Project
PRJNA438478: RNAseq of wild type zebrafish germline
Navigation
Downloads
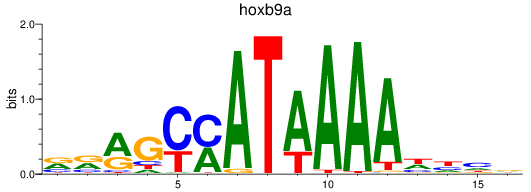
Results for hoxb9a
Z-value: 0.35
Transcription factors associated with hoxb9a
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
hoxb9a
|
ENSDARG00000056023 | homeobox B9a |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| hoxb9a | dr11_v1_chr3_+_23677351_23677351 | -0.79 | 8.9e-05 | Click! |
Activity profile of hoxb9a motif
Sorted Z-values of hoxb9a motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr10_+_44699734 | 1.14 |
ENSDART00000167952
ENSDART00000158681 ENSDART00000190188 ENSDART00000168276 |
scarb1
|
scavenger receptor class B, member 1 |
| chr24_-_9997948 | 1.09 |
ENSDART00000136274
|
si:ch211-146l10.7
|
si:ch211-146l10.7 |
| chr1_-_9485939 | 1.08 |
ENSDART00000157814
|
micall2b
|
mical-like 2b |
| chr10_+_44700103 | 1.01 |
ENSDART00000165999
|
scarb1
|
scavenger receptor class B, member 1 |
| chr20_+_27087539 | 0.93 |
ENSDART00000062094
|
tmem251
|
transmembrane protein 251 |
| chr21_-_19918286 | 0.88 |
ENSDART00000180816
|
ppp1r3b
|
protein phosphatase 1, regulatory subunit 3B |
| chr17_+_24320861 | 0.61 |
ENSDART00000179858
|
otx1
|
orthodenticle homeobox 1 |
| chr20_-_31496679 | 0.60 |
ENSDART00000153437
ENSDART00000153193 |
sash1a
|
SAM and SH3 domain containing 1a |
| chr16_+_38940758 | 0.55 |
ENSDART00000102482
ENSDART00000136215 |
eny2
|
enhancer of yellow 2 homolog (Drosophila) |
| chr19_-_47571456 | 0.54 |
ENSDART00000158071
ENSDART00000165841 |
rrm2
|
ribonucleotide reductase M2 polypeptide |
| chr2_-_45663945 | 0.53 |
ENSDART00000075080
|
prpf38b
|
pre-mRNA processing factor 38B |
| chr23_+_30736895 | 0.46 |
ENSDART00000042944
|
asxl1
|
additional sex combs like transcriptional regulator 1 |
| chr6_+_19948043 | 0.44 |
ENSDART00000182636
|
pik3r5
|
phosphoinositide-3-kinase, regulatory subunit 5 |
| chr21_-_32781612 | 0.44 |
ENSDART00000031028
|
cnot6a
|
CCR4-NOT transcription complex, subunit 6a |
| chr5_-_36597612 | 0.44 |
ENSDART00000031270
ENSDART00000122098 |
rhogc
|
ras homolog gene family, member Gc |
| chr17_-_30652738 | 0.43 |
ENSDART00000154960
|
sh3yl1
|
SH3 and SYLF domain containing 1 |
| chr3_+_36646054 | 0.43 |
ENSDART00000170013
ENSDART00000159948 |
gspt1l
|
G1 to S phase transition 1, like |
| chr11_+_24314148 | 0.42 |
ENSDART00000171491
|
rem1
|
RAS (RAD and GEM)-like GTP-binding 1 |
| chr15_-_26931541 | 0.41 |
ENSDART00000027563
|
ccdc9
|
coiled-coil domain containing 9 |
| chr12_-_33817114 | 0.41 |
ENSDART00000161265
|
twnk
|
twinkle mtDNA helicase |
| chr8_+_3431671 | 0.40 |
ENSDART00000017850
|
ctu1
|
cytosolic thiouridylase subunit 1 homolog (S. pombe) |
| chr13_-_15793585 | 0.40 |
ENSDART00000145914
ENSDART00000010286 |
bag5
|
BCL2 associated athanogene 5 |
| chr15_+_29727799 | 0.40 |
ENSDART00000182006
|
zgc:153372
|
zgc:153372 |
| chr15_+_32405959 | 0.39 |
ENSDART00000177269
|
si:ch211-162k9.6
|
si:ch211-162k9.6 |
| chr14_+_24840669 | 0.38 |
ENSDART00000106039
|
arhgef37
|
Rho guanine nucleotide exchange factor (GEF) 37 |
| chr5_-_32336613 | 0.38 |
ENSDART00000139732
|
dab2
|
Dab, mitogen-responsive phosphoprotein, homolog 2 (Drosophila) |
| chr20_-_29499363 | 0.38 |
ENSDART00000152889
ENSDART00000153252 ENSDART00000170972 ENSDART00000166420 ENSDART00000163079 |
rrm2
|
ribonucleotide reductase M2 polypeptide |
| chr3_+_33745014 | 0.37 |
