Project
PRJNA438478: RNAseq of wild type zebrafish germline
Navigation
Downloads
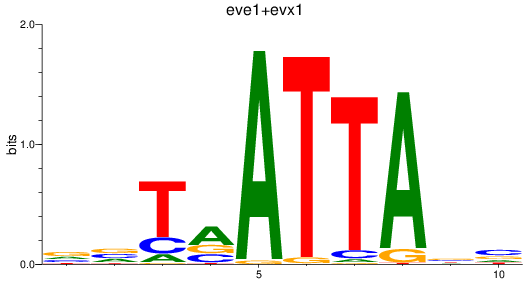
Results for eve1+evx1
Z-value: 0.24
Transcription factors associated with eve1+evx1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
eve1
|
ENSDARG00000056012 | even-skipped-like1 |
|
evx1
|
ENSDARG00000099365 | even-skipped homeobox 1 |
|
evx1
|
ENSDARG00000100087 | even-skipped homeobox 1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| evx1 | dr11_v1_chr19_-_19871211_19871211 | -0.64 | 4.5e-03 | Click! |
| eve1 | dr11_v1_chr3_-_23643751_23643830 | 0.63 | 4.7e-03 | Click! |
Activity profile of eve1+evx1 motif
Sorted Z-values of eve1+evx1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr8_+_2438638 | 0.43 |
ENSDART00000141263
|
ENKD1
|
si:ch211-220d9.3 |
| chr7_-_1504382 | 0.40 |
ENSDART00000172770
|
si:zfos-405g10.4
|
si:zfos-405g10.4 |
| chr8_-_21372446 | 0.37 |
ENSDART00000061481
ENSDART00000079293 |
ela2l
|
elastase 2 like |
| chr6_-_40352215 | 0.34 |
ENSDART00000103992
|
ttll3
|
tubulin tyrosine ligase-like family, member 3 |
| chr17_+_21295132 | 0.32 |
ENSDART00000103845
|
eno4
|
enolase family member 4 |
| chr10_+_31222433 | 0.30 |
ENSDART00000185080
|
tmem218
|
transmembrane protein 218 |
| chr15_+_36309070 | 0.29 |
ENSDART00000157034
|
gmnc
|
geminin coiled-coil domain containing |
| chr21_+_43404945 | 0.26 |
ENSDART00000142234
|
frmd7
|
FERM domain containing 7 |
| chr1_-_46981134 | 0.26 |
ENSDART00000130607
|
pknox1.2
|
pbx/knotted 1 homeobox 1.2 |
| chr21_+_30355767 | 0.24 |
ENSDART00000189948
|
CR749164.1
|
|
| chr1_-_17715493 | 0.24 |
ENSDART00000133027
|
si:dkey-256e7.8
|
si:dkey-256e7.8 |
| chr13_+_33117528 | 0.23 |
ENSDART00000085719
|
si:ch211-10a23.2
|
si:ch211-10a23.2 |
| chr16_-_54455573 | 0.23 |
ENSDART00000075275
|
pklr
|
pyruvate kinase L/R |
| chr17_-_25649079 | 0.23 |
ENSDART00000130955
|
ppp1cb
|
protein phosphatase 1, catalytic subunit, beta isozyme |
| chr23_-_33361425 | 0.21 |
ENSDART00000031638
|
slc48a1a
|
solute carrier family 48 (heme transporter), member 1a |
| chr17_-_31695217 | 0.21 |
ENSDART00000104332
ENSDART00000143090 |
lin52
|
lin-52 DREAM MuvB core complex component |
| chr25_+_3327071 | 0.20 |
ENSDART00000136131
ENSDART00000133243 |
ldhbb
|
lactate dehydrogenase Bb |
| chr7_-_24046999 | 0.19 |
ENSDART00000144616
ENSDART00000124653 ENSDART00000127813 |
dhrs4
|
dehydrogenase/reductase (SDR family) member 4 |
| chr11_-_44876005 | 0.19 |
ENSDART00000192006
|
opn6a
|
opsin 6, group member a |
| chrM_+_12897 | 0.18 |
ENSDART00000093622
|
mt-nd5
|
NADH dehydrogenase 5, mitochondrial |
| chr4_+_11384891 | 0.18 |
ENSDART00000092381
ENSDART00000186577 ENSDART00000191054 ENSDART00000191584 |
pcloa
|
piccolo presynaptic cytomatrix protein a |
| chr9_+_13120419 | 0.18 |
ENSDART00000141005
|
fam117bb
|
family with sequence similarity 117, member Bb |
| chr9_-_53666031 | 0.18 |
ENSDART00000126314
|
pcdh8
|
protocadherin 8 |
| chr13_+_27232848 | 0.17 |
ENSDART00000138043
|
rin2
|
Ras and Rab interactor 2 |
| chr24_+_1023839 | 0.17 |
ENSDART00000082526
|
zgc:111976
|
zgc:111976 |
| chr6_+_515181 | 0.17 |
ENSDART00000171374
|
CACNA1I (1 of many)
|
si:ch73-379f7.5 |
| chr19_+_712127 | 0.17 |
ENSDART00000093281
ENSDART00000180002 ENSDART00000146050 |
fhod3a
|
formin homology 2 domain containing 3a |
| chr21_-_26490186 | 0.17 |
ENSDART00000009889
|
zgc:110540
|
zgc:110540 |
| chr10_+_35257651 | 0.16 |
ENSDART00000028940
|
styxl1
|
serine/threonine/tyrosine interacting-like 1 |
| chr22_-_36996856 | 0.16 |
ENSDART00000165923
|
ATP11B
|
si:dkey-211e20.10 |
| chr5_+_38263240 | 0.16 |
ENSDART00000051231
|
gnb2
|
guanine nucleotide binding protein (G protein), beta polypeptide 2 |
| chr1_-_47071979 | 0.16 |
ENSDART00000160817
|
itsn1
|
intersectin 1 (SH3 domain protein) |
| chr10_-_41664427 | 0.15 |
ENSDART00000150213
|
ggt1b
|
gamma-glutamyltransferase 1b |
| chr21_+_26726936 | 0.15 |
ENSDART00000065392
|
calm2a
|
calmodulin 2a (phosphorylase kinase, delta) |
| chr4_+_306036 | 0.14 |
ENSDART00000103659
|
msgn1
|
mesogenin 1 |
| chr16_+_11029762 | 0.14 |
ENSDART00000091183
|
erfl3
|
Ets2 repressor factor like 3 |
| chr12_-_35830625 | 0.14 |
ENSDART00000180028
|
CU459056.1
|
|
| chr20_-_46362606 | 0.14 |
ENSDART00000153087
|
bmf2
|
BCL2 modifying factor 2 |
| chr3_+_18398876 | 0.14 |
ENSDART00000141100
ENSDART00000138107 |
rps2
|
ribosomal protein S2 |
| chr15_-_34785594 | 0.13 |
ENSDART00000154256
|
gabbr1a
|
gamma-aminobutyric acid (GABA) B receptor, 1a |
| chr5_+_23242370 | 0.13 |
ENSDART00000051532
|
agtr2
|
angiotensin II receptor, type 2 |
| chr6_+_41191482 | 0.13 |
ENSDART00000000877
|
opn1mw3
|
opsin 1 (cone pigments), medium-wave-sensitive, 3 |
| chr19_+_9033376 | 0.13 |
ENSDART00000192298
ENSDART00000052915 |
ash1l
|
ash1 (absent, small, or homeotic)-like (Drosophila) |
| chr13_+_30903816 | 0.13 |
ENSDART00000191727
|
ercc6
|
excision repair cross-complementation group 6 |
| chr12_-_19007834 | 0.12 |
ENSDART00000153248
|
chadlb
|
chondroadherin-like b |
| chr7_+_17816006 | 0.12 |
ENSDART00000080834
|
eml3
|
echinoderm microtubule associated protein like 3 |
| chr10_-_2971407 | 0.12 |
ENSDART00000132526
|
marveld2a
|
MARVEL domain containing 2a |
| chr2_-_11258547 | 0.12 |
ENSDART00000165803
ENSDART00000193817 |
slc44a5a
|
solute carrier family 44, member 5a |
| chr7_-_25895189 | 0.12 |
ENSDART00000173599
ENSDART00000079235 ENSDART00000173786 ENSDART00000173602 ENSDART00000079245 ENSDART00000187568 ENSDART00000173505 |
cd99l2
|
CD99 molecule-like 2 |
| chr15_-_23793641 | 0.12 |
ENSDART00000122891
|
tmem97
|
transmembrane protein 97 |
| chr21_-_5205617 | 0.11 |
ENSDART00000145554
ENSDART00000045284 |
rpl37
|
ribosomal protein L37 |
| chr9_-_202805 | 0.11 |
ENSDART00000182260
|
CABZ01067557.1
|
|
| chr23_+_40460333 | 0.10 |
ENSDART00000184658
|
soga3b
|
SOGA family member 3b |
| chr22_+_20427170 | 0.10 |
ENSDART00000136744
|
foxq2
|
forkhead box Q2 |
| chr19_+_40856534 | 0.10 |
ENSDART00000051950
|
gngt1
|
guanine nucleotide binding protein (G protein), gamma transducing activity polypeptide 1 |
| chr4_-_67969695 | 0.10 |
ENSDART00000190016
|
si:ch211-223k15.1
|
si:ch211-223k15.1 |
| chr22_-_881080 | 0.10 |
ENSDART00000185489
|
cept1b
|
choline/ethanolamine phosphotransferase 1b |
| chr2_-_17235891 | 0.10 |
ENSDART00000144251
|
artnb
|
artemin b |
| chr3_+_13624815 | 0.10 |
ENSDART00000161451
|
pglyrp6
|
peptidoglycan recognition protein 6 |
| chr25_+_35553542 | 0.10 |
ENSDART00000113723
|
spi1a
|
Spi-1 proto-oncogene a |
| chr22_-_10121880 | 0.10 |
ENSDART00000002348
|
rdh5
|
retinol dehydrogenase 5 (11-cis/9-cis) |
| chr2_-_23172708 | 0.09 |
ENSDART00000041365
|
prrx1a
|
paired related homeobox 1a |
| chr10_+_41765944 | 0.09 |
ENSDART00000171484
|
rnf34b
|
ring finger protein 34b |
| chr7_+_17816470 | 0.09 |
ENSDART00000173807
|
eml3
|
echinoderm microtubule associated protein like 3 |
| chr7_-_40145097 | 0.09 |
ENSDART00000173634
|
wdr60
|
WD repeat domain 60 |
| chr4_+_9177997 | 0.09 |
ENSDART00000057254
ENSDART00000154614 |
nfyba
|
nuclear transcription factor Y, beta a |
| chr17_-_23446760 | 0.09 |
ENSDART00000104715
|
pcgf5a
|
polycomb group ring finger 5a |
| chr16_+_21330634 | 0.09 |
ENSDART00000191285
ENSDART00000183267 |
osbpl3b
|
oxysterol binding protein-like 3b |
| chr1_+_17676745 | 0.09 |
ENSDART00000030665
|
slc25a4
|
solute carrier family 25 (mitochondrial carrier; adenine nucleotide translocator), member 4 |
| chr3_-_34027178 | 0.09 |
ENSDART00000170201
ENSDART00000151408 |
ighv1-4
ighv14-1
|
immunoglobulin heavy variable 1-4 immunoglobulin heavy variable 14-1 |
| chr18_-_43884044 | 0.08 |
ENSDART00000087382
|
ddx6
|
DEAD (Asp-Glu-Ala-Asp) box helicase 6 |
| chr10_+_36184751 | 0.08 |
ENSDART00000162639
|
or108-2
|
odorant receptor, family D, subfamily 108, member 2 |
| chr9_-_6372535 | 0.08 |
ENSDART00000149189
|
ecrg4a
|
esophageal cancer related gene 4a |
| chr1_+_33647682 | 0.08 |
ENSDART00000114384
|
znf654
|
zinc finger protein 654 |
| chr12_-_24060358 | 0.08 |
ENSDART00000050831
|
chac2
|
ChaC, cation transport regulator homolog 2 (E. coli) |
| chr16_-_36093776 | 0.08 |
ENSDART00000159134
|
OSCP1
|
organic solute carrier partner 1 |
| chr2_+_56213694 | 0.08 |
ENSDART00000162582
|
upf1
|
upf1 regulator of nonsense transcripts homolog (yeast) |
| chr10_-_44482911 | 0.07 |
ENSDART00000085556
|
hip1ra
|
huntingtin interacting protein 1 related a |
| chr15_-_22074315 | 0.07 |
ENSDART00000149830
|
drd2a
|
dopamine receptor D2a |
| chr10_+_3049636 | 0.07 |
ENSDART00000081794
ENSDART00000183167 ENSDART00000191634 ENSDART00000183514 |
rasgrf2a
|
Ras protein-specific guanine nucleotide-releasing factor 2a |
| chr1_-_10647484 | 0.07 |
ENSDART00000164541
ENSDART00000188958 ENSDART00000190904 |
si:dkey-31e10.1
|
si:dkey-31e10.1 |
| chr11_+_28476298 | 0.07 |
ENSDART00000122319
|
lrrc38b
|
leucine rich repeat containing 38b |
| chr16_-_2414063 | 0.07 |
ENSDART00000073621
|
zgc:152945
|
zgc:152945 |
| chr3_+_39663987 | 0.07 |
ENSDART00000184614
ENSDART00000184573 ENSDART00000183127 |
epn2
|
epsin 2 |
| chr18_+_11506561 | 0.07 |
ENSDART00000121647
|
PRMT8
|
protein arginine methyltransferase 8 |
| chr8_+_53423408 | 0.07 |
ENSDART00000164792
|
cacna1db
|
calcium channel, voltage-dependent, L type, alpha 1D subunit, b |
| chr11_-_29768054 | 0.06 |
ENSDART00000079117
|
plekha3
|
pleckstrin homology domain containing, family A (phosphoinositide binding specific) member 3 |
| chr17_-_28198099 | 0.06 |
ENSDART00000156143
|
htr1d
|
5-hydroxytryptamine (serotonin) receptor 1D, G protein-coupled |
| chr15_+_5360407 | 0.06 |
ENSDART00000110420
|
or112-1
|
odorant receptor, family A, subfamily 112, member 1 |
| chr2_-_10896745 | 0.06 |
ENSDART00000114609
|
cdcp2
|
CUB domain containing protein 2 |
| chr15_-_1844048 | 0.06 |
ENSDART00000102410
|
taf15
|
TAF15 RNA polymerase II, TATA box binding protein (TBP)-associated factor |
| chr13_+_35637048 | 0.06 |
ENSDART00000085037
|
thbs2a
|
thrombospondin 2a |
| chr8_-_27656765 | 0.06 |
ENSDART00000078491
|
mov10b.2
|
Moloney leukemia virus 10b, tandem duplicate 2 |
| chr10_+_21867307 | 0.06 |
ENSDART00000126629
|
cbln17
|
cerebellin 17 |
| chr10_+_19017146 | 0.06 |
ENSDART00000038674
|
tmem230a
|
transmembrane protein 230a |
| chr8_+_48484455 | 0.06 |
ENSDART00000122737
|
PRDM16
|
si:ch211-263k4.2 |
| chr21_-_18993110 | 0.06 |
ENSDART00000144086
|
si:ch211-222n4.6
|
si:ch211-222n4.6 |
| chr2_+_32826235 | 0.05 |
ENSDART00000143127
|
si:dkey-154p10.3
|
si:dkey-154p10.3 |
| chr16_+_31804590 | 0.05 |
ENSDART00000167321
|
wnt4b
|
wingless-type MMTV integration site family, member 4b |
| chr12_+_20352400 | 0.05 |
ENSDART00000066383
|
hbae5
|
hemoglobin, alpha embryonic 5 |
| chr18_-_1185772 | 0.05 |
ENSDART00000143245
|
nptnb
|
neuroplastin b |
| chr2_+_31492662 | 0.05 |
ENSDART00000123495
|
cacnb2b
|
calcium channel, voltage-dependent, beta 2b |
| chr14_-_34044369 | 0.05 |
ENSDART00000149396
ENSDART00000123607 ENSDART00000190746 |
cyfip2
|
cytoplasmic FMR1 interacting protein 2 |
| chr24_-_1341543 | 0.05 |
ENSDART00000169341
|
nrp1a
|
neuropilin 1a |
| chr6_+_39232245 | 0.05 |
ENSDART00000187351
|
b4galnt1b
|
beta-1,4-N-acetyl-galactosaminyl transferase 1b |
| chr23_+_31913292 | 0.05 |
ENSDART00000136910
|
armc1l
|
armadillo repeat containing 1, like |
| chr24_+_39518774 | 0.05 |
ENSDART00000132939
|
dcun1d3
|
defective in cullin neddylation 1 domain containing 3 |
| chr3_+_27786601 | 0.05 |
ENSDART00000086994
|
nat15
|
N-acetyltransferase 15 (GCN5-related, putative) |
| chr9_-_19161982 | 0.05 |
ENSDART00000081878
|
pou1f1
|
POU class 1 homeobox 1 |
| chr9_-_21918963 | 0.05 |
ENSDART00000090782
|
lmo7a
|
LIM domain 7a |
| chr7_-_19642417 | 0.05 |
ENSDART00000160936
|
si:ch211-212k18.4
|
si:ch211-212k18.4 |
| chr21_-_27185915 | 0.05 |
ENSDART00000135052
|
slc8a4a
|
solute carrier family 8 (sodium/calcium exchanger), member 4a |
| chr19_+_40856807 | 0.05 |
ENSDART00000139083
|
gngt1
|
guanine nucleotide binding protein (G protein), gamma transducing activity polypeptide 1 |
| chr18_+_16986903 | 0.04 |
ENSDART00000142088
|
si:ch211-218c6.8
|
si:ch211-218c6.8 |
| chr20_-_40755614 | 0.04 |
ENSDART00000061247
|
cx32.3
|
connexin 32.3 |
| chr11_-_45152702 | 0.04 |
ENSDART00000168066
|
afmid
|
arylformamidase |
| chr25_-_25384045 | 0.04 |
ENSDART00000150631
|
zgc:123278
|
zgc:123278 |
| chr4_+_12612145 | 0.04 |
ENSDART00000181201
|
lmo3
|
LIM domain only 3 |
| chr1_-_9195629 | 0.04 |
ENSDART00000143587
ENSDART00000192174 |
ern2
|
endoplasmic reticulum to nucleus signaling 2 |
| chr25_+_36405021 | 0.04 |
ENSDART00000152801
|
ankrd27
|
ankyrin repeat domain 27 (VPS9 domain) |
| chr11_+_38280454 | 0.04 |
ENSDART00000171496
|
CDK18
|
si:dkey-166c18.1 |
| chr23_-_19230627 | 0.04 |
ENSDART00000007122
|
guca1b
|
guanylate cyclase activator 1B |
| chr18_+_29402623 | 0.04 |
ENSDART00000014703
|
mafa
|
v-maf avian musculoaponeurotic fibrosarcoma oncogene homolog a (paralog a) |
| chr9_+_38372216 | 0.04 |
ENSDART00000141895
|
plcd4b
|
phospholipase C, delta 4b |
| chr16_-_29458806 | 0.04 |
ENSDART00000047931
|
lingo4b
|
leucine rich repeat and Ig domain containing 4b |
| chr6_+_9870192 | 0.04 |
ENSDART00000150894
|
MPP4 (1 of many)
|
si:ch211-222n4.6 |
| chr14_+_23717165 | 0.04 |
ENSDART00000006373
|
ndfip1
|
Nedd4 family interacting protein 1 |
| chr5_-_42904329 | 0.04 |
ENSDART00000112807
|
cxcl20
|
chemokine (C-X-C motif) ligand 20 |
| chr16_+_28994709 | 0.04 |
ENSDART00000088023
|
gon4l
|
gon-4-like (C. elegans) |
| chr25_+_14507567 | 0.04 |
ENSDART00000015681
|
dbx1b
|
developing brain homeobox 1b |
| chr19_-_44054930 | 0.04 |
ENSDART00000151084
ENSDART00000150991 ENSDART00000005191 |
uqcrb
|
ubiquinol-cytochrome c reductase binding protein |
| chr22_-_881725 | 0.03 |
ENSDART00000035514
|
cept1b
|
choline/ethanolamine phosphotransferase 1b |
| chr8_-_31107537 | 0.03 |
ENSDART00000098925
|
vgll4l
|
vestigial like 4 like |
| chr1_-_10647307 | 0.03 |
ENSDART00000103548
|
si:dkey-31e10.1
|
si:dkey-31e10.1 |
| chr23_-_44219902 | 0.03 |
ENSDART00000185874
|
zgc:158659
|
zgc:158659 |
| chr17_+_24687338 | 0.03 |
ENSDART00000135794
|
selenon
|
selenoprotein N |
| chr6_+_41039166 | 0.03 |
ENSDART00000125659
|
entpd8
|
ectonucleoside triphosphate diphosphohydrolase 8 |
| chr11_+_24703108 | 0.03 |
ENSDART00000159173
|
gpr25
|
G protein-coupled receptor 25 |
| chr7_+_30787903 | 0.03 |
ENSDART00000174000
|
apba2b
|
amyloid beta (A4) precursor protein-binding, family A, member 2b |
| chr8_-_24252933 | 0.03 |
ENSDART00000057624
|
zgc:110353
|
zgc:110353 |
| chr5_+_64732036 | 0.03 |
ENSDART00000073950
|
olfm1a
|
olfactomedin 1a |
| chr10_-_21955231 | 0.03 |
ENSDART00000183695
|
FO744833.2
|
|
| chr6_-_6487876 | 0.03 |
ENSDART00000137642
|
cep170ab
|
centrosomal protein 170Ab |
| chr25_-_37262220 | 0.03 |
ENSDART00000153789
ENSDART00000155182 |
rfwd3
|
ring finger and WD repeat domain 3 |
| chr4_+_52356485 | 0.03 |
ENSDART00000170639
|
zgc:173705
|
zgc:173705 |
| chr6_-_43283122 | 0.03 |
ENSDART00000186022
|
frmd4ba
|
FERM domain containing 4Ba |
| chr4_+_9400012 | 0.02 |
ENSDART00000191960
|
tmtc1
|
transmembrane and tetratricopeptide repeat containing 1 |
| chr23_-_31913231 | 0.02 |
ENSDART00000146852
ENSDART00000085054 |
mtfr2
|
mitochondrial fission regulator 2 |
| chr2_-_26720854 | 0.02 |
ENSDART00000148110
|
si:dkey-181m9.8
|
si:dkey-181m9.8 |
| chr19_-_657439 | 0.02 |
ENSDART00000167100
|
slc6a18
|
solute carrier family 6 (neutral amino acid transporter), member 18 |
| chr5_+_64732270 | 0.02 |
ENSDART00000134241
|
olfm1a
|
olfactomedin 1a |
| chr21_-_3853204 | 0.02 |
ENSDART00000188829
|
st6galnac4
|
ST6 (alpha-N-acetyl-neuraminyl-2,3-beta-galactosyl-1,3)-N-acetylgalactosaminide alpha-2,6-sialyltransferase 4 |
| chr12_-_32426685 | 0.02 |
ENSDART00000105574
|
enpp7.2
|
ectonucleotide pyrophosphatase/phosphodiesterase 7, tandem duplicate 2 |
| chr6_+_23712911 | 0.02 |
ENSDART00000167795
|
zgc:158654
|
zgc:158654 |
| chr15_+_5299404 | 0.02 |
ENSDART00000155410
|
or122-2
|
odorant receptor, family E, subfamily 122, member 2 |
| chr23_+_44374041 | 0.02 |
ENSDART00000136056
|
ephb4b
|
eph receptor B4b |
| chr17_-_23616626 | 0.02 |
ENSDART00000104730
|
ifit14
|
interferon-induced protein with tetratricopeptide repeats 14 |
| chr15_+_7086327 | 0.02 |
ENSDART00000114560
|
pik3cb
|
phosphatidylinositol-4,5-bisphosphate 3-kinase, catalytic subunit beta |
| chr10_+_40606084 | 0.02 |
ENSDART00000133119
|
si:ch211-238p8.24
|
si:ch211-238p8.24 |
| chr19_+_5543072 | 0.02 |
ENSDART00000082080
|
jupb
|
junction plakoglobin b |
| chr2_-_1443692 | 0.02 |
ENSDART00000004533
|
rpe65a
|
retinal pigment epithelium-specific protein 65a |
| chr18_-_12052132 | 0.02 |
ENSDART00000074361
|
zgc:110789
|
zgc:110789 |
| chr16_+_5774977 | 0.02 |
ENSDART00000134202
|
ccka
|
cholecystokinin a |
| chr24_+_36636208 | 0.02 |
ENSDART00000139211
|
si:ch73-334d15.4
|
si:ch73-334d15.4 |
| chr5_-_30615901 | 0.01 |
ENSDART00000147769
|
si:ch211-117m20.5
|
si:ch211-117m20.5 |
| chr6_+_41200500 | 0.01 |
ENSDART00000000678
|
opn1mw4
|
opsin 1 (cone pigments), medium-wave-sensitive, 4 |
| chr18_+_50461981 | 0.01 |
ENSDART00000158761
|
CU896640.1
|
|
| chr18_+_1837668 | 0.01 |
ENSDART00000164210
|
CABZ01079192.1
|
|
| chr14_+_924876 | 0.01 |
ENSDART00000183908
|
myoz3a
|
myozenin 3a |
| chr23_-_18130264 | 0.01 |
ENSDART00000016976
|
nucks1b
|
nuclear casein kinase and cyclin-dependent kinase substrate 1b |
| chr21_+_10773522 | 0.01 |
ENSDART00000143257
|
znf532
|
zinc finger protein 532 |
| chr16_-_21038015 | 0.01 |
ENSDART00000059239
|
snx10b
|
sorting nexin 10b |
| chr8_+_52503340 | 0.01 |
ENSDART00000168838
|
si:ch1073-392o20.1
|
si:ch1073-392o20.1 |
| chr15_-_2497568 | 0.01 |
ENSDART00000080398
|
neu4
|
sialidase 4 |
| chr9_+_41156818 | 0.01 |
ENSDART00000105764
ENSDART00000147052 |
stat4
|
signal transducer and activator of transcription 4 |
| chr14_+_45406299 | 0.01 |
ENSDART00000173142
ENSDART00000112377 |
map1lc3cl
|
microtubule-associated protein 1 light chain 3 gamma, like |
| chr2_+_30465102 | 0.01 |
ENSDART00000188404
|
neto1
|
neuropilin (NRP) and tolloid (TLL)-like 1 |
| chr20_+_37294112 | 0.01 |
ENSDART00000076293
ENSDART00000140450 |
cx23
|
connexin 23 |
| chr5_-_68782641 | 0.01 |
ENSDART00000141699
|
mepce
|
methylphosphate capping enzyme |
| chr23_-_24394719 | 0.01 |
ENSDART00000044918
|
epha2b
|
eph receptor A2 b |
| chr12_-_4408828 | 0.01 |
ENSDART00000152447
|
si:ch211-173d10.1
|
si:ch211-173d10.1 |
| chr2_+_29795833 | 0.01 |
ENSDART00000056750
|
si:ch211-207d6.2
|
si:ch211-207d6.2 |
| chr2_+_21229909 | 0.01 |
ENSDART00000046127
|
vipr1a
|
vasoactive intestinal peptide receptor 1a |
| chr22_-_27709692 | 0.01 |
ENSDART00000172458
|
CR547131.1
|
|
| chr2_+_30464829 | 0.01 |
ENSDART00000181432
|
neto1
|
neuropilin (NRP) and tolloid (TLL)-like 1 |
| chr3_+_33341640 | 0.01 |
ENSDART00000186352
|
pyya
|
peptide YYa |
| chr6_-_40713183 | 0.01 |
ENSDART00000157113
ENSDART00000154810 ENSDART00000153702 |
si:ch211-157b11.12
|
si:ch211-157b11.12 |
| chr18_+_28106139 | 0.01 |
ENSDART00000089615
|
kiaa1549lb
|
KIAA1549-like b |
| chr10_+_40568735 | 0.01 |
ENSDART00000136468
ENSDART00000182841 |
taar18i
|
trace amine associated receptor 18i |
| chr5_+_26765275 | 0.01 |
ENSDART00000144169
|
si:ch211-102c2.8
|
si:ch211-102c2.8 |
| chr22_+_17205608 | 0.01 |
ENSDART00000181267
|
rab3b
|
RAB3B, member RAS oncogene family |
| chr18_+_11970987 | 0.01 |
ENSDART00000144111
|
si:dkeyp-2c8.3
|
si:dkeyp-2c8.3 |
| chr13_-_21660203 | 0.00 |
ENSDART00000100925
|
mxtx1
|
mix-type homeobox gene 1 |
| chr22_-_12746539 | 0.00 |
ENSDART00000175374
|
plcd4a
|
phospholipase C, delta 4a |
| chr11_+_33818179 | 0.00 |
ENSDART00000109418
|
spoplb
|
speckle-type POZ protein-like b |
| chr7_-_17591007 | 0.00 |
ENSDART00000171023
|
CU672228.1
|
|
| chr15_+_22014029 | 0.00 |
ENSDART00000079504
|
ankk1
|
ankyrin repeat and kinase domain containing 1 |
| chr4_+_28355058 | 0.00 |
ENSDART00000150799
|
si:ch73-263o4.3
|
si:ch73-263o4.3 |
| chr21_-_31013817 | 0.00 |
ENSDART00000065504
|
ncbp3
|
nuclear cap binding subunit 3 |
| chr10_+_25726694 | 0.00 |
ENSDART00000140308
|
ugt5d1
|
UDP glucuronosyltransferase 5 family, polypeptide D1 |
Network of associatons between targets according to the STRING database.
First level regulatory network of eve1+evx1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:1903251 | multi-ciliated epithelial cell differentiation(GO:1903251) |
| 0.1 | 0.2 | GO:0097067 | cellular response to thyroid hormone stimulus(GO:0097067) |
| 0.0 | 0.1 | GO:0048341 | paraxial mesoderm morphogenesis(GO:0048340) paraxial mesoderm formation(GO:0048341) |
| 0.0 | 0.2 | GO:0015988 | energy coupled proton transmembrane transport, against electrochemical gradient(GO:0015988) electron transport coupled proton transport(GO:0015990) |
| 0.0 | 0.2 | GO:2001244 | positive regulation of intrinsic apoptotic signaling pathway(GO:2001244) |
| 0.0 | 0.3 | GO:0032889 | regulation of vacuole fusion, non-autophagic(GO:0032889) |
| 0.0 | 0.1 | GO:0016045 | detection of biotic stimulus(GO:0009595) detection of bacterium(GO:0016045) detection of other organism(GO:0098543) detection of external biotic stimulus(GO:0098581) |
| 0.0 | 0.1 | GO:0070314 | G1 to G0 transition(GO:0070314) |
| 0.0 | 0.1 | GO:0015868 | intracellular nucleoside transport(GO:0015859) purine nucleoside transmembrane transport(GO:0015860) purine nucleotide transport(GO:0015865) ATP transport(GO:0015867) purine ribonucleotide transport(GO:0015868) adenine nucleotide transport(GO:0051503) mitochondrial ATP transmembrane transport(GO:1990544) |
| 0.0 | 0.2 | GO:0006751 | glutathione catabolic process(GO:0006751) |
| 0.0 | 0.2 | GO:0015886 | heme transport(GO:0015886) iron coordination entity transport(GO:1901678) |
| 0.0 | 0.1 | GO:0014707 | branchiomeric skeletal muscle development(GO:0014707) |
| 0.0 | 0.1 | GO:0006283 | transcription-coupled nucleotide-excision repair(GO:0006283) |
| 0.0 | 0.2 | GO:1904071 | presynaptic active zone assembly(GO:1904071) |
| 0.0 | 0.1 | GO:0042362 | fat-soluble vitamin biosynthetic process(GO:0042362) |
| 0.0 | 0.1 | GO:2001271 | negative regulation of execution phase of apoptosis(GO:1900118) regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:2001270) negative regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:2001271) |
| 0.0 | 0.1 | GO:0098795 | mRNA cleavage involved in gene silencing by miRNA(GO:0035279) mRNA cleavage involved in gene silencing(GO:0098795) |
| 0.0 | 0.1 | GO:0006646 | phosphatidylethanolamine biosynthetic process(GO:0006646) |
| 0.0 | 0.1 | GO:0086002 | cardiac muscle cell action potential involved in contraction(GO:0086002) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | GO:0000015 | phosphopyruvate hydratase complex(GO:0000015) |
| 0.0 | 0.1 | GO:0061689 | tricellular tight junction(GO:0061689) |
| 0.0 | 0.2 | GO:0098982 | GABA-ergic synapse(GO:0098982) |
| 0.0 | 0.1 | GO:0038037 | G-protein coupled receptor dimeric complex(GO:0038037) G-protein coupled receptor heterodimeric complex(GO:0038039) G-protein coupled receptor complex(GO:0097648) |
| 0.0 | 0.1 | GO:0016586 | RSC complex(GO:0016586) |
| 0.0 | 0.0 | GO:1990604 | IRE1-TRAF2-ASK1 complex(GO:1990604) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0070738 | protein-glycine ligase activity(GO:0070735) tubulin-glycine ligase activity(GO:0070738) |
| 0.0 | 0.3 | GO:0004634 | phosphopyruvate hydratase activity(GO:0004634) |
| 0.0 | 0.2 | GO:0004743 | pyruvate kinase activity(GO:0004743) |
| 0.0 | 0.2 | GO:0042809 | vitamin D receptor binding(GO:0042809) |
| 0.0 | 0.1 | GO:0016019 | peptidoglycan receptor activity(GO:0016019) |
| 0.0 | 0.1 | GO:0004945 | angiotensin type II receptor activity(GO:0004945) |
| 0.0 | 0.2 | GO:0004459 | L-lactate dehydrogenase activity(GO:0004459) |
| 0.0 | 0.1 | GO:0004307 | diacylglycerol cholinephosphotransferase activity(GO:0004142) ethanolaminephosphotransferase activity(GO:0004307) |
| 0.0 | 0.2 | GO:0015232 | heme transporter activity(GO:0015232) |
| 0.0 | 0.2 | GO:0019136 | deoxynucleoside kinase activity(GO:0019136) |
| 0.0 | 0.1 | GO:0005471 | ATP:ADP antiporter activity(GO:0005471) |
| 0.0 | 0.1 | GO:0003839 | gamma-glutamylcyclotransferase activity(GO:0003839) |
| 0.0 | 0.1 | GO:0032574 | 5'-3' RNA helicase activity(GO:0032574) |
| 0.0 | 0.1 | GO:0001217 | bacterial-type RNA polymerase transcription factor activity, sequence-specific DNA binding(GO:0001130) bacterial-type RNA polymerase transcriptional repressor activity, sequence-specific DNA binding(GO:0001217) |
| 0.0 | 0.2 | GO:0000048 | peptidyltransferase activity(GO:0000048) glutathione hydrolase activity(GO:0036374) |
| 0.0 | 0.1 | GO:0004965 | G-protein coupled GABA receptor activity(GO:0004965) |
| 0.0 | 0.0 | GO:0004061 | arylformamidase activity(GO:0004061) |
| 0.0 | 0.1 | GO:1902388 | ceramide transporter activity(GO:0035620) ceramide 1-phosphate binding(GO:1902387) ceramide 1-phosphate transporter activity(GO:1902388) |
| 0.0 | 0.2 | GO:0031682 | G-protein gamma-subunit binding(GO:0031682) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | PID RHODOPSIN PATHWAY | Visual signal transduction: Rods |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | REACTOME OLFACTORY SIGNALING PATHWAY | Genes involved in Olfactory Signaling Pathway |
| 0.0 | 0.2 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 0.0 | 0.3 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.0 | 0.2 | REACTOME THROMBOXANE SIGNALLING THROUGH TP RECEPTOR | Genes involved in Thromboxane signalling through TP receptor |