Project
PRJNA438478: RNAseq of wild type zebrafish germline
Navigation
Downloads
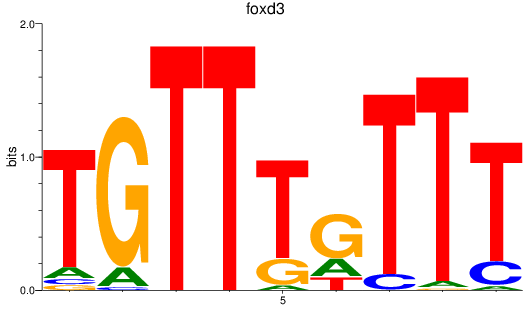
Results for foxd3
Z-value: 2.45
Transcription factors associated with foxd3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
foxd3
|
ENSDARG00000021032 | forkhead box D3 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| foxd3 | dr11_v1_chr6_-_32093830_32093830 | -0.68 | 2.0e-03 | Click! |
Activity profile of foxd3 motif
Sorted Z-values of foxd3 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr12_+_28888975 | 10.09 |
ENSDART00000076362
|
phkg2
|
phosphorylase kinase, gamma 2 (testis) |
| chr22_+_24559947 | 8.20 |
ENSDART00000169847
|
wdr47b
|
WD repeat domain 47b |
| chr6_-_25201810 | 6.52 |
ENSDART00000168683
|
lrrc8c
|
leucine rich repeat containing 8 VRAC subunit C |
| chr17_+_14965570 | 6.51 |
ENSDART00000066604
|
gpr137c
|
G protein-coupled receptor 137c |
| chr15_+_38299385 | 6.36 |
ENSDART00000142403
|
si:dkey-24p1.6
|
si:dkey-24p1.6 |
| chr15_+_38299563 | 5.94 |
ENSDART00000099375
|
si:dkey-24p1.6
|
si:dkey-24p1.6 |
| chr1_-_33647138 | 5.60 |
ENSDART00000142111
ENSDART00000015547 |
cldng
|
claudin g |
| chr14_-_16082806 | 5.27 |
ENSDART00000165656
|
mxd3
|
MAX dimerization protein 3 |
| chr17_-_4245902 | 5.23 |
ENSDART00000151851
|
gdf3
|
growth differentiation factor 3 |
| chr14_+_32926385 | 4.96 |
ENSDART00000139159
|
lnx2b
|
ligand of numb-protein X 2b |
| chr15_-_16076399 | 4.90 |
ENSDART00000135658
ENSDART00000133755 ENSDART00000080413 |
srsf1a
|
serine/arginine-rich splicing factor 1a |
| chr5_+_57924611 | 4.82 |
ENSDART00000050949
|
btg4
|
B-cell translocation gene 4 |
| chr3_+_53116172 | 4.72 |
ENSDART00000115117
|
brd4
|
bromodomain containing 4 |
| chr2_-_44777592 | 4.58 |
ENSDART00000113351
ENSDART00000169310 |
ncapd2
|
non-SMC condensin I complex, subunit D2 |
| chr12_-_11349899 | 4.42 |
ENSDART00000079645
|
zgc:174164
|
zgc:174164 |
| chr6_+_4255319 | 4.39 |
ENSDART00000170351
|
nbeal1
|
neurobeachin-like 1 |
| chr22_-_28777557 | 4.33 |
ENSDART00000135214
ENSDART00000131761 ENSDART00000005112 |
si:dkeyp-34c12.1
|
si:dkeyp-34c12.1 |
| chr15_+_19990068 | 4.33 |
ENSDART00000154033
ENSDART00000054428 |
zgc:112083
|
zgc:112083 |
| chr19_+_25971000 | 4.22 |
ENSDART00000089836
|
jarid2b
|
jumonji, AT rich interactive domain 2b |
| chr22_-_28777374 | 4.08 |
ENSDART00000188206
|
si:dkeyp-34c12.1
|
si:dkeyp-34c12.1 |
| chr20_-_4793450 | 4.02 |
ENSDART00000053870
|
galca
|
galactosylceramidase a |
| chr11_+_45286911 | 4.02 |
ENSDART00000181763
|
pycr1b
|
pyrroline-5-carboxylate reductase 1b |
| chr13_+_11828516 | 3.99 |
ENSDART00000110141
|
sufu
|
suppressor of fused homolog (Drosophila) |
| chr22_-_22337382 | 3.95 |
ENSDART00000144684
|
si:ch211-129c21.1
|
si:ch211-129c21.1 |
| chr15_-_44052927 | 3.93 |
ENSDART00000166209
|
wu:fb44b02
|
wu:fb44b02 |
| chr18_+_15271993 | 3.90 |
ENSDART00000099777
|
si:dkey-103i16.6
|
si:dkey-103i16.6 |
| chr1_-_6028876 | 3.87 |
ENSDART00000168117
|
si:ch1073-345a8.1
|
si:ch1073-345a8.1 |
| chr3_-_34561624 | 3.83 |
ENSDART00000129313
|
sept9a
|
septin 9a |
| chr3_-_43650189 | 3.82 |
ENSDART00000161127
|
axin1
|
axin 1 |
| chr14_-_34633960 | 3.82 |
ENSDART00000128869
ENSDART00000179977 |
afap1l1a
|
actin filament associated protein 1-like 1a |
| chr12_-_17863467 | 3.81 |
ENSDART00000042006
|
baiap2l1a
|
BAI1-associated protein 2-like 1a |
| chr16_+_23487051 | 3.69 |
ENSDART00000145496
|
icn2
|
ictacalcin 2 |
| chr1_-_33645967 | 3.67 |
ENSDART00000192758
|
cldng
|
claudin g |
| chr5_+_31214341 | 3.67 |
ENSDART00000133432
|
atp2a3
|
ATPase sarcoplasmic/endoplasmic reticulum Ca2+ transporting 3 |
| chr20_+_29209767 | 3.66 |
ENSDART00000141252
|
katnbl1
|
katanin p80 subunit B-like 1 |
| chr11_+_18612421 | 3.59 |
ENSDART00000110621
|
ncoa3
|
nuclear receptor coactivator 3 |
| chr17_+_32622933 | 3.57 |
ENSDART00000077418
|
ctsba
|
cathepsin Ba |
| chr5_-_13766651 | 3.56 |
ENSDART00000134064
|
mxd1
|
MAX dimerization protein 1 |
| chr5_+_3891485 | 3.52 |
ENSDART00000129329
ENSDART00000091711 |
rpain
|
RPA interacting protein |
| chr3_-_34528306 | 3.51 |
ENSDART00000023039
|
sept9a
|
septin 9a |
| chr22_-_17671348 | 3.47 |
ENSDART00000137995
|
tjp3
|
tight junction protein 3 |
| chr21_-_13856689 | 3.46 |
ENSDART00000102197
|
fam129ba
|
family with sequence similarity 129, member Ba |
| chr20_+_29209926 | 3.43 |
ENSDART00000152949
ENSDART00000153016 |
katnbl1
|
katanin p80 subunit B-like 1 |
| chr10_-_14929392 | 3.42 |
ENSDART00000137430
|
smad2
|
SMAD family member 2 |
| chr7_+_24023653 | 3.40 |
ENSDART00000141165
|
tinf2
|
TERF1 (TRF1)-interacting nuclear factor 2 |
| chr8_+_10862353 | 3.40 |
ENSDART00000140717
|
brpf3b
|
bromodomain and PHD finger containing, 3b |
| chr23_+_36730713 | 3.39 |
ENSDART00000113179
|
tspan31
|
tetraspanin 31 |
| chr19_+_1510971 | 3.38 |
ENSDART00000157721
|
SLC45A4 (1 of many)
|
solute carrier family 45 member 4 |
| chr3_-_26191960 | 3.38 |
ENSDART00000113843
|
ypel3
|
yippee-like 3 |
| chr6_-_41135215 | 3.37 |
ENSDART00000001861
|
slc6a22.1
|
solute carrier family 6 member 22, tandem duplicate 1 |
| chr13_-_22961605 | 3.36 |
ENSDART00000143112
ENSDART00000057641 |
tspan15
|
tetraspanin 15 |
| chr20_+_29209615 | 3.34 |
ENSDART00000062350
|
katnbl1
|
katanin p80 subunit B-like 1 |
| chr17_-_40956035 | 3.29 |
ENSDART00000124715
|
si:dkey-16j16.4
|
si:dkey-16j16.4 |
| chr7_-_51773166 | 3.23 |
ENSDART00000054591
|
bmp15
|
bone morphogenetic protein 15 |
| chr23_+_22873415 | 3.22 |
ENSDART00000135130
|
rerea
|
arginine-glutamic acid dipeptide (RE) repeats a |
| chr16_+_43077909 | 3.18 |
ENSDART00000014140
|
rundc3b
|
RUN domain containing 3b |
| chr6_+_2174082 | 3.18 |
ENSDART00000073936
|
acvr1bb
|
activin A receptor type 1Bb |
| chr12_+_20691310 | 3.17 |
ENSDART00000064335
|
st6galnac1.2
|
ST6 (alpha-N-acetyl-neuraminyl-2,3-beta-galactosyl-1,3)-N-acetylgalactosaminide alpha-2,6-sialyltransferase 1, tandem duplicate 2 |
| chr17_+_25856671 | 3.17 |
ENSDART00000064817
|
wapla
|
WAPL cohesin release factor a |
| chr9_+_27720428 | 3.16 |
ENSDART00000112415
|
lcmt2
|
leucine carboxyl methyltransferase 2 |
| chr13_+_25397098 | 3.15 |
ENSDART00000132953
|
gsto2
|
glutathione S-transferase omega 2 |
| chr12_-_35505610 | 3.13 |
ENSDART00000105518
|
camk2g1
|
calcium/calmodulin-dependent protein kinase (CaM kinase) II gamma 1 |
| chr13_+_25396896 | 3.12 |
ENSDART00000041257
|
gsto2
|
glutathione S-transferase omega 2 |
| chr12_-_25380028 | 3.11 |
ENSDART00000142674
|
zfp36l2
|
zinc finger protein 36, C3H type-like 2 |
| chr2_+_31437547 | 3.11 |
ENSDART00000141170
|
stam
|
signal transducing adaptor molecule (SH3 domain and ITAM motif) 1 |
| chr6_-_40922971 | 3.11 |
ENSDART00000155363
|
sfi1
|
SFI1 centrin binding protein |
| chr1_+_45707219 | 3.10 |
ENSDART00000143363
|
si:ch211-214c7.4
|
si:ch211-214c7.4 |
| chr2_+_15069011 | 3.09 |
ENSDART00000145893
|
cnn3b
|
calponin 3, acidic b |
| chr12_+_11650146 | 3.08 |
ENSDART00000150191
|
waplb
|
WAPL cohesin release factor b |
| chr7_+_52712807 | 3.07 |
ENSDART00000174095
ENSDART00000174377 ENSDART00000174061 ENSDART00000174094 ENSDART00000110906 ENSDART00000174071 ENSDART00000174238 |
znf280d
|
zinc finger protein 280D |
| chr5_+_13472234 | 3.06 |
ENSDART00000114069
ENSDART00000132406 |
cnnm4b
|
cyclin and CBS domain divalent metal cation transport mediator 4b |
| chr14_-_21660548 | 3.05 |
ENSDART00000161713
ENSDART00000089845 |
kdm3b
|
lysine (K)-specific demethylase 3B |
| chr15_-_37104165 | 3.05 |
ENSDART00000165867
|
zmp:0000001114
|
zmp:0000001114 |
| chr21_-_25295087 | 3.04 |
ENSDART00000087910
ENSDART00000147860 |
st14b
|
suppression of tumorigenicity 14 (colon carcinoma) b |
| chr15_+_23657051 | 3.02 |
ENSDART00000078336
|
klc3
|
kinesin light chain 3 |
| chr20_+_26940178 | 3.02 |
ENSDART00000190888
|
cdca4
|
cell division cycle associated 4 |
| chr13_-_18011168 | 2.98 |
ENSDART00000144813
|
march8
|
membrane-associated ring finger (C3HC4) 8 |
| chr3_-_2623176 | 2.96 |
ENSDART00000179792
ENSDART00000123512 |
si:dkey-217f16.6
|
si:dkey-217f16.6 |
| chr1_-_55015439 | 2.95 |
ENSDART00000161029
|
vps54
|
vacuolar protein sorting 54 homolog (S. cerevisiae) |
| chr7_+_38808027 | 2.90 |
ENSDART00000052323
|
harbi1
|
harbinger transposase derived 1 |
| chr24_-_23716097 | 2.86 |
ENSDART00000084954
ENSDART00000129028 |
pign
|
phosphatidylinositol glycan anchor biosynthesis, class N |
| chr2_-_57110477 | 2.86 |
ENSDART00000181132
|
slc25a42
|
solute carrier family 25, member 42 |
| chr5_-_48070779 | 2.85 |
ENSDART00000078401
|
tmem161b
|
transmembrane protein 161B |
| chr7_-_55633475 | 2.85 |
ENSDART00000149478
|
galns
|
galactosamine (N-acetyl)-6-sulfatase |
| chr10_+_3428194 | 2.82 |
ENSDART00000081599
|
ptpn11a
|
protein tyrosine phosphatase, non-receptor type 11, a |
| chr5_-_3960161 | 2.80 |
ENSDART00000111453
|
myo19
|
myosin XIX |
| chr19_-_6193448 | 2.79 |
ENSDART00000151405
|
erf
|
Ets2 repressor factor |
| chr25_+_16214854 | 2.79 |
ENSDART00000109672
ENSDART00000190093 |
mical2b
|
microtubule associated monooxygenase, calponin and LIM domain containing 2b |
| chr19_-_2822372 | 2.78 |
ENSDART00000109130
ENSDART00000122385 |
recql4
|
RecQ helicase-like 4 |
| chr13_+_5978809 | 2.75 |
ENSDART00000102563
ENSDART00000121598 |
phf10
|
PHD finger protein 10 |
| chr23_+_28378543 | 2.75 |
ENSDART00000145327
|
zgc:153867
|
zgc:153867 |
| chr8_+_30742898 | 2.74 |
ENSDART00000018475
|
snrpd3
|
small nuclear ribonucleoprotein D3 polypeptide |
| chr23_-_36670369 | 2.73 |
ENSDART00000006881
|
zbtb39
|
zinc finger and BTB domain containing 39 |
| chr10_-_15854743 | 2.73 |
ENSDART00000092343
|
tjp2a
|
tight junction protein 2a (zona occludens 2) |
| chr12_+_22404108 | 2.69 |
ENSDART00000153055
|
hdlbpb
|
high density lipoprotein binding protein b |
| chr24_+_36317544 | 2.67 |
ENSDART00000048640
ENSDART00000156096 |
pus3
|
pseudouridylate synthase 3 |
| chr2_-_42492201 | 2.67 |
ENSDART00000180762
ENSDART00000009093 |
esyt2a
|
extended synaptotagmin-like protein 2a |
| chr8_-_25566347 | 2.66 |
ENSDART00000138289
ENSDART00000078022 |
prex1
|
phosphatidylinositol-3,4,5-trisphosphate-dependent Rac exchange factor 1 |
| chr18_+_38775277 | 2.66 |
ENSDART00000186129
|
fam214a
|
family with sequence similarity 214, member A |
| chr7_-_48251234 | 2.66 |
ENSDART00000024062
ENSDART00000098904 |
cpeb1b
|
cytoplasmic polyadenylation element binding protein 1b |
| chr19_+_41479990 | 2.66 |
ENSDART00000087187
|
ago2
|
argonaute RISC catalytic component 2 |
| chr6_-_37745508 | 2.66 |
ENSDART00000078316
|
nipa2
|
non imprinted in Prader-Willi/Angelman syndrome 2 (human) |
| chr2_-_41723487 | 2.65 |
ENSDART00000170171
|
zgc:110158
|
zgc:110158 |
| chr11_+_31864921 | 2.65 |
ENSDART00000180252
|
diaph3
|
diaphanous-related formin 3 |
| chr16_+_35924188 | 2.64 |
ENSDART00000165847
|
sh3d21
|
SH3 domain containing 21 |
| chr1_+_11107688 | 2.64 |
ENSDART00000109858
|
knstrn
|
kinetochore-localized astrin/SPAG5 binding protein |
| chr11_+_506465 | 2.64 |
ENSDART00000082519
|
raf1b
|
Raf-1 proto-oncogene, serine/threonine kinase b |
| chr18_-_40913294 | 2.63 |
ENSDART00000059196
ENSDART00000098878 |
polr2i
|
polymerase (RNA) II (DNA directed) polypeptide I |
| chr2_+_44512324 | 2.62 |
ENSDART00000155017
ENSDART00000156310 ENSDART00000156686 |
pask
|
PAS domain containing serine/threonine kinase |
| chr7_-_49594995 | 2.61 |
ENSDART00000174161
ENSDART00000109147 |
brsk2b
|
BR serine/threonine kinase 2b |
| chr6_+_32393057 | 2.61 |
ENSDART00000190765
|
dock7
|
dedicator of cytokinesis 7 |
| chr7_+_34487833 | 2.57 |
ENSDART00000173854
|
cln6a
|
CLN6, transmembrane ER protein a |
| chr9_+_24088062 | 2.57 |
ENSDART00000126198
|
lrrfip1a
|
leucine rich repeat (in FLII) interacting protein 1a |
| chr2_-_42492445 | 2.57 |
ENSDART00000139929
|
esyt2a
|
extended synaptotagmin-like protein 2a |
| chr11_-_43226255 | 2.56 |
ENSDART00000172929
|
sptbn1
|
spectrin, beta, non-erythrocytic 1 |
| chr2_+_1988036 | 2.56 |
ENSDART00000155956
|
ssx2ipa
|
synovial sarcoma, X breakpoint 2 interacting protein a |
| chr14_-_36437249 | 2.55 |
ENSDART00000016728
|
aga
|
aspartylglucosaminidase |
| chr7_+_21787507 | 2.55 |
ENSDART00000100936
|
tmem88b
|
transmembrane protein 88 b |
| chr9_+_38458193 | 2.55 |
ENSDART00000008053
|
mcm3ap
|
minichromosome maintenance complex component 3 associated protein |
| chr16_-_41488023 | 2.53 |
ENSDART00000169312
|
cmtm6
|
CKLF-like MARVEL transmembrane domain containing 6 |
| chr25_-_14087377 | 2.53 |
ENSDART00000124140
|
zgc:101566
|
zgc:101566 |
| chr13_+_37653851 | 2.53 |
ENSDART00000141988
ENSDART00000126902 ENSDART00000100352 |
phf3
|
PHD finger protein 3 |
| chr13_-_36761379 | 2.52 |
ENSDART00000131534
ENSDART00000029824 |
map4k5
|
mitogen-activated protein kinase kinase kinase kinase 5 |
| chr10_-_33251876 | 2.51 |
ENSDART00000184565
|
bcl7ba
|
BCL tumor suppressor 7Ba |
| chr5_-_49951106 | 2.51 |
ENSDART00000135954
|
fam172a
|
family with sequence similarity 172, member A |
| chr18_+_20047374 | 2.50 |
ENSDART00000146957
|
uacaa
|
uveal autoantigen with coiled-coil domains and ankyrin repeats a |
| chr5_-_22052852 | 2.48 |
ENSDART00000002938
|
mtmr8
|
myotubularin related protein 8 |
| chr5_+_61361815 | 2.48 |
ENSDART00000009507
|
gatsl2
|
GATS protein-like 2 |
| chr11_-_24523161 | 2.46 |
ENSDART00000191936
|
plekhg5a
|
pleckstrin homology domain containing, family G (with RhoGef domain) member 5a |
| chr12_-_28791969 | 2.46 |
ENSDART00000153073
|
osbpl7
|
oxysterol binding protein-like 7 |
| chr25_-_13789955 | 2.45 |
ENSDART00000167742
ENSDART00000165116 ENSDART00000171461 |
ckap5
|
cytoskeleton associated protein 5 |
| chr18_+_17534627 | 2.44 |
ENSDART00000061007
|
mt2
|
metallothionein 2 |
| chr8_-_53896982 | 2.43 |
ENSDART00000169206
|
MIB2
|
mindbomb E3 ubiquitin protein ligase 2 |
| chr16_+_39146696 | 2.43 |
ENSDART00000121756
ENSDART00000084381 |
sybu
|
syntabulin (syntaxin-interacting) |
| chr3_+_28502419 | 2.41 |
ENSDART00000151081
|
sept12
|
septin 12 |
| chr3_-_15119856 | 2.40 |
ENSDART00000138328
|
xpo6
|
exportin 6 |
| chr20_-_28842524 | 2.40 |
ENSDART00000046035
ENSDART00000139843 ENSDART00000129858 ENSDART00000137425 ENSDART00000135720 |
max
|
myc associated factor X |
| chr3_-_55650771 | 2.40 |
ENSDART00000162413
|
axin2
|
axin 2 (conductin, axil) |
| chr22_-_38934989 | 2.39 |
ENSDART00000008365
|
ncbp2
|
nuclear cap binding protein subunit 2 |
| chr2_+_44518636 | 2.39 |
ENSDART00000153733
|
pask
|
PAS domain containing serine/threonine kinase |
| chr1_-_40519340 | 2.38 |
ENSDART00000114659
|
maml3
|
mastermind-like transcriptional coactivator 3 |
| chr22_-_17688868 | 2.38 |
ENSDART00000012336
ENSDART00000147070 |
tjp3
|
tight junction protein 3 |
| chr17_-_23709347 | 2.37 |
ENSDART00000124661
|
papss2a
|
3'-phosphoadenosine 5'-phosphosulfate synthase 2a |
| chr16_+_25259313 | 2.37 |
ENSDART00000058938
|
fbxo32
|
F-box protein 32 |
| chr18_-_21725638 | 2.36 |
ENSDART00000089853
|
fan1
|
FANCD2/FANCI-associated nuclease 1 |
| chr7_+_26549846 | 2.35 |
ENSDART00000141353
|
tnk1
|
tyrosine kinase, non-receptor, 1 |
| chr5_-_23517747 | 2.35 |
ENSDART00000137655
|
stag2a
|
stromal antigen 2a |
| chr5_+_44805269 | 2.34 |
ENSDART00000136965
|
ctsla
|
cathepsin La |
| chr10_+_3153973 | 2.34 |
ENSDART00000183223
|
hic2
|
hypermethylated in cancer 2 |
| chr12_-_4301234 | 2.34 |
ENSDART00000152377
ENSDART00000152521 |
ca15b
|
carbonic anhydrase XVb |
| chr14_+_21699129 | 2.33 |
ENSDART00000073707
|
stx3a
|
syntaxin 3A |
| chr15_-_16946124 | 2.33 |
ENSDART00000154923
|
hip1
|
huntingtin interacting protein 1 |
| chr3_-_4501026 | 2.33 |
ENSDART00000163052
|
zgc:162198
|
zgc:162198 |
| chr19_-_12965020 | 2.32 |
ENSDART00000128975
|
slc25a32a
|
solute carrier family 25 (mitochondrial folate carrier), member 32a |
| chr5_+_44805028 | 2.32 |
ENSDART00000141198
|
ctsla
|
cathepsin La |
| chr1_+_44173245 | 2.32 |
ENSDART00000159450
ENSDART00000106048 ENSDART00000157763 |
ctnnd1
|
catenin (cadherin-associated protein), delta 1 |
| chr16_-_35532937 | 2.31 |
ENSDART00000193209
|
ctps1b
|
CTP synthase 1b |
| chr9_+_54039006 | 2.31 |
ENSDART00000112441
|
tlr7
|
toll-like receptor 7 |
| chr22_-_5171829 | 2.30 |
ENSDART00000140313
|
tnfaip8l1
|
tumor necrosis factor, alpha-induced protein 8-like 1 |
| chr8_-_537716 | 2.29 |
ENSDART00000051777
|
zgc:101664
|
zgc:101664 |
| chr23_+_38171186 | 2.28 |
ENSDART00000148188
|
zgc:112994
|
zgc:112994 |
| chr22_-_10752471 | 2.27 |
ENSDART00000081191
|
sass6
|
SAS-6 centriolar assembly protein |
| chr16_+_47207691 | 2.27 |
ENSDART00000062507
|
ica1
|
islet cell autoantigen 1 |
| chr20_-_23876291 | 2.27 |
ENSDART00000043316
|
katna1
|
katanin p60 (ATPase containing) subunit A 1 |
| chr16_-_31791165 | 2.27 |
ENSDART00000148389
|
chd4b
|
chromodomain helicase DNA binding protein 4b |
| chr22_-_3299100 | 2.26 |
ENSDART00000160305
|
si:zfos-943e10.1
|
si:zfos-943e10.1 |
| chr4_-_1720648 | 2.26 |
ENSDART00000103484
|
gas2l3
|
growth arrest-specific 2 like 3 |
| chr6_+_20954400 | 2.25 |
ENSDART00000143248
ENSDART00000165806 |
stk11ip
|
serine/threonine kinase 11 interacting protein |
| chr7_+_13830052 | 2.24 |
ENSDART00000191360
|
abhd2a
|
abhydrolase domain containing 2a |
| chr3_-_15131438 | 2.23 |
ENSDART00000131720
|
xpo6
|
exportin 6 |
| chr23_-_26784736 | 2.23 |
ENSDART00000024064
ENSDART00000131615 |
galnt6
|
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 6 (GalNAc-T6) |
| chr21_+_4116437 | 2.23 |
ENSDART00000167791
ENSDART00000123759 |
rapgef1b
|
Rap guanine nucleotide exchange factor (GEF) 1b |
| chr7_+_13824150 | 2.23 |
ENSDART00000035067
|
abhd2a
|
abhydrolase domain containing 2a |
| chr25_+_7494181 | 2.22 |
ENSDART00000165005
|
cat
|
catalase |
| chr1_+_12302073 | 2.22 |
ENSDART00000164045
|
gne
|
glucosamine (UDP-N-acetyl)-2-epimerase/N-acetylmannosamine kinase |
| chr16_-_47381519 | 2.22 |
ENSDART00000032188
ENSDART00000150136 |
si:dkey-256h2.1
|
si:dkey-256h2.1 |
| chr17_-_14966384 | 2.22 |
ENSDART00000105064
|
txndc16
|
thioredoxin domain containing 16 |
| chr23_+_4709607 | 2.21 |
ENSDART00000166503
ENSDART00000158752 ENSDART00000163860 ENSDART00000172739 |
raf1a
raf1a
|
Raf-1 proto-oncogene, serine/threonine kinase a Raf-1 proto-oncogene, serine/threonine kinase a |
| chr8_+_8643901 | 2.21 |
ENSDART00000142076
ENSDART00000075624 |
usp11
|
ubiquitin specific peptidase 11 |
| chr2_+_30740095 | 2.21 |
ENSDART00000186387
|
tcea1
|
transcription elongation factor A (SII), 1 |
| chr18_+_26429428 | 2.20 |
ENSDART00000142686
|
blm
|
Bloom syndrome, RecQ helicase-like |
| chr24_+_33462800 | 2.19 |
ENSDART00000166666
ENSDART00000050826 |
rmc1
|
regulator of MON1-CCZ1 |
| chr2_+_34967022 | 2.19 |
ENSDART00000134926
|
astn1
|
astrotactin 1 |
| chr3_+_31662126 | 2.19 |
ENSDART00000113441
|
mylk5
|
myosin, light chain kinase 5 |
| chr13_-_23051766 | 2.19 |
ENSDART00000111774
|
supv3l1
|
SUV3-like helicase |
| chr17_+_50261603 | 2.18 |
ENSDART00000154503
ENSDART00000154467 |
syncripl
|
synaptotagmin binding, cytoplasmic RNA interacting protein, like |
| chr1_+_26676758 | 2.18 |
ENSDART00000152299
|
si:dkey-25o16.4
|
si:dkey-25o16.4 |
| chr4_+_14981854 | 2.18 |
ENSDART00000067046
|
cax1
|
cation/H+ exchanger protein 1 |
| chr10_-_44017642 | 2.18 |
ENSDART00000135240
ENSDART00000014669 |
acads
|
acyl-CoA dehydrogenase short chain |
| chr8_+_42917515 | 2.17 |
ENSDART00000021715
|
slc23a2
|
solute carrier family 23 (ascorbic acid transporter), member 2 |
| chr11_-_38554394 | 2.16 |
ENSDART00000102858
|
nucks1a
|
nuclear casein kinase and cyclin-dependent kinase substrate 1a |
| chr21_-_30082414 | 2.15 |
ENSDART00000157307
ENSDART00000155188 |
ccnjl
|
cyclin J-like |
| chr17_-_29311835 | 2.14 |
ENSDART00000104224
|
tecpr2
|
tectonin beta-propeller repeat containing 2 |
| chr3_+_31925067 | 2.14 |
ENSDART00000127330
ENSDART00000055279 |
snrnp70
|
small nuclear ribonucleoprotein 70 (U1) |
| chr7_-_32629458 | 2.13 |
ENSDART00000001376
|
arl14ep
|
ADP-ribosylation factor-like 14 effector protein |
| chr25_+_28158352 | 2.13 |
ENSDART00000151854
|
cadps2
|
Ca++-dependent secretion activator 2 |
| chr9_-_30220818 | 2.13 |
ENSDART00000140929
|
si:dkey-100n23.3
|
si:dkey-100n23.3 |
| chr17_-_29271359 | 2.12 |
ENSDART00000104219
|
rcor1
|
REST corepressor 1 |
| chr19_-_6983002 | 2.11 |
ENSDART00000104891
|
znf384l
|
zinc finger protein 384 like |
| chr13_-_12667220 | 2.10 |
ENSDART00000079594
|
fam241a
|
family with sequence similarity 241 member A |
Network of associatons between targets according to the STRING database.
First level regulatory network of foxd3
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.1 | 6.2 | GO:0060623 | regulation of chromosome condensation(GO:0060623) |
| 1.9 | 9.3 | GO:0016338 | calcium-independent cell-cell adhesion via plasma membrane cell-adhesion molecules(GO:0016338) |
| 1.6 | 4.7 | GO:1901407 | regulation of phosphorylation of RNA polymerase II C-terminal domain(GO:1901407) |
| 1.4 | 4.2 | GO:0051574 | positive regulation of histone H3-K9 methylation(GO:0051574) |
| 1.2 | 3.7 | GO:0015882 | L-ascorbic acid transport(GO:0015882) L-ascorbic acid metabolic process(GO:0019852) |
| 1.1 | 4.6 | GO:0010032 | meiotic chromosome condensation(GO:0010032) |
| 1.1 | 3.2 | GO:0031591 | wybutosine metabolic process(GO:0031590) wybutosine biosynthetic process(GO:0031591) |
| 1.0 | 5.2 | GO:0038107 | nodal signaling pathway involved in determination of left/right asymmetry(GO:0038107) regulation of nodal signaling pathway involved in determination of left/right asymmetry(GO:1900145) |
| 1.0 | 3.1 | GO:0045040 | outer mitochondrial membrane organization(GO:0007008) protein import into mitochondrial outer membrane(GO:0045040) |
| 1.0 | 3.1 | GO:0071896 | protein localization to adherens junction(GO:0071896) |
| 1.0 | 4.0 | GO:0042308 | regulation of protein import into nucleus(GO:0042306) negative regulation of protein import into nucleus(GO:0042308) negative regulation of nucleocytoplasmic transport(GO:0046823) negative regulation of protein localization to nucleus(GO:1900181) regulation of protein import(GO:1904589) negative regulation of protein import(GO:1904590) |
| 1.0 | 4.8 | GO:0051988 | regulation of attachment of spindle microtubules to kinetochore(GO:0051988) |
| 1.0 | 3.8 | GO:0090244 | Wnt signaling pathway involved in somitogenesis(GO:0090244) |
| 0.9 | 4.5 | GO:0051792 | medium-chain fatty acid biosynthetic process(GO:0051792) |
| 0.8 | 4.2 | GO:0098967 | exocytic insertion of neurotransmitter receptor to plasma membrane(GO:0098881) exocytic insertion of neurotransmitter receptor to postsynaptic membrane(GO:0098967) |
| 0.8 | 2.5 | GO:1901223 | negative regulation of NIK/NF-kappaB signaling(GO:1901223) |
| 0.8 | 3.2 | GO:0060283 | negative regulation of oocyte development(GO:0060283) |
| 0.8 | 2.4 | GO:0046833 | regulation of nucleobase-containing compound transport(GO:0032239) positive regulation of nucleobase-containing compound transport(GO:0032241) positive regulation of nucleocytoplasmic transport(GO:0046824) regulation of RNA export from nucleus(GO:0046831) positive regulation of RNA export from nucleus(GO:0046833) |
| 0.8 | 3.9 | GO:0010693 | regulation of alkaline phosphatase activity(GO:0010692) negative regulation of alkaline phosphatase activity(GO:0010693) |
| 0.8 | 4.7 | GO:0033169 | histone H3-K9 demethylation(GO:0033169) |
| 0.8 | 2.3 | GO:0034154 | toll-like receptor 7 signaling pathway(GO:0034154) |
| 0.8 | 2.3 | GO:0045830 | regulation of isotype switching(GO:0045191) positive regulation of isotype switching(GO:0045830) |
| 0.7 | 3.0 | GO:0090155 | regulation of sphingolipid biosynthetic process(GO:0090153) negative regulation of sphingolipid biosynthetic process(GO:0090155) negative regulation of ceramide biosynthetic process(GO:1900060) regulation of membrane lipid metabolic process(GO:1905038) regulation of ceramide biosynthetic process(GO:2000303) |
| 0.7 | 3.0 | GO:0002495 | antigen processing and presentation of peptide antigen via MHC class II(GO:0002495) |
| 0.7 | 4.4 | GO:0060282 | positive regulation of oocyte development(GO:0060282) |
| 0.7 | 5.5 | GO:0070874 | negative regulation of glycogen biosynthetic process(GO:0045719) negative regulation of glycogen metabolic process(GO:0070874) |
| 0.7 | 2.0 | GO:0034214 | protein hexamerization(GO:0034214) mitotic spindle disassembly(GO:0051228) spindle disassembly(GO:0051230) |
| 0.7 | 6.0 | GO:0006561 | proline biosynthetic process(GO:0006561) L-proline biosynthetic process(GO:0055129) |
| 0.7 | 2.0 | GO:0070537 | histone H2A K63-linked deubiquitination(GO:0070537) |
| 0.7 | 2.0 | GO:1905072 | cardiac jelly development(GO:1905072) |
| 0.6 | 1.9 | GO:0010633 | negative regulation of epithelial cell migration(GO:0010633) |
| 0.6 | 3.7 | GO:0016559 | peroxisome fission(GO:0016559) |
| 0.6 | 2.4 | GO:0030951 | establishment or maintenance of microtubule cytoskeleton polarity(GO:0030951) |
| 0.6 | 2.4 | GO:0060074 | synapse maturation(GO:0060074) |
| 0.6 | 5.8 | GO:0030104 | water homeostasis(GO:0030104) |
| 0.6 | 2.3 | GO:0044210 | 'de novo' CTP biosynthetic process(GO:0044210) |
| 0.6 | 5.1 | GO:0043328 | protein targeting to vacuole involved in ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway(GO:0043328) |
| 0.5 | 1.6 | GO:0051580 | regulation of neurotransmitter uptake(GO:0051580) negative regulation of anion transport(GO:1903792) |
| 0.5 | 2.2 | GO:0019626 | short-chain fatty acid catabolic process(GO:0019626) |
| 0.5 | 1.6 | GO:0006451 | selenocysteine incorporation(GO:0001514) translational readthrough(GO:0006451) |
| 0.5 | 4.3 | GO:0031063 | regulation of histone deacetylation(GO:0031063) |
| 0.5 | 2.7 | GO:1990481 | mRNA pseudouridine synthesis(GO:1990481) |
| 0.5 | 2.7 | GO:0045974 | miRNA mediated inhibition of translation(GO:0035278) negative regulation of translation, ncRNA-mediated(GO:0040033) regulation of translation, ncRNA-mediated(GO:0045974) |
| 0.5 | 2.1 | GO:0044806 | G-quadruplex DNA unwinding(GO:0044806) |
| 0.5 | 1.6 | GO:0000390 | spliceosomal complex disassembly(GO:0000390) |
| 0.5 | 6.3 | GO:1904263 | positive regulation of TORC1 signaling(GO:1904263) |
| 0.5 | 2.0 | GO:0046900 | tetrahydrofolylpolyglutamate metabolic process(GO:0046900) |
| 0.5 | 3.4 | GO:0006542 | glutamine biosynthetic process(GO:0006542) |
| 0.5 | 1.0 | GO:0006882 | cellular zinc ion homeostasis(GO:0006882) zinc ion homeostasis(GO:0055069) |
| 0.5 | 2.4 | GO:0007221 | positive regulation of transcription of Notch receptor target(GO:0007221) |
| 0.5 | 3.8 | GO:0046477 | glycosylceramide catabolic process(GO:0046477) |
| 0.5 | 1.4 | GO:0032570 | response to progesterone(GO:0032570) |
| 0.5 | 3.3 | GO:0006283 | transcription-coupled nucleotide-excision repair(GO:0006283) |
| 0.5 | 1.4 | GO:2000055 | positive regulation of Wnt signaling pathway involved in dorsal/ventral axis specification(GO:2000055) |
| 0.4 | 1.3 | GO:2000402 | response to chemokine(GO:1990868) cellular response to chemokine(GO:1990869) negative regulation of lymphocyte migration(GO:2000402) negative regulation of T cell migration(GO:2000405) |
| 0.4 | 1.7 | GO:0070131 | regulation of mitochondrial translation(GO:0070129) positive regulation of mitochondrial translation(GO:0070131) |
| 0.4 | 2.5 | GO:0010735 | positive regulation of transcription via serum response element binding(GO:0010735) |
| 0.4 | 1.2 | GO:0070914 | UV-damage excision repair(GO:0070914) |
| 0.4 | 4.2 | GO:2000054 | negative regulation of Wnt signaling pathway involved in dorsal/ventral axis specification(GO:2000054) |
| 0.4 | 0.4 | GO:1903430 | negative regulation of cell maturation(GO:1903430) |
| 0.4 | 1.6 | GO:0046951 | ketone body biosynthetic process(GO:0046951) |
| 0.4 | 1.6 | GO:0042754 | negative regulation of circadian rhythm(GO:0042754) |
| 0.4 | 1.2 | GO:1900274 | positive regulation of phospholipase C activity(GO:0010863) regulation of phospholipase C activity(GO:1900274) |
| 0.4 | 7.5 | GO:0031442 | positive regulation of mRNA 3'-end processing(GO:0031442) |
| 0.4 | 1.2 | GO:0030857 | negative regulation of epithelial cell differentiation(GO:0030857) regulation of endothelial cell differentiation(GO:0045601) |
| 0.4 | 2.0 | GO:2001106 | regulation of guanyl-nucleotide exchange factor activity(GO:1905097) regulation of Rho guanyl-nucleotide exchange factor activity(GO:2001106) |
| 0.4 | 3.1 | GO:0051013 | microtubule severing(GO:0051013) |
| 0.4 | 1.1 | GO:1905133 | meiotic cell cycle phase transition(GO:0044771) metaphase/anaphase transition of meiotic cell cycle(GO:0044785) regulation of meiotic cell cycle phase transition(GO:1901993) negative regulation of meiotic cell cycle phase transition(GO:1901994) regulation of metaphase/anaphase transition of meiotic cell cycle(GO:1902102) negative regulation of metaphase/anaphase transition of meiotic cell cycle(GO:1902103) regulation of meiotic chromosome separation(GO:1905132) negative regulation of meiotic chromosome separation(GO:1905133) |
| 0.4 | 1.5 | GO:0034164 | regulation of toll-like receptor 9 signaling pathway(GO:0034163) negative regulation of toll-like receptor 9 signaling pathway(GO:0034164) |
| 0.4 | 1.5 | GO:0045668 | negative regulation of osteoblast differentiation(GO:0045668) |
| 0.4 | 2.2 | GO:0070934 | CRD-mediated mRNA stabilization(GO:0070934) |
| 0.4 | 2.5 | GO:0031048 | chromatin silencing by small RNA(GO:0031048) |
| 0.4 | 6.1 | GO:0015693 | magnesium ion transport(GO:0015693) |
| 0.4 | 1.8 | GO:0036037 | CD8-positive, alpha-beta T cell activation(GO:0036037) |
| 0.4 | 3.9 | GO:0015014 | heparan sulfate proteoglycan biosynthetic process, polysaccharide chain biosynthetic process(GO:0015014) |
| 0.3 | 1.0 | GO:0034035 | sulfate assimilation(GO:0000103) purine ribonucleoside bisphosphate metabolic process(GO:0034035) purine ribonucleoside bisphosphate biosynthetic process(GO:0034036) 3'-phosphoadenosine 5'-phosphosulfate metabolic process(GO:0050427) 3'-phosphoadenosine 5'-phosphosulfate biosynthetic process(GO:0050428) |
| 0.3 | 1.0 | GO:0051026 | chiasma assembly(GO:0051026) |
| 0.3 | 1.0 | GO:0006041 | glucosamine metabolic process(GO:0006041) glucosamine catabolic process(GO:0006043) |
| 0.3 | 1.0 | GO:1901836 | regulation of transcription of nuclear large rRNA transcript from RNA polymerase I promoter(GO:1901836) |
| 0.3 | 2.3 | GO:0040016 | embryonic cleavage(GO:0040016) |
| 0.3 | 1.6 | GO:0002159 | desmosome assembly(GO:0002159) |
| 0.3 | 1.3 | GO:0032119 | sequestering of zinc ion(GO:0032119) regulation of sequestering of zinc ion(GO:0061088) |
| 0.3 | 5.3 | GO:0051764 | actin crosslink formation(GO:0051764) |
| 0.3 | 2.2 | GO:0070584 | mitochondrion morphogenesis(GO:0070584) |
| 0.3 | 4.6 | GO:0048096 | chromatin-mediated maintenance of transcription(GO:0048096) |
| 0.3 | 1.8 | GO:0032964 | collagen biosynthetic process(GO:0032964) |
| 0.3 | 2.5 | GO:0000727 | double-strand break repair via break-induced replication(GO:0000727) |
| 0.3 | 2.4 | GO:0046716 | muscle cell cellular homeostasis(GO:0046716) |
| 0.3 | 3.6 | GO:1904356 | regulation of telomere maintenance via telomere lengthening(GO:1904356) |
| 0.3 | 1.5 | GO:0097355 | protein localization to heterochromatin(GO:0097355) |
| 0.3 | 1.2 | GO:0038093 | Fc receptor signaling pathway(GO:0038093) |
| 0.3 | 1.5 | GO:0006011 | UDP-glucose metabolic process(GO:0006011) |
| 0.3 | 1.4 | GO:0034551 | respiratory chain complex III assembly(GO:0017062) mitochondrial respiratory chain complex III assembly(GO:0034551) mitochondrial respiratory chain complex III biogenesis(GO:0097033) |
| 0.3 | 1.7 | GO:0016578 | histone deubiquitination(GO:0016578) |
| 0.3 | 3.1 | GO:1900052 | regulation of isoprenoid metabolic process(GO:0019747) regulation of vitamin metabolic process(GO:0030656) regulation of retinoic acid biosynthetic process(GO:1900052) |
| 0.3 | 2.8 | GO:0006268 | DNA unwinding involved in DNA replication(GO:0006268) |
| 0.3 | 1.3 | GO:0051145 | smooth muscle cell differentiation(GO:0051145) |
| 0.3 | 1.9 | GO:0006501 | C-terminal protein lipidation(GO:0006501) |
| 0.3 | 3.9 | GO:0010574 | vascular endothelial growth factor production(GO:0010573) regulation of vascular endothelial growth factor production(GO:0010574) positive regulation of vascular endothelial growth factor production(GO:0010575) |
| 0.3 | 1.8 | GO:1902315 | cell cycle DNA replication initiation(GO:1902292) nuclear cell cycle DNA replication initiation(GO:1902315) mitotic DNA replication initiation(GO:1902975) |
| 0.3 | 0.8 | GO:1903646 | regulation of protein folding(GO:1903332) positive regulation of protein folding(GO:1903334) regulation of chaperone-mediated protein folding(GO:1903644) positive regulation of chaperone-mediated protein folding(GO:1903646) |
| 0.3 | 2.6 | GO:0006517 | protein deglycosylation(GO:0006517) |
| 0.3 | 2.0 | GO:0070933 | histone H4 deacetylation(GO:0070933) |
| 0.2 | 1.2 | GO:0043508 | negative regulation of JUN kinase activity(GO:0043508) |
| 0.2 | 8.7 | GO:0005978 | glycogen biosynthetic process(GO:0005978) glucan biosynthetic process(GO:0009250) |
| 0.2 | 1.7 | GO:0070073 | clustering of voltage-gated calcium channels(GO:0070073) |
| 0.2 | 1.9 | GO:1901224 | positive regulation of NIK/NF-kappaB signaling(GO:1901224) |
| 0.2 | 2.8 | GO:0000055 | ribosomal large subunit export from nucleus(GO:0000055) |
| 0.2 | 2.4 | GO:0006108 | malate metabolic process(GO:0006108) |
| 0.2 | 2.4 | GO:0051189 | Mo-molybdopterin cofactor biosynthetic process(GO:0006777) Mo-molybdopterin cofactor metabolic process(GO:0019720) molybdopterin cofactor biosynthetic process(GO:0032324) molybdopterin cofactor metabolic process(GO:0043545) prosthetic group metabolic process(GO:0051189) |
| 0.2 | 0.7 | GO:0051570 | regulation of histone H3-K9 methylation(GO:0051570) |
| 0.2 | 2.1 | GO:0043928 | exonucleolytic nuclear-transcribed mRNA catabolic process involved in deadenylation-dependent decay(GO:0043928) |
| 0.2 | 2.2 | GO:0072393 | microtubule anchoring at microtubule organizing center(GO:0072393) |
| 0.2 | 1.5 | GO:0045053 | protein retention in Golgi apparatus(GO:0045053) |
| 0.2 | 1.5 | GO:0002686 | negative regulation of leukocyte migration(GO:0002686) |
| 0.2 | 0.9 | GO:0070084 | protein initiator methionine removal(GO:0070084) |
| 0.2 | 1.9 | GO:0070498 | interleukin-1-mediated signaling pathway(GO:0070498) |
| 0.2 | 0.6 | GO:0019262 | N-acetylneuraminate catabolic process(GO:0019262) |
| 0.2 | 4.2 | GO:0006896 | Golgi to vacuole transport(GO:0006896) |
| 0.2 | 1.2 | GO:0048011 | neurotrophin TRK receptor signaling pathway(GO:0048011) |
| 0.2 | 2.8 | GO:0045292 | mRNA cis splicing, via spliceosome(GO:0045292) |
| 0.2 | 0.6 | GO:0045598 | regulation of fat cell differentiation(GO:0045598) |
| 0.2 | 1.8 | GO:0044837 | assembly of actomyosin apparatus involved in cytokinesis(GO:0000912) actomyosin contractile ring assembly(GO:0000915) actomyosin contractile ring organization(GO:0044837) |
| 0.2 | 1.0 | GO:0097039 | protein linear polyubiquitination(GO:0097039) |
| 0.2 | 1.4 | GO:2001045 | negative regulation of integrin-mediated signaling pathway(GO:2001045) |
| 0.2 | 1.0 | GO:0008635 | activation of cysteine-type endopeptidase activity involved in apoptotic process by cytochrome c(GO:0008635) |
| 0.2 | 6.2 | GO:0048268 | clathrin coat assembly(GO:0048268) |
| 0.2 | 4.0 | GO:0007520 | myoblast fusion(GO:0007520) |
| 0.2 | 1.3 | GO:1902902 | negative regulation of autophagosome maturation(GO:1901097) negative regulation of autophagosome assembly(GO:1902902) |
| 0.2 | 3.2 | GO:0008354 | germ cell migration(GO:0008354) |
| 0.2 | 3.7 | GO:0090557 | establishment of endothelial intestinal barrier(GO:0090557) |
| 0.2 | 0.6 | GO:0070932 | histone H3 deacetylation(GO:0070932) |
| 0.2 | 1.4 | GO:0044211 | CTP salvage(GO:0044211) |
| 0.2 | 0.9 | GO:0009450 | gamma-aminobutyric acid catabolic process(GO:0009450) |
| 0.2 | 0.3 | GO:0021516 | dorsal spinal cord development(GO:0021516) |
| 0.2 | 1.5 | GO:0007220 | Notch receptor processing(GO:0007220) |
| 0.2 | 14.9 | GO:0061640 | cytoskeleton-dependent cytokinesis(GO:0061640) |
| 0.2 | 1.5 | GO:0043981 | histone H4-K5 acetylation(GO:0043981) histone H4-K8 acetylation(GO:0043982) |
| 0.2 | 1.0 | GO:0010172 | embryonic body morphogenesis(GO:0010172) |
| 0.2 | 0.8 | GO:0046485 | ether lipid metabolic process(GO:0046485) |
| 0.2 | 1.0 | GO:0033146 | regulation of intracellular estrogen receptor signaling pathway(GO:0033146) |
| 0.2 | 2.2 | GO:0032008 | positive regulation of TOR signaling(GO:0032008) |
| 0.2 | 0.6 | GO:0071033 | nuclear retention of pre-mRNA at the site of transcription(GO:0071033) |
| 0.2 | 2.5 | GO:0010507 | negative regulation of autophagy(GO:0010507) |
| 0.2 | 5.5 | GO:0070534 | protein K63-linked ubiquitination(GO:0070534) |
| 0.2 | 2.3 | GO:0019985 | translesion synthesis(GO:0019985) |
| 0.2 | 2.3 | GO:0045116 | protein neddylation(GO:0045116) positive regulation of ubiquitin-protein transferase activity(GO:0051443) |
| 0.2 | 1.2 | GO:0021634 | optic nerve formation(GO:0021634) |
| 0.2 | 2.3 | GO:0051310 | metaphase plate congression(GO:0051310) |
| 0.2 | 2.1 | GO:0042542 | response to hydrogen peroxide(GO:0042542) |
| 0.1 | 1.2 | GO:0000083 | regulation of transcription involved in G1/S transition of mitotic cell cycle(GO:0000083) |
| 0.1 | 0.9 | GO:0016048 | detection of temperature stimulus(GO:0016048) sensory perception of temperature stimulus(GO:0050951) |
| 0.1 | 2.2 | GO:0033173 | calcineurin-NFAT signaling cascade(GO:0033173) |
| 0.1 | 0.9 | GO:0035313 | wound healing, spreading of epidermal cells(GO:0035313) negative regulation of ruffle assembly(GO:1900028) |
| 0.1 | 0.7 | GO:2000042 | negative regulation of double-strand break repair via homologous recombination(GO:2000042) |
| 0.1 | 1.4 | GO:0045003 | double-strand break repair via synthesis-dependent strand annealing(GO:0045003) |
| 0.1 | 1.9 | GO:1990504 | dense core granule exocytosis(GO:1990504) |
| 0.1 | 1.3 | GO:0061316 | canonical Wnt signaling pathway involved in heart development(GO:0061316) |
| 0.1 | 0.6 | GO:0030220 | platelet formation(GO:0030220) platelet morphogenesis(GO:0036344) |
| 0.1 | 0.3 | GO:0070228 | B cell apoptotic process(GO:0001783) regulation of B cell apoptotic process(GO:0002902) regulation of lymphocyte apoptotic process(GO:0070228) |
| 0.1 | 1.4 | GO:1901678 | heme transport(GO:0015886) iron coordination entity transport(GO:1901678) |
| 0.1 | 2.1 | GO:0090110 | cargo loading into COPII-coated vesicle(GO:0090110) |
| 0.1 | 1.8 | GO:0044154 | histone H3-K14 acetylation(GO:0044154) |
| 0.1 | 2.9 | GO:0050796 | regulation of insulin secretion(GO:0050796) |
| 0.1 | 0.6 | GO:0034334 | adherens junction maintenance(GO:0034334) |
| 0.1 | 1.2 | GO:0033387 | putrescine biosynthetic process from ornithine(GO:0033387) |
| 0.1 | 4.1 | GO:0006506 | GPI anchor biosynthetic process(GO:0006506) |
| 0.1 | 0.7 | GO:0036498 | IRE1-mediated unfolded protein response(GO:0036498) |
| 0.1 | 1.3 | GO:0051601 | exocyst localization(GO:0051601) |
| 0.1 | 2.1 | GO:0016926 | protein desumoylation(GO:0016926) |
| 0.1 | 0.8 | GO:1990126 | retrograde transport, endosome to plasma membrane(GO:1990126) |
| 0.1 | 1.5 | GO:0097369 | sodium ion import(GO:0097369) sodium ion import across plasma membrane(GO:0098719) sodium ion import into cell(GO:1990118) |
| 0.1 | 2.2 | GO:0006047 | UDP-N-acetylglucosamine metabolic process(GO:0006047) |
| 0.1 | 1.4 | GO:0006611 | protein export from nucleus(GO:0006611) |
| 0.1 | 2.7 | GO:0034315 | regulation of Arp2/3 complex-mediated actin nucleation(GO:0034315) |
| 0.1 | 1.9 | GO:0090279 | regulation of calcium ion import(GO:0090279) |
| 0.1 | 1.9 | GO:0048922 | posterior lateral line neuromast deposition(GO:0048922) regulation of BMP signaling pathway involved in heart jogging(GO:2000223) |
| 0.1 | 0.5 | GO:0015722 | canalicular bile acid transport(GO:0015722) bile acid secretion(GO:0032782) |
| 0.1 | 2.9 | GO:0048263 | determination of dorsal identity(GO:0048263) |
| 0.1 | 1.9 | GO:0000186 | activation of MAPKK activity(GO:0000186) |
| 0.1 | 4.4 | GO:0032924 | activin receptor signaling pathway(GO:0032924) |
| 0.1 | 5.6 | GO:0003146 | heart jogging(GO:0003146) |
| 0.1 | 2.0 | GO:0008156 | negative regulation of DNA replication(GO:0008156) |
| 0.1 | 2.3 | GO:0008078 | mesodermal cell migration(GO:0008078) |
| 0.1 | 2.2 | GO:0007158 | neuron cell-cell adhesion(GO:0007158) |
| 0.1 | 0.6 | GO:0002431 | Fc receptor mediated stimulatory signaling pathway(GO:0002431) regulation of Fc receptor mediated stimulatory signaling pathway(GO:0060368) |
| 0.1 | 2.2 | GO:0048264 | determination of ventral identity(GO:0048264) |
| 0.1 | 2.7 | GO:0006509 | membrane protein ectodomain proteolysis(GO:0006509) |
| 0.1 | 2.1 | GO:0048026 | positive regulation of mRNA splicing, via spliceosome(GO:0048026) |
| 0.1 | 0.3 | GO:0009229 | thiamine diphosphate biosynthetic process(GO:0009229) |
| 0.1 | 1.2 | GO:0046548 | retinal rod cell development(GO:0046548) |
| 0.1 | 1.1 | GO:0006488 | dolichol-linked oligosaccharide biosynthetic process(GO:0006488) |
| 0.1 | 1.3 | GO:0043403 | skeletal muscle tissue regeneration(GO:0043403) |
| 0.1 | 2.7 | GO:0043666 | regulation of phosphoprotein phosphatase activity(GO:0043666) |
| 0.1 | 0.6 | GO:0015910 | peroxisomal long-chain fatty acid import(GO:0015910) |
| 0.1 | 0.9 | GO:0002082 | regulation of oxidative phosphorylation(GO:0002082) |
| 0.1 | 0.6 | GO:0060217 | hemangioblast cell differentiation(GO:0060217) |
| 0.1 | 0.7 | GO:0043649 | dicarboxylic acid catabolic process(GO:0043649) |
| 0.1 | 0.5 | GO:0090199 | regulation of release of cytochrome c from mitochondria(GO:0090199) positive regulation of release of cytochrome c from mitochondria(GO:0090200) |
| 0.1 | 1.6 | GO:0071577 | zinc II ion transmembrane transport(GO:0071577) |
| 0.1 | 0.9 | GO:0070262 | peptidyl-serine dephosphorylation(GO:0070262) |
| 0.1 | 2.0 | GO:0006743 | ubiquinone metabolic process(GO:0006743) ubiquinone biosynthetic process(GO:0006744) quinone biosynthetic process(GO:1901663) |
| 0.1 | 0.5 | GO:0051659 | maintenance of mitochondrion location(GO:0051659) |
| 0.1 | 0.8 | GO:0044090 | positive regulation of vacuole organization(GO:0044090) |
| 0.1 | 3.9 | GO:0016575 | histone deacetylation(GO:0016575) |
| 0.1 | 0.7 | GO:0045109 | intermediate filament organization(GO:0045109) intermediate filament bundle assembly(GO:0045110) |
| 0.1 | 1.7 | GO:0034032 | nucleoside bisphosphate metabolic process(GO:0033865) ribonucleoside bisphosphate metabolic process(GO:0033875) purine nucleoside bisphosphate metabolic process(GO:0034032) |
| 0.1 | 0.6 | GO:0045050 | protein insertion into ER membrane by stop-transfer membrane-anchor sequence(GO:0045050) |
| 0.1 | 2.6 | GO:0000387 | spliceosomal snRNP assembly(GO:0000387) |
| 0.1 | 1.0 | GO:0042407 | cristae formation(GO:0042407) |
| 0.1 | 1.4 | GO:0006003 | fructose 2,6-bisphosphate metabolic process(GO:0006003) |
| 0.1 | 0.9 | GO:0032781 | positive regulation of ATPase activity(GO:0032781) |
| 0.1 | 1.6 | GO:0031952 | regulation of protein autophosphorylation(GO:0031952) |
| 0.1 | 0.9 | GO:0006616 | SRP-dependent cotranslational protein targeting to membrane, translocation(GO:0006616) |
| 0.1 | 0.3 | GO:0036149 | phosphatidylinositol acyl-chain remodeling(GO:0036149) |
| 0.1 | 0.9 | GO:0042694 | muscle cell fate specification(GO:0042694) |
| 0.1 | 1.8 | GO:0060173 | limb development(GO:0060173) |
| 0.1 | 1.1 | GO:0003406 | retinal pigment epithelium development(GO:0003406) |
| 0.1 | 0.8 | GO:0007076 | mitotic chromosome condensation(GO:0007076) |
| 0.1 | 0.4 | GO:0090646 | mitochondrial tRNA processing(GO:0090646) |
| 0.1 | 1.8 | GO:0001966 | thigmotaxis(GO:0001966) |
| 0.1 | 1.3 | GO:0034314 | Arp2/3 complex-mediated actin nucleation(GO:0034314) |
| 0.1 | 0.5 | GO:0044341 | sodium-dependent phosphate transport(GO:0044341) |
| 0.1 | 0.5 | GO:0045743 | positive regulation of fibroblast growth factor receptor signaling pathway(GO:0045743) |
| 0.1 | 0.7 | GO:1900077 | negative regulation of insulin receptor signaling pathway(GO:0046627) negative regulation of cellular response to insulin stimulus(GO:1900077) |
| 0.1 | 6.6 | GO:0090504 | epiboly(GO:0090504) |
| 0.1 | 0.9 | GO:0030259 | lipid glycosylation(GO:0030259) |
| 0.1 | 0.6 | GO:0060573 | ventral spinal cord interneuron specification(GO:0021521) cell fate specification involved in pattern specification(GO:0060573) |
| 0.1 | 0.2 | GO:0090141 | positive regulation of mitochondrial fission(GO:0090141) |
| 0.1 | 0.3 | GO:0010896 | regulation of triglyceride catabolic process(GO:0010896) positive regulation of triglyceride catabolic process(GO:0010898) |
| 0.1 | 4.3 | GO:0051693 | actin filament capping(GO:0051693) |
| 0.1 | 0.2 | GO:0002076 | osteoblast development(GO:0002076) |
| 0.1 | 0.8 | GO:0051865 | protein autoubiquitination(GO:0051865) |
| 0.1 | 0.5 | GO:0006348 | chromatin silencing at telomere(GO:0006348) |
| 0.1 | 1.4 | GO:0046475 | glycerophospholipid catabolic process(GO:0046475) |
| 0.1 | 0.6 | GO:0042214 | carotene metabolic process(GO:0016119) carotene catabolic process(GO:0016121) terpene metabolic process(GO:0042214) terpene catabolic process(GO:0046247) |
| 0.1 | 0.5 | GO:0046958 | nonassociative learning(GO:0046958) habituation(GO:0046959) |
| 0.1 | 2.1 | GO:0007040 | lysosome organization(GO:0007040) lytic vacuole organization(GO:0080171) |
| 0.1 | 2.6 | GO:0032456 | endocytic recycling(GO:0032456) |
| 0.1 | 2.1 | GO:0071427 | mRNA export from nucleus(GO:0006406) mRNA-containing ribonucleoprotein complex export from nucleus(GO:0071427) |
| 0.1 | 0.7 | GO:0070212 | protein poly-ADP-ribosylation(GO:0070212) |
| 0.1 | 2.8 | GO:0030490 | maturation of SSU-rRNA(GO:0030490) |
| 0.1 | 0.7 | GO:0021702 | cerebellar Purkinje cell layer formation(GO:0021694) cerebellar Purkinje cell differentiation(GO:0021702) |
| 0.1 | 2.7 | GO:0043534 | blood vessel endothelial cell migration(GO:0043534) |
| 0.1 | 0.1 | GO:0051099 | positive regulation of DNA binding(GO:0043388) positive regulation of binding(GO:0051099) regulation of DNA binding(GO:0051101) |
| 0.1 | 1.1 | GO:0021555 | midbrain-hindbrain boundary morphogenesis(GO:0021555) |
| 0.1 | 3.4 | GO:0090090 | negative regulation of canonical Wnt signaling pathway(GO:0090090) |
| 0.1 | 0.2 | GO:0001112 | DNA-templated transcriptional open complex formation(GO:0001112) transcriptional open complex formation at RNA polymerase II promoter(GO:0001113) protein-DNA complex remodeling(GO:0001120) |
| 0.1 | 3.8 | GO:0000724 | double-strand break repair via homologous recombination(GO:0000724) |
| 0.1 | 1.9 | GO:0031532 | actin cytoskeleton reorganization(GO:0031532) |
| 0.1 | 0.5 | GO:1904825 | protein localization to microtubule(GO:0035372) protein localization to microtubule plus-end(GO:1904825) |
| 0.1 | 0.4 | GO:0009584 | detection of visible light(GO:0009584) |
| 0.1 | 0.2 | GO:0030520 | intracellular estrogen receptor signaling pathway(GO:0030520) |
| 0.1 | 0.9 | GO:0030513 | positive regulation of BMP signaling pathway(GO:0030513) |
| 0.1 | 0.7 | GO:0060325 | face morphogenesis(GO:0060325) |
| 0.1 | 1.8 | GO:0043244 | regulation of protein complex disassembly(GO:0043244) |
| 0.0 | 0.5 | GO:0007045 | cell-substrate adherens junction assembly(GO:0007045) focal adhesion assembly(GO:0048041) regulation of focal adhesion assembly(GO:0051893) regulation of cell-substrate junction assembly(GO:0090109) regulation of adherens junction organization(GO:1903391) |
| 0.0 | 1.0 | GO:0051092 | positive regulation of NF-kappaB transcription factor activity(GO:0051092) |
| 0.0 | 6.7 | GO:1990778 | protein localization to plasma membrane(GO:0072659) protein localization to cell periphery(GO:1990778) |
| 0.0 | 2.2 | GO:0007062 | sister chromatid cohesion(GO:0007062) |
| 0.0 | 1.4 | GO:0032011 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.0 | 0.2 | GO:0071543 | diphosphoinositol polyphosphate metabolic process(GO:0071543) diadenosine pentaphosphate metabolic process(GO:1901906) diadenosine pentaphosphate catabolic process(GO:1901907) diadenosine hexaphosphate metabolic process(GO:1901908) diadenosine hexaphosphate catabolic process(GO:1901909) adenosine 5'-(hexahydrogen pentaphosphate) metabolic process(GO:1901910) adenosine 5'-(hexahydrogen pentaphosphate) catabolic process(GO:1901911) |
| 0.0 | 2.3 | GO:0030010 | establishment of cell polarity(GO:0030010) |
| 0.0 | 0.3 | GO:0007175 | negative regulation of epidermal growth factor-activated receptor activity(GO:0007175) |
| 0.0 | 0.2 | GO:0031571 | mitotic G1 DNA damage checkpoint(GO:0031571) |
| 0.0 | 1.7 | GO:0000245 | spliceosomal complex assembly(GO:0000245) |
| 0.0 | 1.6 | GO:0032007 | negative regulation of TOR signaling(GO:0032007) |
| 0.0 | 0.6 | GO:0045324 | late endosome to vacuole transport(GO:0045324) |
| 0.0 | 0.4 | GO:0051877 | pigment granule aggregation in cell center(GO:0051877) |
| 0.0 | 0.6 | GO:0043507 | activation of JUN kinase activity(GO:0007257) regulation of JUN kinase activity(GO:0043506) positive regulation of JUN kinase activity(GO:0043507) |
| 0.0 | 1.0 | GO:0006418 | tRNA aminoacylation for protein translation(GO:0006418) |
| 0.0 | 0.7 | GO:0001731 | formation of translation preinitiation complex(GO:0001731) |
| 0.0 | 0.5 | GO:0038061 | NIK/NF-kappaB signaling(GO:0038061) |
| 0.0 | 2.0 | GO:0042773 | ATP synthesis coupled electron transport(GO:0042773) |
| 0.0 | 0.9 | GO:0007205 | protein kinase C-activating G-protein coupled receptor signaling pathway(GO:0007205) |
| 0.0 | 1.2 | GO:0043652 | engulfment of apoptotic cell(GO:0043652) |
| 0.0 | 0.4 | GO:0035307 | positive regulation of dephosphorylation(GO:0035306) positive regulation of protein dephosphorylation(GO:0035307) |
| 0.0 | 0.9 | GO:0008286 | insulin receptor signaling pathway(GO:0008286) |
| 0.0 | 1.4 | GO:0001676 | long-chain fatty acid metabolic process(GO:0001676) |
| 0.0 | 0.4 | GO:0031468 | nuclear envelope reassembly(GO:0031468) |
| 0.0 | 0.8 | GO:0072666 | establishment of protein localization to vacuole(GO:0072666) |
| 0.0 | 2.0 | GO:0007528 | neuromuscular junction development(GO:0007528) |
| 0.0 | 1.7 | GO:0099515 | vesicle transport along actin filament(GO:0030050) actin filament-based transport(GO:0099515) |
| 0.0 | 0.2 | GO:1900027 | regulation of ruffle assembly(GO:1900027) positive regulation of ruffle assembly(GO:1900029) |
| 0.0 | 0.3 | GO:0061709 | reticulophagy(GO:0061709) |
| 0.0 | 0.7 | GO:0035329 | hippo signaling(GO:0035329) |
| 0.0 | 2.4 | GO:0016573 | histone acetylation(GO:0016573) |
| 0.0 | 1.0 | GO:0007340 | acrosome reaction(GO:0007340) |
| 0.0 | 1.5 | GO:0030514 | negative regulation of BMP signaling pathway(GO:0030514) |
| 0.0 | 0.2 | GO:0014829 | vascular smooth muscle contraction(GO:0014829) |
| 0.0 | 0.5 | GO:0030512 | negative regulation of transforming growth factor beta receptor signaling pathway(GO:0030512) negative regulation of cellular response to transforming growth factor beta stimulus(GO:1903845) |
| 0.0 | 0.9 | GO:0007006 | mitochondrial membrane organization(GO:0007006) |
| 0.0 | 0.8 | GO:0007031 | peroxisome organization(GO:0007031) |
| 0.0 | 0.7 | GO:0000027 | ribosomal large subunit assembly(GO:0000027) |
| 0.0 | 0.6 | GO:0034472 | snRNA 3'-end processing(GO:0034472) |
| 0.0 | 0.7 | GO:0048538 | thymus development(GO:0048538) |
| 0.0 | 0.6 | GO:0043486 | histone exchange(GO:0043486) |
| 0.0 | 0.7 | GO:0014029 | neural crest formation(GO:0014029) |
| 0.0 | 0.4 | GO:1905150 | regulation of voltage-gated sodium channel activity(GO:1905150) |
| 0.0 | 1.0 | GO:0050906 | detection of stimulus involved in sensory perception(GO:0050906) |
| 0.0 | 1.5 | GO:0007098 | centrosome cycle(GO:0007098) |
| 0.0 | 0.6 | GO:0032786 | positive regulation of DNA-templated transcription, elongation(GO:0032786) |
| 0.0 | 0.5 | GO:0043113 | receptor clustering(GO:0043113) |
| 0.0 | 0.6 | GO:2001236 | regulation of extrinsic apoptotic signaling pathway(GO:2001236) |
| 0.0 | 0.6 | GO:0043171 | peptide catabolic process(GO:0043171) |
| 0.0 | 0.8 | GO:0051090 | regulation of sequence-specific DNA binding transcription factor activity(GO:0051090) |
| 0.0 | 0.8 | GO:0031290 | retinal ganglion cell axon guidance(GO:0031290) |
| 0.0 | 0.5 | GO:0009214 | cyclic nucleotide catabolic process(GO:0009214) |
| 0.0 | 1.4 | GO:0006909 | phagocytosis(GO:0006909) |
| 0.0 | 4.9 | GO:0006869 | lipid transport(GO:0006869) |
| 0.0 | 3.5 | GO:0006814 | sodium ion transport(GO:0006814) |
| 0.0 | 0.1 | GO:0009649 | entrainment of circadian clock(GO:0009649) entrainment of circadian clock by photoperiod(GO:0043153) |
| 0.0 | 0.4 | GO:0048794 | swim bladder development(GO:0048794) |
| 0.0 | 0.7 | GO:0032922 | circadian regulation of gene expression(GO:0032922) |
| 0.0 | 0.2 | GO:0045920 | negative regulation of exocytosis(GO:0045920) |
| 0.0 | 1.9 | GO:0090630 | activation of GTPase activity(GO:0090630) |
| 0.0 | 0.1 | GO:0007608 | sensory perception of smell(GO:0007608) |
| 0.0 | 0.2 | GO:0038007 | netrin-activated signaling pathway(GO:0038007) |
| 0.0 | 1.5 | GO:0006338 | chromatin remodeling(GO:0006338) |
| 0.0 | 0.3 | GO:0003094 | glomerular filtration(GO:0003094) |
| 0.0 | 0.7 | GO:0018107 | peptidyl-threonine phosphorylation(GO:0018107) |
| 0.0 | 8.9 | GO:0030029 | actin filament-based process(GO:0030029) |
| 0.0 | 0.0 | GO:0043504 | mitochondrial DNA repair(GO:0043504) |
| 0.0 | 0.1 | GO:0035778 | pronephric nephron tubule epithelial cell differentiation(GO:0035778) cell differentiation involved in pronephros development(GO:0039014) nephron tubule epithelial cell differentiation(GO:0072160) |
| 0.0 | 0.1 | GO:0010960 | magnesium ion homeostasis(GO:0010960) |
| 0.0 | 0.3 | GO:0070936 | protein K48-linked ubiquitination(GO:0070936) |
| 0.0 | 0.0 | GO:0002407 | dendritic cell chemotaxis(GO:0002407) dendritic cell migration(GO:0036336) |
| 0.0 | 0.0 | GO:0010430 | fatty acid omega-oxidation(GO:0010430) |
| 0.0 | 0.1 | GO:0097241 | hematopoietic stem cell migration to bone marrow(GO:0097241) |
| 0.0 | 0.3 | GO:0006541 | glutamine metabolic process(GO:0006541) |
| 0.0 | 0.4 | GO:0000910 | cytokinesis(GO:0000910) regulation of cytokinesis(GO:0032465) |
| 0.0 | 0.1 | GO:0001837 | epithelial to mesenchymal transition(GO:0001837) |
| 0.0 | 1.4 | GO:0008033 | tRNA processing(GO:0008033) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.3 | 6.6 | GO:0070390 | transcription export complex 2(GO:0070390) |
| 1.1 | 4.5 | GO:0097524 | sperm plasma membrane(GO:0097524) |
| 1.0 | 4.0 | GO:0035301 | Hedgehog signaling complex(GO:0035301) |
| 0.8 | 10.1 | GO:0005964 | phosphorylase kinase complex(GO:0005964) |
| 0.8 | 2.3 | GO:0098536 | deuterosome(GO:0098536) |
| 0.7 | 3.0 | GO:0035339 | SPOTS complex(GO:0035339) |
| 0.7 | 5.0 | GO:0034518 | RNA cap binding complex(GO:0034518) |
| 0.7 | 2.9 | GO:0098890 | extrinsic component of postsynaptic membrane(GO:0098890) |
| 0.6 | 1.9 | GO:0031213 | RSF complex(GO:0031213) |
| 0.6 | 1.8 | GO:0030686 | 90S preribosome(GO:0030686) |
| 0.6 | 2.3 | GO:0097268 | cytoophidium(GO:0097268) |
| 0.5 | 3.8 | GO:0030877 | beta-catenin destruction complex(GO:0030877) |
| 0.5 | 2.6 | GO:0008091 | spectrin(GO:0008091) |
| 0.5 | 2.0 | GO:1990498 | mitotic spindle microtubule(GO:1990498) |
| 0.5 | 2.9 | GO:0000938 | GARP complex(GO:0000938) |
| 0.5 | 3.8 | GO:0001401 | mitochondrial sorting and assembly machinery complex(GO:0001401) |
| 0.5 | 3.7 | GO:0031231 | integral component of peroxisomal membrane(GO:0005779) intrinsic component of peroxisomal membrane(GO:0031231) |
| 0.4 | 2.1 | GO:1990131 | EGO complex(GO:0034448) Gtr1-Gtr2 GTPase complex(GO:1990131) |
| 0.4 | 2.4 | GO:0019008 | molybdopterin synthase complex(GO:0019008) |
| 0.4 | 9.3 | GO:0071141 | SMAD protein complex(GO:0071141) |
| 0.4 | 0.7 | GO:1990745 | EARP complex(GO:1990745) |
| 0.4 | 1.4 | GO:0043540 | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase complex(GO:0043540) |
| 0.3 | 1.4 | GO:0005672 | transcription factor TFIIA complex(GO:0005672) |
| 0.3 | 3.6 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.3 | 2.3 | GO:0072669 | tRNA-splicing ligase complex(GO:0072669) |
| 0.3 | 1.2 | GO:0000110 | nucleotide-excision repair factor 1 complex(GO:0000110) |
| 0.3 | 1.2 | GO:0032584 | growth cone membrane(GO:0032584) |
| 0.3 | 2.7 | GO:0089701 | U2AF(GO:0089701) |
| 0.3 | 11.0 | GO:0031105 | septin ring(GO:0005940) septin complex(GO:0031105) septin cytoskeleton(GO:0032156) |
| 0.3 | 1.7 | GO:0030132 | clathrin coat of coated pit(GO:0030132) |
| 0.3 | 3.9 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.3 | 2.2 | GO:0009295 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.3 | 3.8 | GO:0002102 | podosome(GO:0002102) |
| 0.3 | 2.2 | GO:0070937 | CRD-mediated mRNA stability complex(GO:0070937) |
| 0.3 | 0.8 | GO:0072380 | TRC complex(GO:0072380) |
| 0.2 | 1.7 | GO:0000445 | THO complex(GO:0000347) THO complex part of transcription export complex(GO:0000445) |
| 0.2 | 3.2 | GO:0030015 | CCR4-NOT core complex(GO:0030015) |
| 0.2 | 2.6 | GO:0005665 | DNA-directed RNA polymerase II, core complex(GO:0005665) |
| 0.2 | 3.6 | GO:0070187 | telosome(GO:0070187) |
| 0.2 | 1.9 | GO:0034045 | pre-autophagosomal structure membrane(GO:0034045) |
| 0.2 | 0.7 | GO:1990604 | IRE1-TRAF2-ASK1 complex(GO:1990604) |
| 0.2 | 3.2 | GO:0048179 | activin receptor complex(GO:0048179) |
| 0.2 | 3.6 | GO:0071564 | npBAF complex(GO:0071564) |
| 0.2 | 2.8 | GO:0071007 | U2-type catalytic step 2 spliceosome(GO:0071007) |
| 0.2 | 2.4 | GO:0098835 | presynaptic endocytic zone(GO:0098833) presynaptic endocytic zone membrane(GO:0098835) extrinsic component of presynaptic membrane(GO:0098888) extrinsic component of presynaptic endocytic zone(GO:0098894) |
| 0.2 | 5.2 | GO:0048787 | presynaptic active zone membrane(GO:0048787) |
| 0.2 | 1.3 | GO:0000836 | ER ubiquitin ligase complex(GO:0000835) Hrd1p ubiquitin ligase complex(GO:0000836) |
| 0.2 | 1.7 | GO:0035032 | phosphatidylinositol 3-kinase complex, class III(GO:0035032) |
| 0.2 | 3.9 | GO:0016581 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) |
| 0.2 | 8.2 | GO:0030496 | midbody(GO:0030496) |
| 0.2 | 1.5 | GO:0070765 | gamma-secretase complex(GO:0070765) |
| 0.2 | 1.7 | GO:0032593 | insulin-responsive compartment(GO:0032593) |
| 0.2 | 2.6 | GO:0097038 | perinuclear endoplasmic reticulum(GO:0097038) |
| 0.2 | 11.3 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.2 | 21.3 | GO:0005923 | bicellular tight junction(GO:0005923) occluding junction(GO:0070160) |
| 0.2 | 1.0 | GO:0031428 | box C/D snoRNP complex(GO:0031428) |
| 0.1 | 1.5 | GO:0032300 | mismatch repair complex(GO:0032300) |
| 0.1 | 1.0 | GO:0000782 | telomere cap complex(GO:0000782) nuclear telomere cap complex(GO:0000783) nuclear chromosome, telomeric region(GO:0000784) |
| 0.1 | 1.0 | GO:0031464 | Cul4A-RING E3 ubiquitin ligase complex(GO:0031464) |
| 0.1 | 8.7 | GO:0000118 | histone deacetylase complex(GO:0000118) |
| 0.1 | 0.7 | GO:0033596 | TSC1-TSC2 complex(GO:0033596) |
| 0.1 | 1.5 | GO:0035101 | FACT complex(GO:0035101) |
| 0.1 | 1.8 | GO:0042555 | MCM complex(GO:0042555) |
| 0.1 | 2.3 | GO:0030057 | desmosome(GO:0030057) |
| 0.1 | 0.5 | GO:0016460 | myosin II complex(GO:0016460) |
| 0.1 | 2.0 | GO:0000813 | ESCRT I complex(GO:0000813) |
| 0.1 | 3.1 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.1 | 1.0 | GO:0071797 | LUBAC complex(GO:0071797) |
| 0.1 | 0.7 | GO:0030914 | STAGA complex(GO:0030914) |
| 0.1 | 0.7 | GO:0048500 | signal recognition particle, endoplasmic reticulum targeting(GO:0005786) signal recognition particle(GO:0048500) |
| 0.1 | 3.4 | GO:0001725 | stress fiber(GO:0001725) contractile actin filament bundle(GO:0097517) |
| 0.1 | 5.0 | GO:0031941 | filamentous actin(GO:0031941) |
| 0.1 | 1.8 | GO:0070776 | H3 histone acetyltransferase complex(GO:0070775) MOZ/MORF histone acetyltransferase complex(GO:0070776) |
| 0.1 | 1.0 | GO:0000812 | Swr1 complex(GO:0000812) |
| 0.1 | 2.1 | GO:0030127 | COPII vesicle coat(GO:0030127) |
| 0.1 | 2.0 | GO:0005685 | U1 snRNP(GO:0005685) |
| 0.1 | 24.8 | GO:0005764 | lysosome(GO:0005764) |
| 0.1 | 7.9 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.1 | 5.3 | GO:0030139 | endocytic vesicle(GO:0030139) |
| 0.1 | 5.9 | GO:0000779 | condensed chromosome, centromeric region(GO:0000779) |
| 0.1 | 1.0 | GO:0061617 | MICOS complex(GO:0061617) |
| 0.1 | 1.0 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.1 | 2.0 | GO:0035102 | PRC1 complex(GO:0035102) |
| 0.1 | 1.3 | GO:1990124 | messenger ribonucleoprotein complex(GO:1990124) |
| 0.1 | 2.7 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.1 | 1.9 | GO:0030667 | secretory granule membrane(GO:0030667) |
| 0.1 | 1.4 | GO:0032588 | trans-Golgi network membrane(GO:0032588) |
| 0.1 | 7.3 | GO:0030027 | lamellipodium(GO:0030027) |
| 0.1 | 0.5 | GO:0005614 | interstitial matrix(GO:0005614) |
| 0.1 | 1.9 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.1 | 1.2 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.1 | 0.4 | GO:1902636 | kinociliary basal body(GO:1902636) |
| 0.1 | 0.9 | GO:0000815 | ESCRT III complex(GO:0000815) |
| 0.1 | 4.8 | GO:0000793 | condensed chromosome(GO:0000793) |
| 0.1 | 1.2 | GO:0044666 | MLL3/4 complex(GO:0044666) |
| 0.1 | 15.8 | GO:0005813 | centrosome(GO:0005813) |
| 0.1 | 8.6 | GO:0031234 | extrinsic component of cytoplasmic side of plasma membrane(GO:0031234) |
| 0.1 | 0.6 | GO:0019908 | cyclin/CDK positive transcription elongation factor complex(GO:0008024) nuclear cyclin-dependent protein kinase holoenzyme complex(GO:0019908) |
| 0.1 | 3.6 | GO:0005793 | endoplasmic reticulum-Golgi intermediate compartment(GO:0005793) |
| 0.1 | 2.0 | GO:0016605 | PML body(GO:0016605) |
| 0.1 | 1.0 | GO:0030134 | ER to Golgi transport vesicle(GO:0030134) |
| 0.1 | 15.4 | GO:0005730 | nucleolus(GO:0005730) |
| 0.1 | 5.0 | GO:0016607 | nuclear speck(GO:0016607) |
| 0.0 | 1.3 | GO:0000145 | exocyst(GO:0000145) |
| 0.0 | 2.0 | GO:0005637 | nuclear inner membrane(GO:0005637) |
| 0.0 | 0.6 | GO:1902562 | NuA4 histone acetyltransferase complex(GO:0035267) H4/H2A histone acetyltransferase complex(GO:0043189) H4 histone acetyltransferase complex(GO:1902562) |
| 0.0 | 2.0 | GO:0016342 | catenin complex(GO:0016342) |
| 0.0 | 1.4 | GO:0005844 | polysome(GO:0005844) |
| 0.0 | 2.4 | GO:0016592 | mediator complex(GO:0016592) |
| 0.0 | 0.4 | GO:0036449 | microtubule minus-end(GO:0036449) |
| 0.0 | 4.3 | GO:0042579 | peroxisome(GO:0005777) microbody(GO:0042579) |
| 0.0 | 4.8 | GO:0005681 | spliceosomal complex(GO:0005681) |
| 0.0 | 0.2 | GO:0005673 | transcription factor TFIIE complex(GO:0005673) |
| 0.0 | 0.5 | GO:0005903 | brush border(GO:0005903) |
| 0.0 | 0.5 | GO:0005885 | Arp2/3 protein complex(GO:0005885) |
| 0.0 | 0.6 | GO:0032039 | integrator complex(GO:0032039) |
| 0.0 | 2.0 | GO:0005769 | early endosome(GO:0005769) |
| 0.0 | 2.4 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.0 | 1.7 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.0 | 0.7 | GO:0030135 | coated vesicle(GO:0030135) |
| 0.0 | 0.5 | GO:0000776 | kinetochore(GO:0000776) |
| 0.0 | 0.2 | GO:0031010 | ISWI-type complex(GO:0031010) |
| 0.0 | 1.6 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.0 | 1.5 | GO:0005884 | actin filament(GO:0005884) |
| 0.0 | 1.2 | GO:0000123 | histone acetyltransferase complex(GO:0000123) |
| 0.0 | 2.1 | GO:0005925 | focal adhesion(GO:0005925) |
| 0.0 | 0.1 | GO:0033503 | HULC complex(GO:0033503) |
| 0.0 | 0.4 | GO:0031083 | BLOC-1 complex(GO:0031083) |
| 0.0 | 0.3 | GO:0017119 | Golgi transport complex(GO:0017119) |
| 0.0 | 4.4 | GO:0005768 | endosome(GO:0005768) |
| 0.0 | 0.2 | GO:0017053 | transcriptional repressor complex(GO:0017053) |
| 0.0 | 1.1 | GO:0031228 | integral component of Golgi membrane(GO:0030173) intrinsic component of Golgi membrane(GO:0031228) |
| 0.0 | 0.1 | GO:0016586 | RSC complex(GO:0016586) |
| 0.0 | 0.1 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.0 | 9.4 | GO:0005794 | Golgi apparatus(GO:0005794) |
| 0.0 | 0.6 | GO:0098839 | postsynaptic density membrane(GO:0098839) postsynaptic specialization membrane(GO:0099634) |
| 0.0 | 0.2 | GO:0000775 | chromosome, centromeric region(GO:0000775) |
| 0.0 | 1.3 | GO:0016459 | myosin complex(GO:0016459) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.1 | 6.3 | GO:0045174 | glutathione dehydrogenase (ascorbate) activity(GO:0045174) |
| 2.0 | 10.1 | GO:0004689 | phosphorylase kinase activity(GO:0004689) |
| 1.2 | 6.0 | GO:0004735 | pyrroline-5-carboxylate reductase activity(GO:0004735) |
| 1.0 | 5.2 | GO:0035197 | siRNA binding(GO:0035197) |
| 1.0 | 2.9 | GO:0080122 | coenzyme A transmembrane transporter activity(GO:0015228) adenosine 3',5'-bisphosphate transmembrane transporter activity(GO:0071077) AMP transmembrane transporter activity(GO:0080122) |
| 0.8 | 2.4 | GO:0070336 | flap-structured DNA binding(GO:0070336) |
| 0.8 | 2.3 | GO:0008517 | folic acid transporter activity(GO:0008517) FAD transmembrane transporter activity(GO:0015230) |
| 0.7 | 4.5 | GO:0008126 | acetylesterase activity(GO:0008126) |
| 0.7 | 2.2 | GO:0008761 | UDP-N-acetylglucosamine 2-epimerase activity(GO:0008761) N-acylmannosamine kinase activity(GO:0009384) |
| 0.7 | 2.9 | GO:0051377 | mannose-ethanolamine phosphotransferase activity(GO:0051377) |
| 0.7 | 6.3 | GO:0009378 | four-way junction helicase activity(GO:0009378) |
| 0.6 | 1.8 | GO:1902945 | metalloendopeptidase activity involved in amyloid precursor protein catabolic process(GO:1902945) |
| 0.6 | 2.4 | GO:0004473 | malic enzyme activity(GO:0004470) malate dehydrogenase (decarboxylating) (NAD+) activity(GO:0004471) malate dehydrogenase (decarboxylating) (NADP+) activity(GO:0004473) |
| 0.6 | 2.4 | GO:0030366 | molybdopterin synthase activity(GO:0030366) |
| 0.6 | 1.7 | GO:0008440 | inositol-1,4,5-trisphosphate 3-kinase activity(GO:0008440) |
| 0.6 | 2.3 | GO:0003883 | CTP synthase activity(GO:0003883) |
| 0.6 | 4.5 | GO:0001217 | bacterial-type RNA polymerase transcription factor activity, sequence-specific DNA binding(GO:0001130) bacterial-type RNA polymerase transcriptional repressor activity, sequence-specific DNA binding(GO:0001217) |
| 0.6 | 5.0 | GO:0016922 | ligand-dependent nuclear receptor binding(GO:0016922) |
| 0.5 | 3.2 | GO:0016361 | activin receptor activity, type I(GO:0016361) |
| 0.5 | 2.0 | GO:0048487 | beta-tubulin binding(GO:0048487) |
| 0.5 | 2.0 | GO:0051139 | calcium:proton antiporter activity(GO:0015369) metal ion:proton antiporter activity(GO:0051139) |
| 0.5 | 3.4 | GO:0016211 | glutamate-ammonia ligase activity(GO:0004356) ammonia ligase activity(GO:0016211) acid-ammonia (or amide) ligase activity(GO:0016880) |
| 0.5 | 3.9 | GO:0015016 | [heparan sulfate]-glucosamine N-sulfotransferase activity(GO:0015016) |
| 0.5 | 3.2 | GO:0070698 | type I activin receptor binding(GO:0070698) |
| 0.4 | 2.5 | GO:0043914 | NADPH:sulfur oxidoreductase activity(GO:0043914) |
| 0.4 | 4.2 | GO:0034647 | histone demethylase activity (H3-trimethyl-K4 specific)(GO:0034647) |
| 0.4 | 2.5 | GO:0008349 | MAP kinase kinase kinase kinase activity(GO:0008349) |
| 0.4 | 1.6 | GO:0004419 | hydroxymethylglutaryl-CoA lyase activity(GO:0004419) |
| 0.4 | 1.2 | GO:0043621 | protein self-association(GO:0043621) |
| 0.4 | 3.7 | GO:0045294 | alpha-catenin binding(GO:0045294) |
| 0.4 | 6.2 | GO:0032454 | histone demethylase activity (H3-K9 specific)(GO:0032454) |
| 0.4 | 1.2 | GO:0008454 | alpha-1,3-mannosylglycoprotein 4-beta-N-acetylglucosaminyltransferase activity(GO:0008454) |
| 0.4 | 4.7 | GO:0070577 | lysine-acetylated histone binding(GO:0070577) |
| 0.3 | 1.0 | GO:0004781 | adenylylsulfate kinase activity(GO:0004020) sulfate adenylyltransferase activity(GO:0004779) sulfate adenylyltransferase (ATP) activity(GO:0004781) |
| 0.3 | 6.9 | GO:0070411 | I-SMAD binding(GO:0070411) |
| 0.3 | 5.2 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 0.3 | 1.0 | GO:0071532 | ankyrin repeat binding(GO:0071532) |
| 0.3 | 1.0 | GO:0004342 | glucosamine-6-phosphate deaminase activity(GO:0004342) |
| 0.3 | 6.7 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.3 | 6.5 | GO:0017136 | histone deacetylase activity(GO:0004407) NAD-dependent histone deacetylase activity(GO:0017136) |
| 0.3 | 1.8 | GO:0016715 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced ascorbate as one donor, and incorporation of one atom of oxygen(GO:0016715) |
| 0.3 | 1.5 | GO:0003983 | UTP:glucose-1-phosphate uridylyltransferase activity(GO:0003983) UTP-monosaccharide-1-phosphate uridylyltransferase activity(GO:0051748) |
| 0.3 | 0.8 | GO:0016971 | flavin-linked sulfhydryl oxidase activity(GO:0016971) |
| 0.3 | 2.7 | GO:0030628 | pre-mRNA 3'-splice site binding(GO:0030628) |
| 0.3 | 2.7 | GO:0008568 | microtubule-severing ATPase activity(GO:0008568) |
| 0.3 | 6.4 | GO:0031210 | phosphatidylcholine binding(GO:0031210) |
| 0.3 | 2.1 | GO:0030619 | U1 snRNA binding(GO:0030619) |
| 0.3 | 2.1 | GO:0070139 | ubiquitin-like protein-specific endopeptidase activity(GO:0070137) SUMO-specific endopeptidase activity(GO:0070139) |
| 0.3 | 2.7 | GO:0035925 | AU-rich element binding(GO:0017091) mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.3 | 2.9 | GO:0043325 | phosphatidylinositol-3,4-bisphosphate binding(GO:0043325) |
| 0.3 | 3.4 | GO:0005332 | gamma-aminobutyric acid:sodium symporter activity(GO:0005332) |
| 0.2 | 1.2 | GO:0004679 | AMP-activated protein kinase activity(GO:0004679) |
| 0.2 | 2.5 | GO:0052629 | phosphatidylinositol-3,5-bisphosphate 3-phosphatase activity(GO:0052629) |
| 0.2 | 2.2 | GO:0004096 | catalase activity(GO:0004096) |
| 0.2 | 1.5 | GO:0016212 | kynurenine-oxoglutarate transaminase activity(GO:0016212) kynurenine aminotransferase activity(GO:0036137) |
| 0.2 | 0.9 | GO:0030626 | U12 snRNA binding(GO:0030626) pre-mRNA intronic binding(GO:0097157) |
| 0.2 | 1.5 | GO:0035173 | histone kinase activity(GO:0035173) |
| 0.2 | 3.7 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.2 | 3.2 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.2 | 2.2 | GO:0048027 | mRNA 5'-UTR binding(GO:0048027) |
| 0.2 | 4.3 | GO:0004065 | arylsulfatase activity(GO:0004065) |
| 0.2 | 3.7 | GO:0015299 | solute:proton antiporter activity(GO:0015299) |
| 0.2 | 1.0 | GO:0071074 | eukaryotic initiation factor eIF2 binding(GO:0071074) |
| 0.2 | 0.6 | GO:0004113 | 2',3'-cyclic-nucleotide 3'-phosphodiesterase activity(GO:0004113) |
| 0.2 | 2.7 | GO:0032050 | clathrin heavy chain binding(GO:0032050) |
| 0.2 | 4.8 | GO:0001103 | RNA polymerase II repressing transcription factor binding(GO:0001103) |
| 0.2 | 1.4 | GO:0004849 | uridine kinase activity(GO:0004849) |
| 0.2 | 1.8 | GO:0005078 | MAP-kinase scaffold activity(GO:0005078) |
| 0.2 | 1.4 | GO:0015232 | heme transporter activity(GO:0015232) |
| 0.2 | 0.9 | GO:0016774 | phosphotransferase activity, carboxyl group as acceptor(GO:0016774) |
| 0.2 | 0.9 | GO:0030942 | endoplasmic reticulum signal peptide binding(GO:0030942) |
| 0.2 | 3.6 | GO:0050321 | tau-protein kinase activity(GO:0050321) |
| 0.2 | 2.4 | GO:0000340 | RNA 7-methylguanosine cap binding(GO:0000340) |
| 0.2 | 1.0 | GO:0030331 | estrogen receptor binding(GO:0030331) |
| 0.2 | 3.0 | GO:0072542 | phosphatase activator activity(GO:0019211) protein phosphatase activator activity(GO:0072542) |
| 0.1 | 1.9 | GO:0070491 | repressing transcription factor binding(GO:0070491) |
| 0.1 | 2.2 | GO:0003995 | acyl-CoA dehydrogenase activity(GO:0003995) |
| 0.1 | 1.7 | GO:0042285 | xylosyltransferase activity(GO:0042285) |
| 0.1 | 1.4 | GO:0005049 | nuclear export signal receptor activity(GO:0005049) |
| 0.1 | 2.4 | GO:0019894 | kinesin binding(GO:0019894) |
| 0.1 | 1.2 | GO:0004586 | ornithine decarboxylase activity(GO:0004586) |
| 0.1 | 1.4 | GO:0003727 | single-stranded RNA binding(GO:0003727) |
| 0.1 | 3.7 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.1 | 1.8 | GO:0010484 | H3 histone acetyltransferase activity(GO:0010484) histone acetyltransferase activity (H3-K23 specific)(GO:0043994) |
| 0.1 | 0.4 | GO:0004736 | pyruvate carboxylase activity(GO:0004736) |
| 0.1 | 0.5 | GO:0015432 | canalicular bile acid transmembrane transporter activity(GO:0015126) bile acid-exporting ATPase activity(GO:0015432) |
| 0.1 | 2.4 | GO:0009982 | pseudouridine synthase activity(GO:0009982) |
| 0.1 | 0.5 | GO:0031697 | beta-1 adrenergic receptor binding(GO:0031697) type II activin receptor binding(GO:0070699) |
| 0.1 | 0.5 | GO:0032028 | myosin head/neck binding(GO:0032028) myosin II head/neck binding(GO:0032034) myosin II heavy chain binding(GO:0032038) |
| 0.1 | 0.6 | GO:0070644 | vitamin D response element binding(GO:0070644) |
| 0.1 | 3.1 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 0.1 | 1.2 | GO:0070273 | phosphatidylinositol-4-phosphate binding(GO:0070273) |
| 0.1 | 3.2 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.1 | 1.2 | GO:0005391 | sodium:potassium-exchanging ATPase activity(GO:0005391) potassium-transporting ATPase activity(GO:0008556) |
| 0.1 | 3.6 | GO:0042162 | telomeric DNA binding(GO:0042162) |
| 0.1 | 2.3 | GO:0097602 | cullin family protein binding(GO:0097602) |
| 0.1 | 1.5 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 0.1 | 0.3 | GO:0004788 | thiamine diphosphokinase activity(GO:0004788) thiamine binding(GO:0030975) |
| 0.1 | 2.7 | GO:0032266 | phosphatidylinositol-3-phosphate binding(GO:0032266) |
| 0.1 | 15.9 | GO:0004197 | cysteine-type endopeptidase activity(GO:0004197) |
| 0.1 | 0.6 | GO:0004997 | thyrotropin-releasing hormone receptor activity(GO:0004997) |
| 0.1 | 2.0 | GO:0008242 | omega peptidase activity(GO:0008242) |
| 0.1 | 1.1 | GO:0047498 | calcium-dependent phospholipase A2 activity(GO:0047498) |
| 0.1 | 1.4 | GO:0003873 | 6-phosphofructo-2-kinase activity(GO:0003873) |
| 0.1 | 0.4 | GO:0052905 | tRNA (guanine(9)-N(1))-methyltransferase activity(GO:0052905) |
| 0.1 | 0.9 | GO:0008484 | sulfuric ester hydrolase activity(GO:0008484) |
| 0.1 | 1.3 | GO:0002039 | p53 binding(GO:0002039) |
| 0.1 | 2.6 | GO:0003899 | DNA-directed RNA polymerase activity(GO:0003899) RNA polymerase activity(GO:0034062) |
| 0.1 | 0.3 | GO:0016155 | formyltetrahydrofolate dehydrogenase activity(GO:0016155) |
| 0.1 | 18.4 | GO:0042393 | histone binding(GO:0042393) |
| 0.1 | 1.0 | GO:0050700 | CARD domain binding(GO:0050700) |
| 0.1 | 2.6 | GO:0015248 | sterol transporter activity(GO:0015248) |
| 0.1 | 1.9 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.1 | 1.2 | GO:0005092 | GDP-dissociation inhibitor activity(GO:0005092) |
| 0.1 | 1.2 | GO:0005007 | fibroblast growth factor-activated receptor activity(GO:0005007) |
| 0.1 | 0.8 | GO:0016714 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced pteridine as one donor, and incorporation of one atom of oxygen(GO:0016714) |
| 0.1 | 0.6 | GO:0004528 | phosphodiesterase I activity(GO:0004528) |
| 0.1 | 1.2 | GO:0046975 | histone methyltransferase activity (H3-K36 specific)(GO:0046975) |
| 0.1 | 1.4 | GO:0017025 | TBP-class protein binding(GO:0017025) |
| 0.1 | 1.9 | GO:0000993 | RNA polymerase II core binding(GO:0000993) |
| 0.1 | 2.0 | GO:0004659 | prenyltransferase activity(GO:0004659) |
| 0.1 | 0.6 | GO:0019136 | deoxynucleoside kinase activity(GO:0019136) |
| 0.1 | 0.3 | GO:2001070 | starch binding(GO:2001070) |
| 0.1 | 4.2 | GO:0017124 | SH3 domain binding(GO:0017124) |
| 0.1 | 3.8 | GO:0001085 | RNA polymerase II transcription factor binding(GO:0001085) |
| 0.1 | 6.2 | GO:0003697 | single-stranded DNA binding(GO:0003697) |
| 0.1 | 0.5 | GO:0005436 | sodium:phosphate symporter activity(GO:0005436) |
| 0.1 | 1.0 | GO:0005432 | calcium:sodium antiporter activity(GO:0005432) |
| 0.1 | 3.8 | GO:0051539 | 4 iron, 4 sulfur cluster binding(GO:0051539) |
| 0.1 | 1.9 | GO:0004697 | protein kinase C activity(GO:0004697) calcium-dependent protein kinase C activity(GO:0004698) |
| 0.1 | 0.4 | GO:0043023 | ribosomal large subunit binding(GO:0043023) |
| 0.1 | 3.9 | GO:0003725 | double-stranded RNA binding(GO:0003725) |
| 0.1 | 0.9 | GO:0005158 | insulin receptor binding(GO:0005158) |
| 0.1 | 0.4 | GO:0016312 | inositol bisphosphate phosphatase activity(GO:0016312) |
| 0.1 | 0.4 | GO:0047429 | nucleoside-triphosphate diphosphatase activity(GO:0047429) |
| 0.1 | 1.0 | GO:0019200 | carbohydrate kinase activity(GO:0019200) |
| 0.1 | 3.2 | GO:0008373 | sialyltransferase activity(GO:0008373) |
| 0.1 | 0.9 | GO:0051879 | Hsp90 protein binding(GO:0051879) |
| 0.1 | 0.4 | GO:0015616 | DNA translocase activity(GO:0015616) |
| 0.1 | 0.5 | GO:0004144 | diacylglycerol O-acyltransferase activity(GO:0004144) |
| 0.1 | 2.1 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.1 | 13.9 | GO:0042802 | identical protein binding(GO:0042802) |
| 0.1 | 0.6 | GO:0003834 | beta-carotene 15,15'-monooxygenase activity(GO:0003834) carotenoid dioxygenase activity(GO:0010436) |
| 0.1 | 3.7 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.1 | 0.2 | GO:0017020 | myosin phosphatase regulator activity(GO:0017020) |
| 0.1 | 1.0 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.1 | 0.7 | GO:1990404 | protein ADP-ribosylase activity(GO:1990404) |
| 0.1 | 2.1 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.1 | 2.0 | GO:0008175 | tRNA methyltransferase activity(GO:0008175) |
| 0.1 | 8.5 | GO:0060090 | binding, bridging(GO:0060090) |
| 0.1 | 0.2 | GO:0030273 | melanin-concentrating hormone receptor activity(GO:0030273) |
| 0.1 | 1.6 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.1 | 0.2 | GO:0050135 | NAD(P)+ nucleosidase activity(GO:0050135) |
| 0.1 | 3.5 | GO:0016811 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amides(GO:0016811) |
| 0.1 | 2.9 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.1 | 11.1 | GO:0050839 | cell adhesion molecule binding(GO:0050839) |
| 0.1 | 0.7 | GO:0003886 | DNA (cytosine-5-)-methyltransferase activity(GO:0003886) |
| 0.1 | 1.9 | GO:0004629 | phosphatidylinositol phospholipase C activity(GO:0004435) phospholipase C activity(GO:0004629) |
| 0.1 | 1.6 | GO:0042287 | MHC protein binding(GO:0042287) |
| 0.1 | 0.9 | GO:0030983 | mismatched DNA binding(GO:0030983) |
| 0.1 | 0.6 | GO:0061575 | cyclin-dependent protein serine/threonine kinase activator activity(GO:0061575) |
| 0.1 | 16.6 | GO:0005085 | guanyl-nucleotide exchange factor activity(GO:0005085) |
| 0.1 | 0.9 | GO:0000175 | 3'-5'-exoribonuclease activity(GO:0000175) |
| 0.0 | 0.2 | GO:0004032 | alditol:NADP+ 1-oxidoreductase activity(GO:0004032) |
| 0.0 | 1.2 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.0 | 1.4 | GO:0030145 | manganese ion binding(GO:0030145) |
| 0.0 | 3.6 | GO:0019904 | protein domain specific binding(GO:0019904) |
| 0.0 | 0.4 | GO:0030507 | spectrin binding(GO:0030507) |
| 0.0 | 1.0 | GO:0008474 | palmitoyl-(protein) hydrolase activity(GO:0008474) palmitoyl hydrolase activity(GO:0098599) |
| 0.0 | 0.5 | GO:0070001 | aspartic-type endopeptidase activity(GO:0004190) aspartic-type peptidase activity(GO:0070001) |
| 0.0 | 1.5 | GO:0070006 | metalloaminopeptidase activity(GO:0070006) |
| 0.0 | 0.2 | GO:0008486 | endopolyphosphatase activity(GO:0000298) diphosphoinositol-polyphosphate diphosphatase activity(GO:0008486) bis(5'-adenosyl)-hexaphosphatase activity(GO:0034431) bis(5'-adenosyl)-pentaphosphatase activity(GO:0034432) |
| 0.0 | 6.2 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.0 | 0.7 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.0 | 1.3 | GO:0005227 | calcium activated cation channel activity(GO:0005227) |
| 0.0 | 1.0 | GO:0016875 | aminoacyl-tRNA ligase activity(GO:0004812) ligase activity, forming carbon-oxygen bonds(GO:0016875) ligase activity, forming aminoacyl-tRNA and related compounds(GO:0016876) |
| 0.0 | 1.9 | GO:0016765 | transferase activity, transferring alkyl or aryl (other than methyl) groups(GO:0016765) |
| 0.0 | 0.5 | GO:0002020 | protease binding(GO:0002020) |
| 0.0 | 0.9 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.0 | 6.3 | GO:0000149 | SNARE binding(GO:0000149) |
| 0.0 | 0.2 | GO:1903924 | estradiol binding(GO:1903924) |
| 0.0 | 0.3 | GO:0017134 | fibroblast growth factor binding(GO:0017134) |
| 0.0 | 0.6 | GO:0004724 | magnesium-dependent protein serine/threonine phosphatase activity(GO:0004724) |
| 0.0 | 6.1 | GO:0019901 | protein kinase binding(GO:0019901) |
| 0.0 | 0.9 | GO:0042826 | histone deacetylase binding(GO:0042826) |
| 0.0 | 3.2 | GO:0008134 | transcription factor binding(GO:0008134) |
| 0.0 | 0.5 | GO:0031726 | CCR1 chemokine receptor binding(GO:0031726) CCR4 chemokine receptor binding(GO:0031729) |
| 0.0 | 0.6 | GO:0045504 | dynein heavy chain binding(GO:0045504) |
| 0.0 | 0.5 | GO:0070566 | adenylyltransferase activity(GO:0070566) |
| 0.0 | 4.5 | GO:0008083 | growth factor activity(GO:0008083) |
| 0.0 | 3.6 | GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds(GO:0004553) |
| 0.0 | 0.4 | GO:0031386 | protein tag(GO:0031386) |
| 0.0 | 1.7 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.0 | 0.1 | GO:0055105 | ubiquitin-protein transferase inhibitor activity(GO:0055105) |
| 0.0 | 0.1 | GO:0043024 | ribosomal small subunit binding(GO:0043024) |
| 0.0 | 0.3 | GO:0016864 | protein disulfide isomerase activity(GO:0003756) intramolecular oxidoreductase activity, transposing S-S bonds(GO:0016864) |
| 0.0 | 1.2 | GO:0005544 | calcium-dependent phospholipid binding(GO:0005544) |
| 0.0 | 1.3 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.0 | 1.5 | GO:1901981 | phosphatidylinositol phosphate binding(GO:1901981) |
| 0.0 | 0.8 | GO:0019003 | GDP binding(GO:0019003) |
| 0.0 | 1.8 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.0 | 0.2 | GO:0050694 | galactosylceramide sulfotransferase activity(GO:0001733) galactose 3-O-sulfotransferase activity(GO:0050694) |
| 0.0 | 1.2 | GO:0015296 | anion:cation symporter activity(GO:0015296) |
| 0.0 | 1.7 | GO:0008375 | acetylglucosaminyltransferase activity(GO:0008375) |
| 0.0 | 0.6 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
| 0.0 | 0.3 | GO:0016884 | carbon-nitrogen ligase activity, with glutamine as amido-N-donor(GO:0016884) |
| 0.0 | 0.1 | GO:0016308 | 1-phosphatidylinositol-4-phosphate 5-kinase activity(GO:0016308) |
| 0.0 | 0.1 | GO:0004949 | cannabinoid receptor activity(GO:0004949) |
| 0.0 | 1.0 | GO:0005347 | ATP transmembrane transporter activity(GO:0005347) |
