Project
PRJNA438478: RNAseq of wild type zebrafish germline
Navigation
Downloads
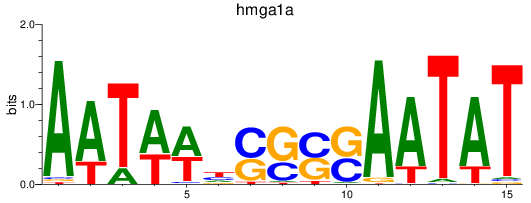
Results for hmga1a
Z-value: 0.52
Transcription factors associated with hmga1a
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
hmga1a
|
ENSDARG00000028335 | high mobility group AT-hook 1a |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| hmga1a | dr11_v1_chr23_-_3759692_3759692 | 0.44 | 7.1e-02 | Click! |
Activity profile of hmga1a motif
Sorted Z-values of hmga1a motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr23_-_45439903 | 0.77 |
ENSDART00000170729
|
NPNT (1 of many)
|
nephronectin |
| chr16_-_29557338 | 0.60 |
ENSDART00000058888
|
hormad1
|
HORMA domain containing 1 |
| chr17_-_5769196 | 0.53 |
ENSDART00000113885
|
si:dkey-100n19.2
|
si:dkey-100n19.2 |
| chr7_-_33130552 | 0.53 |
ENSDART00000127006
|
mns1
|
meiosis-specific nuclear structural 1 |
| chr16_-_42969598 | 0.51 |
ENSDART00000156011
|
si:ch211-135n15.3
|
si:ch211-135n15.3 |
| chr5_+_71924175 | 0.51 |
ENSDART00000115182
ENSDART00000170215 |
nup214
|
nucleoporin 214 |
| chr10_-_26202766 | 0.51 |
ENSDART00000136393
|
fhdc3
|
FH2 domain containing 3 |
| chr14_-_34513103 | 0.49 |
ENSDART00000136306
|
zgc:194246
|
zgc:194246 |
| chr19_-_20403507 | 0.46 |
ENSDART00000052603
ENSDART00000137590 |
dazl
|
deleted in azoospermia-like |
| chr3_+_62161184 | 0.41 |
ENSDART00000090370
ENSDART00000192665 |
noxo1a
|
NADPH oxidase organizer 1a |
| chr16_+_2433696 | 0.38 |
ENSDART00000182803
|
CU928063.1
|
|
| chr10_+_44584614 | 0.37 |
ENSDART00000163523
|
sez6l
|
seizure related 6 homolog (mouse)-like |
| chr11_-_16975190 | 0.35 |
ENSDART00000122222
|
suclg2
|
succinate-CoA ligase, GDP-forming, beta subunit |
| chr4_+_26053044 | 0.34 |
ENSDART00000039877
|
SCYL2
|
si:ch211-244b2.1 |
| chr17_+_31739418 | 0.33 |
ENSDART00000155073
ENSDART00000156180 |
arhgap5
|
Rho GTPase activating protein 5 |
| chr9_+_55455801 | 0.31 |
ENSDART00000144757
ENSDART00000186543 |
mxra5b
|
matrix-remodelling associated 5b |
| chr4_+_13568469 | 0.30 |
ENSDART00000171235
ENSDART00000136152 |
calua
|
calumenin a |
| chr22_+_23359369 | 0.29 |
ENSDART00000170886
|
dennd1b
|
DENN/MADD domain containing 1B |
| chr25_+_581227 | 0.29 |
ENSDART00000126863
|
nell2a
|
neural EGFL like 2a |
| chr11_+_37095165 | 0.27 |
ENSDART00000124997
|
card19
|
caspase recruitment domain family, member 19 |
| chr24_-_15159658 | 0.27 |
ENSDART00000142473
|
rttn
|
rotatin |
| chr19_+_29808699 | 0.26 |
ENSDART00000051799
ENSDART00000164205 |
hdac1
|
histone deacetylase 1 |
| chr10_+_6121558 | 0.26 |
ENSDART00000166799
ENSDART00000157947 |
tln1
|
talin 1 |
| chr12_-_10409961 | 0.26 |
ENSDART00000149521
ENSDART00000052001 |
eef2k
|
eukaryotic elongation factor 2 kinase |