ENSDART00000159966
|
nacc1a
|
nucleus accumbens associated 1, BEN and BTB (POZ) domain containing a |
| chr18_-_48296793 | 0.37 |
ENSDART00000032184
ENSDART00000193076 |
CABZ01069595.1
|
|
| chr25_+_16080181 | 0.37 |
ENSDART00000061753
|
far1
|
fatty acyl CoA reductase 1 |
| chr5_-_37881345 | 0.36 |
ENSDART00000084819
|
arhgap35b
|
Rho GTPase activating protein 35b |
| chr5_-_30074332 | 0.34 |
ENSDART00000147963
|
bco2a
|
beta-carotene oxygenase 2a |
| chr10_-_25561751 | 0.34 |
ENSDART00000147089
|
grik1a
|
glutamate receptor, ionotropic, kainate 1a |
| chr1_+_52481332 | 0.33 |
ENSDART00000074231
|
cldnd1b
|
claudin domain containing 1b |
| chr2_+_16696052 | 0.32 |
ENSDART00000022356
ENSDART00000164329 |
ppp1r7
|
protein phosphatase 1, regulatory (inhibitor) subunit 7 |
| chr24_+_29352039 | 0.32 |
ENSDART00000101641
|
prmt6
|
protein arginine methyltransferase 6 |
| chr16_-_17586883 | 0.32 |
ENSDART00000017142
|
m6pr
|
mannose-6-phosphate receptor (cation dependent) |
| chr22_+_2751887 | 0.31 |
ENSDART00000133652
|
si:dkey-20i20.11
|
si:dkey-20i20.11 |
| chr19_-_7358184 | 0.31 |
ENSDART00000092379
|
oxr1b
|
oxidation resistance 1b |
| chr11_-_38533505 | 0.30 |
ENSDART00000113894
|
slc45a3
|
solute carrier family 45, member 3 |
| chr15_+_29662401 | 0.29 |
ENSDART00000135540
|
nrip1a
|
nuclear receptor interacting protein 1a |
| chr1_+_2321384 | 0.29 |
ENSDART00000157662
|
farp1
|
FERM, RhoGEF (ARHGEF) and pleckstrin domain protein 1 (chondrocyte-derived) |
| chr21_-_11970199 | 0.29 |
ENSDART00000114524
|
nop56
|
NOP56 ribonucleoprotein homolog |
| chr5_-_57723929 | 0.29 |
ENSDART00000144237
|
gig2p
|
grass carp reovirus (GCRV)-induced gene 2p |
| chr8_-_53166975 | 0.28 |
ENSDART00000114683
|
rbsn
|
rabenosyn, RAB effector |
| chr16_-_7793457 | 0.27 |
ENSDART00000113483
|
trim71
|
tripartite motif containing 71, E3 ubiquitin protein ligase |
| chr1_-_9249943 | 0.27 |
ENSDART00000055011
|
zgc:136472
|
zgc:136472 |
| chr13_-_2283176 | 0.26 |
ENSDART00000158462
|
lrrc1
|
leucine rich repeat containing 1 |
| chr11_+_25560632 | 0.25 |
ENSDART00000033914
|
mbd1b
|
methyl-CpG binding domain protein 1b |
| chr16_-_27138478 | 0.25 |
ENSDART00000147438
|
tmem245
|
transmembrane protein 245 |
| chr21_-_39024754 | 0.24 |
ENSDART00000056878
|
traf4b
|
tnf receptor-associated factor 4b |
| chr20_-_31497300 | 0.24 |
ENSDART00000046841
|
sash1a
|
SAM and SH3 domain containing 1a |
| chr7_-_59589547 | 0.23 |
ENSDART00000008554
ENSDART00000101731 |
larp7
|
La ribonucleoprotein domain family, member 7 |
| chr17_+_33375469 | 0.23 |
ENSDART00000032827
|
zgc:162964
|
zgc:162964 |
| chr13_+_43400443 | 0.22 |
ENSDART00000084321
|
dact2
|
dishevelled-binding antagonist of beta-catenin 2 |
| chr5_-_9625459 | 0.22 |
ENSDART00000143347
|
sh2b3
|
SH2B adaptor protein 3 |
| chr14_+_30795559 | 0.22 |
ENSDART00000006132
|
cfl1
|
cofilin 1 |
| chr14_+_14841685 | 0.22 |
ENSDART00000158291
ENSDART00000162039 |
slbp
|
stem-loop binding protein |
| chr24_+_30392834 | 0.22 |
ENSDART00000162555
|
dpyda.1
|
dihydropyrimidine dehydrogenase a, tandem duplicate 1 |
| chr20_-_34670236 | 0.21 |
ENSDART00000033325
|
slc25a24
|
solute carrier family 25 (mitochondrial carrier; phosphate carrier), member 24 |
| chr3_-_25490970 | 0.21 |
ENSDART00000184651
|
grb2b
|
growth factor receptor-bound protein 2b |
| chr10_-_2788668 | 0.21 |
ENSDART00000131749
