Project
PRJNA438478: RNAseq of wild type zebrafish germline
Navigation
Downloads
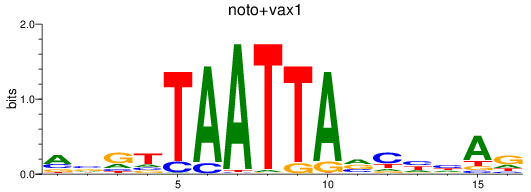
Results for noto+vax1
Z-value: 0.37
Transcription factors associated with noto+vax1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
noto
|
ENSDARG00000021201 | notochord homeobox |
|
vax1
|
ENSDARG00000021916 | ventral anterior homeobox 1 |
|
vax1
|
ENSDARG00000114230 | ventral anterior homeobox 1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| vax1 | dr11_v1_chr17_-_21441464_21441464 | -0.89 | 7.1e-07 | Click! |
| noto | dr11_v1_chr13_+_14976108_14976108 | -0.12 | 6.2e-01 | Click! |
Activity profile of noto+vax1 motif
Sorted Z-values of noto+vax1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr22_-_21897203 | 0.60 |
ENSDART00000158501
ENSDART00000105566 ENSDART00000136795 |
gna11a
|
guanine nucleotide binding protein (G protein), alpha 11a (Gq class) |
| chr13_-_35808904 | 0.55 |
ENSDART00000171667
|
map3k4
|
mitogen-activated protein kinase kinase kinase 4 |
| chr12_+_47446158 | 0.51 |
ENSDART00000152857
|
fmn2b
|
formin 2b |
| chr20_+_29209767 | 0.49 |
ENSDART00000141252
|
katnbl1
|
katanin p80 subunit B-like 1 |
| chr20_+_29209615 | 0.46 |
ENSDART00000062350
|
katnbl1
|
katanin p80 subunit B-like 1 |
| chr20_+_29209926 | 0.46 |
ENSDART00000152949
ENSDART00000153016 |
katnbl1
|
katanin p80 subunit B-like 1 |
| chr19_-_2861444 | 0.46 |
ENSDART00000169053
|
clec3bb
|
C-type lectin domain family 3, member Bb |
| chr10_+_16036246 | 0.45 |
ENSDART00000141586
ENSDART00000135868 ENSDART00000065037 ENSDART00000124502 |
lmnb1
|
lamin B1 |
| chr15_-_43284021 | 0.43 |
ENSDART00000041677
|
serpine2
|
serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 2 |
| chr6_+_32382743 | 0.43 |
ENSDART00000190009
|
dock7
|
dedicator of cytokinesis 7 |
| chr2_-_21819421 | 0.39 |
ENSDART00000121586
|
chd7
|
chromodomain helicase DNA binding protein 7 |
| chr11_+_11303458 | 0.39 |
ENSDART00000162486
ENSDART00000160703 |
si:dkey-23f9.4
|
si:dkey-23f9.4 |
| chr17_+_21902058 | 0.39 |
ENSDART00000142178
|
plekha1a
|
pleckstrin homology domain containing, family A (phosphoinositide binding specific) member 1a |
| chr10_+_28160265 | 0.38 |
ENSDART00000022484
|
rnft1
|
ring finger protein, transmembrane 1 |
| chr5_+_41477954 | 0.38 |
ENSDART00000185871
|
pias2
|
protein inhibitor of activated STAT, 2 |
| chr9_-_54248182 | 0.37 |
ENSDART00000129540
|
larsa
|
leucyl-tRNA synthetase a |
| chr5_+_20257225 | 0.35 |
ENSDART00000127919
|
ssh1a
|
slingshot protein phosphatase 1a |
| chr14_+_51056605 | 0.35 |
ENSDART00000159639
|
CABZ01078593.1
|
|
| chr1_+_513986 | 0.34 |
ENSDART00000109083
ENSDART00000081945 |
txnl4b
|
thioredoxin-like 4B |
| chr19_-_20430892 | 0.34 |
ENSDART00000111409
|
tbc1d5
|
TBC1 domain family, member 5 |
| chr8_-_25034411 | 0.34 |
ENSDART00000135973
|
nfyal
|
nuclear transcription factor Y, alpha, like |
| chr10_+_7593185 | 0.34 |
ENSDART00000162617
ENSDART00000162590 ENSDART00000171744 |
ppp2cb
|
protein phosphatase 2, catalytic subunit, beta isozyme |
| chr5_+_41477526 | 0.33 |
ENSDART00000153567
|
pias2
|
protein inhibitor of activated STAT, 2 |
| chr16_-_22544047 | 0.33 |
ENSDART00000131657
|
cgna
|
cingulin a |
| chr8_-_39822917 | 0.32 |
ENSDART00000067843
|
zgc:162025
|
zgc:162025 |
| chr16_+_13818743 | 0.31 |
ENSDART00000090191
|
flcn
|
folliculin |
| chr5_-_15283509 | 0.31 |
ENSDART00000052712
|
gnb1l
|
guanine nucleotide binding protein (G protein), beta polypeptide 1-like |
| chr16_+_13818500 | 0.31 |
ENSDART00000135245
|
flcn
|
folliculin |
| chr8_-_50888806 | 0.31 |
ENSDART00000053750
|
acsl2
|
acyl-CoA synthetase long chain family member 2 |
| chr24_+_19415124 | 0.31 |
ENSDART00000186931
|
sulf1
|
sulfatase 1 |
| chr21_-_26490186 | 0.29 |
ENSDART00000009889
|
zgc:110540
|
zgc:110540 |
| chr6_-_19271210 | 0.28 |
ENSDART00000163628
ENSDART00000159124 |
zgc:174863
|
zgc:174863 |
| chr21_+_8427059 | 0.27 |
ENSDART00000143151
|
dennd1a
|
DENN/MADD domain containing 1A |
| chr17_+_16046314 | 0.26 |
ENSDART00000154554
ENSDART00000154338 ENSDART00000155336 |
si:ch73-204p21.2
|
si:ch73-204p21.2 |
| chr9_+_30421489 | 0.25 |
ENSDART00000145025
ENSDART00000132058 |
zgc:113314
|
zgc:113314 |
| chr13_-_24257631 | 0.25 |
ENSDART00000146524
|
urb2
|
URB2 ribosome biogenesis 2 homolog (S. cerevisiae) |
| chr4_+_9011448 | 0.24 |
ENSDART00000192357
|
samm50l
|
sorting and assembly machinery component 50 homolog, like |
| chr9_-_14683574 | 0.23 |
ENSDART00000144022
|
pard3bb
|
par-3 family cell polarity regulator beta b |
| chr19_-_18313303 | 0.23 |
ENSDART00000164644
ENSDART00000167480 ENSDART00000163104 |
si:dkey-208k4.2
|
si:dkey-208k4.2 |
| chr6_-_40922971 | 0.22 |
ENSDART00000155363
|
sfi1
|
SFI1 centrin binding protein |
| chr19_+_32401278 | 0.22 |
ENSDART00000184353
|
atxn1a
|
ataxin 1a |
| chr21_-_42876565 | 0.21 |
ENSDART00000126480
|
zmp:0000001268
|
zmp:0000001268 |
| chr19_-_6988837 | 0.21 |
ENSDART00000145741
ENSDART00000167640 |
znf384l
|
zinc finger protein 384 like |
| chr14_-_14640401 | 0.21 |
ENSDART00000168027
ENSDART00000167521 |
znf185
|
zinc finger protein 185 with LIM domain |
| chr12_+_48803098 | 0.18 |
ENSDART00000074768
|
ppifb
|
peptidylprolyl isomerase Fb |
| chr16_-_7828838 | 0.18 |
ENSDART00000191434
ENSDART00000108653 |
tcaim
|
T cell activation inhibitor, mitochondrial |
