Project
PRJNA438478: RNAseq of wild type zebrafish germline
Navigation
Downloads
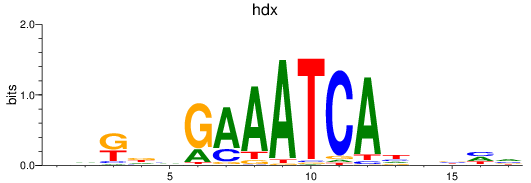
Results for hdx
Z-value: 0.63
Transcription factors associated with hdx
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
hdx
|
ENSDARG00000079382 | highly divergent homeobox |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| hdx | dr11_v1_chr14_-_6225336_6225336 | 0.41 | 9.5e-02 | Click! |
Activity profile of hdx motif
Sorted Z-values of hdx motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_+_24382651 | 2.64 |
ENSDART00000123789
|
qdprb2
|
quinoid dihydropteridine reductase b2 |
| chr17_-_2596125 | 1.54 |
ENSDART00000175740
|
zp3.2
|
zona pellucida glycoprotein 3, tandem duplicate 2 |
| chr2_+_37295088 | 1.48 |
ENSDART00000056519
|
gpr160
|
G protein-coupled receptor 160 |
| chr3_-_3413669 | 1.37 |
ENSDART00000113517
ENSDART00000179861 ENSDART00000115331 |
zgc:171446
|
zgc:171446 |
| chr9_+_8364553 | 1.21 |
ENSDART00000190713
|
si:dkey-90l23.2
|
si:dkey-90l23.2 |
| chr22_+_11972786 | 1.06 |
ENSDART00000105788
|
mgat5
|
mannosyl (alpha-1,6-)-glycoprotein beta-1,6-N-acetyl-glucosaminyltransferase |
| chr24_-_27409599 | 1.02 |
ENSDART00000041770
|
ccl34b.3
|
chemokine (C-C motif) ligand 34b, duplicate 3 |
| chr3_-_53092509 | 0.91 |
ENSDART00000062081
|
lpar2a
|
lysophosphatidic acid receptor 2a |
| chr17_-_6536305 | 0.86 |
ENSDART00000154855
|
cenpo
|
centromere protein O |
| chr23_+_19655301 | 0.79 |
ENSDART00000104441
ENSDART00000135269 |
abhd6b
|
abhydrolase domain containing 6b |
| chr10_-_36793412 | 0.68 |
ENSDART00000185966
|
dhrs13a.2
|
dehydrogenase/reductase (SDR family) member 13a, tandem duplicate 2 |
| chr17_-_6536466 | 0.66 |
ENSDART00000188735
|
cenpo
|
centromere protein O |
| chr19_-_20270178 | 0.61 |
ENSDART00000144891
ENSDART00000090883 |
gpnmb
|
glycoprotein (transmembrane) nmb |
| chr9_-_12888082 | 0.60 |
ENSDART00000133135
ENSDART00000134415 |
si:ch211-167j6.3
|
si:ch211-167j6.3 |
| chr10_-_7734939 | 0.59 |
ENSDART00000075563
|
snx24
|
sorting nexin 24 |
| chr25_-_4313699 | 0.53 |
ENSDART00000154038
|
syt7a
|
synaptotagmin VIIa |
| chr18_-_14691727 | 0.53 |
ENSDART00000010129
|
pdf
|
peptide deformylase, mitochondrial |
| chr21_-_2958422 | 0.50 |
ENSDART00000174091
|
zgc:194215
|
zgc:194215 |
| chr5_-_30535327 | 0.50 |
ENSDART00000040328
|
h2afx
|
H2A histone family, member X |
| chr21_-_7940043 | 0.50 |
ENSDART00000099733
ENSDART00000136671 |
f2rl1.1
|
coagulation factor II (thrombin) receptor-like 1, tandem duplicate 1 |
| chr7_+_31319876 | 0.47 |
ENSDART00000187611
|
fam189a1
|
family with sequence similarity 189, member A1 |
| chr6_+_21005725 | 0.47 |
ENSDART00000041370
|
cx44.2
|
connexin 44.2 |
| chr5_-_18007077 | 0.47 |
ENSDART00000129878
|
zdhhc8b
|
zinc finger, DHHC-type containing 8b |
| chr21_-_2322102 | 0.46 |
ENSDART00000162867
|
zgc:66483
|
zgc:66483 |
| chr5_+_57210237 | 0.46 |
ENSDART00000167660
|
pja2
|
praja ring finger ubiquitin ligase 2 |
| chr1_+_59314675 | 0.45 |
ENSDART00000161872
ENSDART00000160658 ENSDART00000169792 ENSDART00000160735 |
parn
|
poly(A)-specific ribonuclease (deadenylation nuclease) |
| chr12_+_30788912 | 0.45 |
ENSDART00000160422
|
aldh18a1
|
aldehyde dehydrogenase 18 family, member A1 |
| chr9_+_25840720 | 0.44 |
ENSDART00000024572
|
gtdc1
|
glycosyltransferase-like domain containing 1 |
| chr23_-_478201 | 0.44 |
ENSDART00000140749
|
si:ch73-181d5.4
|
si:ch73-181d5.4 |
| chr13_-_9841806 | 0.42 |
ENSDART00000101949
|
sfxn4
|
sideroflexin 4 |
| chr10_-_9346360 | 0.41 |
ENSDART00000039536
|
mrrf
|
mitochondrial ribosome recycling factor |
| chr5_-_63302944 | 0.41 |
ENSDART00000047110
|
gsnb
|
gelsolin b |
| chr14_+_26719691 | 0.41 |
ENSDART00000078522
ENSDART00000172927 |
eef1g
|
eukaryotic translation elongation factor 1 gamma |
| chr23_+_37579107 | 0.41 |
ENSDART00000169376
|
plekhg5b
|
pleckstrin homology domain containing, family G (with RhoGef domain) member 5b |
| chr10_-_18492617 | 0.41 |
ENSDART00000179936
