Project
PRJNA207719: Tissue specific transcriptome profiling
Navigation
Downloads
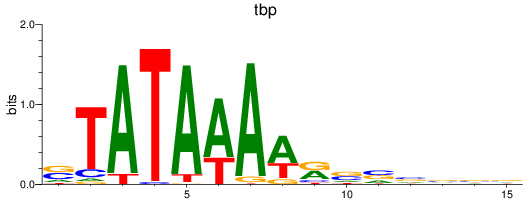
Results for tbp
Z-value: 3.30
Transcription factors associated with tbp
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
tbp
|
ENSDARG00000014994 | TATA box binding protein |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| tbp | dr11_v1_chr13_-_24396199_24396199 | -0.75 | 1.5e-01 | Click! |
Activity profile of tbp motif
Sorted Z-values of tbp motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr9_-_33081978 | 3.64 |
ENSDART00000100918
|
zgc:172053
|
zgc:172053 |
| chr18_-_5527050 | 3.29 |
ENSDART00000145400
ENSDART00000132498 ENSDART00000146209 |
zgc:153317
|
zgc:153317 |
| chr23_-_10177442 | 3.08 |
ENSDART00000144280
ENSDART00000129044 |
krt5
|
keratin 5 |
| chr9_-_33081781 | 3.04 |
ENSDART00000165748
|
zgc:172053
|
zgc:172053 |
| chr3_-_32818607 | 2.97 |
ENSDART00000075465
|
mylpfa
|
myosin light chain, phosphorylatable, fast skeletal muscle a |
| chr1_-_59169815 | 2.84 |
ENSDART00000100163
|
wu:fk65c09
|
wu:fk65c09 |
| chr9_-_33062891 | 2.83 |
ENSDART00000161182
|
si:ch211-125e6.5
|
si:ch211-125e6.5 |
| chr9_-_33063083 | 2.82 |
ENSDART00000048550
|
si:ch211-125e6.5
|
si:ch211-125e6.5 |
| chr21_+_25221940 | 2.62 |
ENSDART00000108972
|
sycn.1
|
syncollin, tandem duplicate 1 |
| chr21_+_5531138 | 2.57 |
ENSDART00000163825
|
ly6m6
|
lymphocyte antigen 6 family member M6 |
| chr19_-_5332784 | 2.56 |
ENSDART00000010373
|
krt1-19d
|
keratin, type 1, gene 19d |
| chr7_+_35068036 | 2.40 |
ENSDART00000022139
|
zgc:136461
|
zgc:136461 |
| chr1_-_43920371 | 2.38 |
ENSDART00000109283
|
scpp7
|
secretory calcium-binding phosphoprotein 7 |
| chr9_+_33207574 | 2.31 |
ENSDART00000055897
ENSDART00000166030 |
si:ch211-125e6.11
|
si:ch211-125e6.11 |
| chr10_+_38593645 | 2.28 |
ENSDART00000011573
|
mmp13a
|
matrix metallopeptidase 13a |
| chr5_-_25583125 | 2.24 |
ENSDART00000031665
ENSDART00000145353 |
anxa1a
|
annexin A1a |
| chr1_-_43920576 | 2.19 |
ENSDART00000191914
|
scpp7
|
secretory calcium-binding phosphoprotein 7 |
| chr1_-_49225890 | 2.11 |
ENSDART00000111598
|
cxcl18b
|
chemokine (C-X-C motif) ligand 18b |
| chr16_-_25085327 | 2.10 |
ENSDART00000077661
|
prss1
|
protease, serine 1 |
| chr2_+_55365727 | 1.98 |
ENSDART00000162943
|
FP245456.1
|
|
| chr19_-_5345930 | 1.91 |
ENSDART00000066620
ENSDART00000151398 |
krtt1c19e
|
keratin type 1 c19e |
| chr14_-_11507211 | 1.91 |
ENSDART00000186873
ENSDART00000109181 ENSDART00000186166 ENSDART00000186986 |
zgc:174917
|
zgc:174917 |
| chr22_-_263117 | 1.90 |
ENSDART00000158134
|
zgc:66156
|
zgc:66156 |
| chr21_-_42876565 | 1.88 |
ENSDART00000126480
|
zmp:0000001268
|
zmp:0000001268 |
| chr9_-_45602978 | 1.85 |
ENSDART00000139019
ENSDART00000085763 |
agr1
|
anterior gradient 1 |
| chr23_+_45456490 | 1.73 |
ENSDART00000036631
|
cyr61
|
cysteine-rich, angiogenic inducer, 61 |
| chr25_-_25550938 | 1.73 |
ENSDART00000150412
ENSDART00000103622 |
irf7
|
interferon regulatory factor 7 |
| chr16_-_45917322 | 1.67 |
ENSDART00000060822
|
afp4
|
antifreeze protein type IV |
| chr22_+_7480465 | 1.67 |
ENSDART00000034545
|
CELA1 (1 of many)
|
zgc:92745 |
| chr5_-_30615901 | 1.66 |
ENSDART00000147769
|
si:ch211-117m20.5
|
si:ch211-117m20.5 |
| chr7_+_8456999 | 1.65 |
ENSDART00000172880
|
jac4
|
jacalin 4 |
| chr22_-_282498 | 1.65 |
ENSDART00000182766
|
CABZ01079178.1
|
|
| chr24_-_27419198 | 1.65 |
ENSDART00000141124
|
ccl34b.4
|
chemokine (C-C motif) ligand 34b, duplicate 4 |
| chr9_+_23770666 | 1.64 |
ENSDART00000182493
|
si:ch211-219a4.3
|
si:ch211-219a4.3 |
| chr13_-_12660318 | 1.62 |
ENSDART00000008498
|
adh8a
|
alcohol dehydrogenase 8a |
| chr14_-_11430566 | 1.61 |
ENSDART00000137154
ENSDART00000091158 |
irg1l
|
immunoresponsive gene 1, like |
| chr23_-_40194732 | 1.58 |
ENSDART00000164931
|
tgm1l2
|
transglutaminase 1 like 2 |
| chr16_+_35401543 | 1.58 |
ENSDART00000171608
|
rab42b
|
RAB42, member RAS oncogene family |
| chr4_+_7817996 | 1.57 |
ENSDART00000166809
|
si:ch1073-67j19.1
|
si:ch1073-67j19.1 |
| chr7_-_7810348 | 1.57 |
ENSDART00000171984
|
cxcl19
|
chemokine (C-X-C motif) ligand 19 |
| chr10_-_7785930 | 1.55 |
ENSDART00000043961
ENSDART00000111058 |
mpx
|
myeloid-specific peroxidase |
| chr25_-_18454016 | 1.55 |
ENSDART00000005877
|
cpa1
|
carboxypeptidase A1 (pancreatic) |
| chr5_+_21181047 | 1.52 |
ENSDART00000088506
|
arhgap25
|
Rho GTPase activating protein 25 |
| chr5_-_36837846 | 1.52 |
ENSDART00000032481
|
ckma
|
creatine kinase, muscle a |
| chr25_-_12982193 | 1.50 |
ENSDART00000159617
|
ccl39.5
|
chemokine (C-C motif) ligand 39, duplicate 5 |
| chr22_-_24248420 | 1.50 |
ENSDART00000165433
|
rgs2
|
regulator of G protein signaling 2 |
| chr10_+_44956660 | 1.49 |
ENSDART00000169225
ENSDART00000189298 |
il1b
|