| 0.0 | 0.4 | GO:0004709 | MAP kinase kinase kinase activity(GO:0004709) |
| 0.0 | 3.4 | GO:0004725 | protein tyrosine phosphatase activity(GO:0004725) |
| 0.0 | 0.5 | GO:0016289 | CoA hydrolase activity(GO:0016289) |
| 0.0 | 5.0 | GO:0044822 | mRNA binding(GO:0003729) poly(A) RNA binding(GO:0044822) |
| 0.0 | 0.1 | GO:0004126 | cytidine deaminase activity(GO:0004126) |
| 0.0 | 1.5 | GO:0035091 | phosphatidylinositol binding(GO:0035091) |
| 0.0 | 0.2 | GO:0005042 | netrin receptor activity(GO:0005042) |
| 0.0 | 1.1 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.0 | 0.0 | GO:0008297 | single-stranded DNA exodeoxyribonuclease activity(GO:0008297) |
| 0.0 | 0.3 | GO:0015279 | store-operated calcium channel activity(GO:0015279) |
| 0.0 | 0.2 | GO:0016907 | G-protein coupled acetylcholine receptor activity(GO:0016907) |
| 0.0 | 0.3 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.0 | 0.3 | GO:0008253 | 5'-nucleotidase activity(GO:0008253) |
| 0.0 | 0.5 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.0 | 5.7 | GO:0003779 | actin binding(GO:0003779) |
| 0.0 | 0.4 | GO:0005548 | phospholipid transporter activity(GO:0005548) |
| 0.0 | 0.1 | GO:0030332 | cyclin binding(GO:0030332) |
| 0.0 | 0.0 | GO:0070643 | vitamin D 25-hydroxylase activity(GO:0070643) |
| 0.0 | 3.1 | GO:0005096 | GTPase activator activity(GO:0005096) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 8.8 | PID BETA CATENIN DEG PATHWAY | Degradation of beta catenin |
| 0.3 | 4.8 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.3 | 1.1 | ST ERK1 ERK2 MAPK PATHWAY | ERK1/ERK2 MAPK Pathway |
| 0.3 | 4.7 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.3 | 9.3 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.2 | 0.7 | SA G2 AND M PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
| 0.2 | 1.9 | PID AR NONGENOMIC PATHWAY | Nongenotropic Androgen signaling |
| 0.2 | 12.2 | PID HDAC CLASSI PATHWAY | Signaling events mediated by HDAC Class I |
| 0.2 | 3.8 | PID ECADHERIN KERATINOCYTE PATHWAY | E-cadherin signaling in keratinocytes |
| 0.1 | 4.0 | PID ERBB2 ERBB3 PATHWAY | ErbB2/ErbB3 signaling events |
| 0.1 | 4.5 | PID CDC42 PATHWAY | CDC42 signaling events |
| 0.1 | 3.3 | PID ERBB1 INTERNALIZATION PATHWAY | Internalization of ErbB1 |
| 0.1 | 0.8 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
| 0.1 | 2.2 | PID INTEGRIN A4B1 PATHWAY | Alpha4 beta1 integrin signaling events |
| 0.1 | 5.3 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 0.1 | 3.6 | SIG INSULIN RECEPTOR PATHWAY IN CARDIAC MYOCYTES | Genes related to the insulin receptor pathway |
| 0.1 | 2.6 | PID IL1 PATHWAY | IL1-mediated signaling events |
| 0.1 | 4.4 | PID RAC1 REG PATHWAY | Regulation of RAC1 activity |
| 0.1 | 1.1 | PID CERAMIDE PATHWAY | Ceramide signaling pathway |
| 0.1 | 5.7 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 0.1 | 2.3 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.1 | 2.4 | PID FOXO PATHWAY | FoxO family signaling |
| 0.1 | 1.8 | PID AURORA A PATHWAY | Aurora A signaling |
| 0.1 | 2.6 | PID HEDGEHOG GLI PATHWAY | Hedgehog signaling events mediated by Gli proteins |
| 0.1 | 2.0 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.1 | 5.8 | PID AR PATHWAY | Coregulation of Androgen receptor activity |
| 0.1 | 2.5 | PID TNF PATHWAY | TNF receptor signaling pathway |
| 0.1 | 2.0 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.1 | 1.6 | PID IL2 1PATHWAY | IL2-mediated signaling events |
| 0.1 | 1.7 | PID BMP PATHWAY | BMP receptor signaling |
| 0.1 | 2.6 | PID TGFBR PATHWAY | TGF-beta receptor signaling |
| 0.1 | 1.2 | PID FCER1 PATHWAY | Fc-epsilon receptor I signaling in mast cells |
| 0.1 | 1.1 | PID FAK PATHWAY | Signaling events mediated by focal adhesion kinase |
| 0.1 | 1.9 | PID LYSOPHOSPHOLIPID PATHWAY | LPA receptor mediated events |
| 0.1 | 1.1 | PID VEGFR1 2 PATHWAY | Signaling events mediated by VEGFR1 and VEGFR2 |
| 0.1 | 0.3 | PID NEPHRIN NEPH1 PATHWAY | Nephrin/Neph1 signaling in the kidney podocyte |
| 0.1 | 1.8 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.1 | 0.4 | ST IL 13 PATHWAY | Interleukin 13 (IL-13) Pathway |
| 0.0 | 1.1 | ST JNK MAPK PATHWAY | JNK MAPK Pathway |
| 0.0 | 0.6 | SIG BCR SIGNALING PATHWAY | Members of the BCR signaling pathway |
| 0.0 | 0.5 | PID ANGIOPOIETIN RECEPTOR PATHWAY | Angiopoietin receptor Tie2-mediated signaling |
| 0.0 | 0.6 | PID HIF2PATHWAY | HIF-2-alpha transcription factor network |
| 0.0 | 0.4 | PID PI3KCI PATHWAY | Class I PI3K signaling events |
| 0.0 | 0.3 | ST TUMOR NECROSIS FACTOR PATHWAY | Tumor Necrosis Factor Pathway. |
| 0.0 | 0.2 | PID EPO PATHWAY | EPO signaling pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.8 | 10.1 | REACTOME GLYCOGEN BREAKDOWN GLYCOGENOLYSIS | Genes involved in Glycogen breakdown (glycogenolysis) |
| 0.5 | 4.8 | REACTOME SIGNALING BY NODAL | Genes involved in Signaling by NODAL |
| 0.4 | 6.3 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.4 | 2.2 | REACTOME PURINE CATABOLISM | Genes involved in Purine catabolism |
| 0.4 | 4.7 | REACTOME SLBP DEPENDENT PROCESSING OF REPLICATION DEPENDENT HISTONE PRE MRNAS | Genes involved in SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs |
| 0.4 | 4.2 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION IN TLR7 8 OR 9 SIGNALING | Genes involved in TRAF6 mediated IRF7 activation in TLR7/8 or 9 signaling |
| 0.3 | 2.1 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |
| 0.3 | 4.6 | REACTOME CTNNB1 PHOSPHORYLATION CASCADE | Genes involved in Beta-catenin phosphorylation cascade |
| 0.3 | 4.6 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |
| 0.3 | 5.0 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.3 | 6.3 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.3 | 3.8 | REACTOME MITOCHONDRIAL FATTY ACID BETA OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.3 | 4.8 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.2 | 2.4 | REACTOME NOTCH HLH TRANSCRIPTION PATHWAY | Genes involved in Notch-HLH transcription pathway |
| 0.2 | 3.4 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.2 | 4.1 | REACTOME SYNTHESIS OF GLYCOSYLPHOSPHATIDYLINOSITOL GPI | Genes involved in Synthesis of glycosylphosphatidylinositol (GPI) |
| 0.2 | 2.4 | REACTOME CYCLIN E ASSOCIATED EVENTS DURING G1 S TRANSITION | Genes involved in Cyclin E associated events during G1/S transition |
| 0.2 | 4.3 | REACTOME PROCESSING OF CAPPED INTRONLESS PRE MRNA | Genes involved in Processing of Capped Intronless Pre-mRNA |
| 0.2 | 1.9 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.2 | 1.5 | REACTOME TRAFFICKING AND PROCESSING OF ENDOSOMAL TLR | Genes involved in Trafficking and processing of endosomal TLR |
| 0.2 | 5.4 | REACTOME EGFR DOWNREGULATION | Genes involved in EGFR downregulation |
| 0.2 | 2.6 | REACTOME VIRAL MESSENGER RNA SYNTHESIS | Genes involved in Viral Messenger RNA Synthesis |
| 0.2 | 2.2 | REACTOME ELONGATION ARREST AND RECOVERY | Genes involved in Elongation arrest and recovery |
| 0.2 | 1.6 | REACTOME MEMBRANE BINDING AND TARGETTING OF GAG PROTEINS | Genes involved in Membrane binding and targetting of GAG proteins |
| 0.2 | 8.0 | REACTOME LOSS OF NLP FROM MITOTIC CENTROSOMES | Genes involved in Loss of Nlp from mitotic centrosomes |
| 0.2 | 2.6 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.2 | 1.4 | REACTOME COPI MEDIATED TRANSPORT | Genes involved in COPI Mediated Transport |
| 0.2 | 8.0 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.1 | 1.9 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |
| 0.1 | 2.0 | REACTOME ASSOCIATION OF LICENSING FACTORS WITH THE PRE REPLICATIVE COMPLEX | Genes involved in Association of licensing factors with the pre-replicative complex |
| 0.1 | 5.5 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.1 | 0.6 | REACTOME PD1 SIGNALING | Genes involved in PD-1 signaling |
| 0.1 | 1.4 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 0.1 | 1.5 | REACTOME SEMA4D INDUCED CELL MIGRATION AND GROWTH CONE COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.1 | 2.5 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.1 | 2.4 | REACTOME ACTIVATED NOTCH1 TRANSMITS SIGNAL TO THE NUCLEUS | Genes involved in Activated NOTCH1 Transmits Signal to the Nucleus |
| 0.1 | 2.5 | REACTOME ACTIVATION OF THE PRE REPLICATIVE COMPLEX | Genes involved in Activation of the pre-replicative complex |
| 0.1 | 1.0 | REACTOME ROLE OF DCC IN REGULATING APOPTOSIS | Genes involved in Role of DCC in regulating apoptosis |
| 0.1 | 1.7 | REACTOME FANCONI ANEMIA PATHWAY | Genes involved in Fanconi Anemia pathway |
| 0.1 | 0.6 | REACTOME RNA POL I PROMOTER OPENING | Genes involved in RNA Polymerase I Promoter Opening |
| 0.1 | 2.8 | REACTOME MEIOTIC RECOMBINATION | Genes involved in Meiotic Recombination |
| 0.1 | 0.7 | REACTOME KERATAN SULFATE BIOSYNTHESIS | Genes involved in Keratan sulfate biosynthesis |
| 0.1 | 0.9 | REACTOME AMINE DERIVED HORMONES | Genes involved in Amine-derived hormones |
| 0.1 | 0.3 | REACTOME SYNTHESIS OF PA | Genes involved in Synthesis of PA |
| 0.1 | 2.5 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.1 | 2.9 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |
| 0.1 | 1.2 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.1 | 0.9 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.1 | 0.6 | REACTOME ALPHA LINOLENIC ACID ALA METABOLISM | Genes involved in alpha-linolenic acid (ALA) metabolism |
| 0.1 | 2.0 | REACTOME NOTCH1 INTRACELLULAR DOMAIN REGULATES TRANSCRIPTION | Genes involved in NOTCH1 Intracellular Domain Regulates Transcription |
| 0.1 | 0.9 | REACTOME GABA SYNTHESIS RELEASE REUPTAKE AND DEGRADATION | Genes involved in GABA synthesis, release, reuptake and degradation |
| 0.1 | 1.2 | REACTOME JNK C JUN KINASES PHOSPHORYLATION AND ACTIVATION MEDIATED BY ACTIVATED HUMAN TAK1 | Genes involved in JNK (c-Jun kinases) phosphorylation and activation mediated by activated human TAK1 |
| 0.1 | 2.0 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |
| 0.1 | 0.4 | REACTOME REGULATION OF MRNA STABILITY BY PROTEINS THAT BIND AU RICH ELEMENTS | Genes involved in Regulation of mRNA Stability by Proteins that Bind AU-rich Elements |
| 0.0 | 0.9 | REACTOME HS GAG DEGRADATION | Genes involved in HS-GAG degradation |
| 0.0 | 0.3 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.0 | 0.6 | REACTOME PRE NOTCH EXPRESSION AND PROCESSING | Genes involved in Pre-NOTCH Expression and Processing |
| 0.0 | 0.3 | REACTOME ACYL CHAIN REMODELLING OF PG | Genes involved in Acyl chain remodelling of PG |
| 0.0 | 0.2 | REACTOME REGULATION OF KIT SIGNALING | Genes involved in Regulation of KIT signaling |
| 0.0 | 1.0 | REACTOME DOWNSTREAM TCR SIGNALING | Genes involved in Downstream TCR signaling |
| 0.0 | 1.0 | REACTOME CYTOSOLIC TRNA AMINOACYLATION | Genes involved in Cytosolic tRNA aminoacylation |
| 0.0 | 0.5 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 0.0 | 0.4 | REACTOME PLATELET CALCIUM HOMEOSTASIS | Genes involved in Platelet calcium homeostasis |
| 0.0 | 2.4 | REACTOME MRNA SPLICING | Genes involved in mRNA Splicing |
| 0.0 | 0.8 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.0 | 0.4 | REACTOME LYSOSOME VESICLE BIOGENESIS | Genes involved in Lysosome Vesicle Biogenesis |
| 0.0 | 0.2 | REACTOME REGULATION OF SIGNALING BY CBL | Genes involved in Regulation of signaling by CBL |
| 0.0 | 0.5 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.0 | 0.3 | REACTOME ANTIGEN ACTIVATES B CELL RECEPTOR LEADING TO GENERATION OF SECOND MESSENGERS | Genes involved in Antigen Activates B Cell Receptor Leading to Generation of Second Messengers |
| 0.0 | 1.0 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.0 | 0.9 | REACTOME SIGNALING BY SCF KIT | Genes involved in Signaling by SCF-KIT |
| 0.0 | 1.6 | REACTOME PPARA ACTIVATES GENE EXPRESSION | Genes involved in PPARA Activates Gene Expression |
| 0.0 | 1.1 | REACTOME G ALPHA1213 SIGNALLING EVENTS | Genes involved in G alpha (12/13) signalling events |
| 0.0 | 0.5 | REACTOME UNFOLDED PROTEIN RESPONSE | Genes involved in Unfolded Protein Response |
| 0.0 | 0.2 | REACTOME BILE SALT AND ORGANIC ANION SLC TRANSPORTERS | Genes involved in Bile salt and organic anion SLC transporters |
| 0.0 | 0.2 | REACTOME IL1 SIGNALING | Genes involved in Interleukin-1 signaling |
| 0.0 | 0.8 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |