Project
PRJNA207719: Tissue specific transcriptome profiling
Navigation
Downloads
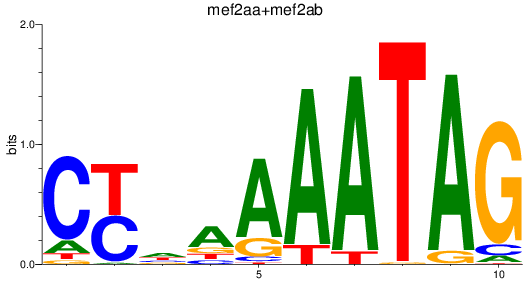
Results for mef2aa+mef2ab
Z-value: 3.67
Transcription factors associated with mef2aa+mef2ab
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
mef2aa
|
ENSDARG00000031756 | myocyte enhancer factor 2aa |
|
mef2ab
|
ENSDARG00000057527 | myocyte enhancer factor 2ab |
|
mef2ab
|
ENSDARG00000110771 | myocyte enhancer factor 2ab |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| mef2aa | dr11_v1_chr18_+_23218980_23218980 | 0.84 | 7.7e-02 | Click! |
| mef2ab | dr11_v1_chr7_+_15659280_15659280 | 0.21 | 7.3e-01 | Click! |
Activity profile of mef2aa+mef2ab motif
Sorted Z-values of mef2aa+mef2ab motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr3_-_61205711 | 3.91 |
ENSDART00000055062
|
pvalb1
|
parvalbumin 1 |
| chr19_-_7450796 | 3.69 |
ENSDART00000104750
|
mllt11
|
MLLT11, transcription factor 7 cofactor |
| chr3_+_32526263 | 3.55 |
ENSDART00000150897
|
si:ch73-367p23.2
|
si:ch73-367p23.2 |
| chr16_+_25245857 | 3.50 |
ENSDART00000155220
|
klhl38b
|
kelch-like family member 38b |
| chr25_+_31264155 | 3.43 |
ENSDART00000012256
|
tnni2a.3
|
troponin I type 2a (skeletal, fast), tandem duplicate 3 |
| chr8_-_11112058 | 3.21 |
ENSDART00000042755
|
ampd1
|
adenosine monophosphate deaminase 1 (isoform M) |
| chr5_-_72125551 | 3.03 |
ENSDART00000149412
|
smyd1a
|
SET and MYND domain containing 1a |
| chr7_+_7048245 | 3.00 |
ENSDART00000001649
|
actn3b
|
actinin alpha 3b |
| chr23_+_24926407 | 2.57 |
ENSDART00000137486
|
klhl21
|
kelch-like family member 21 |
| chr12_-_26064105 | 2.46 |
ENSDART00000168825
|
ldb3b
|
LIM domain binding 3b |
| chr7_+_6969909 | 2.29 |
ENSDART00000189886
|
actn3b
|
actinin alpha 3b |
| chr12_-_17712393 | 2.23 |
ENSDART00000143534
ENSDART00000010144 |
pvalb2
|
parvalbumin 2 |
| chr14_+_49296052 | 2.13 |
ENSDART00000006073
ENSDART00000105346 |
anxa6
|
annexin A6 |
| chr1_+_7546259 | 2.05 |
ENSDART00000015732
|
mylz3
|
myosin, light polypeptide 3, skeletal muscle |
| chr12_-_26064480 | 2.02 |
ENSDART00000158215
ENSDART00000171206 ENSDART00000171212 ENSDART00000182956 ENSDART00000186779 |
ldb3b
|
LIM domain binding 3b |
| chr6_-_18199062 | 2.01 |
ENSDART00000167513
|
ppp1r27b
|
protein phosphatase 1, regulatory subunit 27b |
| chr5_+_64368770 | 2.01 |
ENSDART00000162246
|
plpp7
|
phospholipid phosphatase 7 (inactive) |
| chr5_+_51597677 | 1.99 |
ENSDART00000048210
ENSDART00000184797 |
ckmt2b
|
creatine kinase, mitochondrial 2b (sarcomeric) |
| chr24_+_34089977 | 1.98 |
ENSDART00000157466
|
asb10
|
ankyrin repeat and SOCS box containing 10 |
| chr16_+_29663809 | 1.92 |
ENSDART00000191336
|
tmod4
|
tropomodulin 4 (muscle) |
| chr17_-_5583345 | 1.69 |
ENSDART00000035944
|
clic5a
|
chloride intracellular channel 5a |
| chr9_-_42873700 | 1.69 |
ENSDART00000125953
|
ttn.1
|
titin, tandem duplicate 1 |
| chr21_+_20383837 | 1.67 |
ENSDART00000026430
|
hspb11
|
heat shock protein, alpha-crystallin-related, b11 |
| chr1_-_13271085 | 1.60 |
ENSDART00000193663
|
pcdh18a
|
protocadherin 18a |
| chr6_-_20875111 | 1.59 |
ENSDART00000115118
ENSDART00000159916 |
tns1a
|
tensin 1a |
| chr20_-_26001288 | 1.57 |
ENSDART00000136518
ENSDART00000063177 |
capn3b
|
calpain 3b |
| chr2_-_15324837 | 1.54 |
ENSDART00000015655
|
tecrl2b
|
trans-2,3-enoyl-CoA reductase-like 2b |
| chr23_+_6077503 | 1.53 |
ENSDART00000081714
ENSDART00000139834 |
mybpha
|
myosin binding protein Ha |
| chr11_+_11200550 | 1.53 |
ENSDART00000181339
ENSDART00000187116 |
myom2a
|
myomesin 2a |
| chr5_+_64732270 | 1.51 |
ENSDART00000134241
|
olfm1a
|
olfactomedin 1a |
| chr21_-_131236 | 1.50 |
ENSDART00000160005
|
si:ch1073-398f15.1
|
si:ch1073-398f15.1 |
| chr16_+_34046733 | 1.49 |
ENSDART00000058424
|
fam46ba
|
family with sequence similarity 46, member Ba |
| chr8_+_22930627 | 1.46 |
ENSDART00000187860
|
sypa
|
synaptophysin a |
| chr24_+_9298198 | 1.46 |
ENSDART00000165780
|
otud1
|
OTU deubiquitinase 1 |
| chr25_+_20077225 | 1.46 |
ENSDART00000136543
|
tnni4b.1
|
troponin I4b, tandem duplicate 1 |
| chr17_-_14671098 | 1.40 |
ENSDART00000037371
|
ppp1r13ba
|
protein phosphatase 1, regulatory subunit 13Ba |
| chr19_-_33087246 | 1.37 |
ENSDART00000052078
|
klhl38a
|
kelch-like family member 38a |
| chrM_+_4993 | 1.33 |
ENSDART00000093600
|
mt-nd2
|
NADH dehydrogenase 2, mitochondrial |
| chr16_-_43356018 | 1.32 |
ENSDART00000181683
|
FO704821.1
|
|
| chr15_+_5923851 | 1.30 |
ENSDART00000152520
ENSDART00000145827 ENSDART00000121529 |
sh3bgr
|
SH3 domain binding glutamate-rich protein |
| chr12_-_4781801 | 1.30 |
ENSDART00000167490
ENSDART00000121718 |
mapta
|
microtubule-associated protein tau a |
| chr8_-_26388090 | 1.29 |
ENSDART00000147912
|
si:dkey-20d21.12
|
si:dkey-20d21.12 |
| chr13_-_2215213 | 1.29 |
ENSDART00000129773
|
mlip
|
muscular LMNA-interacting protein |
| chr16_-_17200120 | 1.23 |
ENSDART00000147739
|
gapdh
|
glyceraldehyde-3-phosphate dehydrogenase |
| chr6_+_14301214 | 1.22 |
ENSDART00000129491
|
tmem182b
|
transmembrane protein 182b |
| chr13_+_22480857 | 1.20 |
ENSDART00000078721
ENSDART00000044719 ENSDART00000130957 ENSDART00000078757 ENSDART00000130424 ENSDART00000078747 |
ldb3a
|
LIM domain binding 3a |
| chr5_-_57635664 | 1.16 |
ENSDART00000074306
|
hspb2
|
heat shock protein, alpha-crystallin-related, b2 |
| chr19_-_31035155 | 1.16 |
ENSDART00000161882
|
bzw2
|
basic leucine zipper and W2 domains 2 |
| chr21_-_27507786 | 1.15 |
ENSDART00000077657
ENSDART00000191284 ENSDART00000178229 ENSDART00000186153 |
CAPN1
|
si:dkeyp-50d11.2 |
| chr9_-_23891102 | 1.14 |
ENSDART00000186799
|
asb18
|
ankyrin repeat and SOCS box containing 18 |
| chr5_-_48260145 | 1.14 |
ENSDART00000044083
ENSDART00000163250 ENSDART00000135911 |
mef2cb
|
myocyte enhancer factor 2cb |
| chr12_+_6002715 | 1.13 |
ENSDART00000114961
|
si:ch211-131k2.3
|
si:ch211-131k2.3 |
| chr22_-_7006974 | 1.13 |
ENSDART00000133143
|
gpd1b
|
glycerol-3-phosphate dehydrogenase 1b |
| chr23_-_19953089 | 1.09 |
ENSDART00000153828
|
atp2b3b
|
ATPase plasma membrane Ca2+ transporting 3b |
| chrM_-_15232 | 1.08 |
ENSDART00000093623
|
mt-nd6
|
NADH dehydrogenase 6, mitochondrial |
| chr7_+_69841017 | 1.08 |
ENSDART00000169107
|
FO818704.1
|
|
| chr18_+_23193820 | 1.06 |
ENSDART00000148106
|
mef2aa
|
myocyte enhancer factor 2aa |
| chr1_-_13271569 | 1.04 |
ENSDART00000127838
|
pcdh18a
|
protocadherin 18a |
| chr18_+_23193567 | 1.04 |
ENSDART00000190072
ENSDART00000171594 ENSDART00000181762 |
mef2aa
|
myocyte enhancer factor 2aa |
| chr16_-_29164379 | 1.04 |
ENSDART00000132589
|
mef2d
|
myocyte enhancer factor 2d |
| chr21_-_32487061 | 1.03 |
ENSDART00000114359
ENSDART00000131591 ENSDART00000131477 |
si:dkeyp-72g9.4
|
si:dkeyp-72g9.4 |
| chr6_+_12462079 | 1.03 |
ENSDART00000192029
ENSDART00000065385 |
nr4a2b
|
nuclear receptor subfamily 4, group A, member 2b |
| chr13_-_30149973 | 1.01 |
ENSDART00000041515
|
sar1ab
|
secretion associated, Ras related GTPase 1Ab |
| chr18_+_26719787 | 1.01 |
ENSDART00000141672
|
alpk3a
|
alpha-kinase 3a |
| chr23_+_36601984 | 1.01 |
ENSDART00000128598
|
igfbp6b
|
insulin-like growth factor binding protein 6b |
| chr19_-_31035325 | 1.00 |
ENSDART00000147504
|
bzw2
|
basic leucine zipper and W2 domains 2 |
| chr1_-_11075403 | 1.00 |
ENSDART00000102903
ENSDART00000170290 |
dmd
|
dystrophin |
| chr3_+_40170216 | 0.98 |
ENSDART00000011568
|
syngr3a
|
synaptogyrin 3a |
| chr20_-_36575475 | 0.95 |
ENSDART00000062893
|
enah
|
enabled homolog (Drosophila) |
| chr7_-_23745984 | 0.95 |
ENSDART00000048050
|
ITGB1BP2
|
zgc:92429 |
| chr1_+_6171585 | 0.94 |
ENSDART00000024358
|
prkag3a
|
protein kinase, AMP-activated, gamma 3a non-catalytic subunit |
| chr10_-_22845485 | 0.93 |
ENSDART00000079454
|
vamp2
|
vesicle-associated membrane protein 2 |
| chr25_-_23745765 | 0.92 |
ENSDART00000089278
|
si:ch211-236l14.4
|
si:ch211-236l14.4 |
| chr1_-_42778510 | 0.90 |
ENSDART00000190172
|
lrrtm1
|
leucine rich repeat transmembrane neuronal 1 |
| chr3_-_32170850 | 0.88 |
ENSDART00000055307
ENSDART00000157366 |
tnnt1
|
troponin T type 1 (skeletal, slow) |
| chr1_+_8601935 | 0.87 |
ENSDART00000152367
|
si:ch211-160d14.6
|
si:ch211-160d14.6 |
| chr1_+_33969015 | 0.87 |
ENSDART00000042984
ENSDART00000146530 |
epha6
|
eph receptor A6 |
| chr25_+_14087045 | 0.86 |
ENSDART00000155770
|
actc1c
|
actin, alpha, cardiac muscle 1c |
| chr14_-_24332786 | 0.86 |
ENSDART00000173164
|
fam13b
|
family with sequence similarity 13, member B |
| chr15_-_22139566 | 0.86 |
ENSDART00000149017
|
scn3b
|
sodium channel, voltage-gated, type III, beta |
| chr3_-_52614747 | 0.85 |
ENSDART00000154365
|
trim35-13
|
tripartite motif containing 35-13 |
| chr13_+_28821841 | 0.84 |
ENSDART00000179900
|
CU639469.1
|
|
| chr7_-_66864756 | 0.83 |
ENSDART00000184462
ENSDART00000189424 |
ampd3a
|
adenosine monophosphate deaminase 3a |
| chr17_-_12336987 | 0.83 |
ENSDART00000172001
|
snap25b
|
synaptosomal-associated protein, 25b |
| chr6_+_57541776 | 0.83 |
ENSDART00000157330
|
necab3
|
N-terminal EF-hand calcium binding protein 3 |
| chr3_-_18189283 | 0.82 |
ENSDART00000049240
|
tob1a
|
transducer of ERBB2, 1a |
| chr3_+_59051503 | 0.82 |
ENSDART00000160767
|
rasd4
|
rasd family member 4 |
| chr16_+_26777473 | 0.81 |
ENSDART00000188870
|
cdh17
|
cadherin 17, LI cadherin (liver-intestine) |
| chr7_-_26270014 | 0.80 |
ENSDART00000079347
|
serpine1
|
serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 1 |
| chr23_+_20110086 | 0.80 |
ENSDART00000054664
|
tnnc1b
|
troponin C type 1b (slow) |
| chr14_-_32405387 | 0.76 |
ENSDART00000184647
|
fgf13a
|
fibroblast growth factor 13a |
| chr6_-_18698456 | 0.74 |
ENSDART00000166587
|
rhbdl3
|
rhomboid, veinlet-like 3 (Drosophila) |
| chr23_+_33907899 | 0.71 |
ENSDART00000159445
|
cs
|
citrate synthase |
| chr14_+_22113331 | 0.71 |
ENSDART00000109759
|
tmx2a
|
thioredoxin-related transmembrane protein 2a |
| chr20_-_40717900 | 0.70 |
ENSDART00000181663
|
cx43
|
connexin 43 |
| chr25_+_3677650 | 0.69 |
ENSDART00000154348
|
prnprs3
|
prion protein, related sequence 3 |
| chr2_-_28102264 | 0.68 |
ENSDART00000013638
|
cdh6
|
cadherin 6 |
| chr7_+_40093662 | 0.68 |
ENSDART00000053011
|
si:ch73-174h16.4
|
si:ch73-174h16.4 |
| chr18_+_10820430 | 0.67 |
ENSDART00000161990
|
mical3a
|
microtubule associated monooxygenase, calponin and LIM domain containing 3a |
| chr8_+_26432677 | 0.65 |
ENSDART00000078369
ENSDART00000131925 |
zgc:136971
|
zgc:136971 |
| chr20_+_34455645 | 0.65 |
ENSDART00000135789
|
mettl11b
|
methyltransferase like 11B |
| chr17_-_16069905 | 0.64 |
ENSDART00000110383
|
map7a
|
microtubule-associated protein 7a |
| chr2_+_2957515 | 0.64 |
ENSDART00000160715
|
pik3r3a
|
phosphoinositide-3-kinase, regulatory subunit 3a (gamma) |
| chr21_+_40589770 | 0.63 |
ENSDART00000164650
ENSDART00000161584 ENSDART00000161108 |
pdk3b
|
pyruvate dehydrogenase kinase, isozyme 3b |
| chr13_+_38817871 | 0.63 |
ENSDART00000187708
|
col19a1
|
collagen, type XIX, alpha 1 |
| chr17_+_8184649 | 0.62 |
ENSDART00000091818
|
tulp4b
|
tubby like protein 4b |
| chr17_-_5769196 | 0.62 |
ENSDART00000113885
|
si:dkey-100n19.2
|
si:dkey-100n19.2 |
| chr5_+_69808763 | 0.61 |
ENSDART00000143482
|
fsd1l
|
fibronectin type III and SPRY domain containing 1-like |
| chr9_+_54644626 | 0.60 |
ENSDART00000190609
|
egfl6
|
EGF-like-domain, multiple 6 |
| chr24_+_38155830 | 0.60 |
ENSDART00000152019
|
si:ch211-234p6.5
|
si:ch211-234p6.5 |
| chr20_-_26066020 | 0.59 |
ENSDART00000078559
|
myct1a
|
myc target 1a |
| chr5_-_8711157 | 0.59 |
ENSDART00000062957
|
paip1
|
poly(A) binding protein interacting protein 1 |
| chr21_+_11685009 | 0.58 |
ENSDART00000014668
|
pcsk1
|
proprotein convertase subtilisin/kexin type 1 |
| chr16_+_5774977 | 0.57 |
ENSDART00000134202
|
ccka
|
cholecystokinin a |
| chr3_+_31953145 | 0.57 |
ENSDART00000148861
|
kcnc3a
|
potassium voltage-gated channel, Shaw-related subfamily, member 3a |
| chr14_+_23520986 | 0.55 |
ENSDART00000170473
ENSDART00000175970 |
si:ch211-221f10.2
|
si:ch211-221f10.2 |
| chr23_-_7797207 | 0.55 |
ENSDART00000181611
|
myt1b
|
myelin transcription factor 1b |
| chr4_-_2719128 | 0.54 |
ENSDART00000184693
|
slco1c1
|
solute carrier organic anion transporter family, member 1C1 |
| chr12_+_26632448 | 0.54 |
ENSDART00000185762
|
arhgap12b
|
Rho GTPase activating protein 12b |
| chr14_+_23518110 | 0.54 |
ENSDART00000112930
|
si:ch211-221f10.2
|
si:ch211-221f10.2 |
| chr8_-_18010735 | 0.53 |
ENSDART00000125014
|
acot11b
|
acyl-CoA thioesterase 11b |
| chr12_-_19119176 | 0.53 |
ENSDART00000149180
|
aco2
|
aconitase 2, mitochondrial |
| chr25_+_11389220 | 0.52 |
ENSDART00000187787
|
AGBL1
|
si:ch73-141f14.1 |
| chr11_-_33960318 | 0.52 |
ENSDART00000087597
|
col6a2
|
collagen, type VI, alpha 2 |
| chr23_-_27633730 | 0.51 |
ENSDART00000103639
|
arf3a
|
ADP-ribosylation factor 3a |
| chr19_-_10394931 | 0.51 |
ENSDART00000191549
|
zgc:194578
|
zgc:194578 |
| chr23_+_14097508 | 0.51 |
ENSDART00000087280
|
cacnb3a
|
calcium channel, voltage-dependent, beta 3a |
| chr10_+_36458563 | 0.50 |
ENSDART00000077008
|
alox5ap
|
arachidonate 5-lipoxygenase-activating protein |
| chr19_-_11315224 | 0.50 |
ENSDART00000104933
|
eepd1
|
endonuclease/exonuclease/phosphatase family domain containing 1 |
| chr7_+_72882446 | 0.50 |
ENSDART00000131113
ENSDART00000174223 |
ADGRL3
|
adhesion G protein-coupled receptor L3 |
| chr10_-_24371312 | 0.50 |
ENSDART00000149362
|
pitpnab
|
phosphatidylinositol transfer protein, alpha b |
| chr8_-_11229523 | 0.49 |
ENSDART00000002164
|
unc45b
|
unc-45 myosin chaperone B |
| chr21_+_11684830 | 0.49 |
ENSDART00000147473
|
pcsk1
|
proprotein convertase subtilisin/kexin type 1 |
| chr15_+_15856178 | 0.48 |
ENSDART00000080338
|
dusp14
|
dual specificity phosphatase 14 |
| chr1_+_20084389 | 0.47 |
ENSDART00000140263
|
prss12
|
protease, serine, 12 (neurotrypsin, motopsin) |
| chr16_-_31475904 | 0.47 |
ENSDART00000145691
|
si:ch211-251p5.5
|
si:ch211-251p5.5 |
| chr18_-_38087875 | 0.47 |
ENSDART00000111301
|
luzp2
|
leucine zipper protein 2 |
| chr8_-_17516448 | 0.46 |
ENSDART00000100667
|
skia
|
v-ski avian sarcoma viral oncogene homolog a |
| chr2_-_3678029 | 0.46 |
ENSDART00000146861
|
mmp16b
|
matrix metallopeptidase 16b (membrane-inserted) |
| chr5_-_65153731 | 0.45 |
ENSDART00000171024
|
sh2d3cb
|
SH2 domain containing 3Cb |
| chr10_-_8033468 | 0.44 |
ENSDART00000140476
|
atp6v0a2a
|
ATPase H+ transporting V0 subunit a2a |
| chr16_-_16590489 | 0.43 |
ENSDART00000190021
|
si:ch211-257p13.3
|
si:ch211-257p13.3 |
| chr8_-_12264486 | 0.42 |
ENSDART00000091612
ENSDART00000135812 |
dab2ipa
|
DAB2 interacting protein a |
| chr18_-_16791331 | 0.42 |
ENSDART00000148222
|
ampd3b
|
adenosine monophosphate deaminase 3b |
| chr4_+_5848229 | 0.42 |
ENSDART00000161101
ENSDART00000067357 |
lyrm5a
|
LYR motif containing 5a |
| chr16_-_36798783 | 0.41 |
ENSDART00000145697
|
calb1
|
calbindin 1 |
| chr10_+_21776911 | 0.38 |
ENSDART00000163077
ENSDART00000186093 |
pcdh1g22
|
protocadherin 1 gamma 22 |
| chr6_-_894006 | 0.37 |
ENSDART00000171091
|
zeb2b
|
zinc finger E-box binding homeobox 2b |
| chr1_+_10378706 | 0.36 |
ENSDART00000046283
ENSDART00000103535 ENSDART00000132076 |
dachb
|
dachshund b |
| chr18_+_34601023 | 0.35 |
ENSDART00000149647
|
tiparp
|
TCDD-inducible poly(ADP-ribose) polymerase |
| chr20_-_47732703 | 0.35 |
ENSDART00000193975
|
tfap2d
|
transcription factor AP-2 delta (activating enhancer binding protein 2 delta) |
| chr14_+_30328567 | 0.35 |
ENSDART00000105889
|
mtus1a
|
microtubule associated tumor suppressor 1a |
| chr6_-_10863582 | 0.35 |
ENSDART00000135454
ENSDART00000133893 ENSDART00000014521 ENSDART00000164342 |
lnpk
|
lunapark, ER junction formation factor |
| chr4_+_9279515 | 0.34 |
ENSDART00000048707
|
srgap1b
|
SLIT-ROBO Rho GTPase activating protein 1b |
| chr9_+_29643036 | 0.33 |
ENSDART00000023210
ENSDART00000175160 |
trim13
|
tripartite motif containing 13 |
| chr4_+_9279784 | 0.33 |
ENSDART00000014897
|
srgap1b
|
SLIT-ROBO Rho GTPase activating protein 1b |
| chr22_+_25841786 | 0.33 |
ENSDART00000180863
|
vasna
|
vasorin a |
| chr19_+_30990815 | 0.32 |
ENSDART00000134645
|
sync
|
syncoilin, intermediate filament protein |
| chr2_+_35880600 | 0.31 |
ENSDART00000004277
|
lamc1
|
laminin, gamma 1 |
| chr14_-_46451249 | 0.31 |
ENSDART00000058466
|
fgfbp2a
|
fibroblast growth factor binding protein 2a |
| chr4_-_25908871 | 0.31 |
ENSDART00000066949
|
fgd6
|
FYVE, RhoGEF and PH domain containing 6 |
| chr7_+_39006837 | 0.30 |
ENSDART00000173735
|
dgkza
|
diacylglycerol kinase, zeta a |
| chr10_+_41765616 | 0.30 |
ENSDART00000170682
|
rnf34b
|
ring finger protein 34b |
| chr10_+_32561317 | 0.30 |
ENSDART00000109029
|
map6a
|
microtubule-associated protein 6a |
| chr10_-_26512742 | 0.30 |
ENSDART00000135951
|
si:dkey-5g14.1
|
si:dkey-5g14.1 |
| chr9_+_29040425 | 0.30 |
ENSDART00000150201
|
MRAS
|
si:ch73-116o1.2 |
| chr1_-_35653560 | 0.29 |
ENSDART00000142154
|
frem3
|
Fras1 related extracellular matrix 3 |
| chr24_+_25471196 | 0.29 |
ENSDART00000066625
|
smpx
|
small muscle protein, X-linked |
| chr8_-_42238543 | 0.27 |
ENSDART00000062697
|
gfra2a
|
GDNF family receptor alpha 2a |
| chr16_-_19890303 | 0.27 |
ENSDART00000147161
ENSDART00000079159 |
hdac9b
|
histone deacetylase 9b |
| chr16_-_32837806 | 0.26 |
ENSDART00000003997
|
MCHR2
|
si:dkey-165n16.5 |
| chr7_+_26716321 | 0.25 |
ENSDART00000189750
|
cd82a
|
CD82 molecule a |
| chr12_-_4028079 | 0.25 |
ENSDART00000128676
|
si:ch211-180a12.2
|
si:ch211-180a12.2 |
| chr1_+_35495368 | 0.25 |
ENSDART00000053806
|
gab1
|
GRB2-associated binding protein 1 |
| chr12_-_20340543 | 0.25 |
ENSDART00000055623
|
hbbe3
|
hemoglobin beta embryonic-3 |
| chr7_+_33424044 | 0.24 |
ENSDART00000180260
|
glceb
|
glucuronic acid epimerase b |
| chr1_-_45662774 | 0.24 |
ENSDART00000042158
|
serhl
|
serine hydrolase-like |
| chr7_+_40094081 | 0.24 |
ENSDART00000186054
|
si:ch73-174h16.4
|
si:ch73-174h16.4 |
| chr17_+_33495194 | 0.23 |
ENSDART00000033691
|
pth2
|
parathyroid hormone 2 |
| chr3_-_42981739 | 0.23 |
ENSDART00000167844
|
mafk
|
v-maf avian musculoaponeurotic fibrosarcoma oncogene homolog K |
| chr3_+_32403758 | 0.23 |
ENSDART00000156982
|
si:ch211-195b15.8
|
si:ch211-195b15.8 |
| chr8_-_32497815 | 0.23 |
ENSDART00000122359
|
si:dkey-164f24.2
|
si:dkey-164f24.2 |
| chr15_+_28940352 | 0.22 |
ENSDART00000154085
|
gipr
|
gastric inhibitory polypeptide receptor |
| chr5_-_20678300 | 0.21 |
ENSDART00000088639
|
wscd2
|
WSC domain containing 2 |
| chr9_-_28990649 | 0.21 |
ENSDART00000078823
|
ptpn4a
|
protein tyrosine phosphatase, non-receptor type 4a |
| chr5_-_31926906 | 0.21 |
ENSDART00000187340
|
ssh1b
|
slingshot protein phosphatase 1b |
| chr11_-_18253111 | 0.20 |
ENSDART00000125984
|
mustn1b
|
musculoskeletal, embryonic nuclear protein 1b |
| chr17_+_28340138 | 0.20 |
ENSDART00000033943
|
mdga2a
|
MAM domain containing glycosylphosphatidylinositol anchor 2a |
| chr7_-_22956889 | 0.20 |
ENSDART00000101447
|
tnfsf10l
|
TNF superfamily member 10, like |
| chr1_-_40102836 | 0.20 |
ENSDART00000147317
|
cntf
|
ciliary neurotrophic factor |
| chr12_+_6041575 | 0.20 |
ENSDART00000091868
|
g6pca.2
|
glucose-6-phosphatase a, catalytic subunit, tandem duplicate 2 |
| chr8_+_21159122 | 0.19 |
ENSDART00000033491
|
spryd4
|
SPRY domain containing 4 |
| chr9_+_27876146 | 0.19 |
ENSDART00000133997
|
armc8
|
armadillo repeat containing 8 |
| chr20_+_2134816 | 0.19 |
ENSDART00000039249
|
l3mbtl3
|
l(3)mbt-like 3 (Drosophila) |
| chr22_+_10678141 | 0.19 |
ENSDART00000193341
|
hyal2b
|
hyaluronoglucosaminidase 2b |
| chr24_-_6647275 | 0.19 |
ENSDART00000161494
|
arhgap21a
|
Rho GTPase activating protein 21a |
| chr18_+_45792035 | 0.18 |
ENSDART00000135045
|
abcc5
|
ATP-binding cassette, sub-family C (CFTR/MRP), member 5 |
| chr11_+_41560792 | 0.18 |
ENSDART00000127292
|
kcnab2a
|
potassium voltage-gated channel, shaker-related subfamily, beta member 2 a |
| chr7_+_7511914 | 0.18 |
ENSDART00000172848
|
clcn3
|
chloride channel 3 |
| chr7_+_42206847 | 0.18 |
ENSDART00000149250
|
phkb
|
phosphorylase kinase, beta |
Network of associatons between targets according to the STRING database.
First level regulatory network of mef2aa+mef2ab
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 3.7 | GO:0051881 | regulation of mitochondrial membrane potential(GO:0051881) regulation of release of cytochrome c from mitochondria(GO:0090199) positive regulation of release of cytochrome c from mitochondria(GO:0090200) |
| 0.5 | 2.1 | GO:0003418 | plasma membrane repair(GO:0001778) chondrocyte differentiation involved in endochondral bone morphogenesis(GO:0003413) growth plate cartilage chondrocyte differentiation(GO:0003418) |
| 0.5 | 1.6 | GO:0097264 | self proteolysis(GO:0097264) |
| 0.3 | 7.5 | GO:0031033 | myosin filament organization(GO:0031033) |
| 0.3 | 4.5 | GO:0032264 | IMP salvage(GO:0032264) |
| 0.3 | 0.9 | GO:0070358 | actin polymerization-dependent cell motility(GO:0070358) |
| 0.3 | 1.1 | GO:0046168 | NADH oxidation(GO:0006116) glycerol-3-phosphate catabolic process(GO:0046168) |
| 0.3 | 1.0 | GO:0090113 | regulation of COPII vesicle coating(GO:0003400) regulation of ER to Golgi vesicle-mediated transport by GTP hydrolysis(GO:0090113) |
| 0.2 | 1.9 | GO:0051694 | pointed-end actin filament capping(GO:0051694) |
| 0.2 | 0.9 | GO:0086005 | ventricular cardiac muscle cell action potential(GO:0086005) |
| 0.2 | 0.8 | GO:0010755 | regulation of plasminogen activation(GO:0010755) |
| 0.2 | 0.9 | GO:0021731 | trigeminal motor nucleus development(GO:0021731) |
| 0.2 | 0.9 | GO:0031444 | slow-twitch skeletal muscle fiber contraction(GO:0031444) |
| 0.2 | 1.5 | GO:1902868 | positive regulation of neural retina development(GO:0061075) positive regulation of retina development in camera-type eye(GO:1902868) |
| 0.2 | 2.0 | GO:0006603 | phosphagen metabolic process(GO:0006599) phosphocreatine metabolic process(GO:0006603) phosphagen biosynthetic process(GO:0042396) phosphocreatine biosynthetic process(GO:0046314) |
| 0.2 | 2.6 | GO:0034629 | cellular protein complex localization(GO:0034629) |
| 0.1 | 0.5 | GO:0003322 | pancreatic A cell development(GO:0003322) |
| 0.1 | 5.7 | GO:0003009 | skeletal muscle contraction(GO:0003009) |
| 0.1 | 0.8 | GO:0042983 | amyloid precursor protein biosynthetic process(GO:0042983) regulation of amyloid precursor protein biosynthetic process(GO:0042984) |
| 0.1 | 0.3 | GO:1903373 | regulation of endoplasmic reticulum tubular network organization(GO:1903371) positive regulation of endoplasmic reticulum tubular network organization(GO:1903373) |
| 0.1 | 1.0 | GO:0043435 | response to corticotropin-releasing hormone(GO:0043435) cellular response to corticotropin-releasing hormone stimulus(GO:0071376) |
| 0.1 | 0.4 | GO:1900271 | regulation of presynaptic cytosolic calcium ion concentration(GO:0099509) regulation of long-term synaptic potentiation(GO:1900271) |
| 0.1 | 0.5 | GO:0019370 | leukotriene metabolic process(GO:0006691) leukotriene biosynthetic process(GO:0019370) |
| 0.1 | 3.3 | GO:0016339 | calcium-dependent cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0016339) |
| 0.1 | 2.0 | GO:0010923 | negative regulation of phosphatase activity(GO:0010923) |
| 0.1 | 0.9 | GO:0043567 | regulation of insulin-like growth factor receptor signaling pathway(GO:0043567) |
| 0.1 | 1.0 | GO:0007525 | somatic muscle development(GO:0007525) |
| 0.1 | 1.1 | GO:0051131 | chaperone-mediated protein complex assembly(GO:0051131) |
| 0.1 | 1.3 | GO:0006120 | mitochondrial electron transport, NADH to ubiquinone(GO:0006120) |
| 0.1 | 0.3 | GO:2001238 | positive regulation of extrinsic apoptotic signaling pathway(GO:2001238) |
| 0.1 | 1.1 | GO:0016486 | peptide hormone processing(GO:0016486) |
| 0.1 | 0.9 | GO:0035493 | SNARE complex assembly(GO:0035493) |
| 0.1 | 0.8 | GO:0045162 | clustering of voltage-gated sodium channels(GO:0045162) |
| 0.1 | 0.9 | GO:1901642 | nucleoside transport(GO:0015858) nucleoside transmembrane transport(GO:1901642) |
| 0.1 | 0.8 | GO:1905150 | regulation of voltage-gated sodium channel activity(GO:1905150) |
| 0.1 | 0.4 | GO:0030948 | negative regulation of vascular endothelial growth factor receptor signaling pathway(GO:0030948) reelin-mediated signaling pathway(GO:0038026) |
| 0.1 | 1.5 | GO:0070536 | protein K63-linked deubiquitination(GO:0070536) |
| 0.0 | 0.6 | GO:0043550 | regulation of lipid kinase activity(GO:0043550) regulation of phosphatidylinositol 3-kinase activity(GO:0043551) |
| 0.0 | 0.5 | GO:0061337 | cardiac conduction(GO:0061337) |
| 0.0 | 0.3 | GO:2001270 | negative regulation of execution phase of apoptosis(GO:1900118) regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:2001270) negative regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:2001271) |
| 0.0 | 0.2 | GO:0035988 | chondrocyte proliferation(GO:0035988) |
| 0.0 | 0.7 | GO:0014823 | response to activity(GO:0014823) |
| 0.0 | 1.2 | GO:0050821 | protein stabilization(GO:0050821) |
| 0.0 | 0.7 | GO:0043252 | sodium-independent organic anion transport(GO:0043252) |
| 0.0 | 0.5 | GO:0045851 | vacuolar acidification(GO:0007035) pH reduction(GO:0045851) intracellular pH reduction(GO:0051452) |
| 0.0 | 0.6 | GO:0010811 | positive regulation of cell-substrate adhesion(GO:0010811) |
| 0.0 | 0.2 | GO:0098957 | anterograde axonal transport of mitochondrion(GO:0098957) |
| 0.0 | 0.8 | GO:0042761 | very long-chain fatty acid biosynthetic process(GO:0042761) |
| 0.0 | 3.7 | GO:0001756 | somitogenesis(GO:0001756) |
| 0.0 | 1.1 | GO:0060914 | heart formation(GO:0060914) |
| 0.0 | 0.8 | GO:0042149 | cellular response to glucose starvation(GO:0042149) |
| 0.0 | 0.6 | GO:0006446 | regulation of translational initiation(GO:0006446) |
| 0.0 | 0.2 | GO:0030202 | heparin metabolic process(GO:0030202) heparin biosynthetic process(GO:0030210) |
| 0.0 | 0.2 | GO:0098900 | regulation of action potential(GO:0098900) |
| 0.0 | 0.2 | GO:0050908 | detection of light stimulus involved in visual perception(GO:0050908) detection of light stimulus involved in sensory perception(GO:0050962) |
| 0.0 | 0.2 | GO:0035970 | peptidyl-threonine dephosphorylation(GO:0035970) |
| 0.0 | 0.2 | GO:0030214 | hyaluronan catabolic process(GO:0030214) |
| 0.0 | 1.0 | GO:0010842 | retina layer formation(GO:0010842) |
| 0.0 | 0.5 | GO:0006099 | tricarboxylic acid cycle(GO:0006099) |
| 0.0 | 2.0 | GO:0006821 | chloride transport(GO:0006821) |
| 0.0 | 0.6 | GO:0007586 | digestion(GO:0007586) |
| 0.0 | 0.6 | GO:0010906 | regulation of glucose metabolic process(GO:0010906) |
| 0.0 | 0.2 | GO:0015671 | oxygen transport(GO:0015671) |
| 0.0 | 0.1 | GO:0044528 | regulation of mitochondrial mRNA stability(GO:0044528) |
| 0.0 | 1.1 | GO:0006936 | muscle contraction(GO:0006936) |
| 0.0 | 0.3 | GO:0007205 | protein kinase C-activating G-protein coupled receptor signaling pathway(GO:0007205) |
| 0.0 | 0.1 | GO:0046865 | retinol transport(GO:0034633) isoprenoid transport(GO:0046864) terpenoid transport(GO:0046865) |
| 0.0 | 0.1 | GO:0006072 | glycerol-3-phosphate metabolic process(GO:0006072) |
| 0.0 | 0.7 | GO:0030336 | negative regulation of cell migration(GO:0030336) |
| 0.0 | 5.1 | GO:0061061 | muscle structure development(GO:0061061) |
| 0.0 | 0.1 | GO:0033539 | fatty acid beta-oxidation using acyl-CoA dehydrogenase(GO:0033539) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 4.8 | GO:0031430 | M band(GO:0031430) |
| 0.2 | 1.2 | GO:0009331 | glycerol-3-phosphate dehydrogenase complex(GO:0009331) |
| 0.2 | 6.6 | GO:0005861 | troponin complex(GO:0005861) |
| 0.1 | 0.8 | GO:0070032 | synaptobrevin 2-SNAP-25-syntaxin-1a-complexin I complex(GO:0070032) |
| 0.1 | 1.9 | GO:0036379 | striated muscle thin filament(GO:0005865) myofilament(GO:0036379) |
| 0.1 | 5.7 | GO:0031941 | filamentous actin(GO:0031941) |
| 0.1 | 0.3 | GO:0098826 | endoplasmic reticulum tubular network membrane(GO:0098826) |
| 0.1 | 2.6 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.1 | 0.5 | GO:0043083 | synaptic cleft(GO:0043083) |
| 0.1 | 0.9 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.1 | 0.5 | GO:0031672 | A band(GO:0031672) |
| 0.1 | 2.1 | GO:0042470 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.0 | 0.1 | GO:0033185 | dolichol-phosphate-mannose synthase complex(GO:0033185) |
| 0.0 | 0.5 | GO:0000220 | vacuolar proton-transporting V-type ATPase, V0 domain(GO:0000220) |
| 0.0 | 1.0 | GO:0016010 | dystrophin-associated glycoprotein complex(GO:0016010) glycoprotein complex(GO:0090665) |
| 0.0 | 0.2 | GO:0034657 | GID complex(GO:0034657) |
| 0.0 | 0.9 | GO:0034706 | voltage-gated sodium channel complex(GO:0001518) sodium channel complex(GO:0034706) |
| 0.0 | 1.0 | GO:0031594 | neuromuscular junction(GO:0031594) |
| 0.0 | 1.3 | GO:0016605 | PML body(GO:0016605) |
| 0.0 | 0.6 | GO:0044298 | neuronal cell body membrane(GO:0032809) cell body membrane(GO:0044298) |
| 0.0 | 2.4 | GO:0070469 | respiratory chain(GO:0070469) |
| 0.0 | 1.5 | GO:0016342 | catenin complex(GO:0016342) |
| 0.0 | 1.7 | GO:0034707 | chloride channel complex(GO:0034707) |
| 0.0 | 0.9 | GO:0032154 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.0 | 0.4 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.0 | 1.5 | GO:0099501 | synaptic vesicle membrane(GO:0030672) exocytic vesicle membrane(GO:0099501) |
| 0.0 | 0.2 | GO:0031838 | haptoglobin-hemoglobin complex(GO:0031838) |
| 0.0 | 0.3 | GO:0030127 | COPII vesicle coat(GO:0030127) |
| 0.0 | 0.3 | GO:0005801 | cis-Golgi network(GO:0005801) |
| 0.0 | 0.2 | GO:0005964 | phosphorylase kinase complex(GO:0005964) |
| 0.0 | 0.2 | GO:0044224 | juxtaparanode region of axon(GO:0044224) |
| 0.0 | 0.8 | GO:0031985 | Golgi cisterna(GO:0031985) |
| 0.0 | 0.6 | GO:0005942 | phosphatidylinositol 3-kinase complex(GO:0005942) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 4.5 | GO:0047623 | AMP deaminase activity(GO:0003876) adenosine-phosphate deaminase activity(GO:0047623) |
| 0.3 | 1.3 | GO:0005521 | lamin binding(GO:0005521) |
| 0.3 | 1.2 | GO:0004365 | glyceraldehyde-3-phosphate dehydrogenase (NAD+) (phosphorylating) activity(GO:0004365) glyceraldehyde-3-phosphate dehydrogenase (NAD(P)+) (phosphorylating) activity(GO:0043891) |
| 0.3 | 5.7 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) alpha-actinin binding(GO:0051393) |
| 0.3 | 1.1 | GO:0004367 | glycerol-3-phosphate dehydrogenase [NAD+] activity(GO:0004367) |
| 0.2 | 0.5 | GO:0003994 | aconitate hydratase activity(GO:0003994) |
| 0.2 | 2.0 | GO:0004111 | creatine kinase activity(GO:0004111) phosphotransferase activity, nitrogenous group as acceptor(GO:0016775) |
| 0.1 | 2.8 | GO:0005523 | tropomyosin binding(GO:0005523) |
| 0.1 | 0.5 | GO:0004464 | leukotriene-C4 synthase activity(GO:0004464) |
| 0.1 | 2.4 | GO:0050136 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.1 | 0.9 | GO:0008504 | monoamine transmembrane transporter activity(GO:0008504) |
| 0.1 | 1.6 | GO:0051879 | Hsp90 protein binding(GO:0051879) |
| 0.1 | 0.9 | GO:0019871 | sodium channel inhibitor activity(GO:0019871) |
| 0.1 | 0.9 | GO:0005522 | profilin binding(GO:0005522) |
| 0.1 | 1.0 | GO:0035259 | glucocorticoid receptor binding(GO:0035259) |
| 0.1 | 0.7 | GO:0046912 | transferase activity, transferring acyl groups, acyl groups converted into alkyl on transfer(GO:0046912) |
| 0.1 | 0.9 | GO:0031994 | insulin-like growth factor I binding(GO:0031994) insulin-like growth factor II binding(GO:0031995) |
| 0.1 | 3.2 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.1 | 0.2 | GO:0004377 | GDP-Man:Man3GlcNAc2-PP-Dol alpha-1,2-mannosyltransferase activity(GO:0004377) |
| 0.1 | 0.6 | GO:0004740 | pyruvate dehydrogenase (acetyl-transferring) kinase activity(GO:0004740) |
| 0.1 | 1.4 | GO:0002039 | p53 binding(GO:0002039) |
| 0.1 | 0.5 | GO:0036042 | long-chain fatty acyl-CoA binding(GO:0036042) |
| 0.1 | 0.5 | GO:0008526 | phosphatidylinositol transporter activity(GO:0008526) |
| 0.1 | 0.6 | GO:0008494 | translation activator activity(GO:0008494) |
| 0.1 | 0.2 | GO:0047464 | heparosan-N-sulfate-glucuronate 5-epimerase activity(GO:0047464) |
| 0.1 | 2.9 | GO:0042826 | histone deacetylase binding(GO:0042826) |
| 0.1 | 0.9 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.1 | 1.6 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.0 | 0.2 | GO:0033906 | hyaluronoglucuronidase activity(GO:0033906) |
| 0.0 | 0.1 | GO:0047291 | lactosylceramide alpha-2,3-sialyltransferase activity(GO:0047291) |
| 0.0 | 2.1 | GO:0001786 | phosphatidylserine binding(GO:0001786) |
| 0.0 | 3.0 | GO:0018024 | histone-lysine N-methyltransferase activity(GO:0018024) |
| 0.0 | 0.2 | GO:0016519 | gastric inhibitory peptide receptor activity(GO:0016519) |
| 0.0 | 0.8 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.0 | 1.1 | GO:0030165 | PDZ domain binding(GO:0030165) |
| 0.0 | 0.7 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) MAP kinase phosphatase activity(GO:0033549) |
| 0.0 | 0.7 | GO:0015347 | sodium-independent organic anion transmembrane transporter activity(GO:0015347) |
| 0.0 | 0.9 | GO:0016208 | AMP binding(GO:0016208) |
| 0.0 | 0.1 | GO:0030882 | lipid antigen binding(GO:0030882) |
| 0.0 | 0.5 | GO:0051117 | ATPase binding(GO:0051117) |
| 0.0 | 0.8 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.0 | 0.1 | GO:0004908 | interleukin-1 receptor activity(GO:0004908) |
| 0.0 | 0.9 | GO:0019003 | GDP binding(GO:0019003) |
| 0.0 | 1.9 | GO:0019902 | phosphatase binding(GO:0019902) |
| 0.0 | 0.1 | GO:0004361 | glutaryl-CoA dehydrogenase activity(GO:0004361) |
| 0.0 | 0.2 | GO:0031720 | haptoglobin binding(GO:0031720) |
| 0.0 | 1.4 | GO:0016627 | oxidoreductase activity, acting on the CH-CH group of donors(GO:0016627) |
| 0.0 | 2.0 | GO:0005254 | chloride channel activity(GO:0005254) |
| 0.0 | 0.3 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.0 | 0.3 | GO:0031078 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.0 | 0.6 | GO:0046935 | 1-phosphatidylinositol-3-kinase regulator activity(GO:0046935) |
| 0.0 | 0.2 | GO:0048495 | Roundabout binding(GO:0048495) |
| 0.0 | 18.5 | GO:0005509 | calcium ion binding(GO:0005509) |
| 0.0 | 0.2 | GO:0016868 | intramolecular transferase activity, phosphotransferases(GO:0016868) |
| 0.0 | 0.6 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.0 | 0.7 | GO:0071949 | FAD binding(GO:0071949) |
| 0.0 | 0.1 | GO:0034632 | retinol transporter activity(GO:0034632) |
| 0.0 | 0.6 | GO:0005251 | delayed rectifier potassium channel activity(GO:0005251) |
| 0.0 | 1.5 | GO:0036459 | thiol-dependent ubiquitin-specific protease activity(GO:0004843) thiol-dependent ubiquitinyl hydrolase activity(GO:0036459) ubiquitinyl hydrolase activity(GO:0101005) |
| 0.0 | 0.3 | GO:0008331 | high voltage-gated calcium channel activity(GO:0008331) |
| 0.0 | 2.2 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.2 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.0 | 1.3 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.0 | 0.8 | PID HIF2PATHWAY | HIF-2-alpha transcription factor network |
| 0.0 | 2.3 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 0.9 | PID EPHA FWDPATHWAY | EPHA forward signaling |
| 0.0 | 1.0 | PID ECADHERIN NASCENT AJ PATHWAY | E-cadherin signaling in the nascent adherens junction |
| 0.0 | 0.2 | PID IL5 PATHWAY | IL5-mediated signaling events |
| 0.0 | 0.3 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.0 | 1.2 | PID PTP1B PATHWAY | Signaling events mediated by PTP1B |
| 0.0 | 1.2 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.0 | 0.3 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
| 0.0 | 0.2 | PID RET PATHWAY | Signaling events regulated by Ret tyrosine kinase |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 3.2 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.3 | 0.9 | REACTOME ACETYLCHOLINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Acetylcholine Neurotransmitter Release Cycle |
| 0.1 | 1.1 | REACTOME PEPTIDE HORMONE BIOSYNTHESIS | Genes involved in Peptide hormone biosynthesis |
| 0.1 | 0.9 | REACTOME GENERATION OF SECOND MESSENGER MOLECULES | Genes involved in Generation of second messenger molecules |
| 0.1 | 0.5 | REACTOME TRANSPORT OF ORGANIC ANIONS | Genes involved in Transport of organic anions |
| 0.1 | 1.2 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.1 | 1.2 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.1 | 1.6 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.1 | 0.9 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.0 | 1.8 | REACTOME CIRCADIAN CLOCK | Genes involved in Circadian Clock |
| 0.0 | 0.6 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.0 | 0.7 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.0 | 1.3 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.0 | 0.2 | REACTOME GLYCOGEN BREAKDOWN GLYCOGENOLYSIS | Genes involved in Glycogen breakdown (glycogenolysis) |
| 0.0 | 0.2 | REACTOME HYALURONAN METABOLISM | Genes involved in Hyaluronan metabolism |
| 0.0 | 0.2 | REACTOME SIGNALING BY CONSTITUTIVELY ACTIVE EGFR | Genes involved in Signaling by constitutively active EGFR |