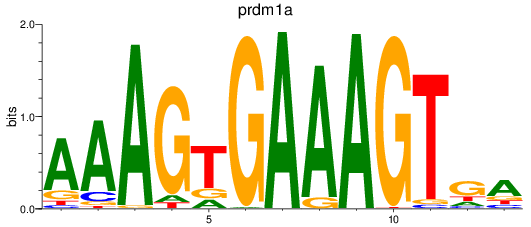
Project
PRJNA207719: Tissue specific transcriptome profiling
Navigation
Downloads
Results for prdm1a
Z-value: 1.71
Transcription factors associated with prdm1a
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
prdm1a
|
ENSDARG00000002445 | PR domain containing 1a, with ZNF domain |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| prdm1a | dr11_v1_chr16_-_7443388_7443388 | 0.92 | 2.8e-02 | Click! |
Activity profile of prdm1a motif
Sorted Z-values of prdm1a motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr5_+_13373593 | 1.34 |
ENSDART00000051668
ENSDART00000183883 |
ccl19a.2
|
chemokine (C-C motif) ligand 19a, tandem duplicate 2 |
| chr25_+_10416583 | 0.89 |
ENSDART00000073907
|
ehf
|
ets homologous factor |
| chr10_+_17776981 | 0.84 |
ENSDART00000141693
|
ccl19b
|
chemokine (C-C motif) ligand 19b |
| chr15_-_47929455 | 0.79 |
ENSDART00000064462
|
psma6l
|
proteasome subunit alpha 6, like |
| chr8_-_50259448 | 0.78 |
ENSDART00000146056
|
nkx3-1
|
NK3 homeobox 1 |
| chr15_+_17752928 | 0.76 |
ENSDART00000155314
|
si:ch211-213d14.2
|
si:ch211-213d14.2 |
| chr6_+_46697710 | 0.71 |
ENSDART00000154969
|
tespa1
|
thymocyte expressed, positive selection associated 1 |
| chr4_-_77216726 | 0.66 |
ENSDART00000099943
|
psmb10
|
proteasome subunit beta 10 |
| chr6_-_40657653 | 0.65 |
ENSDART00000154359
|
ppil1
|
peptidylprolyl isomerase (cyclophilin)-like 1 |
| chr11_-_18705303 | 0.64 |
ENSDART00000059732
|
id1
|
inhibitor of DNA binding 1 |
| chr23_+_642395 | 0.62 |
ENSDART00000186995
|
irf10
|
interferon regulatory factor 10 |
| chr6_-_39313027 | 0.61 |
ENSDART00000012644
|
krt4
|
keratin 4 |
| chr14_+_21159693 | 0.61 |
ENSDART00000192678
|
zgc:136929
|
zgc:136929 |
| chr19_+_7115223 | 0.60 |
ENSDART00000001359
|
psmb12
|
proteasome subunit beta 12 |
| chr1_-_7603734 | 0.60 |
ENSDART00000009315
|
mxb
|
myxovirus (influenza) resistance B |
| chr19_-_7115229 | 0.60 |
ENSDART00000001930
|
psmb13a
|
proteasome subunit beta 13a |
| chr4_+_1461079 | 0.57 |
ENSDART00000082383
|
trim35-14
|
tripartite motif containing 35-14 |
| chr17_-_31058900 | 0.57 |
ENSDART00000134998
ENSDART00000104307 ENSDART00000172721 |
eml1
|
echinoderm microtubule associated protein like 1 |
| chr5_-_38197080 | 0.56 |
ENSDART00000140708
|
si:ch211-284e13.9
|
si:ch211-284e13.9 |
| chr23_+_642001 | 0.54 |
ENSDART00000030643
ENSDART00000124850 |
irf10
|
interferon regulatory factor 10 |
| chr17_-_11439815 | 0.54 |
ENSDART00000130105
|
psma3
|
proteasome subunit alpha 3 |
| chr19_+_18739085 | 0.53 |
ENSDART00000188868
|
spaca4l
|
sperm acrosome associated 4 like |
| chr10_-_45230081 | 0.52 |
ENSDART00000169845
|
pargl
|
poly (ADP-ribose) glycohydrolase, like |
| chr2_-_16254782 | 0.52 |
ENSDART00000133708
|
arhgef4
|
Rho guanine nucleotide exchange factor (GEF) 4 |
| chr1_-_44710937 | 0.50 |
ENSDART00000139036
|
si:dkey-28b4.7
|
si:dkey-28b4.7 |
| chr12_+_17042754 | 0.50 |
ENSDART00000066439
|
ch25h
|
cholesterol 25-hydroxylase |
| chr1_+_31942961 | 0.49 |
ENSDART00000007522
|
anos1a
|
anosmin 1a |
| chr16_-_25535430 | 0.48 |
ENSDART00000077420
|
dtnbp1a
|
dystrobrevin binding protein 1a |
| chr7_+_49695904 | 0.48 |
ENSDART00000183550
ENSDART00000126991 |
ascl1b
|
achaete-scute family bHLH transcription factor 1b |
| chr1_-_31083535 | 0.48 |
ENSDART00000138113
|
ppp1r9alb
|
protein phosphatase 1 regulatory subunit 9A-like B |
| chr16_-_9802998 | 0.46 |
ENSDART00000154217
|
tapbpl
|
TAP binding protein (tapasin)-like |
| chr2_+_42005217 | 0.46 |
ENSDART00000143562
|
gbp2
|
guanylate binding protein 2 |
| chr4_-_76370630 | 0.45 |
ENSDART00000168831
ENSDART00000174313 |
si:ch73-158p21.3
|
si:ch73-158p21.3 |
| chr18_+_8812549 | 0.45 |
