Project
PRJNA438478: RNAseq of wild type zebrafish germline
Navigation
Downloads
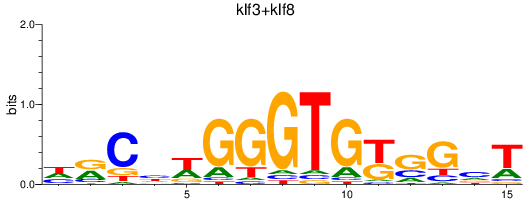
Results for klf3+klf8
Z-value: 0.46
Transcription factors associated with klf3+klf8
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
klf3
|
ENSDARG00000015495 | Kruppel-like factor 3 (basic) |
|
klf8
|
ENSDARG00000060661 | Kruppel-like factor 8 |
|
klf3
|
ENSDARG00000114143 | Kruppel-like factor 3 (basic) |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| klf8 | dr11_v1_chr21_+_38073210_38073216 | -0.94 | 5.2e-09 | Click! |
| klf3 | dr11_v1_chr1_-_18615063_18615063 | -0.37 | 1.3e-01 | Click! |
Activity profile of klf3+klf8 motif
Sorted Z-values of klf3+klf8 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr6_+_120181 | 1.18 |
ENSDART00000151209
ENSDART00000185930 |
cdkn2d
|
cyclin-dependent kinase inhibitor 2D (p19, inhibits CDK4) |
| chr23_-_45049044 | 1.15 |
ENSDART00000067630
|
SH3GL2 (1 of many)
|
SH3 domain containing GRB2 like 2, endophilin A1 |
| chr4_-_193762 | 1.10 |
ENSDART00000169667
|
ptpro
|
protein tyrosine phosphatase, receptor type, O |
| chr16_+_23978978 | 1.03 |
ENSDART00000058964
ENSDART00000135084 |
apoa2
|
apolipoprotein A-II |
| chr20_+_54356272 | 1.02 |
ENSDART00000145735
|
znf410
|
zinc finger protein 410 |
| chr4_-_191736 | 0.86 |
ENSDART00000169187
ENSDART00000192054 |
ptpro
|
protein tyrosine phosphatase, receptor type, O |
| chr3_+_55093581 | 0.84 |
ENSDART00000101709
|
hbaa2
|
hemoglobin, alpha adult 2 |
| chr5_-_24174956 | 0.77 |
ENSDART00000137631
|
si:ch211-137i24.10
|
si:ch211-137i24.10 |
| chr21_-_280769 | 0.76 |
ENSDART00000157753
|
plgrkt
|
plasminogen receptor, C-terminal lysine transmembrane protein |
| chr18_+_5549672 | 0.72 |
ENSDART00000184970
|
nnt2
|
nicotinamide nucleotide transhydrogenase 2 |
| chr19_+_233143 | 0.69 |
ENSDART00000175273
|
syngap1a
|
synaptic Ras GTPase activating protein 1a |
| chr3_-_1190132 | 0.64 |
ENSDART00000149709
|
smdt1a
|
single-pass membrane protein with aspartate-rich tail 1a |
| chr4_+_37992753 | 0.64 |
ENSDART00000193890
|
CR759843.2
|
|
| chr16_-_23471387 | 0.63 |
ENSDART00000180899
|
CR533578.1
|
|
| chr23_-_45439903 | 0.63 |
ENSDART00000170729
|
NPNT (1 of many)
|
nephronectin |
| chr1_-_49434798 | 0.56 |
ENSDART00000150842
ENSDART00000150789 |
ATP5MD
|
si:dkeyp-80c12.10 |
| chr19_+_19786117 | 0.49 |
ENSDART00000167757
ENSDART00000163546 |
hoxa1a
|
homeobox A1a |
| chr7_-_19600181 | 0.48 |
ENSDART00000100757
|
oxa1l
|
oxidase (cytochrome c) assembly 1-like |
| chr1_-_44940830 | 0.48 |
ENSDART00000097500
ENSDART00000134464 ENSDART00000137216 |
tmem176
|
transmembrane protein 176 |
| chr9_-_56459660 | 0.46 |
ENSDART00000162771
|
ubxn4
|
UBX domain protein 4 |
| chr15_-_47822597 | 0.44 |
ENSDART00000193236
