AND SALTS | Genes involved in Recycling of bile acids and salts |
| 0.0 | 0.6 | REACTOME ACTIVATION OF RAC | Genes involved in Activation of Rac |
| 0.0 | 0.3 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |
| 0.0 | 0.2 | REACTOME REGULATION OF IFNG SIGNALING | Genes involved in Regulation of IFNG signaling |
| 0.0 | 0.3 | REACTOME REMOVAL OF THE FLAP INTERMEDIATE FROM THE C STRAND | Genes involved in Removal of the Flap Intermediate from the C-strand |
| 0.0 | 0.5 | REACTOME PROCESSING OF INTRONLESS PRE MRNAS | Genes involved in Processing of Intronless Pre-mRNAs |
| 0.0 | 0.3 | REACTOME FANCONI ANEMIA PATHWAY | Genes involved in Fanconi Anemia pathway |
| 0.0 | 0.1 | REACTOME DCC MEDIATED ATTRACTIVE SIGNALING | Genes involved in DCC mediated attractive signaling |