ENSDART00000124356 ENSDART00000085031 |
ash2l
|
ash2 (absent, small, or homeotic)-like (Drosophila) |
| chr1_+_45663727 | 0.20 |
ENSDART00000038574
ENSDART00000141144 ENSDART00000149565 |
trappc5
|
trafficking protein particle complex 5 |
| chr5_-_31856681 | 0.20 |
ENSDART00000187817
|
pkn3
|
protein kinase N3 |
| chr25_+_32496877 | 0.19 |
ENSDART00000132698
|
ctdspl2a
|
CTD (carboxy-terminal domain, RNA polymerase II, polypeptide A) small phosphatase like 2a |
| chr25_+_22572296 | 0.19 |
ENSDART00000152096
|
stra6
|
stimulated by retinoic acid 6 |
| chr6_-_3982783 | 0.19 |
ENSDART00000171944
|
slc25a12
|
solute carrier family 25 (aspartate/glutamate carrier), member 12 |
| chr5_-_31857345 | 0.18 |
ENSDART00000112546
|
pkn3
|
protein kinase N3 |
| chr11_+_25560072 | 0.18 |
ENSDART00000124131
ENSDART00000147179 |
mbd1b
|
methyl-CpG binding domain protein 1b |
| chr2_-_40135942 | 0.18 |
ENSDART00000176951
ENSDART00000098632 ENSDART00000148563 ENSDART00000149895 |
epha4a
|
eph receptor A4a |
| chr21_-_27251112 | 0.17 |
ENSDART00000137667
|
mark2a
|
MAP/microtubule affinity-regulating kinase 2a |
| chr11_-_36009924 | 0.17 |
ENSDART00000189959
ENSDART00000167472 ENSDART00000191211 ENSDART00000191662 ENSDART00000191780 ENSDART00000192622 ENSDART00000179911 |
itpr1b
|
inositol 1,4,5-trisphosphate receptor, type 1b |
| chr8_-_3312384 | 0.17 |
ENSDART00000035965
|
fut9b
|
fucosyltransferase 9b |
| chr13_+_36585399 | 0.17 |
ENSDART00000030211
|
gmfb
|
glia maturation factor, beta |
| chr12_+_27704015 | 0.17 |
ENSDART00000153256
|
cacna1g
|
calcium channel, voltage-dependent, T type, alpha 1G subunit |
| chr20_-_2713120 | 0.16 |
ENSDART00000138753
ENSDART00000104606 |
rars2
|
arginyl-tRNA synthetase 2, mitochondrial (putative) |
| chr5_+_24087035 | 0.16 |
ENSDART00000183644
|
tp53
|
tumor protein p53 |
| chr22_+_2512154 | 0.15 |
ENSDART00000097363
|
zgc:173726
|
zgc:173726 |
| chr7_-_7692723 | 0.15 |
ENSDART00000183352
|
aadat
|
aminoadipate aminotransferase |
| chr10_-_10969444 | 0.15 |
ENSDART00000138041
|
exd3
|
exonuclease 3'-5' domain containing 3 |
| chr7_+_24522308 | 0.15 |
ENSDART00000173542
|
btr09
|
bloodthirsty-related gene family, member 9 |
| chr8_-_50888806 | 0.14 |
ENSDART00000053750
|
acsl2
|
acyl-CoA synthetase long chain family member 2 |
| chr19_-_7033636 | 0.14 |
ENSDART00000122815
ENSDART00000123443 ENSDART00000124094 |
daxx
|
death-domain associated protein |
| chr2_+_23039041 | 0.14 |
ENSDART00000056921
|
csnk1g2a
|
casein kinase 1, gamma 2a |
| chr21_-_3700334 | 0.14 |
ENSDART00000137844
|
atp8b1
|
ATPase phospholipid transporting 8B1 |
| chr17_-_50050453 | 0.13 |
ENSDART00000182057
|
zgc:100951
|
zgc:100951 |
| chr25_+_20089986 | 0.13 |
ENSDART00000143441
ENSDART00000184073 |
tnni4b.2
|
troponin I4b, tandem duplicate 2 |
| chr10_+_15107886 | 0.13 |
ENSDART00000188047
ENSDART00000164095 |
scpp8
|
secretory calcium-binding phosphoprotein 8 |
| chr7_-_7692992 | 0.13 |
ENSDART00000192619
|
aadat
|
aminoadipate aminotransferase |
| chr17_-_50059664 | 0.13 |
ENSDART00000193940
|
FO704836.1
|
|
| chr4_+_12292274 | 0.13 |
ENSDART00000061070
ENSDART00000150786 |
mkrn1
|
makorin, ring finger protein, 1 |
| chr5_-_67750907 | 0.11 |
ENSDART00000172097
|
b4galt4
|
UDP-Gal:betaGlcNAc beta 1,4- galactosyltransferase, polypeptide 4 |
| chr15_+_44093286 | 0.11 |
ENSDART00000114352
|
zgc:112998
|
zgc:112998 |