ENSDART00000161391 |
CZQB01095947.1
|
|
| chr11_+_37278457 | 0.43 |
ENSDART00000188946
|
creld1a
|
cysteine-rich with EGF-like domains 1a |
| chr3_-_32927516 | 0.42 |
ENSDART00000140117
|
aoc2
|
amine oxidase, copper containing 2 |
| chr8_-_34762163 | 0.42 |
ENSDART00000114080
|
setd1bb
|
SET domain containing 1B, b |
| chr18_+_54354 | 0.41 |
ENSDART00000097163
|
zgc:158482
|
zgc:158482 |
| chr2_-_30135446 | 0.40 |
ENSDART00000141906
|
trpa1a
|
transient receptor potential cation channel, subfamily A, member 1a |
| chr25_+_37348730 | 0.40 |
ENSDART00000156639
|
pamr1
|
peptidase domain containing associated with muscle regeneration 1 |
| chr13_+_681628 | 0.38 |
ENSDART00000016604
|
abcc2
|
ATP-binding cassette, sub-family C (CFTR/MRP), member 2 |
| chr6_+_46258866 | 0.38 |
ENSDART00000134584
|
zgc:162324
|
zgc:162324 |
| chr20_+_50052627 | 0.38 |
ENSDART00000188799
|
cpsf2
|
cleavage and polyadenylation specific factor 2 |
| chr21_-_29197700 | 0.37 |
ENSDART00000172614
|
si:ch211-57b15.2
|
si:ch211-57b15.2 |
| chr18_-_45617146 | 0.37 |
ENSDART00000146543
|
wt1b
|
wilms tumor 1b |
| chr18_-_5103931 | 0.36 |
ENSDART00000188091
|
pdcd10a
|
programmed cell death 10a |
| chr14_-_48260744 | 0.34 |
ENSDART00000183955
|
rxfp1
|
relaxin/insulin-like family peptide receptor 1 |
| chr11_-_8269227 | 0.31 |
ENSDART00000127202
ENSDART00000158079 ENSDART00000158748 |
si:cabz01021067.1
pimr202
|
si:cabz01021067.1 Pim proto-oncogene, serine/threonine kinase, related 202 |
| chr12_+_48851381 | 0.31 |
ENSDART00000187241
ENSDART00000187796 |
dlg5b.1
|
discs, large homolog 5b (Drosophila), tandem duplicate 1 |
| chr7_-_6351021 | 0.30 |
ENSDART00000159542
|
zgc:112234
|
zgc:112234 |
| chr20_-_54377933 | 0.29 |
ENSDART00000182664
|
entpd5b
|
ectonucleoside triphosphate diphosphohydrolase 5b |
| chr7_-_67317119 | 0.29 |
ENSDART00000183915
|
psmd7
|
proteasome 26S subunit, non-ATPase 7 |
| chr18_+_5618368 | 0.28 |
ENSDART00000159945
|
ulk3
|
unc-51 like kinase 3 |
| chr21_+_27448856 | 0.28 |
ENSDART00000100784
|
cfbl
|
complement factor b-like |
| chr22_-_30948457 | 0.28 |
ENSDART00000139887
|
ssuh2.1
|
ssu-2 homolog, tandem duplicate 1 |
| chr12_+_34763795 | 0.28 |
ENSDART00000153097
|
slc38a10
|
solute carrier family 38, member 10 |
| chr19_+_56351 | 0.28 |
ENSDART00000168334
|
col14a1b
|
collagen, type XIV, alpha 1b |
| chr11_+_45461419 | 0.27 |
ENSDART00000173424
|
sos1
|
son of sevenless homolog 1 (Drosophila) |
| chr3_-_15444396 | 0.24 |
ENSDART00000104361
|
si:dkey-56d12.4
|
si:dkey-56d12.4 |
| chr14_-_21218891 | 0.24 |
ENSDART00000158294
|
ppp2r2cb
|
protein phosphatase 2, regulatory subunit B, gamma b |
| chr7_-_67316903 | 0.24 |
ENSDART00000159408
|
psmd7
|
proteasome 26S subunit, non-ATPase 7 |
| chr10_+_466926 | 0.22 |
ENSDART00000145220
