Project
PRJNA207719: Tissue specific transcriptome profiling
Navigation
Downloads
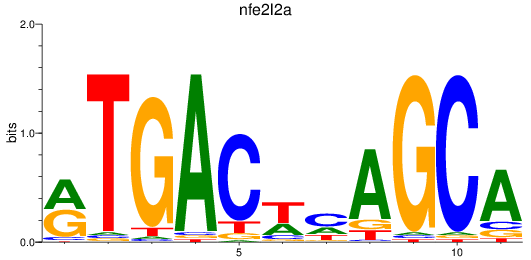
Results for nfe2l2a
Z-value: 1.35
Transcription factors associated with nfe2l2a
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
nfe2l2a
|
ENSDARG00000042824 | nuclear factor, erythroid 2-like 2a |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| nfe2l2a | dr11_v1_chr9_+_1654284_1654284 | -0.19 | 7.6e-01 | Click! |
Activity profile of nfe2l2a motif
Sorted Z-values of nfe2l2a motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr19_-_7450796 | 1.86 |
ENSDART00000104750
|
mllt11
|
MLLT11, transcription factor 7 cofactor |
| chr10_-_7974155 | 1.25 |
ENSDART00000147368
ENSDART00000075524 |
osbp2
|
oxysterol binding protein 2 |
| chr23_-_15878879 | 1.16 |
ENSDART00000010119
|
eef1a2
|
eukaryotic translation elongation factor 1 alpha 2 |
| chr13_+_35746440 | 1.11 |
ENSDART00000187859
|
gpr75
|
G protein-coupled receptor 75 |
| chr22_-_13857729 | 0.91 |
ENSDART00000177971
|
s100b
|
S100 calcium binding protein, beta (neural) |
| chr19_-_6385594 | 0.90 |
ENSDART00000104950
|
atp1a3a
|
ATPase Na+/K+ transporting subunit alpha 3a |
| chr18_+_1703984 | 0.88 |
ENSDART00000114010
|
slitrk3a
|
SLIT and NTRK-like family, member 3a |
| chr15_+_37559570 | 0.80 |
ENSDART00000085522
|
hspb6
|
heat shock protein, alpha-crystallin-related, b6 |
| chr2_+_9822319 | 0.78 |
ENSDART00000144078
ENSDART00000144371 |
anxa13l
|
annexin A13, like |
| chr1_-_40227166 | 0.78 |
ENSDART00000146680
|
si:ch211-113e8.3
|
si:ch211-113e8.3 |
| chr16_-_27640995 | 0.76 |
ENSDART00000019658
|
nacad
|
NAC alpha domain containing |
| chr13_-_21688176 | 0.75 |
ENSDART00000063825
|
sprn
|
shadow of prion protein |
| chr18_-_46763170 | 0.75 |
ENSDART00000171880
|
dner
|
delta/notch-like EGF repeat containing |
| chr11_-_9948487 | 0.72 |
ENSDART00000189677
ENSDART00000113171 |
nlgn1
|
neuroligin 1 |
| chr15_-_16679517 | 0.72 |
ENSDART00000177384
|
caln1
|
calneuron 1 |
| chr24_+_24923166 | 0.71 |
ENSDART00000065288
|
pcyt1ba
|
phosphate cytidylyltransferase 1, choline, beta a |
| chr1_-_53750522 | 0.70 |
ENSDART00000190755
|
akt3b
|
v-akt murine thymoma viral oncogene homolog 3b |
| chr8_+_24861264 | 0.70 |
ENSDART00000099607
|
slc6a17
|
solute carrier family 6 (neutral amino acid transporter), member 17 |
| chr2_+_9821757 | 0.70 |
ENSDART00000018408
ENSDART00000141227 ENSDART00000144681 ENSDART00000148227 |
anxa13l
|
annexin A13, like |
| chr8_-_22660678 | 0.70 |
ENSDART00000181258
|
iqsec2a
|
IQ motif and Sec7 domain 2a |
| chr5_-_31928913 | 0.63 |
ENSDART00000142919
|
ssh1b
|
slingshot protein phosphatase 1b |
| chr1_+_41690402 | 0.62 |
ENSDART00000177298
|
fbxo41
|
F-box protein 41 |
| chr24_-_24146875 | 0.61 |
ENSDART00000173052
|
map7d2b
|
MAP7 domain containing 2b |
| chr4_+_13696537 | 0.61 |
ENSDART00000109195
ENSDART00000122041 ENSDART00000192554 |
nrcama
|
neuronal cell adhesion molecule a |
| chr23_+_23183449 | 0.60 |
ENSDART00000132296
|
klhl17
|
kelch-like family member 17 |
| chr15_-_16070731 | 0.58 |
ENSDART00000122099
|
dynll2a
|
dynein, light chain, LC8-type 2a |
| chr8_-_34051548 | 0.57 |
ENSDART00000105204
|
pbx3b
|
pre-B-cell leukemia homeobox 3b |
| chr14_-_17588345 | 0.57 |
ENSDART00000143486
|
selenot2
|
selenoprotein T, 2 |
| chr3_-_21280373 | 0.57 |
ENSDART00000003939
|
syngr1a
|
synaptogyrin 1a |
| chr10_+_29963518 | 0.56 |
ENSDART00000011317
ENSDART00000099964 ENSDART00000182990 ENSDART00000113912 |
ntm
|
neurotrimin |
| chr5_-_71460556 | 0.56 |
ENSDART00000108804
|
brinp1
|
bone morphogenetic protein/retinoic acid inducible neural-specific 1 |
| chr20_+_38724575 | 0.56 |
ENSDART00000015095
ENSDART00000152972 |
uts1
|
urotensin 1 |
| chr24_+_41940299 | 0.56 |
ENSDART00000022349
|
epb41l3a
|
erythrocyte membrane protein band 4.1-like 3a |
| chr6_-_35446110 | 0.55 |
ENSDART00000058773
|
rgs16
|
regulator of G protein signaling 16 |
| chr4_+_6572364 | 0.54 |
ENSDART00000122574
|
ppp1r3aa
|
protein phosphatase 1, regulatory subunit 3Aa |
| chr3_+_17846890 | 0.54 |
ENSDART00000193384
|
znf385c
|
zinc finger protein 385C |
| chr18_-_19405616 | 0.54 |
ENSDART00000191290
ENSDART00000090855 |
megf11
|
multiple EGF-like-domains 11 |
| chr6_+_21395051 | 0.53 |
ENSDART00000017774
|
cacng5a
|
calcium channel, voltage-dependent, gamma subunit 5a |
| chr16_-_17162485 | 0.53 |
ENSDART00000123011
|
iffo1b
|
intermediate filament family orphan 1b |
| chr24_+_26039464 | 0.53 |
ENSDART00000131017
|
tnk2a
|
tyrosine kinase, non-receptor, 2a |
| chr16_+_25259313 | 0.52 |
ENSDART00000058938
|
fbxo32
|
F-box protein 32 |
| chr23_+_19558574 | 0.50 |
ENSDART00000137811
|
atp6ap1lb
|
ATPase H+ transporting accessory protein 1 like b |
| chr23_-_29376859 | 0.49 |
ENSDART00000146411
|
sst6
|
somatostatin 6 |
| chr8_-_34052019 | 0.49 |
ENSDART00000040126
ENSDART00000159208 ENSDART00000048994 ENSDART00000098822 |
pbx3b
|
pre-B-cell leukemia homeobox 3b |
| chr14_-_48588422 | 0.48 |
ENSDART00000161147
|
si:ch211-154c21.1
|
si:ch211-154c21.1 |
| chr13_-_226109 | 0.48 |
ENSDART00000161705
ENSDART00000172744 ENSDART00000163902 ENSDART00000158208 |
rtn4b
|
reticulon 4b |
| chr17_+_27176243 | 0.48 |
ENSDART00000162527
|
si:ch211-160f23.7
|
si:ch211-160f23.7 |
| chr19_-_17658160 | 0.47 |
ENSDART00000151766
ENSDART00000170790 ENSDART00000186678 ENSDART00000188045 ENSDART00000176980 ENSDART00000166313 ENSDART00000188589 |
thrb
|
thyroid hormone receptor beta |
| chr7_-_52842605 | 0.46 |
ENSDART00000083002
|
map1aa
|
microtubule-associated protein 1Aa |
| chr13_-_226393 | 0.45 |
ENSDART00000172677
|
rtn4b
|
reticulon 4b |
| chr21_+_9576176 | 0.45 |
ENSDART00000161289
ENSDART00000159899 ENSDART00000162834 |
mapk10
|
mitogen-activated protein kinase 10 |
| chr2_-_31800521 | 0.43 |
ENSDART00000112763
|
retreg1
|
reticulophagy regulator 1 |
| chr15_-_25518084 | 0.42 |
ENSDART00000158594
|
hif1al
|
hypoxia-inducible factor 1, alpha subunit, like |
| chr9_+_11202552 | 0.41 |
ENSDART00000151931
|
asic4a
|
acid-sensing (proton-gated) ion channel family member 4a |
| chr7_-_32833153 | 0.40 |
ENSDART00000099871
ENSDART00000099872 |
slc17a6b
|
solute carrier family 17 (vesicular glutamate transporter), member 6b |
| chr14_-_30490763 | 0.40 |
ENSDART00000193166
ENSDART00000183471 ENSDART00000087859 |
micu3b
|
mitochondrial calcium uptake family, member 3b |
| chr20_-_39271844 | 0.39 |
ENSDART00000192708
|
clu
|
clusterin |
| chr5_-_40297003 | 0.39 |
ENSDART00000097526
|
hspb3
|
heat shock protein, alpha-crystallin-related, b3 |
| chr11_+_45300669 | 0.39 |
ENSDART00000172238
|
mafgb
|
v-maf avian musculoaponeurotic fibrosarcoma oncogene homolog Gb |
| chr2_-_9544161 | 0.38 |
ENSDART00000124425
|
slc25a24l
|
solute carrier family 25 (mitochondrial carrier; phosphate carrier), member 24, like |
| chr20_-_8443425 | 0.37 |
ENSDART00000083908
|
dab1a
|
Dab, reelin signal transducer, homolog 1a (Drosophila) |
| chr1_+_51312752 | 0.36 |
ENSDART00000063938
|
mast1a
|
microtubule associated serine/threonine kinase 1a |
| chr21_+_25092300 | 0.36 |
ENSDART00000168505
|
dixdc1b
|
DIX domain containing 1b |
| chr13_+_24519175 | 0.36 |
ENSDART00000190790
|
sertad2b
|
SERTA domain containing 2b |
| chr6_-_12135741 | 0.36 |
ENSDART00000155090
|
tanc1a
|
tetratricopeptide repeat, ankyrin repeat and coiled-coil containing 1a |
| chr20_+_48100261 | 0.36 |
ENSDART00000158604
|
xkr5a
|
XK related 5a |
| chr23_-_18286822 | 0.36 |
ENSDART00000136672
|
fam19a1a
|
family with sequence similarity 19 (chemokine (C-C motif)-like), member A1a |
| chr14_-_30490465 | 0.35 |
ENSDART00000173107
|
micu3b
|
mitochondrial calcium uptake family, member 3b |
| chr10_+_42462577 | 0.35 |
ENSDART00000133463
|
npy8ar
|
neuropeptide Y receptor Y8a |
| chr18_-_42333428 | 0.34 |
ENSDART00000034225
|
cntn5
|
contactin 5 |
| chr5_-_50640171 | 0.34 |
ENSDART00000183353
|
mctp1a
|
multiple C2 domains, transmembrane 1a |
| chr10_-_74408 | 0.33 |
ENSDART00000100073
ENSDART00000141723 |
dyrk1aa
|
dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 1A, a |
| chr21_-_31143903 | 0.33 |
ENSDART00000111571
|
rap1gap2b
|
RAP1 GTPase activating protein 2b |
| chr19_+_31404686 | 0.32 |
ENSDART00000078459
|
pip4p2
|
phosphatidylinositol-4,5-bisphosphate 4-phosphatase 2 |
| chr25_-_16818978 | 0.32 |
ENSDART00000104140
|
dyrk4
|
dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 4 |
| chr7_+_63325819 | 0.32 |
ENSDART00000085612
ENSDART00000161436 |
pcdh7b
|
protocadherin 7b |
| chr17_-_22001303 | 0.31 |
ENSDART00000122190
|
slc22a7b.2
|
solute carrier family 22 (organic anion transporter), member 7b, tandem duplicate 2 |
| chr18_-_5692292 | 0.31 |
ENSDART00000121503
|
cplx3b
|
complexin 3b |
| chr21_-_24865217 | 0.31 |
ENSDART00000101136
|
igsf9bb
|
immunoglobulin superfamily, member 9Bb |
| chr22_-_18116635 | 0.30 |
ENSDART00000005724
|
ncanb
|
neurocan b |
| chr14_+_34068075 | 0.29 |
ENSDART00000135556
|
lonrf1
|
LON peptidase N-terminal domain and ring finger 1 |
| chr15_+_40792667 | 0.29 |
ENSDART00000193186
ENSDART00000184559 |
fat3a
|
FAT atypical cadherin 3a |
| chr24_-_29868151 | 0.28 |
ENSDART00000184802
|
aglb
|
amylo-alpha-1, 6-glucosidase, 4-alpha-glucanotransferase b |
| chr22_-_26323893 | 0.27 |
ENSDART00000105099
|
capn1b
|
calpain 1, (mu/I) large subunit b |
| chr12_+_5209822 | 0.27 |
ENSDART00000152610
|
SLC35G1
|
si:ch211-197g18.2 |
| chr15_-_39820491 | 0.27 |
ENSDART00000097134
|
robo1
|
roundabout, axon guidance receptor, homolog 1 (Drosophila) |
| chr3_+_36160979 | 0.27 |
ENSDART00000038525
|
si:ch211-234h8.7
|
si:ch211-234h8.7 |
| chr16_+_10557504 | 0.27 |
ENSDART00000091241
|
si:ch73-22o12.1
|
si:ch73-22o12.1 |
| chr10_+_29265463 | 0.26 |
ENSDART00000155390
|
crebzf
|
CREB/ATF bZIP transcription factor |
| chr17_-_17948587 | 0.26 |
ENSDART00000090447
|
hhipl1
|
HHIP-like 1 |
| chr3_+_41731527 | 0.26 |
ENSDART00000049007
ENSDART00000187866 |
chst12a
|
carbohydrate (chondroitin 4) sulfotransferase 12a |
| chr21_-_929448 | 0.25 |
ENSDART00000133976
|
txnl1
|
thioredoxin-like 1 |
| chr14_+_24840669 | 0.25 |
ENSDART00000106039
|
arhgef37
|
Rho guanine nucleotide exchange factor (GEF) 37 |
| chr3_-_26017831 | 0.25 |
ENSDART00000179982
|
hmox1a
|
heme oxygenase 1a |
| chr18_+_34478959 | 0.24 |
ENSDART00000059394
|
kcnab1a
|
potassium voltage-gated channel, shaker-related subfamily, beta member 1 a |
| chr22_-_11829436 | 0.23 |
ENSDART00000126784
|
ptpn4b
|
protein tyrosine phosphatase, non-receptor type 4b |
| chr7_+_13582256 | 0.23 |
ENSDART00000158477
|
ankdd1a
|
ankyrin repeat and death domain containing 1A |
| chr17_+_28533102 | 0.23 |
ENSDART00000156218
|
mdga2a
|
MAM domain containing glycosylphosphatidylinositol anchor 2a |
| chr25_-_11378623 | 0.23 |
ENSDART00000166586
|
enc2
|
ectodermal-neural cortex 2 |
| chr17_-_22001935 | 0.22 |
ENSDART00000185429
ENSDART00000193302 |
slc22a7b.2
|
solute carrier family 22 (organic anion transporter), member 7b, tandem duplicate 2 |
| chr21_-_40782393 | 0.22 |
ENSDART00000075808
|
apbb3
|
amyloid beta (A4) precursor protein-binding, family B, member 3 |
| chr11_+_26609110 | 0.22 |
ENSDART00000042322
|
map1lc3a
|
microtubule-associated protein 1 light chain 3 alpha |
| chr10_+_38775959 | 0.22 |
ENSDART00000192990
|
dscama
|
Down syndrome cell adhesion molecule a |
| chr10_+_38775408 | 0.21 |
ENSDART00000125045
|
dscama
|
Down syndrome cell adhesion molecule a |
| chr4_+_25630555 | 0.20 |
ENSDART00000133425
|
acot15
|
acyl-CoA thioesterase 15 |
| chr2_+_37134281 | 0.20 |
ENSDART00000020135
|
pex19
|
peroxisomal biogenesis factor 19 |
| chr21_+_38073210 | 0.20 |
ENSDART00000186309
ENSDART00000142106 |
klf8
|
Kruppel-like factor 8 |
| chr7_+_756942 | 0.20 |
ENSDART00000152224
|
zgc:63470
|
zgc:63470 |
| chr1_+_6640437 | 0.19 |
ENSDART00000147638
|
si:ch211-93g23.2
|
si:ch211-93g23.2 |
| chr15_+_29393519 | 0.19 |
ENSDART00000193488
ENSDART00000112375 |
gdpd5b
|
glycerophosphodiester phosphodiesterase domain containing 5b |
| chr10_+_17235370 | 0.19 |
ENSDART00000038780
|
sppl3
|
signal peptide peptidase 3 |
| chr13_-_22907260 | 0.19 |
ENSDART00000143097
|
rufy2
|
RUN and FYVE domain containing 2 |
| chr18_-_7456378 | 0.19 |
ENSDART00000081459
|
pdp2
|
putative pyruvate dehydrogenase phosphatase isoenzyme 2 |
| chr5_+_51909740 | 0.18 |
ENSDART00000162541
|
thbs4a
|
thrombospondin 4a |
| chr18_+_24922125 | 0.18 |
ENSDART00000180385
|
rgma
|
repulsive guidance molecule family member a |
| chr25_+_19710619 | 0.18 |
ENSDART00000156435
|
mxg
|
myxovirus (influenza virus) resistance G |
| chr14_-_30960470 | 0.18 |
ENSDART00000129989
|
si:ch211-191o15.6
|
si:ch211-191o15.6 |
| chr2_-_24962002 | 0.17 |
ENSDART00000132050
|
hltf
|
helicase-like transcription factor |
| chr9_-_30145080 | 0.17 |
ENSDART00000133746
|
abi3bpa
|
ABI family, member 3 (NESH) binding protein a |
| chr9_+_28688574 | 0.17 |
ENSDART00000101319
|
zgc:162396
|
zgc:162396 |
| chr14_+_34780886 | 0.17 |
ENSDART00000130469
ENSDART00000188806 ENSDART00000172924 ENSDART00000173266 ENSDART00000166377 ENSDART00000173371 |
sh3tc2
|
SH3 domain and tetratricopeptide repeats 2 |
| chr6_-_12644563 | 0.17 |
ENSDART00000153797
|
dock9b
|
dedicator of cytokinesis 9b |
| chr14_-_32486757 | 0.17 |
ENSDART00000148830
|
mcf2a
|
MCF.2 cell line derived transforming sequence a |
| chr7_-_40630698 | 0.17 |
ENSDART00000134547
|
ube3c
|
ubiquitin protein ligase E3C |
| chr14_-_25985698 | 0.17 |
ENSDART00000172909
ENSDART00000123053 |
atox1
|
antioxidant 1 copper chaperone |
| chr14_+_5835134 | 0.17 |
ENSDART00000054867
|
aup1
|
ancient ubiquitous protein 1 |
| chr24_+_35911020 | 0.17 |
ENSDART00000088480
|
abcd4
|
ATP-binding cassette, sub-family D (ALD), member 4 |
| chr2_-_6115688 | 0.17 |
ENSDART00000081663
|
prdx1
|
peroxiredoxin 1 |
| chr1_-_5745420 | 0.16 |
ENSDART00000166779
|
nrp2a
|
neuropilin 2a |
| chr8_+_20438884 | 0.16 |
ENSDART00000016422
ENSDART00000133794 |
mknk2b
|
MAP kinase interacting serine/threonine kinase 2b |
| chr8_+_10835456 | 0.16 |
ENSDART00000151388
|
mapk13
|
mitogen-activated protein kinase 13 |
| chr6_+_43903209 | 0.16 |
ENSDART00000006435
|
gpr27
|
G protein-coupled receptor 27 |
| chr21_-_929293 | 0.15 |
ENSDART00000006419
|
txnl1
|
thioredoxin-like 1 |
| chr5_-_64823750 | 0.15 |
ENSDART00000140305
|
lix1
|
limb and CNS expressed 1 |
| chr11_-_3334248 | 0.15 |
ENSDART00000154314
ENSDART00000121861 |
prph
|
peripherin |
| chr15_+_45591669 | 0.15 |
ENSDART00000157459
|
atg16l1
|
ATG16 autophagy related 16-like 1 (S. cerevisiae) |
| chr13_-_35459928 | 0.15 |
ENSDART00000144109
|
slx4ip
|
SLX4 interacting protein |
| chr3_-_26017592 | 0.15 |
ENSDART00000030890
|
hmox1a
|
heme oxygenase 1a |
| chr9_+_22466218 | 0.15 |
ENSDART00000131214
|
dgkg
|
diacylglycerol kinase, gamma |
| chr25_+_33046060 | 0.14 |
ENSDART00000165345
|
tln2b
|
talin 2b |
| chr10_+_26747755 | 0.14 |
ENSDART00000100329
|
f9b
|
coagulation factor IXb |
| chr14_+_30279391 | 0.13 |
ENSDART00000172794
|
fgl1
|
fibrinogen-like 1 |
| chr18_+_37015185 | 0.13 |
ENSDART00000191305
|
sipa1l3
|
signal-induced proliferation-associated 1 like 3 |
| chr18_-_16315822 | 0.13 |
ENSDART00000136479
|
rassf9
|
Ras association (RalGDS/AF-6) domain family (N-terminal) member 9 |
| chr2_-_2813259 | 0.13 |
ENSDART00000032540
|
usp14
|
ubiquitin specific peptidase 14 (tRNA-guanine transglycosylase) |
| chr22_-_31059670 | 0.13 |
ENSDART00000022445
|
cand2
|
cullin-associated and neddylation-dissociated 2 (putative) |
| chr7_+_17992640 | 0.13 |
ENSDART00000173679
|
ehbp1l1a
|
EH domain binding protein 1-like 1a |
| chr13_+_1430094 | 0.12 |
ENSDART00000169888
|
si:ch211-165e15.1
|
si:ch211-165e15.1 |
| chr3_+_62140077 | 0.12 |
ENSDART00000108945
|
GID4
|
GID complex subunit 4 homolog |
| chr20_-_39103119 | 0.12 |
ENSDART00000143379
|
rcan2
|
regulator of calcineurin 2 |
| chr16_+_45930962 | 0.11 |
ENSDART00000124689
ENSDART00000041811 |
otud7b
|
OTU deubiquitinase 7B |
| chr16_-_13613475 | 0.11 |
ENSDART00000139102
|
dbpb
|
D site albumin promoter binding protein b |
| chr6_+_515181 | 0.11 |
ENSDART00000171374
|
CACNA1I (1 of many)
|
si:ch73-379f7.5 |
| chr8_-_18516385 | 0.11 |
ENSDART00000149446
|
si:dkey-30h22.11
|
si:dkey-30h22.11 |
| chr12_-_1034383 | 0.10 |
ENSDART00000152455
ENSDART00000152346 |
polr3e
|
polymerase (RNA) III (DNA directed) polypeptide E |
| chr2_-_3038904 | 0.10 |
ENSDART00000186795
|
guk1a
|
guanylate kinase 1a |
| chr2_+_31437547 | 0.09 |
ENSDART00000141170
|
stam
|
signal transducing adaptor molecule (SH3 domain and ITAM motif) 1 |
| chr24_-_28229618 | 0.08 |
ENSDART00000145290
|
bcl2a
|
BCL2, apoptosis regulator a |
| chr7_-_62003831 | 0.08 |
ENSDART00000113585
ENSDART00000062704 |
plaa
|
phospholipase A2-activating protein |
| chr4_+_20085114 | 0.08 |
ENSDART00000186698
ENSDART00000188635 |
ppp6r2a
|
protein phosphatase 6, regulatory subunit 2a |
| chr12_-_19151708 | 0.08 |
ENSDART00000057124
|
tefa
|
thyrotrophic embryonic factor a |
| chr14_+_21755469 | 0.08 |
ENSDART00000186326
|
kdm2ab
|
lysine (K)-specific demethylase 2Ab |
| chr2_-_6115389 | 0.08 |
ENSDART00000134921
|
prdx1
|
peroxiredoxin 1 |
| chr2_+_25278107 | 0.07 |
ENSDART00000131977
|
ppp2r3a
|
protein phosphatase 2, regulatory subunit B'', alpha |
| chr11_+_13528437 | 0.07 |
ENSDART00000011362
|
arrdc2
|
arrestin domain containing 2 |
| chr14_-_2004291 | 0.07 |
ENSDART00000114039
|
pcdh2g5
|
protocadherin 2 gamma 5 |
| chr13_+_42309688 | 0.07 |
ENSDART00000158367
|
ide
|
insulin-degrading enzyme |
| chr14_-_34771371 | 0.07 |
ENSDART00000160598
ENSDART00000150413 ENSDART00000168910 |
ablim3
|
actin binding LIM protein family, member 3 |
| chr23_+_21278948 | 0.07 |
ENSDART00000156701
ENSDART00000033970 |
ubr4
|
ubiquitin protein ligase E3 component n-recognin 4 |
| chr7_-_23971497 | 0.07 |
ENSDART00000173603
|
si:dkey-183c6.8
|
si:dkey-183c6.8 |
| chr7_-_44704910 | 0.07 |
ENSDART00000037850
|
dync1li2
|
dynein, cytoplasmic 1, light intermediate chain 2 |
| chr23_+_3731375 | 0.06 |
ENSDART00000141782
|
smim29
|
small integral membrane protein 29 |
| chr6_-_52723901 | 0.06 |
ENSDART00000033949
|
oser1
|
oxidative stress responsive serine-rich 1 |
| chr1_+_41609676 | 0.06 |
ENSDART00000183675
|
mogs
|
mannosyl-oligosaccharide glucosidase |
| chr3_+_23669267 | 0.06 |
ENSDART00000111227
|
hoxb10a
|
homeobox B10a |
| chr10_-_300000 | 0.06 |
ENSDART00000183273
|
emsy
|
EMSY BRCA2-interacting transcriptional repressor |
| chr6_+_20651398 | 0.06 |
ENSDART00000183663
|
slc19a1
|
solute carrier family 19 (folate transporter), member 1 |
| chr22_-_16416882 | 0.05 |
ENSDART00000062749
|
cts12
|
cathepsin 12 |
| chr14_-_48245816 | 0.05 |
ENSDART00000172452
|
rxfp1
|
relaxin/insulin-like family peptide receptor 1 |
| chr9_+_19304697 | 0.05 |
ENSDART00000040449
|
htr1fa
|
5-hydroxytryptamine (serotonin) receptor 1Fa |
| chr1_+_35473219 | 0.05 |
ENSDART00000109678
ENSDART00000181635 |
usp38
|
ubiquitin specific peptidase 38 |
| chr20_-_32981053 | 0.05 |
ENSDART00000138708
|
nbas
|
neuroblastoma amplified sequence |
| chr24_-_27409599 | 0.05 |
ENSDART00000041770
|
ccl34b.3
|
chemokine (C-C motif) ligand 34b, duplicate 3 |
| chr4_+_33012407 | 0.05 |
ENSDART00000151873
|
si:dkey-26h11.2
|
si:dkey-26h11.2 |
| chr2_-_37134169 | 0.04 |
ENSDART00000146123
ENSDART00000146533 ENSDART00000040427 |
elavl1a
|
ELAV like RNA binding protein 1a |
| chr13_-_23007813 | 0.04 |
ENSDART00000057638
|
hk1
|
hexokinase 1 |
| chr22_-_10354381 | 0.04 |
ENSDART00000092050
|
stab1
|
stabilin 1 |
| chr17_+_51744450 | 0.04 |
ENSDART00000190955
ENSDART00000149807 |
odc1
|
ornithine decarboxylase 1 |
| chr7_+_13491452 | 0.04 |
ENSDART00000053535
|
arih1l
|
ariadne homolog, ubiquitin-conjugating enzyme E2 binding protein, 1 like |
| chr1_+_19943803 | 0.04 |
ENSDART00000164661
|
apbb2b
|
amyloid beta (A4) precursor protein-binding, family B, member 2b |
| chr21_+_34686764 | 0.04 |
ENSDART00000005479
|
chm
|
choroideremia (Rab escort protein 1) |
| chr3_+_26064091 | 0.04 |
ENSDART00000143697
|
si:dkeyp-69e1.8
|
si:dkeyp-69e1.8 |
| chr8_-_39884359 | 0.04 |
ENSDART00000131372
|
mlec
|
malectin |
| chr23_-_12906228 | 0.04 |
ENSDART00000138807
|
ndnl2
|
necdin-like 2 |
| chr9_+_23224761 | 0.04 |
ENSDART00000142008
|
map3k19
|
mitogen-activated protein kinase kinase kinase 19 |
| chr7_+_50464500 | 0.03 |
ENSDART00000191356
|
serinc4
|
serine incorporator 4 |
| chr3_+_32832042 | 0.03 |
ENSDART00000132679
ENSDART00000035759 |
cd2bp2
|
CD2 (cytoplasmic tail) binding protein 2 |
| chr2_-_26642831 | 0.03 |
ENSDART00000132854
ENSDART00000087714 ENSDART00000132651 |
u2surp
|
U2 snRNP-associated SURP domain containing |
| chr7_+_62004048 | 0.03 |
ENSDART00000181818
|
smim20
|
small integral membrane protein 20 |
| chr17_+_15674052 | 0.03 |
ENSDART00000156726
|
bach2a
|
BTB and CNC homology 1, basic leucine zipper transcription factor 2a |
Network of associatons between targets according to the STRING database.
First level regulatory network of nfe2l2a
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.9 | GO:0051881 | regulation of mitochondrial membrane potential(GO:0051881) regulation of release of cytochrome c from mitochondria(GO:0090199) positive regulation of release of cytochrome c from mitochondria(GO:0090200) |
| 0.2 | 0.7 | GO:0014014 | negative regulation of gliogenesis(GO:0014014) |
| 0.2 | 0.9 | GO:0021557 | oculomotor nerve development(GO:0021557) |
| 0.2 | 0.7 | GO:0015824 | proline transport(GO:0015824) |
| 0.1 | 0.4 | GO:0043576 | respiratory gaseous exchange(GO:0007585) regulation of respiratory gaseous exchange(GO:0043576) |
| 0.1 | 0.5 | GO:0060220 | retinal cone cell fate determination(GO:0042671) eye photoreceptor cell fate commitment(GO:0042706) photoreceptor cell fate determination(GO:0043703) retinal cone cell fate commitment(GO:0046551) photoreceptor cell fate commitment(GO:0046552) camera-type eye photoreceptor cell fate commitment(GO:0060220) |
| 0.1 | 0.2 | GO:0021611 | facial nerve formation(GO:0021611) |
| 0.1 | 0.7 | GO:0097105 | presynaptic membrane organization(GO:0097090) postsynaptic membrane assembly(GO:0097104) presynaptic membrane assembly(GO:0097105) |
| 0.1 | 0.2 | GO:1901017 | negative regulation of potassium ion transmembrane transporter activity(GO:1901017) negative regulation of potassium ion transmembrane transport(GO:1901380) negative regulation of delayed rectifier potassium channel activity(GO:1902260) negative regulation of voltage-gated potassium channel activity(GO:1903817) |
| 0.1 | 0.5 | GO:0046716 | muscle cell cellular homeostasis(GO:0046716) |
| 0.1 | 0.6 | GO:0071300 | cellular response to retinoic acid(GO:0071300) |
| 0.1 | 0.4 | GO:0045604 | regulation of epidermal cell differentiation(GO:0045604) |
| 0.1 | 0.5 | GO:1904861 | excitatory synapse assembly(GO:1904861) |
| 0.1 | 0.4 | GO:0098700 | neurotransmitter loading into synaptic vesicle(GO:0098700) |
| 0.1 | 0.2 | GO:0015919 | peroxisomal membrane transport(GO:0015919) protein import into peroxisome membrane(GO:0045046) |
| 0.0 | 0.9 | GO:0036376 | cellular potassium ion homeostasis(GO:0030007) sodium ion export from cell(GO:0036376) sodium ion export(GO:0071436) |
| 0.0 | 0.4 | GO:0097476 | spinal cord motor neuron migration(GO:0097476) lateral motor column neuron migration(GO:0097477) |
| 0.0 | 0.2 | GO:1904184 | regulation of pyruvate dehydrogenase activity(GO:1904182) positive regulation of pyruvate dehydrogenase activity(GO:1904184) |
| 0.0 | 0.8 | GO:0051496 | positive regulation of stress fiber assembly(GO:0051496) |
| 0.0 | 0.1 | GO:0010265 | SCF complex assembly(GO:0010265) |
| 0.0 | 0.7 | GO:0051560 | mitochondrial calcium ion homeostasis(GO:0051560) |
| 0.0 | 0.2 | GO:0031293 | membrane protein intracellular domain proteolysis(GO:0031293) |
| 0.0 | 0.3 | GO:0003262 | endocardial progenitor cell migration to the midline involved in heart field formation(GO:0003262) |
| 0.0 | 0.5 | GO:0051968 | positive regulation of synaptic transmission, glutamatergic(GO:0051968) |
| 0.0 | 0.4 | GO:0070782 | phosphatidylserine exposure on apoptotic cell surface(GO:0070782) |
| 0.0 | 0.2 | GO:0019079 | viral genome replication(GO:0019079) negative stranded viral RNA replication(GO:0039689) viral RNA genome replication(GO:0039694) RNA replication(GO:0039703) multi-organism biosynthetic process(GO:0044034) |
| 0.0 | 0.2 | GO:0015910 | peroxisomal long-chain fatty acid import(GO:0015910) |
| 0.0 | 0.3 | GO:0000272 | polysaccharide catabolic process(GO:0000272) glycogen catabolic process(GO:0005980) glucan catabolic process(GO:0009251) cellular polysaccharide catabolic process(GO:0044247) |
| 0.0 | 0.5 | GO:0010962 | regulation of glycogen biosynthetic process(GO:0005979) regulation of glucan biosynthetic process(GO:0010962) |
| 0.0 | 0.2 | GO:0071451 | response to superoxide(GO:0000303) response to oxygen radical(GO:0000305) removal of superoxide radicals(GO:0019430) cellular response to oxygen radical(GO:0071450) cellular response to superoxide(GO:0071451) cellular oxidant detoxification(GO:0098869) cellular detoxification(GO:1990748) |
| 0.0 | 0.2 | GO:0034389 | lipid particle organization(GO:0034389) |
| 0.0 | 0.1 | GO:0071947 | protein deubiquitination involved in ubiquitin-dependent protein catabolic process(GO:0071947) |
| 0.0 | 0.4 | GO:0014823 | response to activity(GO:0014823) |
| 0.0 | 0.2 | GO:0006465 | signal peptide processing(GO:0006465) |
| 0.0 | 0.2 | GO:0006878 | cellular copper ion homeostasis(GO:0006878) |
| 0.0 | 0.3 | GO:0048096 | chromatin-mediated maintenance of transcription(GO:0048096) |
| 0.0 | 0.1 | GO:0098838 | folic acid transport(GO:0015884) drug transport(GO:0015893) methotrexate transport(GO:0051958) reduced folate transmembrane transport(GO:0098838) |
| 0.0 | 0.2 | GO:0021684 | cerebellar granular layer development(GO:0021681) cerebellar granular layer morphogenesis(GO:0021683) cerebellar granular layer formation(GO:0021684) cerebellar Purkinje cell layer formation(GO:0021694) cerebellar Purkinje cell differentiation(GO:0021702) cerebellar granule cell differentiation(GO:0021707) |
| 0.0 | 0.2 | GO:0061709 | reticulophagy(GO:0061709) |
| 0.0 | 0.5 | GO:0031114 | regulation of microtubule depolymerization(GO:0031114) |
| 0.0 | 0.9 | GO:0043123 | positive regulation of I-kappaB kinase/NF-kappaB signaling(GO:0043123) |
| 0.0 | 0.7 | GO:0032011 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.0 | 0.4 | GO:0070593 | dendrite self-avoidance(GO:0070593) |
| 0.0 | 0.1 | GO:0009794 | regulation of mitotic cell cycle, embryonic(GO:0009794) mitotic cell cycle, embryonic(GO:0045448) |
| 0.0 | 0.8 | GO:0006414 | translational elongation(GO:0006414) |
| 0.0 | 0.6 | GO:0046928 | regulation of neurotransmitter secretion(GO:0046928) |
| 0.0 | 0.0 | GO:2000622 | negative regulation of mRNA catabolic process(GO:1902373) regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000622) negative regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000623) |
| 0.0 | 0.2 | GO:0006995 | cellular response to nitrogen starvation(GO:0006995) cellular response to nitrogen levels(GO:0043562) |
| 0.0 | 0.0 | GO:1901492 | positive regulation of lymphangiogenesis(GO:1901492) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.8 | GO:0005854 | nascent polypeptide-associated complex(GO:0005854) |
| 0.1 | 0.5 | GO:0033181 | plasma membrane proton-transporting V-type ATPase complex(GO:0033181) |
| 0.1 | 1.2 | GO:0097038 | perinuclear endoplasmic reticulum(GO:0097038) |
| 0.1 | 0.7 | GO:1990246 | uniplex complex(GO:1990246) |
| 0.0 | 0.2 | GO:0071458 | integral component of cytoplasmic side of endoplasmic reticulum membrane(GO:0071458) integral component of lumenal side of endoplasmic reticulum membrane(GO:0071556) lumenal side of endoplasmic reticulum membrane(GO:0098553) |
| 0.0 | 0.4 | GO:0042583 | chromaffin granule(GO:0042583) |
| 0.0 | 0.2 | GO:0000839 | Hrd1p ubiquitin ligase ERAD-L complex(GO:0000839) |
| 0.0 | 0.5 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.0 | 0.4 | GO:0098563 | integral component of synaptic vesicle membrane(GO:0030285) intrinsic component of synaptic vesicle membrane(GO:0098563) |
| 0.0 | 1.5 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.0 | 0.4 | GO:1990907 | beta-catenin-TCF complex(GO:1990907) |
| 0.0 | 0.6 | GO:0031594 | neuromuscular junction(GO:0031594) |
| 0.0 | 0.2 | GO:0044224 | juxtaparanode region of axon(GO:0044224) |
| 0.0 | 0.6 | GO:0043186 | P granule(GO:0043186) |
| 0.0 | 0.3 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.0 | 0.5 | GO:0098839 | postsynaptic density membrane(GO:0098839) postsynaptic specialization membrane(GO:0099634) |
| 0.0 | 0.1 | GO:0031907 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.9 | GO:0044548 | S100 protein binding(GO:0044548) |
| 0.1 | 0.7 | GO:0004105 | choline-phosphate cytidylyltransferase activity(GO:0004105) |
| 0.1 | 0.5 | GO:0070324 | thyroid hormone binding(GO:0070324) |
| 0.1 | 0.9 | GO:0005391 | sodium:potassium-exchanging ATPase activity(GO:0005391) |
| 0.1 | 0.3 | GO:0004133 | glycogen debranching enzyme activity(GO:0004133) 4-alpha-glucanotransferase activity(GO:0004134) amylo-alpha-1,6-glucosidase activity(GO:0004135) |
| 0.1 | 0.4 | GO:0004392 | heme oxygenase (decyclizing) activity(GO:0004392) |
| 0.1 | 0.6 | GO:0004791 | thioredoxin-disulfide reductase activity(GO:0004791) |
| 0.1 | 0.3 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 0.0 | 0.2 | GO:0004741 | [pyruvate dehydrogenase (lipoamide)] phosphatase activity(GO:0004741) |
| 0.0 | 0.5 | GO:0016909 | JUN kinase activity(GO:0004705) SAP kinase activity(GO:0016909) |
| 0.0 | 1.1 | GO:0015248 | sterol transporter activity(GO:0015248) |
| 0.0 | 0.3 | GO:0034597 | phosphatidylinositol-4,5-bisphosphate 4-phosphatase activity(GO:0034597) |
| 0.0 | 0.5 | GO:2001069 | glycogen binding(GO:2001069) |
| 0.0 | 0.2 | GO:0008379 | thioredoxin peroxidase activity(GO:0008379) |
| 0.0 | 0.4 | GO:0015280 | ligand-gated sodium channel activity(GO:0015280) |
| 0.0 | 0.2 | GO:0016531 | copper chaperone activity(GO:0016531) |
| 0.0 | 0.4 | GO:0005326 | neurotransmitter transporter activity(GO:0005326) |
| 0.0 | 0.7 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 0.0 | 0.2 | GO:0097027 | ubiquitin-protein transferase activator activity(GO:0097027) |
| 0.0 | 1.1 | GO:0016493 | C-C chemokine receptor activity(GO:0016493) |
| 0.0 | 0.2 | GO:0042500 | aspartic endopeptidase activity, intramembrane cleaving(GO:0042500) |
| 0.0 | 1.0 | GO:0004712 | protein serine/threonine/tyrosine kinase activity(GO:0004712) |
| 0.0 | 0.3 | GO:0015129 | lactate transmembrane transporter activity(GO:0015129) |
| 0.0 | 0.2 | GO:0008429 | phosphatidylethanolamine binding(GO:0008429) |
| 0.0 | 0.1 | GO:0004385 | guanylate kinase activity(GO:0004385) |
| 0.0 | 0.3 | GO:0004983 | neuropeptide Y receptor activity(GO:0004983) |
| 0.0 | 0.1 | GO:0008597 | calcium-dependent protein serine/threonine phosphatase regulator activity(GO:0008597) |
| 0.0 | 0.1 | GO:0008518 | reduced folate carrier activity(GO:0008518) methotrexate transporter activity(GO:0015350) |
| 0.0 | 0.2 | GO:0000268 | peroxisome targeting sequence binding(GO:0000268) |
| 0.0 | 0.8 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.0 | 0.1 | GO:1990380 | Lys48-specific deubiquitinase activity(GO:1990380) |
| 0.0 | 1.5 | GO:0005544 | calcium-dependent phospholipid binding(GO:0005544) |
| 0.0 | 0.2 | GO:0005021 | vascular endothelial growth factor-activated receptor activity(GO:0005021) |
| 0.0 | 0.4 | GO:0098632 | protein binding involved in cell-cell adhesion(GO:0098632) |
| 0.0 | 0.2 | GO:0004033 | aldo-keto reductase (NADP) activity(GO:0004033) |
| 0.0 | 0.3 | GO:0051787 | misfolded protein binding(GO:0051787) |
| 0.0 | 0.2 | GO:0005324 | long-chain fatty acid transporter activity(GO:0005324) |
| 0.0 | 0.3 | GO:0001540 | beta-amyloid binding(GO:0001540) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.0 | 1.0 | PID FOXO PATHWAY | FoxO family signaling |
| 0.0 | 0.2 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.0 | 0.7 | PID NOTCH PATHWAY | Notch signaling pathway |
| 0.0 | 0.1 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.9 | REACTOME ADVANCED GLYCOSYLATION ENDPRODUCT RECEPTOR SIGNALING | Genes involved in Advanced glycosylation endproduct receptor signaling |
| 0.1 | 0.5 | REACTOME ACTIVATION OF THE AP1 FAMILY OF TRANSCRIPTION FACTORS | Genes involved in Activation of the AP-1 family of transcription factors |
| 0.0 | 0.7 | REACTOME ACTIVATED NOTCH1 TRANSMITS SIGNAL TO THE NUCLEUS | Genes involved in Activated NOTCH1 Transmits Signal to the Nucleus |
| 0.0 | 0.2 | REACTOME DSCAM INTERACTIONS | Genes involved in DSCAM interactions |
| 0.0 | 0.3 | REACTOME ACTIVATION OF RAC | Genes involved in Activation of Rac |
| 0.0 | 0.2 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.0 | 0.2 | REACTOME REGULATION OF PYRUVATE DEHYDROGENASE PDH COMPLEX | Genes involved in Regulation of pyruvate dehydrogenase (PDH) complex |