Project
PRJNA207719: Tissue specific transcriptome profiling
Navigation
Downloads
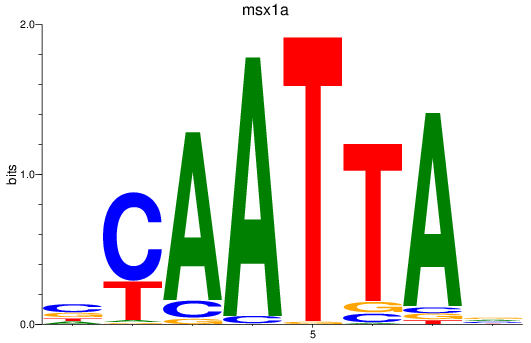
Results for msx1a
Z-value: 1.35
Transcription factors associated with msx1a
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
msx1a
|
ENSDARG00000116118 | muscle segment homeobox 1a |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| msx1a | dr11_v1_chr14_-_12822_12847 | 0.64 | 2.4e-01 | Click! |
Activity profile of msx1a motif
Sorted Z-values of msx1a motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr21_+_6751405 | 0.99 |
ENSDART00000037265
ENSDART00000146371 |
olfm1b
|
olfactomedin 1b |
| chr17_-_12389259 | 0.96 |
ENSDART00000185724
|
snap25b
|
synaptosomal-associated protein, 25b |
| chr15_-_21877726 | 0.83 |
ENSDART00000127819
ENSDART00000145646 ENSDART00000100897 ENSDART00000144739 |
zgc:162608
|
zgc:162608 |
| chr10_+_21867307 | 0.82 |
ENSDART00000126629
|
cbln17
|
cerebellin 17 |
| chr13_-_20381485 | 0.81 |
ENSDART00000131351
|
si:ch211-270n8.1
|
si:ch211-270n8.1 |
| chr21_+_6751760 | 0.75 |
ENSDART00000135914
|
olfm1b
|
olfactomedin 1b |
| chr16_-_29437373 | 0.67 |
ENSDART00000148405
|
si:ch211-113g11.6
|
si:ch211-113g11.6 |
| chr23_+_40460333 | 0.67 |
ENSDART00000184658
|
soga3b
|
SOGA family member 3b |
| chr5_-_23280098 | 0.66 |
ENSDART00000126540
ENSDART00000051533 |
plp1b
|
proteolipid protein 1b |
| chr20_-_23426339 | 0.64 |
ENSDART00000004625
|
zar1
|
zygote arrest 1 |
| chr2_+_6253246 | 0.63 |
ENSDART00000058256
ENSDART00000076700 |
zp3b
|
zona pellucida glycoprotein 3b |
| chr12_+_24342303 | 0.60 |
ENSDART00000111239
|
nrxn1a
|
neurexin 1a |
| chr4_+_12615836 | 0.59 |
ENSDART00000003583
|
lmo3
|
LIM domain only 3 |
| chr20_-_32110882 | 0.58 |
ENSDART00000030324
|
grm1a
|
glutamate receptor, metabotropic 1a |
| chr11_-_6452444 | 0.57 |
ENSDART00000137879
ENSDART00000134957 ENSDART00000004483 |
larp6b
|
La ribonucleoprotein domain family, member 6b |
| chr16_+_39159752 | 0.57 |
ENSDART00000122081
|
sybu
|
syntabulin (syntaxin-interacting) |
| chr9_-_35633827 | 0.56 |
ENSDART00000077745
|
zp2l1
|
zona pellucida glycoprotein 2, like 1 |
| chr17_-_44247707 | 0.55 |
ENSDART00000126097
|
otx2b
|
orthodenticle homeobox 2b |
| chr16_+_5774977 | 0.55 |
ENSDART00000134202
|
ccka
|
cholecystokinin a |
| chr6_+_3828560 | 0.52 |
ENSDART00000185273
ENSDART00000179091 |
gad1b
|
glutamate decarboxylase 1b |
| chr14_+_34490445 | 0.51 |
ENSDART00000132193
ENSDART00000148044 |
wnt8a
|
wingless-type MMTV integration site family, member 8a |
| chr10_-_21362071 | 0.51 |
ENSDART00000125167
|
avd
|
avidin |
| chr21_+_25777425 | 0.50 |
ENSDART00000021620
|
cldnd
|
claudin d |
| chr6_+_39232245 | 0.49 |
ENSDART00000187351
|
b4galnt1b
|
beta-1,4-N-acetyl-galactosaminyl transferase 1b |
| chr24_-_24849091 | 0.49 |
ENSDART00000133649
ENSDART00000038290 |
crhb
|
corticotropin releasing hormone b |
| chr5_+_37032038 | 0.48 |
ENSDART00000045036
|
nfkbib
|
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, beta |
| chr2_+_50626476 | 0.47 |
ENSDART00000018150
|
neurod6b
|
neuronal differentiation 6b |
| chr22_-_23253481 | 0.47 |
ENSDART00000054807
|
lhx9
|
LIM homeobox 9 |
| chr2_+_50608099 | 0.46 |
ENSDART00000185805
ENSDART00000111135 |
neurod6b
|
neuronal differentiation 6b |
| chr20_-_28800999 | 0.45 |
ENSDART00000049462
|
rab15
|
RAB15, member RAS oncogene family |
| chr10_+_17714866 | 0.45 |
ENSDART00000039969
|
slc20a1b
|
solute carrier family 20 (phosphate transporter), member 1b |
| chr20_+_30490682 | 0.45 |
ENSDART00000184871
|
myt1la
|
myelin transcription factor 1-like, a |
| chr15_-_6247775 | 0.44 |
ENSDART00000148350
|
dscamb
|
Down syndrome cell adhesion molecule b |
| chr25_-_32869794 | 0.44 |
ENSDART00000162784
|
tmem266
|
transmembrane protein 266 |
| chr6_-_40352215 | 0.43 |
ENSDART00000103992
|
ttll3
|
tubulin tyrosine ligase-like family, member 3 |
| chr10_-_21362320 | 0.43 |
ENSDART00000189789
|
avd
|
avidin |
| chr13_-_36911118 | 0.42 |
ENSDART00000048739
|
trim9
|
tripartite motif containing 9 |
| chr13_+_38430466 | 0.41 |
ENSDART00000132691
|
adgrb3
|
adhesion G protein-coupled receptor B3 |
| chr11_-_42554290 | 0.41 |
ENSDART00000130573
|
atp6ap1la
|
ATPase H+ transporting accessory protein 1 like a |
| chr18_-_40708537 | 0.41 |
ENSDART00000077577
|
si:ch211-132b12.8
|
si:ch211-132b12.8 |
| chr16_-_27628994 | 0.40 |
ENSDART00000157407
|
nacad
|
NAC alpha domain containing |
| chr7_+_30787903 | 0.39 |
ENSDART00000174000
|
apba2b
|
amyloid beta (A4) precursor protein-binding, family A, member 2b |
| chr23_-_28141419 | 0.39 |
ENSDART00000133039
|
tac3a
|
tachykinin 3a |
| chr7_-_28148310 | 0.39 |
ENSDART00000044208
|
lmo1
|
LIM domain only 1 |
| chr6_-_6487876 | 0.38 |
ENSDART00000137642
|
cep170ab
|
centrosomal protein 170Ab |
| chr10_-_34915886 | 0.38 |
ENSDART00000141201
ENSDART00000002166 |
ccna1
|
cyclin A1 |
| chr1_-_18811517 | 0.37 |
ENSDART00000142026
|
si:dkey-167i21.2
|
si:dkey-167i21.2 |
| chr4_-_15420452 | 0.37 |
ENSDART00000016230
|
plxna4
|
plexin A4 |
| chr20_-_40755614 | 0.37 |
ENSDART00000061247
|
cx32.3
|
connexin 32.3 |
| chr20_+_41021054 | 0.36 |
ENSDART00000146052
|
man1a1
|
mannosidase, alpha, class 1A, member 1 |
| chr23_+_28582865 | 0.36 |
ENSDART00000020296
|
l1cama
|
L1 cell adhesion molecule, paralog a |
| chr8_+_22582146 | 0.35 |
ENSDART00000157655
ENSDART00000189892 |
CT583651.2
|
|
| chr22_-_7129631 | 0.35 |
ENSDART00000171359
|
asic1b
|
acid-sensing (proton-gated) ion channel 1b |
| chr7_+_32722227 | 0.34 |
ENSDART00000126565
|
si:ch211-150g13.3
|
si:ch211-150g13.3 |
| chr4_+_12612723 | 0.34 |
ENSDART00000133767
|
lmo3
|
LIM domain only 3 |
| chr7_+_27603211 | 0.34 |
ENSDART00000148782
|
cyp2r1
|
cytochrome P450, family 2, subfamily R, polypeptide 1 |
| chr14_-_2364761 | 0.33 |
ENSDART00000167322
|
PCDHGC3 (1 of many)
|
si:ch73-233f7.3 |
| chr15_-_25518084 | 0.33 |
ENSDART00000158594
|
hif1al
|
hypoxia-inducible factor 1, alpha subunit, like |
| chr2_-_33645411 | 0.33 |
ENSDART00000114663
|
b4galt2
|
UDP-Gal:betaGlcNAc beta 1,4- galactosyltransferase, polypeptide 2 |
| chr18_+_15644559 | 0.31 |
ENSDART00000061794
|
nr1h4
|
nuclear receptor subfamily 1, group H, member 4 |
| chr14_-_30587814 | 0.31 |
ENSDART00000144912
ENSDART00000149714 |
tmem265
|
transmembrane protein 265 |
| chr3_+_56366395 | 0.31 |
ENSDART00000154367
|
cacng5b
|
calcium channel, voltage-dependent, gamma subunit 5b |
| chr24_-_31846366 | 0.30 |
ENSDART00000155295
|
steap2
|
STEAP family member 2, metalloreductase |
| chr15_-_22074315 | 0.30 |
ENSDART00000149830
|
drd2a
|
dopamine receptor D2a |
| chr12_+_2446837 | 0.30 |
ENSDART00000112032
|
ARHGAP22
|
si:dkey-191m6.4 |
| chr6_+_21001264 | 0.29 |
ENSDART00000044519
ENSDART00000151278 |
cx44.2
|
connexin 44.2 |
| chr19_-_8604429 | 0.28 |
ENSDART00000151165
|
trim46b
|
tripartite motif containing 46b |
| chr12_+_31744217 | 0.28 |
ENSDART00000190361
|
RNF157
|
si:dkey-49c17.3 |
| chr10_-_25217347 | 0.28 |
ENSDART00000036906
|
kpna7
|
karyopherin alpha 7 (importin alpha 8) |
| chr1_-_47071979 | 0.28 |
ENSDART00000160817
|
itsn1
|
intersectin 1 (SH3 domain protein) |
| chr7_-_23996133 | 0.27 |
ENSDART00000173761
|
si:dkey-183c6.8
|
si:dkey-183c6.8 |
| chr1_+_18811679 | 0.27 |
ENSDART00000078610
|
slc25a51a
|
solute carrier family 25, member 51a |
| chr13_-_36184476 | 0.27 |
ENSDART00000057185
|
map3k9