|
arvcfa
|
ARVCF, delta catenin family member a |
| chr19_+_164013 | 0.22 |
ENSDART00000167379
|
carmil1
|
capping protein regulator and myosin 1 linker 1 |
| chr1_-_59571758 | 0.22 |
ENSDART00000193546
ENSDART00000167087 |
wfikkn1
|
WAP, follistatin/kazal, immunoglobulin, kunitz and netrin domain containing 1 |
| chr1_-_625875 | 0.22 |
ENSDART00000167331
|
appa
|
amyloid beta (A4) precursor protein a |
| chr1_+_36194761 | 0.21 |
ENSDART00000053773
ENSDART00000147458 |
lsm6
|
LSM6 homolog, U6 small nuclear RNA and mRNA degradation associated |
| chr4_+_53976731 | 0.21 |
ENSDART00000165813
|
si:ch211-249c2.1
|
si:ch211-249c2.1 |
| chr24_-_40668208 | 0.21 |
ENSDART00000171543
|
smyhc1
|
slow myosin heavy chain 1 |
| chr2_+_10007113 | 0.20 |
ENSDART00000155213
|
slc35a3b
|
solute carrier family 35 (UDP-N-acetylglucosamine (UDP-GlcNAc) transporter), member A3b |
| chr2_-_16217344 | 0.20 |
ENSDART00000152031
|
arhgef4
|
Rho guanine nucleotide exchange factor (GEF) 4 |
| chr22_+_1440702 | 0.19 |
ENSDART00000165677
|
si:dkeyp-53d3.3
|
si:dkeyp-53d3.3 |
| chr25_+_23489979 | 0.19 |
ENSDART00000158200
|
CU024870.1
|
|
| chr1_+_44941031 | 0.19 |
ENSDART00000141145
|
si:dkey-9i23.16
|
si:dkey-9i23.16 |
| chr25_-_28587896 | 0.19 |
ENSDART00000111908
|
si:dkeyp-67e1.6
|
si:dkeyp-67e1.6 |
| chr22_+_1006573 | 0.17 |
ENSDART00000123458
|
pparda
|
peroxisome proliferator-activated receptor delta a |
| chr20_-_39391833 | 0.16 |
ENSDART00000135149
|
si:dkey-217m5.8
|
si:dkey-217m5.8 |
| chr24_-_40667800 | 0.15 |
ENSDART00000169315
|
smyhc1
|
slow myosin heavy chain 1 |
| chr6_+_4084739 | 0.14 |
ENSDART00000087661
|
sema5bb
|
sema domain, seven thrombospondin repeats (type 1 and type 1-like), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 5Bb |
| chr5_-_29587351 | 0.13 |
ENSDART00000136446
ENSDART00000051434 |
entpd2a.1
|
ectonucleoside triphosphate diphosphohydrolase 2a, tandem duplicate 1 |
| chr12_+_19030391 | 0.13 |
ENSDART00000153927
|
si:ch73-139e5.2
|
si:ch73-139e5.2 |
| chr9_-_56420087 | 0.13 |
ENSDART00000112016
|
actr3
|
ARP3 actin related protein 3 homolog |
| chr8_-_1973220 | 0.12 |
ENSDART00000184435
|
si:dkey-178e17.1
|
si:dkey-178e17.1 |
| chr10_+_4875262 | 0.12 |
ENSDART00000165942
|
palm2
|
paralemmin 2 |
| chr8_-_30204650 | 0.12 |
ENSDART00000133209
|
zgc:162939
|
zgc:162939 |
| chr20_-_39391492 | 0.12 |
ENSDART00000184886
|
si:dkey-217m5.8
|
si:dkey-217m5.8 |
| chr14_+_13453130 | 0.11 |
ENSDART00000054849
|
pls3
|
plastin 3 (T isoform) |
| chr15_+_29085955 | 0.11 |
ENSDART00000156799
|
si:ch211-137a8.4
|
si:ch211-137a8.4 |
| chr7_+_35229645 | 0.11 |
ENSDART00000144327
|
tppp3
|
tubulin polymerization-promoting protein family member 3 |
| chr18_+_7309150 | 0.10 |
ENSDART00000052792
|
cers3b
|
ceramide synthase 3b |
| chr19_-_677713 | 0.09 |
ENSDART00000025146
|
slc6a19a.1
|
solute carrier family 6 (neutral amino acid transporter), member 19a, tandem duplicate 1 |
| chr8_+_28065803 | 0.09 |
ENSDART00000178481
|
kcnd3
|
potassium voltage-gated channel, Shal-related subfamily, member 3 |
| chr16_-_20492799 | 0.09 |
ENSDART00000014769
|
chn2
|
chimerin 2 |
| chr17_-_24353462 | 0.08 |
ENSDART00000191021
|
wdpcp
|
WD repeat containing planar cell polarity effector |
| chr6_-_39344259 | 0.08 |
ENSDART00000104074
|
zgc:158846
|
zgc:158846 |
| chr18_+_814328 | 0.08 |
ENSDART00000157801
ENSDART00000188492 ENSDART00000184489 |
fam219b
|
family with sequence similarity 219, member B |
| chr23_-_44233408 | 0.08 |
ENSDART00000149318
ENSDART00000085528 |
zgc:158659
|
zgc:158659 |
| chr7_-_2039060 | 0.08 |
ENSDART00000173879
|
si:cabz01007794.1
|
si:cabz01007794.1 |
| chr18_-_24988645 | 0.06 |
ENSDART00000136434
ENSDART00000085735 |
chd2
|
chromodomain helicase DNA binding protein 2 |
| chr19_-_24218942 | 0.05 |
ENSDART00000189198
|
BX547993.2
|
|
| chr16_-_50918908 | 0.05 |
ENSDART00000108744
|
CU855821.1
|
|
| chr7_-_32599669 | 0.05 |
ENSDART00000173752
|
kcna4
|
potassium voltage-gated channel, shaker-related subfamily, member 4 |
| chr6_+_41181869 | 0.05 |
ENSDART00000002046
|
opn1mw1
|
opsin 1 (cone pigments), medium-wave-sensitive, 1 |
| chr16_-_35329803 | 0.05 |
ENSDART00000161729
ENSDART00000157700 ENSDART00000184584 ENSDART00000174713 ENSDART00000162518 |
ptprub
|
protein tyrosine phosphatase, receptor type, U, b |
| chr25_+_126174 | 0.05 |
ENSDART00000168913
|
cntn1a
|
contactin 1a |
| chr4_+_69577362 | 0.05 |
ENSDART00000160913
|
znf1030
|
zinc finger protein 1030 |
| chr24_-_38644937 | 0.04 |
ENSDART00000170194
|
slc6a16b
|
solute carrier family 6, member 16b |
| chr6_+_41186320 | 0.04 |
ENSDART00000025241
|
opn1mw2
|
opsin 1 (cone pigments), medium-wave-sensitive, 2 |
| chr6_-_52757276 | 0.03 |
ENSDART00000159154
|
AL954861.2
|
|
| chr8_+_54013199 | 0.03 |
ENSDART00000158497
|
CABZ01079663.1
|
|
| chr8_-_3413139 | 0.03 |
ENSDART00000182673
ENSDART00000166741 ENSDART00000169430 ENSDART00000170478 |
FUT9 (1 of many)
FUT9 (1 of many)
fut9b
|
zgc:103510 zgc:165519 fucosyltransferase 9b |
| chr18_-_40508528 | 0.03 |
ENSDART00000185249
|
chrna5
|
cholinergic receptor, nicotinic, alpha 5 |
| chr7_-_1163829 | 0.02 |
ENSDART00000181937
|
CT030017.1
|
|
| chr19_+_7847920 | 0.02 |
ENSDART00000132454
|
si:dkeyp-85e10.1
|
si:dkeyp-85e10.1 |
| chr10_+_4874542 | 0.02 |
ENSDART00000101431
|
palm2
|
paralemmin 2 |
| chr5_-_35953472 | 0.02 |
ENSDART00000143448
|
rxfp2l
|
relaxin/insulin-like family peptide receptor 2, like |
| chr2_-_54639964 | 0.02 |
ENSDART00000100103
|
acss2l
|
acyl-CoA synthetase short chain family member 2 like |
| chr22_-_10019768 | 0.01 |
ENSDART00000162077
|
BX324216.2
|
|
| chr24_+_37461457 | 0.01 |
ENSDART00000165775
|
nlrc3
|
NLR family, CARD domain containing 3 |
| chr22_-_16412496 | 0.01 |
ENSDART00000137497
|
cts12
|
cathepsin 12 |
| chr13_-_40238813 | 0.01 |
ENSDART00000044963
|
loxl4
|
lysyl oxidase-like 4 |
| chr20_+_48129759 | 0.01 |
ENSDART00000148494
|
tram2
|
translocation associated membrane protein 2 |
| chr9_-_11551608 | 0.00 |
ENSDART00000128629
|
fev
|
FEV (ETS oncogene family) |
| chr4_+_64562090 | 0.00 |
ENSDART00000188810
|
si:ch211-223a21.3
|
si:ch211-223a21.3 |
| chr21_+_21811867 | 0.00 |
ENSDART00000185752
|
neu3.4
|
sialidase 3 (membrane sialidase), tandem duplicate 4 |
Network of associatons between targets according to the STRING database.
First level regulatory network of klf3+klf8
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.8 | GO:0010755 | regulation of plasminogen activation(GO:0010755) |
| 0.2 | 0.7 | GO:0006740 | NADPH regeneration(GO:0006740) |
| 0.1 | 0.4 | GO:1903358 | regulation of Golgi organization(GO:1903358) |
| 0.1 | 0.4 | GO:0050955 | thermoception(GO:0050955) detection of temperature stimulus involved in thermoception(GO:0050960) detection of temperature stimulus involved in sensory perception(GO:0050961) |
| 0.1 | 0.5 | GO:0032979 | protein insertion into membrane from inner side(GO:0032978) protein insertion into mitochondrial membrane from inner side(GO:0032979) protein insertion into mitochondrial membrane(GO:0051204) |
| 0.1 | 0.2 | GO:2000812 | regulation of barbed-end actin filament capping(GO:2000812) |
| 0.1 | 0.4 | GO:0006398 | mRNA 3'-end processing by stem-loop binding and cleavage(GO:0006398) |
| 0.1 | 0.4 | GO:0031444 | slow-twitch skeletal muscle fiber contraction(GO:0031444) |
| 0.0 | 2.0 | GO:0021587 | cerebellum morphogenesis(GO:0021587) |
| 0.0 | 0.8 | GO:0015671 | oxygen transport(GO:0015671) |
| 0.0 | 0.6 | GO:0036444 | calcium ion transmembrane import into mitochondrion(GO:0036444) |
| 0.0 | 0.1 | GO:0006843 | mitochondrial citrate transport(GO:0006843) |
| 0.0 | 0.1 | GO:0097623 | potassium ion export(GO:0071435) potassium ion export across plasma membrane(GO:0097623) |
| 0.0 | 0.4 | GO:0045762 | activation of adenylate cyclase activity(GO:0007190) positive regulation of cAMP metabolic process(GO:0030816) positive regulation of cAMP biosynthetic process(GO:0030819) positive regulation of adenylate cyclase activity(GO:0045762) |
| 0.0 | 0.4 | GO:0009134 | nucleoside diphosphate catabolic process(GO:0009134) |
| 0.0 | 0.2 | GO:0070262 | peptidyl-serine dephosphorylation(GO:0070262) |
| 0.0 | 0.5 | GO:0021551 | central nervous system morphogenesis(GO:0021551) |
| 0.0 | 0.2 | GO:0010887 | negative regulation of cholesterol storage(GO:0010887) |