ENSDART00000193799 ENSDART00000193198 |
si:dkey-28o19.1
|
si:dkey-28o19.1 |
| chr21_+_17051478 | 0.40 |
ENSDART00000047201
ENSDART00000161650 ENSDART00000167298 |
atp2a2a
|
ATPase sarcoplasmic/endoplasmic reticulum Ca2+ transporting 2a |
| chr12_-_1405228 | 0.39 |
ENSDART00000152127
|
MED9
|
si:ch73-105m5.1 |
| chr24_-_11057305 | 0.39 |
ENSDART00000186494
|
asap1b
|
ArfGAP with SH3 domain, ankyrin repeat and PH domain 1b |
| chr9_-_5318873 | 0.38 |
ENSDART00000129308
|
ACVR1C
|
activin A receptor type 1C |
| chr12_+_23912074 | 0.38 |
ENSDART00000152864
|
svila
|
supervillin a |
| chr1_-_8928841 | 0.38 |
ENSDART00000103652
|
grin2ab
|
glutamate receptor, ionotropic, N-methyl D-aspartate 2A, b |
| chr22_-_9896180 | 0.35 |
ENSDART00000138343
|
znf990
|
zinc finger protein 990 |
| chr22_-_23267893 | 0.35 |
ENSDART00000105613
|
si:dkey-121a9.3
|
si:dkey-121a9.3 |
| chr17_-_6535941 | 0.35 |
ENSDART00000109249
|
cenpo
|
centromere protein O |
| chr1_-_23268569 | 0.34 |
ENSDART00000143948
|
rfc1
|
replication factor C (activator 1) 1 |
| chr4_+_77060861 | 0.33 |
ENSDART00000174271
ENSDART00000174393 ENSDART00000150450 |
si:dkey-240n22.8
|
si:dkey-240n22.8 |
| chr8_-_16592491 | 0.32 |
ENSDART00000101655
|
calr
|
calreticulin |
| chr15_-_6946286 | 0.31 |
ENSDART00000019330
|
ech1
|
enoyl CoA hydratase 1, peroxisomal |
| chr16_+_19029297 | 0.31 |
ENSDART00000115263
ENSDART00000114954 |
rapgef5b
|
Rap guanine nucleotide exchange factor (GEF) 5b |
| chr6_-_6295513 | 0.30 |
ENSDART00000092172
|
mtif2
|
mitochondrial translational initiation factor 2 |
| chr2_+_45696743 | 0.29 |
ENSDART00000114225
ENSDART00000169279 |
CU467828.1
|
|
| chr22_+_2100774 | 0.28 |
ENSDART00000106436
|
si:dkey-1b17.9
|
si:dkey-1b17.9 |
| chr19_+_26424734 | 0.28 |
ENSDART00000187402
|
npr1a
|
natriuretic peptide receptor 1a |
| chr6_+_18569453 | 0.27 |
ENSDART00000171338
|
rhot1b
|
ras homolog family member T1 |
| chr1_-_29761086 | 0.27 |
ENSDART00000136760
|
alg11
|
asparagine-linked glycosylation 11 (alpha-1,2-mannosyltransferase) |
| chr10_-_22797959 | 0.27 |
ENSDART00000183269
|
pcolcea
|
procollagen C-endopeptidase enhancer a |
| chr19_+_19989380 | 0.26 |
ENSDART00000142841
|
osbpl3a
|
oxysterol binding protein-like 3a |
| chr24_+_9881219 | 0.26 |
ENSDART00000036204
|
cdv3
|
carnitine deficiency-associated gene expressed in ventricle 3 |
| chr14_+_34495216 | 0.26 |
ENSDART00000147756
|
wnt8a
|
wingless-type MMTV integration site family, member 8a |
| chr8_+_47099033 | 0.26 |
ENSDART00000142979
|
arhgef16
|
Rho guanine nucleotide exchange factor (GEF) 16 |
| chr5_+_50953240 | 0.25 |
ENSDART00000148501
ENSDART00000149892 ENSDART00000190312 |
col4a3bpa
|
collagen, type IV, alpha 3 (Goodpasture antigen) binding protein a |
| chr2_-_40890004 | 0.25 |
ENSDART00000191746
|
uggt1
|
UDP-glucose glycoprotein glucosyltransferase 1 |
| chr13_-_36582341 | 0.25 |
ENSDART00000137335
|
lgals3a
|
lectin, galactoside binding soluble 3a |
| chr7_+_56889879 | 0.25 |
ENSDART00000039810
|
myo9aa
|
myosin IXAa |
| chr4_-_16706776 | 0.24 |
ENSDART00000079461
|
dennd5b
|
DENN/MADD domain containing 5B |
| chr7_+_13684012 | 0.24 |
ENSDART00000056893
|
pdcd7
|
programmed cell death 7 |
| chr22_+_10201826 | 0.24 |
ENSDART00000006513
ENSDART00000132641 |
pdhb
|
pyruvate dehydrogenase E1 beta subunit |
| chr10_-_25570647 | 0.24 |
ENSDART00000134215
|
grik1a
|
glutamate receptor, ionotropic, kainate 1a |
| chr22_+_1786230 | 0.24 |
ENSDART00000169318
ENSDART00000164948 |
znf1154
|
zinc finger protein 1154 |
| chr5_+_27421025 | 0.24 |
ENSDART00000098590
|
cyb561a3a
|
cytochrome b561 family, member A3a |
| chr6_+_8652310 | 0.23 |
ENSDART00000105098
|
usp40
|
ubiquitin specific peptidase 40 |
| chr13_+_31550185 | 0.23 |
ENSDART00000127843
|
ppm1aa
|
protein phosphatase, Mg2+/Mn2+ dependent, 1Aa |
| chr2_-_40890264 | 0.23 |
ENSDART00000123886
|
uggt1
|
UDP-glucose glycoprotein glucosyltransferase 1 |
| chr3_-_27072179 | 0.23 |
ENSDART00000156556
|
atf7ip2
|
activating transcription factor 7 interacting protein 2 |