| chr10_+_20364009 | 0.25 |
ENSDART00000186139
ENSDART00000080395 |
golga7
|
golgin A7 |
| chr19_+_29808471 | 0.25 |
ENSDART00000186428
|
hdac1
|
histone deacetylase 1 |
| chr25_-_34670413 | 0.24 |
ENSDART00000073440
|
DNAJA4
|
DnaJ heat shock protein family (Hsp40) member A4 |
| chr4_+_17245005 | 0.24 |
ENSDART00000027645
|
casc1
|
cancer susceptibility candidate 1 |
| chr4_+_17245217 | 0.23 |
ENSDART00000184215
ENSDART00000110908 |
casc1
|
cancer susceptibility candidate 1 |
| chr25_+_15287036 | 0.23 |
ENSDART00000147572
|
hipk3a
|
homeodomain interacting protein kinase 3a |
| chr6_-_2222707 | 0.22 |
ENSDART00000022179
|
prkag1
|
protein kinase, AMP-activated, gamma 1 non-catalytic subunit |
| chr1_-_14258409 | 0.22 |
ENSDART00000079359
|
pde5aa
|
phosphodiesterase 5A, cGMP-specific, a |
| chr8_+_49975160 | 0.22 |
ENSDART00000156403
ENSDART00000080135 |
gfpt1
|
glutamine--fructose-6-phosphate transaminase 1 |
| chr7_+_72469724 | 0.21 |
ENSDART00000187629
|
HECTD4
|
HECT domain E3 ubiquitin protein ligase 4 |
| chr23_+_45512825 | 0.20 |
ENSDART00000064846
|
prelid1b
|
PRELI domain containing 1b |
| chr13_-_41917050 | 0.20 |
ENSDART00000110549
|
cisd1
|
CDGSH iron sulfur domain 1 |
| chr24_-_16975960 | 0.20 |
ENSDART00000180687
|
klhl15
|
kelch-like family member 15 |
| chr24_-_35534273 | 0.19 |
ENSDART00000026578
|
ube2v2
|
ubiquitin-conjugating enzyme E2 variant 2 |
| chr25_-_13789955 | 0.19 |
ENSDART00000167742
ENSDART00000165116 ENSDART00000171461 |
ckap5
|
cytoskeleton associated protein 5 |
| chr15_-_33807758 | 0.18 |
ENSDART00000158445
|
pds5b
|
PDS5 cohesin associated factor B |
| chr4_+_73872345 | 0.18 |
ENSDART00000174075
|
CABZ01069022.1
|
|
| chr22_-_16416882 | 0.17 |
ENSDART00000062749
|
cts12
|
cathepsin 12 |
| chr16_-_12236362 | 0.17 |
ENSDART00000114759
|
lpcat3
|
lysophosphatidylcholine acyltransferase 3 |
| chr22_-_784110 | 0.17 |
ENSDART00000061775
|
rbbp5
|
retinoblastoma binding protein 5 |
| chr3_-_34136368 | 0.17 |
ENSDART00000136900
ENSDART00000186125 |
clpp
|
caseinolytic mitochondrial matrix peptidase proteolytic subunit |
| chr16_-_26232411 | 0.17 |
ENSDART00000139355
|
arhgef1b
|
Rho guanine nucleotide exchange factor (GEF) 1b |
| chr1_+_44360973 | 0.17 |
ENSDART00000167568
|
si:ch211-165a10.5
|
si:ch211-165a10.5 |
| chr9_+_25568839 | 0.16 |
ENSDART00000177342
|
zeb2a
|
zinc finger E-box binding homeobox 2a |
| chr11_+_5817202 | 0.16 |
ENSDART00000126084
|
ctdspl3
|
CTD (carboxy-terminal domain, RNA polymerase II, polypeptide A) small phosphatase like 3 |
| chr23_+_38940700 | 0.16 |
ENSDART00000065331
|
sall4
|
spalt-like transcription factor 4 |
| chr14_+_35237613 | 0.15 |
ENSDART00000163465
|
ebf3a
|
early B cell factor 3a |
| chr23_-_27633730 | 0.15 |
ENSDART00000103639
|
arf3a
|
ADP-ribosylation factor 3a |
| chr3_+_26019426 | 0.14 |
ENSDART00000135389
ENSDART00000182411 |
foxred2
|
FAD-dependent oxidoreductase domain containing 2 |
| chr6_+_30533504 | 0.13 |
ENSDART00000155842
|
wwc3
|
WWC family member 3 |
| chr25_-_13549577 | 0.12 |
ENSDART00000166772
|
ano10b
|
anoctamin 10b |
| chr7_-_22956889 | 0.12 |
ENSDART00000101447
|
tnfsf10l
|
TNF superfamily member 10, like |
| chr8_-_49725430 | 0.12 |
ENSDART00000135675
|
gkap1
|
G kinase anchoring protein 1 |
| chr14_+_26566936 | 0.12 |
ENSDART00000173056
|
arap3
|
ArfGAP with RhoGAP domain, ankyrin repeat and PH domain 3 |
| chr17_+_51520073 | 0.12 |
ENSDART00000189646
ENSDART00000170951 ENSDART00000189492 |
pxdn
|
peroxidasin |
| chr22_-_10580194 | 0.11 |
ENSDART00000105848
|
SHISA4 (1 of many)
|
si:dkey-42i9.7 |
| chr12_+_46841918 | 0.11 |
ENSDART00000157464
|
adkb
|
adenosine kinase b |
| chr8_+_42917515 | 0.11 |
ENSDART00000021715
|
slc23a2
|
solute carrier family 23 (ascorbic acid transporter), member 2 |
| chr1_+_41690402 | 0.11 |
ENSDART00000177298
|
fbxo41
|
F-box protein 41 |
| chr14_-_38827442 | 0.11 |
ENSDART00000160000
|
spdl1
|
spindle apparatus coiled-coil protein 1 |
| chr9_-_27738110 | 0.10 |
ENSDART00000060347
|
crygs2
|
crystallin, gamma S2 |
| chr17_-_23221504 | 0.10 |
ENSDART00000156411
|
fam98a
|
family with sequence similarity 98, member A |
| chr7_+_33130639 | 0.10 |
ENSDART00000142450
ENSDART00000173967 ENSDART00000173832 |
zgc:153219
si:ch211-194p6.7
|
zgc:153219 si:ch211-194p6.7 |
| chr1_-_54971968 | 0.09 |
ENSDART00000140016
|
khsrp
|
KH-type splicing regulatory protein |
| chr14_-_38828057 | 0.09 |
ENSDART00000186088
|
spdl1
|
spindle apparatus coiled-coil protein 1 |
| chr6_-_39518489 | 0.09 |
ENSDART00000185446
|
atf1
|
activating transcription factor 1 |
| chr7_+_19615056 | 0.08 |
ENSDART00000124752
ENSDART00000190297 |
si:ch211-212k18.15
|
si:ch211-212k18.15 |
| chr19_-_45650994 | 0.08 |
ENSDART00000163504
|
trps1
|
trichorhinophalangeal syndrome I |
| chr25_-_17910714 | 0.08 |
ENSDART00000191586
|
arntl1a
|
aryl hydrocarbon receptor nuclear translocator-like 1a |
| chr13_+_42406883 | 0.08 |
ENSDART00000084354
|
cpeb3
|
cytoplasmic polyadenylation element binding protein 3 |
| chr1_-_54972170 | 0.08 |
ENSDART00000150548
ENSDART00000038330 |
khsrp
|
KH-type splicing regulatory protein |
| chr24_-_31090948 | 0.08 |
ENSDART00000176799
|
hccsb
|
holocytochrome c synthase b |
| chr6_-_7123210 | 0.08 |
ENSDART00000041304
|
atg3
|
autophagy related 3 |
| chr6_-_48094342 | 0.08 |
ENSDART00000137458
|
slc2a1b
|
solute carrier family 2 (facilitated glucose transporter), member 1b |
| chr4_+_19127973 | 0.08 |
ENSDART00000136611
|
si:dkey-21o22.2
|
si:dkey-21o22.2 |
| chr7_-_69521481 | 0.08 |
ENSDART00000148465
|
slc1a1
|
solute carrier family 1 (neuronal/epithelial high affinity glutamate transporter, system Xag), member 1 |
| chr3_+_36515376 | 0.07 |
ENSDART00000161652
|
si:dkeyp-72e1.9
|
si:dkeyp-72e1.9 |
| chr24_+_39137001 | 0.07 |
ENSDART00000181086
ENSDART00000183724 ENSDART00000193466 |
tbc1d24
|
TBC1 domain family, member 24 |
| chr5_-_20814576 | 0.06 |
ENSDART00000098682
ENSDART00000147639 |
si:ch211-225b11.1
|
si:ch211-225b11.1 |
| chr7_+_48999723 | 0.06 |
ENSDART00000182699
ENSDART00000166329 |
si:ch211-288d18.1
|
si:ch211-288d18.1 |
| chr9_-_2892250 | 0.05 |
ENSDART00000140695
|
cdca7a
|
cell division cycle associated 7a |
| chr15_+_47440477 | 0.05 |
ENSDART00000002384
|
phox2a
|
paired-like homeobox 2a |
| chr5_+_41476443 | 0.05 |
ENSDART00000145228
ENSDART00000137981 ENSDART00000142538 |
pias2
|
protein inhibitor of activated STAT, 2 |
| chr18_-_35407289 | 0.05 |
ENSDART00000012018
|
snrpa
|
small nuclear ribonucleoprotein polypeptide A |
| chr6_+_3827751 | 0.05 |
ENSDART00000003008
ENSDART00000122348 |
gad1b
|
glutamate decarboxylase 1b |
| chr16_+_11724230 | 0.05 |
ENSDART00000060266
|
ceacam1
|
carcinoembryonic antigen-related cell adhesion molecule 1 |
| chr20_-_874807 | 0.05 |
ENSDART00000020506
|
snx14
|
sorting nexin 14 |
| chr18_+_14595546 | 0.05 |
ENSDART00000080788
|
wfdc1
|
WAP four-disulfide core domain 1 |
| chr17_+_5768608 | 0.05 |
ENSDART00000157039
|
rp1l1a
|
retinitis pigmentosa 1-like 1a |
| chr15_+_15856178 | 0.05 |
ENSDART00000080338
|
dusp14
|
dual specificity phosphatase 14 |
| chr1_+_49814461 | 0.05 |
ENSDART00000132405
|
lef1
|
lymphoid enhancer-binding factor 1 |
| chr10_-_22106331 | 0.04 |
ENSDART00000187654
|
BX005027.2
|
|
| chr21_+_21743599 | 0.04 |
ENSDART00000101700
|
pold3
|
polymerase (DNA-directed), delta 3, accessory subunit |
| chr18_-_35407695 | 0.04 |
ENSDART00000191845
ENSDART00000141703 |
snrpa
|
small nuclear ribonucleoprotein polypeptide A |
| chr10_+_3049636 | 0.04 |
ENSDART00000081794
ENSDART00000183167 ENSDART00000191634 ENSDART00000183514 |
rasgrf2a
|
Ras protein-specific guanine nucleotide-releasing factor 2a |
| chr4_-_73372362 | 0.04 |
ENSDART00000182312
ENSDART00000158404 ENSDART00000174367 |
si:cabz01021430.2
zgc:171727
si:cabz01021426.2
|
si:cabz01021430.2 zgc:171727 si:cabz01021426.2 |
| chr20_+_26349002 | 0.04 |
ENSDART00000152842
|
syne1a
|
spectrin repeat containing, nuclear envelope 1a |
| chr5_+_32831561 | 0.04 |
ENSDART00000169358
ENSDART00000192078 |
si:ch211-208h16.4
|
si:ch211-208h16.4 |
| chr14_+_8710568 | 0.03 |
ENSDART00000169505
|
kcnk4a
|
potassium channel, subfamily K, member 4a |
| chr11_-_12051502 | 0.03 |
ENSDART00000183462
|
socs7
|
suppressor of cytokine signaling 7 |
| chr6_+_22679610 | 0.03 |
ENSDART00000102701
|
zmp:0000000634
|
zmp:0000000634 |
| chr15_-_29387446 | 0.03 |
ENSDART00000145976
ENSDART00000035096 |
serpinh1b
|
serpin peptidase inhibitor, clade H (heat shock protein 47), member 1b |
| chr6_+_3828560 | 0.03 |
ENSDART00000185273
ENSDART00000179091 |
gad1b
|
glutamate decarboxylase 1b |
| chr6_+_28294113 | 0.03 |
ENSDART00000136898
|
lpp
|
LIM domain containing preferred translocation partner in lipoma |
| chr5_-_51830997 | 0.02 |
ENSDART00000163616
|
homer1b
|
homer scaffolding protein 1b |
| chr20_+_34400715 | 0.02 |
ENSDART00000061632
|
fam129aa
|
family with sequence similarity 129, member Aa |
| chr9_+_36314867 | 0.02 |
ENSDART00000176763
|
lrp1bb
|
low density lipoprotein receptor-related protein 1Bb |
| chr11_-_12051283 | 0.02 |
ENSDART00000170516
|
socs7
|
suppressor of cytokine signaling 7 |
| chr15_+_44135726 | 0.02 |
ENSDART00000192151
|
CU655961.2
|
|
| chr6_-_10740365 | 0.02 |
ENSDART00000150942
|
sp3b
|
Sp3b transcription factor |
| chr16_+_1383914 | 0.02 |
ENSDART00000185089
|
cers2b
|
ceramide synthase 2b |
| chr14_+_46020282 | 0.02 |
ENSDART00000190087
|
c1qtnf2
|
C1q and TNF related 2 |
| chr1_+_2112726 | 0.02 |
ENSDART00000131714
ENSDART00000138396 |
mbnl2
|
muscleblind-like splicing regulator 2 |
| chr14_+_46020571 | 0.02 |
ENSDART00000157617
ENSDART00000083928 |
c1qtnf2
|
C1q and TNF related 2 |
| chr3_+_57820913 | 0.01 |
ENSDART00000168101
|
CU571328.1
|
|
| chr1_+_31658011 | 0.01 |
ENSDART00000192203
|
poll
|
polymerase (DNA directed), lambda |
| chr8_+_40644838 | 0.01 |
ENSDART00000169311
|
adra2b
|
adrenoceptor alpha 2B |
| chr5_-_38451082 | 0.01 |
ENSDART00000136428
|
chrne
|
cholinergic receptor, nicotinic, epsilon |
| chr2_+_45548890 | 0.01 |
ENSDART00000113994
|
fndc7a
|
fibronectin type III domain containing 7a |
| chr11_+_36683859 | 0.01 |
ENSDART00000170102
|
si:ch211-11c3.12
|
si:ch211-11c3.12 |
| chr17_-_15201031 | 0.00 |
ENSDART00000124426
|
socs4
|
suppressor of cytokine signaling 4 |
| chr5_-_39805874 | 0.00 |
ENSDART00000176202
ENSDART00000191683 |
rasgef1ba
|
RasGEF domain family, member 1Ba |
| chr9_-_12885201 | 0.00 |
ENSDART00000124957
|
ankzf1
|
ankyrin repeat and zinc finger domain containing 1 |
| chr7_-_22956716 | 0.00 |
ENSDART00000122113
|
tnfsf10l
|
TNF superfamily member 10, like |
| chr8_-_50981175 | 0.00 |
ENSDART00000004065
|
zgc:91909
|
zgc:91909 |
| chr21_-_20765338 | 0.00 |
ENSDART00000135940
|
ghrb
|
growth hormone receptor b |
| chr18_-_35407530 | 0.00 |
ENSDART00000137663
|
snrpa
|
small nuclear ribonucleoprotein polypeptide A |
| chr15_+_888704 | 0.00 |
ENSDART00000182796
|
si:dkey-7i4.9
|
si:dkey-7i4.9 |
Network of associatons between targets according to the STRING database.
First level regulatory network of hmga1a
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.6 | GO:0060631 | regulation of meiosis I(GO:0060631) |
| 0.1 | 0.5 | GO:0060967 | negative regulation of posttranscriptional gene silencing(GO:0060149) negative regulation of gene silencing by miRNA(GO:0060965) negative regulation of gene silencing by RNA(GO:0060967) |
| 0.1 | 0.5 | GO:0033687 | osteoblast proliferation(GO:0033687) |
| 0.0 | 0.2 | GO:0030951 | establishment or maintenance of microtubule cytoskeleton polarity(GO:0030951) |
| 0.0 | 0.3 | GO:0010457 | centriole-centriole cohesion(GO:0010457) |
| 0.0 | 0.1 | GO:0019852 | L-ascorbic acid transport(GO:0015882) L-ascorbic acid metabolic process(GO:0019852) |
| 0.0 | 0.5 | GO:0070193 | synaptonemal complex assembly(GO:0007130) synaptonemal complex organization(GO:0070193) |
| 0.0 | 0.1 | GO:0009449 | gamma-aminobutyric acid biosynthetic process(GO:0009449) |
| 0.0 | 0.2 | GO:0090243 | fibroblast growth factor receptor signaling pathway involved in somitogenesis(GO:0090243) |
| 0.0 | 0.2 | GO:0007080 | mitotic metaphase plate congression(GO:0007080) |
| 0.0 | 0.2 | GO:0006656 | phosphatidylcholine biosynthetic process(GO:0006656) |
| 0.0 | 0.1 | GO:0044209 | AMP salvage(GO:0044209) |
| 0.0 | 0.3 | GO:0061300 | cerebellum vasculature development(GO:0061300) |
| 0.0 | 0.2 | GO:0006048 | UDP-N-acetylglucosamine biosynthetic process(GO:0006048) |
| 0.0 | 0.1 | GO:0017003 | protein-heme linkage(GO:0017003) protein-tetrapyrrole linkage(GO:0017006) cytochrome c-heme linkage(GO:0018063) |
| 0.0 | 0.3 | GO:0043001 | Golgi to plasma membrane protein transport(GO:0043001) |
| 0.0 | 0.4 | GO:0006801 | superoxide metabolic process(GO:0006801) |
| 0.0 | 0.0 | GO:0070914 | UV-damage excision repair(GO:0070914) |
| 0.0 | 0.1 | GO:0043653 | mitochondrial fragmentation involved in apoptotic process(GO:0043653) |
| 0.0 | 0.1 | GO:2001238 | positive regulation of extrinsic apoptotic signaling pathway(GO:2001238) |
| 0.0 | 0.2 | GO:0007064 | mitotic sister chromatid cohesion(GO:0007064) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0042709 | succinate-CoA ligase complex(GO:0042709) |
| 0.0 | 0.3 | GO:0033018 | sarcoplasmic reticulum lumen(GO:0033018) |
| 0.0 | 0.2 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.0 | 0.3 | GO:0002178 | palmitoyltransferase complex(GO:0002178) |
| 0.0 | 0.1 | GO:0072669 | tRNA-splicing ligase complex(GO:0072669) |
| 0.0 | 0.2 | GO:0000940 | condensed chromosome outer kinetochore(GO:0000940) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0004775 | succinate-CoA ligase activity(GO:0004774) succinate-CoA ligase (ADP-forming) activity(GO:0004775) succinate-CoA ligase (GDP-forming) activity(GO:0004776) |
| 0.1 | 0.5 | GO:0008494 | translation activator activity(GO:0008494) |
| 0.0 | 0.4 | GO:0016176 | superoxide-generating NADPH oxidase activator activity(GO:0016176) |
| 0.0 | 0.2 | GO:0070548 | L-glutamine aminotransferase activity(GO:0070548) |
| 0.0 | 0.1 | GO:0005547 | phosphatidylinositol-3,4,5-trisphosphate binding(GO:0005547) |
| 0.0 | 0.5 | GO:0031078 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.0 | 0.2 | GO:0047184 | 1-acylglycerophosphocholine O-acyltransferase activity(GO:0047184) |
| 0.0 | 0.2 | GO:1990050 | phosphatidic acid transporter activity(GO:1990050) |
| 0.0 | 0.1 | GO:0019777 | Atg12 transferase activity(GO:0019777) |
| 0.0 | 0.1 | GO:0004351 | glutamate decarboxylase activity(GO:0004351) |
| 0.0 | 0.5 | GO:0008139 | nuclear localization sequence binding(GO:0008139) |
| 0.0 | 0.2 | GO:0043515 | kinetochore binding(GO:0043515) |
| 0.0 | 0.1 | GO:0004001 | adenosine kinase activity(GO:0004001) |
| 0.0 | 0.2 | GO:0004176 | ATP-dependent peptidase activity(GO:0004176) |
| 0.0 | 0.2 | GO:0017091 | AU-rich element binding(GO:0017091) mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.0 | 0.2 | GO:0047555 | 3',5'-cyclic-GMP phosphodiesterase activity(GO:0047555) |
| 0.0 | 0.1 | GO:0004408 | holocytochrome-c synthase activity(GO:0004408) |
| 0.0 | 0.3 | GO:0005080 | protein kinase C binding(GO:0005080) |
| 0.0 | 0.1 | GO:0030619 | U1 snRNA binding(GO:0030619) |
| 0.0 | 0.1 | GO:0055056 | D-glucose transmembrane transporter activity(GO:0055056) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.6 | PID RANBP2 PATHWAY | Sumoylation by RanBP2 regulates transcriptional repression |
| 0.0 | 0.5 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.0 | 0.3 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.0 | 0.3 | PID INTEGRIN A4B1 PATHWAY | Alpha4 beta1 integrin signaling events |
| 0.0 | 0.4 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |
| 0.0 | 0.2 | REACTOME ACYL CHAIN REMODELLING OF PS | Genes involved in Acyl chain remodelling of PS |
| 0.0 | 0.3 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.0 | 0.3 | REACTOME MTORC1 MEDIATED SIGNALLING | Genes involved in mTORC1-mediated signalling |
| 0.0 | 0.5 | REACTOME DOWNREGULATION OF SMAD2 3 SMAD4 TRANSCRIPTIONAL ACTIVITY | Genes involved in Downregulation of SMAD2/3:SMAD4 transcriptional activity |
| 0.0 | 0.4 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.0 | 0.5 | REACTOME TRANSPORT OF RIBONUCLEOPROTEINS INTO THE HOST NUCLEUS | Genes involved in Transport of Ribonucleoproteins into the Host Nucleus |
| 0.0 | 0.1 | REACTOME GLUTAMATE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Glutamate Neurotransmitter Release Cycle |