| chr17_-_21057617 | 0.10 |
ENSDART00000148095
ENSDART00000048853 |
ube2d1a
|
ubiquitin-conjugating enzyme E2D 1a |
| chr24_-_32025637 | 0.10 |
ENSDART00000180448
ENSDART00000159034 |
rsu1
|
Ras suppressor protein 1 |
| chr8_-_45867358 | 0.10 |
ENSDART00000132810
|
adam9
|
ADAM metallopeptidase domain 9 |
| chr2_+_2967255 | 0.10 |
ENSDART00000167649
ENSDART00000166449 |
pik3r3a
|
phosphoinositide-3-kinase, regulatory subunit 3a (gamma) |
| chr1_+_26105141 | 0.10 |
ENSDART00000102379
ENSDART00000127154 |
toporsa
|
topoisomerase I binding, arginine/serine-rich a |
| chr25_-_12923482 | 0.10 |
ENSDART00000161754
|
CR450808.1
|
|
| chr5_-_54424019 | 0.09 |
ENSDART00000169270
ENSDART00000164067 |
coq4
|
coenzyme Q4 homolog (S. cerevisiae) |
| chr23_-_7799184 | 0.09 |
ENSDART00000190946
ENSDART00000165427 |
myt1b
|
myelin transcription factor 1b |
| chr22_+_1421212 | 0.09 |
ENSDART00000161813
|
zgc:101130
|
zgc:101130 |
| chr17_-_44249720 | 0.09 |
ENSDART00000156648
|
otx2b
|
orthodenticle homeobox 2b |
| chr3_-_53091946 | 0.09 |
ENSDART00000187297
|
lpar2a
|
lysophosphatidic acid receptor 2a |
| chr21_-_2415808 | 0.08 |
ENSDART00000171179
|
si:ch211-241b2.5
|
si:ch211-241b2.5 |
| chr3_+_58119571 | 0.08 |
ENSDART00000108979
|
si:ch211-256e16.4
|
si:ch211-256e16.4 |
| chr12_+_10053852 | 0.08 |
ENSDART00000078497
|
slc4a1b
|
solute carrier family 4 (anion exchanger), member 1b (Diego blood group) |
| chr10_+_11282883 | 0.08 |
ENSDART00000135355
|
si:ch211-126i22.5
|
si:ch211-126i22.5 |
| chr7_+_31145386 | 0.07 |
ENSDART00000075407
ENSDART00000169462 |
fam189a1
|
family with sequence similarity 189, member A1 |
| chr11_+_45323455 | 0.07 |
ENSDART00000189249
|
mafgb
|
v-maf avian musculoaponeurotic fibrosarcoma oncogene homolog Gb |
| chr16_-_21915223 | 0.07 |
ENSDART00000170634
|
setdb1b
|
SET domain, bifurcated 1b |
| chr13_-_4018888 | 0.06 |
ENSDART00000058238
|
tjap1
|
tight junction associated protein 1 (peripheral) |
| chr25_-_31898552 | 0.06 |
ENSDART00000156128
|
si:ch73-330k17.3
|
si:ch73-330k17.3 |
| chr11_-_15090118 | 0.06 |
ENSDART00000171118
|
slc1a8a
|
solute carrier family 1 (glutamate transporter), member 8a |
| chr2_+_19785540 | 0.06 |
ENSDART00000149789
|
ptbp2b
|
polypyrimidine tract binding protein 2b |
| chr23_+_13124085 | 0.05 |
ENSDART00000139475
|
samd10b
|
sterile alpha motif domain containing 10b |
| chr9_-_9980704 | 0.05 |
ENSDART00000130243
ENSDART00000193475 |
ugt1ab
|
UDP glucuronosyltransferase 1 family a, b |
| chr4_-_71108793 | 0.05 |
ENSDART00000189856
|
si:dkeyp-80d11.1
|
si:dkeyp-80d11.1 |
| chr23_+_3625083 | 0.04 |
ENSDART00000184958
|
si:dkey-9l20.3
|
si:dkey-9l20.3 |
| chr8_+_45004666 | 0.04 |
ENSDART00000145348
|
si:ch211-163b2.4
|
si:ch211-163b2.4 |
| chr16_-_55028740 | 0.04 |
ENSDART00000156368
ENSDART00000161704 |
zgc:114181
|
zgc:114181 |
| chr17_-_23727978 | 0.04 |
ENSDART00000079600
|
minpp1a
|
multiple inositol-polyphosphate phosphatase 1a |
| chr1_+_56463494 | 0.04 |
ENSDART00000097964
|
zgc:171452
|
zgc:171452 |
| chr13_+_1182257 | 0.04 |
ENSDART00000033528
ENSDART00000183702 ENSDART00000147959 |
tnfaip3
|
tumor necrosis factor, alpha-induced protein 3 |
| chr13_+_31286076 | 0.04 |
ENSDART00000142725
|
gdf10a
|
growth differentiation factor 10a |
| chr15_+_14826966 | 0.04 |
ENSDART00000157912
ENSDART00000171133 |
or130-1
|
odorant receptor, family H, subfamily 130, member 1 |
