Project
PRJNA438478: RNAseq of wild type zebrafish germline
Navigation
Downloads
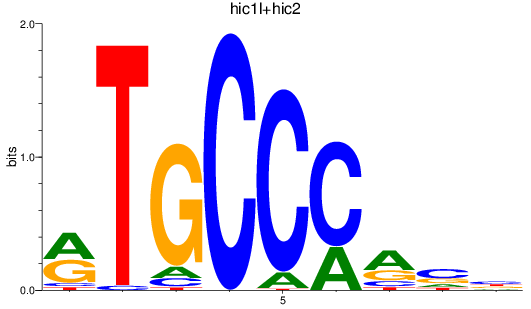
Results for hic1l+hic2
Z-value: 0.41
Transcription factors associated with hic1l+hic2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
hic1l
|
ENSDARG00000045660 | hypermethylated in cancer 1 like |
|
hic2
|
ENSDARG00000100497 | hypermethylated in cancer 2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| hic1l | dr11_v1_chr25_-_25575241_25575241 | -0.55 | 1.9e-02 | Click! |
| hic2 | dr11_v1_chr10_+_3153973_3153973 | -0.34 | 1.6e-01 | Click! |
Activity profile of hic1l+hic2 motif
Sorted Z-values of hic1l+hic2 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr22_-_25612680 | 1.57 |
ENSDART00000114167
|
si:ch211-12h2.8
|
si:ch211-12h2.8 |
| chr19_+_14109348 | 0.69 |
ENSDART00000159015
|
zgc:175136
|
zgc:175136 |
| chr4_-_8030583 | 0.66 |
ENSDART00000113628
|
si:ch211-240l19.8
|
si:ch211-240l19.8 |
| chr3_+_24207243 | 0.48 |
ENSDART00000023454
ENSDART00000136400 |
adsl
|
adenylosuccinate lyase |
| chr6_-_29402578 | 0.40 |
ENSDART00000115031
|
mrpl47
|
mitochondrial ribosomal protein L47 |
| chr4_-_8040436 | 0.38 |
ENSDART00000113033
|
si:ch211-240l19.6
|
si:ch211-240l19.6 |
| chr8_-_52715911 | 0.38 |
ENSDART00000168241
|
tubb2b
|
tubulin, beta 2b |
| chr24_-_9989634 | 0.37 |
ENSDART00000115275
|
zgc:152652
|
zgc:152652 |
| chr3_-_1387292 | 0.31 |
ENSDART00000163535
|
ddx47
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 47 |
| chr21_-_38717854 | 0.31 |
ENSDART00000065169
ENSDART00000113813 |
siah2l
|
seven in absentia homolog 2 (Drosophila)-like |
| chr20_+_54280549 | 0.31 |
ENSDART00000151048
|
actr10
|
ARP10 actin related protein 10 homolog |
| chr11_-_16021424 | 0.29 |
ENSDART00000193291
ENSDART00000170731 ENSDART00000104107 |
zgc:173544
|
zgc:173544 |
| chr2_+_24936766 | 0.28 |
ENSDART00000025962
|
gyg1a
|
glycogenin 1a |
| chr5_-_57820873 | 0.27 |
ENSDART00000089961
|
sik2a
|
salt-inducible kinase 2a |
| chr21_-_22681534 | 0.26 |
ENSDART00000159233
|
gig2f
|
grass carp reovirus (GCRV)-induced gene 2f |
| chr22_-_7129631 | 0.25 |
ENSDART00000171359
|
asic1b
|
acid-sensing (proton-gated) ion channel 1b |
| chr22_+_7742211 | 0.23 |
ENSDART00000140896
|
CELA1 (1 of many)
|
zgc:92511 |
| chr22_+_7497319 | 0.22 |
ENSDART00000034564
|
CELA1 (1 of many)
|
zgc:92511 |
| chr16_-_53800047 | 0.20 |
ENSDART00000158047
|
znrf2b
|
zinc and ring finger 2b |
| chr3_-_11828206 | 0.20 |
ENSDART00000018159
|
si:ch211-262e15.1
|
si:ch211-262e15.1 |
| chr21_+_21612214 | 0.19 |
ENSDART00000008099
|
b9d2
|
B9 domain containing 2 |
| chr7_-_30143092 | 0.19 |
ENSDART00000173636
|
frmd5
|
FERM domain containing 5 |
| chr23_-_20126257 | 0.18 |
ENSDART00000005021
|
tktb
|
transketolase b |
| chr21_-_36619599 | 0.18 |
ENSDART00000065208
|
nop16
|
NOP16 nucleolar protein homolog (yeast) |
| chr8_+_36570791 | 0.17 |
ENSDART00000145566
ENSDART00000180527 |
pold2
|
polymerase (DNA directed), delta 2, regulatory subunit |
| chr23_+_24957008 | 0.17 |
ENSDART00000181069
|
nol9
|
nucleolar protein 9 |
| chr21_+_18877130 | 0.16 |
ENSDART00000136893
|
si:dkey-65l23.2
|
si:dkey-65l23.2 |
| chr2_-_30912922 | 0.15 |
ENSDART00000141669
|
myl12.2
|
myosin, light chain 12, genome duplicate 2 |
| chr7_-_38689562 | 0.15 |
ENSDART00000167209
|
aplnr2
|
apelin receptor 2 |
| chr18_-_5103931 | 0.15 |
ENSDART00000188091
|
pdcd10a
|
programmed cell death 10a |
| chr7_+_21887307 | 0.15 |
ENSDART00000052871
|
pop7
|
POP7 homolog, ribonuclease P/MRP subunit |
| chr23_+_2669 | 0.15 |
ENSDART00000011146
|
twist3
|
twist3 |
| chr11_-_41220794 | 0.15 |
ENSDART00000192895
|
mrps16
|
mitochondrial ribosomal protein S16 |
| chr13_-_7573670 | 0.14 |
ENSDART00000102538
|
pitx3
|
paired-like homeodomain 3 |
| chr5_-_25733745 | 0.13 |
ENSDART00000051566
|
zgc:101016
|
zgc:101016 |
| chr5_+_15992655 | 0.13 |
ENSDART00000182148
|
znrf3
|
zinc and ring finger 3 |
| chr11_+_13032483 | 0.13 |
ENSDART00000108968
|
ptgfr
|
prostaglandin F receptor (FP) |
| chr23_-_6765653 | 0.12 |
ENSDART00000192310
|
FP102169.1
|
|
| chr10_+_41199660 | 0.12 |
ENSDART00000125314
|
adrb3b
|
adrenoceptor beta 3b |
| chr3_+_34821327 | 0.11 |
ENSDART00000055262
|
cdk5r1a
|
cyclin-dependent kinase 5, regulatory subunit 1a (p35) |
| chr15_-_28082310 | 0.11 |
ENSDART00000152620
|
dhrs13a.3
|
dehydrogenase/reductase (SDR family) member 13a, duplicate 3 |
| chr13_+_21870269 | 0.11 |
ENSDART00000144612
|
zswim8
|
zinc finger, SWIM-type containing 8 |
| chr19_+_19241372 | 0.11 |
ENSDART00000184392
ENSDART00000165008 |
ptpn23b
|
protein tyrosine phosphatase, non-receptor type 23, b |
| chr14_-_897874 | 0.11 |
ENSDART00000167395
|
rgs14a
|
regulator of G protein signaling 14a |
| chr4_-_25485404 | 0.11 |
ENSDART00000041402
|
pfkfb3
|
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 3 |
| chr4_+_72235562 | 0.11 |
ENSDART00000168547
|
si:cabz01071912.2
|
si:cabz01071912.2 |
| chr11_+_20057182 | 0.11 |
ENSDART00000184979
|
fezf2
|
FEZ family zinc finger 2 |
| chr2_-_43605556 | 0.11 |
ENSDART00000084223
|
epc1b
|
enhancer of polycomb homolog 1 (Drosophila) b |
| chr25_-_16146851 | 0.11 |
ENSDART00000104043
|
dkk3b
|
dickkopf WNT signaling pathway inhibitor 3b |
| chr2_+_11206317 | 0.10 |
ENSDART00000144982
|
lhx8a
|
LIM homeobox 8a |
| chr16_+_43347966 | 0.10 |
ENSDART00000171308
|
zmp:0000000930
|
zmp:0000000930 |
| chr21_-_26520629 | 0.10 |
ENSDART00000142731
|
rce1b
|
Ras converting CAAX endopeptidase 1b |
| chr21_-_25250594 | 0.10 |
ENSDART00000163862
|
nfrkb
|
nuclear factor related to kappaB binding protein |
| chr18_-_16123222 | 0.10 |
ENSDART00000061189
|
sspn
|
sarcospan (Kras oncogene-associated gene) |
| chr18_-_50845804 | 0.10 |
ENSDART00000158517
|
PDPR
|
si:cabz01113374.3 |
| chr12_+_28995942 | 0.09 |
ENSDART00000076334
|
valopb
|
vertebrate ancient long opsin b |
| chr18_-_19004247 | 0.09 |
ENSDART00000135800
|
ints14
|
integrator complex subunit 14 |
| chr3_-_61592417 | 0.09 |
ENSDART00000155082
|
nptx2a
|
neuronal pentraxin 2a |
| chr5_+_61738276 | 0.09 |
ENSDART00000186256
|
RASL10B
|
RAS like family 10 member B |
| chr2_+_31475772 | 0.08 |
ENSDART00000130722
|
cacnb2b
|
calcium channel, voltage-dependent, beta 2b |
| chr10_+_42691210 | 0.08 |
ENSDART00000193813
|
rhobtb2b
|
Rho-related BTB domain containing 2b |
| chr23_-_32404022 | 0.08 |
ENSDART00000156387
ENSDART00000155508 |
si:ch211-66i15.4
|
si:ch211-66i15.4 |
| chr21_-_31210749 | 0.08 |
ENSDART00000185356
|
zgc:152891
|
zgc:152891 |
| chr10_+_15608326 | 0.07 |
ENSDART00000188770
|
zfand5b
|
zinc finger, AN1-type domain 5b |
| chr17_+_15845765 | 0.07 |
ENSDART00000130881
ENSDART00000074936 |
gabrr2a
|
gamma-aminobutyric acid (GABA) A receptor, rho 2a |
| chr14_-_21064199 | 0.07 |
ENSDART00000172099
|
si:dkey-74k8.3
|
si:dkey-74k8.3 |
| chr15_-_16704417 | 0.07 |
ENSDART00000155163
|
caln1
|
calneuron 1 |
| chr12_+_22680115 | 0.07 |
ENSDART00000152879
|
ablim2
|
actin binding LIM protein family, member 2 |
| chr17_+_53294228 | 0.07 |
ENSDART00000158172
|
si:ch1073-416d2.3
|
si:ch1073-416d2.3 |
| chr25_-_31423493 | 0.07 |
ENSDART00000027661
|
myod1
|
myogenic differentiation 1 |
| chr5_-_26181863 | 0.07 |
ENSDART00000098500
|
ccdc125
|
coiled-coil domain containing 125 |
| chr8_+_29267093 | 0.07 |
ENSDART00000077647
|
grid2
|
glutamate receptor, ionotropic, delta 2 |
| chr12_+_35654749 | 0.07 |
ENSDART00000169889
ENSDART00000167873 |
baiap2b
|
BAI1-associated protein 2b |
| chr7_-_46782264 | 0.07 |
ENSDART00000166565
|
tshz3b
|
teashirt zinc finger homeobox 3b |
| chr8_-_38616712 | 0.07 |
ENSDART00000141827
|
si:ch211-198d23.1
|
si:ch211-198d23.1 |
| chr18_+_20677090 | 0.06 |
ENSDART00000060243
|
cftr
|
cystic fibrosis transmembrane conductance regulator (ATP-binding cassette sub-family C, member 7) |
| chr1_-_54425791 | 0.06 |
ENSDART00000039911
|
pkd1a
|
polycystic kidney disease 1a |
| chr2_-_32666174 | 0.06 |
ENSDART00000133660
|
puf60a
|
poly-U binding splicing factor a |
| chr15_+_25158104 | 0.06 |
ENSDART00000128267
|
slc35f2l
|
info solute carrier family 35, member F2, like |
| chr13_+_32148338 | 0.06 |
ENSDART00000188591
|
osr1
|
odd-skipped related transciption factor 1 |
| chr8_+_8973425 | 0.06 |
ENSDART00000066107
|
bcap31
|
B cell receptor associated protein 31 |
| chr14_-_5677979 | 0.06 |
ENSDART00000182156
|
tlx2
|
T cell leukemia homeobox 2 |
| chr16_+_31853919 | 0.06 |
ENSDART00000133886
|
atn1
|
atrophin 1 |
| chr6_-_57655030 | 0.06 |
ENSDART00000155438
|
cbfa2t2
|
core-binding factor, runt domain, alpha subunit 2; translocated to, 2 |
| chr2_+_54827391 | 0.06 |
ENSDART00000167269
|
twsg1a
|
twisted gastrulation BMP signaling modulator 1a |
| chr14_-_448182 | 0.05 |
ENSDART00000180018
|
FAT4
|
FAT atypical cadherin 4 |
| chr21_+_45733871 | 0.05 |
ENSDART00000187285
ENSDART00000193018 |
zgc:77058
|
zgc:77058 |
| chr20_+_27003713 | 0.05 |
ENSDART00000153232
|
si:dkey-177p2.18
|
si:dkey-177p2.18 |
| chr15_-_38202630 | 0.05 |
ENSDART00000183772
|
rhoga
|
ras homolog family member Ga |
| chr13_-_33822550 | 0.05 |
ENSDART00000143703
|
flrt3
|
fibronectin leucine rich transmembrane 3 |
| chr2_-_14798295 | 0.05 |
ENSDART00000143430
ENSDART00000145869 |
si:ch73-366i20.1
|
si:ch73-366i20.1 |
| chr23_-_21797517 | 0.05 |
ENSDART00000110041
|
lrrc38a
|
leucine rich repeat containing 38a |
| chr9_-_19610436 | 0.05 |
ENSDART00000108697
|
fgf9
|
fibroblast growth factor 9 |
| chr7_-_65236649 | 0.05 |
ENSDART00000185070
|
pkd1l2a
|
polycystic kidney disease 1 like 2a |
| chr7_-_37812176 | 0.05 |
ENSDART00000164485
|
adcy7
|
adenylate cyclase 7 |
| chr9_+_17862858 | 0.05 |
ENSDART00000166566
|
dgkh
|
diacylglycerol kinase, eta |
| chr15_-_13254480 | 0.05 |
ENSDART00000190499
|
zgc:172282
|
zgc:172282 |
| chr20_+_15600167 | 0.04 |
ENSDART00000171991
|
faslg
|
Fas ligand (TNF superfamily, member 6) |
| chr20_-_25481306 | 0.04 |
ENSDART00000182495
|
si:dkey-183n20.15
|
si:dkey-183n20.15 |
| chr2_-_37889111 | 0.04 |
ENSDART00000168939
ENSDART00000098529 |
mbl2
|
mannose binding lectin 2 |
| chr17_+_52823015 | 0.04 |
ENSDART00000160507
ENSDART00000186979 |
meis2a
|
Meis homeobox 2a |
| chr3_+_9588705 | 0.04 |
ENSDART00000172543
ENSDART00000104875 |
trap1
|
TNF receptor-associated protein 1 |
| chr19_+_4463389 | 0.04 |
ENSDART00000168805
|
kcnk9
|
potassium channel, subfamily K, member 9 |
| chr14_+_50807558 | 0.04 |
ENSDART00000183649
|
si:ch211-59p23.1
|
si:ch211-59p23.1 |
| chr20_-_54435287 | 0.04 |
ENSDART00000148632
|
yy1b
|
YY1 transcription factor b |
| chr16_-_29458806 | 0.04 |
ENSDART00000047931
|
lingo4b
|
leucine rich repeat and Ig domain containing 4b |
| chr5_-_28679135 | 0.04 |
ENSDART00000193585
|
tnc
|
tenascin C |
| chr12_+_31729498 | 0.04 |
ENSDART00000188546
ENSDART00000182562 ENSDART00000186147 |
RNF157
|
si:dkey-49c17.3 |
| chr3_-_61065055 | 0.04 |
ENSDART00000074199
|
cacng1b
|
calcium channel, voltage-dependent, gamma subunit 1b |
| chr19_-_24958393 | 0.04 |
ENSDART00000098592
|
si:ch211-195b13.6
|
si:ch211-195b13.6 |
| chr7_+_29952719 | 0.04 |
ENSDART00000173737
|
tpma
|
alpha-tropomyosin |
| chr9_-_33107237 | 0.04 |
ENSDART00000013918
|
casq2
|
calsequestrin 2 |
| chr12_-_22524388 | 0.04 |
ENSDART00000020942
|
shbg
|
sex hormone-binding globulin |
| chr5_+_57641554 | 0.04 |
ENSDART00000171309
ENSDART00000157992 ENSDART00000164742 |
cryabb
|
crystallin, alpha B, b |
| chr17_+_27134806 | 0.04 |
ENSDART00000151901
|
rps6ka1
|
ribosomal protein S6 kinase a, polypeptide 1 |
| chr13_+_7292061 | 0.04 |
ENSDART00000179504
|
CABZ01072077.1
|
Danio rerio neuroblast differentiation-associated protein AHNAK-like (LOC795051), mRNA. |
| chr15_-_7337148 | 0.04 |
ENSDART00000182568
|
SLC7A1 (1 of many)
|
high affinity cationic amino acid transporter 1 |
| chr2_-_25159309 | 0.04 |
ENSDART00000137290
|
si:dkey-223d7.6
|
si:dkey-223d7.6 |
| chr12_+_42574148 | 0.04 |
ENSDART00000157855
|
ebf3a
|
early B cell factor 3a |
| chr13_+_15800742 | 0.04 |
ENSDART00000146234
|
apopt1
|
apoptogenic 1, mitochondrial |
| chr23_+_27740788 | 0.04 |
ENSDART00000053871
|
dhh
|
desert hedgehog |
| chr8_+_15254564 | 0.04 |
ENSDART00000024433
|
slc5a9
|
solute carrier family 5 (sodium/sugar cotransporter), member 9 |
| chr3_+_13830599 | 0.04 |
ENSDART00000159282
|
ptger1b
|
prostaglandin E receptor 1b (subtype EP1) |
| chr5_-_54148271 | 0.04 |
ENSDART00000163034
|
arhgef15
|
Rho guanine nucleotide exchange factor (GEF) 15 |
| chr8_+_35664152 | 0.04 |
ENSDART00000144520
|
si:dkeyp-14d3.1
|
si:dkeyp-14d3.1 |
| chr15_+_31471808 | 0.03 |
ENSDART00000110078
|
or102-3
|
odorant receptor, family C, subfamily 102, member 3 |
| chr21_+_25375172 | 0.03 |
ENSDART00000078748
|
or132-5
|
odorant receptor, family H, subfamily 132, member 5 |
| chr7_-_65191937 | 0.03 |
ENSDART00000173234
|
pkd1l2a
|
polycystic kidney disease 1 like 2a |
| chr10_-_26179805 | 0.03 |
ENSDART00000174797
|
trim3b
|
tripartite motif containing 3b |
| chr12_+_28367557 | 0.03 |
ENSDART00000066294
|
cdk5r1b
|
cyclin-dependent kinase 5, regulatory subunit 1b (p35) |
| chr1_+_47585700 | 0.03 |
ENSDART00000153746
ENSDART00000084457 |
sh3pxd2aa
|
SH3 and PX domains 2Aa |
| chr22_-_9618867 | 0.03 |
ENSDART00000131246
|
si:ch73-286h23.4
|
si:ch73-286h23.4 |
| chr1_+_44992207 | 0.03 |
ENSDART00000172357
|
tmem132a
|
transmembrane protein 132A |
| chr3_-_32170850 | 0.03 |
ENSDART00000055307
ENSDART00000157366 |
tnnt1
|
troponin T type 1 (skeletal, slow) |
| chr25_-_7686201 | 0.03 |
ENSDART00000157267
ENSDART00000155094 |
si:ch211-286c4.6
|
si:ch211-286c4.6 |
| chr1_-_55810730 | 0.03 |
ENSDART00000100551
|
zgc:136908
|
zgc:136908 |
| chr21_-_8370500 | 0.03 |
ENSDART00000055328
|
nek6
|
NIMA-related kinase 6 |
| chr25_+_19681662 | 0.03 |
ENSDART00000104347
|
si:dkeyp-110c12.3
|
si:dkeyp-110c12.3 |
| chr12_-_20362041 | 0.03 |
ENSDART00000184145
ENSDART00000105952 |
aqp8a.2
|
aquaporin 8a, tandem duplicate 2 |
| chr7_+_29952169 | 0.03 |
ENSDART00000173540
ENSDART00000173940 ENSDART00000173906 ENSDART00000173772 ENSDART00000173506 ENSDART00000039657 |
tpma
|
alpha-tropomyosin |
| chr4_-_8014463 | 0.03 |
ENSDART00000036153
|
ccdc3a
|
coiled-coil domain containing 3a |
| chr16_-_25606889 | 0.03 |
ENSDART00000077447
ENSDART00000131528 |
zgc:110410
|
zgc:110410 |
| chr15_-_37425468 | 0.03 |
ENSDART00000059630
|
si:ch211-113j13.2
|
si:ch211-113j13.2 |
| chr22_-_24285432 | 0.03 |
ENSDART00000164083
|
si:ch211-117l17.4
|
si:ch211-117l17.4 |
| chr2_+_17181777 | 0.03 |
ENSDART00000112063
|
ptger4c
|
prostaglandin E receptor 4 (subtype EP4) c |
| chr6_+_13726844 | 0.03 |
ENSDART00000055833
|
wnt6a
|
wingless-type MMTV integration site family, member 6a |
| chr20_+_48543492 | 0.03 |
ENSDART00000175480
|
LO018254.1
|
|
| chr2_-_44720551 | 0.03 |
ENSDART00000146380
|
map6d1
|
MAP6 domain containing 1 |
| chr13_-_45811137 | 0.03 |
ENSDART00000190512
|
fndc5a
|
fibronectin type III domain containing 5a |
| chr15_+_33851280 | 0.03 |
ENSDART00000162308
|
inpp5ka
|
inositol polyphosphate-5-phosphatase Ka |
| chr22_-_5006119 | 0.03 |
ENSDART00000187998
|
rx1
|
retinal homeobox gene 1 |
| chr3_+_23692462 | 0.03 |
ENSDART00000145934
|
hoxb7a
|
homeobox B7a |
| chr23_-_1144425 | 0.03 |
ENSDART00000098271
|
psmb11b
|
proteasome subunit beta11b |
| chr12_-_44010532 | 0.03 |
ENSDART00000183875
|
si:ch211-182p11.1
|
si:ch211-182p11.1 |
| chr23_-_1587955 | 0.03 |
ENSDART00000136037
|
fndc7b
|
fibronectin type III domain containing 7b |
| chr10_+_44437080 | 0.03 |
ENSDART00000181517
ENSDART00000054153 |
denr
|
density-regulated protein |
| chr25_-_27541621 | 0.03 |
ENSDART00000130678
|
spam1
|
sperm adhesion molecule 1 |
| chr6_-_49634787 | 0.03 |
ENSDART00000188538
|
BX323459.1
|
|
| chr16_-_25606235 | 0.03 |
ENSDART00000192741
|
zgc:110410
|
zgc:110410 |
| chr24_-_39858710 | 0.03 |
ENSDART00000134251
|
slc12a7b
|
solute carrier family 12 (potassium/chloride transporter), member 7b |
| chr7_+_34786591 | 0.03 |
ENSDART00000173700
|
si:dkey-148a17.5
|
si:dkey-148a17.5 |
| chr5_-_32158198 | 0.03 |
ENSDART00000187215
ENSDART00000137503 |
m17
|
IL-6 subfamily cytokine M17 |
| chr21_+_5169154 | 0.03 |
ENSDART00000102559
|
zgc:122979
|
zgc:122979 |
| chr5_-_67145505 | 0.03 |
ENSDART00000011295
|
rom1a
|
retinal outer segment membrane protein 1a |
| chr4_+_54519511 | 0.03 |
ENSDART00000161653
|
znf974
|
zinc finger protein 974 |
| chr2_-_37883256 | 0.03 |
ENSDART00000035685
|
hbl4
|
hexose-binding lectin 4 |
| chr14_+_25464681 | 0.03 |
ENSDART00000067500
ENSDART00000187601 |
si:dkey-280e21.3
|
si:dkey-280e21.3 |
| chr25_+_31222069 | 0.03 |
ENSDART00000159373
|
tnni2a.1
|
troponin I type 2a (skeletal, fast), tandem duplicate 1 |
| chr4_+_74396786 | 0.03 |
ENSDART00000127501
ENSDART00000174347 |
zmp:0000001020
|
zmp:0000001020 |
| chr23_+_4324625 | 0.03 |
ENSDART00000146302
ENSDART00000136792 ENSDART00000135027 ENSDART00000179819 |
sgk2a
|
serum/glucocorticoid regulated kinase 2a |
| chr12_-_1951233 | 0.03 |
ENSDART00000005676
ENSDART00000127937 |
sox9a
|
SRY (sex determining region Y)-box 9a |
| chr18_+_33100606 | 0.03 |
ENSDART00000003907
|
si:ch211-229c8.13
|
si:ch211-229c8.13 |
| chr8_-_53548145 | 0.03 |
ENSDART00000180665
|
parapinopsina
|
parapinopsin a |
| chr11_+_40043569 | 0.03 |
ENSDART00000130278
|
si:dkey-264d12.1
|
si:dkey-264d12.1 |
| chr24_-_33291784 | 0.03 |
ENSDART00000124938
|
si:ch1073-406l10.2
|
si:ch1073-406l10.2 |
| chr7_+_16963091 | 0.03 |
ENSDART00000173770
|
nav2a
|
neuron navigator 2a |
| chr19_+_17642356 | 0.02 |
ENSDART00000176431
|
CR382334.1
|
|
| chr14_+_6159162 | 0.02 |
ENSDART00000128638
|
bscl2l
|
Bernardinelli-Seip congenital lipodystrophy 2, like |
| chr16_-_41840668 | 0.02 |
ENSDART00000146150
|
si:dkey-199f5.4
|
si:dkey-199f5.4 |
| chr5_-_6508250 | 0.02 |
ENSDART00000060535
|
crybb3
|
crystallin, beta B3 |
| chr16_-_41807397 | 0.02 |
ENSDART00000138577
|
si:dkey-199f5.7
|
si:dkey-199f5.7 |
| chr7_+_35268880 | 0.02 |
ENSDART00000182231
|
dpep2
|
dipeptidase 2 |
| chr19_+_23298037 | 0.02 |
ENSDART00000183948
|
irgf1
|
immunity-related GTPase family, f1 |
| chr16_-_52916248 | 0.02 |
ENSDART00000113931
|
zdhhc11
|
zinc finger, DHHC-type containing 11 |
| chr15_-_669476 | 0.02 |
ENSDART00000153687
ENSDART00000030603 |
si:ch211-210b2.2
|
si:ch211-210b2.2 |
| chr15_-_11956981 | 0.02 |
ENSDART00000164163
|
si:dkey-202l22.3
|
si:dkey-202l22.3 |
| chr22_+_5123479 | 0.02 |
ENSDART00000111822
ENSDART00000081910 |
loxl5a
|
lysyl oxidase-like 5a |
| chr5_+_39537195 | 0.02 |
ENSDART00000051256
|
fgf5
|
fibroblast growth factor 5 |
| chr24_-_30862168 | 0.02 |
ENSDART00000168540
|
ptbp2a
|
polypyrimidine tract binding protein 2a |
| chr3_-_53533128 | 0.02 |
ENSDART00000183591
|
notch3
|
notch 3 |
| chr25_-_3087556 | 0.02 |
ENSDART00000193249
|
best1
|
bestrophin 1 |
| chr8_+_18101608 | 0.02 |
ENSDART00000140941
|
glis1b
|
GLIS family zinc finger 1b |
| chr1_+_32791920 | 0.02 |
ENSDART00000109369
|
zgc:174320
|
zgc:174320 |
| chr7_+_26172682 | 0.02 |
ENSDART00000173631
|
si:ch211-196f2.7
|
si:ch211-196f2.7 |
| chr19_+_47311869 | 0.02 |
ENSDART00000136647
|
ext1c
|
exostoses (multiple) 1c |
| chr10_-_34867401 | 0.02 |
ENSDART00000145545
|
dclk1a
|
doublecortin-like kinase 1a |
| chr23_-_27875140 | 0.02 |
ENSDART00000143662
|
ankrd33aa
|
ankyrin repeat domain 33Aa |
| chr12_+_31729075 | 0.02 |
ENSDART00000152973
|
RNF157
|
si:dkey-49c17.3 |
| chr17_-_24890843 | 0.02 |
ENSDART00000184984
ENSDART00000135569 ENSDART00000193661 |
gale
|
UDP-galactose-4-epimerase |
| chr17_-_12926456 | 0.02 |
ENSDART00000044126
|
INSM2
|
INSM transcriptional repressor 2 |
Network of associatons between targets according to the STRING database.
First level regulatory network of hic1l+hic2
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0032289 | central nervous system myelin formation(GO:0032289) |
| 0.1 | 0.5 | GO:0006167 | AMP biosynthetic process(GO:0006167) |
| 0.0 | 0.1 | GO:1903358 | regulation of Golgi organization(GO:1903358) |
| 0.0 | 0.2 | GO:0000448 | cleavage in ITS2 between 5.8S rRNA and LSU-rRNA of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000448) |
| 0.0 | 0.2 | GO:0070207 | protein homotrimerization(GO:0070207) |
| 0.0 | 0.1 | GO:0035994 | response to muscle stretch(GO:0035994) |
| 0.0 | 0.1 | GO:0071586 | CAAX-box protein processing(GO:0071586) CAAX-box protein maturation(GO:0080120) |
| 0.0 | 0.1 | GO:0032801 | receptor catabolic process(GO:0032801) |
| 0.0 | 0.2 | GO:0036372 | opsin transport(GO:0036372) |
| 0.0 | 0.1 | GO:0002025 | vasodilation by norepinephrine-epinephrine involved in regulation of systemic arterial blood pressure(GO:0002025) |
| 0.0 | 0.1 | GO:0002074 | extraocular skeletal muscle development(GO:0002074) |
| 0.0 | 0.1 | GO:0048389 | intermediate mesoderm development(GO:0048389) |
| 0.0 | 0.1 | GO:0021767 | mammillary body development(GO:0021767) |
| 0.0 | 0.1 | GO:0045730 | respiratory burst(GO:0045730) |
| 0.0 | 0.1 | GO:0044246 | regulation of collagen metabolic process(GO:0010712) regulation of collagen biosynthetic process(GO:0032965) regulation of multicellular organismal metabolic process(GO:0044246) |
| 0.0 | 0.0 | GO:0008582 | regulation of synaptic growth at neuromuscular junction(GO:0008582) axon regeneration at neuromuscular junction(GO:0014814) positive regulation of synaptic growth at neuromuscular junction(GO:0045887) |
| 0.0 | 0.0 | GO:0010882 | regulation of cardiac muscle contraction by calcium ion signaling(GO:0010882) |
| 0.0 | 0.1 | GO:0070973 | protein localization to endoplasmic reticulum exit site(GO:0070973) |
| 0.0 | 0.2 | GO:0043951 | negative regulation of cAMP-mediated signaling(GO:0043951) |
| 0.0 | 0.4 | GO:0032543 | mitochondrial translation(GO:0032543) |
| 0.0 | 0.0 | GO:0051228 | mitotic spindle disassembly(GO:0051228) spindle disassembly(GO:0051230) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0090443 | FAR/SIN/STRIPAK complex(GO:0090443) |
| 0.0 | 0.1 | GO:0032777 | Piccolo NuA4 histone acetyltransferase complex(GO:0032777) |
| 0.0 | 0.2 | GO:0043625 | delta DNA polymerase complex(GO:0043625) |
| 0.0 | 0.1 | GO:0016533 | cyclin-dependent protein kinase 5 holoenzyme complex(GO:0016533) |
| 0.0 | 0.3 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.0 | 0.2 | GO:0036038 | MKS complex(GO:0036038) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0016842 | amidine-lyase activity(GO:0016842) |
| 0.0 | 0.2 | GO:0044736 | acid-sensing ion channel activity(GO:0044736) |
| 0.0 | 0.2 | GO:0060182 | apelin receptor activity(GO:0060182) |
| 0.0 | 0.1 | GO:0051380 | norepinephrine binding(GO:0051380) |
| 0.0 | 0.2 | GO:0016744 | transferase activity, transferring aldehyde or ketonic groups(GO:0016744) |
| 0.0 | 0.2 | GO:0004960 | thromboxane receptor activity(GO:0004960) |
| 0.0 | 0.1 | GO:0016165 | linoleate 13S-lipoxygenase activity(GO:0016165) |
| 0.0 | 0.2 | GO:0051731 | polynucleotide 5'-hydroxyl-kinase activity(GO:0051731) |
| 0.0 | 0.1 | GO:0039706 | co-receptor binding(GO:0039706) |
| 0.0 | 0.1 | GO:0050253 | retinyl-palmitate esterase activity(GO:0050253) |
| 0.0 | 0.1 | GO:0004526 | ribonuclease P activity(GO:0004526) |
| 0.0 | 0.2 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 0.0 | 0.1 | GO:0061575 | cyclin-dependent protein serine/threonine kinase activator activity(GO:0061575) |
| 0.0 | 0.1 | GO:0052650 | NADP-retinol dehydrogenase activity(GO:0052650) |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | REACTOME PURINE RIBONUCLEOSIDE MONOPHOSPHATE BIOSYNTHESIS | Genes involved in Purine ribonucleoside monophosphate biosynthesis |
| 0.0 | 0.1 | REACTOME PROSTANOID LIGAND RECEPTORS | Genes involved in Prostanoid ligand receptors |
| 0.0 | 0.4 | REACTOME POST CHAPERONIN TUBULIN FOLDING PATHWAY | Genes involved in Post-chaperonin tubulin folding pathway |
| 0.0 | 0.2 | REACTOME REMOVAL OF THE FLAP INTERMEDIATE FROM THE C STRAND | Genes involved in Removal of the Flap Intermediate from the C-strand |
| 0.0 | 0.1 | REACTOME SIGNALING BY ACTIVATED POINT MUTANTS OF FGFR1 | Genes involved in Signaling by activated point mutants of FGFR1 |