Project
PRJNA438478: RNAseq of wild type zebrafish germline
Navigation
Downloads
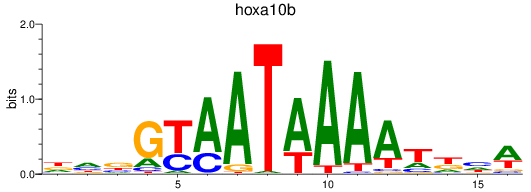
Results for hoxa10b
Z-value: 0.69
Transcription factors associated with hoxa10b
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
hoxa10b
|
ENSDARG00000031337 | homeobox A10b |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| hoxa10b | dr11_v1_chr16_+_20910186_20910186 | -0.36 | 1.4e-01 | Click! |
Activity profile of hoxa10b motif
Sorted Z-values of hoxa10b motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr12_-_25887864 | 1.95 |
ENSDART00000152983
|
si:dkey-193p11.2
|
si:dkey-193p11.2 |
| chr1_-_9485939 | 1.70 |
ENSDART00000157814
|
micall2b
|
mical-like 2b |
| chr11_-_25257045 | 1.63 |
ENSDART00000130477
|
snai1a
|
snail family zinc finger 1a |
| chr7_+_1473929 | 1.57 |
ENSDART00000050687
|
lpcat4
|
lysophosphatidylcholine acyltransferase 4 |
| chr24_-_21090447 | 1.35 |
ENSDART00000136507
ENSDART00000140786 ENSDART00000184841 |
qtrt2
|
queuine tRNA-ribosyltransferase accessory subunit 2 |
| chr2_-_17115256 | 1.32 |
ENSDART00000190488
|
pif1
|
PIF1 5'-to-3' DNA helicase homolog (S. cerevisiae) |
| chr13_-_24717618 | 1.28 |
ENSDART00000172156
|
erlin1
|
ER lipid raft associated 1 |
| chr17_+_50701748 | 1.23 |
ENSDART00000191938
ENSDART00000183220 ENSDART00000049464 |
fermt2
|
fermitin family member 2 |
| chr13_-_24717365 | 1.18 |
ENSDART00000137934
ENSDART00000003922 |
erlin1
|
ER lipid raft associated 1 |
| chr2_+_51818039 | 1.18 |
ENSDART00000170353
|
acvr2bb
|
activin A receptor type 2Bb |
| chr14_+_45028062 | 1.16 |
ENSDART00000184717
ENSDART00000185481 |
ATP8A1
|
ATPase phospholipid transporting 8A1 |
| chr10_+_15340768 | 1.15 |
ENSDART00000046274
ENSDART00000168909 |
trappc13
|
trafficking protein particle complex 13 |
| chr22_-_881725 | 1.12 |
ENSDART00000035514
|
cept1b
|
choline/ethanolamine phosphotransferase 1b |
| chr12_-_4243268 | 0.99 |
ENSDART00000131275
|
zgc:92313
|
zgc:92313 |
| chr11_+_11303458 | 0.98 |
ENSDART00000162486
ENSDART00000160703 |
si:dkey-23f9.4
|
si:dkey-23f9.4 |
| chr12_-_18961289 | 0.95 |
ENSDART00000168405
|
ep300a
|
E1A binding protein p300 a |
| chr14_-_16082806 | 0.95 |
ENSDART00000165656
|
mxd3
|
MAX dimerization protein 3 |
| chr15_-_43978141 | 0.94 |
ENSDART00000041249
|
chordc1a
|
cysteine and histidine-rich domain (CHORD) containing 1a |
| chr21_-_14878220 | 0.92 |
ENSDART00000131237
|
ulk1b
|
unc-51 like autophagy activating kinase 1 |
| chr3_+_39579393 | 0.87 |
ENSDART00000055170
|
cln3
|
ceroid-lipofuscinosis, neuronal 3 |
| chr19_-_47571456 | 0.86 |
ENSDART00000158071
ENSDART00000165841 |
rrm2
|
ribonucleotide reductase M2 polypeptide |
| chr16_-_44945224 | 0.85 |
ENSDART00000156921
|
ncam3
|
neural cell adhesion molecule 3 |
| chr2_+_22409249 | 0.85 |
ENSDART00000182915
|
zgc:56628
|
zgc:56628 |
| chr12_-_10512911 | 0.85 |
ENSDART00000124562
ENSDART00000106163 |
zgc:152977
|
zgc:152977 |
| chr6_-_7107868 | 0.84 |
ENSDART00000171123
|
nhej1
|
nonhomologous end-joining factor 1 |
| chr22_-_20950448 | 0.84 |
ENSDART00000002029
|
fkbp8
|
FK506 binding protein 8 |
| chr22_+_15959844 | 0.81 |
ENSDART00000182201
|
stil
|
scl/tal1 interrupting locus |
| chr10_-_31805923 | 0.81 |
ENSDART00000077785
|
vps26bl
|
vacuolar protein sorting 26 homolog B, like |
| chr13_+_8892784 | 0.80 |
ENSDART00000075054
ENSDART00000143705 |
thada
|
thyroid adenoma associated |
| chr17_-_30652738 | 0.79 |
ENSDART00000154960
|
sh3yl1
|
SH3 and SYLF domain containing 1 |
| chr21_+_43178831 | 0.78 |
ENSDART00000151512
|
aff4
|
AF4/FMR2 family, member 4 |
| chr14_+_22132388 | 0.77 |
ENSDART00000109065
|
ccng1
|
cyclin G1 |
| chr11_-_25257595 | 0.76 |
ENSDART00000123567
|
snai1a
|
snail family zinc finger 1a |
| chr12_-_18961491 | 0.75 |
ENSDART00000172574
|
ep300a
|
E1A binding protein p300 a |
| chr13_-_17860307 | 0.75 |
ENSDART00000135920
ENSDART00000054579 |
march8
|
membrane-associated ring finger (C3HC4) 8 |
| chr8_+_2656231 | 0.74 |
ENSDART00000160833
|
fam102aa
|
family with sequence similarity 102, member Aa |
| chr25_+_3549841 | 0.73 |
ENSDART00000164030
|
ccdc77
|
coiled-coil domain containing 77 |
| chr20_-_29499363 | 0.73 |
ENSDART00000152889
ENSDART00000153252 ENSDART00000170972 ENSDART00000166420 ENSDART00000163079 |
rrm2
|
ribonucleotide reductase M2 polypeptide |
| chr5_-_65121747 | 0.71 |
ENSDART00000165556
|
tor2a
|
torsin family 2, member A |
| chr15_+_22390076 | 0.71 |
ENSDART00000183764
|
oafa
|
OAF homolog a (Drosophila) |
| chr15_-_1843831 | 0.70 |
ENSDART00000156718
ENSDART00000154175 |
taf15
|
TAF15 RNA polymerase II, TATA box binding protein (TBP)-associated factor |
| chr14_-_50028428 | 0.69 |
ENSDART00000163543
ENSDART00000168510 |
abhd18
|
abhydrolase domain containing 18 |
| chr1_-_14506759 | 0.69 |
ENSDART00000057044
|
si:dkey-194g4.1
|
si:dkey-194g4.1 |
| chr13_+_7575563 | 0.68 |
ENSDART00000146592
|
gbf1
|
golgi brefeldin A resistant guanine nucleotide exchange factor 1 |
| chr8_-_18899427 | 0.68 |
ENSDART00000079840
|
rorca
|
RAR-related orphan receptor C a |
| chr6_-_9282080 | 0.68 |
ENSDART00000159506
|
ccdc14
|
coiled-coil domain containing 14 |
| chr2_-_10877765 | 0.67 |
ENSDART00000100607
|
cdc7
|
cell division cycle 7 homolog (S. cerevisiae) |
| chr13_+_33462232 | 0.67 |
ENSDART00000177841
|
zgc:136302
|
zgc:136302 |
| chr19_-_47571797 | 0.66 |
ENSDART00000166180
ENSDART00000168134 |
rrm2
|
ribonucleotide reductase M2 polypeptide |
| chr12_+_19199735 | 0.65 |
ENSDART00000066393
|
pdap1a
|
pdgfa associated protein 1a |
| chr16_+_35916371 | 0.65 |
ENSDART00000167208
|
sh3d21
|
SH3 domain containing 21 |
| chr8_-_11324143 | 0.65 |
ENSDART00000008215
|
pip5k1bb
|
phosphatidylinositol-4-phosphate 5-kinase, type I, beta b |
| chr12_-_11560794 | 0.65 |
ENSDART00000149098
ENSDART00000169975 |
plekha1b
|
pleckstrin homology domain containing, family A (phosphoinositide binding specific) member 1b |
| chr23_-_29751730 | 0.64 |
ENSDART00000056865
|
ctnnbip1
|
catenin, beta interacting protein 1 |
| chr9_-_36924388 | 0.63 |
ENSDART00000059756
|
ralba
|
v-ral simian leukemia viral oncogene homolog Ba (ras related) |
| chr13_+_9559461 | 0.63 |
ENSDART00000047740
|
wdr32
|
WD repeat domain 32 |
| chr18_-_39288894 | 0.63 |
ENSDART00000186216
|
mapk6
|
mitogen-activated protein kinase 6 |
| chr8_+_35964482 | 0.63 |
ENSDART00000129357
ENSDART00000154953 |
glt1d1
|
glycosyltransferase 1 domain containing 1 |
| chr21_-_14830353 | 0.62 |
ENSDART00000134687
|
pus1
|
pseudouridylate synthase 1 |
| chr22_-_12337781 | 0.62 |
ENSDART00000188357
ENSDART00000123574 |
zranb3
|
zinc finger, RAN-binding domain containing 3 |
| chr22_+_23430688 | 0.61 |
ENSDART00000160457
|
dennd1b
|
DENN/MADD domain containing 1B |
| chr2_-_40890004 | 0.61 |
ENSDART00000191746
|
uggt1
|
UDP-glucose glycoprotein glucosyltransferase 1 |
| chr15_-_1844048 | 0.60 |
ENSDART00000102410
|
taf15
|
TAF15 RNA polymerase II, TATA box binding protein (TBP)-associated factor |
| chr12_+_33361948 | 0.60 |
ENSDART00000124982
|
fasn
|
fatty acid synthase |
| chr2_-_40889465 | 0.60 |
ENSDART00000192631
ENSDART00000180824 |
uggt1
|
UDP-glucose glycoprotein glucosyltransferase 1 |
| chr2_+_49805892 | 0.60 |
ENSDART00000056248
|
wdr48b
|
WD repeat domain 48b |
| chr19_+_27970292 | 0.60 |
ENSDART00000103887
ENSDART00000145549 |
med10
|
mediator complex subunit 10 |
| chr2_-_37458527 | 0.59 |
ENSDART00000146820
|
si:dkey-57k2.7
|
si:dkey-57k2.7 |
| chr22_+_5574952 | 0.59 |
ENSDART00000171774
|
zgc:171566
|
zgc:171566 |
| chr1_+_45663727 | 0.59 |
ENSDART00000038574
ENSDART00000141144 ENSDART00000149565 |
trappc5
|
trafficking protein particle complex 5 |
| chr1_+_12394205 | 0.58 |
ENSDART00000138622
ENSDART00000136421 ENSDART00000139440 ENSDART00000184296 ENSDART00000008127 |
zgc:77739
|
zgc:77739 |
| chr1_+_2321384 | 0.58 |
ENSDART00000157662
|
farp1
|
FERM, RhoGEF (ARHGEF) and pleckstrin domain protein 1 (chondrocyte-derived) |
| chr12_+_3262564 | 0.56 |
ENSDART00000184264
|
tmem101
|
transmembrane protein 101 |
| chr15_-_5901514 | 0.56 |
ENSDART00000155252
|
si:ch73-281n10.2
|
si:ch73-281n10.2 |
| chr12_-_20409794 | 0.55 |
ENSDART00000077936
|
lcmt1
|
leucine carboxyl methyltransferase 1 |
| chr23_+_45200481 | 0.55 |
ENSDART00000004357
ENSDART00000111126 ENSDART00000193560 ENSDART00000190476 |
psip1b
|
PC4 and SFRS1 interacting protein 1b |
| chr19_+_7424347 | 0.54 |
ENSDART00000004622
|
sf3b4
|
splicing factor 3b, subunit 4 |
| chr14_+_22467672 | 0.54 |
ENSDART00000079409
ENSDART00000136597 |
nudt22
|
nudix (nucleoside diphosphate linked moiety X)-type motif 22 |
| chr20_-_34670236 | 0.54 |
ENSDART00000033325
|
slc25a24
|
solute carrier family 25 (mitochondrial carrier; phosphate carrier), member 24 |
| chr3_-_60589292 | 0.53 |
ENSDART00000157822
|
jmjd6
|
jumonji domain containing 6 |
| chr7_-_30280934 | 0.53 |
ENSDART00000126741
|
serf2
|
small EDRK-rich factor 2 |
| chr13_-_25745089 | 0.52 |
ENSDART00000189333
ENSDART00000147420 |
sgpl1
|
sphingosine-1-phosphate lyase 1 |
| chr13_+_13945218 | 0.52 |
ENSDART00000089501
ENSDART00000142997 |
eif2ak3
|
eukaryotic translation initiation factor 2-alpha kinase 3 |
| chr21_-_11970199 | 0.52 |
ENSDART00000114524
|
nop56
|
NOP56 ribonucleoprotein homolog |
| chr22_+_1953096 | 0.51 |
ENSDART00000166630
ENSDART00000186234 |
si:dkey-15h8.16
|
si:dkey-15h8.16 |
| chr8_-_31416403 | 0.51 |
ENSDART00000098912
|
znf131
|
zinc finger protein 131 |
| chr23_-_24253200 | 0.51 |
ENSDART00000114840
|
zbtb17
|
zinc finger and BTB domain containing 17 |
| chr12_+_13205955 | 0.51 |
ENSDART00000092906
|
ppp1cab
|
protein phosphatase 1, catalytic subunit, alpha isozyme b |
| chr18_+_15876385 | 0.51 |
ENSDART00000142527
|
eea1
|
early endosome antigen 1 |
| chr6_+_40554551 | 0.50 |
ENSDART00000017859
ENSDART00000155928 |
ddi2
|
DNA-damage inducible protein 2 |
| chr16_-_32671998 | 0.50 |
ENSDART00000161395
|
pnisr
|
PNN-interacting serine/arginine-rich protein |
| chr20_+_26943072 | 0.49 |
ENSDART00000153215
|
cdca4
|
cell division cycle associated 4 |
| chr20_-_23219964 | 0.49 |
ENSDART00000144933
|
dcun1d4
|
DCN1, defective in cullin neddylation 1, domain containing 4 (S. cerevisiae) |
| chr7_+_17907883 | 0.49 |
ENSDART00000147564
|
mta2
|
metastasis associated 1 family, member 2 |
| chr16_+_46401756 | 0.48 |
ENSDART00000147370
ENSDART00000144000 |
rpz2
|
rapunzel 2 |
| chr3_-_4501026 | 0.48 |
ENSDART00000163052
|
zgc:162198
|
zgc:162198 |
| chr2_-_50272077 | 0.47 |
ENSDART00000127623
|
cul1a
|
cullin 1a |
| chr11_-_25461336 | 0.47 |
ENSDART00000014945
|
hcfc1a
|
host cell factor C1a |
| chr23_+_20431140 | 0.47 |
ENSDART00000193950
|
slc35c2
|
solute carrier family 35 (GDP-fucose transporter), member C2 |
| chr15_-_26931541 | 0.47 |
ENSDART00000027563
|
ccdc9
|
coiled-coil domain containing 9 |
| chr2_+_49457449 | 0.46 |
ENSDART00000185470
|
sh3gl1a
|
SH3-domain GRB2-like 1a |
| chr2_-_49627099 | 0.46 |
ENSDART00000083710
ENSDART00000147421 |
cactin
|
cactin |
| chr13_+_33268657 | 0.46 |
ENSDART00000002095
|
tmem39b
|
transmembrane protein 39B |
| chr10_-_20588787 | 0.45 |
ENSDART00000138045
ENSDART00000181885 ENSDART00000091115 |
nsd3
|
nuclear receptor binding SET domain protein 3 |
| chr21_+_43702016 | 0.44 |
ENSDART00000017176
|
dkc1
|
dyskeratosis congenita 1, dyskerin |
| chr13_+_15580758 | 0.43 |
ENSDART00000087194
ENSDART00000013525 |
mark3a
|
MAP/microtubule affinity-regulating kinase 3a |
| chr22_+_30195257 | 0.42 |
ENSDART00000027803
ENSDART00000172496 |
add3a
|
adducin 3 (gamma) a |
| chr25_+_3549401 | 0.42 |
ENSDART00000166312
|
ccdc77
|
coiled-coil domain containing 77 |
| chr3_-_45778123 | 0.41 |
ENSDART00000146211
|
h3f3b.1
|
H3 histone, family 3B.1 |
| chr17_+_17764979 | 0.40 |
ENSDART00000105013
|
alkbh1
|
alkB homolog 1, histone H2A dioxygenase |
| chr24_-_3407507 | 0.40 |
ENSDART00000132648
|
nck1b
|
NCK adaptor protein 1b |
| chr25_-_29074064 | 0.40 |
ENSDART00000165603
|
arid3b
|
AT rich interactive domain 3B (BRIGHT-like) |
| chr5_-_68074592 | 0.40 |
ENSDART00000165052
ENSDART00000018792 |
spag7
|
sperm associated antigen 7 |
| chr14_-_238931 | 0.39 |
ENSDART00000161122
|
bod1l1
|
biorientation of chromosomes in cell division 1-like 1 |
| chr20_-_36416922 | 0.39 |
ENSDART00000019145
|
lbr
|
lamin B receptor |
| chr19_-_30447611 | 0.39 |
ENSDART00000073705
ENSDART00000048977 ENSDART00000191237 |
abcf1
|
ATP-binding cassette, sub-family F (GCN20), member 1 |
| chr14_-_47963115 | 0.38 |
ENSDART00000003826
|
rapgef2
|
Rap guanine nucleotide exchange factor (GEF) 2 |
| chr7_-_8712148 | 0.38 |
ENSDART00000065488
|
tex261
|
testis expressed 261 |
| chr1_+_41596099 | 0.38 |
ENSDART00000111367
|
si:dkey-56e3.3
|
si:dkey-56e3.3 |
| chr16_-_32671782 | 0.38 |
ENSDART00000123980
|
pnisr
|
PNN-interacting serine/arginine-rich protein |
| chr13_+_1131748 | 0.37 |
ENSDART00000054318
|
wdr92
|
WD repeat domain 92 |
| chr7_-_7692992 | 0.36 |
ENSDART00000192619
|
aadat
|
aminoadipate aminotransferase |
| chr17_+_15535501 | 0.36 |
ENSDART00000002932
|
marcksb
|
myristoylated alanine-rich protein kinase C substrate b |
| chr6_-_10728921 | 0.36 |
ENSDART00000151484
|
sp3b
|
Sp3b transcription factor |
| chr9_+_30090656 | 0.36 |
ENSDART00000102981
|
col8a1a
|
collagen, type VIII, alpha 1a |
| chr7_-_7692723 | 0.35 |
ENSDART00000183352
|
aadat
|
aminoadipate aminotransferase |
| chr24_-_15159658 | 0.35 |
ENSDART00000142473
|
rttn
|
rotatin |
| chr20_-_34801181 | 0.35 |
ENSDART00000048375
ENSDART00000132426 |
stmn4
|
stathmin-like 4 |
| chr17_-_25630822 | 0.35 |
ENSDART00000126201
ENSDART00000105503 ENSDART00000151878 |
rab3gap2
|
RAB3 GTPase activating protein subunit 2 (non-catalytic) |
| chr10_+_6013076 | 0.34 |
ENSDART00000167613
ENSDART00000159216 |
hmgcs1
|
3-hydroxy-3-methylglutaryl-CoA synthase 1 (soluble) |
| chr3_+_31093455 | 0.34 |
ENSDART00000153074
|
si:dkey-66i24.9
|
si:dkey-66i24.9 |
| chr22_+_1947494 | 0.34 |
ENSDART00000159121
|
si:dkey-15h8.15
|
si:dkey-15h8.15 |
| chr5_-_57289872 | 0.34 |
ENSDART00000189893
ENSDART00000050957 |
fer
|
fer (fps/fes related) tyrosine kinase |
| chr20_-_45709990 | 0.33 |
ENSDART00000027482
|
gpcpd1
|
glycerophosphocholine phosphodiesterase 1 |
| chr14_+_26439227 | 0.33 |
ENSDART00000054183
|
gpr137
|
G protein-coupled receptor 137 |
| chr12_-_33817114 | 0.32 |
ENSDART00000161265
|
twnk
|
twinkle mtDNA helicase |
| chr12_+_27285994 | 0.32 |
ENSDART00000087204
|
dusp3a
|
dual specificity phosphatase 3a |
| chr24_-_23784701 | 0.32 |
ENSDART00000090368
|
sgk3
|
serum/glucocorticoid regulated kinase family, member 3 |
| chr1_-_41374117 | 0.32 |
ENSDART00000074777
|
htt
|
huntingtin |
| chr6_-_30485009 | 0.31 |
ENSDART00000025698
|
zgc:153311
|
zgc:153311 |
| chr4_-_11053543 | 0.31 |
ENSDART00000067262
|
mettl25
|
methyltransferase like 25 |
| chr5_+_20453874 | 0.31 |
ENSDART00000124545
ENSDART00000008402 |
sart3
|
squamous cell carcinoma antigen recognized by T cells 3 |
| chr4_-_17353972 | 0.31 |
ENSDART00000041529
|
parpbp
|
PARP1 binding protein |
| chr2_+_21048661 | 0.30 |
ENSDART00000156876
|
rreb1b
|
ras responsive element binding protein 1b |
| chr5_+_27137473 | 0.30 |
ENSDART00000181833
|
unc5db
|
unc-5 netrin receptor Db |
| chr13_+_29292011 | 0.30 |
ENSDART00000115023
|
parga
|
poly (ADP-ribose) glycohydrolase a |
| chr25_+_9013342 | 0.29 |
ENSDART00000154207
ENSDART00000153705 |
im:7145024
|
im:7145024 |
| chr15_-_20731638 | 0.29 |
ENSDART00000170616
|
tpst1
|
tyrosylprotein sulfotransferase 1 |
| chr23_-_29505645 | 0.29 |
ENSDART00000146458
|
kif1b
|
kinesin family member 1B |
| chr10_-_33297864 | 0.29 |
ENSDART00000163360
|
PRDM15
|
PR/SET domain 15 |
| chr3_+_60589157 | 0.29 |
ENSDART00000165367
|
mettl23
|
methyltransferase like 23 |
| chr18_+_6857071 | 0.29 |
ENSDART00000018735
ENSDART00000181969 |
dnaja2l
|
DnaJ (Hsp40) homolog, subfamily A, member 2, like |
| chr14_+_45608576 | 0.29 |
ENSDART00000173248
|
si:ch211-276i12.11
|
si:ch211-276i12.11 |
| chr1_+_27808916 | 0.28 |
ENSDART00000102335
|
pspc1
|
paraspeckle component 1 |
| chr20_-_25709247 | 0.28 |
ENSDART00000146711
|
si:dkeyp-117h8.2
|
si:dkeyp-117h8.2 |
| chr21_-_26691959 | 0.28 |
ENSDART00000149702
ENSDART00000149840 |
bscl2
|
Berardinelli-Seip congenital lipodystrophy 2 (seipin) |
| chr25_+_20089986 | 0.28 |
ENSDART00000143441
ENSDART00000184073 |
tnni4b.2
|
troponin I4b, tandem duplicate 2 |
| chr15_-_47857687 | 0.27 |