| chr11_-_15090564 | 0.04 |
ENSDART00000162079
|
slc1a8a
|
solute carrier family 1 (glutamate transporter), member 8a |
| chr15_-_13254480 | 0.04 |
ENSDART00000190499
|
zgc:172282
|
zgc:172282 |
| chr5_+_72194444 | 0.04 |
ENSDART00000165436
|
ddx54
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 54 |
| chr1_-_56556326 | 0.04 |
ENSDART00000188760
|
CABZ01059408.1
|
|
| chr12_-_13081229 | 0.04 |
ENSDART00000152510
ENSDART00000126866 |
ccser2b
|
coiled-coil serine-rich protein 2b |
| chr8_-_51507144 | 0.04 |
ENSDART00000024882
ENSDART00000135166 |
fgfr1a
|
fibroblast growth factor receptor 1a |
| chr15_-_16704417 | 0.03 |
ENSDART00000155163
|
caln1
|
calneuron 1 |
| chr5_-_39698224 | 0.03 |
ENSDART00000076929
|
prkg2
|
protein kinase, cGMP-dependent, type II |
| chr4_+_33574463 | 0.03 |
ENSDART00000150255
|
si:dkey-84h14.1
|
si:dkey-84h14.1 |
| chr15_+_21254800 | 0.03 |
ENSDART00000142902
|
usf1
|
upstream transcription factor 1 |
| chr12_-_43428542 | 0.03 |
ENSDART00000192266
|
ptprea
|
protein tyrosine phosphatase, receptor type, E, a |
| chr4_-_46915962 | 0.03 |
ENSDART00000169555
|
si:ch211-134c10.1
|
si:ch211-134c10.1 |
| chr2_-_3158919 | 0.03 |
ENSDART00000098394
|
wnt3a
|
wingless-type MMTV integration site family, member 3A |
| chr4_-_11122638 | 0.03 |
ENSDART00000150600
|
si:dkey-21h14.10
|
si:dkey-21h14.10 |
| chr4_-_64142389 | 0.03 |
ENSDART00000172126
|
BX914205.3
|
|
| chr22_+_6321038 | 0.03 |
ENSDART00000179961
|
si:rp71-1i20.1
|
si:rp71-1i20.1 |
| chr3_-_29910547 | 0.03 |
ENSDART00000151501
|
RUNDC1
|
si:dkey-151m15.5 |
| chr17_+_15788100 | 0.02 |
ENSDART00000027667
|
rragd
|
ras-related GTP binding D |
| chr22_-_10605045 | 0.02 |
ENSDART00000184812
|
bap1
|
BRCA1 associated protein-1 (ubiquitin carboxy-terminal hydrolase) |
| chr3_-_49554912 | 0.02 |
ENSDART00000159392
|
CR847534.1
|
|
| chr20_-_47270519 | 0.02 |
ENSDART00000153329
|
si:dkeyp-104f11.8
|
si:dkeyp-104f11.8 |
| chr23_+_10352921 | 0.02 |
ENSDART00000081193
|
KRT18 (1 of many)
|
si:ch211-133j6.3 |
| chr6_+_41956355 | 0.02 |
ENSDART00000176856
|
oxtr
|
oxytocin receptor |
| chr15_-_2841677 | 0.02 |
ENSDART00000026145
ENSDART00000180290 |
AMOTL1
|
angiomotin like 1 |
| chr11_-_18253111 | 0.02 |
ENSDART00000125984
|
mustn1b
|
musculoskeletal, embryonic nuclear protein 1b |
| chr10_+_40660772 | 0.02 |
ENSDART00000148007
|
taar19l
|
trace amine associated receptor 19l |
| chr15_+_32643873 | 0.02 |
ENSDART00000189433
|
trpc4b
|
transient receptor potential cation channel, subfamily C, member 4b |
| chr4_+_59711338 | 0.02 |
ENSDART00000150849
|
si:dkey-149m13.4
|
si:dkey-149m13.4 |
| chr4_-_77300601 | 0.02 |
ENSDART00000163015
ENSDART00000174015 |
slco1f2
|
solute carrier organic anion transporter family, member 1F2 |
| chr23_-_21215311 | 0.02 |
ENSDART00000112424
|
megf6a
|
multiple EGF-like-domains 6a |
| chr14_-_41388178 | 0.02 |
ENSDART00000124532
ENSDART00000125016 ENSDART00000169247 |
cstf2
|
cleavage stimulation factor, 3' pre-RNA, subunit 2 |
| chr7_+_33152723 | 0.02 |
ENSDART00000132658
|
si:ch211-194p6.10
|
si:ch211-194p6.10 |
| chr2_+_15586632 | 0.02 |
ENSDART00000164903
|
BX510342.1
|
|
| chr9_-_27442339 | 0.01 |
ENSDART00000138602
|
stxbp5l
|
syntaxin binding protein 5-like |
| chr1_+_17900306 | 0.01 |
ENSDART00000089480
|
cyp4v8
|
cytochrome P450, family 4, subfamily V, polypeptide 8 |
| chr10_+_40700311 | 0.01 |
ENSDART00000157650
