Project
PRJNA438478: RNAseq of wild type zebrafish germline
Navigation
Downloads
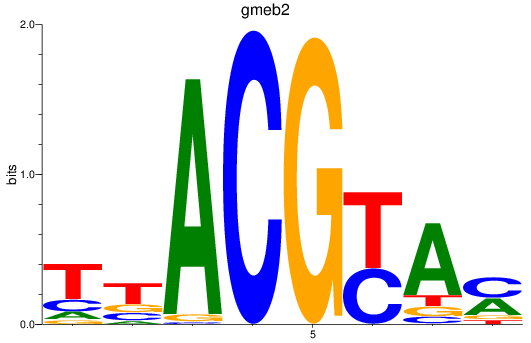
Results for gmeb2
Z-value: 0.43
Transcription factors associated with gmeb2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
gmeb2
|
ENSDARG00000093240 | si_ch73-302a13.2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| GMEB2 | dr11_v1_chr23_-_41824460_41824460 | 0.41 | 9.5e-02 | Click! |
Activity profile of gmeb2 motif
Sorted Z-values of gmeb2 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr3_-_49382896 | 0.60 |
ENSDART00000169115
|
si:ch73-167f10.1
|
si:ch73-167f10.1 |
| chr5_-_67365333 | 0.52 |
ENSDART00000133438
|
unga
|
uracil DNA glycosylase a |
| chr5_-_6567464 | 0.42 |
ENSDART00000184985
|
tnks1bp1
|
tankyrase 1 binding protein 1 |
| chr23_-_33558161 | 0.40 |
ENSDART00000018301
|
itga5
|
integrin, alpha 5 (fibronectin receptor, alpha polypeptide) |
| chr9_+_24920677 | 0.36 |
ENSDART00000037025
|
slc39a10
|
solute carrier family 39 (zinc transporter), member 10 |
| chr16_+_41015163 | 0.35 |
ENSDART00000058586
|
dek
|
DEK proto-oncogene |
| chr16_-_47427016 | 0.33 |
ENSDART00000074575
|
sept7b
|
septin 7b |
| chr24_+_37484661 | 0.31 |
ENSDART00000165125
ENSDART00000109221 |
wdr90
|
WD repeat domain 90 |
| chr8_+_13064750 | 0.31 |
ENSDART00000039878
|
sap30bp
|
SAP30 binding protein |
| chr15_+_46357080 | 0.30 |
ENSDART00000155571
ENSDART00000156541 |
wu:fb18f06
|
wu:fb18f06 |
| chr8_-_5267442 | 0.30 |
ENSDART00000155816
|
ppp2r2aa
|
protein phosphatase 2, regulatory subunit B, alpha a |
| chr3_+_32789605 | 0.29 |
ENSDART00000171895
|
tbc1d10b
|
TBC1 domain family, member 10b |
| chr17_-_6451801 | 0.28 |
ENSDART00000064700
|
fuca2
|
alpha-L-fucosidase 2 |
| chr12_+_11650146 | 0.26 |
ENSDART00000150191
|
waplb
|
WAPL cohesin release factor b |
| chr15_+_46356879 | 0.26 |
ENSDART00000154388
|
wu:fb18f06
|
wu:fb18f06 |
| chr12_+_17603528 | 0.25 |
ENSDART00000111565
|
pms2
|
PMS1 homolog 2, mismatch repair system component |
| chr20_+_38837238 | 0.25 |
ENSDART00000061334
|
ift172
|
intraflagellar transport 172 |
| chr2_+_2168547 | 0.25 |
ENSDART00000029347
|
higd1a
|
HIG1 hypoxia inducible domain family, member 1A |
| chr6_-_60079551 | 0.23 |
ENSDART00000154753
|
pmepa1
|
prostate transmembrane protein, androgen induced 1 |
| chr8_-_1219815 | 0.22 |
ENSDART00000016800
ENSDART00000149969 |
znf367
|
zinc finger protein 367 |
| chr2_+_56012016 | 0.22 |
ENSDART00000146160
ENSDART00000188702 |
loxl5b
|
lysyl oxidase-like 5b |
| chr14_-_22495604 | 0.22 |
ENSDART00000137167
|
si:ch211-107m4.1
|
si:ch211-107m4.1 |
| chr8_+_53311965 | 0.21 |
ENSDART00000130104
|
gnb1a
|
guanine nucleotide binding protein (G protein), beta polypeptide 1a |
| chr13_+_42309688 | 0.21 |
ENSDART00000158367
|
ide
|
insulin-degrading enzyme |
| chr8_+_47897734 | 0.20 |
ENSDART00000140266
|
mfn2
|
mitofusin 2 |
| chr11_+_11120532 | 0.20 |
ENSDART00000026135
ENSDART00000189872 |
ly75
|
lymphocyte antigen 75 |
| chr8_-_38022298 | 0.20 |
ENSDART00000067809
|
rab11fip1a
|
RAB11 family interacting protein 1 (class I) a |
| chr5_-_13076779 | 0.20 |
ENSDART00000192826
|
ypel1
|
yippee-like 1 |
| chr15_+_32798333 | 0.19 |
ENSDART00000162370
ENSDART00000166525 |
spartb
|
spartin b |
| chr20_-_30938184 | 0.19 |
ENSDART00000147359
ENSDART00000062552 |
wtap
|
WT1 associated protein |
| chr12_-_27588299 | 0.19 |
ENSDART00000178023
ENSDART00000066282 |
dhx8
|
DEAH (Asp-Glu-Ala-His) box polypeptide 8 |
| chr10_-_14545658 | 0.19 |
ENSDART00000057865
|
ier3ip1
|
immediate early response 3 interacting protein 1 |
| chr16_+_23397785 | 0.18 |
ENSDART00000148961
|
s100a10b
|
S100 calcium binding protein A10b |
| chr20_-_9199721 | 0.18 |
ENSDART00000064140
|
ylpm1
|
YLP motif containing 1 |
| chr11_-_41132296 | 0.18 |
ENSDART00000162944
|
dnajc11b
|