interleukin 1, beta |
| chr3_-_36606884 | 1.49 |
ENSDART00000172003
|
si:dkeyp-72e1.6
|
si:dkeyp-72e1.6 |
| chr22_+_16497670 | 1.47 |
ENSDART00000014330
|
ier5
|
immediate early response 5 |
| chr21_+_25226558 | 1.47 |
ENSDART00000168480
|
sycn.2
|
syncollin, tandem duplicate 2 |
| chr25_+_35020529 | 1.47 |
ENSDART00000158016
|
flnca
|
filamin C, gamma a (actin binding protein 280) |
| chr15_-_5815006 | 1.46 |
ENSDART00000102459
|
rbp2a
|
retinol binding protein 2a, cellular |
| chr9_-_30264415 | 1.46 |
ENSDART00000060150
|
mid1ip1a
|
MID1 interacting protein 1a |
| chr2_+_24507770 | 1.45 |
ENSDART00000154802
ENSDART00000052063 |
rps28
|
ribosomal protein S28 |
| chr3_-_16784280 | 1.44 |
ENSDART00000137108
ENSDART00000137276 |
si:dkey-30j10.5
|
si:dkey-30j10.5 |
| chr11_+_6136220 | 1.43 |
ENSDART00000082223
|
tax1bp3
|
Tax1 (human T-cell leukemia virus type I) binding protein 3 |
| chr18_-_26854959 | 1.41 |
ENSDART00000057553
|
ch25hl1.1
|
cholesterol 25-hydroxylase like 1, tandem duplicate 1 |
| chr5_+_37978501 | 1.37 |
ENSDART00000012050
|
apoa1a
|
apolipoprotein A-Ia |
| chr24_-_25691020 | 1.36 |
ENSDART00000015391
|
chrnd
|
cholinergic receptor, nicotinic, delta (muscle) |
| chr5_-_71722257 | 1.33 |
ENSDART00000013404
|
ak1
|
adenylate kinase 1 |
| chr22_-_294700 | 1.33 |
ENSDART00000189179
|
CABZ01079179.1
|
|
| chr3_+_8825382 | 1.32 |
ENSDART00000158893
|
si:dkeyp-30d6.2
|
si:dkeyp-30d6.2 |
| chr21_+_25777425 | 1.32 |
ENSDART00000021620
|
cldnd
|
claudin d |
| chr23_+_25305431 | 1.32 |
ENSDART00000143291
|
RPL41
|
si:dkey-151g10.6 |
| chr3_+_15505275 | 1.32 |
ENSDART00000141714
|
nupr1
|
nuclear protein 1 |
| chr13_-_2215213 | 1.31 |
ENSDART00000129773
|
mlip
|
muscular LMNA-interacting protein |
| chr24_-_10021341 | 1.30 |
ENSDART00000137250
|
zgc:173856
|
zgc:173856 |
| chr8_+_52442622 | 1.29 |
ENSDART00000012758
|
zgc:77112
|
zgc:77112 |
| chr6_-_426041 | 1.29 |
ENSDART00000162789
|
fam83fb
|
family with sequence similarity 83, member Fb |
| chr2_+_16780643 | 1.28 |
ENSDART00000125647
ENSDART00000108611 ENSDART00000181245 ENSDART00000163194 |
tfa
|
transferrin-a |
| chr12_-_17712393 | 1.28 |
ENSDART00000143534
ENSDART00000010144 |
pvalb2
|
parvalbumin 2 |
| chr6_+_269204 | 1.28 |
ENSDART00000191678
|
atf4a
|
activating transcription factor 4a |
| chr11_+_45153104 | 1.27 |
ENSDART00000159204
ENSDART00000177585 |
tk1
|
thymidine kinase 1, soluble |
| chr23_+_27675581 | 1.27 |
ENSDART00000127198
|
rps26
|
ribosomal protein S26 |
| chr2_+_16781015 | 1.26 |
ENSDART00000155147
ENSDART00000003845 |
tfa
|
transferrin-a |
| chr12_+_38929663 | 1.26 |
ENSDART00000156334
|
si:dkey-239b22.1
|
si:dkey-239b22.1 |
| chr11_+_25477643 | 1.25 |
ENSDART00000065941
|
opn1lw1
|
opsin 1 (cone pigments), long-wave-sensitive, 1 |
| chr22_+_5118361 | 1.23 |
ENSDART00000168371
ENSDART00000170222 ENSDART00000158846 |
mibp
|
muscle-specific beta 1 integrin binding protein |
| chr4_-_8035520 | 1.21 |
ENSDART00000146146
|
si:ch211-240l19.7
|
si:ch211-240l19.7 |
| chr6_-_54180699 | 1.20 |
ENSDART00000045901
|
rps10
|
ribosomal protein S10 |
| chr13_+_30692669 | 1.19 |
ENSDART00000187818
|
CR762483.1
|
|
| chr2_+_24507597 | 1.19 |
ENSDART00000133109
|
rps28
|
ribosomal protein S28 |
| chr5_-_7834582 | 1.18 |
ENSDART00000162626
ENSDART00000157661 |
pdlim5a
|
PDZ and LIM domain 5a |
| chr7_-_73717082 | 1.18 |
ENSDART00000164301
ENSDART00000082625 |
BX664721.2
|
|
| chr7_+_29951997 | 1.18 |
ENSDART00000173453
|
tpma
|
alpha-tropomyosin |
| chr2_-_6482240 | 1.17 |
ENSDART00000132623
|
rgs13
|
regulator of G protein signaling 13 |
| chr13_-_12645584 | 1.16 |
ENSDART00000176216
|
adh8b
|
alcohol dehydrogenase 8b |
| chr21_+_5560040 | 1.16 |
ENSDART00000163205
|
si:ch211-134a4.6
|
si:ch211-134a4.6 |
| chr8_+_999421 | 1.16 |
ENSDART00000149528
|
fabp1b.1
|
fatty acid binding protein 1b, tandem duplicate 1 |
| chr4_+_306036 | 1.16 |
ENSDART00000103659
|
msgn1
|
mesogenin 1 |
| chr17_+_27456804 | 1.15 |
ENSDART00000017756
ENSDART00000181461 ENSDART00000180178 |
ctsl.1
|
cathepsin L.1 |
| chr5_-_32274383 | 1.13 |
ENSDART00000122889
|
myhz1.3
|
myosin, heavy polypeptide 1.3, skeletal muscle |
| chr3_-_57425961 | 1.11 |
ENSDART00000033716
|
socs3a
|
suppressor of cytokine signaling 3a |
| chr1_-_22678471 | 1.11 |
ENSDART00000128918
|
fgfbp1b
|
fibroblast growth factor binding protein 1b |
| chr21_+_20383837 | 1.10 |
ENSDART00000026430
|
hspb11
|
heat shock protein, alpha-crystallin-related, b11 |
| chr12_+_20641102 | 1.10 |
ENSDART00000152964
|
calcoco2
|
calcium binding and coiled-coil domain 2 |
| chr21_-_11646878 | 1.09 |
ENSDART00000162426
ENSDART00000135937 ENSDART00000131448 ENSDART00000148097 ENSDART00000133443 |
cast
|
calpastatin |
| chr16_-_45917683 | 1.06 |
ENSDART00000184289
|
afp4
|
antifreeze protein type IV |
| chr11_+_3575963 | 1.06 |
ENSDART00000077305
|
si:dkey-33m11.8
|
si:dkey-33m11.8 |
| chr9_+_23900703 | 1.04 |
ENSDART00000127859
|
trim63b
|
tripartite motif containing 63b |
| chr25_+_8356707 | 1.04 |
ENSDART00000153708
|
muc5.1
|
mucin 5.1, oligomeric mucus/gel-forming |
| chr20_-_26846028 | 1.04 |
ENSDART00000136687
|
mylk4b
|
myosin light chain kinase family, member 4b |
| chr5_-_60885935 | 1.04 |
ENSDART00000128350
|
rad51d
|
RAD51 paralog D |
| chr6_+_49052741 | 1.03 |
ENSDART00000011876
|
sycp1
|
synaptonemal complex protein 1 |
| chr5_-_33259079 | 1.01 |
ENSDART00000132223
|
ifitm1
|
interferon induced transmembrane protein 1 |
| chr24_+_3307857 | 1.01 |
ENSDART00000106527
|
gyg1b
|
glycogenin 1b |
| chr22_+_18530395 | 1.01 |
ENSDART00000105415
ENSDART00000183958 |
si:ch211-212d10.1
|
si:ch211-212d10.1 |
| chr24_+_17005647 | 0.99 |
ENSDART00000149149
|
zfx
|
zinc finger protein, X-linked |
| chr15_+_714203 | 0.99 |
ENSDART00000153847
|
si:dkey-7i4.24
|
si:dkey-7i4.24 |
| chr15_-_23814330 | 0.97 |
ENSDART00000153843
|
si:ch211-167j9.5
|
si:ch211-167j9.5 |
| chr8_-_14209852 | 0.97 |
ENSDART00000005359
|
def6a
|
differentially expressed in FDCP 6a homolog (mouse) |
| chr8_-_46486009 | 0.97 |
ENSDART00000140431
|
sult1st9
|
sulfotransferase family 1, cytosolic sulfotransferase 9 |
| chr6_+_41181869 | 0.95 |
ENSDART00000002046
|
opn1mw1
|
opsin 1 (cone pigments), medium-wave-sensitive, 1 |
| chr7_-_73846995 | 0.94 |
ENSDART00000188079
|
FP236812.4
|
|
| chr3_-_40276057 | 0.94 |
ENSDART00000132225
ENSDART00000074737 |
shmt1
|
serine hydroxymethyltransferase 1 (soluble) |
| chr21_-_28523548 | 0.93 |
ENSDART00000077910
|
epdl2
|
ependymin-like 2 |
| chr13_-_15702672 | 0.93 |
ENSDART00000144445
ENSDART00000168950 |
ckba
|
creatine kinase, brain a |
| chr16_+_31542645 | 0.92 |
ENSDART00000163724
|
SLA (1 of many)
|
Src like adaptor |
| chr10_+_17026870 | 0.92 |
ENSDART00000184529
ENSDART00000157480 |
CR855996.2
|
|
| chr24_-_9991153 | 0.91 |
ENSDART00000137794
ENSDART00000106252 ENSDART00000188309 ENSDART00000188266 ENSDART00000188660 ENSDART00000185713 ENSDART00000179773 |
zgc:152652
|
zgc:152652 |
| chr13_+_17694845 | 0.91 |
ENSDART00000079778
|
ifit8
|
interferon-induced protein with tetratricopeptide repeats 8 |
| chr12_+_17042754 | 0.90 |
ENSDART00000066439
|
ch25h
|
cholesterol 25-hydroxylase |
| chr6_+_52918537 | 0.89 |
ENSDART00000174229
|
or137-1
|
odorant receptor, family H, subfamily 137, member 1 |
| chr2_+_55665095 | 0.87 |
ENSDART00000059188
|
klf2b
|
Kruppel-like factor 2b |
| chr2_-_30182353 | 0.86 |
ENSDART00000019149
|
rpl7
|
ribosomal protein L7 |
| chr16_+_1471 | 0.86 |
ENSDART00000181972
|
CABZ01090785.1
|
|
| chr21_-_280769 | 0.85 |
ENSDART00000157753
|
plgrkt
|
plasminogen receptor, C-terminal lysine transmembrane protein |
| chr7_-_73851280 | 0.85 |
ENSDART00000190053
|
FP236812.3
|
|
| chr14_-_33334065 | 0.85 |
ENSDART00000052761
|
rpl39
|
ribosomal protein L39 |
| chr1_-_53880639 | 0.84 |
ENSDART00000010543
|
ltv1
|
LTV1 ribosome biogenesis factor |
| chr15_+_19682013 | 0.84 |
ENSDART00000127368
|
si:dkey-4p15.5
|
si:dkey-4p15.5 |
| chr12_-_36268723 | 0.83 |
ENSDART00000113740
|
kcnj16
|
potassium inwardly-rectifying channel, subfamily J, member 16 |
| chr7_+_5906327 | 0.83 |
ENSDART00000173160
|
zgc:112234
|
zgc:112234 |
| chr23_+_18103080 | 0.82 |
ENSDART00000010270
|
mfsd4ab
|
major facilitator superfamily domain containing 4Ab |
| chr2_+_17181777 | 0.82 |
ENSDART00000112063
|
ptger4c
|
prostaglandin E receptor 4 (subtype EP4) c |
| chr1_+_40199074 | 0.82 |
ENSDART00000179679
ENSDART00000136849 |
si:ch211-113e8.5
|
si:ch211-113e8.5 |
| chr11_+_25472758 | 0.81 |
ENSDART00000011178
|
opn1sw2
|
opsin 1 (cone pigments), short-wave-sensitive 2 |
| chr17_+_51743908 | 0.81 |
ENSDART00000149039
ENSDART00000148869 |
odc1
|
ornithine decarboxylase 1 |
| chr22_-_817479 | 0.81 |
ENSDART00000123487
|
zgc:153675
|
zgc:153675 |
| chr21_-_5077715 | 0.80 |
ENSDART00000081954
|
haus1
|
HAUS augmin-like complex, subunit 1 |
| chr25_+_8407892 | 0.80 |
ENSDART00000153536
|
muc5.2
|
mucin 5.2 |
| chr9_+_48108174 | 0.80 |
ENSDART00000077260
|
cxcr2
|
chemokine (C-X-C motif) receptor 2 |
| chr11_-_24016761 | 0.80 |
ENSDART00000153601
ENSDART00000067817 ENSDART00000170531 |
chia.3
|
chitinase, acidic.3 |
| chr22_+_19264181 | 0.78 |
ENSDART00000192750
ENSDART00000137591 ENSDART00000136197 |
si:dkey-78l4.5
si:dkey-21e2.11
|
si:dkey-78l4.5 si:dkey-21e2.11 |
| chr8_-_46005359 | 0.78 |
ENSDART00000084131
|
fam160b2
|
family with sequence similarity 160, member B2 |
| chr22_-_10165446 | 0.78 |
ENSDART00000142012
|
rbck1
|
RanBP-type and C3HC4-type zinc finger containing 1 |
| chr12_-_20665164 | 0.77 |
ENSDART00000105352
|
gip
|
gastric inhibitory polypeptide |
| chr24_-_9985019 | 0.77 |
ENSDART00000193536
ENSDART00000189595 |
zgc:171977
|
zgc:171977 |
| chr24_-_9979342 | 0.76 |
ENSDART00000138576
ENSDART00000191206 |
zgc:171977
|
zgc:171977 |
| chr22_+_8612462 | 0.76 |
ENSDART00000114586
|
CR450686.1
|
|
| chr7_+_5960491 | 0.75 |
ENSDART00000145370
|
zgc:112234
|
zgc:112234 |
| chr24_+_21621654 | 0.75 |
ENSDART00000002595
|
rpl21
|
ribosomal protein L21 |
| chr16_+_32736588 | 0.74 |
ENSDART00000075191
ENSDART00000168358 |
zgc:172323
|
zgc:172323 |
| chr20_-_9980318 | 0.74 |
ENSDART00000080664
|
ACTC1
|
zgc:86709 |
| chr23_+_11374168 | 0.73 |
ENSDART00000163083
|
chl1a
|
cell adhesion molecule L1-like a |
| chr3_-_6709938 | 0.73 |
ENSDART00000172196
|
atg4db
|
autophagy related 4D, cysteine peptidase b |
| chr7_+_22637515 | 0.72 |
ENSDART00000158698
|
si:dkey-112a7.5
|
si:dkey-112a7.5 |
| chr7_-_7823662 | 0.72 |
ENSDART00000167652
|
cxcl8b.3
|
chemokine (C-X-C motif) ligand 8b, duplicate 3 |
| chr23_+_32011768 | 0.71 |
ENSDART00000053509
|
plagx
|
pleiomorphic adenoma gene X |
| chr16_+_53455638 | 0.71 |
ENSDART00000045792
ENSDART00000154189 |
rbm24b
|
RNA binding motif protein 24b |
| chr15_+_45643787 | 0.71 |
ENSDART00000055995
ENSDART00000157750 |
sagb
|
S-antigen; retina and pineal gland (arrestin) b |
| chr7_-_31441420 | 0.70 |
ENSDART00000075398
|
cilp
|
cartilage intermediate layer protein, nucleotide pyrophosphohydrolase |
| chr2_-_28420415 | 0.70 |
ENSDART00000183857
|
CABZ01056051.1
|
|
| chr3_+_26135502 | 0.69 |
ENSDART00000146979
|
atp2a1
|
ATPase sarcoplasmic/endoplasmic reticulum Ca2+ transporting 1 |
| chr5_-_26181863 | 0.69 |
ENSDART00000098500
|
ccdc125
|
coiled-coil domain containing 125 |
| chr7_-_73845736 | 0.69 |
ENSDART00000193414
|
zgc:173552
|
zgc:173552 |
| chr6_-_346084 | 0.68 |
ENSDART00000162599
|
pde6ha
|
phosphodiesterase 6H, cGMP-specific, cone, gamma, paralog a |
| chr25_+_35132090 | 0.68 |
ENSDART00000154377
|
si:dkey-261m9.6
|
si:dkey-261m9.6 |
| chr13_+_14976108 | 0.67 |
ENSDART00000011520
|
noto
|
notochord homeobox |
| chr1_+_23560216 | 0.67 |
ENSDART00000187824
|
ncapg
|
non-SMC condensin I complex, subunit G |
| chr17_+_14711765 | 0.66 |
ENSDART00000012889
|
cx28.6
|
connexin 28.6 |
| chr25_-_13050959 | 0.66 |
ENSDART00000169041
|
ccl35.1
|
chemokine (C-C motif) ligand 35, duplicate 1 |
| chr19_+_48290786 | 0.66 |
ENSDART00000165986
|
zgc:101785
|
zgc:101785 |
| chr22_+_6096303 | 0.65 |
ENSDART00000139276
|
zgc:171887
|
zgc:171887 |
| chr23_+_19655301 | 0.64 |
ENSDART00000104441
ENSDART00000135269 |
abhd6b
|
abhydrolase domain containing 6b |
| chr21_+_6197223 | 0.64 |
ENSDART00000147716
|
si:dkey-93m18.3
|
si:dkey-93m18.3 |
| chr13_+_40501455 | 0.64 |
ENSDART00000114985
|
hpse2
|
heparanase 2 |
| chr18_-_50799510 | 0.64 |
ENSDART00000174373
|
taldo1
|
transaldolase 1 |
| chr2_+_55665322 | 0.64 |
ENSDART00000183636
ENSDART00000183814 |
klf2b
|
Kruppel-like factor 2b |
| chr6_-_53426773 | 0.62 |
ENSDART00000162791
|
mst1
|
macrophage stimulating 1 |
| chr5_+_1933131 | 0.62 |
ENSDART00000061693
|
si:ch73-55i23.1
|
si:ch73-55i23.1 |
| chr25_+_16312258 | 0.62 |
ENSDART00000064187
|
parvaa
|
parvin, alpha a |
| chr11_+_24002503 | 0.61 |
ENSDART00000164702
|
chia.2
|
chitinase, acidic.2 |
| chr15_-_33834577 | 0.61 |
ENSDART00000163354
|
mmp13b
|
matrix metallopeptidase 13b |
| chr16_-_31598771 | 0.61 |
ENSDART00000016386
|
tg
|
thyroglobulin |
| chr6_-_40449399 | 0.61 |
ENSDART00000103879
|
tatdn2
|
TatD DNase domain containing 2 |
| chr3_+_5575313 | 0.60 |
ENSDART00000134693
ENSDART00000101807 |
si:ch211-106h11.3
|
si:ch211-106h11.3 |
| chr9_-_22272181 | 0.60 |
ENSDART00000113174
|
crygm2d7
|
crystallin, gamma M2d7 |
| chr1_+_55137943 | 0.60 |
ENSDART00000138070
ENSDART00000150510 ENSDART00000133472 ENSDART00000136378 |
mb
|
myoglobin |
| chr5_+_38685089 | 0.59 |
ENSDART00000139743
|
si:dkey-58f10.10
|
si:dkey-58f10.10 |
| chr3_+_1037946 | 0.59 |
ENSDART00000167590
ENSDART00000011111 |
zgc:153921
zgc:153921
|
zgc:153921 zgc:153921 |
| chr13_-_20381485 | 0.59 |
ENSDART00000131351
|
si:ch211-270n8.1
|
si:ch211-270n8.1 |
| chr12_-_1361517 | 0.59 |
ENSDART00000188297
|
LO018020.2
|
|
| chr15_-_18232712 | 0.58 |
ENSDART00000081199
|
wu:fj20b03
|
wu:fj20b03 |
| chr5_-_33035295 | 0.57 |
ENSDART00000061153
|
glipr2
|
GLI pathogenesis-related 2 |
| chr24_-_26854032 | 0.57 |
ENSDART00000087991
|
fndc3bb
|
fibronectin type III domain containing 3Bb |
| chr11_-_45152702 | 0.57 |
ENSDART00000168066
|
afmid
|
arylformamidase |
| chr18_-_46010 | 0.56 |
ENSDART00000052641
|
gatm
|
glycine amidinotransferase (L-arginine:glycine amidinotransferase) |
| chr22_-_22337382 | 0.56 |
ENSDART00000144684
|
si:ch211-129c21.1
|
si:ch211-129c21.1 |
| chr8_+_37111643 | 0.55 |
ENSDART00000061336
|
ren
|
renin |
Network of associatons between targets according to the STRING database.
First level regulatory network of tbp
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 1.7 | GO:1901533 | negative regulation of hematopoietic progenitor cell differentiation(GO:1901533) |
| 0.5 | 1.5 | GO:0003223 | ventricular compact myocardium morphogenesis(GO:0003223) |
| 0.5 | 1.4 | GO:2000009 | regulation of protein localization to cell surface(GO:2000008) negative regulation of protein localization to cell surface(GO:2000009) |
| 0.5 | 0.9 | GO:0048247 | lymphocyte chemotaxis(GO:0048247) |
| 0.4 | 1.3 | GO:0046125 | deoxyribonucleoside metabolic process(GO:0009120) thymidine metabolic process(GO:0046104) pyrimidine deoxyribonucleoside metabolic process(GO:0046125) |
| 0.4 | 2.8 | GO:0046292 | formaldehyde metabolic process(GO:0046292) formaldehyde catabolic process(GO:0046294) |
| 0.4 | 1.2 | GO:0048340 | paraxial mesoderm morphogenesis(GO:0048340) paraxial mesoderm formation(GO:0048341) |
| 0.4 | 1.5 | GO:0090299 | regulation of neural crest formation(GO:0090299) |
| 0.4 | 2.2 | GO:0031394 | neutrophil apoptotic process(GO:0001781) regulation of T-helper 1 type immune response(GO:0002825) positive regulation of T-helper 1 type immune response(GO:0002827) negative regulation of type 2 immune response(GO:0002829) inflammatory cell apoptotic process(GO:0006925) regulation of prostaglandin biosynthetic process(GO:0031392) positive regulation of prostaglandin biosynthetic process(GO:0031394) regulation of phospholipase A2 activity(GO:0032429) interleukin-2 production(GO:0032623) regulation of interleukin-2 production(GO:0032663) positive regulation of interleukin-2 production(GO:0032743) myeloid cell apoptotic process(GO:0033028) regulation of neutrophil apoptotic process(GO:0033029) positive regulation of neutrophil apoptotic process(GO:0033031) regulation of myeloid cell apoptotic process(GO:0033032) positive regulation of myeloid cell apoptotic process(GO:0033034) T-helper 1 type immune response(GO:0042088) positive regulation of CD4-positive, alpha-beta T cell differentiation(GO:0043372) T-helper 1 cell differentiation(GO:0045063) positive regulation of T-helper cell differentiation(GO:0045624) regulation of T-helper 1 cell differentiation(GO:0045625) positive regulation of T-helper 1 cell differentiation(GO:0045627) regulation of T-helper 2 cell differentiation(GO:0045628) negative regulation of T-helper 2 cell differentiation(GO:0045629) positive regulation of fatty acid biosynthetic process(GO:0045723) positive regulation of alpha-beta T cell differentiation(GO:0046638) neutrophil clearance(GO:0097350) negative regulation of phospholipase A2 activity(GO:1900138) positive regulation of CD4-positive, alpha-beta T cell activation(GO:2000516) regulation of unsaturated fatty acid biosynthetic process(GO:2001279) positive regulation of unsaturated fatty acid biosynthetic process(GO:2001280) |
| 0.4 | 1.5 | GO:2000659 | regulation of interleukin-1-mediated signaling pathway(GO:2000659) negative regulation of interleukin-1-mediated signaling pathway(GO:2000660) |
| 0.3 | 1.0 | GO:0042148 | strand invasion(GO:0042148) |
| 0.3 | 1.0 | GO:0051026 | meiotic DNA repair synthesis(GO:0000711) chiasma assembly(GO:0051026) |
| 0.3 | 0.9 | GO:0006565 | L-serine catabolic process(GO:0006565) glycine biosynthetic process from serine(GO:0019264) |
| 0.3 | 1.4 | GO:0010872 | regulation of cholesterol esterification(GO:0010872) positive regulation of cholesterol esterification(GO:0010873) |
| 0.3 | 1.9 | GO:0001561 | fatty acid alpha-oxidation(GO:0001561) |
| 0.3 | 1.3 | GO:0016338 | calcium-independent cell-cell adhesion via plasma membrane cell-adhesion molecules(GO:0016338) |
| 0.3 | 1.0 | GO:0042304 | regulation of fatty acid biosynthetic process(GO:0042304) |
| 0.2 | 2.5 | GO:0019731 | antibacterial humoral response(GO:0019731) |
| 0.2 | 0.7 | GO:0048322 | axial mesoderm formation(GO:0048320) axial mesodermal cell differentiation(GO:0048321) axial mesodermal cell fate commitment(GO:0048322) axial mesodermal cell fate specification(GO:0048327) |
| 0.2 | 0.9 | GO:0010755 | regulation of plasminogen activation(GO:0010755) |
| 0.2 | 0.6 | GO:0006590 | thyroid hormone generation(GO:0006590) |
| 0.2 | 1.6 | GO:0090594 | inflammatory response to wounding(GO:0090594) |
| 0.2 | 0.8 | GO:0002523 | leukocyte migration involved in inflammatory response(GO:0002523) |
| 0.2 | 1.3 | GO:0006172 | ADP biosynthetic process(GO:0006172) purine nucleoside diphosphate biosynthetic process(GO:0009136) purine ribonucleoside diphosphate biosynthetic process(GO:0009180) |
| 0.2 | 2.5 | GO:0006603 | phosphagen metabolic process(GO:0006599) phosphocreatine metabolic process(GO:0006603) phosphagen biosynthetic process(GO:0042396) phosphocreatine biosynthetic process(GO:0046314) |
| 0.2 | 0.6 | GO:0006601 | creatine metabolic process(GO:0006600) creatine biosynthetic process(GO:0006601) |
| 0.2 | 0.7 | GO:0003228 | atrial cardiac muscle tissue development(GO:0003228) |
| 0.2 | 1.9 | GO:0006032 | chitin catabolic process(GO:0006032) |
| 0.2 | 0.7 | GO:0031446 | regulation of twitch skeletal muscle contraction(GO:0014724) regulation of fast-twitch skeletal muscle fiber contraction(GO:0031446) positive regulation of fast-twitch skeletal muscle fiber contraction(GO:0031448) negative regulation of striated muscle contraction(GO:0045988) relaxation of skeletal muscle(GO:0090076) |
| 0.2 | 3.8 | GO:0000028 | ribosomal small subunit assembly(GO:0000028) |
| 0.2 | 0.8 | GO:0034695 | response to prostaglandin E(GO:0034695) cellular response to prostaglandin E stimulus(GO:0071380) |
| 0.2 | 1.5 | GO:0046716 | muscle cell cellular homeostasis(GO:0046716) |
| 0.2 | 0.5 | GO:0065001 | specification of axis polarity(GO:0065001) |
| 0.2 | 0.2 | GO:0060063 | Spemann organizer formation at the embryonic shield(GO:0060063) |
| 0.2 | 8.8 | GO:0070098 | chemokine-mediated signaling pathway(GO:0070098) |
| 0.2 | 0.8 | GO:0097039 | protein linear polyubiquitination(GO:0097039) |
| 0.2 | 1.1 | GO:0097341 | inhibition of cysteine-type endopeptidase activity(GO:0097340) zymogen inhibition(GO:0097341) |
| 0.1 | 1.1 | GO:0048681 | negative regulation of axon regeneration(GO:0048681) negative regulation of neuron projection regeneration(GO:0070571) |
| 0.1 | 0.6 | GO:0009052 | pentose-phosphate shunt, non-oxidative branch(GO:0009052) |
| 0.1 | 1.5 | GO:2000377 | regulation of reactive oxygen species metabolic process(GO:2000377) |
| 0.1 | 0.6 | GO:0010693 | regulation of alkaline phosphatase activity(GO:0010692) negative regulation of alkaline phosphatase activity(GO:0010693) |
| 0.1 | 0.2 | GO:0002138 | retinoic acid biosynthetic process(GO:0002138) |
| 0.1 | 1.3 | GO:0070307 | intrinsic apoptotic signaling pathway in response to endoplasmic reticulum stress(GO:0070059) lens fiber cell development(GO:0070307) lens fiber cell morphogenesis(GO:0070309) |
| 0.1 | 2.3 | GO:0048246 | macrophage chemotaxis(GO:0048246) |
| 0.1 | 0.5 | GO:0006700 | C21-steroid hormone biosynthetic process(GO:0006700) |
| 0.1 | 0.3 | GO:0015695 | organic cation transport(GO:0015695) |
| 0.1 | 0.8 | GO:0000056 | ribosomal small subunit export from nucleus(GO:0000056) |
| 0.1 | 0.5 | GO:0006953 | acute-phase response(GO:0006953) |
| 0.1 | 2.7 | GO:0060030 | dorsal convergence(GO:0060030) |
| 0.1 | 0.3 | GO:0042776 | mitochondrial ATP synthesis coupled proton transport(GO:0042776) |
| 0.1 | 0.3 | GO:0007638 | mechanosensory behavior(GO:0007638) |
| 0.1 | 0.2 | GO:0035994 | response to muscle stretch(GO:0035994) |
| 0.1 | 1.6 | GO:0018149 | peptide cross-linking(GO:0018149) |
| 0.1 | 1.6 | GO:0005979 | regulation of glycogen biosynthetic process(GO:0005979) regulation of glucan biosynthetic process(GO:0010962) |
| 0.1 | 2.2 | GO:0051058 | negative regulation of small GTPase mediated signal transduction(GO:0051058) |
| 0.1 | 3.2 | GO:0018298 | protein-chromophore linkage(GO:0018298) |
| 0.1 | 0.5 | GO:0019441 | tryptophan catabolic process to kynurenine(GO:0019441) |
| 0.1 | 0.5 | GO:0045338 | farnesyl diphosphate metabolic process(GO:0045338) |
| 0.1 | 0.7 | GO:0002031 | G-protein coupled receptor internalization(GO:0002031) |
| 0.1 | 0.3 | GO:1901741 | positive regulation of myoblast fusion(GO:1901741) |
| 0.1 | 0.5 | GO:2000251 | positive regulation of actin cytoskeleton reorganization(GO:2000251) |
| 0.1 | 0.4 | GO:1902262 | apoptotic process involved in patterning of blood vessels(GO:1902262) |
| 0.1 | 0.5 | GO:0006857 | oligopeptide transport(GO:0006857) |
| 0.1 | 0.2 | GO:0034380 | high-density lipoprotein particle assembly(GO:0034380) |
| 0.1 | 0.9 | GO:0000463 | maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000463) |
| 0.1 | 0.2 | GO:0006391 | transcription initiation from mitochondrial promoter(GO:0006391) |
| 0.0 | 0.4 | GO:0060005 | reflex(GO:0060004) vestibular reflex(GO:0060005) |
| 0.0 | 0.9 | GO:0031033 | myosin filament organization(GO:0031033) |
| 0.0 | 1.0 | GO:0046475 | glycerophospholipid catabolic process(GO:0046475) |
| 0.0 | 1.4 | GO:0016126 | sterol biosynthetic process(GO:0016126) |
| 0.0 | 0.2 | GO:0060405 | copulation(GO:0007620) regulation of epinephrine secretion(GO:0014060) positive regulation of epinephrine secretion(GO:0032812) positive regulation of catecholamine secretion(GO:0033605) penile erection(GO:0043084) epinephrine transport(GO:0048241) epinephrine secretion(GO:0048242) regulation of penile erection(GO:0060405) positive regulation of penile erection(GO:0060406) prolactin secretion(GO:0070459) |
| 0.0 | 1.5 | GO:0046890 | regulation of lipid biosynthetic process(GO:0046890) |
| 0.0 | 0.2 | GO:0070920 | regulation of production of small RNA involved in gene silencing by RNA(GO:0070920) regulation of production of miRNAs involved in gene silencing by miRNA(GO:1903798) |
| 0.0 | 1.0 | GO:0009250 | glycogen biosynthetic process(GO:0005978) glucan biosynthetic process(GO:0009250) |
| 0.0 | 0.6 | GO:0016486 | peptide hormone processing(GO:0016486) |
| 0.0 | 0.3 | GO:0034501 | protein localization to kinetochore(GO:0034501) protein localization to chromosome, centromeric region(GO:0071459) |
| 0.0 | 0.2 | GO:0009253 | peptidoglycan metabolic process(GO:0000270) peptidoglycan catabolic process(GO:0009253) |
| 0.0 | 1.0 | GO:0051923 | sulfation(GO:0051923) |
| 0.0 | 3.6 | GO:0031101 | fin regeneration(GO:0031101) |
| 0.0 | 0.6 | GO:0014823 | response to activity(GO:0014823) |
| 0.0 | 0.8 | GO:0010507 | negative regulation of autophagy(GO:0010507) |
| 0.0 | 0.5 | GO:0032088 | negative regulation of NF-kappaB transcription factor activity(GO:0032088) |
| 0.0 | 0.2 | GO:0032732 | positive regulation of interleukin-1 production(GO:0032732) |
| 0.0 | 0.4 | GO:0007198 | adenylate cyclase-inhibiting serotonin receptor signaling pathway(GO:0007198) |
| 0.0 | 0.1 | GO:0045745 | positive regulation of G-protein coupled receptor protein signaling pathway(GO:0045745) |
| 0.0 | 0.3 | GO:0030719 | P granule organization(GO:0030719) |
| 0.0 | 2.0 | GO:0002088 | lens development in camera-type eye(GO:0002088) |
| 0.0 | 0.1 | GO:0097428 | protein maturation by iron-sulfur cluster transfer(GO:0097428) |
| 0.0 | 0.5 | GO:0008156 | negative regulation of DNA replication(GO:0008156) |
| 0.0 | 1.0 | GO:0000079 | regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0000079) regulation of cyclin-dependent protein kinase activity(GO:1904029) |
| 0.0 | 0.6 | GO:0030574 | collagen catabolic process(GO:0030574) multicellular organism catabolic process(GO:0044243) |
| 0.0 | 0.2 | GO:0033387 | putrescine biosynthetic process from ornithine(GO:0033387) |
| 0.0 | 0.6 | GO:0048263 | determination of dorsal identity(GO:0048263) |
| 0.0 | 1.3 | GO:0042074 | cell migration involved in gastrulation(GO:0042074) |
| 0.0 | 1.1 | GO:0034446 | substrate adhesion-dependent cell spreading(GO:0034446) |
| 0.0 | 0.2 | GO:0048672 | positive regulation of collateral sprouting(GO:0048672) |
| 0.0 | 1.0 | GO:0051607 | defense response to virus(GO:0051607) |
| 0.0 | 0.2 | GO:0045116 | protein neddylation(GO:0045116) |
| 0.0 | 0.6 | GO:0030490 | maturation of SSU-rRNA(GO:0030490) |
| 0.0 | 1.1 | GO:0061515 | myeloid cell development(GO:0061515) |
| 0.0 | 1.0 | GO:1990573 | potassium ion import across plasma membrane(GO:1990573) |
| 0.0 | 0.6 | GO:0030282 | bone mineralization(GO:0030282) |
| 0.0 | 0.8 | GO:0007098 | centrosome cycle(GO:0007098) |
| 0.0 | 0.3 | GO:0051209 | release of sequestered calcium ion into cytosol(GO:0051209) |
| 0.0 | 0.5 | GO:0032456 | endocytic recycling(GO:0032456) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 2.6 | GO:0098556 | cytoplasmic side of rough endoplasmic reticulum membrane(GO:0098556) |
| 0.3 | 3.1 | GO:0045095 | keratin filament(GO:0045095) |
| 0.3 | 1.0 | GO:0043073 | germ cell nucleus(GO:0043073) |
| 0.2 | 3.3 | GO:0005751 | mitochondrial respiratory chain complex IV(GO:0005751) |
| 0.2 | 1.5 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.2 | 1.0 | GO:0033063 | Rad51B-Rad51C-Rad51D-XRCC2 complex(GO:0033063) |
| 0.2 | 4.4 | GO:0030667 | secretory granule membrane(GO:0030667) |
| 0.2 | 0.6 | GO:0097124 | cyclin A2-CDK2 complex(GO:0097124) |
| 0.2 | 0.7 | GO:0031673 | H zone(GO:0031673) |
| 0.1 | 2.2 | GO:0016328 | lateral plasma membrane(GO:0016328) |
| 0.1 | 1.2 | GO:0030688 | preribosome, small subunit precursor(GO:0030688) |
| 0.1 | 1.4 | GO:0042627 | chylomicron(GO:0042627) |
| 0.1 | 1.3 | GO:1990589 | ATF4-CREB1 transcription factor complex(GO:1990589) |
| 0.1 | 4.1 | GO:0001750 | photoreceptor outer segment(GO:0001750) |
| 0.1 | 0.8 | GO:0070652 | HAUS complex(GO:0070652) |
| 0.1 | 0.3 | GO:0033391 | chromatoid body(GO:0033391) |
| 0.1 | 0.8 | GO:0071797 | LUBAC complex(GO:0071797) |
| 0.1 | 0.7 | GO:0000796 | condensin complex(GO:0000796) |
| 0.1 | 1.3 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.1 | 0.2 | GO:0034362 | low-density lipoprotein particle(GO:0034362) |
| 0.1 | 2.4 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.0 | 0.3 | GO:0045261 | proton-transporting ATP synthase complex, catalytic core F(1)(GO:0045261) |
| 0.0 | 0.3 | GO:0005790 | smooth endoplasmic reticulum(GO:0005790) |
| 0.0 | 2.5 | GO:0022626 | cytosolic large ribosomal subunit(GO:0022625) cytosolic ribosome(GO:0022626) |
| 0.0 | 1.3 | GO:0016605 | PML body(GO:0016605) |
| 0.0 | 3.1 | GO:0016459 | myosin complex(GO:0016459) |
| 0.0 | 1.2 | GO:0001725 | stress fiber(GO:0001725) contractile actin filament bundle(GO:0097517) |
| 0.0 | 0.5 | GO:0000783 | telomere cap complex(GO:0000782) nuclear telomere cap complex(GO:0000783) nuclear chromosome, telomeric region(GO:0000784) |
| 0.0 | 2.3 | GO:0044815 | nucleosome(GO:0000786) DNA packaging complex(GO:0044815) |
| 0.0 | 0.3 | GO:0000940 | condensed chromosome outer kinetochore(GO:0000940) |
| 0.0 | 0.2 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.0 | 16.6 | GO:0005615 | extracellular space(GO:0005615) |
| 0.0 | 0.0 | GO:0070545 | PeBoW complex(GO:0070545) |
| 0.0 | 0.3 | GO:0043256 | basal lamina(GO:0005605) laminin complex(GO:0043256) |
| 0.0 | 5.4 | GO:0031012 | extracellular matrix(GO:0031012) |
| 0.0 | 0.7 | GO:0005942 | phosphatidylinositol 3-kinase complex(GO:0005942) |
| 0.0 | 0.4 | GO:0035869 | ciliary transition zone(GO:0035869) |
| 0.0 | 0.5 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.0 | 1.1 | GO:0005884 | actin filament(GO:0005884) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 2.9 | GO:0051916 | granulocyte colony-stimulating factor binding(GO:0051916) |
| 0.6 | 2.8 | GO:0004024 | alcohol dehydrogenase activity, zinc-dependent(GO:0004024) S-(hydroxymethyl)glutathione dehydrogenase activity(GO:0051903) |
| 0.4 | 2.2 | GO:0019834 | phospholipase A2 inhibitor activity(GO:0019834) |
| 0.3 | 2.3 | GO:0000254 | C-4 methylsterol oxidase activity(GO:0000254) |
| 0.3 | 1.3 | GO:0005521 | lamin binding(GO:0005521) |
| 0.3 | 0.9 | GO:0004372 | glycine hydroxymethyltransferase activity(GO:0004372) serine binding(GO:0070905) |
| 0.3 | 1.4 | GO:0060228 | phosphatidylcholine-sterol O-acyltransferase activator activity(GO:0060228) |
| 0.3 | 1.3 | GO:0031769 | glucagon receptor binding(GO:0031769) |
| 0.3 | 3.8 | GO:0001594 | trace-amine receptor activity(GO:0001594) |
| 0.2 | 2.3 | GO:0045236 | CXCR chemokine receptor binding(GO:0045236) |
| 0.2 | 2.5 | GO:0004111 | creatine kinase activity(GO:0004111) phosphotransferase activity, nitrogenous group as acceptor(GO:0016775) |
| 0.2 | 1.5 | GO:0048019 | receptor antagonist activity(GO:0048019) |
| 0.2 | 1.3 | GO:0008140 | cAMP response element binding protein binding(GO:0008140) |
| 0.2 | 1.3 | GO:0019136 | deoxynucleoside kinase activity(GO:0019136) |
| 0.2 | 1.2 | GO:0050262 | ribosylnicotinamide kinase activity(GO:0050262) |
| 0.2 | 1.9 | GO:0004568 | chitinase activity(GO:0004568) |
| 0.2 | 0.5 | GO:0004061 | arylformamidase activity(GO:0004061) |
| 0.2 | 0.5 | GO:0071532 | ankyrin repeat binding(GO:0071532) |
| 0.2 | 0.5 | GO:0016713 | cholesterol monooxygenase (side-chain-cleaving) activity(GO:0008386) oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced iron-sulfur protein as one donor, and incorporation of one atom of oxygen(GO:0016713) |
| 0.2 | 0.5 | GO:0004311 | farnesyltranstransferase activity(GO:0004311) |
| 0.2 | 2.6 | GO:0031729 | CCR1 chemokine receptor binding(GO:0031726) CCR4 chemokine receptor binding(GO:0031729) |
| 0.2 | 1.1 | GO:0010859 | calcium-dependent cysteine-type endopeptidase inhibitor activity(GO:0010859) |
| 0.1 | 0.9 | GO:0034632 | retinol transporter activity(GO:0034632) |
| 0.1 | 3.3 | GO:0016675 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors(GO:0016675) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.1 | 1.0 | GO:0004062 | aryl sulfotransferase activity(GO:0004062) |
| 0.1 | 0.6 | GO:0016744 | transferase activity, transferring aldehyde or ketonic groups(GO:0016744) |
| 0.1 | 1.0 | GO:0000400 | four-way junction DNA binding(GO:0000400) |
| 0.1 | 1.3 | GO:0004017 | adenylate kinase activity(GO:0004017) |
| 0.1 | 0.4 | GO:0070513 | death domain binding(GO:0070513) |
| 0.1 | 1.3 | GO:2001069 | glycogen binding(GO:2001069) |
| 0.1 | 0.8 | GO:0004957 | prostaglandin E receptor activity(GO:0004957) |
| 0.1 | 1.0 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 0.1 | 1.6 | GO:0003810 | protein-glutamine gamma-glutamyltransferase activity(GO:0003810) |
| 0.1 | 1.8 | GO:0022848 | acetylcholine-gated cation channel activity(GO:0022848) |
| 0.1 | 1.2 | GO:0005504 | fatty acid binding(GO:0005504) |
| 0.1 | 1.5 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) alpha-actinin binding(GO:0051393) |
| 0.1 | 1.9 | GO:0048020 | CCR chemokine receptor binding(GO:0048020) |
| 0.1 | 0.5 | GO:0042936 | dipeptide transporter activity(GO:0042936) dipeptide transmembrane transporter activity(GO:0071916) |
| 0.1 | 3.2 | GO:0008020 | G-protein coupled photoreceptor activity(GO:0008020) |
| 0.1 | 0.3 | GO:0005219 | ryanodine-sensitive calcium-release channel activity(GO:0005219) calcium-induced calcium release activity(GO:0048763) |
| 0.1 | 0.2 | GO:0008745 | N-acetylmuramoyl-L-alanine amidase activity(GO:0008745) |
| 0.1 | 1.5 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.1 | 0.6 | GO:0047498 | calcium-dependent phospholipase A2 activity(GO:0047498) |
| 0.1 | 0.2 | GO:0043395 | heparan sulfate proteoglycan binding(GO:0043395) |
| 0.0 | 1.1 | GO:0016888 | endodeoxyribonuclease activity, producing 5'-phosphomonoesters(GO:0016888) |
| 0.0 | 0.1 | GO:0015462 | protein-transmembrane transporting ATPase activity(GO:0015462) |
| 0.0 | 0.8 | GO:0015145 | glucose transmembrane transporter activity(GO:0005355) monosaccharide transmembrane transporter activity(GO:0015145) hexose transmembrane transporter activity(GO:0015149) |
| 0.0 | 0.3 | GO:0004719 | protein-L-isoaspartate (D-aspartate) O-methyltransferase activity(GO:0004719) |
| 0.0 | 0.3 | GO:0016312 | inositol bisphosphate phosphatase activity(GO:0016312) |
| 0.0 | 0.7 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.0 | 1.8 | GO:0016831 | carboxy-lyase activity(GO:0016831) |
| 0.0 | 7.6 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.0 | 1.9 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.0 | 0.5 | GO:0001608 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 0.0 | 1.1 | GO:0046935 | 1-phosphatidylinositol-3-kinase regulator activity(GO:0046935) |
| 0.0 | 1.5 | GO:0004601 | peroxidase activity(GO:0004601) |
| 0.0 | 0.5 | GO:0097199 | cysteine-type endopeptidase activity involved in apoptotic signaling pathway(GO:0097199) |
| 0.0 | 0.6 | GO:0019825 | oxygen transporter activity(GO:0005344) oxygen binding(GO:0019825) |
| 0.0 | 0.3 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 0.0 | 0.6 | GO:0070001 | aspartic-type endopeptidase activity(GO:0004190) aspartic-type peptidase activity(GO:0070001) |
| 0.0 | 1.0 | GO:0005242 | inward rectifier potassium channel activity(GO:0005242) |
| 0.0 | 6.0 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.0 | 0.6 | GO:0005520 | insulin-like growth factor binding(GO:0005520) |
| 0.0 | 0.3 | GO:0015101 | organic cation transmembrane transporter activity(GO:0015101) |
| 0.0 | 0.1 | GO:0036402 | proteasome-activating ATPase activity(GO:0036402) |
| 0.0 | 0.4 | GO:0043176 | amine binding(GO:0043176) serotonin binding(GO:0051378) |
| 0.0 | 0.2 | GO:0052650 | NADP-retinol dehydrogenase activity(GO:0052650) |
| 0.0 | 0.4 | GO:0004622 | lysophospholipase activity(GO:0004622) |
| 0.0 | 0.2 | GO:0042805 | actinin binding(GO:0042805) |
| 0.0 | 0.4 | GO:0008253 | 5'-nucleotidase activity(GO:0008253) |
| 0.0 | 1.2 | GO:0016706 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, 2-oxoglutarate as one donor, and incorporation of one atom each of oxygen into both donors(GO:0016706) |
| 0.0 | 0.6 | GO:0016769 | transferase activity, transferring nitrogenous groups(GO:0016769) |
| 0.0 | 1.0 | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity(GO:0016538) |
| 0.0 | 0.1 | GO:0070137 | ubiquitin-like protein-specific endopeptidase activity(GO:0070137) SUMO-specific endopeptidase activity(GO:0070139) |
| 0.0 | 0.1 | GO:0032052 | bile acid binding(GO:0032052) |
| 0.0 | 0.4 | GO:0004438 | phosphatidylinositol-3-phosphatase activity(GO:0004438) |
| 0.0 | 2.3 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.0 | 1.9 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.0 | 0.2 | GO:0005163 | nerve growth factor receptor binding(GO:0005163) neurotrophin receptor binding(GO:0005165) |
| 0.0 | 2.9 | GO:0046982 | protein heterodimerization activity(GO:0046982) |
| 0.0 | 0.4 | GO:0008381 | mechanically-gated ion channel activity(GO:0008381) mechanically gated channel activity(GO:0022833) |
| 0.0 | 0.4 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.0 | 1.0 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.0 | 0.2 | GO:0008301 | DNA binding, bending(GO:0008301) |
| 0.0 | 0.1 | GO:0047555 | 3',5'-cyclic-GMP phosphodiesterase activity(GO:0047555) |
| 0.0 | 0.2 | GO:0030544 | Hsp70 protein binding(GO:0030544) |
| 0.0 | 0.3 | GO:0017147 | Wnt-protein binding(GO:0017147) |
| 0.0 | 0.2 | GO:0097602 | cullin family protein binding(GO:0097602) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.5 | SA MMP CYTOKINE CONNECTION | Cytokines can induce activation of matrix metalloproteinases, which degrade extracellular matrix. |
| 0.2 | 1.7 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.1 | 3.1 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.1 | 0.5 | ST STAT3 PATHWAY | STAT3 Pathway |
| 0.0 | 2.6 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.0 | 0.6 | SA REG CASCADE OF CYCLIN EXPR | Expression of cyclins regulates progression through the cell cycle by activating cyclin-dependent kinases. |
| 0.0 | 0.8 | ST GRANULE CELL SURVIVAL PATHWAY | Granule Cell Survival Pathway is a specific case of more general PAC1 Receptor Pathway. |
| 0.0 | 1.7 | PID SMAD2 3NUCLEAR PATHWAY | Regulation of nuclear SMAD2/3 signaling |
| 0.0 | 0.5 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.0 | 0.4 | PID TXA2PATHWAY | Thromboxane A2 receptor signaling |
| 0.0 | 2.0 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 0.5 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.0 | 0.5 | PID P53 REGULATION PATHWAY | p53 pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 2.1 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.2 | 0.8 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.2 | 1.4 | REACTOME PRESYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | Genes involved in Presynaptic nicotinic acetylcholine receptors |
| 0.1 | 1.7 | REACTOME ACTIVATION OF IRF3 IRF7 MEDIATED BY TBK1 IKK EPSILON | Genes involved in Activation of IRF3/IRF7 mediated by TBK1/IKK epsilon |
| 0.1 | 0.5 | REACTOME HIGHLY CALCIUM PERMEABLE POSTSYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | Genes involved in Highly calcium permeable postsynaptic nicotinic acetylcholine receptors |
| 0.1 | 5.1 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.1 | 0.3 | REACTOME ABACAVIR TRANSPORT AND METABOLISM | Genes involved in Abacavir transport and metabolism |
| 0.1 | 0.9 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS | Genes involved in Synthesis of bile acids and bile salts |
| 0.1 | 1.3 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.1 | 1.0 | REACTOME IMMUNOREGULATORY INTERACTIONS BETWEEN A LYMPHOID AND A NON LYMPHOID CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 0.1 | 0.7 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GIP | Genes involved in Synthesis, Secretion, and Inactivation of Glucose-dependent Insulinotropic Polypeptide (GIP) |
| 0.1 | 0.8 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |
| 0.1 | 3.8 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 0.0 | 1.5 | REACTOME MEIOTIC SYNAPSIS | Genes involved in Meiotic Synapsis |
| 0.0 | 0.5 | REACTOME IL 6 SIGNALING | Genes involved in Interleukin-6 signaling |
| 0.0 | 0.8 | REACTOME INHIBITION OF VOLTAGE GATED CA2 CHANNELS VIA GBETA GAMMA SUBUNITS | Genes involved in Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits |
| 0.0 | 0.9 | REACTOME PYRIMIDINE METABOLISM | Genes involved in Pyrimidine metabolism |
| 0.0 | 0.5 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 0.0 | 1.5 | REACTOME IL1 SIGNALING | Genes involved in Interleukin-1 signaling |
| 0.0 | 0.7 | REACTOME PLATELET CALCIUM HOMEOSTASIS | Genes involved in Platelet calcium homeostasis |
| 0.0 | 0.6 | REACTOME HS GAG DEGRADATION | Genes involved in HS-GAG degradation |
| 0.0 | 0.6 | REACTOME CYCLIN A B1 ASSOCIATED EVENTS DURING G2 M TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 0.0 | 0.2 | REACTOME OPSINS | Genes involved in Opsins |
| 0.0 | 1.1 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |
| 0.0 | 0.3 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.0 | 0.5 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.0 | 0.1 | REACTOME BILE ACID AND BILE SALT METABOLISM | Genes involved in Bile acid and bile salt metabolism |