ENSDART00000017619
|
impdh1a
|
IMP (inosine 5'-monophosphate) dehydrogenase 1a |
| chr3_+_1942219 | 0.45 |
ENSDART00000114520
|
zgc:165583
|
zgc:165583 |
| chr16_-_51271962 | 0.42 |
ENSDART00000164021
ENSDART00000046420 |
serpinb1l1
|
serpin peptidase inhibitor, clade B (ovalbumin), member 1, like 1 |
| chr3_+_59935606 | 0.42 |
ENSDART00000154157
|
arhgdia
|
Rho GDP dissociation inhibitor (GDI) alpha |
| chr17_+_1063988 | 0.41 |
ENSDART00000160400
|
gchfr
|
GTP cyclohydrolase I feedback regulator |
| chr10_-_45229877 | 0.40 |
ENSDART00000169281
|
pargl
|
poly (ADP-ribose) glycohydrolase, like |
| chr5_-_57723929 | 0.39 |
ENSDART00000144237
|
gig2p
|
grass carp reovirus (GCRV)-induced gene 2p |
| chr1_-_45636362 | 0.39 |
ENSDART00000181757
|
sept3
|
septin 3 |
| chr3_-_8035737 | 0.39 |
ENSDART00000170642
|
trim35-19
|
tripartite motif containing 35-19 |
| chr3_+_32833661 | 0.38 |
ENSDART00000183878
|
cd2bp2
|
CD2 (cytoplasmic tail) binding protein 2 |
| chr4_+_38550788 | 0.38 |
ENSDART00000157412
|
si:ch211-209n20.1
|
si:ch211-209n20.1 |
| chr18_-_16953978 | 0.38 |
ENSDART00000100126
|
akip1
|
A kinase (PRKA) interacting protein 1 |
| chr21_-_26715270 | 0.38 |
ENSDART00000053794
|
banf1
|
barrier to autointegration factor 1 |
| chr24_+_26402110 | 0.37 |
ENSDART00000133684
|
si:ch211-230g15.5
|
si:ch211-230g15.5 |
| chr15_-_25126842 | 0.36 |
ENSDART00000193670
|
vps53
|
vacuolar protein sorting 53 homolog (S. cerevisiae) |
| chr15_+_24572926 | 0.35 |
ENSDART00000155636
ENSDART00000187800 |
dhrs13b
|
dehydrogenase/reductase (SDR family) member 13b |
| chr4_+_25558849 | 0.35 |
ENSDART00000113663
ENSDART00000100755 ENSDART00000111416 ENSDART00000127840 ENSDART00000168618 ENSDART00000111820 ENSDART00000113866 ENSDART00000110107 ENSDART00000111344 ENSDART00000108548 |
zgc:195175
|
zgc:195175 |
| chr18_+_47313899 | 0.35 |
ENSDART00000192389
ENSDART00000189592 ENSDART00000184281 |
barx2
|
BARX homeobox 2 |
| chr24_+_12945803 | 0.35 |
ENSDART00000005105
|
psme1
|
proteasome activator subunit 1 |
| chr11_-_21528056 | 0.35 |
ENSDART00000181626
|
srgap2
|
SLIT-ROBO Rho GTPase activating protein 2 |
| chr3_-_7948799 | 0.35 |
ENSDART00000163714
|
trim35-24
|
tripartite motif containing 35-24 |
| chr18_+_47313715 | 0.34 |
ENSDART00000138806
|
barx2
|
BARX homeobox 2 |
| chr3_+_823084 | 0.34 |
ENSDART00000153693
|
si:ch73-166o21.1
|
si:ch73-166o21.1 |
| chr1_-_8966046 | 0.33 |
ENSDART00000136198
|
socs1b
|
suppressor of cytokine signaling 1b |
| chr20_+_27096430 | 0.33 |
ENSDART00000153358
|
ubr7
|
ubiquitin protein ligase E3 component n-recognin 7 |
| chr25_+_245438 | 0.32 |
ENSDART00000004689
|
zgc:92481
|
zgc:92481 |
| chr25_+_245018 | 0.32 |
ENSDART00000155344
|
zgc:92481
|
zgc:92481 |
| chr19_-_6774871 | 0.32 |
ENSDART00000170952
|
pvrl2l
|
poliovirus receptor-related 2 like |
| chr2_+_42005475 | 0.32 |
ENSDART00000056461
|
gbp2
|
guanylate binding protein 2 |
| chr6_-_58149165 | 0.31 |
ENSDART00000170752
|
tox2
|
TOX high mobility group box family member 2 |
| chr14_+_45566074 | 0.31 |
ENSDART00000133477
ENSDART00000185899 |
zgc:92249
|
zgc:92249 |
| chr3_+_24618012 | 0.31 |
ENSDART00000111997
|
zgc:171506
|
zgc:171506 |
| chr4_+_686157 | 0.30 |
ENSDART00000169748
|
fbxo7
|
F-box protein 7 |
| chr24_-_39610585 | 0.30 |
ENSDART00000066506
|
cox6b1
|
cytochrome c oxidase subunit VIb polypeptide 1 |
| chr16_-_9802449 | 0.29 |
ENSDART00000081208
|
tapbpl
|
TAP binding protein (tapasin)-like |
| chr7_+_11197940 | 0.29 |
ENSDART00000081346
|
cemip
|
cell migration inducing protein, hyaluronan binding |
| chr5_-_26247973 | 0.29 |
ENSDART00000098527
|
erap1b
|
endoplasmic reticulum aminopeptidase 1b |
| chr21_+_30306369 | 0.29 |
ENSDART00000145050
ENSDART00000059420 |
lcp2a
|
lymphocyte cytosolic protein 2a |
| chr5_-_50781623 | 0.28 |
ENSDART00000114950
|
zgc:194908
|
zgc:194908 |
| chr10_-_210703 | 0.28 |
ENSDART00000059476
|
psmg1
|
proteasome (prosome, macropain) assembly chaperone 1 |
| chr13_-_1151410 | 0.28 |
ENSDART00000007231
|
psmb1
|
proteasome subunit beta 1 |
| chr13_-_377256 | 0.28 |
ENSDART00000137398
|
ch1073-291c23.1
|
ch1073-291c23.1 |
| chr5_+_22406672 | 0.27 |
ENSDART00000141385
|
si:dkey-27p18.3
|
si:dkey-27p18.3 |
| chr22_-_18164671 | 0.26 |
ENSDART00000014057
|
rfxank
|
regulatory factor X-associated ankyrin-containing protein |
| chr4_-_17263210 | 0.26 |
ENSDART00000147853
|
lrmp
|
lymphoid-restricted membrane protein |
| chr3_+_832788 | 0.26 |
ENSDART00000193025
|
FP102888.2
|
|
| chr13_-_35892243 | 0.26 |
ENSDART00000002750
ENSDART00000122810 ENSDART00000162399 |
tacc3
|
transforming, acidic coiled-coil containing protein 3 |
| chr5_-_26247671 | 0.26 |
ENSDART00000145187
|
erap1b
|
endoplasmic reticulum aminopeptidase 1b |
| chr7_+_26167420 | 0.26 |
ENSDART00000173941
|
si:ch211-196f2.6
|
si:ch211-196f2.6 |
| chr8_+_39570615 | 0.25 |
ENSDART00000142557
|
lzts1
|
leucine zipper, putative tumor suppressor 1 |
| chr14_+_31493306 | 0.24 |
ENSDART00000138341
|
phf6
|
PHD finger protein 6 |
| chr22_-_36926342 | 0.24 |
ENSDART00000151804
|
si:dkey-37m8.11
|
si:dkey-37m8.11 |
| chr22_+_36719821 | 0.23 |
ENSDART00000149093
|
TRIM35 (1 of many)
|
si:ch1073-324l1.1 |
| chr24_+_39641991 | 0.23 |
ENSDART00000142182
|
luc7l
|
LUC7-like (S. cerevisiae) |
| chr16_-_43344859 | 0.23 |
ENSDART00000058680
|
psma2
|
proteasome subunit alpha 2 |
| chr12_-_2869565 | 0.23 |
ENSDART00000152772
|
zdhhc16b
|
zinc finger, DHHC-type containing 16b |
| chr18_-_12858016 | 0.22 |
ENSDART00000130343
|
parp12a
|
poly (ADP-ribose) polymerase family, member 12a |
| chr10_+_9222476 | 0.22 |
ENSDART00000064973
ENSDART00000145879 |
paqr3b
|
progestin and adipoQ receptor family member IIIb |
| chr19_+_23296616 | 0.21 |
ENSDART00000134567
|
irgf1
|
immunity-related GTPase family, f1 |
| chr24_+_17260329 | 0.21 |
ENSDART00000129554
|
bmi1a
|
bmi1 polycomb ring finger oncogene 1a |
| chr20_-_26936887 | 0.20 |
ENSDART00000160827
|
ftr79
|
finTRIM family, member 79 |
| chr22_+_36902352 | 0.20 |
ENSDART00000036273
|
trim35-20
|
tripartite motif containing 35-20 |
| chr10_+_10210455 | 0.20 |
ENSDART00000144214
|
sh2d3ca
|
SH2 domain containing 3Ca |
| chr2_-_7605121 | 0.20 |
ENSDART00000182099
|
CABZ01021599.1
|
|
| chr7_-_3693303 | 0.19 |
ENSDART00000166949
|
si:ch211-282j17.13
|
si:ch211-282j17.13 |
| chr22_+_36704764 | 0.19 |
ENSDART00000146662
|
trim35-33
|
tripartite motif containing 35-33 |
| chr1_+_24557414 | 0.19 |
ENSDART00000076519
|
dctpp1
|
dCTP pyrophosphatase 1 |
| chr22_-_23253252 | 0.18 |
ENSDART00000175556
|
lhx9
|
LIM homeobox 9 |
| chr16_+_42830152 | 0.18 |
ENSDART00000159730
|
polr3glb
|
polymerase (RNA) III (DNA directed) polypeptide G like b |
| chr7_-_32782430 | 0.18 |
ENSDART00000173808
|
gas2b
|
growth arrest-specific 2b |
| chr23_-_6982966 | 0.18 |
ENSDART00000033774
ENSDART00000169965 |
edem2
|
ER degradation enhancer, mannosidase alpha-like 2 |
| chr9_+_33158191 | 0.17 |
ENSDART00000180786
|
dopey2
|
dopey family member 2 |
| chr14_-_4120636 | 0.17 |
ENSDART00000059230
|
irf2
|
interferon regulatory factor 2 |
| chr1_+_16574312 | 0.17 |
ENSDART00000187067
|
mtus1b
|
microtubule associated tumor suppressor 1b |
| chr23_+_35504824 | 0.17 |
ENSDART00000082647
ENSDART00000159218 |
acp1
|
acid phosphatase 1 |
| chr20_-_6196989 | 0.17 |
ENSDART00000013343
|
b4galt6
|
UDP-Gal:betaGlcNAc beta 1,4- galactosyltransferase, polypeptide 6 |
| chr7_+_36552725 | 0.16 |
ENSDART00000173682
|
chd9
|
chromodomain helicase DNA binding protein 9 |
| chr16_+_13855039 | 0.16 |
ENSDART00000113764
ENSDART00000143983 |
zgc:174888
|
zgc:174888 |
| chr22_-_18164835 | 0.16 |
ENSDART00000143189
|
rfxank
|
regulatory factor X-associated ankyrin-containing protein |
| chr13_+_42011287 | 0.16 |
ENSDART00000131147
|
cyp1b1
|
cytochrome P450, family 1, subfamily B, polypeptide 1 |
| chr3_-_7961226 | 0.16 |
ENSDART00000159801
|
trim35-23
|
tripartite motif containing 35-23 |
| chr23_+_33718602 | 0.16 |
ENSDART00000024695
|
dazap2
|
DAZ associated protein 2 |
| chr18_+_14693682 | 0.16 |
ENSDART00000132249
|
uri1
|
URI1, prefoldin-like chaperone |
| chr23_+_24705424 | 0.16 |
ENSDART00000104029
|
c1qtnf12
|
C1q and TNF related 12 |
| chr15_+_38025319 | 0.15 |
ENSDART00000155000
|
si:ch73-380l3.3
|
si:ch73-380l3.3 |
| chr16_+_42829735 | 0.15 |
ENSDART00000014956
|
polr3glb
|
polymerase (RNA) III (DNA directed) polypeptide G like b |
| chr12_-_4028079 | 0.15 |
ENSDART00000128676
|
si:ch211-180a12.2
|
si:ch211-180a12.2 |
| chr11_-_36350421 | 0.15 |
ENSDART00000141477
|
psma5
|
proteasome subunit alpha 5 |
| chr13_+_11550454 | 0.15 |
ENSDART00000034935
ENSDART00000166908 |
desi2
|
desumoylating isopeptidase 2 |
| chr15_-_2857961 | 0.15 |
ENSDART00000033263
|
ankrd49
|
ankyrin repeat domain 49 |
| chr20_-_4793450 | 0.14 |
ENSDART00000053870
|
galca
|
galactosylceramidase a |
| chr3_+_39663987 | 0.14 |
ENSDART00000184614
ENSDART00000184573 ENSDART00000183127 |
epn2
|
epsin 2 |
| chr22_+_19230434 | 0.14 |
ENSDART00000132929
|
si:dkey-21e2.8
|
si:dkey-21e2.8 |
| chr20_-_26929685 | 0.14 |
ENSDART00000132556
|
ftr79
|
finTRIM family, member 79 |
| chr21_+_30010061 | 0.14 |
ENSDART00000182163
|
ttc1
|
tetratricopeptide repeat domain 1 |
| chr3_-_42016693 | 0.14 |
ENSDART00000184741
|
ttyh3a
|
tweety family member 3a |
| chr13_-_50234122 | 0.14 |
ENSDART00000164338
|
eif2ak2
|
eukaryotic translation initiation factor 2-alpha kinase 2 |
| chr16_-_41488023 | 0.14 |
ENSDART00000169312
|
cmtm6
|
CKLF-like MARVEL transmembrane domain containing 6 |
| chr6_+_38896158 | 0.14 |
ENSDART00000029930
ENSDART00000131347 |
slc48a1b
|
solute carrier family 48 (heme transporter), member 1b |
| chr7_-_3669868 | 0.13 |
ENSDART00000172907
|
si:ch211-282j17.12
|
si:ch211-282j17.12 |
| chr18_-_44935174 | 0.13 |
ENSDART00000081025
|
pex16
|
peroxisomal biogenesis factor 16 |
| chr23_-_33709964 | 0.13 |
ENSDART00000143333
ENSDART00000130338 |
pou6f1
|
POU class 6 homeobox 1 |
| chr20_+_715739 | 0.13 |
ENSDART00000136768
|
myo6a
|
myosin VIa |
| chr3_+_12593558 | 0.13 |
ENSDART00000186891
ENSDART00000159252 |
abca3b
|
ATP-binding cassette, sub-family A (ABC1), member 3b |
| chr3_+_938438 | 0.12 |
ENSDART00000165671
|
irgf4
|
immunity-related GTPase family, f4 |
| chr5_-_64454459 | 0.12 |
ENSDART00000172321
ENSDART00000168030 |
brd3b
|
bromodomain containing 3b |
| chr17_+_45404758 | 0.12 |
ENSDART00000147557
|
tagapa
|
T cell activation RhoGTPase activating protein a |
| chr19_+_42716542 | 0.12 |
ENSDART00000144557
|
clasp2
|
cytoplasmic linker associated protein 2 |
| chr25_+_28693451 | 0.12 |
ENSDART00000148366
|
cnot2
|
CCR4-NOT transcription complex, subunit 2 |
| chr14_+_21686207 | 0.11 |
ENSDART00000034438
|
ran
|
RAN, member RAS oncogene family |
| chr11_-_26040594 | 0.11 |
ENSDART00000144115
|
kaznb
|
kazrin, periplakin interacting protein b |
| chr19_+_2275019 | 0.11 |
ENSDART00000136138
|
itgb8
|
integrin, beta 8 |
| chr4_+_17524330 | 0.10 |
ENSDART00000172169
ENSDART00000021610 |
recql
|
RecQ helicase-like |
| chr8_-_46457233 | 0.10 |
ENSDART00000113214
|
sult1st7
|
sulfotransferase family 1, cytosolic sulfotransferase 7 |
| chr5_+_59494079 | 0.10 |
ENSDART00000148727
|
gtf2ird1
|
GTF2I repeat domain containing 1 |