|
mitogen-activated protein kinase kinase kinase 9 |
| chr15_-_20024205 | 0.26 |
ENSDART00000161379
|
auts2b
|
autism susceptibility candidate 2b |
| chr6_-_39649504 | 0.26 |
ENSDART00000179960
ENSDART00000190951 |
larp4ab
|
La ribonucleoprotein domain family, member 4Ab |
| chr19_-_38830582 | 0.26 |
ENSDART00000189966
ENSDART00000183055 |
adgrb2
|
adhesion G protein-coupled receptor B2 |
| chr8_+_45334255 | 0.26 |
ENSDART00000126848
ENSDART00000134161 ENSDART00000142322 ENSDART00000145011 ENSDART00000183560 |
pabpc1l
|
poly(A) binding protein, cytoplasmic 1-like |
| chr4_+_11384891 | 0.25 |
ENSDART00000092381
ENSDART00000186577 ENSDART00000191054 ENSDART00000191584 |
pcloa
|
piccolo presynaptic cytomatrix protein a |
| chr4_+_12612145 | 0.25 |
ENSDART00000181201
|
lmo3
|
LIM domain only 3 |
| chr14_-_2933185 | 0.24 |
ENSDART00000161677
ENSDART00000162446 ENSDART00000109378 |
si:dkey-201i24.6
|
si:dkey-201i24.6 |
| chr3_+_28860283 | 0.23 |
ENSDART00000077235
|
alg1
|
ALG1, chitobiosyldiphosphodolichol beta-mannosyltransferase |
| chr10_+_42690374 | 0.23 |
ENSDART00000123496
|
rhobtb2b
|
Rho-related BTB domain containing 2b |
| chr11_-_44801968 | 0.22 |
ENSDART00000161846
|
map1lc3c
|
microtubule-associated protein 1 light chain 3 gamma |
| chr11_-_29563437 | 0.22 |
ENSDART00000163958
|
arhgef10la
|
Rho guanine nucleotide exchange factor (GEF) 10-like a |
| chr9_+_34331368 | 0.22 |
ENSDART00000147913
|
pou2f1b
|
POU class 2 homeobox 1b |
| chr9_-_20372977 | 0.22 |
ENSDART00000113418
|
igsf3
|
immunoglobulin superfamily, member 3 |
| chr24_+_12835935 | 0.22 |
ENSDART00000114762
|
nanog
|
nanog homeobox |
| chr18_-_2433011 | 0.22 |
ENSDART00000181922
ENSDART00000193276 |
CR769778.1
|
|
| chr10_-_34002185 | 0.21 |
ENSDART00000046599
|
zar1l
|
zygote arrest 1-like |
| chr5_-_12219572 | 0.21 |
ENSDART00000167834
|
nos1
|
nitric oxide synthase 1 (neuronal) |
| chr2_-_57076687 | 0.21 |
ENSDART00000161523
|
slc25a42
|
solute carrier family 25, member 42 |
| chr17_+_24821627 | 0.20 |
ENSDART00000112389
|
wdr43
|
WD repeat domain 43 |
| chr7_-_31941330 | 0.20 |
ENSDART00000144682
|
bdnf
|
brain-derived neurotrophic factor |
| chr17_+_31914877 | 0.20 |
ENSDART00000177801
|
FAM196A (1 of many)
|
family with sequence similarity 196 member A |
| chr3_+_18807006 | 0.19 |
ENSDART00000180091
|
tnpo2
|
transportin 2 (importin 3, karyopherin beta 2b) |
| chr9_-_52962521 | 0.19 |
ENSDART00000170419
|
CU855885.1
|
|
| chr16_+_54263921 | 0.19 |
ENSDART00000002856
|
drd2l
|
dopamine receptor D2 like |
| chr10_-_34916208 | 0.19 |
ENSDART00000187371
|
ccna1
|
cyclin A1 |
| chr24_-_30843250 | 0.19 |
ENSDART00000162920
|
ptbp2a
|
polypyrimidine tract binding protein 2a |
| chr7_-_31940590 | 0.19 |
ENSDART00000131009
|
bdnf
|
brain-derived neurotrophic factor |
| chr24_+_40860320 | 0.19 |
ENSDART00000161351
|
gorasp1b
|
golgi reassembly stacking protein 1b |
| chr11_+_18183220 | 0.19 |
ENSDART00000113468
|
LO018315.10
|
|
| chr15_+_30323491 | 0.18 |
ENSDART00000048847
|
nos2b
|
nitric oxide synthase 2b, inducible |
| chr10_+_6884627 | 0.18 |
ENSDART00000125262
ENSDART00000121729 ENSDART00000105384 |
ddx4
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 4 |
| chr8_+_26125218 | 0.18 |
ENSDART00000145095
|
celsr3
|
cadherin, EGF LAG seven-pass G-type receptor 3 |
| chr11_+_18130300 | 0.17 |
ENSDART00000169146
|
zgc:175135
|
zgc:175135 |
| chr19_-_8768564 | 0.17 |
ENSDART00000170416
|
si:ch73-350k19.1
|
si:ch73-350k19.1 |
| chr20_+_19238382 | 0.17 |
ENSDART00000136757
|
fndc4a
|
fibronectin type III domain containing 4a |
| chr20_-_19511016 | 0.17 |
ENSDART00000168521
|
snx17
|
sorting nexin 17 |
| chr11_+_30244356 | 0.16 |
ENSDART00000036050
ENSDART00000150080 |
rs1a
|
retinoschisin 1a |
| chr13_+_2625150 | 0.16 |
ENSDART00000164177
|
plpp4
|
phospholipid phosphatase 4 |
| chr18_+_24921587 | 0.15 |
ENSDART00000191345
|
rgma
|
repulsive guidance molecule family member a |
| chr6_-_44711942 | 0.15 |
ENSDART00000055035
|
cntn3b
|
contactin 3b |
| chr1_-_55248496 | 0.15 |
ENSDART00000098615
|
nanos3
|
nanos homolog 3 |
| chr5_-_67629263 | 0.14 |
ENSDART00000133753
|
zbtb20
|
zinc finger and BTB domain containing 20 |
| chr20_-_46362606 | 0.14 |
ENSDART00000153087
|
bmf2
|
BCL2 modifying factor 2 |
| chr11_+_18157260 | 0.14 |
ENSDART00000144659
|
zgc:173545
|
zgc:173545 |
| chr7_-_23768234 | 0.14 |
ENSDART00000173981
|
si:ch211-200p22.4
|
si:ch211-200p22.4 |
| chr8_+_26253252 | 0.13 |
ENSDART00000142031
|
slc26a6
|
solute carrier family 26, member 6 |
| chr18_-_25568994 | 0.13 |
ENSDART00000133029
|
si:ch211-13k12.2
|
si:ch211-13k12.2 |
| chr2_-_55298075 | 0.13 |
ENSDART00000186404
ENSDART00000149062 |
rab8a
|
RAB8A, member RAS oncogene family |
| chr16_+_35661771 | 0.12 |
ENSDART00000161393
|
map7d1a
|
MAP7 domain containing 1a |
| chr1_-_50859053 | 0.12 |
ENSDART00000132779
ENSDART00000137648 |
si:dkeyp-123h10.2
|
si:dkeyp-123h10.2 |
| chr12_+_21298317 | 0.11 |
ENSDART00000178562
|
ca10a
|
carbonic anhydrase Xa |
| chr7_+_42460302 | 0.11 |
ENSDART00000004120
|
adamts18
|
ADAM metallopeptidase with thrombospondin type 1 motif, 18 |
| chr15_+_1796313 | 0.11 |
ENSDART00000126253
|
fam124b
|
family with sequence similarity 124B |
| chr19_-_20403845 | 0.11 |
ENSDART00000151265
ENSDART00000147911 ENSDART00000151356 |
dazl
|
deleted in azoospermia-like |
| chr6_-_37744430 | 0.11 |
ENSDART00000150177
ENSDART00000149722 |
nipa2
|
non imprinted in Prader-Willi/Angelman syndrome 2 (human) |
| chr19_-_15855427 | 0.11 |
ENSDART00000133059
|
cited4a
|
Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 4a |
| chr14_-_34044369 | 0.11 |
ENSDART00000149396
ENSDART00000123607 ENSDART00000190746 |
cyfip2
|
cytoplasmic FMR1 interacting protein 2 |
| chr21_-_3853204 | 0.10 |
ENSDART00000188829
|
st6galnac4
|
ST6 (alpha-N-acetyl-neuraminyl-2,3-beta-galactosyl-1,3)-N-acetylgalactosaminide alpha-2,6-sialyltransferase 4 |
| chr23_+_33963619 | 0.10 |
ENSDART00000140666
ENSDART00000084792 |
plpbp
|
pyridoxal phosphate binding protein |
| chr7_-_12464412 | 0.10 |
ENSDART00000178723
|
adamtsl3
|
ADAMTS-like 3 |
| chr15_-_12270857 | 0.10 |
ENSDART00000170093
|
si:dkey-36i7.3
|
si:dkey-36i7.3 |
| chr1_-_36152131 | 0.10 |
ENSDART00000182113
ENSDART00000182904 |
znf827
|
zinc finger protein 827 |
| chr17_+_16046132 | 0.09 |
ENSDART00000155005
|
si:ch73-204p21.2
|
si:ch73-204p21.2 |
| chr10_-_28513861 | 0.09 |
ENSDART00000177781
|
bbx
|
bobby sox homolog (Drosophila) |
| chr20_-_11178022 | 0.08 |
ENSDART00000152246
|
flrt2
|
fibronectin leucine rich transmembrane protein 2 |
| chr19_-_19556778 | 0.08 |
ENSDART00000164060
|
tax1bp1a
|
Tax1 (human T-cell leukemia virus type I) binding protein 1a |
| chr3_+_32365811 | 0.07 |
ENSDART00000155967
|
ap2a1
|
adaptor-related protein complex 2, alpha 1 subunit |
| chr13_+_38990939 | 0.07 |
ENSDART00000145979
|
col19a1
|
collagen, type XIX, alpha 1 |
| chr5_+_6954162 | 0.07 |
ENSDART00000086666
|
stpg2
|
sperm-tail PG-rich repeat containing 2 |
| chr17_+_21295132 | 0.07 |
ENSDART00000103845
|
eno4
|
enolase family member 4 |
| chr18_+_9362455 | 0.07 |
ENSDART00000187025
|
sema3ab
|
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3Ab |
| chr16_+_33902006 | 0.07 |
ENSDART00000161807
ENSDART00000159474 |
gnl2
|
guanine nucleotide binding protein-like 2 (nucleolar) |
| chr10_+_3049636 | 0.07 |
ENSDART00000081794
ENSDART00000183167 ENSDART00000191634 ENSDART00000183514 |
rasgrf2a
|
Ras protein-specific guanine nucleotide-releasing factor 2a |
| chr2_+_20793982 | 0.06 |
ENSDART00000014785
|
prg4a
|
proteoglycan 4a |
| chr5_+_60590796 | 0.06 |
ENSDART00000159859
|
tmem132e
|
transmembrane protein 132E |
| chr10_+_43039947 | 0.05 |
ENSDART00000193434
|
atg10
|
ATG10 autophagy related 10 homolog (S. cerevisiae) |
| chr16_+_31804590 | 0.05 |
ENSDART00000167321
|
wnt4b
|
wingless-type MMTV integration site family, member 4b |
| chr19_-_19871211 | 0.05 |
ENSDART00000170980
|
evx1
|
even-skipped homeobox 1 |
| chr4_-_72638972 | 0.05 |
ENSDART00000193312
|
CABZ01054394.1
|
|
| chr17_+_46818521 | 0.05 |
ENSDART00000156022
|
pimr14
|
Pim proto-oncogene, serine/threonine kinase, related 14 |
| chr12_-_18872927 | 0.05 |
ENSDART00000187717
|
shisa8b
|
shisa family member 8b |
| chr2_+_31804582 | 0.04 |
ENSDART00000086646
|
rnf182
|
ring finger protein 182 |
| chr14_-_32631013 | 0.04 |
ENSDART00000176815
|
atp11c
|
ATPase phospholipid transporting 11C |
| chr6_+_23887314 | 0.04 |
ENSDART00000163188
|
znf648
|
zinc finger protein 648 |
| chr17_+_33311784 | 0.04 |
ENSDART00000156229
|
si:ch211-132f19.7
|
si:ch211-132f19.7 |
| chr20_-_29864390 | 0.04 |
ENSDART00000161834
ENSDART00000132278 |
rnf144ab
|
ring finger protein 144ab |
| chr11_+_33312601 | 0.04 |
ENSDART00000188024
|
cntnap5l
|
contactin associated protein-like 5 like |
| chr21_+_34088110 | 0.03 |
ENSDART00000145123
ENSDART00000029599 ENSDART00000147519 |
mtmr1b
|
myotubularin related protein 1b |
| chr23_+_30967686 | 0.03 |
ENSDART00000144485
|
si:ch211-197l9.2
|
si:ch211-197l9.2 |
| chr13_+_25449681 | 0.03 |
ENSDART00000101328
|
atoh7
|
atonal bHLH transcription factor 7 |
| chr11_-_30158191 | 0.03 |
ENSDART00000155278
ENSDART00000156121 |
scml2
|
Scm polycomb group protein like 2 |
| chr11_+_31864921 | 0.03 |
ENSDART00000180252
|
diaph3
|
diaphanous-related formin 3 |
| chr15_-_5172170 | 0.03 |
ENSDART00000062830
|
or126-5
|
odorant receptor, family E, subfamily 126, member 5 |
| chr15_-_26931541 | 0.03 |
ENSDART00000027563
|
ccdc9
|
coiled-coil domain containing 9 |
| chr16_-_47483142 | 0.03 |
ENSDART00000147072
|
cthrc1b
|
collagen triple helix repeat containing 1b |
| chr21_+_30746348 | 0.03 |
ENSDART00000050172
|
trpc2b
|
transient receptor potential cation channel, subfamily C, member 2b |
| chr21_-_8085635 | 0.02 |
ENSDART00000082790
|
si:dkey-163m14.2
|
si:dkey-163m14.2 |
| chr19_-_20403507 | 0.02 |
ENSDART00000052603
ENSDART00000137590 |
dazl
|
deleted in azoospermia-like |
| chr24_-_25144441 | 0.02 |
ENSDART00000152104
|
phldb2b
|
pleckstrin homology-like domain, family B, member 2b |
| chr22_-_13649588 | 0.02 |
ENSDART00000131877
|
si:ch211-279g13.1
|
si:ch211-279g13.1 |
| chr5_+_68807170 | 0.01 |
ENSDART00000017849
|
her7
|
hairy and enhancer of split related-7 |
| chr6_+_9204099 | 0.01 |
ENSDART00000150167
ENSDART00000115394 |
tbx19
|
T-box 19 |
| chr2_+_21229909 | 0.01 |
ENSDART00000046127
|
vipr1a
|
vasoactive intestinal peptide receptor 1a |
| chr15_-_16177603 | 0.01 |
ENSDART00000156352
|
si:ch211-259g3.4
|
si:ch211-259g3.4 |
| chr14_-_25034142 | 0.01 |
ENSDART00000000392
|
pde6a
|
phosphodiesterase 6A, cGMP-specific, rod, alpha |
| chr19_+_28187480 | 0.01 |
ENSDART00000183825
|
irx4b
|
iroquois homeobox 4b |
| chr24_+_26742226 | 0.00 |
ENSDART00000079721
|
ghsrb
|
growth hormone secretagogue receptor b |
| chr17_+_26543690 | 0.00 |
ENSDART00000156417
|
sparcl2
|
SPARC-like 2 |
| chr13_-_44630111 | 0.00 |
ENSDART00000110092
|
mdga1
|
MAM domain containing glycosylphosphatidylinositol anchor 1 |
| chr2_+_37227011 | 0.00 |
ENSDART00000126587
ENSDART00000084958 |
samd7
|
sterile alpha motif domain containing 7 |
Network of associatons between targets according to the STRING database.
First level regulatory network of msx1a
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.5 | GO:0009449 | gamma-aminobutyric acid biosynthetic process(GO:0009449) |
| 0.2 | 0.5 | GO:0039015 | spinal cord anterior/posterior patterning(GO:0021512) cell proliferation involved in pronephros development(GO:0039015) cell proliferation involved in kidney development(GO:0072111) |
| 0.2 | 0.8 | GO:0010872 | regulation of cholesterol esterification(GO:0010872) positive regulation of cholesterol esterification(GO:0010873) |
| 0.2 | 0.5 | GO:0035902 | response to immobilization stress(GO:0035902) |
| 0.2 | 0.5 | GO:0097376 | interneuron axon guidance(GO:0097376) spinal cord interneuron axon guidance(GO:0097377) dorsal spinal cord interneuron axon guidance(GO:0097378) |
| 0.1 | 0.6 | GO:0060074 | synapse maturation(GO:0060074) |
| 0.1 | 0.4 | GO:0007414 | axonal defasciculation(GO:0007414) |
| 0.1 | 0.5 | GO:0016338 | calcium-independent cell-cell adhesion via plasma membrane cell-adhesion molecules(GO:0016338) |
| 0.1 | 0.4 | GO:0016322 | neuron remodeling(GO:0016322) |
| 0.1 | 0.4 | GO:0007217 | tachykinin receptor signaling pathway(GO:0007217) |
| 0.1 | 0.3 | GO:0015677 | copper ion import(GO:0015677) |
| 0.1 | 0.2 | GO:0098924 | retrograde trans-synaptic signaling by soluble gas(GO:0098923) retrograde trans-synaptic signaling by nitric oxide(GO:0098924) |
| 0.1 | 1.0 | GO:0045162 | clustering of voltage-gated sodium channels(GO:0045162) |
| 0.1 | 0.2 | GO:0030828 | regulation of cGMP metabolic process(GO:0030823) positive regulation of cGMP metabolic process(GO:0030825) regulation of cGMP biosynthetic process(GO:0030826) positive regulation of cGMP biosynthetic process(GO:0030828) regulation of guanylate cyclase activity(GO:0031282) positive regulation of guanylate cyclase activity(GO:0031284) |
| 0.1 | 0.3 | GO:0071305 | vitamin D3 metabolic process(GO:0070640) cellular response to vitamin D(GO:0071305) |
| 0.1 | 0.4 | GO:0070207 | protein homotrimerization(GO:0070207) |
| 0.0 | 0.6 | GO:0001574 | ganglioside biosynthetic process(GO:0001574) |
| 0.0 | 0.6 | GO:2000344 | binding of sperm to zona pellucida(GO:0007339) sperm-egg recognition(GO:0035036) egg coat formation(GO:0035803) regulation of acrosome reaction(GO:0060046) positive regulation of acrosome reaction(GO:2000344) |
| 0.0 | 0.1 | GO:0030718 | germ-line stem cell population maintenance(GO:0030718) |
| 0.0 | 0.5 | GO:0007195 | adenylate cyclase-inhibiting dopamine receptor signaling pathway(GO:0007195) negative regulation of cytosolic calcium ion concentration(GO:0051481) negative regulation of synaptic transmission, glutamatergic(GO:0051967) |
| 0.0 | 0.7 | GO:0032291 | central nervous system myelination(GO:0022010) axon ensheathment in central nervous system(GO:0032291) |
| 0.0 | 0.6 | GO:0003406 | retinal pigment epithelium development(GO:0003406) |
| 0.0 | 0.4 | GO:0048672 | positive regulation of collateral sprouting(GO:0048672) |
| 0.0 | 0.1 | GO:0048210 | Golgi vesicle fusion to target membrane(GO:0048210) |
| 0.0 | 0.2 | GO:0031293 | membrane protein intracellular domain proteolysis(GO:0031293) |
| 0.0 | 0.3 | GO:1904071 | presynaptic active zone assembly(GO:1904071) |
| 0.0 | 0.3 | GO:0006607 | NLS-bearing protein import into nucleus(GO:0006607) |
| 0.0 | 0.1 | GO:0045852 | pH elevation(GO:0045852) intracellular pH elevation(GO:0051454) |
| 0.0 | 0.3 | GO:0014823 | response to activity(GO:0014823) |
| 0.0 | 0.3 | GO:0050728 | negative regulation of inflammatory response(GO:0050728) |
| 0.0 | 0.4 | GO:0035435 | phosphate ion transmembrane transport(GO:0035435) |
| 0.0 | 0.3 | GO:0098943 | neurotransmitter receptor transport, postsynaptic endosome to lysosome(GO:0098943) |
| 0.0 | 0.1 | GO:0060965 | negative regulation of posttranscriptional gene silencing(GO:0060149) negative regulation of gene silencing by miRNA(GO:0060965) negative regulation of gene silencing by RNA(GO:0060967) |
| 0.0 | 0.6 | GO:0007216 | G-protein coupled glutamate receptor signaling pathway(GO:0007216) |
| 0.0 | 0.2 | GO:0071501 | response to sterol depletion(GO:0006991) SREBP signaling pathway(GO:0032933) cellular response to sterol depletion(GO:0071501) |
| 0.0 | 0.1 | GO:0000448 | cleavage in ITS2 between 5.8S rRNA and LSU-rRNA of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000448) |
| 0.0 | 0.4 | GO:0006491 | N-glycan processing(GO:0006491) |
| 0.0 | 0.4 | GO:0045773 | positive regulation of axon extension(GO:0045773) |
| 0.0 | 0.4 | GO:0070593 | dendrite self-avoidance(GO:0070593) |
| 0.0 | 0.5 | GO:0032482 | Rab protein signal transduction(GO:0032482) |
| 0.0 | 0.6 | GO:0007586 | digestion(GO:0007586) |
| 0.0 | 0.3 | GO:0007257 | activation of JUN kinase activity(GO:0007257) positive regulation of JUN kinase activity(GO:0043507) |
| 0.0 | 0.2 | GO:0006995 | cellular response to nitrogen starvation(GO:0006995) cellular response to nitrogen levels(GO:0043562) |
| 0.0 | 0.1 | GO:0048755 | branching morphogenesis of a nerve(GO:0048755) positive regulation of neuron migration(GO:2001224) |
| 0.0 | 0.6 | GO:1904029 | regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0000079) regulation of cyclin-dependent protein kinase activity(GO:1904029) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.6 | GO:0097124 | cyclin A2-CDK2 complex(GO:0097124) |
| 0.2 | 0.5 | GO:0097189 | apoptotic body(GO:0097189) |
| 0.2 | 1.0 | GO:0070032 | synaptobrevin 2-SNAP-25-syntaxin-1a-complexin I complex(GO:0070032) |
| 0.1 | 0.4 | GO:0043083 | synaptic cleft(GO:0043083) |
| 0.1 | 0.4 | GO:0033181 | plasma membrane proton-transporting V-type ATPase complex(GO:0033181) |
| 0.1 | 0.4 | GO:0005854 | nascent polypeptide-associated complex(GO:0005854) |
| 0.1 | 0.8 | GO:0042627 | chylomicron(GO:0042627) |
| 0.0 | 0.5 | GO:0032593 | insulin-responsive compartment(GO:0032593) |
| 0.0 | 0.7 | GO:0043209 | myelin sheath(GO:0043209) |
| 0.0 | 0.4 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.0 | 0.3 | GO:0098982 | GABA-ergic synapse(GO:0098982) |
| 0.0 | 0.4 | GO:0002116 | semaphorin receptor complex(GO:0002116) |
| 0.0 | 0.9 | GO:0099634 | postsynaptic density membrane(GO:0098839) postsynaptic specialization membrane(GO:0099634) |
| 0.0 | 0.3 | GO:1990907 | beta-catenin-TCF complex(GO:1990907) |
| 0.0 | 0.5 | GO:0098978 | glutamatergic synapse(GO:0098978) |
| 0.0 | 0.6 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.0 | 0.1 | GO:0098888 | presynaptic endocytic zone(GO:0098833) presynaptic endocytic zone membrane(GO:0098835) extrinsic component of presynaptic membrane(GO:0098888) extrinsic component of presynaptic endocytic zone(GO:0098894) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.9 | GO:0009374 | biotin binding(GO:0009374) |
| 0.2 | 0.5 | GO:0004351 | glutamate decarboxylase activity(GO:0004351) |
| 0.2 | 0.8 | GO:0060228 | phosphatidylcholine-sterol O-acyltransferase activator activity(GO:0060228) |
| 0.1 | 0.4 | GO:0070735 | protein-glycine ligase activity(GO:0070735) tubulin-glycine ligase activity(GO:0070738) |
| 0.1 | 0.6 | GO:0099530 | G-protein coupled receptor activity involved in regulation of postsynaptic membrane potential(GO:0099530) |
| 0.1 | 0.3 | GO:0052851 | cupric reductase activity(GO:0008823) ferric-chelate reductase (NADPH) activity(GO:0052851) |
| 0.1 | 0.4 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 0.1 | 0.4 | GO:0044736 | acid-sensing ion channel activity(GO:0044736) |
| 0.1 | 0.2 | GO:0071077 | coenzyme A transmembrane transporter activity(GO:0015228) adenosine 3',5'-bisphosphate transmembrane transporter activity(GO:0071077) AMP transmembrane transporter activity(GO:0080122) |
| 0.1 | 0.6 | GO:0035804 | structural constituent of egg coat(GO:0035804) |
| 0.1 | 0.4 | GO:0004517 | nitric-oxide synthase activity(GO:0004517) |
| 0.1 | 0.3 | GO:0032052 | bile acid binding(GO:0032052) |
| 0.0 | 0.7 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.0 | 1.5 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.0 | 0.4 | GO:0004571 | mannosyl-oligosaccharide 1,2-alpha-mannosidase activity(GO:0004571) mannosyl-oligosaccharide mannosidase activity(GO:0015924) |
| 0.0 | 0.5 | GO:0001591 | dopamine neurotransmitter receptor activity, coupled via Gi/Go(GO:0001591) |
| 0.0 | 0.3 | GO:0004563 | beta-N-acetylhexosaminidase activity(GO:0004563) |
| 0.0 | 0.4 | GO:0005163 | nerve growth factor receptor binding(GO:0005163) neurotrophin receptor binding(GO:0005165) |
| 0.0 | 0.4 | GO:0005315 | inorganic phosphate transmembrane transporter activity(GO:0005315) |
| 0.0 | 0.3 | GO:0004706 | JUN kinase kinase kinase activity(GO:0004706) |
| 0.0 | 0.5 | GO:0061608 | nuclear import signal receptor activity(GO:0061608) |
| 0.0 | 0.3 | GO:0008266 | poly(U) RNA binding(GO:0008266) |
| 0.0 | 0.1 | GO:0019777 | Atg12 transferase activity(GO:0019777) |
| 0.0 | 0.4 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.0 | 0.3 | GO:0098882 | structural constituent of presynaptic active zone(GO:0098882) |
| 0.0 | 0.4 | GO:0098632 | protein binding involved in cell-cell adhesion(GO:0098632) |
| 0.0 | 0.6 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.0 | 0.6 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.0 | 0.1 | GO:0001665 | alpha-N-acetylgalactosaminide alpha-2,6-sialyltransferase activity(GO:0001665) |
| 0.0 | 0.3 | GO:0008378 | galactosyltransferase activity(GO:0008378) |
| 0.0 | 0.4 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.0 | 0.1 | GO:0008494 | translation activator activity(GO:0008494) |
| 0.0 | 0.5 | GO:0008376 | acetylgalactosaminyltransferase activity(GO:0008376) |
| 0.0 | 0.1 | GO:0005545 | 1-phosphatidylinositol binding(GO:0005545) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.6 | SA REG CASCADE OF CYCLIN EXPR | Expression of cyclins regulates progression through the cell cycle by activating cyclin-dependent kinases. |
| 0.0 | 0.3 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.0 | 0.5 | ST TUMOR NECROSIS FACTOR PATHWAY | Tumor Necrosis Factor Pathway. |
| 0.0 | 0.3 | PID EPHB FWD PATHWAY | EPHB forward signaling |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.6 | REACTOME CYCLIN A B1 ASSOCIATED EVENTS DURING G2 M TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 0.0 | 0.5 | REACTOME RIP MEDIATED NFKB ACTIVATION VIA DAI | Genes involved in RIP-mediated NFkB activation via DAI |
| 0.0 | 0.1 | REACTOME TERMINATION OF O GLYCAN BIOSYNTHESIS | Genes involved in Termination of O-glycan biosynthesis |
| 0.0 | 0.3 | REACTOME CYTOCHROME P450 ARRANGED BY SUBSTRATE TYPE | Genes involved in Cytochrome P450 - arranged by substrate type |
| 0.0 | 0.7 | REACTOME TRANSPORT TO THE GOLGI AND SUBSEQUENT MODIFICATION | Genes involved in Transport to the Golgi and subsequent modification |
| 0.0 | 0.2 | REACTOME LATENT INFECTION OF HOMO SAPIENS WITH MYCOBACTERIUM TUBERCULOSIS | Genes involved in Latent infection of Homo sapiens with Mycobacterium tuberculosis |