| chr25_+_28825657 | 0.17 |
ENSDART00000153625
|
nfybb
|
nuclear transcription factor Y, beta b |
| chr20_-_1635922 | 0.16 |
ENSDART00000181502
|
CR846082.1
|
|
| chr6_-_57539141 | 0.16 |
ENSDART00000156967
|
itcha
|
itchy E3 ubiquitin protein ligase a |
| chr5_-_66397688 | 0.14 |
ENSDART00000161483
|
hip1rb
|
huntingtin interacting protein 1 related b |
| chr11_+_43043171 | 0.13 |
ENSDART00000180344
|
CABZ01092982.1
|
|
| chr15_+_31899312 | 0.13 |
ENSDART00000155315
|
frya
|
furry homolog a (Drosophila) |
| chr15_+_31701716 | 0.12 |
ENSDART00000113370
|
b3glcta
|
beta 3-glucosyltransferase a |
| chr4_+_21129752 | 0.12 |
ENSDART00000169764
|
syt1a
|
synaptotagmin Ia |
| chr1_-_51719110 | 0.11 |
ENSDART00000190574
|
rnaseh2a
|
ribonuclease H2, subunit A |
| chr19_+_2631565 | 0.11 |
ENSDART00000171487
|
fam126a
|
family with sequence similarity 126, member A |
| chr7_-_12464412 | 0.09 |
ENSDART00000178723
|
adamtsl3
|
ADAMTS-like 3 |
| chr6_-_30954268 | 0.09 |
ENSDART00000154523
|
pde4ba
|
phosphodiesterase 4B, cAMP-specific a |
| chr2_-_26720854 | 0.08 |
ENSDART00000148110
|
si:dkey-181m9.8
|
si:dkey-181m9.8 |
| chr18_-_2727764 | 0.08 |
ENSDART00000160841
|
ARHGEF17
|
si:ch211-248g20.5 |
| chr1_-_31534089 | 0.07 |
ENSDART00000007770
|
lbx1b
|
ladybird homeobox 1b |
| chr22_-_392224 | 0.06 |
ENSDART00000124720
|
CABZ01072989.1
|
|
| chr4_-_77506362 | 0.06 |
ENSDART00000174387
ENSDART00000181181 |
CABZ01087415.1
|
|
| chr10_-_2524917 | 0.05 |
ENSDART00000188642
|
CU856539.1
|
|
| chr13_+_35339182 | 0.04 |
ENSDART00000019323
|
jag1b
|
jagged 1b |
| chr9_-_22310919 | 0.04 |
ENSDART00000108719
|
crygm2d10
|
crystallin, gamma M2d10 |
| chr18_+_9362455 | 0.03 |
ENSDART00000187025
|
sema3ab
|
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3Ab |
| chr22_+_19218733 | 0.02 |
ENSDART00000183212
ENSDART00000133595 |
si:dkey-21e2.7
|
si:dkey-21e2.7 |
| chr4_+_73085993 | 0.01 |
ENSDART00000165749
|
si:ch73-170d6.2
|
si:ch73-170d6.2 |
| chr19_-_7690975 | 0.01 |
ENSDART00000151384
|
si:dkey-204a24.10
|
si:dkey-204a24.10 |
| chr2_-_39558643 | 0.01 |
ENSDART00000139860
ENSDART00000145231 ENSDART00000141721 |
cbln7
|
cerebellin 7 |
| chr19_+_9295244 | 0.01 |
ENSDART00000132255
ENSDART00000144299 |
si:ch73-15n24.1
|
si:ch73-15n24.1 |
| chr5_+_30635309 | 0.01 |
ENSDART00000183769
|
abcg4a
|
ATP-binding cassette, sub-family G (WHITE), member 4a |
| chr15_+_5360407 | 0.00 |
ENSDART00000110420
|
or112-1
|
odorant receptor, family A, subfamily 112, member 1 |
| chr3_-_50443607 | 0.00 |
ENSDART00000074036
|
rcvrna
|
recoverin a |
Network of associatons between targets according to the STRING database.
First level regulatory network of noto+vax1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | GO:0030511 | positive regulation of transforming growth factor beta receptor signaling pathway(GO:0030511) positive regulation of cellular response to transforming growth factor beta stimulus(GO:1903846) |
| 0.1 | 0.2 | GO:0007008 | outer mitochondrial membrane organization(GO:0007008) protein import into mitochondrial outer membrane(GO:0045040) |
| 0.1 | 0.4 | GO:1904292 | regulation of ERAD pathway(GO:1904292) |
| 0.1 | 0.2 | GO:0051660 | establishment of centrosome localization(GO:0051660) |
| 0.1 | 0.3 | GO:0097065 | anterior head development(GO:0097065) |
| 0.0 | 0.3 | GO:2000290 | smoothened signaling pathway involved in ventral spinal cord patterning(GO:0021910) regulation of myotome development(GO:2000290) |
| 0.0 | 0.4 | GO:0009303 | rRNA transcription(GO:0009303) swim bladder maturation(GO:0048796) swim bladder inflation(GO:0048798) |
| 0.0 | 0.7 | GO:0016925 | protein sumoylation(GO:0016925) |
| 0.0 | 0.1 | GO:0043137 | DNA replication, removal of RNA primer(GO:0043137) |
| 0.0 | 0.2 | GO:0070301 | cellular response to hydrogen peroxide(GO:0070301) |
| 0.0 | 0.3 | GO:0060158 | phospholipase C-activating dopamine receptor signaling pathway(GO:0060158) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | GO:0016602 | CCAAT-binding factor complex(GO:0016602) |
| 0.0 | 0.2 | GO:0001401 | mitochondrial sorting and assembly machinery complex(GO:0001401) |
| 0.0 | 0.1 | GO:0032299 | ribonuclease H2 complex(GO:0032299) |
| 0.0 | 0.1 | GO:0042584 | chromaffin granule membrane(GO:0042584) |
| 0.0 | 0.3 | GO:0005682 | U5 snRNP(GO:0005682) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | GO:0031826 | type 2A serotonin receptor binding(GO:0031826) |
| 0.1 | 0.7 | GO:0061665 | SUMO ligase activity(GO:0061665) |
| 0.0 | 0.3 | GO:0019136 | deoxynucleoside kinase activity(GO:0019136) |
| 0.0 | 0.3 | GO:0047676 | arachidonate-CoA ligase activity(GO:0047676) |
| 0.0 | 0.3 | GO:0008449 | N-acetylglucosamine-6-sulfatase activity(GO:0008449) |
| 0.0 | 0.1 | GO:0004523 | RNA-DNA hybrid ribonuclease activity(GO:0004523) |
| 0.0 | 0.4 | GO:0030165 | PDZ domain binding(GO:0030165) |
| 0.0 | 0.1 | GO:0043325 | phosphatidylinositol-3,4-bisphosphate binding(GO:0043325) |
| 0.0 | 0.2 | GO:0016018 | cyclosporin A binding(GO:0016018) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.7 | PID IL12 STAT4 PATHWAY | IL12 signaling mediated by STAT4 |
| 0.0 | 0.5 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
| 0.0 | 0.4 | PID WNT NONCANONICAL PATHWAY | Noncanonical Wnt signaling pathway |
| 0.0 | 0.5 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | REACTOME ERKS ARE INACTIVATED | Genes involved in ERKs are inactivated |
| 0.0 | 0.5 | REACTOME APOPTOTIC CLEAVAGE OF CELLULAR PROTEINS | Genes involved in Apoptotic cleavage of cellular proteins |