ENSDART00000138342 |
taar19n
|
trace amine associated receptor 19n |
| chr2_-_32637592 | 0.01 |
ENSDART00000136353
|
si:dkeyp-73d8.8
|
si:dkeyp-73d8.8 |
| chr19_+_19747430 | 0.01 |
ENSDART00000166129
|
hoxa9a
|
homeobox A9a |
| chr6_+_39536596 | 0.00 |
ENSDART00000185742
|
CU467856.3
|
|
| chr14_+_5861435 | 0.00 |
ENSDART00000041279
ENSDART00000147341 |
tubb4b
|
tubulin, beta 4B class IVb |
| chr23_+_20931030 | 0.00 |
ENSDART00000167014
|
pax7b
|
paired box 7b |
| chr4_+_17336557 | 0.00 |
ENSDART00000111650
|
pmch
|
pro-melanin-concentrating hormone |
Network of associatons between targets according to the STRING database.
First level regulatory network of hoxb9a
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 2.1 | GO:0043654 | recognition of apoptotic cell(GO:0043654) |
| 0.3 | 0.9 | GO:0005981 | regulation of glycogen catabolic process(GO:0005981) |
| 0.1 | 0.4 | GO:0018872 | arsonoacetate metabolic process(GO:0018872) |
| 0.1 | 0.4 | GO:0002184 | cytoplasmic translational termination(GO:0002184) |
| 0.1 | 0.4 | GO:1901842 | regulation of high voltage-gated calcium channel activity(GO:1901841) negative regulation of high voltage-gated calcium channel activity(GO:1901842) |
| 0.1 | 0.9 | GO:0009263 | deoxyribonucleotide biosynthetic process(GO:0009263) |
| 0.1 | 0.3 | GO:0034969 | histone arginine methylation(GO:0034969) |
| 0.1 | 0.2 | GO:1900182 | positive regulation of protein localization to nucleus(GO:1900182) regulation of protein localization to Cajal body(GO:1904869) positive regulation of protein localization to Cajal body(GO:1904871) |
| 0.1 | 0.2 | GO:0030043 | actin filament fragmentation(GO:0030043) |
| 0.1 | 0.4 | GO:0032447 | tRNA wobble position uridine thiolation(GO:0002143) protein urmylation(GO:0032447) |
| 0.1 | 0.6 | GO:0016578 | histone deubiquitination(GO:0016578) |
| 0.1 | 0.2 | GO:0007250 | activation of NF-kappaB-inducing kinase activity(GO:0007250) |
| 0.1 | 0.3 | GO:0010586 | miRNA metabolic process(GO:0010586) |
| 0.1 | 0.2 | GO:0043525 | regulation of transposition, RNA-mediated(GO:0010525) negative regulation of transposition, RNA-mediated(GO:0010526) transposition, RNA-mediated(GO:0032197) positive regulation of neuron apoptotic process(GO:0043525) positive regulation of neuron death(GO:1901216) |
| 0.0 | 0.4 | GO:1905207 | regulation of cardiocyte differentiation(GO:1905207) regulation of cardiac muscle cell differentiation(GO:2000725) |
| 0.0 | 0.4 | GO:0006264 | mitochondrial DNA replication(GO:0006264) |
| 0.0 | 0.2 | GO:0019860 | uracil catabolic process(GO:0006212) uracil metabolic process(GO:0019860) |
| 0.0 | 0.7 | GO:0003406 | retinal pigment epithelium development(GO:0003406) |
| 0.0 | 0.2 | GO:0071939 | vitamin A transport(GO:0071938) vitamin A import(GO:0071939) |
| 0.0 | 0.3 | GO:0016119 | carotene metabolic process(GO:0016119) carotene catabolic process(GO:0016121) terpene metabolic process(GO:0042214) terpene catabolic process(GO:0046247) |
| 0.0 | 0.2 | GO:1900108 | negative regulation of nodal signaling pathway(GO:1900108) |
| 0.0 | 0.2 | GO:0006398 | mRNA 3'-end processing by stem-loop binding and cleavage(GO:0006398) |
| 0.0 | 0.2 | GO:0003404 | optic vesicle morphogenesis(GO:0003404) |
| 0.0 | 0.3 | GO:0006622 | protein targeting to lysosome(GO:0006622) |
| 0.0 | 0.1 | GO:0003322 | pancreatic A cell development(GO:0003322) |