| chr10_+_29850330 | 0.23 |
ENSDART00000168898
|
hspa8
|
heat shock protein 8 |
| chr10_+_29849977 | 0.22 |
ENSDART00000180242
|
hspa8
|
heat shock protein 8 |
| chr25_-_23583101 | 0.22 |
ENSDART00000149107
ENSDART00000103704 ENSDART00000184903 |
nap1l4a
|
nucleosome assembly protein 1-like 4a |
| chr16_+_27383717 | 0.22 |
ENSDART00000132329
ENSDART00000136256 |
stx17
|
syntaxin 17 |
| chr22_-_5171362 | 0.21 |
ENSDART00000124889
|
tnfaip8l1
|
tumor necrosis factor, alpha-induced protein 8-like 1 |
| chr22_+_35131890 | 0.21 |
ENSDART00000003303
ENSDART00000130581 |
rnf13
|
ring finger protein 13 |
| chr14_+_11290828 | 0.21 |
ENSDART00000184078
|
rlim
|
ring finger protein, LIM domain interacting |
| chr19_-_2420990 | 0.21 |
ENSDART00000181498
|
TMEM196 (1 of many)
|
transmembrane protein 196 |
| chr19_-_26964348 | 0.21 |
ENSDART00000103955
|
zbtb12.2
|
zinc finger and BTB domain containing 12, tandem duplicate 2 |
| chr3_+_33918510 | 0.20 |
ENSDART00000055248
|
alkbh7
|
alkB homolog 7 |
| chr21_+_25071805 | 0.20 |
ENSDART00000078651
|
dixdc1b
|
DIX domain containing 1b |
| chr7_-_32875552 | 0.20 |
ENSDART00000132504
|
ano5b
|
anoctamin 5b |
| chr7_+_22399669 | 0.19 |
ENSDART00000144375
|
fgf11a
|
fibroblast growth factor 11a |
| chr14_+_30398546 | 0.19 |
ENSDART00000053925
|
mtmr7a
|
myotubularin related protein 7a |
| chr10_+_43406796 | 0.19 |
ENSDART00000184337
ENSDART00000183034 ENSDART00000180623 ENSDART00000132486 |
rasa1b
|
RAS p21 protein activator (GTPase activating protein) 1b |
| chr20_+_32552912 | 0.19 |
ENSDART00000009691
|
scml4
|
Scm polycomb group protein like 4 |
| chr4_-_13156971 | 0.19 |
ENSDART00000182164
|
grip1
|
glutamate receptor interacting protein 1 |
| chr19_-_3488860 | 0.19 |
ENSDART00000172520
|
hivep1
|
human immunodeficiency virus type I enhancer binding protein 1 |
| chr3_+_17951790 | 0.18 |
ENSDART00000164663
|
aclya
|
ATP citrate lyase a |
| chr20_-_46554440 | 0.18 |
ENSDART00000043298
ENSDART00000060680 |
fosab
|
v-fos FBJ murine osteosarcoma viral oncogene homolog Ab |
| chr17_+_6536152 | 0.18 |
ENSDART00000062952
ENSDART00000121789 |
slc5a6b
|
solute carrier family 5 (sodium/multivitamin and iodide cotransporter), member 6 |
| chr11_-_18557277 | 0.18 |
ENSDART00000103950
|
dido1
|
death inducer-obliterator 1 |
| chr21_+_43882274 | 0.18 |
ENSDART00000075672
|
sra1
|
steroid receptor RNA activator 1 |
| chr21_-_43952958 | 0.17 |
ENSDART00000039571
|
camk2a
|
calcium/calmodulin-dependent protein kinase II alpha |
| chr13_-_11986754 | 0.17 |
ENSDART00000164214
|
npm3
|
nucleophosmin/nucleoplasmin, 3 |
| chr4_-_16706941 | 0.17 |
ENSDART00000033053
ENSDART00000140510 ENSDART00000141119 |
dennd5b
|
DENN/MADD domain containing 5B |
| chr11_+_5565082 | 0.17 |
ENSDART00000112590
ENSDART00000183207 |
si:ch73-40i7.5
|
si:ch73-40i7.5 |
| chr16_+_4181105 | 0.16 |
ENSDART00000180921
|
yrdc
|
yrdC N(6)-threonylcarbamoyltransferase domain containing |
| chr25_+_22587306 | 0.16 |
ENSDART00000067479
|
stra6
|
stimulated by retinoic acid 6 |
| chr25_+_36333993 | 0.16 |
ENSDART00000184379
|
CR354435.2
|
|
| chr13_+_39297802 | 0.16 |
ENSDART00000133636
|
si:dkey-85a20.4
|
si:dkey-85a20.4 |
| chr18_+_808911 | 0.16 |
ENSDART00000172518
|
cox5ab
|
cytochrome c oxidase subunit Vab |
| chr10_-_36808348 | 0.16 |
ENSDART00000099320
|
dhrs13a.1
|
dehydrogenase/reductase (SDR family) member 13a, tandem duplicate 1 |
| chr17_+_6535694 | 0.15 |
ENSDART00000191681
|
slc5a6b
|
solute carrier family 5 (sodium/multivitamin and iodide cotransporter), member 6 |
| chr23_-_31633201 | 0.15 |
ENSDART00000143335
ENSDART00000053531 |
slc2a12
|
solute carrier family 2 (facilitated glucose transporter), member 12 |
| chr21_+_39336285 | 0.15 |
ENSDART00000139677
|
si:ch211-274p24.4
|
si:ch211-274p24.4 |
| chr2_+_34967210 | 0.15 |
ENSDART00000141796
|
astn1
|
astrotactin 1 |
| chr19_-_12324514 | 0.15 |
ENSDART00000122192
|
ncaldb
|
neurocalcin delta b |
| chr17_-_6618574 | 0.15 |
ENSDART00000184486
|
si:ch211-189e2.3
|
si:ch211-189e2.3 |
| chr17_-_20143946 | 0.15 |
ENSDART00000138911
|
actn2b
|
actinin, alpha 2b |
| chr9_-_12886108 | 0.15 |
ENSDART00000177283
|
ankzf1
|
ankyrin repeat and zinc finger domain containing 1 |
| chr17_-_6641535 | 0.15 |
ENSDART00000154540
ENSDART00000180384 |
si:ch211-189e2.3
|
si:ch211-189e2.3 |
| chr1_-_28950366 | 0.14 |
ENSDART00000110270
|
pwp2h
|
PWP2 periodic tryptophan protein homolog (yeast) |
| chr14_-_2213660 | 0.14 |
ENSDART00000162537
|
pcdh2ab3
|
protocadherin 2 alpha b 3 |
| chr3_+_52078798 | 0.14 |
ENSDART00000156882
|
si:dkey-88e12.3
|
si:dkey-88e12.3 |
| chr20_+_19562383 | 0.13 |
ENSDART00000103798
|
nme6
|
NME/NM23 nucleoside diphosphate kinase 6 |
| chr13_-_36581875 | 0.13 |
ENSDART00000113204
|
lgals3a
|
lectin, galactoside binding soluble 3a |
| chr4_+_1530287 | 0.13 |
ENSDART00000067446
|
slc38a4
|
solute carrier family 38, member 4 |
| chr16_-_26820634 | 0.13 |
ENSDART00000111156
|
pdp1
|
pyruvate dehyrogenase phosphatase catalytic subunit 1 |
| chr18_+_9810642 | 0.12 |
ENSDART00000143965
|
CACNA2D1
|
si:dkey-266j7.2 |
| chr10_-_8197049 | 0.12 |
ENSDART00000129467
|
dhx29
|
DEAH (Asp-Glu-Ala-His) box polypeptide 29 |
| chr16_+_51154918 | 0.12 |
ENSDART00000058290
|
dhdds
|
dehydrodolichyl diphosphate synthase |
| chr25_+_33072676 | 0.12 |
ENSDART00000182885
ENSDART00000181516 |
tln2b
|
talin 2b |
| chr18_+_814328 | 0.12 |
ENSDART00000157801
ENSDART00000188492 ENSDART00000184489 |
fam219b
|
family with sequence similarity 219, member B |
| chr4_-_38281909 | 0.11 |
ENSDART00000162448
|
si:ch211-229l10.9
|
si:ch211-229l10.9 |
| chr6_-_55399214 | 0.11 |
ENSDART00000168367
|
ctsa
|
cathepsin A |
| chr3_-_25086986 | 0.11 |
ENSDART00000050245
|
xpnpep3
|
X-prolyl aminopeptidase 3, mitochondrial |
| chr19_+_2685779 | 0.11 |
ENSDART00000160533
ENSDART00000097531 |
tomm7
|
translocase of outer mitochondrial membrane 7 homolog (yeast) |
| chr9_-_23152092 | 0.11 |
ENSDART00000180155
ENSDART00000186935 |
lypd6b
|
LY6/PLAUR domain containing 6B |
| chr19_-_10286228 | 0.11 |
ENSDART00000027316
|
cnot3b
|
CCR4-NOT transcription complex, subunit 3b |
| chr20_-_25644131 | 0.11 |
ENSDART00000138997
|
si:dkeyp-117h8.4
|
si:dkeyp-117h8.4 |
| chr1_-_11519934 | 0.10 |
ENSDART00000162060
|
sdk1b
|
sidekick cell adhesion molecule 1b |
| chr8_-_36370552 | 0.10 |
ENSDART00000097932
ENSDART00000148323 |
si:busm1-104n07.3
|
si:busm1-104n07.3 |
| chr13_-_24257631 | 0.10 |
ENSDART00000146524
|
urb2
|
URB2 ribosome biogenesis 2 homolog (S. cerevisiae) |
| chr20_+_14789148 | 0.10 |
ENSDART00000164761
|
tmed5
|
transmembrane p24 trafficking protein 5 |
| chr3_+_16724614 | 0.10 |
ENSDART00000182135
|
gys1
|
glycogen synthase 1 (muscle) |
| chr25_+_10811551 | 0.09 |
ENSDART00000167730
|
anpepb
|
alanyl (membrane) aminopeptidase b |
| chr8_-_33568964 | 0.09 |
ENSDART00000138921
|
mvb12bb
|
multivesicular body subunit 12Bb |
| chr10_+_44373349 | 0.09 |
ENSDART00000172191
|
snrnp35
|
small nuclear ribonucleoprotein 35 (U11/U12) |
| chr21_+_1320767 | 0.09 |
ENSDART00000187486
ENSDART00000193483 ENSDART00000182549 |
TCF4
|
transcription factor 4 |
| chr20_+_50052116 | 0.09 |
ENSDART00000047212
|
cpsf2
|
cleavage and polyadenylation specific factor 2 |
| chr14_-_8890437 | 0.09 |
ENSDART00000167242
|
ATG2A
|
si:ch73-45o6.2 |
| chr13_+_9842276 | 0.09 |
ENSDART00000041700
ENSDART00000141406 |
prdx3
|
peroxiredoxin 3 |
| chr6_-_31323984 | 0.09 |
ENSDART00000183660
ENSDART00000184146 ENSDART00000180156 |
dnajc6
|
DnaJ (Hsp40) homolog, subfamily C, member 6 |
| chr9_-_16877456 | 0.08 |
ENSDART00000161105
ENSDART00000160869 |
fbxl3a
|
F-box and leucine-rich repeat protein 3a |
| chr11_-_24532988 | 0.08 |
ENSDART00000067078
|
plekhg5a
|
pleckstrin homology domain containing, family G (with RhoGef domain) member 5a |
| chr10_-_32663759 | 0.08 |
ENSDART00000126727
|
atg101
|
autophagy related 101 |
| chr18_-_7112882 | 0.08 |
ENSDART00000176292
|
si:dkey-88e18.8
|
si:dkey-88e18.8 |
| chr3_+_25154078 | 0.08 |
ENSDART00000156973
|
si:ch211-256m1.8
|
si:ch211-256m1.8 |
| chr2_-_36497113 | 0.08 |
ENSDART00000109291
|
BX901889.4
|
|
| chr17_-_2834764 | 0.08 |
ENSDART00000024027
|
bdkrb1
|
bradykinin receptor B1 |
| chr21_+_21679086 | 0.08 |
ENSDART00000146225
|
or125-5
|
odorant receptor, family E, subfamily 125, member 5 |
| chr16_+_42471455 | 0.08 |
ENSDART00000166640
|
si:ch211-215k15.5
|
si:ch211-215k15.5 |
| chr10_+_36328702 | 0.08 |
ENSDART00000166676
|
or106-2
|
odorant receptor, family G, subfamily 106, member 2 |
| chr16_+_23495600 | 0.08 |
ENSDART00000021092
|
snx27b
|
sorting nexin family member 27b |
| chr8_+_44475793 | 0.07 |
ENSDART00000190118
|
si:ch73-211l2.3
|
si:ch73-211l2.3 |
| chr8_-_14067517 | 0.07 |
ENSDART00000140948
|
dedd
|
death effector domain containing |
| chr18_-_40481028 | 0.07 |
ENSDART00000134177
|
zgc:101040
|
zgc:101040 |
| chr21_-_2415808 | 0.07 |
ENSDART00000171179
|
si:ch211-241b2.5
|
si:ch211-241b2.5 |
| chr23_+_32029304 | 0.07 |
ENSDART00000185217
|
tpx2
|
TPX2, microtubule-associated, homolog (Xenopus laevis) |
| chr6_-_24143923 | 0.07 |
ENSDART00000157948
|
si:ch73-389b16.1
|
si:ch73-389b16.1 |
| chr18_-_6742287 | 0.07 |
ENSDART00000140752
|
miox
|
myo-inositol oxygenase |
| chr15_-_917274 | 0.07 |
ENSDART00000156624
|
si:dkey-77f5.15
|
si:dkey-77f5.15 |
| chr7_+_41146560 | 0.07 |
ENSDART00000143285
ENSDART00000173852 ENSDART00000174003 ENSDART00000038487 ENSDART00000173463 ENSDART00000166448 ENSDART00000052274 |
puf60b
|
poly-U binding splicing factor b |
| chr11_+_6752468 | 0.07 |
ENSDART00000187046
ENSDART00000182089 |
pde4cb
|
phosphodiesterase 4C, cAMP-specific b |
| chr24_-_14212521 | 0.07 |
ENSDART00000130825
|
xkr9
|
XK, Kell blood group complex subunit-related family, member 9 |
| chr10_+_9346383 | 0.07 |
ENSDART00000064975
ENSDART00000141869 |
rbm18
|
RNA binding motif protein 18 |
| chr7_+_31074636 | 0.07 |
ENSDART00000173805
|
tjp1a
|
tight junction protein 1a |
| chr18_-_17077596 | 0.07 |
ENSDART00000111821
|
il17c
|
interleukin 17c |
| chr2_-_47923803 | 0.07 |
ENSDART00000056306
|
ftr24
|
finTRIM family, member 24 |
| chr7_-_3011514 | 0.06 |
ENSDART00000169589
|
zgc:171695
|
zgc:171695 |
| chr9_+_30720048 | 0.06 |
ENSDART00000146115
|
klf12b
|
Kruppel-like factor 12b |
| chr19_+_23298037 | 0.06 |
ENSDART00000183948
|
irgf1
|
immunity-related GTPase family, f1 |
| chr2_-_51634431 | 0.06 |
ENSDART00000165568
|
pigr
|
polymeric immunoglobulin receptor |
| chr23_-_12788113 | 0.06 |
ENSDART00000103231
ENSDART00000146640 ENSDART00000180549 |
si:dkey-96f10.1
|
si:dkey-96f10.1 |
| chr14_+_12316581 | 0.06 |
ENSDART00000170115
ENSDART00000149757 |
si:ch211-125c23.3
cx31.7
|
si:ch211-125c23.3 connexin 31.7 |
| chr12_+_29173523 | 0.06 |
ENSDART00000153212
|
adgra1a
|
adhesion G protein-coupled receptor A1a |
| chr5_+_27267186 | 0.06 |
ENSDART00000182238
ENSDART00000087857 |
unc5db
|
unc-5 netrin receptor Db |
| chr16_+_26735894 | 0.06 |
ENSDART00000191605
ENSDART00000185175 |
rad54b
|
RAD54 homolog B (S. cerevisiae) |
| chr14_-_42997145 | 0.06 |
ENSDART00000172801
|
pcdh10b
|
protocadherin 10b |
| chr13_-_9895564 | 0.06 |
ENSDART00000169831
ENSDART00000142629 |
si:ch211-117n7.6
|
si:ch211-117n7.6 |
| chr3_-_33029938 | 0.06 |
ENSDART00000181730
|
CT583642.1
|
|
| chr15_+_5088210 | 0.06 |
ENSDART00000183423
|
mxf
|
myxovirus (influenza virus) resistance F |
| chr22_+_6843363 | 0.06 |
ENSDART00000126044
|
FO904898.3
|
|
| chr7_-_17268111 | 0.06 |
ENSDART00000169319
|
nitr10a
|
novel immune-type receptor 10a |
| chr15_+_31428332 | 0.06 |
ENSDART00000141429
|
or103-5
|
odorant receptor, family C, subfamily 103, member 5 |
| chr12_+_26877381 | 0.06 |
ENSDART00000087329
|
znf438
|
zinc finger protein 438 |
| chr7_+_49865049 | 0.06 |
ENSDART00000164165
|
slc1a2a
|
solute carrier family 1 (glial high affinity glutamate transporter), member 2a |
| chr17_+_21546993 | 0.06 |
ENSDART00000182387
|
chst15
|
carbohydrate (N-acetylgalactosamine 4-sulfate 6-O) sulfotransferase 15 |
| chr2_+_5300405 | 0.06 |
ENSDART00000019925
|
GNB4
|
G protein subunit beta 4 |
| chr20_+_2039518 | 0.06 |
ENSDART00000043157
|
CABZ01088134.1
|
|
| chr17_-_40110782 | 0.06 |
ENSDART00000126929
|
si:dkey-187k19.2
|
si:dkey-187k19.2 |
| chr18_-_19491162 | 0.06 |
ENSDART00000090413
|
snapc5
|
small nuclear RNA activating complex, polypeptide 5 |
| chr7_+_65387374 | 0.06 |
ENSDART00000188680
|
CLEC3A
|
C-type lectin domain family 3 member A |
| chr20_-_35052823 | 0.05 |
ENSDART00000153033
|
kif26bb
|
kinesin family member 26Bb |
| chr18_-_8885792 | 0.05 |
ENSDART00000143619
|
si:dkey-95h12.1
|
si:dkey-95h12.1 |
Network of associatons between targets according to the STRING database.
First level regulatory network of hdx
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 2.6 | GO:0046146 | tetrahydrobiopterin biosynthetic process(GO:0006729) tetrahydrobiopterin metabolic process(GO:0046146) |
| 0.2 | 0.9 | GO:0045647 | negative regulation of erythrocyte differentiation(GO:0045647) |
| 0.1 | 1.5 | GO:0007339 | binding of sperm to zona pellucida(GO:0007339) egg coat formation(GO:0035803) regulation of acrosome reaction(GO:0060046) positive regulation of acrosome reaction(GO:2000344) |
| 0.1 | 0.3 | GO:0021512 | spinal cord anterior/posterior patterning(GO:0021512) cell proliferation involved in pronephros development(GO:0039015) cell proliferation involved in kidney development(GO:0072111) |
| 0.1 | 0.3 | GO:0015887 | biotin transport(GO:0015878) pantothenate transmembrane transport(GO:0015887) coenzyme transport(GO:0051182) |
| 0.1 | 0.4 | GO:0048245 | eosinophil chemotaxis(GO:0048245) eosinophil migration(GO:0072677) |
| 0.1 | 0.5 | GO:0048260 | positive regulation of receptor-mediated endocytosis(GO:0048260) protein localization to early endosome(GO:1902946) |
| 0.1 | 0.5 | GO:0018206 | peptidyl-methionine modification(GO:0018206) |
| 0.1 | 0.5 | GO:0070493 | thrombin receptor signaling pathway(GO:0070493) |
| 0.1 | 1.1 | GO:0070654 | sensory epithelium regeneration(GO:0070654) epithelium regeneration(GO:1990399) |
| 0.0 | 0.2 | GO:0098529 | neuromuscular junction development, skeletal muscle fiber(GO:0098529) |
| 0.0 | 0.4 | GO:0006561 | proline biosynthetic process(GO:0006561) L-proline biosynthetic process(GO:0055129) |
| 0.0 | 0.8 | GO:0042574 | retinal metabolic process(GO:0042574) |
| 0.0 | 0.2 | GO:0006086 | acetyl-CoA biosynthetic process from pyruvate(GO:0006086) |
| 0.0 | 0.2 | GO:0035461 | vitamin transmembrane transport(GO:0035461) vitamin A transport(GO:0071938) vitamin A import(GO:0071939) |
| 0.0 | 0.5 | GO:0071712 | ER-associated misfolded protein catabolic process(GO:0071712) |
| 0.0 | 0.1 | GO:1904182 | regulation of pyruvate dehydrogenase activity(GO:1904182) positive regulation of pyruvate dehydrogenase activity(GO:1904184) |
| 0.0 | 0.5 | GO:0010738 | regulation of protein kinase A signaling(GO:0010738) |
| 0.0 | 0.8 | GO:0051014 | actin filament severing(GO:0051014) |
| 0.0 | 0.1 | GO:0016094 | polyprenol biosynthetic process(GO:0016094) |
| 0.0 | 0.1 | GO:1902592 | virion assembly(GO:0019068) virus maturation(GO:0019075) multi-organism membrane organization(GO:0044803) viral budding(GO:0046755) multi-organism organelle organization(GO:1902590) multi-organism membrane budding(GO:1902592) |
| 0.0 | 0.9 | GO:0002548 | monocyte chemotaxis(GO:0002548) |
| 0.0 | 0.2 | GO:0006085 | acetyl-CoA biosynthetic process(GO:0006085) |
| 0.0 | 0.2 | GO:0061026 | cardiac muscle tissue regeneration(GO:0061026) |
| 0.0 | 0.2 | GO:0046902 | regulation of mitochondrial membrane permeability(GO:0046902) |
| 0.0 | 0.4 | GO:0060079 | excitatory postsynaptic potential(GO:0060079) |
| 0.0 | 0.2 | GO:0046323 | glucose import(GO:0046323) |
| 0.0 | 0.3 | GO:0006490 | oligosaccharide-lipid intermediate biosynthetic process(GO:0006490) |
| 0.0 | 0.1 | GO:0016191 | synaptic vesicle uncoating(GO:0016191) |
| 0.0 | 0.2 | GO:0099638 | neurotransmitter receptor transport, endosome to postsynaptic membrane(GO:0098887) endosome to plasma membrane protein transport(GO:0099638) neurotransmitter receptor transport, endosome to plasma membrane(GO:0099639) |
