Project
PRJNA438478: RNAseq of wild type zebrafish germline
Navigation
Downloads
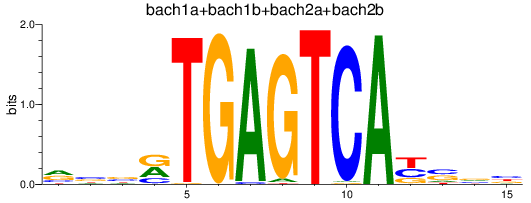
Results for bach1a+bach1b+bach2a+bach2b
Z-value: 0.64
Transcription factors associated with bach1a+bach1b+bach2a+bach2b
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
bach1b
|
ENSDARG00000002196 | BTB and CNC homology 1, basic leucine zipper transcription factor 1 b |
|
bach2b
|
ENSDARG00000004074 | BTB and CNC homology 1, basic leucine zipper transcription factor 2b |
|
bach2a
|
ENSDARG00000036569 | BTB and CNC homology 1, basic leucine zipper transcription factor 2a |
|
bach1a
|
ENSDARG00000062553 | BTB and CNC homology 1, basic leucine zipper transcription factor 1 a |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| bach1a | dr11_v1_chr15_-_8191992_8191992 | 0.90 | 4.6e-07 | Click! |
| bach2a | dr11_v1_chr17_+_15674052_15674052 | 0.86 | 3.8e-06 | Click! |
| bach2b | dr11_v1_chr20_-_24122881_24122881 | 0.82 | 2.5e-05 | Click! |
| bach1b | dr11_v1_chr10_+_25369254_25369254 | -0.43 | 7.8e-02 | Click! |
Activity profile of bach1a+bach1b+bach2a+bach2b motif
Sorted Z-values of bach1a+bach1b+bach2a+bach2b motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr24_-_33366188 | 1.59 |
ENSDART00000074161
|
slc4a2b
|
solute carrier family 4 (anion exchanger), member 2b |
| chr16_-_24832038 | 1.54 |
ENSDART00000153731
|
si:dkey-79d12.5
|
si:dkey-79d12.5 |
| chr16_+_23403602 | 1.47 |
ENSDART00000159848
|
s100w
|
S100 calcium binding protein W |
| chr1_+_46598764 | 1.29 |
ENSDART00000053240
|
cab39l
|
calcium binding protein 39-like |
| chr7_+_26649319 | 1.27 |
ENSDART00000173823
ENSDART00000101053 |
tp53i11a
|
tumor protein p53 inducible protein 11a |
| chr10_-_32877348 | 1.20 |
ENSDART00000018977
ENSDART00000133421 |
rabgef1
|
RAB guanine nucleotide exchange factor (GEF) 1 |
| chr24_-_21973365 | 1.13 |
ENSDART00000081204
ENSDART00000030592 |
acot9.1
|
acyl-CoA thioesterase 9, tandem duplicate 1 |
| chr5_+_44804791 | 1.08 |
ENSDART00000122288
|
ctsla
|
cathepsin La |
| chr1_+_46598502 | 1.00 |
ENSDART00000132861
|
cab39l
|
calcium binding protein 39-like |
| chr11_+_37638873 | 1.00 |
ENSDART00000186384
ENSDART00000184291 ENSDART00000131782 ENSDART00000140502 |
sh2d5
|
SH2 domain containing 5 |
| chr19_+_37120491 | 1.00 |
ENSDART00000032341
|
pef1
|
penta-EF-hand domain containing 1 |
| chr5_+_44805028 | 0.96 |
ENSDART00000141198
|
ctsla
|
cathepsin La |
| chr5_+_44805269 | 0.94 |
ENSDART00000136965
|
ctsla
|
cathepsin La |
| chr8_+_8671229 | 0.89 |
ENSDART00000131963
|
usp11
|
ubiquitin specific peptidase 11 |
| chr2_-_23391266 | 0.88 |
ENSDART00000159048
|
ivns1abpb
|
influenza virus NS1A binding protein b |
| chr21_+_25765734 | 0.84 |
ENSDART00000021664
|
cldnb
|
claudin b |
| chr18_+_3169579 | 0.83 |
ENSDART00000164724
ENSDART00000186340 ENSDART00000181247 ENSDART00000168056 |
pak1
|
p21 protein (Cdc42/Rac)-activated kinase 1 |
| chr18_+_3243292 | 0.83 |
ENSDART00000166580
|
pak1
|
p21 protein (Cdc42/Rac)-activated kinase 1 |
| chr23_+_2728095 | 0.82 |
ENSDART00000066086
|
zgc:114123
|
zgc:114123 |
| chr5_-_49951106 | 0.81 |
ENSDART00000135954
|
fam172a
|
family with sequence similarity 172, member A |
| chr11_+_45219558 | 0.81 |
ENSDART00000167828
|
tmc6b
|
transmembrane channel-like 6b |
| chr5_-_42180205 | 0.79 |
ENSDART00000145247
|
fam222ba
|
family with sequence similarity 222, member Ba |
| chr16_+_27543893 | 0.79 |
ENSDART00000182421
|
si:ch211-197h24.6
|
si:ch211-197h24.6 |
| chr22_+_17399124 | 0.78 |
ENSDART00000145769
|
rabgap1l
|
RAB GTPase activating protein 1-like |
| chr21_+_13233377 | 0.76 |
ENSDART00000142569
|
specc1lb
|
sperm antigen with calponin homology and coiled-coil domains 1-like b |
| chr5_+_9037650 | 0.76 |
ENSDART00000158226
|
si:ch211-155m12.1
|
si:ch211-155m12.1 |
| chr15_+_25489406 | 0.76 |
ENSDART00000162482
|
zgc:152863
|
zgc:152863 |
| chr19_+_551963 | 0.75 |
ENSDART00000110495
|
akap9
|
A kinase (PRKA) anchor protein 9 |
| chr12_-_3077395 | 0.75 |
ENSDART00000002867
ENSDART00000126315 |
rfng
|
RFNG O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase |
| chr6_+_51713076 | 0.74 |
ENSDART00000146281
|
ripor3
|
RIPOR family member 3 |
| chr16_+_13818743 | 0.73 |
ENSDART00000090191
|
flcn
|
folliculin |
| chr3_+_35498119 | 0.71 |
ENSDART00000178963
|
tnrc6a
|
trinucleotide repeat containing 6a |
| chr15_-_31177324 | 0.70 |
ENSDART00000008854
|
wsb1
|
WD repeat and SOCS box containing 1 |
| chr2_+_9821757 | 0.70 |
ENSDART00000018408
ENSDART00000141227 ENSDART00000144681 ENSDART00000148227 |
anxa13l
|
annexin A13, like |
| chr21_+_37477001 | 0.70 |
ENSDART00000114778
|
amot
|
angiomotin |
| chr17_+_19626479 | 0.70 |
ENSDART00000044993
ENSDART00000131863 |
rgs7a
|
regulator of G protein signaling 7a |
| chr12_-_27212596 | 0.69 |
ENSDART00000153101
|
psme3
|
proteasome activator subunit 3 |
| chr16_+_13818500 | 0.69 |
ENSDART00000135245
|
flcn
|
folliculin |
| chr9_+_28140089 | 0.68 |
ENSDART00000046880
|
plekhm3
|
pleckstrin homology domain containing, family M, member 3 |
| chr12_-_17147473 | 0.68 |
ENSDART00000106035
|
stambpl1
|
STAM binding protein-like 1 |
| chr17_+_32623931 | 0.67 |
ENSDART00000144217
|
ctsba
|
cathepsin Ba |
| chr19_+_26072624 | 0.67 |
ENSDART00000147627
|
jarid2b
|
jumonji, AT rich interactive domain 2b |
| chr18_+_15841449 | 0.66 |
ENSDART00000141800
ENSDART00000091349 |
eea1
|
early endosome antigen 1 |
| chr19_-_31035155 | 0.66 |
ENSDART00000161882
|
bzw2
|
basic leucine zipper and W2 domains 2 |
| chr15_+_17343319 | 0.64 |
ENSDART00000018461
|
vmp1
|
vacuole membrane protein 1 |
| chr19_-_5805923 | 0.62 |
ENSDART00000134340
|
si:ch211-264f5.8
|
si:ch211-264f5.8 |
| chr2_+_15100742 | 0.62 |
ENSDART00000027171
|
f3b
|
coagulation factor IIIb |
| chr23_+_24501918 | 0.61 |
ENSDART00000078824
|
szrd1
|
SUZ RNA binding domain containing 1 |
| chr22_-_6562618 | 0.61 |
ENSDART00000106100
|
zgc:171490
|
zgc:171490 |
| chr2_-_26642831 | 0.60 |
ENSDART00000132854
ENSDART00000087714 ENSDART00000132651 |
u2surp
|
U2 snRNP-associated SURP domain containing |
| chr11_-_11965033 | 0.59 |
ENSDART00000193683
|
abcc10
|
ATP-binding cassette, sub-family C (CFTR/MRP), member 10 |
| chr13_+_7575563 | 0.59 |
ENSDART00000146592
|
gbf1
|
golgi brefeldin A resistant guanine nucleotide exchange factor 1 |
| chr12_-_35386910 | 0.59 |
ENSDART00000153453
|
camk2g1
|
calcium/calmodulin-dependent protein kinase (CaM kinase) II gamma 1 |
| chr20_-_34069956 | 0.58 |
ENSDART00000017941
|
tprb
|
translocated promoter region b, nuclear basket protein |
| chr19_-_31035325 | 0.57 |
ENSDART00000147504
|
bzw2
|
basic leucine zipper and W2 domains 2 |
| chr4_-_4535189 | 0.57 |
ENSDART00000057519
|
zgc:194209
|
zgc:194209 |
| chr11_-_39118882 | 0.57 |
ENSDART00000113185
ENSDART00000156526 |
ap5b1
|
adaptor-related protein complex 5, beta 1 subunit |
| chr9_+_41024973 | 0.56 |
ENSDART00000014660
ENSDART00000144467 |
ormdl1
|
ORMDL sphingolipid biosynthesis regulator 1 |
| chr25_+_14697247 | 0.56 |
ENSDART00000180747
|
mpped2
|
metallophosphoesterase domain containing 2b |
| chr13_+_7241170 | 0.56 |
ENSDART00000109434
|
aifm2
|
apoptosis-inducing factor, mitochondrion-associated, 2 |
| chr3_+_19687217 | 0.56 |
ENSDART00000141937
|
tlk2
|
tousled-like kinase 2 |
| chr14_-_36763302 | 0.55 |
ENSDART00000074786
|
ctso
|
cathepsin O |
| chr25_+_18583877 | 0.55 |
ENSDART00000148741
|
met
|
MET proto-oncogene, receptor tyrosine kinase |
| chr20_+_42537768 | 0.55 |
ENSDART00000134066
ENSDART00000153434 |
si:dkeyp-93d12.1
|
si:dkeyp-93d12.1 |
| chr5_+_35786141 | 0.54 |
ENSDART00000022043
ENSDART00000127383 |
stard8
|
StAR-related lipid transfer (START) domain containing 8 |
| chr4_+_25181572 | 0.53 |
ENSDART00000078529
ENSDART00000136643 |
kin
|
Kin17 DNA and RNA binding protein |
| chr5_+_34549845 | 0.51 |
ENSDART00000139317
|
aif1l
|
allograft inflammatory factor 1-like |
| chr4_-_1801519 | 0.51 |
ENSDART00000188604
ENSDART00000135749 |
nudt4b
|
nudix (nucleoside diphosphate linked moiety X)-type motif 4b |
| chr14_-_493029 | 0.49 |
ENSDART00000115093
|
zgc:172215
|
zgc:172215 |
| chr12_-_27212880 | 0.49 |
ENSDART00000002835
|
psme3
|
proteasome activator subunit 3 |
| chr15_+_40188076 | 0.48 |
ENSDART00000063779
|
efhd1
|
EF-hand domain family, member D1 |
| chr24_-_5691956 | 0.48 |
ENSDART00000189112
|
dia1b
|
deleted in autism 1b |
| chr10_+_32050906 | 0.48 |
ENSDART00000137373
|
si:ch211-266i6.3
|
si:ch211-266i6.3 |
| chr19_+_48359259 | 0.47 |
ENSDART00000167353
|
sgo1
|
shugoshin 1 |
| chr24_-_31090948 | 0.47 |
ENSDART00000176799
|
hccsb
|
holocytochrome c synthase b |
| chr2_+_37837249 | 0.46 |
ENSDART00000113337
|
parp2
|
poly (ADP-ribose) polymerase 2 |
| chr18_-_37355666 | 0.46 |
ENSDART00000098914
|
yap1
|
Yes-associated protein 1 |
| chr23_+_45200481 | 0.45 |
ENSDART00000004357
ENSDART00000111126 ENSDART00000193560 ENSDART00000190476 |
psip1b
|
PC4 and SFRS1 interacting protein 1b |
| chr4_+_5317483 | 0.45 |
ENSDART00000150366
|
si:ch211-214j24.10
|
si:ch211-214j24.10 |
| chr24_-_21973163 | 0.45 |
ENSDART00000131406
|
acot9.1
|
acyl-CoA thioesterase 9, tandem duplicate 1 |
| chr23_+_45339684 | 0.45 |
ENSDART00000149410
ENSDART00000102441 |
psip1b
|
PC4 and SFRS1 interacting protein 1b |
| chr2_+_9822319 | 0.43 |
ENSDART00000144078
ENSDART00000144371 |
anxa13l
|
annexin A13, like |
| chr7_-_29723761 | 0.43 |
ENSDART00000173560
|
vps13c
|
vacuolar protein sorting 13 homolog C (S. cerevisiae) |
| chr2_-_39759059 | 0.42 |
ENSDART00000007333
|
slc25a36a
|
solute carrier family 25 (pyrimidine nucleotide carrier ), member 36a |
| chr7_+_34549198 | 0.42 |
ENSDART00000173784
|
fhod1
|
formin homology 2 domain containing 1 |
| chr14_+_28464736 | 0.42 |
ENSDART00000019003
|
psmd10
|
proteasome 26S subunit, non-ATPase 10 |
| chr6_-_35309661 | 0.42 |
ENSDART00000114960
|
nos1apa
|
nitric oxide synthase 1 (neuronal) adaptor protein a |
| chr8_-_34402839 | 0.40 |
ENSDART00000180173
ENSDART00000191588 ENSDART00000186376 |
gapvd1
|
GTPase activating protein and VPS9 domains 1 |
| chr25_-_22187397 | 0.40 |
ENSDART00000123211
ENSDART00000139110 |
pkp3a
|
plakophilin 3a |
| chr15_+_24704798 | 0.40 |
ENSDART00000192470
|
LRRC75A
|
si:dkey-151p21.7 |
| chr13_+_2357637 | 0.40 |
ENSDART00000017148
|
gclc
|
glutamate-cysteine ligase, catalytic subunit |
| chr16_+_46410520 | 0.40 |
ENSDART00000131072
|
rpz2
|
rapunzel 2 |
| chr22_+_26600834 | 0.39 |
ENSDART00000157411
|
adcy9
|
adenylate cyclase 9 |
| chr3_+_17616201 | 0.38 |
ENSDART00000156775
|
rab5c
|
RAB5C, member RAS oncogene family |
| chr3_+_46762703 | 0.38 |
ENSDART00000133283
|
prkcsh
|
protein kinase C substrate 80K-H |
| chr22_-_3595439 | 0.37 |
ENSDART00000083308
|
ptprsa
|
protein tyrosine phosphatase, receptor type, s, a |
| chr7_+_34549377 | 0.37 |
ENSDART00000191814
|
fhod1
|
formin homology 2 domain containing 1 |
| chr3_-_24205339 | 0.37 |
ENSDART00000157135
|
si:dkey-110g7.8
|
si:dkey-110g7.8 |
| chr1_+_47165842 | 0.36 |
ENSDART00000053152
ENSDART00000167051 |
cbr1
|
carbonyl reductase 1 |
| chr7_-_5487593 | 0.36 |
ENSDART00000136594
|
arhgef11
|
Rho guanine nucleotide exchange factor (GEF) 11 |
| chr25_+_10410620 | 0.35 |
ENSDART00000151886
|
ehf
|
ets homologous factor |
| chr21_+_10866421 | 0.34 |
ENSDART00000137858
|
alpk2
|
alpha-kinase 2 |
| chr9_+_7030016 | 0.34 |
ENSDART00000148047
ENSDART00000148181 |
inpp4aa
|
inositol polyphosphate-4-phosphatase type I Aa |
| chr5_-_20678300 | 0.34 |
ENSDART00000088639
|
wscd2
|
WSC domain containing 2 |
| chr5_+_20035284 | 0.34 |
ENSDART00000191808
|
sgsm1a
|
small G protein signaling modulator 1a |
| chr20_-_16548912 | 0.33 |
ENSDART00000137601
|
ches1
|
checkpoint suppressor 1 |
| chr5_-_42123794 | 0.33 |
ENSDART00000051135
|
emc6
|
ER membrane protein complex subunit 6 |
| chr11_+_19370717 | 0.33 |
ENSDART00000165906
|
prickle2b
|
prickle homolog 2b |
| chr1_+_41690402 | 0.32 |
ENSDART00000177298
|
fbxo41
|
F-box protein 41 |
| chr22_+_31023205 | 0.31 |
ENSDART00000111561
|
zmp:0000000735
|
zmp:0000000735 |
| chr11_+_807153 | 0.30 |
ENSDART00000173289
|
vgll4b
|
vestigial-like family member 4b |
| chr8_-_11170114 | 0.30 |
ENSDART00000133532
|
si:ch211-204d2.4
|
si:ch211-204d2.4 |
| chr21_+_6328801 | 0.29 |
ENSDART00000163577
|
fnbp1b
|
formin binding protein 1b |
| chr15_-_43978141 | 0.28 |
ENSDART00000041249
|
chordc1a
|
cysteine and histidine-rich domain (CHORD) containing 1a |
| chr18_+_44849809 | 0.27 |
ENSDART00000097328
|
arfgap2
|
ADP-ribosylation factor GTPase activating protein 2 |
| chr10_-_13239367 | 0.27 |
ENSDART00000001253
|
si:busm1-57f23.1
|
si:busm1-57f23.1 |
| chr23_-_4705110 | 0.27 |
ENSDART00000144536
ENSDART00000129050 ENSDART00000136399 |
cnbpa
|
CCHC-type zinc finger, nucleic acid binding protein a |
| chr7_-_19638319 | 0.26 |
ENSDART00000163686
|
si:ch211-212k18.4
|
si:ch211-212k18.4 |
| chr24_+_80653 | 0.26 |
ENSDART00000158473
ENSDART00000129135 |
reck
|
reversion-inducing-cysteine-rich protein with kazal motifs |
| chr18_+_618005 | 0.24 |
ENSDART00000189667
|
prtga
|
protogenin homolog a (Gallus gallus) |
| chr11_-_14102131 | 0.24 |
ENSDART00000085158
ENSDART00000191962 |
tmem259
|
transmembrane protein 259 |
| chr6_+_56147812 | 0.23 |
ENSDART00000150219
|
tfap2c
|
transcription factor AP-2 gamma (activating enhancer binding protein 2 gamma) |
| chr2_+_47944967 | 0.23 |
ENSDART00000135584
|
ftr19
|
finTRIM family, member 19 |
| chr20_-_21994901 | 0.22 |
ENSDART00000004984
|
daam1b
|
dishevelled associated activator of morphogenesis 1b |
| chr23_-_4704938 | 0.22 |
ENSDART00000067293
|
cnbpa
|
CCHC-type zinc finger, nucleic acid binding protein a |
| chr21_-_30254185 | 0.21 |
ENSDART00000101054
|
dnajc18
|
DnaJ (Hsp40) homolog, subfamily C, member 18 |
| chr14_+_34486629 | 0.21 |
ENSDART00000131861
|
tmsb2
|
thymosin beta 2 |
| chr7_-_5396154 | 0.21 |
ENSDART00000172980
|
arhgef11
|
Rho guanine nucleotide exchange factor (GEF) 11 |
| chr9_-_22057658 | 0.20 |
ENSDART00000101944
|
crygmxl1
|
crystallin, gamma MX, like 1 |
| chr20_-_16849306 | 0.19 |
ENSDART00000131395
ENSDART00000027582 |
brms1lb
|
breast cancer metastasis-suppressor 1-like b |
| chr7_+_38680034 | 0.19 |
ENSDART00000171691
ENSDART00000007913 |
psmc3
|
proteasome 26S subunit, ATPase 3 |
| chr17_+_58211 | 0.18 |
ENSDART00000157642
|
si:ch1073-209e23.1
|
si:ch1073-209e23.1 |
| chr18_-_15155962 | 0.18 |
ENSDART00000111333
|
mterf2
|
mitochondrial transcription termination factor 2 |
| chr7_+_7696665 | 0.18 |
ENSDART00000091099
|
ino80b
|
INO80 complex subunit B |
| chr15_-_36727462 | 0.17 |
ENSDART00000085971
|
nphs1
|
nephrosis 1, congenital, Finnish type (nephrin) |
| chr20_-_34767472 | 0.16 |
ENSDART00000186710
|
paqr8
|
progestin and adipoQ receptor family member VIII |
| chr11_+_19370447 | 0.16 |
ENSDART00000186154
|
prickle2b
|
prickle homolog 2b |
| chr2_+_36109002 | 0.15 |
ENSDART00000158978
|
traj28
|
T-cell receptor alpha joining 28 |
| chr24_-_24797455 | 0.15 |
ENSDART00000138741
|
pde7a
|
phosphodiesterase 7A |
| chr14_+_30279391 | 0.14 |
ENSDART00000172794
|
fgl1
|
fibrinogen-like 1 |
| chr2_+_47945359 | 0.13 |
ENSDART00000098054
|
ftr19
|
finTRIM family, member 19 |
| chr3_-_16493528 | 0.12 |
ENSDART00000142099
|
si:ch211-23l10.3
|
si:ch211-23l10.3 |
| chr11_+_11267829 | 0.12 |
ENSDART00000026814
ENSDART00000173346 ENSDART00000151926 |
ptp4a1
|
protein tyrosine phosphatase type IVA, member 1 |
| chr19_-_7272921 | 0.12 |
ENSDART00000102075
ENSDART00000132887 ENSDART00000130234 ENSDART00000193535 ENSDART00000136528 |
rxrba
|
retinoid x receptor, beta a |
| chr5_-_14564878 | 0.12 |
ENSDART00000160511
|
sema4c
|
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4C |
| chr1_-_53750522 | 0.12 |
ENSDART00000190755
|
akt3b
|
v-akt murine thymoma viral oncogene homolog 3b |
| chr4_-_77506362 | 0.11 |
ENSDART00000174387
ENSDART00000181181 |
CABZ01087415.1
|
|
| chr2_-_898899 | 0.11 |
ENSDART00000058289
|
dusp22b
|
dual specificity phosphatase 22b |
| chr12_-_29233738 | 0.10 |
ENSDART00000153175
|
si:ch211-214e3.5
|
si:ch211-214e3.5 |
| chr4_+_43408004 | 0.09 |
ENSDART00000150476
|
si:dkeyp-53e4.2
|
si:dkeyp-53e4.2 |
| chr10_+_4235998 | 0.09 |
ENSDART00000169328
|
plppr1
|
phospholipid phosphatase related 1 |
| chr5_+_27488975 | 0.09 |
ENSDART00000123635
|
sfrp1a
|
secreted frizzled-related protein 1a |
| chr8_+_48965767 | 0.09 |
ENSDART00000008058
|
aak1a
|
AP2 associated kinase 1a |
| chr18_+_17537344 | 0.09 |
ENSDART00000025782
|
nup93
|
nucleoporin 93 |
| chr19_-_10395683 | 0.09 |
ENSDART00000109488
|
zgc:194578
|
zgc:194578 |
| chr23_+_7548797 | 0.09 |
ENSDART00000006765
|
tm9sf4
|
transmembrane 9 superfamily protein member 4 |
| chr3_-_51142942 | 0.09 |
ENSDART00000160789
|
si:ch211-148f13.1
|
si:ch211-148f13.1 |
| chr21_+_25688388 | 0.09 |
ENSDART00000125709
|
bicdl2
|
bicaudal-D-related protein 2 |
| chr23_-_36441693 | 0.09 |
ENSDART00000024354
|
csad
|
cysteine sulfinic acid decarboxylase |
| chr11_+_11267493 | 0.09 |
ENSDART00000148425
|
ptp4a1
|
protein tyrosine phosphatase type IVA, member 1 |
| chr25_+_32474031 | 0.08 |
ENSDART00000152124
|
sqor
|
sulfide quinone oxidoreductase |
| chr5_-_30615901 | 0.08 |
ENSDART00000147769
|
si:ch211-117m20.5
|
si:ch211-117m20.5 |
| chr5_+_42064144 | 0.08 |
ENSDART00000035235
|
si:ch211-202a12.4
|
si:ch211-202a12.4 |
| chr9_-_9705512 | 0.08 |
ENSDART00000149021
|
gpr156
|
G protein-coupled receptor 156 |
| chr4_+_49196098 | 0.07 |
ENSDART00000150678
|
si:dkey-40n15.1
|
si:dkey-40n15.1 |
| chr24_-_23998897 | 0.07 |
ENSDART00000130053
|
zmp:0000000991
|
zmp:0000000991 |
| chr8_+_48966165 | 0.07 |
ENSDART00000165425
|
aak1a
|
AP2 associated kinase 1a |
| chr6_-_35439406 | 0.06 |
ENSDART00000073784
|
rgs5a
|
regulator of G protein signaling 5a |
| chr19_+_4139065 | 0.06 |
ENSDART00000172524
|
si:dkey-218f9.10
|
si:dkey-218f9.10 |
| chr8_+_39511932 | 0.06 |
ENSDART00000113511
|
lzts1
|
leucine zipper, putative tumor suppressor 1 |
| chr23_+_17522867 | 0.06 |
ENSDART00000002714
|
slc17a9b
|
solute carrier family 17 (vesicular nucleotide transporter), member 9b |
| chr7_+_34786591 | 0.06 |
ENSDART00000173700
|
si:dkey-148a17.5
|
si:dkey-148a17.5 |
| chr24_+_30215475 | 0.06 |
ENSDART00000164717
|
si:ch73-358j7.2
|
si:ch73-358j7.2 |
| chr17_-_45247332 | 0.05 |
ENSDART00000016815
|
ttbk2a
|
tau tubulin kinase 2a |
| chr13_+_1430094 | 0.05 |
ENSDART00000169888
|
si:ch211-165e15.1
|
si:ch211-165e15.1 |
| chr4_+_7822773 | 0.05 |
ENSDART00000171391
|
si:ch1073-67j19.2
|
si:ch1073-67j19.2 |
| chr17_-_19626357 | 0.05 |
ENSDART00000011432
|
reep3a
|
receptor accessory protein 3a |
| chr9_+_22388686 | 0.04 |
ENSDART00000182731
ENSDART00000181462 |
dgkg
|
diacylglycerol kinase, gamma |
| chr15_-_38129845 | 0.04 |
ENSDART00000057095
|
si:dkey-24p1.1
|
si:dkey-24p1.1 |
| chr22_-_26595027 | 0.04 |
ENSDART00000184162
|
CABZ01072309.1
|
|
| chr18_-_13360106 | 0.04 |
ENSDART00000091512
|
cmip
|
c-Maf inducing protein |
| chr3_-_584950 | 0.04 |
ENSDART00000164752
|
dicp1.1
|
diverse immunoglobulin domain-containing protein 1.1 |
| chr15_+_28989573 | 0.04 |
ENSDART00000076648
|
clip3
|
CAP-GLY domain containing linker protein 3 |
| chr3_-_34816893 | 0.04 |
ENSDART00000084448
ENSDART00000154696 |
psmd11a
|
proteasome 26S subunit, non-ATPase 11a |
| chr4_+_54899568 | 0.04 |
ENSDART00000162786
|
si:dkey-56m15.8
|
si:dkey-56m15.8 |
| chr3_-_48259289 | 0.04 |
ENSDART00000160717
|
znf750
|
zinc finger protein 750 |
| chr10_+_22771176 | 0.04 |
ENSDART00000192046
|
tmem88a
|
transmembrane protein 88 a |
| chr8_-_44463985 | 0.03 |
ENSDART00000016845
|
mhc1lba
|
major histocompatibility complex class I LBA |
| chr16_+_48714048 | 0.03 |
ENSDART00000148709
ENSDART00000150121 |
brd2b
|
bromodomain containing 2b |
| chr22_+_8800432 | 0.03 |
ENSDART00000139940
|
si:dkey-182g1.6
|
si:dkey-182g1.6 |
| chr21_+_26620867 | 0.03 |
ENSDART00000176008
|
esrra
|
estrogen-related receptor alpha |
| chr10_-_20445549 | 0.03 |
ENSDART00000064613
|
loxl2a
|
lysyl oxidase-like 2a |
| chr23_-_28239750 | 0.03 |
ENSDART00000003548
|
znf385a
|
zinc finger protein 385A |
| chr20_-_47270519 | 0.03 |
ENSDART00000153329
|
si:dkeyp-104f11.8
|
si:dkeyp-104f11.8 |
| chr6_+_10534752 | 0.02 |
ENSDART00000090874
|
kcnh7
|
potassium channel, voltage gated eag related subfamily H, member 7 |
| chr4_-_71913556 | 0.02 |
ENSDART00000167608
|
si:dkey-92c21.1
|
si:dkey-92c21.1 |
| chr2_+_25278107 | 0.02 |
ENSDART00000131977
|
ppp2r3a
|
protein phosphatase 2, regulatory subunit B'', alpha |
| chr4_-_64604842 | 0.02 |
ENSDART00000169862
|
znf1099
|
zinc finger protein 1099 |
| chr4_+_69624049 | 0.02 |
ENSDART00000170052
|
znf1030
|
zinc finger protein 1030 |
| chr6_-_21534301 | 0.02 |
ENSDART00000126186
|
psmd12
|
proteasome 26S subunit, non-ATPase 12 |
Network of associatons between targets according to the STRING database.