ENSDART00000098982
ENSDART00000151594 |
h3f3b.1
|
H3 histone, family 3B.1 |
| chr5_+_37032038 | 0.27 |
ENSDART00000045036
|
nfkbib
|
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, beta |
| chr11_-_26362294 | 0.27 |
ENSDART00000184654
ENSDART00000115037 |
foxj3
|
forkhead box J3 |
| chr9_-_40765868 | 0.27 |
ENSDART00000138634
|
abca12
|
ATP-binding cassette, sub-family A (ABC1), member 12 |
| chr4_+_23125689 | 0.26 |
ENSDART00000077854
|
mdm2
|
MDM2 oncogene, E3 ubiquitin protein ligase |
| chr20_+_32552912 | 0.26 |
ENSDART00000009691
|
scml4
|
Scm polycomb group protein like 4 |
| chr6_+_45692026 | 0.26 |
ENSDART00000164759
|
cntn4
|
contactin 4 |
| chr24_+_10202718 | 0.26 |
ENSDART00000126668
|
pou6f2
|
POU class 6 homeobox 2 |
| chr5_-_37881345 | 0.26 |
ENSDART00000084819
|
arhgap35b
|
Rho GTPase activating protein 35b |
| chr2_+_54086436 | 0.26 |
ENSDART00000174581
|
CU179656.1
|
|
| chr16_+_52966812 | 0.25 |
ENSDART00000148435
ENSDART00000049099 ENSDART00000150117 |
trip13
|
thyroid hormone receptor interactor 13 |
| chr17_+_43595692 | 0.25 |
ENSDART00000156271
|
cfap99
|
cilia and flagella associated protein 99 |
| chr16_+_54829293 | 0.25 |
ENSDART00000024729
|
pabpc1a
|
poly(A) binding protein, cytoplasmic 1a |
| chr12_+_23912074 | 0.25 |
ENSDART00000152864
|
svila
|
supervillin a |
| chr25_+_28823952 | 0.24 |
ENSDART00000067072
|
nfybb
|
nuclear transcription factor Y, beta b |
| chr1_+_52392511 | 0.24 |
ENSDART00000144025
|
si:ch211-217k17.8
|
si:ch211-217k17.8 |
| chr15_-_46718759 | 0.24 |
ENSDART00000085926
ENSDART00000154577 |
zgc:162872
|
zgc:162872 |
| chr11_+_31730680 | 0.24 |
ENSDART00000145497
|
diaph3
|
diaphanous-related formin 3 |
| chr2_+_25315591 | 0.24 |
ENSDART00000161386
|
fndc3ba
|
fibronectin type III domain containing 3Ba |
| chr6_+_8315050 | 0.24 |
ENSDART00000189987
|
gcdha
|
glutaryl-CoA dehydrogenase a |
| chr16_-_52966811 | 0.23 |
ENSDART00000153758
ENSDART00000009256 |
brd9
|
bromodomain containing 9 |
| chr14_+_10625112 | 0.23 |
ENSDART00000143377
ENSDART00000136480 |
nup62l
|
nucleoporin 62 like |
| chr25_-_14433503 | 0.23 |
ENSDART00000103957
|
exoc3l1
|
exocyst complex component 3-like 1 |
| chr22_+_10606573 | 0.22 |
ENSDART00000192638
|
rad54l2
|
RAD54 like 2 |
| chr25_+_22571475 | 0.22 |
ENSDART00000161559
|
stra6
|
stimulated by retinoic acid 6 |
| chr8_-_10947205 | 0.22 |
ENSDART00000164467
|
pqlc2
|
PQ loop repeat containing 2 |
| chr24_+_14581864 | 0.22 |
ENSDART00000134536
|
thtpa
|
thiamine triphosphatase |
| chr1_-_55873178 | 0.22 |
ENSDART00000019936
|
prkacab
|
protein kinase, cAMP-dependent, catalytic, alpha, genome duplicate b |
| chr23_-_21215311 | 0.22 |
ENSDART00000112424
|
megf6a
|
multiple EGF-like-domains 6a |
| chr8_+_3820134 | 0.22 |
ENSDART00000122454
|
citb
|
citron rho-interacting serine/threonine kinase b |
| chr19_+_40069524 | 0.21 |
ENSDART00000151365
ENSDART00000140926 |
zmym4
|
zinc finger, MYM-type 4 |
| chr12_+_32368574 | 0.21 |
ENSDART00000086389
|
ANKFN1
|
si:ch211-277e21.2 |
| chr4_-_22749553 | 0.20 |
ENSDART00000040033
|
nup107
|
nucleoporin 107 |
| chr2_+_19163965 | 0.19 |
ENSDART00000166073
|
elovl1a
|
ELOVL fatty acid elongase 1a |
| chr12_-_10476448 | 0.19 |
ENSDART00000106172
|
rac1a
|
Rac family small GTPase 1a |
| chr13_+_4888351 | 0.19 |
ENSDART00000145940
|
micu1
|
mitochondrial calcium uptake 1 |
| chr18_+_21951551 | 0.19 |
ENSDART00000146261
|
ranbp10
|
RAN binding protein 10 |
| chr3_-_5277041 | 0.19 |
ENSDART00000046995
|
txn2
|
thioredoxin 2 |
| chr23_-_21215520 | 0.19 |
ENSDART00000143206
|
megf6a
|
multiple EGF-like-domains 6a |
| chr2_+_35733335 | 0.19 |
ENSDART00000113489
|
rasal2
|
RAS protein activator like 2 |
| chr5_+_50913034 | 0.18 |
ENSDART00000149787
|
col4a3bpa
|
collagen, type IV, alpha 3 (Goodpasture antigen) binding protein a |
| chr2_+_20967673 | 0.18 |
ENSDART00000057174
|
arpc5a
|
actin related protein 2/3 complex, subunit 5A |
| chr15_+_29727799 | 0.18 |
ENSDART00000182006
|
zgc:153372
|
zgc:153372 |
Network of associatons between targets according to the STRING database.
First level regulatory network of hoxa10b
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.9 | GO:1903003 | positive regulation of protein deubiquitination(GO:1903003) |
| 0.2 | 1.1 | GO:1990481 | mRNA pseudouridine synthesis(GO:1990481) |
| 0.2 | 2.3 | GO:0009263 | deoxyribonucleotide biosynthetic process(GO:0009263) |
| 0.2 | 0.6 | GO:1905072 | cardiac jelly development(GO:1905072) |
| 0.2 | 2.5 | GO:0071501 | response to sterol depletion(GO:0006991) SREBP signaling pathway(GO:0032933) cellular response to sterol depletion(GO:0071501) |
| 0.2 | 0.7 | GO:0002495 | antigen processing and presentation of peptide antigen via MHC class II(GO:0002495) |
| 0.2 | 0.5 | GO:2000434 | regulation of protein neddylation(GO:2000434) |
| 0.1 | 0.6 | GO:0046900 | tetrahydrofolylpolyglutamate metabolic process(GO:0046900) |
| 0.1 | 1.6 | GO:0000002 | mitochondrial genome maintenance(GO:0000002) |
| 0.1 | 0.5 | GO:0017185 | peptidyl-lysine hydroxylation(GO:0017185) |
| 0.1 | 0.9 | GO:0015809 | arginine transport(GO:0015809) |
| 0.1 | 0.3 | GO:1902767 | farnesyl diphosphate biosynthetic process, mevalonate pathway(GO:0010142) isoprenoid biosynthetic process via mevalonate(GO:1902767) |
| 0.1 | 2.4 | GO:0008078 | mesodermal cell migration(GO:0008078) |
| 0.1 | 0.3 | GO:0048917 | posterior lateral line ganglion development(GO:0048917) |
| 0.1 | 1.1 | GO:0006646 | phosphatidylethanolamine biosynthetic process(GO:0006646) |
| 0.1 | 0.8 | GO:0071539 | protein localization to centrosome(GO:0071539) |
| 0.1 | 0.6 | GO:0048478 | replication fork protection(GO:0048478) |
| 0.1 | 0.3 | GO:0038093 | Fc receptor signaling pathway(GO:0038093) |
| 0.1 | 0.4 | GO:0035513 | oxidative RNA demethylation(GO:0035513) |
| 0.1 | 1.2 | GO:0071712 | ER-associated misfolded protein catabolic process(GO:0071712) |
| 0.1 | 0.2 | GO:0000973 | posttranscriptional tethering of RNA polymerase II gene DNA at nuclear periphery(GO:0000973) |
| 0.1 | 0.3 | GO:0033313 | meiotic cell cycle checkpoint(GO:0033313) |
| 0.1 | 0.2 | GO:0018872 | arsonoacetate metabolic process(GO:0018872) |
| 0.1 | 0.4 | GO:0010457 | centriole-centriole cohesion(GO:0010457) |
| 0.1 | 0.3 | GO:1904036 | negative regulation of epithelial cell apoptotic process(GO:1904036) |
| 0.1 | 0.3 | GO:2000042 | negative regulation of double-strand break repair via homologous recombination(GO:2000042) |
| 0.1 | 0.6 | GO:0018410 | C-terminal protein amino acid modification(GO:0018410) |
| 0.0 | 0.4 | GO:0032515 | negative regulation of phosphoprotein phosphatase activity(GO:0032515) |
| 0.0 | 0.3 | GO:0034389 | lipid particle organization(GO:0034389) |
| 0.0 | 0.3 | GO:0001839 | neural plate morphogenesis(GO:0001839) |
| 0.0 | 0.2 | GO:0071938 | vitamin A transport(GO:0071938) vitamin A import(GO:0071939) |
| 0.0 | 0.1 | GO:1902626 | mature ribosome assembly(GO:0042256) assembly of large subunit precursor of preribosome(GO:1902626) |
| 0.0 | 1.2 | GO:0055008 | cardiac muscle tissue morphogenesis(GO:0055008) |
| 0.0 | 0.7 | GO:0000727 | double-strand break repair via break-induced replication(GO:0000727) |
| 0.0 | 0.2 | GO:0051561 | positive regulation of mitochondrial calcium ion concentration(GO:0051561) |
| 0.0 | 0.2 | GO:0042723 | thiamine metabolic process(GO:0006772) thiamine-containing compound metabolic process(GO:0042723) |
| 0.0 | 0.3 | GO:0006478 | protein sulfation(GO:0006477) peptidyl-tyrosine sulfation(GO:0006478) |
| 0.0 | 0.4 | GO:0050714 | positive regulation of protein secretion(GO:0050714) |
| 0.0 | 0.8 | GO:1990798 | pancreas regeneration(GO:1990798) |
| 0.0 | 0.3 | GO:0048730 | epidermis morphogenesis(GO:0048730) |
| 0.0 | 0.5 | GO:0048096 | chromatin-mediated maintenance of transcription(GO:0048096) |
| 0.0 | 0.5 | GO:0045824 | negative regulation of innate immune response(GO:0045824) |
| 0.0 | 1.7 | GO:0036269 | swimming behavior(GO:0036269) |
| 0.0 | 0.4 | GO:0014028 | notochord formation(GO:0014028) |
| 0.0 | 0.5 | GO:2000404 | regulation of T cell migration(GO:2000404) |
| 0.0 | 0.2 | GO:0090200 | regulation of release of cytochrome c from mitochondria(GO:0090199) positive regulation of release of cytochrome c from mitochondria(GO:0090200) |
| 0.0 | 0.5 | GO:0048026 | positive regulation of mRNA splicing, via spliceosome(GO:0048026) |
| 0.0 | 0.8 | GO:0006303 | double-strand break repair via nonhomologous end joining(GO:0006303) |
| 0.0 | 0.2 | GO:0033539 | fatty acid beta-oxidation using acyl-CoA dehydrogenase(GO:0033539) |
| 0.0 | 0.1 | GO:1901166 | neural crest cell migration involved in autonomic nervous system development(GO:1901166) |
| 0.0 | 0.3 | GO:1904263 | positive regulation of TORC1 signaling(GO:1904263) |
| 0.0 | 0.9 | GO:0051298 | centrosome duplication(GO:0051298) |
| 0.0 | 0.7 | GO:0051085 | chaperone mediated protein folding requiring cofactor(GO:0051085) |
| 0.0 | 1.2 | GO:0032924 | activin receptor signaling pathway(GO:0032924) |
| 0.0 | 0.1 | GO:0006824 | cobalt ion transport(GO:0006824) cobalamin transport(GO:0015889) |
| 0.0 | 0.9 | GO:0006998 | nuclear envelope organization(GO:0006998) |
| 0.0 | 0.2 | GO:0018904 | glycerol ether metabolic process(GO:0006662) ether metabolic process(GO:0018904) |
| 0.0 | 0.2 | GO:0051601 | exocyst localization(GO:0051601) |
| 0.0 | 0.5 | GO:0038061 | NIK/NF-kappaB signaling(GO:0038061) |
| 0.0 | 0.6 | GO:0003094 | glomerular filtration(GO:0003094) |
| 0.0 | 1.4 | GO:0033339 | pectoral fin development(GO:0033339) |
| 0.0 | 0.8 | GO:0042147 | retrograde transport, endosome to Golgi(GO:0042147) |
| 0.0 | 0.4 | GO:0006695 | cholesterol biosynthetic process(GO:0006695) secondary alcohol biosynthetic process(GO:1902653) |
| 0.0 | 0.7 | GO:0032012 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.0 | 0.8 | GO:1904029 | regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0000079) regulation of cyclin-dependent protein kinase activity(GO:1904029) |
| 0.0 | 1.0 | GO:0071241 | cellular response to inorganic substance(GO:0071241) |
| 0.0 | 0.1 | GO:2000290 | smoothened signaling pathway involved in ventral spinal cord patterning(GO:0021910) regulation of myotome development(GO:2000290) |
| 0.0 | 0.7 | GO:0051016 | barbed-end actin filament capping(GO:0051016) |
| 0.0 | 0.3 | GO:0046475 | glycerophospholipid catabolic process(GO:0046475) |
| 0.0 | 0.6 | GO:0032456 | endocytic recycling(GO:0032456) |
| 0.0 | 1.1 | GO:0006400 | tRNA modification(GO:0006400) |
| 0.0 | 0.5 | GO:0046466 | sphingolipid catabolic process(GO:0030149) membrane lipid catabolic process(GO:0046466) |
| 0.0 | 0.1 | GO:0070979 | protein K11-linked ubiquitination(GO:0070979) |
| 0.0 | 0.1 | GO:0019730 | antimicrobial humoral response(GO:0019730) |
| 0.0 | 0.2 | GO:0010737 | protein kinase A signaling(GO:0010737) |
| 0.0 | 0.1 | GO:0090594 | nitric oxide mediated signal transduction(GO:0007263) regulation of cGMP metabolic process(GO:0030823) positive regulation of cGMP metabolic process(GO:0030825) regulation of cGMP biosynthetic process(GO:0030826) positive regulation of cGMP biosynthetic process(GO:0030828) regulation of guanylate cyclase activity(GO:0031282) positive regulation of guanylate cyclase activity(GO:0031284) inflammatory response to wounding(GO:0090594) |
| 0.0 | 0.3 | GO:0007340 | acrosome reaction(GO:0007340) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 1.7 | GO:1990072 | TRAPPIII protein complex(GO:1990072) |
| 0.2 | 1.2 | GO:0031258 | lamellipodium membrane(GO:0031258) |
| 0.2 | 0.8 | GO:0032807 | DNA ligase IV complex(GO:0032807) |
| 0.1 | 0.4 | GO:0072588 | box H/ACA snoRNP complex(GO:0031429) box H/ACA RNP complex(GO:0072588) |
| 0.1 | 0.8 | GO:0032783 | ELL-EAF complex(GO:0032783) |
| 0.1 | 0.6 | GO:0030877 | beta-catenin destruction complex(GO:0030877) |
| 0.1 | 0.5 | GO:0031428 | box C/D snoRNP complex(GO:0031428) |
| 0.1 | 1.2 | GO:0048179 | activin receptor complex(GO:0048179) |
| 0.1 | 0.8 | GO:0030904 | retromer complex(GO:0030904) |
| 0.0 | 0.9 | GO:0034993 | microtubule organizing center attachment site(GO:0034992) LINC complex(GO:0034993) |
| 0.0 | 1.9 | GO:0005657 | replication fork(GO:0005657) |
| 0.0 | 1.9 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.0 | 0.4 | GO:0000940 | condensed chromosome outer kinetochore(GO:0000940) |
| 0.0 | 2.1 | GO:0005814 | centriole(GO:0005814) |
| 0.0 | 0.5 | GO:0048188 | Set1C/COMPASS complex(GO:0048188) |
| 0.0 | 0.5 | GO:0016581 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) |
| 0.0 | 0.2 | GO:0044613 | nuclear pore central transport channel(GO:0044613) |
| 0.0 | 0.5 | GO:0005689 | U12-type spliceosomal complex(GO:0005689) |
| 0.0 | 0.1 | GO:0016600 | flotillin complex(GO:0016600) |
| 0.0 | 0.2 | GO:0016602 | CCAAT-binding factor complex(GO:0016602) |
| 0.0 | 0.9 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.0 | 0.3 | GO:0015030 | Cajal body(GO:0015030) |
| 0.0 | 1.7 | GO:0000123 | histone acetyltransferase complex(GO:0000123) |
| 0.0 | 0.8 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.0 | 0.5 | GO:0080008 | Cul4-RING E3 ubiquitin ligase complex(GO:0080008) |
| 0.0 | 0.2 | GO:1990246 | uniplex complex(GO:1990246) |
| 0.0 | 0.3 | GO:0016363 | nuclear matrix(GO:0016363) |
| 0.0 | 0.9 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.0 | 1.5 | GO:0016607 | nuclear speck(GO:0016607) |
| 0.0 | 1.4 | GO:0005741 | mitochondrial outer membrane(GO:0005741) |
| 0.0 | 0.6 | GO:0016592 | mediator complex(GO:0016592) |
| 0.0 | 0.4 | GO:0005637 | nuclear inner membrane(GO:0005637) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 2.4 | GO:0051059 | NF-kappaB binding(GO:0051059) |
| 0.3 | 2.3 | GO:0016728 | ribonucleoside-diphosphate reductase activity, thioredoxin disulfide as acceptor(GO:0004748) oxidoreductase activity, acting on CH or CH2 groups, disulfide as acceptor(GO:0016728) ribonucleoside-diphosphate reductase activity(GO:0061731) |
| 0.3 | 1.3 | GO:0043139 | 5'-3' DNA helicase activity(GO:0043139) |
| 0.2 | 1.2 | GO:0003980 | UDP-glucose:glycoprotein glucosyltransferase activity(GO:0003980) |
| 0.2 | 1.1 | GO:0004142 | diacylglycerol cholinephosphotransferase activity(GO:0004142) ethanolaminephosphotransferase activity(GO:0004307) |
| 0.2 | 0.5 | GO:0070815 | peptidyl-lysine 5-dioxygenase activity(GO:0070815) |
| 0.2 | 0.9 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 0.1 | 0.5 | GO:0008117 | sphinganine-1-phosphate aldolase activity(GO:0008117) |
| 0.1 | 0.5 | GO:0004694 | eukaryotic translation initiation factor 2alpha kinase activity(GO:0004694) |
| 0.1 | 0.3 | GO:0004421 | hydroxymethylglutaryl-CoA synthase activity(GO:0004421) |
| 0.1 | 0.7 | GO:0016212 | kynurenine-oxoglutarate transaminase activity(GO:0016212) kynurenine aminotransferase activity(GO:0036137) |
| 0.1 | 0.4 | GO:0035516 | oxidative DNA demethylase activity(GO:0035516) |
| 0.1 | 0.3 | GO:2001070 | starch binding(GO:2001070) |
| 0.1 | 0.3 | GO:0004649 | poly(ADP-ribose) glycohydrolase activity(GO:0004649) |
| 0.1 | 1.2 | GO:0017002 | activin-activated receptor activity(GO:0017002) |
| 0.1 | 0.9 | GO:0043495 | protein anchor(GO:0043495) |
| 0.1 | 0.6 | GO:0010340 | carboxyl-O-methyltransferase activity(GO:0010340) protein carboxyl O-methyltransferase activity(GO:0051998) |
| 0.1 | 0.2 | GO:0004361 | glutaryl-CoA dehydrogenase activity(GO:0004361) |
| 0.1 | 0.2 | GO:0030792 | arsenite methyltransferase activity(GO:0030791) methylarsonite methyltransferase activity(GO:0030792) |
| 0.1 | 0.2 | GO:0050333 | thiamin-triphosphatase activity(GO:0050333) |
| 0.0 | 0.1 | GO:0042806 | fucose binding(GO:0042806) |
| 0.0 | 0.9 | GO:0009982 | pseudouridine synthase activity(GO:0009982) |
| 0.0 | 2.5 | GO:0015485 | cholesterol binding(GO:0015485) |
| 0.0 | 0.2 | GO:0004679 | AMP-activated protein kinase activity(GO:0004679) |
| 0.0 | 0.3 | GO:0017070 | U6 snRNA binding(GO:0017070) |
| 0.0 | 0.3 | GO:0008476 | protein-tyrosine sulfotransferase activity(GO:0008476) |
| 0.0 | 3.1 | GO:0043566 | structure-specific DNA binding(GO:0043566) |
| 0.0 | 0.2 | GO:0034632 | retinol transporter activity(GO:0034632) |
| 0.0 | 0.6 | GO:0008242 | omega peptidase activity(GO:0008242) |
| 0.0 | 0.4 | GO:0046975 | histone methyltransferase activity (H3-K36 specific)(GO:0046975) |
| 0.0 | 0.5 | GO:0070001 | aspartic-type endopeptidase activity(GO:0004190) aspartic-type peptidase activity(GO:0070001) |
| 0.0 | 0.4 | GO:0051721 | protein phosphatase 2A binding(GO:0051721) |
| 0.0 | 0.5 | GO:0030515 | snoRNA binding(GO:0030515) |
| 0.0 | 0.5 | GO:0097602 | cullin family protein binding(GO:0097602) |
| 0.0 | 0.7 | GO:0042287 | MHC protein binding(GO:0042287) |
| 0.0 | 0.1 | GO:0043023 | ribosomal large subunit binding(GO:0043023) |
| 0.0 | 1.2 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 0.4 | GO:0050321 | tau-protein kinase activity(GO:0050321) |
| 0.0 | 0.5 | GO:0001103 | RNA polymerase II repressing transcription factor binding(GO:0001103) |
| 0.0 | 0.2 | GO:0008266 | poly(U) RNA binding(GO:0008266) |
| 0.0 | 0.2 | GO:0070577 | lysine-acetylated histone binding(GO:0070577) |
| 0.0 | 0.6 | GO:0004707 | MAP kinase activity(GO:0004707) |
| 0.0 | 0.6 | GO:0016307 | phosphatidylinositol phosphate kinase activity(GO:0016307) |
| 0.0 | 0.6 | GO:0019003 | GDP binding(GO:0019003) |
| 0.0 | 0.5 | GO:0030165 | PDZ domain binding(GO:0030165) |
| 0.0 | 0.8 | GO:0003755 | peptidyl-prolyl cis-trans isomerase activity(GO:0003755) |
| 0.0 | 1.3 | GO:0016763 | transferase activity, transferring pentosyl groups(GO:0016763) |
| 0.0 | 0.4 | GO:0016628 | oxidoreductase activity, acting on the CH-CH group of donors, NAD or NADP as acceptor(GO:0016628) |
| 0.0 | 0.3 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.0 | 0.6 | GO:0008013 | beta-catenin binding(GO:0008013) |
| 0.0 | 0.6 | GO:0001786 | phosphatidylserine binding(GO:0001786) |
| 0.0 | 0.8 | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity(GO:0016538) |
| 0.0 | 0.6 | GO:0004520 | endodeoxyribonuclease activity(GO:0004520) |
| 0.0 | 0.5 | GO:0005347 | ATP transmembrane transporter activity(GO:0005347) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.8 | PID DNA PK PATHWAY | DNA-PK pathway in nonhomologous end joining |
| 0.0 | 0.8 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.0 | 0.5 | PID S1P META PATHWAY | Sphingosine 1-phosphate (S1P) pathway |
| 0.0 | 0.7 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.0 | 0.5 | PID MYC PATHWAY | C-MYC pathway |
| 0.0 | 1.9 | PID E2F PATHWAY | E2F transcription factor network |
| 0.0 | 0.8 | PID NFAT 3PATHWAY | Role of Calcineurin-dependent NFAT signaling in lymphocytes |
| 0.0 | 1.1 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.0 | 1.0 | PID P53 REGULATION PATHWAY | p53 pathway |
| 0.0 | 0.6 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
| 0.0 | 0.4 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.0 | 0.3 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.0 | 0.7 | PID P73PATHWAY | p73 transcription factor network |
| 0.0 | 0.3 | ST GA13 PATHWAY | G alpha 13 Pathway |
| 0.0 | 0.3 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.6 | REACTOME ACYL CHAIN REMODELLING OF PS | Genes involved in Acyl chain remodelling of PS |
| 0.1 | 2.3 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.1 | 1.2 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.1 | 1.2 | REACTOME CALNEXIN CALRETICULIN CYCLE | Genes involved in Calnexin/calreticulin cycle |
| 0.1 | 0.7 | REACTOME COPI MEDIATED TRANSPORT | Genes involved in COPI Mediated Transport |
| 0.1 | 0.7 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 0.1 | 0.6 | REACTOME VITAMIN B5 PANTOTHENATE METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |
| 0.0 | 0.7 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.0 | 0.3 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.0 | 0.6 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.0 | 0.5 | REACTOME PERK REGULATED GENE EXPRESSION | Genes involved in PERK regulated gene expression |
| 0.0 | 0.3 | REACTOME RIP MEDIATED NFKB ACTIVATION VIA DAI | Genes involved in RIP-mediated NFkB activation via DAI |
| 0.0 | 0.7 | REACTOME ACTIVATION OF THE PRE REPLICATIVE COMPLEX | Genes involved in Activation of the pre-replicative complex |
| 0.0 | 0.3 | REACTOME AKT PHOSPHORYLATES TARGETS IN THE CYTOSOL | Genes involved in AKT phosphorylates targets in the cytosol |
| 0.0 | 0.5 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.0 | 0.5 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |