Project
PRJNA207719: Tissue specific transcriptome profiling
Navigation
Downloads
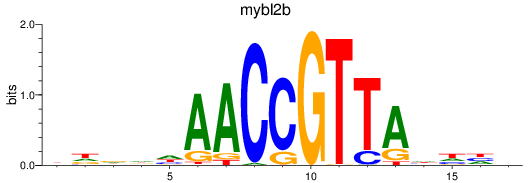
Results for mybl2b
Z-value: 1.59
Transcription factors associated with mybl2b
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
mybl2b
|
ENSDARG00000032264 | v-myb avian myeloblastosis viral oncogene homolog-like 2b |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| mybl2b | dr11_v1_chr11_+_1551603_1551663 | 0.50 | 3.9e-01 | Click! |
Activity profile of mybl2b motif
Sorted Z-values of mybl2b motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr4_-_2380173 | 1.28 |
ENSDART00000177727
|
nap1l1
|
nucleosome assembly protein 1-like 1 |
| chr21_-_43656125 | 1.05 |
ENSDART00000136392
ENSDART00000192805 ENSDART00000181960 |
si:ch211-263m18.4
si:dkey-229d11.5
|
si:ch211-263m18.4 si:dkey-229d11.5 |
| chr6_-_55297274 | 0.96 |
ENSDART00000184283
|
ube2c
|
ubiquitin-conjugating enzyme E2C |
| chr9_-_2892045 | 0.95 |
ENSDART00000137201
|
cdca7a
|
cell division cycle associated 7a |
| chr15_-_17074393 | 0.90 |
ENSDART00000155526
|
si:ch211-24o10.6
|
si:ch211-24o10.6 |
| chr14_+_52563794 | 0.79 |
ENSDART00000168874
|
rpl26
|
ribosomal protein L26 |
| chr9_-_2892250 | 0.78 |
ENSDART00000140695
|
cdca7a
|
cell division cycle associated 7a |
| chr7_+_1579510 | 0.77 |
ENSDART00000190525
|
supt16h
|
SPT16 homolog, facilitates chromatin remodeling subunit |
| chr21_-_22681534 | 0.75 |
ENSDART00000159233
|
gig2f
|
grass carp reovirus (GCRV)-induced gene 2f |
| chr11_-_24015845 | 0.70 |
ENSDART00000191783
|
chia.3
|
chitinase, acidic.3 |
| chr6_+_9191981 | 0.69 |
ENSDART00000150916
|
zgc:112392
|
zgc:112392 |
| chr21_-_1635268 | 0.68 |
ENSDART00000151258
|
zgc:152948
|
zgc:152948 |
| chr11_-_40519886 | 0.66 |
ENSDART00000172819
|
miip
|
migration and invasion inhibitory protein |
| chr9_-_23253870 | 0.65 |
ENSDART00000143657
ENSDART00000169911 |
acmsd
|
aminocarboxymuconate semialdehyde decarboxylase |
| chr1_-_8020589 | 0.65 |
ENSDART00000143881
|
si:dkeyp-9d4.2
|
si:dkeyp-9d4.2 |
| chr19_-_47571456 | 0.62 |
ENSDART00000158071
ENSDART00000165841 |
rrm2
|
ribonucleotide reductase M2 polypeptide |
| chr22_-_23590069 | 0.60 |
ENSDART00000172067
|
f13b
|
coagulation factor XIII, B polypeptide |
| chr24_+_24809877 | 0.59 |
ENSDART00000185117
|
dnajc5b
|
DnaJ (Hsp40) homolog, subfamily C, member 5 beta |
| chr22_-_5655680 | 0.59 |
ENSDART00000159629
|
mcm2
|
minichromosome maintenance complex component 2 |
| chr1_-_49225890 | 0.59 |
ENSDART00000111598
|
cxcl18b
|
chemokine (C-X-C motif) ligand 18b |
| chr23_+_36095260 | 0.58 |
ENSDART00000127384
|
hoxc9a
|
homeobox C9a |
| chr5_-_69212184 | 0.58 |
ENSDART00000053963
|
mat2ab
|
methionine adenosyltransferase II, alpha b |
| chr10_-_26202766 | 0.58 |
ENSDART00000136393
|
fhdc3
|
FH2 domain containing 3 |
| chr17_-_21832119 | 0.57 |
ENSDART00000153539
ENSDART00000079008 |
si:ch211-208g24.8
|
si:ch211-208g24.8 |
| chr5_-_24127310 | 0.57 |
ENSDART00000182700
ENSDART00000154313 |
capga
|
capping protein (actin filament), gelsolin-like a |
| chr5_+_32882688 | 0.56 |
ENSDART00000008807
ENSDART00000185317 |
rpl12
|
ribosomal protein L12 |
| chr13_-_50139916 | 0.55 |
ENSDART00000099475
|
nid1a
|
nidogen 1a |
| chr10_-_35257458 | 0.55 |
ENSDART00000143890
ENSDART00000139107 ENSDART00000082445 |
prr11
|
proline rich 11 |
| chr14_+_41345175 | 0.53 |
ENSDART00000086104
|
nox1
|
NADPH oxidase 1 |
| chr19_-_24555935 | 0.51 |
ENSDART00000132660
ENSDART00000162801 |
polr3gla
|
polymerase (RNA) III (DNA directed) polypeptide G like a |
| chr25_-_13403726 | 0.49 |
ENSDART00000056723
|
gins3
|
GINS complex subunit 3 |
| chr2_-_51511494 | 0.48 |
ENSDART00000161392
ENSDART00000160877 |
si:ch211-9d9.8
|
si:ch211-9d9.8 |
| chr3_+_14641962 | 0.47 |
ENSDART00000091070
|
zgc:158403
|
zgc:158403 |
| chr25_-_37319117 | 0.46 |
ENSDART00000044499
|
igl4v9
|
immunoglobulin light 4 variable 9 |
| chr19_-_24555623 | 0.45 |
ENSDART00000176022
|
polr3gla
|
polymerase (RNA) III (DNA directed) polypeptide G like a |
| chr8_+_25629941 | 0.44 |
ENSDART00000186925
|
wasa
|
Wiskott-Aldrich syndrome (eczema-thrombocytopenia) a |
| chr22_-_15578402 | 0.43 |
ENSDART00000062986
|
hsh2d
|
hematopoietic SH2 domain containing |
| chr19_+_43579786 | 0.43 |
ENSDART00000138404
|
si:ch211-199g17.2
|
si:ch211-199g17.2 |
| chr7_+_5909271 | 0.43 |
ENSDART00000173363
|
zgc:165555
|
zgc:165555 |
| chr3_+_1211242 | 0.43 |
ENSDART00000171287
ENSDART00000165769 |
poldip3
|
polymerase (DNA-directed), delta interacting protein 3 |
| chr25_+_3099073 | 0.42 |
ENSDART00000022506
|
rab3il1
|
RAB3A interacting protein (rabin3)-like 1 |
| chr16_-_14397003 | 0.42 |
ENSDART00000170957
|
crabp2a
|
cellular retinoic acid binding protein 2, a |
| chr24_-_25166720 | 0.41 |
ENSDART00000141601
|
phldb2b
|
pleckstrin homology-like domain, family B, member 2b |
| chr24_-_21150100 | 0.41 |
ENSDART00000154249
ENSDART00000193256 |
gramd1c
|
GRAM domain containing 1c |
| chr22_-_4649238 | 0.39 |
ENSDART00000125302
|
fbn2b
|
fibrillin 2b |
| chr24_-_25166416 | 0.39 |
ENSDART00000111552
ENSDART00000169495 |
phldb2b
|
pleckstrin homology-like domain, family B, member 2b |
| chr4_-_12790886 | 0.39 |
ENSDART00000182535
ENSDART00000067131 ENSDART00000186426 |
irak3
|
interleukin-1 receptor-associated kinase 3 |
| chr6_+_3473657 | 0.39 |
ENSDART00000011785
|
rad54l
|
RAD54 like |
| chr7_-_6352569 | 0.39 |
ENSDART00000173074
|
zgc:165555
|
zgc:165555 |
| chr6_+_57741039 | 0.38 |
ENSDART00000169175
|
SNTA1
|
syntrophin alpha 1 |
| chr23_-_10722664 | 0.37 |
ENSDART00000146526
ENSDART00000129022 ENSDART00000104985 |
foxp1a
|
forkhead box P1a |
| chr5_-_1869982 | 0.37 |
ENSDART00000055878
|
rcl1
|
RNA terminal phosphate cyclase-like 1 |
| chr2_+_22495274 | 0.36 |
ENSDART00000167915
|
lrrc8da
|
leucine rich repeat containing 8 VRAC subunit Da |
| chr7_+_1579236 | 0.36 |
ENSDART00000172830
|
supt16h
|
SPT16 homolog, facilitates chromatin remodeling subunit |
| chr8_+_23382568 | 0.36 |
ENSDART00000129167
|
mapre1a
|
microtubule-associated protein, RP/EB family, member 1a |
| chr2_-_8648440 | 0.36 |
ENSDART00000135743
|
si:ch211-71m22.3
|
si:ch211-71m22.3 |
| chr25_-_16552135 | 0.35 |
ENSDART00000125836
|
si:ch211-266k8.4
|
si:ch211-266k8.4 |
| chr2_+_6253246 | 0.35 |
ENSDART00000058256
ENSDART00000076700 |
zp3b
|
zona pellucida glycoprotein 3b |
| chr18_+_29898955 | 0.35 |
ENSDART00000064080
|
cenpn
|
centromere protein N |
| chr23_+_36308717 | 0.34 |
ENSDART00000131711
|
hnrnpa1b
|
heterogeneous nuclear ribonucleoprotein A1b |
| chr6_-_54180699 | 0.34 |
ENSDART00000045901
|
rps10
|
ribosomal protein S10 |
| chr2_-_51757328 | 0.32 |
ENSDART00000189286
|
si:ch211-9d9.1
|
si:ch211-9d9.1 |
| chr23_+_20771899 | 0.32 |
ENSDART00000123460
|
si:ch211-112c15.8
|
si:ch211-112c15.8 |
| chr1_-_23293261 | 0.32 |
ENSDART00000122648
|
ugdh
|
UDP-glucose 6-dehydrogenase |
| chr18_-_18587745 | 0.32 |
ENSDART00000191973
|
sf3b3
|
splicing factor 3b, subunit 3 |
| chr16_+_19537073 | 0.31 |
ENSDART00000190590
|
sp8b
|
sp8 transcription factor b |
| chr8_-_12432604 | 0.31 |
ENSDART00000133350
ENSDART00000140699 ENSDART00000101174 |
traf1
|
TNF receptor-associated factor 1 |
| chr14_-_733565 | 0.30 |
ENSDART00000158097
|
tlr1
|
toll-like receptor 1 |
| chr4_-_5831036 | 0.30 |
ENSDART00000166232
|
foxm1
|
forkhead box M1 |
| chr5_+_4366431 | 0.30 |
ENSDART00000168560
ENSDART00000149185 |
sat1a.2
|
spermidine/spermine N1-acetyltransferase 1a, duplicate 2 |
| chr4_-_73572030 | 0.29 |
ENSDART00000121652
|
znf1015
|
zinc finger protein 1015 |
| chr1_+_23559478 | 0.29 |
ENSDART00000133659
|
ncapg
|
non-SMC condensin I complex, subunit G |
| chr21_-_41838284 | 0.28 |
ENSDART00000141067
|
cct6a
|
chaperonin containing TCP1, subunit 6A (zeta 1) |
| chr5_-_23855447 | 0.28 |
ENSDART00000051541
|
gbgt1l3
|
globoside alpha-1,3-N-acetylgalactosaminyltransferase 1, like 3 |
| chr13_+_15701849 | 0.28 |
ENSDART00000003517
|
trmt61a
|
tRNA methyltransferase 61A |
| chr5_-_25084318 | 0.28 |
ENSDART00000089339
|
dph7
|
diphthamide biosynthesis 7 |
| chr18_-_18821929 | 0.27 |
ENSDART00000178875
|
aars
|
alanyl-tRNA synthetase |
| chr16_+_19536614 | 0.27 |
ENSDART00000112894
ENSDART00000079201 ENSDART00000139357 |
sp8b
|
sp8 transcription factor b |
| chr22_+_21153031 | 0.27 |
ENSDART00000166342
|
crlf1b
|
cytokine receptor-like factor 1b |
| chr13_+_45524475 | 0.26 |
ENSDART00000074567
ENSDART00000019113 |
maco1b
|
macoilin 1b |
| chr10_-_45029041 | 0.26 |
ENSDART00000167878
|
polm
|
polymerase (DNA directed), mu |
| chr5_-_67499279 | 0.26 |
ENSDART00000128050
|
si:dkey-251i10.3
|
si:dkey-251i10.3 |
| chr10_+_6121558 | 0.26 |
ENSDART00000166799
ENSDART00000157947 |
tln1
|
talin 1 |
| chr22_+_2306386 | 0.26 |
ENSDART00000148300
ENSDART00000147644 |
si:dkey-4c15.16
|
si:dkey-4c15.16 |
| chr3_+_23029484 | 0.25 |
ENSDART00000187900
|
nags
|
N-acetylglutamate synthase |
| chr12_+_26491637 | 0.25 |
ENSDART00000087036
|
armc7
|
armadillo repeat containing 7 |
| chr3_-_36823643 | 0.25 |
ENSDART00000131340
ENSDART00000185075 |
abcc6b.2
|
ATP-binding cassette, sub-family C (CFTR/MRP), member 6b, tandem duplicate 2 |
| chr15_+_17030941 | 0.25 |
ENSDART00000062069
|
plin2
|
perilipin 2 |
| chr4_+_39742119 | 0.25 |
ENSDART00000176004
|
CR749167.2
|
|
| chr23_+_20689255 | 0.25 |
ENSDART00000182420
|
usp21
|
ubiquitin specific peptidase 21 |
| chr17_-_22573311 | 0.25 |
ENSDART00000141523
ENSDART00000140022 ENSDART00000079390 ENSDART00000188644 |
exo1
|
exonuclease 1 |
| chr2_+_26655744 | 0.24 |
ENSDART00000174928
|
asph
|
aspartate beta-hydroxylase |
| chr25_+_35134393 | 0.24 |
ENSDART00000185379
|
CR762436.2
|
|
| chr11_-_20988238 | 0.24 |
ENSDART00000155238
|
taf4a
|
TAF4A RNA polymerase II, TATA box binding protein (TBP)-associated factor |
| chr7_+_17106160 | 0.24 |
ENSDART00000190048
ENSDART00000180004 ENSDART00000013409 |
prmt3
|
protein arginine methyltransferase 3 |
| chr19_+_7124337 | 0.24 |
ENSDART00000031380
|
tap2a
|
transporter associated with antigen processing, subunit type a |
| chr24_+_20325312 | 0.24 |
ENSDART00000007997
|
acaa1
|
acetyl-CoA acyltransferase 1 |
| chr10_+_43037064 | 0.23 |
ENSDART00000160159
|
atg10
|
ATG10 autophagy related 10 homolog (S. cerevisiae) |
| chr19_-_46566430 | 0.23 |
ENSDART00000166668
|
ext1b
|
exostosin glycosyltransferase 1b |
| chr22_+_21152847 | 0.23 |
ENSDART00000007321
|
crlf1b
|
cytokine receptor-like factor 1b |
| chr10_+_16501699 | 0.23 |
ENSDART00000121864
|
slc27a6
|
solute carrier family 27 (fatty acid transporter), member 6 |
| chr4_-_26035770 | 0.23 |
ENSDART00000124514
|
usp44
|
ubiquitin specific peptidase 44 |
| chr8_-_47206114 | 0.22 |
ENSDART00000147606
ENSDART00000060883 ENSDART00000060880 |
ppih
|
peptidylprolyl isomerase H (cyclophilin H) |
| chr19_+_16068445 | 0.22 |
ENSDART00000138709
|
gmeb1
|
glucocorticoid modulatory element binding protein 1 |
| chr6_+_52212927 | 0.22 |
ENSDART00000143458
|
ywhaba
|
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, beta polypeptide a |
| chr8_-_47206399 | 0.21 |
ENSDART00000060877
|
ppih
|
peptidylprolyl isomerase H (cyclophilin H) |
| chr2_-_51507540 | 0.21 |
ENSDART00000166605
ENSDART00000161093 |
pigrl2.3
|
polymeric immunoglobulin receptor-like 2.3 |
| chr20_+_51478939 | 0.21 |
ENSDART00000149758
|
tlr5a
|
toll-like receptor 5a |
| chr20_-_22798794 | 0.20 |
ENSDART00000148084
|
fip1l1a
|
FIP1 like 1a (S. cerevisiae) |
| chr15_+_17030473 | 0.20 |
ENSDART00000129407
|
plin2
|
perilipin 2 |
| chr2_+_25315591 | 0.20 |
ENSDART00000161386
|
fndc3ba
|
fibronectin type III domain containing 3Ba |
| chr20_-_52271262 | 0.20 |
ENSDART00000135463
|
rhpn1
|
rhophilin, Rho GTPase binding protein 1 |
| chr16_+_52343905 | 0.20 |
ENSDART00000131051
|
ifnlr1
|
interferon lambda receptor 1 |
| chr3_+_23029934 | 0.20 |
ENSDART00000110343
|
nags
|
N-acetylglutamate synthase |
| chr20_+_13781779 | 0.19 |
ENSDART00000142999
ENSDART00000152471 |
lpgat1
|
lysophosphatidylglycerol acyltransferase 1 |
| chr16_-_24708586 | 0.19 |
ENSDART00000178614
|
BX005256.1
|
|
| chr20_-_52271015 | 0.19 |
ENSDART00000074307
|
rhpn1
|
rhophilin, Rho GTPase binding protein 1 |
| chr21_+_6212844 | 0.19 |
ENSDART00000150301
|
fnbp1b
|
formin binding protein 1b |
| chr22_+_14117078 | 0.18 |
ENSDART00000013575
|
bzw1a
|
basic leucine zipper and W2 domains 1a |
| chr4_-_58448785 | 0.18 |
ENSDART00000172359
|
si:ch211-212k5.3
|
si:ch211-212k5.3 |
| chr23_-_40766518 | 0.18 |
ENSDART00000127420
|
si:dkeyp-27c8.2
|
si:dkeyp-27c8.2 |
| chr24_-_12832499 | 0.17 |
ENSDART00000182027
ENSDART00000039312 |
ipo4
|
importin 4 |
| chr3_-_60175470 | 0.17 |
ENSDART00000156597
|
si:ch73-364h19.1
|
si:ch73-364h19.1 |
| chr12_+_48803098 | 0.17 |
ENSDART00000074768
|
ppifb
|
peptidylprolyl isomerase Fb |
| chr21_-_5799122 | 0.17 |
ENSDART00000129351
ENSDART00000151202 |
ccni
|
cyclin I |
| chr22_+_34691020 | 0.17 |
ENSDART00000192202
|
hykk.1
|
hydroxylysine kinase, tandem duplicate 1 |
| chr10_-_17083180 | 0.16 |
ENSDART00000170083
|
fam166b
|
family with sequence similarity 166, member B |
| chr17_+_24843401 | 0.16 |
ENSDART00000110179
|
cx34.4
|
connexin 34.4 |
| chr2_+_10007113 | 0.16 |
ENSDART00000155213
|
slc35a3b
|
solute carrier family 35 (UDP-N-acetylglucosamine (UDP-GlcNAc) transporter), member A3b |
| chr20_+_51479263 | 0.16 |
ENSDART00000148798
|
tlr5a
|
toll-like receptor 5a |
| chr17_+_12408188 | 0.16 |
ENSDART00000105218
|
khk
|
ketohexokinase |
| chr3_+_32080403 | 0.15 |
ENSDART00000124943
|
aldh16a1
|
aldehyde dehydrogenase 16 family, member A1 |
| chr19_+_31904836 | 0.15 |
ENSDART00000162297
ENSDART00000088340 ENSDART00000151280 ENSDART00000151218 |
tpd52
|
tumor protein D52 |
| chr4_-_54100377 | 0.15 |
ENSDART00000188064
|
si:dkey-199m13.7
|
si:dkey-199m13.7 |
| chr8_+_23381892 | 0.14 |
ENSDART00000180950
ENSDART00000063010 ENSDART00000074241 ENSDART00000142783 |
mapre1a
|
microtubule-associated protein, RP/EB family, member 1a |
| chr3_-_5394800 | 0.14 |
ENSDART00000181938
|
si:ch73-106l15.2
|
si:ch73-106l15.2 |
| chr8_-_53735399 | 0.14 |
ENSDART00000187005
|
tlr9
|
toll-like receptor 9 |
| chr8_+_28259347 | 0.14 |
ENSDART00000110857
|
fam212b
|
family with sequence similarity 212, member B |
| chr7_+_33130639 | 0.14 |
ENSDART00000142450
ENSDART00000173967 ENSDART00000173832 |
zgc:153219
si:ch211-194p6.7
|
zgc:153219 si:ch211-194p6.7 |
| chr25_-_4192637 | 0.14 |
ENSDART00000153832
|
ppp1r32
|
protein phosphatase 1, regulatory subunit 32 |
| chr25_-_3192405 | 0.13 |
ENSDART00000104835
|
hps5
|
Hermansky-Pudlak syndrome 5 |
| chr5_+_59234421 | 0.13 |
ENSDART00000045518
|
prkrip1
|
PRKR interacting protein 1 |
| chr6_-_49078263 | 0.13 |
ENSDART00000032982
|
slc5a8l
|
solute carrier family 5 (iodide transporter), member 8-like |
| chr20_+_42563618 | 0.13 |
ENSDART00000153441
|
igf2r
|
insulin-like growth factor 2 receptor |
| chr7_+_17467551 | 0.13 |
ENSDART00000101968
|
nitr1b
|
novel immune-type receptor 1b |
| chr2_+_9970942 | 0.13 |
ENSDART00000153063
|
sec22ba
|
SEC22 homolog B, vesicle trafficking protein a |
| chr2_-_2451181 | 0.13 |
ENSDART00000101033
|
pth1ra
|
parathyroid hormone 1 receptor a |
| chr5_+_43870389 | 0.13 |
ENSDART00000141002
|
zgc:112966
|
zgc:112966 |
| chr15_+_5277761 | 0.12 |
ENSDART00000153954
|
si:ch1073-166e24.4
|
si:ch1073-166e24.4 |
| chr22_+_1501566 | 0.12 |
ENSDART00000167788
|
AL954741.3
|
|
| chr4_+_13568469 | 0.12 |
ENSDART00000171235
ENSDART00000136152 |
calua
|
calumenin a |
| chr5_-_37247654 | 0.12 |
ENSDART00000112312
|
lrch2
|
leucine-rich repeats and calponin homology (CH) domain containing 2 |
| chr3_-_9444749 | 0.12 |
ENSDART00000182191
|
FO904885.3
|
|
| chr20_-_35052464 | 0.12 |
ENSDART00000037195
|
kif26bb
|
kinesin family member 26Bb |
| chr15_+_17031111 | 0.12 |
ENSDART00000175378
|
plin2
|
perilipin 2 |
| chr9_-_34308113 | 0.11 |
ENSDART00000137806
|
gpa33
|
glycoprotein A33 (transmembrane) |
| chr9_+_18829360 | 0.11 |
ENSDART00000006514
|
gtf2f2b
|
general transcription factor IIF, polypeptide 2b |
| chr2_+_10006839 | 0.11 |
ENSDART00000160304
|
slc35a3b
|
solute carrier family 35 (UDP-N-acetylglucosamine (UDP-GlcNAc) transporter), member A3b |
| chr17_-_11466700 | 0.11 |
ENSDART00000091159
|
adpgk2
|
ADP-dependent glucokinase 2 |
| chr20_-_284865 | 0.11 |
ENSDART00000104806
|
wisp3
|
WNT1 inducible signaling pathway protein 3 |
| chr18_+_5308392 | 0.10 |
ENSDART00000179072
|
DUT
|
deoxyuridine triphosphatase |
| chr14_-_46310549 | 0.10 |
ENSDART00000123832
|
crybb1l1
|
crystallin, beta B1, like 1 |
| chr17_-_45383699 | 0.10 |
ENSDART00000141182
|
tmem206
|
transmembrane protein 206 |
| chr2_+_307665 | 0.10 |
ENSDART00000082083
|
zgc:113452
|
zgc:113452 |
| chr16_-_31342356 | 0.10 |
ENSDART00000136648
|
mroh1
|
maestro heat-like repeat family member 1 |
| chr18_+_33591704 | 0.09 |
ENSDART00000185631
ENSDART00000057845 |
si:dkey-47k20.5
|
si:dkey-47k20.5 |
| chr17_-_12408109 | 0.09 |
ENSDART00000155509
|
ankef1b
|
ankyrin repeat and EF-hand domain containing 1b |
| chr5_-_61588998 | 0.09 |
ENSDART00000050912
|
pex12
|
peroxisomal biogenesis factor 12 |
| chr1_-_354115 | 0.09 |
ENSDART00000141590
ENSDART00000098627 |
pros1
|
protein S |
| chr6_+_49995781 | 0.09 |
ENSDART00000075009
|
eif2s2
|
eukaryotic translation initiation factor 2, subunit 2 beta |
| chr6_-_9563534 | 0.09 |
ENSDART00000184765
|
map3k2
|
mitogen-activated protein kinase kinase kinase 2 |
| chr15_+_17074185 | 0.08 |
ENSDART00000046648
|
clptm1
|
cleft lip and palate associated transmembrane protein 1 |
| chr10_-_16868211 | 0.08 |
ENSDART00000171755
|
stoml2
|
stomatin (EPB72)-like 2 |
| chr15_+_45640906 | 0.08 |
ENSDART00000149361
ENSDART00000149079 |
sagb
|
S-antigen; retina and pineal gland (arrestin) b |
| chr4_+_308697 | 0.08 |
ENSDART00000174021
ENSDART00000131419 ENSDART00000067489 |
si:ch211-51e12.7
|
si:ch211-51e12.7 |
| chr20_+_21595244 | 0.08 |
ENSDART00000010643
|
esr2a
|
estrogen receptor 2a |
| chr23_+_45025909 | 0.08 |
ENSDART00000188105
|
hmgb2b
|
high mobility group box 2b |
| chr4_-_48578324 | 0.07 |
ENSDART00000122707
|
znf980
|
zinc finger protein 980 |
| chr25_+_10843985 | 0.07 |
ENSDART00000090386
ENSDART00000154975 |
si:ch211-147g22.4
|
si:ch211-147g22.4 |
| chr1_+_30946231 | 0.07 |
ENSDART00000022841
|
metap1d
|
methionyl aminopeptidase type 1D (mitochondrial) |
| chr3_+_13929860 | 0.06 |
ENSDART00000164179
|
syce2
|
synaptonemal complex central element protein 2 |
| chr12_-_27242498 | 0.06 |
ENSDART00000152609
ENSDART00000152170 |
si:dkey-11c5.11
|
si:dkey-11c5.11 |
| chr9_-_950004 | 0.06 |
ENSDART00000168917
|
SCTR
|
secretin receptor |
| chr23_+_41679586 | 0.05 |
ENSDART00000067662
|
CU914487.1
|
|
| chr4_-_1684335 | 0.05 |
ENSDART00000019144
ENSDART00000113360 |
arid2
|
AT rich interactive domain 2 (ARID, RFX-like) |
| chr12_-_4220713 | 0.05 |
ENSDART00000129427
|
vkorc1
|
vitamin K epoxide reductase complex, subunit 1 |
| chr24_-_41180149 | 0.05 |
ENSDART00000019975
|
acvr2ba
|
activin A receptor type 2Ba |
| chr12_+_19138452 | 0.04 |
ENSDART00000141346
ENSDART00000066397 |
phf5a
|
PHD finger protein 5A |
| chr8_+_26226044 | 0.04 |
ENSDART00000186503
|
wdr6
|
WD repeat domain 6 |
| chr25_+_4979446 | 0.04 |
ENSDART00000154131
ENSDART00000155537 |
si:ch73-265h17.1
|
si:ch73-265h17.1 |
| chr3_+_46444999 | 0.04 |
ENSDART00000186530
|
si:ch211-66e2.3
|
si:ch211-66e2.3 |
| chr18_-_41160579 | 0.04 |
ENSDART00000181480
|
CABZ01005876.1
|
|
| chr7_-_63637644 | 0.04 |
ENSDART00000145168
|
si:ch211-208g1.1
|
si:ch211-208g1.1 |
| chr12_+_6117682 | 0.04 |
ENSDART00000146650
|
asah2
|
N-acylsphingosine amidohydrolase 2 |
| chr4_+_28356606 | 0.04 |
ENSDART00000192995
|
si:ch73-263o4.4
|
si:ch73-263o4.4 |
| chr8_-_16675464 | 0.04 |
ENSDART00000191982
|
osbpl9
|
oxysterol binding protein-like 9 |
| chr18_-_26797723 | 0.04 |
ENSDART00000008013
|
sec11a
|
SEC11 homolog A, signal peptidase complex subunit |
| chr6_+_9870192 | 0.04 |
ENSDART00000150894
|
MPP4 (1 of many)
|
si:ch211-222n4.6 |
| chr16_-_28268201 | 0.04 |
ENSDART00000121671
ENSDART00000141911 |
si:dkey-12j5.1
|
si:dkey-12j5.1 |
| chr2_+_38025260 | 0.04 |
ENSDART00000075905
|
hnrnpc
|
heterogeneous nuclear ribonucleoprotein C |
Network of associatons between targets according to the STRING database.
First level regulatory network of mybl2b
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.5 | GO:0071706 | tumor necrosis factor production(GO:0032640) tumor necrosis factor superfamily cytokine production(GO:0071706) |
| 0.2 | 1.1 | GO:1902315 | cell cycle DNA replication initiation(GO:1902292) nuclear cell cycle DNA replication initiation(GO:1902315) mitotic DNA replication initiation(GO:1902975) |
| 0.1 | 0.6 | GO:1901492 | positive regulation of lymphangiogenesis(GO:1901492) |
| 0.1 | 0.4 | GO:0003250 | cell proliferation involved in heart valve morphogenesis(GO:0003249) regulation of cell proliferation involved in heart valve morphogenesis(GO:0003250) endocardial cell development(GO:0060958) cell proliferation involved in heart valve development(GO:2000793) |
| 0.1 | 0.6 | GO:0030326 | embryonic limb morphogenesis(GO:0030326) |
| 0.1 | 0.5 | GO:0006526 | arginine biosynthetic process(GO:0006526) |
| 0.1 | 0.6 | GO:0019805 | quinolinate biosynthetic process(GO:0019805) quinolinate metabolic process(GO:0046874) |
| 0.1 | 0.4 | GO:0034146 | toll-like receptor 5 signaling pathway(GO:0034146) |
| 0.1 | 0.6 | GO:0006556 | S-adenosylmethionine biosynthetic process(GO:0006556) |
| 0.1 | 1.1 | GO:0034724 | DNA replication-independent nucleosome organization(GO:0034724) |
| 0.1 | 0.3 | GO:0009447 | polyamine catabolic process(GO:0006598) putrescine catabolic process(GO:0009447) |
| 0.1 | 0.6 | GO:0010890 | positive regulation of sequestering of triglyceride(GO:0010890) |
| 0.1 | 0.5 | GO:0042554 | superoxide anion generation(GO:0042554) |
| 0.1 | 0.7 | GO:0006032 | chitin catabolic process(GO:0006032) |
| 0.1 | 0.2 | GO:0051352 | negative regulation of ligase activity(GO:0051352) negative regulation of ubiquitin-protein transferase activity(GO:0051444) negative regulation of ubiquitin protein ligase activity(GO:1904667) |
| 0.1 | 0.6 | GO:0009263 | deoxyribonucleotide biosynthetic process(GO:0009263) |
| 0.1 | 0.5 | GO:0035372 | protein localization to microtubule(GO:0035372) protein localization to microtubule plus-end(GO:1904825) |
| 0.1 | 0.3 | GO:0023041 | neuronal signal transduction(GO:0023041) |
| 0.0 | 0.2 | GO:0018197 | peptidyl-aspartic acid modification(GO:0018197) peptidyl-aspartic acid hydroxylation(GO:0042264) |
| 0.0 | 0.2 | GO:0036149 | phosphatidylinositol acyl-chain remodeling(GO:0036149) |
| 0.0 | 0.3 | GO:0003188 | heart valve formation(GO:0003188) atrioventricular valve formation(GO:0003190) |
| 0.0 | 0.2 | GO:0015074 | DNA integration(GO:0015074) |
| 0.0 | 1.7 | GO:0048538 | thymus development(GO:0048538) |
| 0.0 | 0.4 | GO:0045003 | double-strand break repair via synthesis-dependent strand annealing(GO:0045003) |
| 0.0 | 0.5 | GO:0042531 | positive regulation of tyrosine phosphorylation of STAT protein(GO:0042531) |
| 0.0 | 0.3 | GO:0050729 | positive regulation of inflammatory response(GO:0050729) |
| 0.0 | 0.7 | GO:0010972 | negative regulation of G2/M transition of mitotic cell cycle(GO:0010972) |
| 0.0 | 0.1 | GO:0042730 | fibrinolysis(GO:0042730) |
| 0.0 | 1.1 | GO:0001878 | response to yeast(GO:0001878) |
| 0.0 | 0.4 | GO:2000601 | positive regulation of actin nucleation(GO:0051127) positive regulation of Arp2/3 complex-mediated actin nucleation(GO:2000601) |
| 0.0 | 0.4 | GO:0000479 | endonucleolytic cleavage involved in rRNA processing(GO:0000478) endonucleolytic cleavage of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000479) |
| 0.0 | 0.4 | GO:0048385 | regulation of retinoic acid receptor signaling pathway(GO:0048385) |
| 0.0 | 0.8 | GO:0006383 | transcription from RNA polymerase III promoter(GO:0006383) |
| 0.0 | 0.2 | GO:0098787 | mRNA cleavage involved in mRNA processing(GO:0098787) pre-mRNA cleavage required for polyadenylation(GO:0098789) |
| 0.0 | 0.3 | GO:0007076 | mitotic chromosome condensation(GO:0007076) |
| 0.0 | 0.3 | GO:0051382 | kinetochore assembly(GO:0051382) |
| 0.0 | 0.2 | GO:0042537 | benzene-containing compound metabolic process(GO:0042537) |
| 0.0 | 0.3 | GO:0071156 | regulation of cell cycle arrest(GO:0071156) |
| 0.0 | 0.3 | GO:0007339 | binding of sperm to zona pellucida(GO:0007339) sperm-egg recognition(GO:0035036) egg coat formation(GO:0035803) regulation of acrosome reaction(GO:0060046) positive regulation of acrosome reaction(GO:2000344) |
| 0.0 | 0.6 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.0 | 0.1 | GO:0048069 | eye pigmentation(GO:0048069) |
| 0.0 | 0.6 | GO:0051014 | actin filament severing(GO:0051014) |
| 0.0 | 0.3 | GO:0090481 | pyrimidine nucleotide-sugar transport(GO:0015781) pyrimidine nucleotide-sugar transmembrane transport(GO:0090481) |
| 0.0 | 0.3 | GO:0030259 | lipid glycosylation(GO:0030259) |
| 0.0 | 0.1 | GO:0070084 | protein initiator methionine removal(GO:0070084) |
| 0.0 | 0.3 | GO:0033209 | tumor necrosis factor-mediated signaling pathway(GO:0033209) |
| 0.0 | 0.5 | GO:0007050 | cell cycle arrest(GO:0007050) |
| 0.0 | 0.3 | GO:0000028 | ribosomal small subunit assembly(GO:0000028) |
| 0.0 | 0.6 | GO:0071222 | cellular response to molecule of bacterial origin(GO:0071219) cellular response to lipopolysaccharide(GO:0071222) |
| 0.0 | 0.0 | GO:0042373 | vitamin K metabolic process(GO:0042373) |
| 0.0 | 0.5 | GO:0002377 | immunoglobulin production(GO:0002377) |
| 0.0 | 0.1 | GO:0048280 | vesicle fusion with Golgi apparatus(GO:0048280) |
| 0.0 | 0.2 | GO:0018216 | peptidyl-arginine methylation(GO:0018216) |
| 0.0 | 0.2 | GO:0046686 | response to cadmium ion(GO:0046686) |
| 0.0 | 0.2 | GO:0006298 | mismatch repair(GO:0006298) |
| 0.0 | 0.2 | GO:0021551 | central nervous system morphogenesis(GO:0021551) |
| 0.0 | 0.4 | GO:0017121 | phospholipid scrambling(GO:0017121) |
| 0.0 | 0.9 | GO:0006352 | DNA-templated transcription, initiation(GO:0006352) |
| 0.0 | 0.2 | GO:0010043 | response to zinc ion(GO:0010043) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.5 | GO:0097058 | CRLF-CLCF1 complex(GO:0097058) |
| 0.1 | 0.6 | GO:0048269 | methionine adenosyltransferase complex(GO:0048269) |
| 0.1 | 0.4 | GO:0070319 | Golgi to plasma membrane transport vesicle(GO:0070319) |
| 0.1 | 1.1 | GO:0035101 | FACT complex(GO:0035101) |
| 0.1 | 0.5 | GO:0000811 | GINS complex(GO:0000811) |
| 0.1 | 0.5 | GO:0043020 | NADPH oxidase complex(GO:0043020) |
| 0.1 | 0.3 | GO:0031515 | tRNA (m1A) methyltransferase complex(GO:0031515) |
| 0.0 | 0.6 | GO:0042555 | MCM complex(GO:0042555) |
| 0.0 | 0.3 | GO:0000796 | condensin complex(GO:0000796) |
| 0.0 | 0.3 | GO:0071005 | U2-type precatalytic spliceosome(GO:0071005) |
| 0.0 | 0.8 | GO:0005666 | DNA-directed RNA polymerase III complex(GO:0005666) |
| 0.0 | 0.8 | GO:0045180 | basal cortex(GO:0045180) |
| 0.0 | 0.1 | GO:1990415 | Pex17p-Pex14p docking complex(GO:1990415) peroxisomal importomer complex(GO:1990429) |
| 0.0 | 0.3 | GO:0005832 | chaperonin-containing T-complex(GO:0005832) |
| 0.0 | 0.5 | GO:0035371 | microtubule plus-end(GO:0035371) |
| 0.0 | 0.1 | GO:0005850 | eukaryotic translation initiation factor 2 complex(GO:0005850) |
| 0.0 | 0.1 | GO:0033018 | sarcoplasmic reticulum lumen(GO:0033018) |
| 0.0 | 0.2 | GO:0005847 | mRNA cleavage and polyadenylation specificity factor complex(GO:0005847) |
| 0.0 | 0.1 | GO:0000801 | central element(GO:0000801) |
| 0.0 | 0.3 | GO:0030867 | rough endoplasmic reticulum membrane(GO:0030867) |
| 0.0 | 0.6 | GO:0005811 | lipid particle(GO:0005811) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.5 | GO:0034618 | arginine binding(GO:0034618) |
| 0.1 | 0.6 | GO:0051916 | granulocyte colony-stimulating factor binding(GO:0051916) |
| 0.1 | 0.6 | GO:0004478 | methionine adenosyltransferase activity(GO:0004478) |
| 0.1 | 0.6 | GO:0070180 | large ribosomal subunit rRNA binding(GO:0070180) |
| 0.1 | 0.6 | GO:0016728 | ribonucleoside-diphosphate reductase activity, thioredoxin disulfide as acceptor(GO:0004748) oxidoreductase activity, acting on CH or CH2 groups, disulfide as acceptor(GO:0016728) ribonucleoside-diphosphate reductase activity(GO:0061731) |
| 0.1 | 0.2 | GO:0045145 | single-stranded DNA 5'-3' exodeoxyribonuclease activity(GO:0045145) |
| 0.1 | 0.3 | GO:0003979 | UDP-glucose 6-dehydrogenase activity(GO:0003979) |
| 0.1 | 0.2 | GO:0019777 | Atg12 transferase activity(GO:0019777) |
| 0.1 | 0.4 | GO:0015616 | DNA translocase activity(GO:0015616) |
| 0.1 | 0.3 | GO:0005031 | tumor necrosis factor-activated receptor activity(GO:0005031) |
| 0.1 | 0.3 | GO:0030620 | U2 snRNA binding(GO:0030620) |
| 0.1 | 0.7 | GO:0004568 | chitinase activity(GO:0004568) |
| 0.1 | 0.2 | GO:0004597 | peptide-aspartate beta-dioxygenase activity(GO:0004597) |
| 0.1 | 0.5 | GO:0016175 | superoxide-generating NADPH oxidase activity(GO:0016175) |
| 0.1 | 0.2 | GO:0003988 | acetyl-CoA C-acyltransferase activity(GO:0003988) |
| 0.1 | 0.2 | GO:0050508 | glucuronosyl-N-acetylglucosaminyl-proteoglycan 4-alpha-N-acetylglucosaminyltransferase activity(GO:0050508) |
| 0.1 | 0.3 | GO:0016429 | tRNA (adenine-N1-)-methyltransferase activity(GO:0016429) |
| 0.1 | 0.6 | GO:0043142 | single-stranded DNA-dependent ATP-dependent DNA helicase activity(GO:0017116) single-stranded DNA-dependent ATPase activity(GO:0043142) |
| 0.1 | 0.4 | GO:0001972 | retinoic acid binding(GO:0001972) |
| 0.0 | 0.3 | GO:0019808 | polyamine binding(GO:0019808) spermidine binding(GO:0019809) |
| 0.0 | 2.1 | GO:0031491 | nucleosome binding(GO:0031491) |
| 0.0 | 0.6 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 0.0 | 0.2 | GO:0031957 | very long-chain fatty acid-CoA ligase activity(GO:0031957) |
| 0.0 | 0.1 | GO:0015355 | secondary active monocarboxylate transmembrane transporter activity(GO:0015355) |
| 0.0 | 0.3 | GO:0035804 | structural constituent of egg coat(GO:0035804) |
| 0.0 | 0.1 | GO:0008022 | protein C-terminus binding(GO:0008022) |
| 0.0 | 0.4 | GO:0017128 | phospholipid scramblase activity(GO:0017128) |
| 0.0 | 0.2 | GO:0019202 | amino acid kinase activity(GO:0019202) |
| 0.0 | 0.3 | GO:0005459 | UDP-galactose transmembrane transporter activity(GO:0005459) |
| 0.0 | 0.2 | GO:0003993 | acid phosphatase activity(GO:0003993) |
| 0.0 | 0.8 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.0 | 0.2 | GO:0016274 | arginine N-methyltransferase activity(GO:0016273) protein-arginine N-methyltransferase activity(GO:0016274) |
| 0.0 | 0.6 | GO:0016831 | carboxy-lyase activity(GO:0016831) |
| 0.0 | 0.3 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.0 | 0.1 | GO:1903924 | estradiol binding(GO:1903924) |
| 0.0 | 0.6 | GO:0005546 | phosphatidylinositol-4,5-bisphosphate binding(GO:0005546) |
| 0.0 | 0.2 | GO:0061608 | nuclear import signal receptor activity(GO:0061608) |
| 0.0 | 0.1 | GO:0005537 | mannose binding(GO:0005537) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.0 | 0.3 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.0 | 0.4 | PID IL1 PATHWAY | IL1-mediated signaling events |
| 0.0 | 0.5 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 0.0 | 0.6 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.0 | 0.3 | PID NECTIN PATHWAY | Nectin adhesion pathway |
| 0.0 | 0.3 | PID CD40 PATHWAY | CD40/CD40L signaling |
| 0.0 | 0.6 | PID ATR PATHWAY | ATR signaling pathway |
| 0.0 | 0.3 | PID DNA PK PATHWAY | DNA-PK pathway in nonhomologous end joining |
| 0.0 | 0.3 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | REACTOME BETA DEFENSINS | Genes involved in Beta defensins |
| 0.1 | 0.7 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.0 | 0.6 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 0.0 | 1.0 | REACTOME CONVERSION FROM APC C CDC20 TO APC C CDH1 IN LATE ANAPHASE | Genes involved in Conversion from APC/C:Cdc20 to APC/C:Cdh1 in late anaphase |
| 0.0 | 0.6 | REACTOME UNWINDING OF DNA | Genes involved in Unwinding of DNA |
| 0.0 | 1.1 | REACTOME ELONGATION ARREST AND RECOVERY | Genes involved in Elongation arrest and recovery |
| 0.0 | 0.6 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.0 | 0.3 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.0 | 0.2 | REACTOME ACYL CHAIN REMODELLING OF PG | Genes involved in Acyl chain remodelling of PG |
| 0.0 | 0.2 | REACTOME ALPHA LINOLENIC ACID ALA METABOLISM | Genes involved in alpha-linolenic acid (ALA) metabolism |
| 0.0 | 0.3 | REACTOME DEPOSITION OF NEW CENPA CONTAINING NUCLEOSOMES AT THE CENTROMERE | Genes involved in Deposition of New CENPA-containing Nucleosomes at the Centromere |
| 0.0 | 1.7 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 0.0 | 0.3 | REACTOME FORMATION OF TUBULIN FOLDING INTERMEDIATES BY CCT TRIC | Genes involved in Formation of tubulin folding intermediates by CCT/TriC |
| 0.0 | 0.3 | REACTOME PHASE II CONJUGATION | Genes involved in Phase II conjugation |
| 0.0 | 0.3 | REACTOME MRNA SPLICING MINOR PATHWAY | Genes involved in mRNA Splicing - Minor Pathway |
| 0.0 | 0.1 | REACTOME TRAFFICKING AND PROCESSING OF ENDOSOMAL TLR | Genes involved in Trafficking and processing of endosomal TLR |
| 0.0 | 0.4 | REACTOME IL1 SIGNALING | Genes involved in Interleukin-1 signaling |