First level regulatory network of bach1a+bach1b+bach2a+bach2b
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.7 | GO:0003232 | bulbus arteriosus development(GO:0003232) |
| 0.3 | 0.8 | GO:0097095 | frontonasal suture morphogenesis(GO:0097095) |
| 0.2 | 0.7 | GO:0051574 | positive regulation of histone H3-K9 methylation(GO:0051574) |
| 0.2 | 1.4 | GO:0030511 | positive regulation of transforming growth factor beta receptor signaling pathway(GO:0030511) positive regulation of cellular response to transforming growth factor beta stimulus(GO:1903846) |
| 0.1 | 0.4 | GO:1990519 | pyrimidine nucleotide transport(GO:0006864) mitochondrial pyrimidine nucleotide import(GO:1990519) |
| 0.1 | 0.6 | GO:2000303 | regulation of sphingolipid biosynthetic process(GO:0090153) negative regulation of sphingolipid biosynthetic process(GO:0090155) negative regulation of ceramide biosynthetic process(GO:1900060) regulation of membrane lipid metabolic process(GO:1905038) regulation of ceramide biosynthetic process(GO:2000303) |
| 0.1 | 0.5 | GO:0071962 | mitotic sister chromatid cohesion, centromeric(GO:0071962) |
| 0.1 | 0.8 | GO:0031048 | chromatin silencing by small RNA(GO:0031048) |
| 0.1 | 1.0 | GO:0048207 | vesicle targeting, rough ER to cis-Golgi(GO:0048207) COPII vesicle coating(GO:0048208) |
| 0.1 | 0.8 | GO:0040038 | meiotic cytokinesis(GO:0033206) polar body extrusion after meiotic divisions(GO:0040038) |
| 0.1 | 0.5 | GO:1901911 | diphosphoinositol polyphosphate metabolic process(GO:0071543) diadenosine pentaphosphate metabolic process(GO:1901906) diadenosine pentaphosphate catabolic process(GO:1901907) diadenosine hexaphosphate metabolic process(GO:1901908) diadenosine hexaphosphate catabolic process(GO:1901909) adenosine 5'-(hexahydrogen pentaphosphate) metabolic process(GO:1901910) adenosine 5'-(hexahydrogen pentaphosphate) catabolic process(GO:1901911) |
| 0.1 | 0.4 | GO:0006750 | glutathione biosynthetic process(GO:0006750) |
| 0.1 | 0.5 | GO:2000767 | positive regulation of cytoplasmic translation(GO:2000767) |
| 0.1 | 0.5 | GO:0017006 | protein-heme linkage(GO:0017003) protein-tetrapyrrole linkage(GO:0017006) cytochrome c-heme linkage(GO:0018063) |
| 0.1 | 0.5 | GO:0003173 | ventriculo bulbo valve development(GO:0003173) |
| 0.1 | 0.3 | GO:0006901 | vesicle coating(GO:0006901) |
| 0.1 | 0.7 | GO:0003365 | establishment of cell polarity involved in ameboidal cell migration(GO:0003365) |
| 0.1 | 0.3 | GO:1905048 | regulation of metallopeptidase activity(GO:1905048) |
| 0.1 | 0.6 | GO:1901673 | regulation of mitotic spindle assembly(GO:1901673) |
| 0.1 | 0.6 | GO:0001672 | regulation of chromatin assembly or disassembly(GO:0001672) |
| 0.1 | 1.6 | GO:0015701 | bicarbonate transport(GO:0015701) |
| 0.1 | 0.8 | GO:0035283 | central nervous system segmentation(GO:0035283) brain segmentation(GO:0035284) |
| 0.1 | 0.4 | GO:0045053 | protein retention in Golgi apparatus(GO:0045053) |
| 0.1 | 0.8 | GO:0051639 | actin filament network formation(GO:0051639) |
| 0.1 | 0.6 | GO:0021681 | cerebellar granular layer development(GO:0021681) cerebellar granular layer morphogenesis(GO:0021683) cerebellar granular layer formation(GO:0021684) cerebellar granule cell differentiation(GO:0021707) |
| 0.0 | 0.3 | GO:0045050 | protein insertion into ER membrane by stop-transfer membrane-anchor sequence(GO:0045050) |
| 0.0 | 0.2 | GO:1904292 | regulation of ERAD pathway(GO:1904292) |
| 0.0 | 0.7 | GO:0040033 | miRNA mediated inhibition of translation(GO:0035278) negative regulation of translation, ncRNA-mediated(GO:0040033) regulation of translation, ncRNA-mediated(GO:0045974) |
| 0.0 | 0.5 | GO:0070212 | protein poly-ADP-ribosylation(GO:0070212) |
| 0.0 | 0.6 | GO:0042953 | lipoprotein transport(GO:0042953) lipoprotein localization(GO:0044872) |
| 0.0 | 0.2 | GO:0045898 | regulation of RNA polymerase II transcriptional preinitiation complex assembly(GO:0045898) positive regulation of RNA polymerase II transcriptional preinitiation complex assembly(GO:0045899) |
| 0.0 | 0.2 | GO:0014034 | neural crest cell fate commitment(GO:0014034) neural crest cell fate specification(GO:0014036) |
| 0.0 | 0.4 | GO:0099560 | synaptic membrane adhesion(GO:0099560) |
| 0.0 | 1.2 | GO:2000045 | regulation of G1/S transition of mitotic cell cycle(GO:2000045) |
| 0.0 | 0.1 | GO:0019418 | sulfide oxidation(GO:0019418) sulfide oxidation, using sulfide:quinone oxidoreductase(GO:0070221) |
| 0.0 | 0.5 | GO:0097178 | ruffle assembly(GO:0097178) |
| 0.0 | 0.7 | GO:0070536 | protein K63-linked deubiquitination(GO:0070536) |
| 0.0 | 0.2 | GO:0006390 | transcription from mitochondrial promoter(GO:0006390) |
| 0.0 | 0.1 | GO:0055107 | Golgi to secretory granule transport(GO:0055107) |
| 0.0 | 0.2 | GO:0042989 | sequestering of actin monomers(GO:0042989) |
| 0.0 | 0.1 | GO:0051292 | nuclear pore complex assembly(GO:0051292) |
| 0.0 | 0.6 | GO:0008637 | apoptotic mitochondrial changes(GO:0008637) |
| 0.0 | 0.2 | GO:2000369 | regulation of clathrin-mediated endocytosis(GO:2000369) |
| 0.0 | 0.6 | GO:0072310 | glomerular visceral epithelial cell development(GO:0072015) glomerular epithelial cell development(GO:0072310) |
| 0.0 | 0.4 | GO:0006491 | N-glycan processing(GO:0006491) |
| 0.0 | 0.8 | GO:0070830 | apical junction assembly(GO:0043297) bicellular tight junction assembly(GO:0070830) |
| 0.0 | 0.6 | GO:0072114 | pronephros morphogenesis(GO:0072114) |
| 0.0 | 1.6 | GO:0006637 | acyl-CoA metabolic process(GO:0006637) thioester metabolic process(GO:0035383) |
| 0.0 | 1.5 | GO:0045930 | negative regulation of mitotic cell cycle(GO:0045930) |
| 0.0 | 0.6 | GO:0003351 | epithelial cilium movement(GO:0003351) |
| 0.0 | 0.5 | GO:0051904 | melanosome transport(GO:0032402) pigment granule transport(GO:0051904) |
| 0.0 | 0.6 | GO:0032011 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.0 | 0.3 | GO:0035329 | hippo signaling(GO:0035329) |
| 0.0 | 0.0 | GO:1901296 | negative regulation of striated muscle cell differentiation(GO:0051154) negative regulation of cardiac muscle tissue development(GO:0055026) cardiac muscle cell fate commitment(GO:0060923) canonical Wnt signaling pathway involved in cardiac muscle cell fate commitment(GO:0061317) regulation of canonical Wnt signaling pathway involved in cardiac muscle cell fate commitment(GO:1901295) negative regulation of canonical Wnt signaling pathway involved in cardiac muscle cell fate commitment(GO:1901296) negative regulation of cardiocyte differentiation(GO:1905208) negative regulation of cardiac muscle cell differentiation(GO:2000726) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.2 | GO:0008537 | proteasome activator complex(GO:0008537) |
| 0.1 | 0.6 | GO:0035339 | SPOTS complex(GO:0035339) |
| 0.1 | 0.4 | GO:0017177 | glucosidase II complex(GO:0017177) |
| 0.1 | 0.2 | GO:1990909 | Wnt signalosome(GO:1990909) |
| 0.0 | 0.7 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.0 | 1.0 | GO:0030127 | COPII vesicle coat(GO:0030127) |
| 0.0 | 0.4 | GO:0042788 | polysomal ribosome(GO:0042788) |
| 0.0 | 0.8 | GO:0005921 | gap junction(GO:0005921) |
| 0.0 | 0.3 | GO:0072546 | ER membrane protein complex(GO:0072546) |
| 0.0 | 0.6 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.0 | 0.6 | GO:0033116 | endoplasmic reticulum-Golgi intermediate compartment membrane(GO:0033116) |
| 0.0 | 1.9 | GO:0016323 | basolateral plasma membrane(GO:0016323) |
| 0.0 | 0.4 | GO:0030057 | desmosome(GO:0030057) |
| 0.0 | 5.6 | GO:0005764 | lysosome(GO:0005764) |
| 0.0 | 0.6 | GO:0030119 | AP-type membrane coat adaptor complex(GO:0030119) |
| 0.0 | 0.2 | GO:0008540 | proteasome regulatory particle, base subcomplex(GO:0008540) |
| 0.0 | 1.5 | GO:0070160 | bicellular tight junction(GO:0005923) occluding junction(GO:0070160) |
| 0.0 | 0.7 | GO:0005643 | nuclear pore(GO:0005643) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.0 | GO:1990756 | protein binding, bridging involved in substrate recognition for ubiquitination(GO:1990756) |
| 0.2 | 1.2 | GO:0061133 | endopeptidase activator activity(GO:0061133) |
| 0.2 | 0.8 | GO:0033829 | O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase activity(GO:0033829) |
| 0.2 | 0.6 | GO:0016649 | electron-transferring-flavoprotein dehydrogenase activity(GO:0004174) oxidoreductase activity, acting on the CH-NH group of donors, quinone or similar compound as acceptor(GO:0016649) |
| 0.2 | 0.8 | GO:0035197 | siRNA binding(GO:0035197) |
| 0.1 | 0.4 | GO:0015218 | pyrimidine nucleotide transmembrane transporter activity(GO:0015218) |
| 0.1 | 0.5 | GO:0034432 | endopolyphosphatase activity(GO:0000298) diphosphoinositol-polyphosphate diphosphatase activity(GO:0008486) bis(5'-adenosyl)-hexaphosphatase activity(GO:0034431) bis(5'-adenosyl)-pentaphosphatase activity(GO:0034432) |
| 0.1 | 0.5 | GO:0004408 | holocytochrome-c synthase activity(GO:0004408) |
| 0.1 | 1.6 | GO:0005452 | inorganic anion exchanger activity(GO:0005452) |
| 0.1 | 2.3 | GO:0043539 | protein serine/threonine kinase activator activity(GO:0043539) |
| 0.1 | 0.4 | GO:0050998 | nitric-oxide synthase binding(GO:0050998) |
| 0.1 | 0.7 | GO:0034647 | histone demethylase activity (H3-trimethyl-K4 specific)(GO:0034647) |
| 0.1 | 0.4 | GO:0043531 | ADP binding(GO:0043531) |
| 0.1 | 1.6 | GO:0047617 | acyl-CoA hydrolase activity(GO:0047617) |
| 0.1 | 0.3 | GO:0001223 | transcription coactivator binding(GO:0001223) |
| 0.1 | 1.5 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.1 | 0.6 | GO:0016840 | carbon-nitrogen lyase activity(GO:0016840) |
| 0.0 | 0.3 | GO:0016316 | phosphatidylinositol-3,4-bisphosphate 4-phosphatase activity(GO:0016316) |
| 0.0 | 0.6 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.0 | 0.7 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.0 | 0.5 | GO:1990404 | protein ADP-ribosylase activity(GO:1990404) |
| 0.0 | 0.3 | GO:0032977 | membrane insertase activity(GO:0032977) |
| 0.0 | 4.2 | GO:0004197 | cysteine-type endopeptidase activity(GO:0004197) |
| 0.0 | 0.1 | GO:0070224 | sulfide:quinone oxidoreductase activity(GO:0070224) |
| 0.0 | 0.8 | GO:0022833 | mechanically-gated ion channel activity(GO:0008381) mechanically gated channel activity(GO:0022833) |
| 0.0 | 0.7 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.0 | 0.3 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.0 | 0.6 | GO:0099529 | neurotransmitter receptor activity involved in regulation of postsynaptic membrane potential(GO:0099529) transmitter-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1904315) |
| 0.0 | 0.6 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.0 | 0.2 | GO:0030544 | Hsp70 protein binding(GO:0030544) |
| 0.0 | 0.1 | GO:0044323 | retinoic acid-responsive element binding(GO:0044323) |
| 0.0 | 0.2 | GO:0035612 | AP-2 adaptor complex binding(GO:0035612) |
| 0.0 | 0.3 | GO:0004016 | adenylate cyclase activity(GO:0004016) |
| 0.0 | 0.6 | GO:0004683 | calmodulin-dependent protein kinase activity(GO:0004683) |
| 0.0 | 0.3 | GO:0030165 | PDZ domain binding(GO:0030165) |
| 0.0 | 0.5 | GO:0045182 | translation regulator activity(GO:0045182) |
| 0.0 | 0.2 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.0 | 1.0 | GO:0005544 | calcium-dependent phospholipid binding(GO:0005544) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.7 | PID S1P S1P2 PATHWAY | S1P2 pathway |
| 0.0 | 1.2 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.0 | 0.6 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.0 | 0.4 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 0.0 | 0.6 | PID ARF6 PATHWAY | Arf6 signaling events |
| 0.0 | 0.6 | ST GA13 PATHWAY | G alpha 13 Pathway |
| 0.0 | 0.2 | PID TAP63 PATHWAY | Validated transcriptional targets of TAp63 isoforms |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.8 | REACTOME CREB PHOSPHORYLATION THROUGH THE ACTIVATION OF CAMKII | Genes involved in CREB phosphorylation through the activation of CaMKII |
| 0.2 | 1.7 | REACTOME CD28 DEPENDENT VAV1 PATHWAY | Genes involved in CD28 dependent Vav1 pathway |
| 0.2 | 2.3 | REACTOME REGULATION OF AMPK ACTIVITY VIA LKB1 | Genes involved in Regulation of AMPK activity via LKB1 |
| 0.1 | 0.6 | REACTOME COPI MEDIATED TRANSPORT | Genes involved in COPI Mediated Transport |
| 0.1 | 1.2 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.1 | 1.1 | REACTOME SEMA4D IN SEMAPHORIN SIGNALING | Genes involved in Sema4D in semaphorin signaling |
| 0.0 | 0.4 | REACTOME ADVANCED GLYCOSYLATION ENDPRODUCT RECEPTOR SIGNALING | Genes involved in Advanced glycosylation endproduct receptor signaling |
| 0.0 | 0.8 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.0 | 0.7 | REACTOME PRE NOTCH TRANSCRIPTION AND TRANSLATION | Genes involved in Pre-NOTCH Transcription and Translation |
| 0.0 | 0.4 | REACTOME ADENYLATE CYCLASE ACTIVATING PATHWAY | Genes involved in Adenylate cyclase activating pathway |
| 0.0 | 0.4 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.0 | 0.9 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.0 | 0.6 | REACTOME CROSS PRESENTATION OF SOLUBLE EXOGENOUS ANTIGENS ENDOSOMES | Genes involved in Cross-presentation of soluble exogenous antigens (endosomes) |