| 0.0 | 0.4 | GO:0051568 | histone H3-K4 methylation(GO:0051568) |
| 0.0 | 0.4 | GO:0048246 | macrophage chemotaxis(GO:0048246) |
| 0.0 | 0.2 | GO:0015781 | pyrimidine nucleotide-sugar transport(GO:0015781) pyrimidine nucleotide-sugar transmembrane transport(GO:0090481) |
| 0.0 | 1.0 | GO:0055113 | epiboly involved in gastrulation with mouth forming second(GO:0055113) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0090443 | FAR/SIN/STRIPAK complex(GO:0090443) |
| 0.1 | 0.8 | GO:0031838 | haptoglobin-hemoglobin complex(GO:0031838) |
| 0.1 | 0.6 | GO:1990246 | uniplex complex(GO:1990246) |
| 0.0 | 0.1 | GO:0097541 | axonemal basal plate(GO:0097541) |
| 0.0 | 0.4 | GO:0005847 | mRNA cleavage and polyadenylation specificity factor complex(GO:0005847) |
| 0.0 | 0.4 | GO:0048188 | Set1C/COMPASS complex(GO:0048188) |
| 0.0 | 0.2 | GO:0005688 | U6 snRNP(GO:0005688) |
| 0.0 | 0.5 | GO:0005838 | proteasome regulatory particle(GO:0005838) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.7 | GO:0008746 | NAD(P)+ transhydrogenase activity(GO:0008746) oxidoreductase activity, acting on NAD(P)H, NAD(P) as acceptor(GO:0016652) |
| 0.1 | 0.4 | GO:0008131 | primary amine oxidase activity(GO:0008131) |
| 0.1 | 0.8 | GO:0031720 | haptoglobin binding(GO:0031720) |
| 0.1 | 0.5 | GO:0032977 | membrane insertase activity(GO:0032977) |
| 0.0 | 0.5 | GO:0070122 | isopeptidase activity(GO:0070122) |
| 0.0 | 0.4 | GO:0005324 | long-chain fatty acid transporter activity(GO:0005324) |
| 0.0 | 0.4 | GO:0045134 | guanosine-diphosphatase activity(GO:0004382) uridine-diphosphatase activity(GO:0045134) |
| 0.0 | 0.2 | GO:0005459 | UDP-galactose transmembrane transporter activity(GO:0005459) |
| 0.0 | 0.1 | GO:0005250 | A-type (transient outward) potassium channel activity(GO:0005250) |
| 0.0 | 0.4 | GO:0015278 | calcium-release channel activity(GO:0015278) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.2 | PID KIT PATHWAY | Signaling events mediated by Stem cell factor receptor (c-Kit) |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.0 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.0 | 1.2 | REACTOME G1 PHASE | Genes involved in G1 Phase |
| 0.0 | 0.3 | REACTOME SOS MEDIATED SIGNALLING | Genes involved in SOS-mediated signalling |
| 0.0 | 0.4 | REACTOME PROCESSING OF INTRONLESS PRE MRNAS | Genes involved in Processing of Intronless Pre-mRNAs |
| 0.0 | 0.4 | REACTOME ABC FAMILY PROTEINS MEDIATED TRANSPORT | Genes involved in ABC-family proteins mediated transport |
| 0.0 | 0.5 | REACTOME CROSS PRESENTATION OF SOLUBLE EXOGENOUS ANTIGENS ENDOSOMES | Genes involved in Cross-presentation of soluble exogenous antigens (endosomes) |
| 0.0 | 0.2 | REACTOME MRNA DECAY BY 5 TO 3 EXORIBONUCLEASE | Genes involved in mRNA Decay by 5' to 3' Exoribonuclease |