DnaJ (Hsp40) homolog, subfamily C, member 11b |
| chr4_+_23126558 | 0.18 |
ENSDART00000162859
|
mdm2
|
MDM2 oncogene, E3 ubiquitin protein ligase |
| chr10_-_41352502 | 0.17 |
ENSDART00000052971
ENSDART00000128156 |
rab11fip1b
|
RAB11 family interacting protein 1 (class I) b |
| chr16_-_47426482 | 0.17 |
ENSDART00000148631
ENSDART00000149723 |
sept7b
|
septin 7b |
| chr5_-_2112030 | 0.17 |
ENSDART00000091932
|
gusb
|
glucuronidase, beta |
| chr22_-_26353916 | 0.17 |
ENSDART00000077958
|
capn2b
|
calpain 2, (m/II) large subunit b |
| chr6_-_25201810 | 0.17 |
ENSDART00000168683
|
lrrc8c
|
leucine rich repeat containing 8 VRAC subunit C |
| chr1_-_31516091 | 0.17 |
ENSDART00000139828
ENSDART00000146567 |
cenpk
|
centromere protein K |
| chr25_+_21098990 | 0.16 |
ENSDART00000017488
|
rad52
|
RAD52 homolog, DNA repair protein |
| chr14_+_8947282 | 0.16 |
ENSDART00000047993
|
rps6kal
|
ribosomal protein S6 kinase a, like |
| chr16_-_29557338 | 0.16 |
ENSDART00000058888
|
hormad1
|
HORMA domain containing 1 |
| chr10_-_41907213 | 0.15 |
ENSDART00000167004
|
kdm2bb
|
lysine (K)-specific demethylase 2Bb |
| chr25_+_21098675 | 0.15 |
ENSDART00000079012
|
rad52
|
RAD52 homolog, DNA repair protein |
| chr11_+_2506516 | 0.15 |
ENSDART00000130886
ENSDART00000189767 |
NABP2
|
si:ch73-190f16.2 |
| chr13_+_30035253 | 0.15 |
ENSDART00000181303
ENSDART00000057525 ENSDART00000136622 |
dnajb12a
|
DnaJ (Hsp40) homolog, subfamily B, member 12a |
| chr5_-_51198430 | 0.15 |
ENSDART00000132503
ENSDART00000097473 ENSDART00000165870 |
snapc4
|
small nuclear RNA activating complex, polypeptide 4 |
| chr4_+_9011825 | 0.15 |
ENSDART00000058007
|
samm50l
|
sorting and assembly machinery component 50 homolog, like |
| chr25_-_28668776 | 0.15 |
ENSDART00000126490
|
fnbp4
|
formin binding protein 4 |
| chr10_+_6884123 | 0.15 |
ENSDART00000149095
ENSDART00000148772 ENSDART00000149334 |
ddx4
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 4 |
| chr24_+_20968308 | 0.15 |
ENSDART00000130568
|
drd3
|
dopamine receptor D3 |
| chr1_+_23274710 | 0.15 |
ENSDART00000036448
|
lias
|
lipoic acid synthetase |
| chr6_-_37745508 | 0.15 |
ENSDART00000078316
|
nipa2
|
non imprinted in Prader-Willi/Angelman syndrome 2 (human) |
| chr8_-_4760723 | 0.14 |
ENSDART00000064201
|
cdc45
|
CDC45 cell division cycle 45 homolog (S. cerevisiae) |
| chr3_+_53156813 | 0.14 |
ENSDART00000114343
|
brd4
|
bromodomain containing 4 |
| chr19_+_46259619 | 0.14 |
ENSDART00000158032
|
grinaa
|
glutamate receptor, ionotropic, N-methyl D-aspartate-associated protein 1a (glutamate binding) |
| chr15_-_43284021 | 0.14 |
ENSDART00000041677
|
serpine2
|
serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 2 |
| chr21_-_44772738 | 0.13 |
ENSDART00000026178
|
kif4
|
kinesin family member 4 |
| chr18_-_14879135 | 0.13 |
ENSDART00000099701
|
selenoo1
|
selenoprotein O1 |
| chr13_+_36923052 | 0.13 |
ENSDART00000026313
|
tmx1
|
thioredoxin-related transmembrane protein 1 |
| chr6_-_17849786 | 0.13 |
ENSDART00000172709
|
rptor
|
regulatory associated protein of MTOR, complex 1 |
| chr2_+_26647472 | 0.13 |
ENSDART00000145415
ENSDART00000157409 |
ttpa
|
tocopherol (alpha) transfer protein |
| chr2_-_32826108 | 0.13 |
ENSDART00000098834
|
prpf4ba
|
pre-mRNA processing factor 4Ba |
| chr1_+_51479128 | 0.13 |
ENSDART00000028018
|
meis1a
|
Meis homeobox 1 a |
| chr2_-_21170517 | 0.13 |
ENSDART00000135417
|
bmi1b
|
bmi1 polycomb ring finger oncogene 1b |
| chr25_+_36045072 | 0.13 |
ENSDART00000126326
|
rpgrip1l
|
RPGRIP1-like |
| chr21_+_18405585 | 0.12 |
ENSDART00000139318
|
si:dkey-1d7.3
|
si:dkey-1d7.3 |
| chr2_-_54387550 | 0.12 |
ENSDART00000097388
|
napgb
|
N-ethylmaleimide-sensitive factor attachment protein, gamma b |
| chr1_+_26105141 | 0.12 |
ENSDART00000102379
ENSDART00000127154 |
toporsa
|
topoisomerase I binding, arginine/serine-rich a |
| chr20_-_37933237 | 0.12 |
ENSDART00000142567
ENSDART00000036371 ENSDART00000061445 |
angel2
|
angel homolog 2 (Drosophila) |
| chr23_+_2703044 | 0.12 |
ENSDART00000182512
ENSDART00000105286 |
ncoa6
|
nuclear receptor coactivator 6 |
| chr13_-_36535128 | 0.12 |
ENSDART00000043312
|
srsf5a
|
serine/arginine-rich splicing factor 5a |
| chr12_+_20552190 | 0.12 |
ENSDART00000113224
|
gna13a
|
guanine nucleotide binding protein (G protein), alpha 13a |
| chr3_-_8765165 | 0.12 |
ENSDART00000191131
|
CABZ01058333.1
|
|
| chr11_+_24298432 | 0.12 |
ENSDART00000138487
|
si:dkey-76p14.2
|
si:dkey-76p14.2 |
| chr6_-_40744720 | 0.12 |
ENSDART00000154916
ENSDART00000186922 |
p4htm
|
prolyl 4-hydroxylase, transmembrane (endoplasmic reticulum) |
| chr22_-_22301672 | 0.12 |
ENSDART00000111711
|
chaf1a
|
chromatin assembly factor 1, subunit A (p150) |
| chr5_+_26138313 | 0.12 |
ENSDART00000010041
|
dhfr
|
dihydrofolate reductase |
| chr17_-_48805159 | 0.11 |
ENSDART00000103662
|
kif6
|
kinesin family member 6 |
| chr19_-_31402429 | 0.11 |
ENSDART00000137292
|
tmem106bb
|
transmembrane protein 106Bb |
| chr7_+_51795667 | 0.11 |
ENSDART00000174201
ENSDART00000073839 |
slc38a7
|
solute carrier family 38, member 7 |
| chr16_+_40131473 | 0.11 |
ENSDART00000155421
ENSDART00000134732 ENSDART00000138699 |
cenpw
si:ch211-195p4.4
|
centromere protein W si:ch211-195p4.4 |
| chr19_-_11977783 | 0.11 |
ENSDART00000183337
|
si:ch1073-296d18.1
|
si:ch1073-296d18.1 |
| chr9_+_21819082 | 0.11 |
ENSDART00000136902
ENSDART00000101991 |
txndc9
|
thioredoxin domain containing 9 |
| chr4_+_9011448 | 0.11 |
ENSDART00000192357
|
samm50l
|
sorting and assembly machinery component 50 homolog, like |
| chr22_-_22301929 | 0.11 |
ENSDART00000142027
|
chaf1a
|
chromatin assembly factor 1, subunit A (p150) |
| chr7_+_13824150 | 0.11 |
ENSDART00000035067
|
abhd2a
|
abhydrolase domain containing 2a |
| chr5_-_67365006 | 0.11 |
ENSDART00000136116
|
unga
|
uracil DNA glycosylase a |
| chr2_-_17114852 | 0.11 |
ENSDART00000006549
|
pif1
|
PIF1 5'-to-3' DNA helicase homolog (S. cerevisiae) |
| chr4_+_45148652 | 0.11 |
ENSDART00000150798
|
si:dkey-51d8.9
|
si:dkey-51d8.9 |
| chr9_+_12948511 | 0.11 |
ENSDART00000135797
|
si:dkey-230p4.1
|
si:dkey-230p4.1 |
| chr17_-_37195163 | 0.10 |
ENSDART00000108514
|
asxl2
|
additional sex combs like transcriptional regulator 2 |
| chr15_+_5116179 | 0.10 |
ENSDART00000101937
|
pgm2l1
|
phosphoglucomutase 2-like 1 |
| chr19_-_7110617 | 0.10 |
ENSDART00000104838
|
psmb8a
|
proteasome subunit beta 8A |
| chr22_-_621888 | 0.10 |
ENSDART00000135829
|
srsf3b
|
serine/arginine-rich splicing factor 3b |
| chr25_-_32464149 | 0.10 |
ENSDART00000152787
|
atp8b4
|
ATPase phospholipid transporting 8B4 |
| chr10_+_43088270 | 0.10 |
ENSDART00000191221
|
xrcc4
|
X-ray repair complementing defective repair in Chinese hamster cells 4 |
| chr2_-_17115256 | 0.10 |
ENSDART00000190488
|
pif1
|
PIF1 5'-to-3' DNA helicase homolog (S. cerevisiae) |
| chr20_+_25486206 | 0.10 |
ENSDART00000172076
|
hook1
|
hook microtubule-tethering protein 1 |
| chr3_-_46410387 | 0.10 |
ENSDART00000156822
|
cdip1
|
cell death-inducing p53 target 1 |
| chr7_+_6652967 | 0.10 |
ENSDART00000102681
|
pnp5a
|
purine nucleoside phosphorylase 5a |
| chr20_+_25486021 | 0.10 |
ENSDART00000063052
|
hook1
|
hook microtubule-tethering protein 1 |
| chr24_+_5208171 | 0.10 |
ENSDART00000155926
ENSDART00000154464 |
si:ch73-206p6.1
|
si:ch73-206p6.1 |
| chr20_-_39367895 | 0.10 |
ENSDART00000136476
ENSDART00000021788 ENSDART00000180784 |
pbk
|
PDZ binding kinase |
| chr6_-_37469775 | 0.09 |
ENSDART00000156546
|
plcxd1
|
phosphatidylinositol-specific phospholipase C, X domain containing 1 |
| chr20_+_42761881 | 0.09 |
ENSDART00000113625
|
pimr113
|
Pim proto-oncogene, serine/threonine kinase, related 113 |
| chr24_+_39638555 | 0.09 |
ENSDART00000078313
|
luc7l
|
LUC7-like (S. cerevisiae) |
| chr14_-_24095414 | 0.09 |
ENSDART00000172747
ENSDART00000173146 |
si:ch211-277c7.7
|
si:ch211-277c7.7 |
| chr10_+_22034477 | 0.09 |
ENSDART00000133304
ENSDART00000134189 ENSDART00000021240 ENSDART00000100526 |
npm1a
|
nucleophosmin 1a |
| chr7_-_38570878 | 0.09 |
ENSDART00000139187
ENSDART00000134570 ENSDART00000041055 |
celf1
|
cugbp, Elav-like family member 1 |
| chr20_-_27381691 | 0.09 |
ENSDART00000010293
|
serpina10a
|
serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 10a |
| chr19_+_40026041 | 0.09 |
ENSDART00000017359
|
sfpq
|
splicing factor proline/glutamine-rich |
| chr5_-_23596339 | 0.08 |
ENSDART00000024815
|
fam76b
|
family with sequence similarity 76, member B |
| chr4_-_11810799 | 0.08 |
ENSDART00000014153
|
cyb5r3
|
cytochrome b5 reductase 3 |
| chr9_-_7655243 | 0.08 |
ENSDART00000102706
|
dnajb2
|
DnaJ (Hsp40) homolog, subfamily B, member 2 |
| chr12_-_10409961 | 0.08 |
ENSDART00000149521
ENSDART00000052001 |
eef2k
|
eukaryotic elongation factor 2 kinase |
| chr2_+_10821579 | 0.08 |
ENSDART00000179694
|
glmna
|
glomulin, FKBP associated protein a |
| chr4_-_12914163 | 0.08 |
ENSDART00000140002
ENSDART00000145917 ENSDART00000141355 ENSDART00000067135 |
msrb3
|
methionine sulfoxide reductase B3 |
| chr8_-_30944465 | 0.08 |
ENSDART00000128792
ENSDART00000191717 ENSDART00000049944 |
smarcb1a
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily b, member 1a |
| chr18_+_5917625 | 0.08 |
ENSDART00000169100
|
glg1b
|
golgi glycoprotein 1b |
| chr25_-_6389713 | 0.08 |
ENSDART00000083539
|
sin3aa
|
SIN3 transcription regulator family member Aa |
| chr2_+_10821127 | 0.08 |
ENSDART00000145770
ENSDART00000174629 ENSDART00000081094 |
glmna
|
glomulin, FKBP associated protein a |
| chr17_+_1514711 | 0.08 |
ENSDART00000165641
|
akt1
|
v-akt murine thymoma viral oncogene homolog 1 |
| chr1_+_25801648 | 0.08 |
ENSDART00000129471
|
gucy1b1
|
guanylate cyclase 1 soluble subunit beta 1 |
| chr16_-_20294236 | 0.07 |
ENSDART00000059623
|
plekha8
|
pleckstrin homology domain containing, family A (phosphoinositide binding specific) member 8 |
| chr11_-_45138857 | 0.07 |
ENSDART00000166501
|
cant1b
|
calcium activated nucleotidase 1b |
| chr22_-_621609 | 0.07 |
ENSDART00000137264
ENSDART00000106636 |
srsf3b
|
serine/arginine-rich splicing factor 3b |
| chr9_+_35860975 | 0.07 |
ENSDART00000134447
|
rcan1a
|
regulator of calcineurin 1a |
| chr19_-_20163197 | 0.07 |
ENSDART00000148179
ENSDART00000151646 |
fam221a
|
family with sequence similarity 221, member A |
| chr8_-_19467011 | 0.07 |
ENSDART00000162010
|
zgc:92140
|
zgc:92140 |
| chr7_+_69528850 | 0.07 |
ENSDART00000109507
|
RAP1GDS1
|
Rap1 GTPase-GDP dissociation stimulator 1 |
| chr12_-_7824291 | 0.07 |
ENSDART00000148673
ENSDART00000149453 |
ank3b
|
ankyrin 3b |
| chr23_+_20640484 | 0.07 |
ENSDART00000054691
|
uba1
|
ubiquitin-like modifier activating enzyme 1 |
| chr19_-_432083 | 0.07 |
ENSDART00000165371
|
dus3l
|
dihydrouridine synthase 3-like (S. cerevisiae) |
| chr12_-_48960308 | 0.07 |
ENSDART00000176247
|
CABZ01112647.1
|
|
| chr25_-_8513255 | 0.07 |
ENSDART00000150129
|
polg
|
polymerase (DNA directed), gamma |
| chr11_-_36475124 | 0.07 |
ENSDART00000165203
|
usp48
|
ubiquitin specific peptidase 48 |
| chr13_-_293250 | 0.07 |
ENSDART00000138581
|
chs1
|
chitin synthase 1 |
| chr12_-_9132682 | 0.07 |
ENSDART00000066471
|
adam8b
|
ADAM metallopeptidase domain 8b |
| chr19_-_20162980 | 0.06 |
ENSDART00000184605
|
fam221a
|
family with sequence similarity 221, member A |
| chr2_-_31614277 | 0.06 |
ENSDART00000137054
ENSDART00000131507 ENSDART00000137674 ENSDART00000147041 |
si:ch211-106h4.5
|
si:ch211-106h4.5 |
| chr7_+_66634167 | 0.06 |
ENSDART00000027616
|
eif4g2a
|
eukaryotic translation initiation factor 4, gamma 2a |
| chr6_-_51541488 | 0.06 |
ENSDART00000156336
|
si:dkey-6e2.2
|
si:dkey-6e2.2 |
| chr4_-_11810365 | 0.06 |
ENSDART00000150529
|
cyb5r3
|
cytochrome b5 reductase 3 |
| chr13_-_12486076 | 0.06 |
ENSDART00000146195
|
lamtor3
|
late endosomal/lysosomal adaptor, MAPK and MTOR activator 3 |
| chr11_+_29770966 | 0.05 |
ENSDART00000088624
ENSDART00000124471 |
rpgrb
|
retinitis pigmentosa GTPase regulator b |
| chr25_-_2723682 | 0.05 |
ENSDART00000113382
|
adpgk
|
ADP-dependent glucokinase |
| chr14_-_8795081 | 0.05 |
ENSDART00000106671
|
tgfb2l
|
transforming growth factor, beta 2, like |
| chr17_-_49407091 | 0.05 |
ENSDART00000021950
|
mthfd1b
|
methylenetetrahydrofolate dehydrogenase (NADP+ dependent) 1b |
| chr6_-_52675630 | 0.05 |
ENSDART00000083830
|
sdc4
|
syndecan 4 |
| chr10_+_28428222 | 0.05 |
ENSDART00000135003
|
si:ch211-222e20.4
|
si:ch211-222e20.4 |
| chr13_+_21676235 | 0.05 |
ENSDART00000137804
ENSDART00000134950 ENSDART00000129653 |
mtg1
|
mitochondrial ribosome-associated GTPase 1 |
| chr7_-_39751540 | 0.05 |
ENSDART00000016803
|
grpel1
|
GrpE-like 1, mitochondrial |
| chr17_+_12058509 | 0.04 |
ENSDART00000150209
|
tfb2m
|
transcription factor B2, mitochondrial |
| chr23_-_33738945 | 0.04 |
ENSDART00000136386
|
si:ch211-210c8.7
|
si:ch211-210c8.7 |
| chr7_-_7493758 | 0.04 |
ENSDART00000036703
|
pfdn2
|
prefoldin subunit 2 |
| chr23_-_33738570 | 0.04 |
ENSDART00000131680
|
si:ch211-210c8.7
|
si:ch211-210c8.7 |
| chr25_+_32473277 | 0.04 |
ENSDART00000146451
|
sqor
|
sulfide quinone oxidoreductase |
| chr22_+_20145036 | 0.04 |
ENSDART00000137624
|
eef2a.2
|
eukaryotic translation elongation factor 2a, tandem duplicate 2 |
| chr19_-_10395683 | 0.04 |
ENSDART00000109488
|
zgc:194578
|
zgc:194578 |
| chr9_-_42484444 | 0.04 |
ENSDART00000048320
ENSDART00000047653 |
tfpia
|
tissue factor pathway inhibitor a |
| chr24_-_23998897 | 0.03 |
ENSDART00000130053
|
zmp:0000000991
|
zmp:0000000991 |
| chr5_-_12063381 | 0.03 |
ENSDART00000026749
|
nipsnap1
|
nipsnap homolog 1 (C. elegans) |
| chr9_-_10769097 | 0.03 |
ENSDART00000189967
|
tmbim1b
|
transmembrane BAX inhibitor motif containing 1b |
| chr21_-_37733571 | 0.03 |
ENSDART00000176214
|
mpp1
|
membrane protein, palmitoylated 1 |
| chr16_+_28596555 | 0.03 |
ENSDART00000046209
ENSDART00000141708 |
acbd7
|
acyl-CoA binding domain containing 7 |
| chr8_-_4618653 | 0.03 |
ENSDART00000025535
|
sept5a
|
septin 5a |
| chr4_+_77053467 | 0.03 |
ENSDART00000187643
ENSDART00000150848 |
znf1009
|
zinc finger protein 1009 |
| chr3_+_32933663 | 0.03 |
ENSDART00000112742
|
nbr1b
|
neighbor of brca1 gene 1b |
| chr3_-_30186296 | 0.03 |
ENSDART00000134395
ENSDART00000077057 ENSDART00000017422 |
tbc1d17
|
TBC1 domain family, member 17 |
| chr3_+_46764278 | 0.03 |
ENSDART00000136051
ENSDART00000164930 |
prkcsh
|
protein kinase C substrate 80K-H |
| chr17_+_37932433 | 0.03 |
ENSDART00000185349
|
plekhh1
|
pleckstrin homology domain containing, family H (with MyTH4 domain) member 1 |
| chr22_-_817479 | 0.02 |
ENSDART00000123487
|
zgc:153675
|
zgc:153675 |
| chr20_-_45661049 | 0.02 |
ENSDART00000124582
ENSDART00000131251 |
napbb
|
N-ethylmaleimide-sensitive factor attachment protein, beta b |
| chr9_+_19529951 | 0.02 |
ENSDART00000125416
|
pknox1.1
|
pbx/knotted 1 homeobox 1.1 |
| chr10_-_16868211 | 0.02 |
ENSDART00000171755
|
stoml2
|
stomatin (EPB72)-like 2 |
| chr7_-_37895771 | 0.02 |
ENSDART00000084282
|
papd5
|
PAP associated domain containing 5 |
| chr3_-_11089062 | 0.02 |
ENSDART00000168938
|
mif4gdb
|
MIF4G domain containing b |
| chr8_+_1239 | 0.02 |
ENSDART00000067666
ENSDART00000192564 ENSDART00000192612 ENSDART00000190683 |
tmed7
|
transmembrane p24 trafficking protein 7 |
| chr10_-_21941443 | 0.02 |
ENSDART00000174954
|
FO744833.1
|
|
| chr14_+_6605675 | 0.02 |
ENSDART00000143179
|
adam19b
|
ADAM metallopeptidase domain 19b |
| chr3_+_46764022 | 0.02 |
ENSDART00000023814
|
prkcsh
|
protein kinase C substrate 80K-H |
| chr23_-_38330746 | 0.02 |
ENSDART00000192262
|
FO904868.1
|
|
| chr6_+_9163007 | 0.02 |
ENSDART00000183054
ENSDART00000157552 |
zgc:112023
|
zgc:112023 |
| chr22_-_11493236 | 0.02 |
ENSDART00000002691
|
tspan7b
|
tetraspanin 7b |
| chr11_-_18799827 | 0.02 |
ENSDART00000185438
ENSDART00000189116 ENSDART00000180504 |
pfkfb2b
|
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 2b |
| chr11_-_22372072 | 0.02 |
ENSDART00000065996
|
tmem183a
|
transmembrane protein 183A |
| chr21_-_43040780 | 0.02 |
ENSDART00000187037
|
jakmip2
|
janus kinase and microtubule interacting protein 2 |
| chr6_-_24053404 | 0.01 |
ENSDART00000168511
|
si:dkey-44g17.6
|
si:dkey-44g17.6 |
| chr3_+_22335030 | 0.01 |
ENSDART00000055676
|
zgc:103564
|
zgc:103564 |
| chr1_-_6494384 | 0.01 |
ENSDART00000109356
|
klf7a
|
Kruppel-like factor 7a |
| chr1_-_26062798 | 0.01 |
ENSDART00000185348
|
pdcd4a
|
programmed cell death 4a |
| chr2_-_22927958 | 0.01 |
ENSDART00000141621
|
myo7bb
|
myosin VIIBb |
| chr12_+_28956374 | 0.01 |
ENSDART00000033878
|
znf668
|
zinc finger protein 668 |
| chr18_+_30847237 | 0.01 |
ENSDART00000012374
|
foxf1
|
forkhead box F1 |
| chr18_+_11970987 | 0.01 |
ENSDART00000144111
|
si:dkeyp-2c8.3
|
si:dkeyp-2c8.3 |
| chr20_-_44576949 | 0.01 |
ENSDART00000148639
|
ubxn2a
|
UBX domain protein 2A |
Network of associatons between targets according to the STRING database.
First level regulatory network of gmeb2
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | GO:0097510 | base-excision repair, AP site formation via deaminated base removal(GO:0097510) |
| 0.1 | 0.4 | GO:0050861 | positive regulation of B cell receptor signaling pathway(GO:0050861) |
| 0.1 | 0.3 | GO:0045002 | double-strand break repair via single-strand annealing(GO:0045002) |
| 0.1 | 0.3 | GO:0060623 | regulation of chromosome condensation(GO:0060623) |
| 0.1 | 0.3 | GO:0007008 | outer mitochondrial membrane organization(GO:0007008) protein import into mitochondrial outer membrane(GO:0045040) |
| 0.1 | 0.4 | GO:0090245 | axis elongation involved in somitogenesis(GO:0090245) |
| 0.1 | 0.5 | GO:0016137 | glycoside metabolic process(GO:0016137) glycoside catabolic process(GO:0016139) |
| 0.1 | 0.2 | GO:0000390 | spliceosomal complex disassembly(GO:0000390) |
| 0.1 | 0.2 | GO:0060631 | regulation of meiosis I(GO:0060631) |
| 0.0 | 0.1 | GO:1901407 | regulation of phosphorylation of RNA polymerase II C-terminal domain(GO:1901407) |
| 0.0 | 0.2 | GO:0097250 | mitochondrial respiratory chain supercomplex assembly(GO:0097250) |
| 0.0 | 0.2 | GO:1904036 | negative regulation of epithelial cell apoptotic process(GO:1904036) |
| 0.0 | 0.2 | GO:0060394 | negative regulation of pathway-restricted SMAD protein phosphorylation(GO:0060394) |
| 0.0 | 0.2 | GO:0080009 | mRNA methylation(GO:0080009) |
| 0.0 | 0.2 | GO:0042772 | DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator(GO:0006978) DNA damage response, signal transduction resulting in transcription(GO:0042772) |
| 0.0 | 0.3 | GO:0070262 | peptidyl-serine dephosphorylation(GO:0070262) |
| 0.0 | 0.1 | GO:2000048 | activation of phospholipase D activity(GO:0031584) negative regulation of cell-cell adhesion mediated by cadherin(GO:2000048) |
| 0.0 | 0.2 | GO:0048660 | regulation of smooth muscle cell proliferation(GO:0048660) negative regulation of smooth muscle cell proliferation(GO:0048662) |
| 0.0 | 0.1 | GO:0046654 | tetrahydrofolate biosynthetic process(GO:0046654) |
| 0.0 | 0.1 | GO:0051103 | DNA ligation(GO:0006266) DNA ligation involved in DNA repair(GO:0051103) |
| 0.0 | 0.3 | GO:0000002 | mitochondrial genome maintenance(GO:0000002) |
| 0.0 | 0.1 | GO:0051792 | medium-chain fatty acid biosynthetic process(GO:0051792) |
| 0.0 | 0.1 | GO:0061113 | endodermal digestive tract morphogenesis(GO:0061031) pancreas morphogenesis(GO:0061113) |
| 0.0 | 0.1 | GO:0051044 | positive regulation of membrane protein ectodomain proteolysis(GO:0051044) |
| 0.0 | 0.2 | GO:1902624 | positive regulation of neutrophil migration(GO:1902624) |
| 0.0 | 0.1 | GO:0009249 | protein lipoylation(GO:0009249) |
| 0.0 | 0.1 | GO:0030091 | protein repair(GO:0030091) |
| 0.0 | 0.2 | GO:0050435 | beta-amyloid metabolic process(GO:0050435) |
| 0.0 | 0.1 | GO:0099548 | trans-synaptic signaling by soluble gas(GO:0099543) trans-synaptic signaling by nitric oxide(GO:0099548) |
| 0.0 | 0.2 | GO:0046548 | retinal rod cell development(GO:0046548) |
| 0.0 | 0.1 | GO:0006348 | chromatin silencing at telomere(GO:0006348) |
| 0.0 | 0.0 | GO:0019418 | sulfide oxidation(GO:0019418) sulfide oxidation, using sulfide:quinone oxidoreductase(GO:0070221) |
| 0.0 | 0.2 | GO:0042407 | cristae formation(GO:0042407) |
| 0.0 | 0.1 | GO:0070935 | 3'-UTR-mediated mRNA stabilization(GO:0070935) |
| 0.0 | 0.1 | GO:0006031 | chitin biosynthetic process(GO:0006031) glucosamine-containing compound biosynthetic process(GO:1901073) |
| 0.0 | 0.1 | GO:0048103 | neuronal stem cell division(GO:0036445) somatic stem cell division(GO:0048103) |
| 0.0 | 0.2 | GO:0018057 | peptidyl-lysine oxidation(GO:0018057) |
| 0.0 | 0.2 | GO:0098927 | early endosome to late endosome transport(GO:0045022) vesicle-mediated transport between endosomal compartments(GO:0098927) |
| 0.0 | 0.1 | GO:1902108 | regulation of mitochondrial membrane permeability involved in apoptotic process(GO:1902108) |
| 0.0 | 0.2 | GO:0009303 | rRNA transcription(GO:0009303) |
| 0.0 | 0.0 | GO:0006391 | transcription initiation from mitochondrial promoter(GO:0006391) |
| 0.0 | 0.1 | GO:0035627 | ceramide transport(GO:0035627) |
| 0.0 | 0.1 | GO:0048681 | negative regulation of axon regeneration(GO:0048681) negative regulation of neuron projection regeneration(GO:0070571) |
| 0.0 | 0.1 | GO:0051481 | negative regulation of cytosolic calcium ion concentration(GO:0051481) negative regulation of synaptic transmission, glutamatergic(GO:0051967) |
| 0.0 | 0.3 | GO:0006298 | mismatch repair(GO:0006298) |
| 0.0 | 0.1 | GO:0000056 | ribosomal small subunit export from nucleus(GO:0000056) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.2 | GO:0033186 | CAF-1 complex(GO:0033186) |
| 0.1 | 0.2 | GO:0000941 | condensed nuclear chromosome inner kinetochore(GO:0000941) |
| 0.0 | 0.2 | GO:0045293 | MIS complex(GO:0036396) mRNA editing complex(GO:0045293) |
| 0.0 | 0.2 | GO:0031907 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.0 | 0.1 | GO:0005958 | DNA-dependent protein kinase-DNA ligase 4 complex(GO:0005958) |
| 0.0 | 0.3 | GO:0001401 | mitochondrial sorting and assembly machinery complex(GO:0001401) |
| 0.0 | 0.2 | GO:0070695 | FHF complex(GO:0070695) |
| 0.0 | 0.1 | GO:0097524 | sperm plasma membrane(GO:0097524) |
| 0.0 | 0.3 | GO:0032300 | mismatch repair complex(GO:0032300) |
| 0.0 | 0.1 | GO:0071065 | alpha9-beta1 integrin-vascular cell adhesion molecule-1 complex(GO:0071065) |
| 0.0 | 0.1 | GO:0031261 | DNA replication preinitiation complex(GO:0031261) |
| 0.0 | 0.2 | GO:0061617 | MICOS complex(GO:0061617) |
| 0.0 | 0.1 | GO:0031931 | TORC1 complex(GO:0031931) |
| 0.0 | 0.1 | GO:0035517 | PR-DUB complex(GO:0035517) |
| 0.0 | 0.5 | GO:0005940 | septin ring(GO:0005940) septin complex(GO:0031105) septin cytoskeleton(GO:0032156) |
| 0.0 | 0.3 | GO:0030992 | intraciliary transport particle B(GO:0030992) |
| 0.0 | 0.1 | GO:0031526 | brush border membrane(GO:0031526) |
| 0.0 | 0.2 | GO:0031306 | intrinsic component of mitochondrial outer membrane(GO:0031306) |
| 0.0 | 0.0 | GO:0017177 | glucosidase II complex(GO:0017177) |
| 0.0 | 0.0 | GO:0001405 | presequence translocase-associated import motor(GO:0001405) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.6 | GO:0043739 | G/U mismatch-specific uracil-DNA glycosylase activity(GO:0043739) |
| 0.1 | 0.3 | GO:0015928 | alpha-L-fucosidase activity(GO:0004560) fucosidase activity(GO:0015928) |
| 0.1 | 0.2 | GO:0055105 | ubiquitin-protein transferase inhibitor activity(GO:0055105) |
| 0.0 | 0.2 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.0 | 0.2 | GO:0043139 | 5'-3' DNA helicase activity(GO:0043139) |
| 0.0 | 0.1 | GO:0031752 | D5 dopamine receptor binding(GO:0031752) |
| 0.0 | 0.1 | GO:0000099 | sulfur amino acid transmembrane transporter activity(GO:0000099) |
| 0.0 | 0.1 | GO:0005483 | soluble NSF attachment protein activity(GO:0005483) |
| 0.0 | 0.1 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.0 | 0.1 | GO:0004128 | cytochrome-b5 reductase activity, acting on NAD(P)H(GO:0004128) |
| 0.0 | 0.4 | GO:0001968 | fibronectin binding(GO:0001968) |
| 0.0 | 0.1 | GO:0008126 | acetylesterase activity(GO:0008126) |
| 0.0 | 0.2 | GO:0004720 | protein-lysine 6-oxidase activity(GO:0004720) |
| 0.0 | 0.0 | GO:0070224 | sulfide:quinone oxidoreductase activity(GO:0070224) |
| 0.0 | 0.1 | GO:1902387 | ceramide transporter activity(GO:0035620) ceramide 1-phosphate binding(GO:1902387) ceramide 1-phosphate transporter activity(GO:1902388) |
| 0.0 | 0.0 | GO:0034246 | core DNA-dependent RNA polymerase binding promoter specificity activity(GO:0000996) mitochondrial RNA polymerase binding promoter specificity activity(GO:0034246) |
| 0.0 | 0.2 | GO:0031682 | G-protein gamma-subunit binding(GO:0031682) |
| 0.0 | 0.3 | GO:0030983 | mismatched DNA binding(GO:0030983) |
| 0.0 | 0.4 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.0 | 0.2 | GO:0004711 | ribosomal protein S6 kinase activity(GO:0004711) |
| 0.0 | 0.1 | GO:0033743 | peptide-methionine (R)-S-oxide reductase activity(GO:0033743) |
| 0.0 | 0.1 | GO:0008597 | calcium-dependent protein serine/threonine phosphatase regulator activity(GO:0008597) |
| 0.0 | 0.1 | GO:0017150 | tRNA dihydrouridine synthase activity(GO:0017150) |
| 0.0 | 0.1 | GO:0070577 | lysine-acetylated histone binding(GO:0070577) |
| 0.0 | 0.1 | GO:0004731 | purine-nucleoside phosphorylase activity(GO:0004731) |
| 0.0 | 0.1 | GO:0001591 | dopamine neurotransmitter receptor activity, coupled via Gi/Go(GO:0001591) |
| 0.0 | 0.2 | GO:0030544 | Hsp70 protein binding(GO:0030544) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | PID SYNDECAN 4 PATHWAY | Syndecan-4-mediated signaling events |
| 0.0 | 0.2 | PID RANBP2 PATHWAY | Sumoylation by RanBP2 regulates transcriptional repression |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |
| 0.0 | 0.4 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.0 | 0.4 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.0 | 0.2 | REACTOME MTORC1 MEDIATED SIGNALLING | Genes involved in mTORC1-mediated signalling |
| 0.0 | 0.2 | REACTOME HYALURONAN UPTAKE AND DEGRADATION | Genes involved in Hyaluronan uptake and degradation |
| 0.0 | 0.1 | REACTOME INTEGRATION OF PROVIRUS | Genes involved in Integration of provirus |
| 0.0 | 0.3 | REACTOME HOMOLOGOUS RECOMBINATION REPAIR OF REPLICATION INDEPENDENT DOUBLE STRAND BREAKS | Genes involved in Homologous recombination repair of replication-independent double-strand breaks |
| 0.0 | 0.4 | REACTOME SIGNAL TRANSDUCTION BY L1 | Genes involved in Signal transduction by L1 |
| 0.0 | 0.2 | REACTOME DOWNREGULATION OF TGF BETA RECEPTOR SIGNALING | Genes involved in Downregulation of TGF-beta receptor signaling |
| 0.0 | 0.2 | REACTOME AKT PHOSPHORYLATES TARGETS IN THE CYTOSOL | Genes involved in AKT phosphorylates targets in the cytosol |
| 0.0 | 0.1 | REACTOME AMINE LIGAND BINDING RECEPTORS | Genes involved in Amine ligand-binding receptors |
| 0.0 | 0.1 | REACTOME UNWINDING OF DNA | Genes involved in Unwinding of DNA |