| chr21_+_40695345 | 0.10 |
ENSDART00000143594
|
ccdc82
|
coiled-coil domain containing 82 |
| chr19_+_12350069 | 0.10 |
ENSDART00000052240
|
psmg2
|
proteasome (prosome, macropain) assembly chaperone 2 |
| chr11_+_19433936 | 0.10 |
ENSDART00000162081
|
prickle2b
|
prickle homolog 2b |
| chr2_+_33051730 | 0.09 |
ENSDART00000177902
ENSDART00000187903 |
rnf220a
|
ring finger protein 220a |
| chr2_+_21486529 | 0.09 |
ENSDART00000047468
|
inhbab
|
inhibin, beta Ab |
| chr15_+_45591669 | 0.09 |
ENSDART00000157459
|
atg16l1
|
ATG16 autophagy related 16-like 1 (S. cerevisiae) |
| chr1_+_44710955 | 0.09 |
ENSDART00000131296
ENSDART00000142187 |
ssrp1b
|
structure specific recognition protein 1b |
| chr4_+_70967639 | 0.09 |
ENSDART00000159230
|
si:dkeyp-80d11.12
|
si:dkeyp-80d11.12 |
| chr1_+_51903413 | 0.09 |
ENSDART00000135861
|
nfixa
|
nuclear factor I/Xa |
| chr13_-_21739142 | 0.09 |
ENSDART00000078460
|
si:dkey-191g9.5
|
si:dkey-191g9.5 |
| chr14_-_4121052 | 0.09 |
ENSDART00000167074
|
irf2
|
interferon regulatory factor 2 |
| chr6_-_6423885 | 0.08 |
ENSDART00000092257
|
si:ch211-194e18.2
|
si:ch211-194e18.2 |
| chr7_-_28148310 | 0.08 |
ENSDART00000044208
|
lmo1
|
LIM domain only 1 |
| chr2_+_30969029 | 0.08 |
ENSDART00000085242
|
lpin2
|
lipin 2 |
| chr15_+_7992906 | 0.08 |
ENSDART00000090790
|
cadm2b
|
cell adhesion molecule 2b |
| chr24_+_39027481 | 0.08 |
ENSDART00000085565
|
capn15
|
calpain 15 |
| chr3_+_6291635 | 0.08 |
ENSDART00000185055
ENSDART00000157707 |
si:ch211-12p12.2
|
si:ch211-12p12.2 |
| chr15_-_1003553 | 0.08 |
ENSDART00000154195
|
si:dkey-77f5.6
|
si:dkey-77f5.6 |
| chr21_-_31143903 | 0.08 |
ENSDART00000111571
|
rap1gap2b
|
RAP1 GTPase activating protein 2b |
| chr6_-_1587291 | 0.08 |
ENSDART00000067592
ENSDART00000178877 |
zgc:123305
|
zgc:123305 |
| chr1_+_58360935 | 0.07 |
ENSDART00000140754
|
si:dkey-222h21.1
|
si:dkey-222h21.1 |
| chr4_+_70791887 | 0.07 |
ENSDART00000164234
|
si:ch211-127b6.2
|
si:ch211-127b6.2 |
| chr23_-_37835794 | 0.07 |
ENSDART00000137358
|
cd40
|
CD40 molecule, TNF receptor superfamily member 5 |
| chr1_-_54938137 | 0.07 |
ENSDART00000147212
|
crtac1a
|
cartilage acidic protein 1a |
| chr14_-_1543834 | 0.07 |
ENSDART00000185876
|
CABZ01109480.1
|
|
| chr10_+_28587446 | 0.07 |
ENSDART00000030138
ENSDART00000137508 |
cblb
|
Cbl proto-oncogene B, E3 ubiquitin protein ligase |
| chr4_-_75869738 | 0.07 |
ENSDART00000170963
|
si:dkey-261j11.2
|
si:dkey-261j11.2 |
| chr23_+_2825940 | 0.07 |
ENSDART00000135781
|
plcg1
|
phospholipase C, gamma 1 |
| chr24_+_38216340 | 0.06 |
ENSDART00000188561
|
CU896602.4
|
|
| chr11_+_37144328 | 0.06 |
ENSDART00000162830
|
wnk2
|
WNK lysine deficient protein kinase 2 |
| chr18_-_6803424 | 0.06 |
ENSDART00000142647
|
si:dkey-266m15.5
|
si:dkey-266m15.5 |
| chr1_-_21599219 | 0.06 |
ENSDART00000148327
|
adamtsl7
|
ADAMTS-like 7 |
| chr5_-_64213253 | 0.06 |
ENSDART00000171711
|
gpsm1a
|
G protein signaling modulator 1a |
| chr18_-_8857137 | 0.06 |
ENSDART00000126331
|
prrt4
|
proline-rich transmembrane protein 4 |
| chr3_-_22366562 | 0.06 |
ENSDART00000129447
|
ifnphi1
|
interferon phi 1 |
| chr3_-_34586403 | 0.06 |
ENSDART00000151515
|
sept9a
|
septin 9a |
| chr6_+_29861288 | 0.06 |
ENSDART00000166782
|
dlg1
|
discs, large homolog 1 (Drosophila) |
| chr6_-_8580857 | 0.06 |
ENSDART00000138858
ENSDART00000041142 |
myh11a
|
myosin, heavy chain 11a, smooth muscle |
| chr17_-_29902187 | 0.06 |
ENSDART00000009104
|
esrrb
|
estrogen-related receptor beta |
| chr23_-_27123171 | 0.06 |
ENSDART00000023218
ENSDART00000146467 |
stat6
|
signal transducer and activator of transcription 6, interleukin-4 induced |
| chr19_+_15485556 | 0.06 |
ENSDART00000079014
|
pdik1l
|
PDLIM1 interacting kinase 1 like |
| chr25_+_8955530 | 0.06 |
ENSDART00000156444
|
si:ch211-256a21.4
|
si:ch211-256a21.4 |
| chr12_+_5190049 | 0.05 |
ENSDART00000126667
|
lgi1b
|
leucine-rich, glioma inactivated 1b |
| chr16_-_9383629 | 0.05 |
ENSDART00000084264
ENSDART00000166958 |
adcy2a
|
adenylate cyclase 2a |
| chr4_-_46948863 | 0.05 |
ENSDART00000168835
|
si:dkey-16p6.1
|
si:dkey-16p6.1 |
| chr11_-_29623380 | 0.05 |
ENSDART00000162587
ENSDART00000193935 ENSDART00000191646 |
chd5
|
chromodomain helicase DNA binding protein 5 |
| chr20_-_20402500 | 0.05 |
ENSDART00000190747
ENSDART00000185065 |
prkchb
|
protein kinase C, eta, b |
| chr16_+_33121106 | 0.05 |
ENSDART00000110195
|
akirin1
|
akirin 1 |
| chr23_-_27479558 | 0.05 |
ENSDART00000013563
|
atf7a
|
activating transcription factor 7a |
| chr18_-_37252036 | 0.04 |
ENSDART00000132230
|
six5
|
SIX homeobox 5 |
| chr2_+_33052169 | 0.04 |
ENSDART00000180008
|
rnf220a
|
ring finger protein 220a |
| chr13_+_26703922 | 0.04 |
ENSDART00000020946
|
fancl
|
Fanconi anemia, complementation group L |
Network of associatons between targets according to the STRING database.
First level regulatory network of prdm1a
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.0 | GO:0050862 | positive regulation of T cell receptor signaling pathway(GO:0050862) |
| 0.1 | 3.8 | GO:0010499 | proteasomal ubiquitin-independent protein catabolic process(GO:0010499) |
| 0.1 | 0.5 | GO:0060155 | secretory granule organization(GO:0033363) platelet dense granule organization(GO:0060155) |
| 0.1 | 1.3 | GO:0019885 | antigen processing and presentation of endogenous peptide antigen(GO:0002483) antigen processing and presentation of endogenous antigen(GO:0019883) antigen processing and presentation of endogenous peptide antigen via MHC class I(GO:0019885) |
| 0.1 | 0.3 | GO:0035046 | pronuclear migration(GO:0035046) |
| 0.1 | 0.9 | GO:1900407 | regulation of cellular response to oxidative stress(GO:1900407) |
| 0.1 | 0.3 | GO:2001223 | negative regulation of neuron migration(GO:2001223) |
| 0.1 | 2.7 | GO:0048247 | lymphocyte chemotaxis(GO:0048247) |
| 0.1 | 0.2 | GO:0009211 | pyrimidine deoxyribonucleoside triphosphate metabolic process(GO:0009211) |
| 0.1 | 0.2 | GO:0097376 | interneuron axon guidance(GO:0097376) spinal cord interneuron axon guidance(GO:0097377) dorsal spinal cord interneuron axon guidance(GO:0097378) |
| 0.1 | 0.3 | GO:0060330 | regulation of response to interferon-gamma(GO:0060330) regulation of interferon-gamma-mediated signaling pathway(GO:0060334) |
| 0.1 | 0.6 | GO:0036368 | cone photoresponse recovery(GO:0036368) |
| 0.0 | 0.4 | GO:0046037 | GMP biosynthetic process(GO:0006177) GMP metabolic process(GO:0046037) |
| 0.0 | 0.2 | GO:0046324 | regulation of glucose import(GO:0046324) |
| 0.0 | 0.6 | GO:0098884 | postsynaptic neurotransmitter receptor internalization(GO:0098884) |
| 0.0 | 0.1 | GO:0006404 | RNA import into nucleus(GO:0006404) snRNA import into nucleus(GO:0061015) |
| 0.0 | 0.5 | GO:0048922 | posterior lateral line neuromast deposition(GO:0048922) |
| 0.0 | 0.4 | GO:0043248 | proteasome assembly(GO:0043248) chaperone-mediated protein complex assembly(GO:0051131) |
| 0.0 | 0.1 | GO:0002714 | response to protozoan(GO:0001562) positive regulation of B cell mediated immunity(GO:0002714) positive regulation of immunoglobulin mediated immune response(GO:0002891) TRIF-dependent toll-like receptor signaling pathway(GO:0035666) defense response to protozoan(GO:0042832) regulation of isotype switching(GO:0045191) positive regulation of isotype switching(GO:0045830) |
| 0.0 | 0.5 | GO:0000731 | DNA synthesis involved in DNA repair(GO:0000731) |
| 0.0 | 0.9 | GO:0009225 | nucleotide-sugar metabolic process(GO:0009225) |
| 0.0 | 0.4 | GO:1901222 | regulation of NIK/NF-kappaB signaling(GO:1901222) |
| 0.0 | 0.1 | GO:0035477 | regulation of angioblast cell migration involved in selective angioblast sprouting(GO:0035477) |
| 0.0 | 0.6 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.0 | 0.5 | GO:0007286 | spermatid development(GO:0007286) spermatid differentiation(GO:0048515) |
| 0.0 | 0.1 | GO:1902745 | regulation of macrophage chemotaxis(GO:0010758) positive regulation of macrophage chemotaxis(GO:0010759) skeletal muscle cell proliferation(GO:0014856) regulation of skeletal muscle cell proliferation(GO:0014857) mononuclear cell migration(GO:0071674) regulation of mononuclear cell migration(GO:0071675) positive regulation of lamellipodium organization(GO:1902745) |
| 0.0 | 0.1 | GO:0019079 | viral genome replication(GO:0019079) negative stranded viral RNA replication(GO:0039689) viral RNA genome replication(GO:0039694) RNA replication(GO:0039703) multi-organism biosynthetic process(GO:0044034) |
| 0.0 | 0.1 | GO:0046477 | glycosylceramide catabolic process(GO:0046477) |
| 0.0 | 0.1 | GO:1901678 | heme transport(GO:0015886) iron coordination entity transport(GO:1901678) |
| 0.0 | 0.2 | GO:0036353 | histone H2A-K119 monoubiquitination(GO:0036353) |
| 0.0 | 0.6 | GO:0032922 | circadian regulation of gene expression(GO:0032922) |
| 0.0 | 0.1 | GO:0035479 | angioblast cell migration from lateral mesoderm to midline(GO:0035479) |
| 0.0 | 0.0 | GO:0042148 | strand invasion(GO:0042148) |
| 0.0 | 0.2 | GO:0042572 | retinol metabolic process(GO:0042572) |
| 0.0 | 1.1 | GO:0050673 | epithelial cell proliferation(GO:0050673) |
| 0.0 | 0.1 | GO:0046685 | response to arsenic-containing substance(GO:0046685) |
| 0.0 | 0.1 | GO:2000036 | regulation of stem cell population maintenance(GO:2000036) |
| 0.0 | 0.0 | GO:0045649 | regulation of macrophage differentiation(GO:0045649) |
| 0.0 | 0.1 | GO:0007175 | negative regulation of epidermal growth factor-activated receptor activity(GO:0007175) |
| 0.0 | 0.3 | GO:0006383 | transcription from RNA polymerase III promoter(GO:0006383) |
| 0.0 | 0.2 | GO:0051764 | actin crosslink formation(GO:0051764) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.7 | GO:0019773 | proteasome core complex, alpha-subunit complex(GO:0019773) |
| 0.1 | 2.1 | GO:0005839 | proteasome core complex(GO:0005839) |
| 0.1 | 0.5 | GO:0043625 | delta DNA polymerase complex(GO:0043625) |
| 0.1 | 0.3 | GO:0008537 | proteasome activator complex(GO:0008537) |
| 0.1 | 0.4 | GO:0000938 | GARP complex(GO:0000938) |
| 0.1 | 0.6 | GO:0045095 | keratin filament(GO:0045095) |
| 0.0 | 0.9 | GO:0044327 | dendritic spine head(GO:0044327) |
| 0.0 | 0.5 | GO:0033162 | melanosome membrane(GO:0033162) chitosome(GO:0045009) |
| 0.0 | 0.6 | GO:0035102 | PRC1 complex(GO:0035102) |
| 0.0 | 0.4 | GO:0005682 | U5 snRNP(GO:0005682) |
| 0.0 | 0.3 | GO:0097431 | mitotic spindle pole(GO:0097431) |
| 0.0 | 0.2 | GO:0005685 | U1 snRNP(GO:0005685) |
| 0.0 | 0.3 | GO:0005666 | DNA-directed RNA polymerase III complex(GO:0005666) |
| 0.0 | 0.4 | GO:0099569 | presynaptic cytoskeleton(GO:0099569) |
| 0.0 | 0.7 | GO:0072686 | mitotic spindle(GO:0072686) |
| 0.0 | 0.1 | GO:0030128 | clathrin coat of endocytic vesicle(GO:0030128) clathrin-coated endocytic vesicle membrane(GO:0030669) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.9 | GO:0004649 | poly(ADP-ribose) glycohydrolase activity(GO:0004649) |
| 0.1 | 2.1 | GO:0070003 | threonine-type endopeptidase activity(GO:0004298) threonine-type peptidase activity(GO:0070003) |
| 0.1 | 0.4 | GO:0005094 | Rho GDP-dissociation inhibitor activity(GO:0005094) |
| 0.1 | 0.4 | GO:0003938 | IMP dehydrogenase activity(GO:0003938) |
| 0.1 | 0.5 | GO:0000254 | C-4 methylsterol oxidase activity(GO:0000254) |
| 0.1 | 0.3 | GO:0061133 | endopeptidase activator activity(GO:0061133) |
| 0.1 | 0.2 | GO:0008489 | UDP-galactose:glucosylceramide beta-1,4-galactosyltransferase activity(GO:0008489) |
| 0.1 | 2.2 | GO:0048020 | CCR chemokine receptor binding(GO:0048020) |
| 0.0 | 0.1 | GO:0036310 | annealing helicase activity(GO:0036310) |
| 0.0 | 0.1 | GO:0004694 | eukaryotic translation initiation factor 2alpha kinase activity(GO:0004694) |
| 0.0 | 0.2 | GO:0004726 | non-membrane spanning protein tyrosine phosphatase activity(GO:0004726) |
| 0.0 | 0.2 | GO:0047429 | nucleoside-triphosphate diphosphatase activity(GO:0047429) |
| 0.0 | 0.2 | GO:0070330 | aromatase activity(GO:0070330) |
| 0.0 | 0.3 | GO:0070628 | proteasome binding(GO:0070628) |
| 0.0 | 0.1 | GO:0047690 | aspartyltransferase activity(GO:0047690) |
| 0.0 | 0.5 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.0 | 0.1 | GO:0005132 | type I interferon receptor binding(GO:0005132) |
| 0.0 | 0.1 | GO:0015232 | heme transporter activity(GO:0015232) |
| 0.0 | 0.2 | GO:0015924 | mannosyl-oligosaccharide 1,2-alpha-mannosidase activity(GO:0004571) mannosyl-oligosaccharide mannosidase activity(GO:0015924) |
| 0.0 | 0.5 | GO:0004675 | transmembrane receptor protein serine/threonine kinase activity(GO:0004675) |
| 0.0 | 0.5 | GO:0070006 | metalloaminopeptidase activity(GO:0070006) |
| 0.0 | 0.1 | GO:0072320 | volume-sensitive chloride channel activity(GO:0072320) |
| 0.0 | 0.3 | GO:0031267 | small GTPase binding(GO:0031267) |
| 0.0 | 0.6 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
| 0.0 | 0.6 | GO:0003755 | peptidyl-prolyl cis-trans isomerase activity(GO:0003755) |
| 0.0 | 0.6 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.0 | 0.2 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 0.0 | 0.1 | GO:0004062 | aryl sulfotransferase activity(GO:0004062) |
| 0.0 | 0.3 | GO:0046935 | 1-phosphatidylinositol-3-kinase regulator activity(GO:0046935) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.6 | PID IL3 PATHWAY | IL3-mediated signaling events |
| 0.0 | 0.5 | PID HDAC CLASSII PATHWAY | Signaling events mediated by HDAC Class II |
| 0.0 | 0.4 | PID THROMBIN PAR1 PATHWAY | PAR1-mediated thrombin signaling events |
| 0.0 | 0.5 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.0 | 0.2 | PID EPHA2 FWD PATHWAY | EPHA2 forward signaling |
| 0.0 | 0.5 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.0 | 0.3 | PID AURORA A PATHWAY | Aurora A signaling |
| 0.0 | 0.6 | PID MET PATHWAY | Signaling events mediated by Hepatocyte Growth Factor Receptor (c-Met) |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | REACTOME INTEGRATION OF PROVIRUS | Genes involved in Integration of provirus |
| 0.1 | 0.4 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |
| 0.1 | 2.1 | REACTOME CROSS PRESENTATION OF SOLUBLE EXOGENOUS ANTIGENS ENDOSOMES | Genes involved in Cross-presentation of soluble exogenous antigens (endosomes) |
| 0.0 | 0.5 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS | Genes involved in Synthesis of bile acids and bile salts |
| 0.0 | 1.0 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 0.0 | 0.2 | REACTOME ENDOGENOUS STEROLS | Genes involved in Endogenous sterols |
| 0.0 | 0.1 | REACTOME ROLE OF SECOND MESSENGERS IN NETRIN1 SIGNALING | Genes involved in Role of second messengers in netrin-1 signaling |
| 0.0 | 0.3 | REACTOME RNA POL III CHAIN ELONGATION | Genes involved in RNA Polymerase III Chain Elongation |
| 0.0 | 0.6 | REACTOME NRAGE SIGNALS DEATH THROUGH JNK | Genes involved in NRAGE signals death through JNK |
| 0.0 | 0.4 | REACTOME P75 NTR RECEPTOR MEDIATED SIGNALLING | Genes involved in p75 NTR receptor-mediated signalling |