| 0.0 | 0.6 | GO:0000289 | nuclear-transcribed mRNA poly(A) tail shortening(GO:0000289) |
| 0.0 | 0.2 | GO:0006450 | regulation of translational fidelity(GO:0006450) |
| 0.0 | 0.3 | GO:0051654 | establishment of mitochondrion localization, microtubule-mediated(GO:0034643) mitochondrion transport along microtubule(GO:0047497) establishment of mitochondrion localization(GO:0051654) |
| 0.0 | 0.2 | GO:1905150 | regulation of voltage-gated sodium channel activity(GO:1905150) |
| 0.0 | 0.2 | GO:0043162 | ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway(GO:0043162) |
| 0.0 | 0.0 | GO:0003250 | cell proliferation involved in heart valve morphogenesis(GO:0003249) regulation of cell proliferation involved in heart valve morphogenesis(GO:0003250) endocardial cell development(GO:0060958) cell proliferation involved in heart valve development(GO:2000793) |
| 0.0 | 0.3 | GO:0060038 | cardiac muscle cell proliferation(GO:0060038) |
| 0.0 | 0.5 | GO:0018230 | peptidyl-L-cysteine S-palmitoylation(GO:0018230) peptidyl-S-diacylglycerol-L-cysteine biosynthetic process from peptidyl-cysteine(GO:0018231) |
| 0.0 | 0.1 | GO:0006398 | mRNA 3'-end processing by stem-loop binding and cleavage(GO:0006398) |
| 0.0 | 0.1 | GO:0045948 | positive regulation of translational initiation(GO:0045948) |
| 0.0 | 0.1 | GO:0006660 | phosphatidylserine catabolic process(GO:0006660) monoacylglycerol metabolic process(GO:0046462) monoacylglycerol catabolic process(GO:0052651) |
| 0.0 | 0.2 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
| 0.0 | 0.0 | GO:1903232 | melanosome assembly(GO:1903232) |
| 0.0 | 0.5 | GO:0014059 | dopamine secretion(GO:0014046) regulation of dopamine secretion(GO:0014059) |
| 0.0 | 0.0 | GO:0015874 | norepinephrine transport(GO:0015874) |
| 0.0 | 0.1 | GO:0006384 | transcription initiation from RNA polymerase III promoter(GO:0006384) |
| 0.0 | 1.3 | GO:0010466 | negative regulation of peptidase activity(GO:0010466) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.9 | GO:0031511 | Mis6-Sim4 complex(GO:0031511) |
| 0.1 | 0.3 | GO:0097189 | apoptotic body(GO:0097189) |
| 0.0 | 0.4 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.0 | 0.2 | GO:0005967 | mitochondrial pyruvate dehydrogenase complex(GO:0005967) |
| 0.0 | 0.1 | GO:1904423 | dehydrodolichyl diphosphate synthase complex(GO:1904423) |
| 0.0 | 0.4 | GO:0017146 | NMDA selective glutamate receptor complex(GO:0017146) |
| 0.0 | 0.3 | GO:0031307 | integral component of mitochondrial outer membrane(GO:0031307) |
| 0.0 | 0.1 | GO:0034388 | Pwp2p-containing subcomplex of 90S preribosome(GO:0034388) |
| 0.0 | 0.3 | GO:0000813 | ESCRT I complex(GO:0000813) |
| 0.0 | 0.1 | GO:0005880 | nuclear microtubule(GO:0005880) |
| 0.0 | 0.3 | GO:0097038 | perinuclear endoplasmic reticulum(GO:0097038) |
| 0.0 | 0.4 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.0 | 0.1 | GO:0043540 | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase complex(GO:0043540) |
| 0.0 | 0.1 | GO:0005742 | mitochondrial outer membrane translocase complex(GO:0005742) |
| 0.0 | 0.8 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.0 | 0.2 | GO:0005751 | mitochondrial respiratory chain complex IV(GO:0005751) |
| 0.0 | 0.0 | GO:0031085 | BLOC-3 complex(GO:0031085) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 2.6 | GO:0004155 | 6,7-dihydropteridine reductase activity(GO:0004155) NADH binding(GO:0070404) |
| 0.1 | 1.5 | GO:0035804 | structural constituent of egg coat(GO:0035804) |
| 0.1 | 0.5 | GO:0003980 | UDP-glucose:glycoprotein glucosyltransferase activity(GO:0003980) |
| 0.1 | 0.3 | GO:0004377 | GDP-Man:Man3GlcNAc2-PP-Dol alpha-1,2-mannosyltransferase activity(GO:0004377) |
| 0.1 | 0.4 | GO:0016774 | phosphotransferase activity, carboxyl group as acceptor(GO:0016774) |
| 0.1 | 0.8 | GO:0052650 | NADP-retinol dehydrogenase activity(GO:0052650) |
| 0.1 | 0.3 | GO:0008523 | sodium-dependent multivitamin transmembrane transporter activity(GO:0008523) |
| 0.1 | 0.4 | GO:0019863 | IgE binding(GO:0019863) immunoglobulin binding(GO:0019865) |
| 0.1 | 0.4 | GO:0043023 | ribosomal large subunit binding(GO:0043023) |
| 0.1 | 0.9 | GO:0070915 | lysophosphatidic acid receptor activity(GO:0070915) |
| 0.1 | 0.5 | GO:0015057 | thrombin receptor activity(GO:0015057) |
| 0.0 | 0.2 | GO:0004739 | pyruvate dehydrogenase (acetyl-transferring) activity(GO:0004739) |
| 0.0 | 0.3 | GO:0016863 | intramolecular oxidoreductase activity, transposing C=C bonds(GO:0016863) |
| 0.0 | 0.1 | GO:0002094 | polyprenyltransferase activity(GO:0002094) |
| 0.0 | 0.3 | GO:0016941 | natriuretic peptide receptor activity(GO:0016941) |
| 0.0 | 0.1 | GO:0004741 | [pyruvate dehydrogenase (lipoamide)] phosphatase activity(GO:0004741) |
| 0.0 | 0.5 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.0 | 0.4 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.0 | 0.1 | GO:0004373 | glycogen (starch) synthase activity(GO:0004373) |
| 0.0 | 0.2 | GO:0034632 | retinol transporter activity(GO:0034632) |
| 0.0 | 0.4 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.0 | 0.2 | GO:0046912 | transferase activity, transferring acyl groups, acyl groups converted into alkyl on transfer(GO:0046912) |
| 0.0 | 0.1 | GO:0004185 | serine-type carboxypeptidase activity(GO:0004185) |
| 0.0 | 0.9 | GO:0048020 | CCR chemokine receptor binding(GO:0048020) |
| 0.0 | 0.4 | GO:0004972 | NMDA glutamate receptor activity(GO:0004972) |
| 0.0 | 0.1 | GO:0099602 | acetylcholine receptor regulator activity(GO:0030548) neurotransmitter receptor regulator activity(GO:0099602) |
| 0.0 | 0.2 | GO:0035259 | glucocorticoid receptor binding(GO:0035259) |
| 0.0 | 0.1 | GO:0008798 | beta-aspartyl-peptidase activity(GO:0008798) |
| 0.0 | 0.8 | GO:0005546 | phosphatidylinositol-4,5-bisphosphate binding(GO:0005546) |
| 0.0 | 1.1 | GO:0008375 | acetylglucosaminyltransferase activity(GO:0008375) |
| 0.0 | 0.5 | GO:0001786 | phosphatidylserine binding(GO:0001786) |
| 0.0 | 0.7 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 0.1 | GO:0004060 | arylamine N-acetyltransferase activity(GO:0004060) |
| 0.0 | 0.0 | GO:0005329 | dopamine transmembrane transporter activity(GO:0005329) dopamine:sodium symporter activity(GO:0005330) norepinephrine transmembrane transporter activity(GO:0005333) norepinephrine:sodium symporter activity(GO:0005334) |
| 0.0 | 0.1 | GO:0046920 | alpha-(1->3)-fucosyltransferase activity(GO:0046920) |
| 0.0 | 1.3 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 0.1 | GO:0015616 | DNA translocase activity(GO:0015616) |
| 0.0 | 0.5 | GO:0019706 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | PID ATM PATHWAY | ATM pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.4 | REACTOME DEPOSITION OF NEW CENPA CONTAINING NUCLEOSOMES AT THE CENTROMERE | Genes involved in Deposition of New CENPA-containing Nucleosomes at the Centromere |
| 0.1 | 0.2 | REACTOME UNBLOCKING OF NMDA RECEPTOR GLUTAMATE BINDING AND ACTIVATION | Genes involved in Unblocking of NMDA receptor, glutamate binding and activation |
| 0.0 | 0.8 | REACTOME CALNEXIN CALRETICULIN CYCLE | Genes involved in Calnexin/calreticulin cycle |
| 0.0 | 0.4 | REACTOME AMINO ACID SYNTHESIS AND INTERCONVERSION TRANSAMINATION | Genes involved in Amino acid synthesis and interconversion (transamination) |
| 0.0 | 1.1 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.0 | 0.4 | REACTOME SIGNALING BY NODAL | Genes involved in Signaling by NODAL |
| 0.0 | 0.5 | REACTOME GABA SYNTHESIS RELEASE REUPTAKE AND DEGRADATION | Genes involved in GABA synthesis, release, reuptake and degradation |
| 0.0 | 0.4 | REACTOME REGULATION OF PYRUVATE DEHYDROGENASE PDH COMPLEX | Genes involved in Regulation of pyruvate dehydrogenase (PDH) complex |
| 0.0 | 0.4 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.0 | 0.2 | REACTOME FACILITATIVE NA INDEPENDENT GLUCOSE TRANSPORTERS | Genes involved in Facilitative Na+-independent glucose transporters |