| 0.0 | 0.4 | GO:0031397 | negative regulation of protein ubiquitination(GO:0031397) |
| 0.0 | 1.1 | GO:0036269 | swimming behavior(GO:0036269) |
| 0.0 | 0.2 | GO:0015810 | acidic amino acid transport(GO:0015800) aspartate transport(GO:0015810) L-glutamate transport(GO:0015813) malate-aspartate shuttle(GO:0043490) |
| 0.0 | 0.1 | GO:0045647 | negative regulation of erythrocyte differentiation(GO:0045647) |
| 0.0 | 0.4 | GO:0016601 | Rac protein signal transduction(GO:0016601) |
| 0.0 | 0.5 | GO:0035336 | long-chain fatty-acyl-CoA metabolic process(GO:0035336) |
| 0.0 | 0.4 | GO:0000289 | nuclear-transcribed mRNA poly(A) tail shortening(GO:0000289) |
| 0.0 | 0.0 | GO:1900274 | positive regulation of phospholipase C activity(GO:0010863) regulation of phospholipase C activity(GO:1900274) |
| 0.0 | 0.2 | GO:0086010 | membrane depolarization during action potential(GO:0086010) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | GO:0070390 | transcription export complex 2(GO:0070390) |
| 0.1 | 0.4 | GO:0018444 | translation release factor complex(GO:0018444) |
| 0.1 | 0.5 | GO:0035517 | PR-DUB complex(GO:0035517) |
| 0.1 | 0.2 | GO:1990072 | TRAPPIII protein complex(GO:1990072) |
| 0.1 | 0.9 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.0 | 2.1 | GO:0044853 | caveola(GO:0005901) plasma membrane raft(GO:0044853) |
| 0.0 | 0.3 | GO:0031428 | box C/D snoRNP complex(GO:0031428) |
| 0.0 | 0.4 | GO:0005944 | phosphatidylinositol 3-kinase complex, class IB(GO:0005944) phosphatidylinositol 3-kinase complex, class I(GO:0097651) |
| 0.0 | 0.4 | GO:0030014 | CCR4-NOT complex(GO:0030014) |
| 0.0 | 0.5 | GO:0071011 | precatalytic spliceosome(GO:0071011) |
| 0.0 | 0.2 | GO:0048188 | Set1C/COMPASS complex(GO:0048188) |
| 0.0 | 0.1 | GO:0031314 | extrinsic component of mitochondrial inner membrane(GO:0031314) |
| 0.0 | 0.4 | GO:0097610 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 2.1 | GO:0030169 | low-density lipoprotein particle binding(GO:0030169) |
| 0.1 | 0.4 | GO:0030792 | arsenite methyltransferase activity(GO:0030791) methylarsonite methyltransferase activity(GO:0030792) |
| 0.1 | 0.9 | GO:0061731 | ribonucleoside-diphosphate reductase activity, thioredoxin disulfide as acceptor(GO:0004748) oxidoreductase activity, acting on CH or CH2 groups, disulfide as acceptor(GO:0016728) ribonucleoside-diphosphate reductase activity(GO:0061731) |
| 0.1 | 0.5 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.1 | 0.3 | GO:0008469 | histone-arginine N-methyltransferase activity(GO:0008469) |
| 0.1 | 0.9 | GO:2001069 | glycogen binding(GO:2001069) |
| 0.1 | 0.4 | GO:0080019 | fatty-acyl-CoA reductase (alcohol-forming) activity(GO:0080019) |
| 0.0 | 0.2 | GO:0002058 | uracil binding(GO:0002058) pyrimidine nucleobase binding(GO:0002061) dihydropyrimidine dehydrogenase (NADP+) activity(GO:0017113) |
| 0.0 | 0.4 | GO:0008079 | translation release factor activity(GO:0003747) translation termination factor activity(GO:0008079) |
| 0.0 | 0.3 | GO:0036137 | kynurenine-oxoglutarate transaminase activity(GO:0016212) kynurenine aminotransferase activity(GO:0036137) |
| 0.0 | 0.4 | GO:0000774 | adenyl-nucleotide exchange factor activity(GO:0000774) |
| 0.0 | 0.3 | GO:0003834 | beta-carotene 15,15'-monooxygenase activity(GO:0003834) carotenoid dioxygenase activity(GO:0010436) |
| 0.0 | 0.3 | GO:0015157 | sugar:proton symporter activity(GO:0005351) cation:sugar symporter activity(GO:0005402) sucrose:proton symporter activity(GO:0008506) sucrose transmembrane transporter activity(GO:0008515) disaccharide transmembrane transporter activity(GO:0015154) oligosaccharide transmembrane transporter activity(GO:0015157) |
| 0.0 | 0.2 | GO:0005068 | transmembrane receptor protein tyrosine kinase adaptor activity(GO:0005068) |
| 0.0 | 0.2 | GO:0017070 | U6 snRNA binding(GO:0017070) |
| 0.0 | 0.4 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.0 | 0.3 | GO:0035198 | miRNA binding(GO:0035198) |
| 0.0 | 0.2 | GO:0034632 | retinol transporter activity(GO:0034632) |
| 0.0 | 0.3 | GO:0005537 | mannose binding(GO:0005537) |
| 0.0 | 0.2 | GO:0008332 | low voltage-gated calcium channel activity(GO:0008332) |
| 0.0 | 0.2 | GO:0071933 | Arp2/3 complex binding(GO:0071933) |
| 0.0 | 0.5 | GO:0046935 | 1-phosphatidylinositol-3-kinase regulator activity(GO:0046935) |
| 0.0 | 0.4 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.0 | 0.2 | GO:0015183 | L-aspartate transmembrane transporter activity(GO:0015183) |
| 0.0 | 0.3 | GO:0030515 | snoRNA binding(GO:0030515) |
| 0.0 | 0.2 | GO:0005220 | inositol 1,4,5-trisphosphate-sensitive calcium-release channel activity(GO:0005220) |
| 0.0 | 0.4 | GO:0004698 | protein kinase C activity(GO:0004697) calcium-dependent protein kinase C activity(GO:0004698) |
| 0.0 | 0.1 | GO:0047676 | arachidonate-CoA ligase activity(GO:0047676) |
| 0.0 | 0.2 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.0 | 0.2 | GO:0036002 | pre-mRNA binding(GO:0036002) |
| 0.0 | 0.0 | GO:0034416 | bisphosphoglycerate phosphatase activity(GO:0034416) |
| 0.0 | 0.3 | GO:0099529 | neurotransmitter receptor activity involved in regulation of postsynaptic membrane potential(GO:0099529) transmitter-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1904315) |
| 0.0 | 0.2 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.0 | 0.4 | PID EPHA FWDPATHWAY | EPHA forward signaling |
| 0.0 | 0.7 | PID TGFBR PATHWAY | TGF-beta receptor signaling |
| 0.0 | 0.2 | SA G1 AND S PHASES | Cdk2, 4, and 6 bind cyclin D in G1, while cdk2/cyclin E promotes the G1/S transition. |
| 0.0 | 0.9 | PID E2F PATHWAY | E2F transcription factor network |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 2.1 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.1 | 0.4 | REACTOME GAP JUNCTION DEGRADATION | Genes involved in Gap junction degradation |
| 0.0 | 0.9 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.0 | 0.3 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 0.0 | 0.4 | REACTOME G BETA GAMMA SIGNALLING THROUGH PI3KGAMMA | Genes involved in G beta:gamma signalling through PI3Kgamma |
| 0.0 | 0.2 | REACTOME SEMA3A PAK DEPENDENT AXON REPULSION | Genes involved in Sema3A PAK dependent Axon repulsion |
| 0.0 | 0.2 | REACTOME SLBP DEPENDENT PROCESSING OF REPLICATION DEPENDENT HISTONE PRE MRNAS | Genes involved in SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs |
| 0.0 | 0.2 | REACTOME ACTIVATION OF BH3 ONLY PROTEINS | Genes involved in Activation of BH3-only proteins |
| 0.0 | 0.4 | REACTOME PEROXISOMAL LIPID METABOLISM | Genes involved in Peroxisomal lipid metabolism |
| 0.0 | 0.2 | REACTOME REGULATION OF KIT SIGNALING | Genes involved in Regulation of KIT signaling |
| 0.0 | 0.3 | REACTOME LYSOSOME VESICLE BIOGENESIS | Genes involved in Lysosome Vesicle Biogenesis |