Project
PRJNA207719: Tissue specific transcriptome profiling
Navigation
Downloads
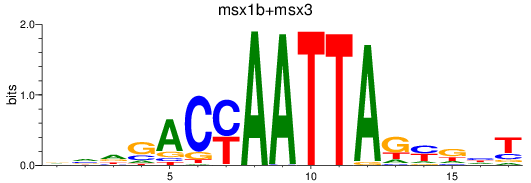
Results for msx1b+msx3
Z-value: 1.56
Transcription factors associated with msx1b+msx3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
msx1b
|
ENSDARG00000008886 | muscle segment homeobox 1b |
|
msx3
|
ENSDARG00000015674 | muscle segment homeobox 3 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| msx3 | dr11_v1_chr13_+_24662238_24662238 | -0.82 | 8.8e-02 | Click! |
| msx1b | dr11_v1_chr1_+_49352900_49352900 | -0.48 | 4.1e-01 | Click! |
Activity profile of msx1b+msx3 motif
Sorted Z-values of msx1b+msx3 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr21_-_22648007 | 1.66 |
ENSDART00000121788
|
gig2l
|
grass carp reovirus (GCRV)-induced gene 2l |
| chr20_+_46040666 | 1.34 |
ENSDART00000060744
|
si:dkey-7c18.24
|
si:dkey-7c18.24 |
| chr15_-_5815006 | 1.33 |
ENSDART00000102459
|
rbp2a
|
retinol binding protein 2a, cellular |
| chr11_-_40457325 | 1.23 |
ENSDART00000128442
|
tnfrsf1b
|
tumor necrosis factor receptor superfamily, member 1B |
| chr21_-_22647695 | 1.20 |
ENSDART00000187553
|
gig2l
|
grass carp reovirus (GCRV)-induced gene 2l |
| chr25_+_29160102 | 1.12 |
ENSDART00000162854
|
pkmb
|
pyruvate kinase M1/2b |
| chr22_+_12770877 | 1.11 |
ENSDART00000044683
|
ftcd
|
formimidoyltransferase cyclodeaminase |
| chr5_-_29534748 | 1.09 |
ENSDART00000159587
|
AL831768.1
|
|
| chr24_+_20373605 | 1.01 |
ENSDART00000112349
|
plcd1a
|
phospholipase C, delta 1a |
| chr6_+_50451337 | 1.00 |
ENSDART00000155051
|
mych
|
myelocytomatosis oncogene homolog |
| chr12_+_28749189 | 1.00 |
ENSDART00000013980
|
tbx21
|
T-box 21 |
| chr4_-_9891874 | 1.00 |
ENSDART00000067193
|
adm2a
|
adrenomedullin 2a |
| chr25_-_25550938 | 0.99 |
ENSDART00000150412
ENSDART00000103622 |
irf7
|
interferon regulatory factor 7 |
| chr14_-_1543834 | 0.98 |
ENSDART00000185876
|
CABZ01109480.1
|
|
| chr13_+_22476742 | 0.93 |
ENSDART00000078759
ENSDART00000130101 ENSDART00000137220 ENSDART00000133065 ENSDART00000147348 |
ldb3a
|
LIM domain binding 3a |
| chr11_-_2478374 | 0.90 |
ENSDART00000173205
|
si:ch73-267c23.10
|
si:ch73-267c23.10 |
| chr3_+_23692462 | 0.89 |
ENSDART00000145934
|
hoxb7a
|
homeobox B7a |
| chr24_-_9979342 | 0.86 |
ENSDART00000138576
ENSDART00000191206 |
zgc:171977
|
zgc:171977 |
| chr5_+_29794058 | 0.86 |
ENSDART00000045410
|
thy1
|
Thy-1 cell surface antigen |
| chr20_-_40755614 | 0.84 |
ENSDART00000061247
|
cx32.3
|
connexin 32.3 |
| chr6_-_19042294 | 0.83 |
ENSDART00000159461
|
si:rp71-81e14.2
|
si:rp71-81e14.2 |
| chr19_-_8768564 | 0.79 |
ENSDART00000170416
|
si:ch73-350k19.1
|
si:ch73-350k19.1 |
| chr10_-_44560165 | 0.79 |
ENSDART00000181217
ENSDART00000076084 |
npm2b
|
nucleophosmin/nucleoplasmin, 2b |
| chr15_-_37683768 | 0.79 |
ENSDART00000059609
|
si:dkey-117a8.3
|
si:dkey-117a8.3 |
| chr17_-_6514962 | 0.78 |
ENSDART00000163514
|
dnajc5gb
|
DnaJ (Hsp40) homolog, subfamily C, member 5 gamma b |
| chr4_+_77943184 | 0.77 |
ENSDART00000159094
|
pacsin2
|
protein kinase C and casein kinase substrate in neurons 2 |
| chr15_+_46344655 | 0.77 |
ENSDART00000155893
|
si:ch1073-340i21.2
|
si:ch1073-340i21.2 |
| chr22_+_5532003 | 0.74 |
ENSDART00000106174
|
si:ch73-256j6.2
|
si:ch73-256j6.2 |
| chr15_-_37767901 | 0.74 |
ENSDART00000154763
|
si:dkey-42l23.4
|
si:dkey-42l23.4 |
| chr7_+_8324506 | 0.73 |
ENSDART00000168110
|
si:dkey-185m8.2
|
si:dkey-185m8.2 |
| chr1_-_19402802 | 0.72 |
ENSDART00000135552
|
rbm47
|
RNA binding motif protein 47 |
| chr7_-_8374950 | 0.71 |
ENSDART00000057101
|
aep1
|
aerolysin-like protein |
| chr10_+_36178713 | 0.70 |
ENSDART00000140816
|
or108-3
|
odorant receptor, family D, subfamily 108, member 3 |
| chr8_-_53680653 | 0.69 |
ENSDART00000164739
|
wnt5a
|
wingless-type MMTV integration site family, member 5a |
| chr1_-_7975755 | 0.69 |
ENSDART00000141765
|
si:dkey-79f11.7
|
si:dkey-79f11.7 |
| chr11_+_24716837 | 0.68 |
ENSDART00000145217
|
zgc:153953
|
zgc:153953 |
| chr24_-_17039638 | 0.66 |
ENSDART00000048823
|
c8g
|
complement component 8, gamma polypeptide |
| chr20_+_11731039 | 0.64 |
ENSDART00000152215
ENSDART00000152585 |
si:ch211-155o21.3
|
si:ch211-155o21.3 |
| chr8_-_27656765 | 0.63 |
ENSDART00000078491
|
mov10b.2
|
Moloney leukemia virus 10b, tandem duplicate 2 |
| chr17_-_50071748 | 0.63 |
ENSDART00000075188
|
zgc:113886
|
zgc:113886 |
| chr1_+_29068654 | 0.63 |
ENSDART00000053932
|
cbsa
|
cystathionine-beta-synthase a |
| chr23_-_16485190 | 0.62 |
ENSDART00000155038
|
si:dkeyp-100a5.4
|
si:dkeyp-100a5.4 |
| chr11_-_25538341 | 0.61 |
ENSDART00000171560
|
si:dkey-245f22.3
|
si:dkey-245f22.3 |
| chr16_-_26855936 | 0.60 |
ENSDART00000167320
ENSDART00000078119 |
ino80c
|
INO80 complex subunit C |
| chr13_+_27328098 | 0.59 |
ENSDART00000037585
|
mb21d1
|
Mab-21 domain containing 1 |
| chr19_-_18127808 | 0.59 |
ENSDART00000108627
|
snx10a
|
sorting nexin 10a |
| chr16_+_41171949 | 0.58 |
ENSDART00000135294
|
nek11
|
NIMA-related kinase 11 |
| chr16_+_33144112 | 0.58 |
ENSDART00000183149
|
rhbdl2
|
rhomboid, veinlet-like 2 (Drosophila) |
| chr1_+_40801696 | 0.57 |
ENSDART00000147497
|
cpz
|
carboxypeptidase Z |
| chr16_+_33144306 | 0.57 |
ENSDART00000101953
|
rhbdl2
|
rhomboid, veinlet-like 2 (Drosophila) |
| chr17_+_24851951 | 0.56 |
ENSDART00000180746
|
cx35.4
|
connexin 35.4 |
| chr24_+_38306010 | 0.56 |
ENSDART00000143184
|
mybpc2b
|
myosin binding protein C, fast type b |
| chr2_+_31804582 | 0.56 |
ENSDART00000086646
|
rnf182
|
ring finger protein 182 |
| chr1_+_55730205 | 0.55 |
ENSDART00000148075
|
adgre10
|
adhesion G protein-coupled receptor E10 |
| chr15_-_37709072 | 0.55 |
ENSDART00000157115
|
si:dkey-117a8.2
|
si:dkey-117a8.2 |
| chr13_+_9468535 | 0.54 |
ENSDART00000135088
ENSDART00000164270 ENSDART00000099619 ENSDART00000164656 |
HTRA2 (1 of many)
|
si:dkey-265c15.6 |
| chr15_-_37733238 | 0.54 |
ENSDART00000115178
|
si:dkey-42l23.7
|
si:dkey-42l23.7 |
| chr18_+_50461981 | 0.54 |
ENSDART00000158761
|
CU896640.1
|
|
| chr19_-_19871211 | 0.54 |
ENSDART00000170980
|
evx1
|
even-skipped homeobox 1 |
| chr1_-_25966068 | 0.52 |
ENSDART00000137869
ENSDART00000134192 |
synpo2b
|
synaptopodin 2b |
| chr19_-_18127629 | 0.51 |
ENSDART00000187722
|
snx10a
|
sorting nexin 10a |
| chr10_+_39212898 | 0.50 |
ENSDART00000159501
|
stt3a
|
STT3A, subunit of the oligosaccharyltransferase complex (catalytic) |
| chr9_-_35633827 | 0.50 |
ENSDART00000077745
|
zp2l1
|
zona pellucida glycoprotein 2, like 1 |
| chr5_+_27897504 | 0.49 |
ENSDART00000130936
|
adam28
|
ADAM metallopeptidase domain 28 |
| chr24_-_25144441 | 0.49 |
ENSDART00000152104
|
phldb2b
|
pleckstrin homology-like domain, family B, member 2b |
| chr18_+_9171778 | 0.48 |
ENSDART00000101192
|
sema3d
|
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3D |
| chr10_-_13343831 | 0.48 |
ENSDART00000135941
|
il11ra
|
interleukin 11 receptor, alpha |
| chr1_+_40801448 | 0.47 |
ENSDART00000187394
|
cpz
|
carboxypeptidase Z |
| chr15_-_37719015 | 0.46 |
ENSDART00000154875
|
si:dkey-117a8.1
|
si:dkey-117a8.1 |
| chr3_+_2719243 | 0.46 |
ENSDART00000189256
|
CR388047.3
|
|
| chr5_+_56023186 | 0.46 |
ENSDART00000156230
|
fzd9a
|
frizzled class receptor 9a |
| chr11_+_31864921 | 0.46 |
ENSDART00000180252
|
diaph3
|
diaphanous-related formin 3 |
| chr16_-_35975254 | 0.46 |
ENSDART00000167537
|
eva1ba
|
eva-1 homolog Ba (C. elegans) |
| chr23_+_16889352 | 0.46 |
ENSDART00000145275
|
si:dkey-147f3.8
|
si:dkey-147f3.8 |
| chr2_+_30032303 | 0.45 |
ENSDART00000151841
|
rbm33b
|
RNA binding motif protein 33b |
| chr6_+_39370587 | 0.45 |
ENSDART00000157165
ENSDART00000155079 |
si:dkey-195m11.8
|
si:dkey-195m11.8 |
| chr15_-_37673733 | 0.45 |
ENSDART00000154513
|
si:dkey-117a8.4
|
si:dkey-117a8.4 |
| chr15_+_5360407 | 0.44 |
ENSDART00000110420
|
or112-1
|
odorant receptor, family A, subfamily 112, member 1 |
| chr17_+_24821627 | 0.44 |
ENSDART00000112389
|
wdr43
|
WD repeat domain 43 |
| chr3_-_31079186 | 0.42 |
ENSDART00000145636
ENSDART00000140569 |
ELOB (1 of many)
elob
|
elongin B elongin B |
| chr6_-_9695294 | 0.40 |
ENSDART00000162728
|
nop58
|
NOP58 ribonucleoprotein homolog (yeast) |
| chr18_-_16801033 | 0.40 |
ENSDART00000100100
|
admb
|
adrenomedullin b |
| chr15_+_27384798 | 0.39 |
ENSDART00000164887
|
tbx4
|
T-box 4 |
| chr17_+_7513673 | 0.38 |
ENSDART00000156674
|
klhl10b.1
|
kelch-like family member 10b, tandem duplicate 1 |
| chr24_-_4782052 | 0.38 |
ENSDART00000149911
|
agtr1b
|
angiotensin II receptor, type 1b |
| chr18_-_46380471 | 0.38 |
ENSDART00000144444
ENSDART00000086772 |
slc19a3b
|
solute carrier family 19 (thiamine transporter), member 3b |
| chr13_+_15701596 | 0.37 |
ENSDART00000130832
|
trmt61a
|
tRNA methyltransferase 61A |
| chr4_-_75681004 | 0.37 |
ENSDART00000186296
|
si:dkey-71l4.5
|
si:dkey-71l4.5 |
| chr4_-_14162327 | 0.35 |
ENSDART00000138656
|
si:dkey-234l24.1
|
si:dkey-234l24.1 |
| chr16_+_35595312 | 0.34 |
ENSDART00000170438
|
si:ch211-1i11.3
|
si:ch211-1i11.3 |
| chr25_+_10947743 | 0.34 |
ENSDART00000073381
|
mhc1lfa
|
major histocompatibility complex class I LFA |
| chr13_+_35955562 | 0.33 |
ENSDART00000137377
|
si:ch211-67f13.7
|
si:ch211-67f13.7 |
| chr12_+_3078221 | 0.33 |
ENSDART00000148835
ENSDART00000149427 |
sgca
|
sarcoglycan, alpha |
| chr24_+_22485710 | 0.31 |
ENSDART00000146058
|
si:dkey-40h20.1
|
si:dkey-40h20.1 |
| chr10_-_40416054 | 0.31 |
ENSDART00000137947
ENSDART00000143606 |
taar20j
|
trace amine associated receptor 20j |
| chr2_+_10771787 | 0.31 |
ENSDART00000187782
|
gfi1aa
|
growth factor independent 1A transcription repressor a |
| chr9_+_21277846 | 0.30 |
ENSDART00000139620
ENSDART00000110996 ENSDART00000111899 |
lats2
|
large tumor suppressor kinase 2 |
| chr22_+_1615691 | 0.30 |
ENSDART00000171555
|
si:ch211-255f4.13
|
si:ch211-255f4.13 |
| chr20_+_25568694 | 0.30 |
ENSDART00000063107
ENSDART00000063128 |
cyp2p7
|
cytochrome P450, family 2, subfamily P, polypeptide 7 |
| chr18_+_48423973 | 0.29 |
ENSDART00000184233
ENSDART00000147074 |
fli1a
|
Fli-1 proto-oncogene, ETS transcription factor a |
| chr4_-_20292821 | 0.29 |
ENSDART00000136069
ENSDART00000192504 |
cacna2d4a
|
calcium channel, voltage-dependent, alpha 2/delta subunit 4a |
| chr22_+_9114416 | 0.29 |
ENSDART00000190169
|
nlrp15
|
NACHT, LRR and PYD domains-containing protein 15 |
| chr23_-_35483163 | 0.29 |
ENSDART00000138660
ENSDART00000113643 ENSDART00000189269 |
fbxo25
|
F-box protein 25 |
| chr14_+_6535426 | 0.28 |
ENSDART00000055961
|
thg1l
|
tRNA-histidine guanylyltransferase 1-like |
| chr25_+_19666621 | 0.28 |
ENSDART00000112746
|
zgc:171459
|
zgc:171459 |
| chr5_+_7564644 | 0.28 |
ENSDART00000192173
|
CABZ01039096.1
|
|
| chr10_+_39212601 | 0.28 |
ENSDART00000168778
|
stt3a
|
STT3A, subunit of the oligosaccharyltransferase complex (catalytic) |
| chr20_-_9095105 | 0.28 |
ENSDART00000140792
|
oma1
|
OMA1 zinc metallopeptidase |
| chr16_+_9713850 | 0.27 |
ENSDART00000164103
|
ecm1b
|
extracellular matrix protein 1b |
| chr9_-_22821901 | 0.27 |
ENSDART00000101711
|
neb
|
nebulin |
| chr15_-_37620167 | 0.27 |
ENSDART00000059582
|
si:ch211-137j23.8
|
si:ch211-137j23.8 |
| chr4_-_42408339 | 0.27 |
ENSDART00000172612
|
si:ch211-59d8.3
|
si:ch211-59d8.3 |
| chr4_+_62576422 | 0.25 |
ENSDART00000180679
|
si:ch211-79g12.2
|
si:ch211-79g12.2 |
| chr5_+_4806851 | 0.24 |
ENSDART00000067599
|
angptl2a
|
angiopoietin-like 2a |
| chr16_-_41714988 | 0.23 |
ENSDART00000138798
|
cep85
|
centrosomal protein 85 |
| chr11_+_37251825 | 0.23 |
ENSDART00000169804
|
il17rc
|
interleukin 17 receptor C |
| chr10_+_21867307 | 0.23 |
ENSDART00000126629
|
cbln17
|
cerebellin 17 |
| chr18_+_43891051 | 0.23 |
ENSDART00000024213
|
cxcr5
|
chemokine (C-X-C motif) receptor 5 |
| chr3_+_32663865 | 0.22 |
ENSDART00000190240
|
si:dkey-16l2.16
|
si:dkey-16l2.16 |
| chr13_+_9521629 | 0.22 |
ENSDART00000149870
ENSDART00000137666 |
si:dkey-19p15.4
|
si:dkey-19p15.4 |
| chr9_+_18829360 | 0.20 |
ENSDART00000006514
|
gtf2f2b
|
general transcription factor IIF, polypeptide 2b |
| chr4_+_54519511 | 0.19 |
ENSDART00000161653
|
znf974
|
zinc finger protein 974 |
| chr4_-_14185225 | 0.19 |
ENSDART00000147061
|
si:dkey-234l24.8
|
si:dkey-234l24.8 |
| chr17_-_41081806 | 0.19 |
ENSDART00000030622
|
rbks
|
ribokinase |
| chr5_-_67629263 | 0.18 |
ENSDART00000133753
|
zbtb20
|
zinc finger and BTB domain containing 20 |
| chr15_-_14212777 | 0.18 |
ENSDART00000165572
|
CR925813.1
|
|
| chr17_-_2834764 | 0.18 |
ENSDART00000024027
|
bdkrb1
|
bradykinin receptor B1 |
| chr13_-_9159852 | 0.18 |
ENSDART00000170185
ENSDART00000158697 ENSDART00000143393 ENSDART00000164973 ENSDART00000159910 |
HTRA2 (1 of many)
|
si:dkey-112g5.16 |
| chr17_-_6618574 | 0.18 |
ENSDART00000184486
|
si:ch211-189e2.3
|
si:ch211-189e2.3 |
| chr25_+_6471401 | 0.17 |
ENSDART00000132927
|
snx33
|
sorting nexin 33 |
| chr21_+_15592426 | 0.17 |
ENSDART00000138207
|
smarcb1b
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily b, member 1b |
| chr20_+_28803977 | 0.17 |
ENSDART00000153351
ENSDART00000038149 |
fntb
|
farnesyltransferase, CAAX box, beta |
| chr8_+_18555559 | 0.17 |
ENSDART00000149523
|
tab3
|
TGF-beta activated kinase 1/MAP3K7 binding protein 3 |
| chr13_-_9045879 | 0.17 |
ENSDART00000155463
ENSDART00000140041 ENSDART00000137454 |
si:dkey-112g5.11
|
si:dkey-112g5.11 |
| chr4_-_62714083 | 0.15 |
ENSDART00000188299
|
si:dkey-28k24.2
|
si:dkey-28k24.2 |
| chr11_+_2855430 | 0.15 |
ENSDART00000172837
|
kif21b
|
kinesin family member 21B |
| chr2_+_30368800 | 0.15 |
ENSDART00000179564
|
pi15b
|
peptidase inhibitor 15b |
| chr3_+_34140507 | 0.14 |
ENSDART00000131802
|
si:dkey-204f11.64
|
si:dkey-204f11.64 |
| chr22_+_9761930 | 0.14 |
ENSDART00000181107
|
BX664625.5
|
|
| chr20_+_46897504 | 0.13 |
ENSDART00000158124
|
si:ch73-21k16.1
|
si:ch73-21k16.1 |
| chr24_-_7957581 | 0.12 |
ENSDART00000145815
|
txndc5
|
thioredoxin domain containing 5 |
| chr13_-_9213207 | 0.12 |
ENSDART00000139861
ENSDART00000140524 |
HTRA2 (1 of many)
|
si:dkey-33c12.11 |
| chr21_-_20328375 | 0.12 |
ENSDART00000079593
|
slc26a1
|
solute carrier family 26 (anion exchanger), member 1 |
| chr5_+_872299 | 0.11 |
ENSDART00000130042
|
fubp3
|
far upstream element (FUSE) binding protein 3 |
| chr8_-_15129573 | 0.10 |
ENSDART00000142358
|
bcar3
|
BCAR3, NSP family adaptor protein |
| chr21_-_13661631 | 0.10 |
ENSDART00000184408
|
pnpla7a
|
patatin-like phospholipase domain containing 7a |
| chr12_-_31457301 | 0.09 |
ENSDART00000043887
ENSDART00000148603 |
acsl5
|
acyl-CoA synthetase long chain family member 5 |
| chr11_-_1935883 | 0.09 |
ENSDART00000172885
|
faim2b
|
Fas apoptotic inhibitory molecule 2b |
| chr9_-_32343673 | 0.09 |
ENSDART00000078499
|
rftn2
|
raftlin family member 2 |
| chr23_-_5729484 | 0.09 |
ENSDART00000000488
|
tnni1a
|
troponin I type 1a (skeletal, slow) |
| chr14_+_20918701 | 0.09 |
ENSDART00000144736
ENSDART00000148204 |
zgc:66433
|
zgc:66433 |
| chr12_-_33357655 | 0.08 |
ENSDART00000066233
ENSDART00000148165 |
slc16a3
|
solute carrier family 16 (monocarboxylate transporter), member 3 |
| chr7_-_8324927 | 0.07 |
ENSDART00000102535
|
f13a1b
|
coagulation factor XIII, A1 polypeptide b |
| chr23_+_18527028 | 0.07 |
ENSDART00000190263
ENSDART00000104545 |
avpr2ab
|
arginine vasopressin receptor 2a, duplicate b |
| chr13_+_18321140 | 0.06 |
ENSDART00000180947
|
eif4e1c
|
eukaryotic translation initiation factor 4E family member 1c |
| chr11_-_6974022 | 0.06 |
ENSDART00000172851
|
COMP
|
si:ch211-43f4.1 |
| chr2_+_30369116 | 0.05 |
ENSDART00000142137
|
pi15b
|
peptidase inhibitor 15b |
| chr15_+_36188171 | 0.04 |
ENSDART00000156598
|
masp1
|
mannan-binding lectin serine peptidase 1 |
| chr16_-_7379328 | 0.04 |
ENSDART00000060447
|
med18
|
mediator of RNA polymerase II transcription, subunit 18 homolog (yeast) |
| chr6_-_18971063 | 0.04 |
ENSDART00000188163
|
sept9b
|
septin 9b |
| chr20_-_51831816 | 0.03 |
ENSDART00000060505
|
mia3
|
melanoma inhibitory activity family, member 3 |
| chr15_-_25518084 | 0.03 |
ENSDART00000158594
|
hif1al
|
hypoxia-inducible factor 1, alpha subunit, like |
| chr16_-_18960613 | 0.03 |
ENSDART00000183197
|
fhod3b
|
formin homology 2 domain containing 3b |
| chr12_-_18883079 | 0.02 |
ENSDART00000178619
|
shisa8b
|
shisa family member 8b |
| chr4_-_67969695 | 0.02 |
ENSDART00000190016
|
si:ch211-223k15.1
|
si:ch211-223k15.1 |
| chr17_+_41081512 | 0.02 |
ENSDART00000056742
|
babam2
|
BRISC and BRCA1 A complex member 2 |
| chr11_+_42726712 | 0.01 |
ENSDART00000028955
|
tdrd3
|
tudor domain containing 3 |
| chr21_-_37973081 | 0.01 |
ENSDART00000136569
|
ripply1
|
ripply transcriptional repressor 1 |
| chr5_-_29512538 | 0.01 |
ENSDART00000098364
|
ehmt1a
|
euchromatic histone-lysine N-methyltransferase 1a |
| chr19_+_9212031 | 0.01 |
ENSDART00000052930
|
ndufv1
|
NADH dehydrogenase (ubiquinone) flavoprotein 1 |
Network of associatons between targets according to the STRING database.
First level regulatory network of msx1b+msx3
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.1 | GO:0019556 | formate metabolic process(GO:0015942) histidine catabolic process to glutamate and formamide(GO:0019556) histidine catabolic process to glutamate and formate(GO:0019557) formamide metabolic process(GO:0043606) |
| 0.3 | 1.0 | GO:1901533 | negative regulation of hematopoietic progenitor cell differentiation(GO:1901533) |
| 0.2 | 0.7 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 0.2 | 1.1 | GO:0070986 | left/right axis specification(GO:0070986) |
| 0.2 | 3.8 | GO:0002430 | complement receptor mediated signaling pathway(GO:0002430) |
| 0.1 | 0.6 | GO:0035279 | mRNA cleavage involved in gene silencing by miRNA(GO:0035279) mRNA cleavage involved in gene silencing(GO:0098795) |
| 0.1 | 0.6 | GO:0070814 | cysteine biosynthetic process from serine(GO:0006535) hydrogen sulfide biosynthetic process(GO:0070814) |
| 0.1 | 1.0 | GO:0072498 | embryonic skeletal joint development(GO:0072498) |
| 0.1 | 0.4 | GO:0030326 | embryonic limb morphogenesis(GO:0030326) |
| 0.1 | 1.2 | GO:1901534 | positive regulation of hematopoietic progenitor cell differentiation(GO:1901534) |
| 0.1 | 0.8 | GO:0044857 | membrane raft assembly(GO:0001765) plasma membrane raft assembly(GO:0044854) plasma membrane raft organization(GO:0044857) caveola assembly(GO:0070836) |
| 0.1 | 0.2 | GO:0019323 | pentose catabolic process(GO:0019323) |
| 0.1 | 0.8 | GO:0045740 | positive regulation of DNA replication(GO:0045740) |
| 0.0 | 0.6 | GO:0031573 | intra-S DNA damage checkpoint(GO:0031573) |
| 0.0 | 0.9 | GO:0006658 | phosphatidylserine metabolic process(GO:0006658) |
| 0.0 | 0.3 | GO:0071691 | cardiac muscle thin filament assembly(GO:0071691) |
| 0.0 | 0.8 | GO:0018279 | protein N-linked glycosylation via asparagine(GO:0018279) post-translational protein modification(GO:0043687) |
| 0.0 | 0.2 | GO:0018343 | protein farnesylation(GO:0018343) |
| 0.0 | 0.3 | GO:1990845 | adaptive thermogenesis(GO:1990845) |
| 0.0 | 0.1 | GO:0010747 | regulation of plasma membrane long-chain fatty acid transport(GO:0010746) positive regulation of plasma membrane long-chain fatty acid transport(GO:0010747) plasma membrane long-chain fatty acid transport(GO:0015911) fatty acid transmembrane transport(GO:1902001) positive regulation of anion transmembrane transport(GO:1903961) regulation of fatty acid transport(GO:2000191) positive regulation of fatty acid transport(GO:2000193) |
| 0.0 | 0.1 | GO:0019532 | oxalate transport(GO:0019532) |
| 0.0 | 0.3 | GO:0003139 | secondary heart field specification(GO:0003139) regulation of organ growth(GO:0046620) |
| 0.0 | 0.3 | GO:0000186 | activation of MAPKK activity(GO:0000186) |
| 0.0 | 0.3 | GO:0046549 | retinal cone cell development(GO:0046549) |
| 0.0 | 0.5 | GO:0006516 | glycoprotein catabolic process(GO:0006516) |
| 0.0 | 0.5 | GO:0032233 | positive regulation of actin filament bundle assembly(GO:0032233) |
| 0.0 | 0.3 | GO:0007198 | adenylate cyclase-inhibiting serotonin receptor signaling pathway(GO:0007198) |
| 0.0 | 1.3 | GO:0032869 | cellular response to insulin stimulus(GO:0032869) |
| 0.0 | 0.4 | GO:0019229 | regulation of vasoconstriction(GO:0019229) |
| 0.0 | 1.0 | GO:0072676 | lymphocyte migration(GO:0072676) |
| 0.0 | 0.3 | GO:0035036 | binding of sperm to zona pellucida(GO:0007339) sperm-egg recognition(GO:0035036) egg coat formation(GO:0035803) regulation of acrosome reaction(GO:0060046) positive regulation of acrosome reaction(GO:2000344) |
| 0.0 | 0.4 | GO:0051180 | vitamin transport(GO:0051180) |
| 0.0 | 1.0 | GO:0048015 | phosphatidylinositol-mediated signaling(GO:0048015) |
| 0.0 | 0.4 | GO:0030488 | tRNA methylation(GO:0030488) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0031515 | tRNA (m1A) methyltransferase complex(GO:0031515) |
| 0.1 | 0.4 | GO:0030891 | VCB complex(GO:0030891) |
| 0.1 | 0.4 | GO:0031428 | box C/D snoRNP complex(GO:0031428) |
| 0.0 | 0.3 | GO:0016012 | sarcoglycan complex(GO:0016012) |
| 0.0 | 0.2 | GO:0005965 | protein farnesyltransferase complex(GO:0005965) |
| 0.0 | 0.6 | GO:0031011 | Ino80 complex(GO:0031011) DNA helicase complex(GO:0033202) |
| 0.0 | 0.7 | GO:0046930 | pore complex(GO:0046930) |
| 0.0 | 0.2 | GO:0005674 | transcription factor TFIIF complex(GO:0005674) |
| 0.0 | 0.2 | GO:0035060 | brahma complex(GO:0035060) |
| 0.0 | 0.5 | GO:0045180 | basal cortex(GO:0045180) |
| 0.0 | 0.9 | GO:0031941 | filamentous actin(GO:0031941) |
| 0.0 | 0.8 | GO:0032587 | ruffle membrane(GO:0032587) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.1 | GO:0016841 | ammonia-lyase activity(GO:0016841) |
| 0.2 | 0.7 | GO:0048030 | disaccharide binding(GO:0048030) |
| 0.2 | 0.6 | GO:0032574 | 5'-3' RNA helicase activity(GO:0032574) |
| 0.2 | 3.8 | GO:0004875 | complement receptor activity(GO:0004875) |
| 0.2 | 0.8 | GO:0004576 | oligosaccharyl transferase activity(GO:0004576) dolichyl-diphosphooligosaccharide-protein glycotransferase activity(GO:0004579) |
| 0.2 | 1.1 | GO:0004743 | pyruvate kinase activity(GO:0004743) |
| 0.1 | 0.6 | GO:0004122 | cystathionine beta-synthase activity(GO:0004122) |
| 0.1 | 0.4 | GO:0004945 | angiotensin type I receptor activity(GO:0001596) angiotensin type II receptor activity(GO:0004945) |
| 0.1 | 0.4 | GO:0016429 | tRNA (adenine-N1-)-methyltransferase activity(GO:0016429) |
| 0.1 | 2.9 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.0 | 0.3 | GO:0070568 | guanylyltransferase activity(GO:0070568) |
| 0.0 | 0.9 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) alpha-actinin binding(GO:0051393) |
| 0.0 | 0.2 | GO:0004660 | protein farnesyltransferase activity(GO:0004660) |
| 0.0 | 1.0 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.0 | 0.5 | GO:0038191 | neuropilin binding(GO:0038191) |
| 0.0 | 0.4 | GO:0090482 | vitamin transmembrane transporter activity(GO:0090482) |
| 0.0 | 1.0 | GO:0004629 | phosphatidylinositol phospholipase C activity(GO:0004435) phospholipase C activity(GO:0004629) |
| 0.0 | 0.5 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.0 | 0.4 | GO:0030515 | snoRNA binding(GO:0030515) |
| 0.0 | 0.2 | GO:0030368 | interleukin-17 receptor activity(GO:0030368) |
| 0.0 | 0.3 | GO:0043176 | amine binding(GO:0043176) serotonin binding(GO:0051378) |
| 0.0 | 0.3 | GO:0032190 | acrosin binding(GO:0032190) |
| 0.0 | 0.3 | GO:0004709 | MAP kinase kinase kinase activity(GO:0004709) |
| 0.0 | 0.9 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 0.5 | GO:0005109 | frizzled binding(GO:0005109) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.2 | SA MMP CYTOKINE CONNECTION | Cytokines can induce activation of matrix metalloproteinases, which degrade extracellular matrix. |
| 0.1 | 1.0 | PID IL12 STAT4 PATHWAY | IL12 signaling mediated by STAT4 |
| 0.1 | 0.9 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.0 | 0.7 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
| 0.0 | 0.5 | PID INTEGRIN A4B1 PATHWAY | Alpha4 beta1 integrin signaling events |
| 0.0 | 1.0 | PID SMAD2 3NUCLEAR PATHWAY | Regulation of nuclear SMAD2/3 signaling |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.0 | REACTOME ACTIVATION OF IRF3 IRF7 MEDIATED BY TBK1 IKK EPSILON | Genes involved in Activation of IRF3/IRF7 mediated by TBK1/IKK epsilon |
| 0.0 | 0.2 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.0 | 0.7 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 0.0 | 0.2 | REACTOME TRAF6 MEDIATED INDUCTION OF TAK1 COMPLEX | Genes involved in TRAF6 mediated induction of TAK1 complex |
| 0.0 | 0.3 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.0 | 0.7 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.0 | 0.1 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |