Project
PRJNA438478: RNAseq of wild type zebrafish germline
Navigation
Downloads
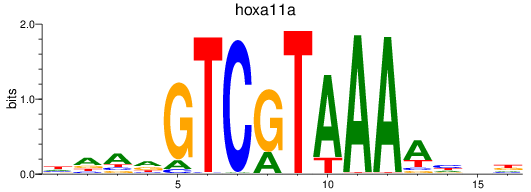
Results for hoxa11a
Z-value: 0.39
Transcription factors associated with hoxa11a
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
hoxa11a
|
ENSDARG00000104162 | homeobox A11a |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| hoxa11a | dr11_v1_chr19_+_19737214_19737300 | 0.09 | 7.2e-01 | Click! |
Activity profile of hoxa11a motif
Sorted Z-values of hoxa11a motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr4_-_20108833 | 0.52 |
ENSDART00000100867
|
fam3c
|
family with sequence similarity 3, member C |
| chr16_-_10316359 | 0.44 |
ENSDART00000104025
|
flot1b
|
flotillin 1b |
| chr21_+_13128180 | 0.43 |
ENSDART00000081426
|
odf2a
|
outer dense fiber of sperm tails 2a |
| chr2_-_37874647 | 0.42 |
ENSDART00000039386
|
zgc:66427
|
zgc:66427 |
| chr1_+_31019165 | 0.40 |
ENSDART00000102210
|
itga6b
|
integrin, alpha 6b |
| chr19_+_7424347 | 0.36 |
ENSDART00000004622
|
sf3b4
|
splicing factor 3b, subunit 4 |
| chr16_-_47426482 | 0.33 |
ENSDART00000148631
ENSDART00000149723 |
sept7b
|
septin 7b |
| chr9_+_20483846 | 0.33 |
ENSDART00000192067
ENSDART00000145111 |
parp4
|
poly (ADP-ribose) polymerase family, member 4 |
| chr8_-_33154677 | 0.29 |
ENSDART00000133300
|
zbtb34
|
zinc finger and BTB domain containing 34 |
| chr20_+_34671386 | 0.29 |
ENSDART00000152836
ENSDART00000138226 |
elp3
|
elongator acetyltransferase complex subunit 3 |
| chr2_+_54696042 | 0.28 |
ENSDART00000074270
|
ankrd12
|
ankyrin repeat domain 12 |
| chr24_-_21090447 | 0.27 |
ENSDART00000136507
ENSDART00000140786 ENSDART00000184841 |
qtrt2
|
queuine tRNA-ribosyltransferase accessory subunit 2 |
| chr17_+_44463230 | 0.25 |
ENSDART00000130311
|
naa30
|
N(alpha)-acetyltransferase 30, NatC catalytic subunit |
| chr15_+_44093286 | 0.22 |
ENSDART00000114352
|
zgc:112998
|
zgc:112998 |
| chr21_-_25618175 | 0.21 |
ENSDART00000133512
|
fosl1b
|
FOS-like antigen 1b |
| chr13_+_42309688 | 0.21 |
ENSDART00000158367
|
ide
|
insulin-degrading enzyme |
| chr7_+_9308625 | 0.21 |
ENSDART00000084598
|
selenos
|
selenoprotein S |
| chr21_+_13127742 | 0.18 |
ENSDART00000179221
|
odf2a
|
outer dense fiber of sperm tails 2a |
| chr4_-_6373735 | 0.18 |
ENSDART00000140100
|
MDFIC
|
si:ch73-156e19.1 |
| chr10_+_2587234 | 0.18 |
ENSDART00000126937
|
wu:fb59d01
|
wu:fb59d01 |
| chr7_-_30624435 | 0.17 |
ENSDART00000173828
|
rnf111
|
ring finger protein 111 |
| chr12_-_11560794 | 0.17 |
ENSDART00000149098
ENSDART00000169975 |
plekha1b
|
pleckstrin homology domain containing, family A (phosphoinositide binding specific) member 1b |
| chr7_-_30280934 | 0.16 |
ENSDART00000126741
|
serf2
|
small EDRK-rich factor 2 |
| chr1_+_36722122 | 0.16 |
ENSDART00000111566
|
tmem184c
|
transmembrane protein 184C |
| chr5_-_32489796 | 0.16 |
ENSDART00000168870
|
gpr107
|
G protein-coupled receptor 107 |
| chr13_+_29925397 | 0.15 |
ENSDART00000123482
|
cuedc2
|
CUE domain containing 2 |
| chr25_-_18002937 | 0.14 |
ENSDART00000149696
|
cep290
|
centrosomal protein 290 |
| chr3_-_40254634 | 0.14 |
ENSDART00000154562
|
top3a
|
DNA topoisomerase III alpha |
| chr6_-_40744720 | 0.14 |
ENSDART00000154916
ENSDART00000186922 |
p4htm
|
prolyl 4-hydroxylase, transmembrane (endoplasmic reticulum) |
| chr25_-_18002498 | 0.13 |
ENSDART00000158688
|
cep290
|
centrosomal protein 290 |
| chr6_+_42338309 | 0.13 |
ENSDART00000015277
|
gpx1b
|
glutathione peroxidase 1b |
| chr16_-_6205790 | 0.12 |
ENSDART00000038495
|
ctnnb1
|
catenin (cadherin-associated protein), beta 1 |
| chr23_+_33752275 | 0.11 |
ENSDART00000007260
|
si:ch211-210c8.6
|
si:ch211-210c8.6 |
| chr1_+_45056371 | 0.11 |
ENSDART00000073689
ENSDART00000167309 |
btr01
|
bloodthirsty-related gene family, member 1 |
| chr18_-_17087138 | 0.10 |
ENSDART00000135597
|
zc3h18
|
zinc finger CCCH-type containing 18 |
| chr11_-_26832685 | 0.10 |
ENSDART00000153519
|
iqsec1b
|
IQ motif and Sec7 domain 1b |
| chr3_+_24190207 | 0.10 |
ENSDART00000034762
|
prr15la
|
proline rich 15-like a |
| chr12_+_33361948 | 0.10 |
ENSDART00000124982
|
fasn
|
fatty acid synthase |
| chr12_-_26851726 | 0.10 |
ENSDART00000047724
|
zeb1b
|
zinc finger E-box binding homeobox 1b |
| chr12_+_27117609 | 0.09 |
ENSDART00000076154
|
hoxb8b
|
homeobox B8b |
| chr11_+_24716837 | 0.09 |
ENSDART00000145217
|
zgc:153953
|
zgc:153953 |
| chr22_-_33916620 | 0.09 |
ENSDART00000191276
|
CR974456.1
|
|
| chr5_+_36611128 | 0.08 |
ENSDART00000097684
|
nova1
|
neuro-oncological ventral antigen 1 |
| chr8_+_23861461 | 0.08 |
ENSDART00000037109
|
srpk1a
|
SRSF protein kinase 1a |
| chr5_+_59397739 | 0.06 |
ENSDART00000148659
|
clip2
|
CAP-GLY domain containing linker protein 2 |
| chr15_-_734203 | 0.06 |
ENSDART00000177867
ENSDART00000154918 |
si:dkey-7i4.21
BX004964.1
|
si:dkey-7i4.21 |
| chr11_+_43751263 | 0.06 |
ENSDART00000163843
|
zgc:153431
|
zgc:153431 |
| chr11_+_25560072 | 0.06 |
ENSDART00000124131
ENSDART00000147179 |
mbd1b
|
methyl-CpG binding domain protein 1b |
| chr17_-_23727978 | 0.05 |
ENSDART00000079600
|
minpp1a
|
multiple inositol-polyphosphate phosphatase 1a |
| chr17_+_17764979 | 0.05 |
ENSDART00000105013
|
alkbh1
|
alkB homolog 1, histone H2A dioxygenase |
| chr9_-_39005317 | 0.05 |
ENSDART00000014207
|
myl1
|
myosin, light chain 1, alkali; skeletal, fast |
| chr4_+_39368978 | 0.05 |
ENSDART00000160640
|
si:dkey-261o4.1
|
si:dkey-261o4.1 |
| chr11_+_25560632 | 0.05 |
ENSDART00000033914
|
mbd1b
|
methyl-CpG binding domain protein 1b |
| chr17_-_31695217 | 0.04 |
ENSDART00000104332
ENSDART00000143090 |
lin52
|
lin-52 DREAM MuvB core complex component |
| chr6_+_49551614 | 0.04 |
ENSDART00000022581
|
rab22a
|
RAB22A, member RAS oncogene family |
| chr8_-_12432604 | 0.04 |
ENSDART00000133350
ENSDART00000140699 ENSDART00000101174 |
traf1
|
TNF receptor-associated factor 1 |
| chr9_-_23922011 | 0.04 |
ENSDART00000145734
|
col6a3
|
collagen, type VI, alpha 3 |
| chr10_+_33990572 | 0.04 |
ENSDART00000138547
|
fryb
|
furry homolog b (Drosophila) |
| chr23_-_33750307 | 0.03 |
ENSDART00000162772
|
bin2a
|
bridging integrator 2a |
| chr23_-_33750135 | 0.03 |
ENSDART00000187641
|
bin2a
|
bridging integrator 2a |
| chr9_-_3671911 | 0.03 |
ENSDART00000102900
|
sp5a
|
Sp5 transcription factor a |
| chr6_-_53334259 | 0.03 |
ENSDART00000172465
|
gnb1b
|
guanine nucleotide binding protein (G protein), beta polypeptide 1b |
| chr7_-_18508815 | 0.03 |
ENSDART00000173539
|
rgs12a
|
regulator of G protein signaling 12a |
| chr10_-_25561751 | 0.03 |
ENSDART00000147089
|
grik1a
|
glutamate receptor, ionotropic, kainate 1a |
| chr13_-_25196758 | 0.03 |
ENSDART00000184722
|
adka
|
adenosine kinase a |
| chr5_+_59397348 | 0.03 |
ENSDART00000175642
|
clip2
|
CAP-GLY domain containing linker protein 2 |
| chr9_-_2892250 | 0.03 |
ENSDART00000140695
|
cdca7a
|
cell division cycle associated 7a |
| chr21_+_11865972 | 0.02 |
ENSDART00000081676
|
ubap1
|
ubiquitin associated protein 1 |
| chr14_-_48348973 | 0.02 |
ENSDART00000185822
|
CABZ01080056.1
|
|
| chr9_-_2892045 | 0.02 |
ENSDART00000137201
|
cdca7a
|
cell division cycle associated 7a |
| chr18_-_2433011 | 0.02 |
ENSDART00000181922
ENSDART00000193276 |
CR769778.1
|
|
| chr3_-_18737126 | 0.02 |
ENSDART00000055767
|
e4f1
|
E4F transcription factor 1 |
| chr1_-_56213723 | 0.02 |
ENSDART00000142505
ENSDART00000137237 |
si:dkey-76b14.2
|
si:dkey-76b14.2 |
| chr17_+_4368859 | 0.01 |
ENSDART00000055385
|
crls1
|
cardiolipin synthase 1 |
| chr20_+_43379029 | 0.01 |
ENSDART00000142486
ENSDART00000186486 |
unc93a
|
unc-93 homolog A |
| chr13_-_9875538 | 0.01 |
ENSDART00000041609
|
tm9sf3
|
transmembrane 9 superfamily member 3 |
| chr7_+_31879649 | 0.01 |
ENSDART00000099789
|
mybpc3
|
myosin binding protein C, cardiac |
| chr22_+_2239974 | 0.00 |
ENSDART00000141993
|
znf1144
|
zinc finger protein 1144 |
| chr12_-_34713888 | 0.00 |
ENSDART00000153272
|
bahcc1b
|
BAH domain and coiled-coil containing 1b |
| chr7_+_31879986 | 0.00 |
ENSDART00000138491
|
mybpc3
|
myosin binding protein C, cardiac |
Network of associatons between targets according to the STRING database.
First level regulatory network of hoxa11a
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:1901890 | positive regulation of cell junction assembly(GO:1901890) |
| 0.1 | 0.3 | GO:0002926 | tRNA wobble base 5-methoxycarbonylmethyl-2-thiouridine biosynthesis.(GO:0002926) |
| 0.1 | 0.5 | GO:0045721 | negative regulation of gluconeogenesis(GO:0045721) |
| 0.0 | 0.3 | GO:0042769 | DNA damage response, detection of DNA damage(GO:0042769) |
| 0.0 | 0.2 | GO:0030970 | retrograde protein transport, ER to cytosol(GO:0030970) |
| 0.0 | 0.1 | GO:2000055 | positive regulation of Wnt signaling pathway involved in dorsal/ventral axis specification(GO:2000055) |
| 0.0 | 0.3 | GO:0017196 | N-terminal peptidyl-methionine acetylation(GO:0017196) |
| 0.0 | 0.2 | GO:0030579 | ubiquitin-dependent SMAD protein catabolic process(GO:0030579) |
| 0.0 | 0.4 | GO:0048026 | positive regulation of mRNA splicing, via spliceosome(GO:0048026) |
| 0.0 | 0.2 | GO:0050435 | beta-amyloid metabolic process(GO:0050435) |
| 0.0 | 0.1 | GO:0071800 | podosome assembly(GO:0071800) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0016600 | flotillin complex(GO:0016600) |
| 0.1 | 0.3 | GO:0031417 | NatC complex(GO:0031417) |
| 0.0 | 0.2 | GO:0031907 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.0 | 0.3 | GO:0033588 | Elongator holoenzyme complex(GO:0033588) |
| 0.0 | 0.3 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.0 | 0.1 | GO:0005915 | cell-cell adherens junction(GO:0005913) zonula adherens(GO:0005915) |
| 0.0 | 0.4 | GO:0005689 | U12-type spliceosomal complex(GO:0005689) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.2 | GO:0032184 | SUMO polymer binding(GO:0032184) |
| 0.0 | 0.3 | GO:0004596 | peptide alpha-N-acetyltransferase activity(GO:0004596) |
| 0.0 | 0.1 | GO:0045294 | alpha-catenin binding(GO:0045294) |
| 0.0 | 0.0 | GO:0034416 | bisphosphoglycerate phosphatase activity(GO:0034416) |
| 0.0 | 0.0 | GO:0035516 | oxidative DNA demethylase activity(GO:0035516) |
| 0.0 | 0.2 | GO:0032050 | clathrin heavy chain binding(GO:0032050) |
| 0.0 | 0.1 | GO:0004656 | procollagen-proline 4-dioxygenase activity(GO:0004656) |
| 0.0 | 0.1 | GO:0003917 | DNA topoisomerase type I activity(GO:0003917) |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |