Project
PRJNA207719: Tissue specific transcriptome profiling
Navigation
Downloads
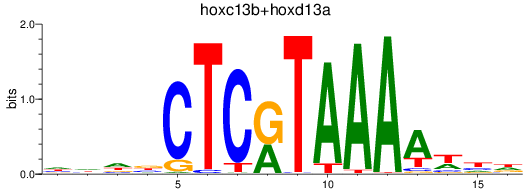
Results for hoxc13b+hoxd13a
Z-value: 1.16
Transcription factors associated with hoxc13b+hoxd13a
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
hoxd13a
|
ENSDARG00000059256 | homeobox D13a |
|
hoxc13b
|
ENSDARG00000113877 | homeobox C13b |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| hoxc13b | dr11_v1_chr11_+_2156430_2156430 | -0.67 | 2.1e-01 | Click! |
Activity profile of hoxc13b+hoxd13a motif
Sorted Z-values of hoxc13b+hoxd13a motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr9_-_33081978 | 1.46 |
ENSDART00000100918
|
zgc:172053
|
zgc:172053 |
| chr9_-_33063083 | 1.30 |
ENSDART00000048550
|
si:ch211-125e6.5
|
si:ch211-125e6.5 |
| chr9_-_33081781 | 1.30 |
ENSDART00000165748
|
zgc:172053
|
zgc:172053 |
| chr9_-_33062891 | 1.12 |
ENSDART00000161182
|
si:ch211-125e6.5
|
si:ch211-125e6.5 |
| chr17_+_2549503 | 1.11 |
ENSDART00000156843
|
si:dkey-248g15.3
|
si:dkey-248g15.3 |
| chr4_-_77551860 | 1.08 |
ENSDART00000188176
|
AL935186.6
|
|
| chr4_+_55758103 | 0.97 |
ENSDART00000185964
|
CT583728.23
|
|
| chr1_+_54650048 | 0.88 |
ENSDART00000141207
|
si:ch211-202h22.7
|
si:ch211-202h22.7 |
| chr22_-_24297510 | 0.88 |
ENSDART00000163297
|
si:ch211-117l17.6
|
si:ch211-117l17.6 |
| chr19_+_20778011 | 0.84 |
ENSDART00000024208
|
nutf2l
|
nuclear transport factor 2, like |
| chr25_-_3867990 | 0.79 |
ENSDART00000075663
|
cracr2b
|
calcium release activated channel regulator 2B |
| chr16_-_12060770 | 0.77 |
ENSDART00000183237
ENSDART00000103948 |
si:ch211-69g19.2
|
si:ch211-69g19.2 |
| chr16_-_12060488 | 0.70 |
ENSDART00000188733
|
si:ch211-69g19.2
|
si:ch211-69g19.2 |
| chr17_-_37395460 | 0.61 |
ENSDART00000148160
ENSDART00000075975 |
crip1
|
cysteine-rich protein 1 |
| chr1_+_45056371 | 0.61 |
ENSDART00000073689
ENSDART00000167309 |
btr01
|
bloodthirsty-related gene family, member 1 |
| chr9_+_23770666 | 0.60 |
ENSDART00000182493
|
si:ch211-219a4.3
|
si:ch211-219a4.3 |
| chr25_+_31277415 | 0.59 |
ENSDART00000036275
|
tnni2a.4
|
troponin I type 2a (skeletal, fast), tandem duplicate 4 |
| chr16_-_16761164 | 0.58 |
ENSDART00000135872
|
si:dkey-27n14.1
|
si:dkey-27n14.1 |
| chr6_+_49771626 | 0.57 |
ENSDART00000134207
|
ctsz
|
cathepsin Z |
| chr1_-_59243542 | 0.55 |
ENSDART00000163021
|
mvb12a
|
multivesicular body subunit 12A |
| chr12_-_30523216 | 0.55 |
ENSDART00000152896
ENSDART00000153191 |
plekhs1
|
pleckstrin homology domain containing, family S member 1 |
| chr9_-_1965727 | 0.55 |
ENSDART00000082354
|
hoxd9a
|
homeobox D9a |
| chr24_+_21621654 | 0.54 |
ENSDART00000002595
|
rpl21
|
ribosomal protein L21 |
| chr18_+_2228737 | 0.54 |
ENSDART00000165301
|
rab27a
|
RAB27A, member RAS oncogene family |
| chr23_+_36083529 | 0.54 |
ENSDART00000053295
ENSDART00000130260 |
hoxc10a
|
homeobox C10a |
| chr6_-_8360918 | 0.54 |
ENSDART00000004716
|
acp5a
|
acid phosphatase 5a, tartrate resistant |
| chr6_+_49771372 | 0.52 |
ENSDART00000063251
|
ctsz
|
cathepsin Z |
| chr16_+_35594670 | 0.51 |
ENSDART00000163275
|
si:ch211-1i11.3
|
si:ch211-1i11.3 |
| chr14_+_10450216 | 0.51 |
ENSDART00000134814
|
cysltr1
|
cysteinyl leukotriene receptor 1 |
| chr5_-_67365006 | 0.51 |
ENSDART00000136116
|
unga
|
uracil DNA glycosylase a |
| chr5_+_57480014 | 0.50 |
ENSDART00000135520
|
si:ch211-202f5.3
|
si:ch211-202f5.3 |
| chr25_+_31276842 | 0.50 |
ENSDART00000187238
|
tnni2a.4
|
troponin I type 2a (skeletal, fast), tandem duplicate 4 |
| chr18_-_43866001 | 0.50 |
ENSDART00000150218
|
treh
|
trehalase (brush-border membrane glycoprotein) |
| chr24_-_40668208 | 0.50 |
ENSDART00000171543
|
smyhc1
|
slow myosin heavy chain 1 |
| chr9_-_9992697 | 0.49 |
ENSDART00000123415
|
ugt1ab
|
UDP glucuronosyltransferase 1 family a, b |
| chr2_+_37295088 | 0.49 |
ENSDART00000056519
|
gpr160
|
G protein-coupled receptor 160 |
| chr13_-_37649595 | 0.48 |
ENSDART00000115354
|
si:dkey-188i13.10
|
si:dkey-188i13.10 |
| chr4_-_30249792 | 0.48 |
ENSDART00000151833
ENSDART00000137564 ENSDART00000168249 |
si:dkey-265e15.2
|
si:dkey-265e15.2 |
| chr16_+_33143503 | 0.48 |
ENSDART00000058471
ENSDART00000179385 |
rhbdl2
|
rhomboid, veinlet-like 2 (Drosophila) |
| chr22_+_24645325 | 0.48 |
ENSDART00000159531
|
lpar3
|
lysophosphatidic acid receptor 3 |
| chr11_+_3254252 | 0.47 |
ENSDART00000123568
|
pmela
|
premelanosome protein a |
| chr25_+_34014523 | 0.47 |
ENSDART00000182856
|
anxa2a
|
annexin A2a |
| chr12_+_13256415 | 0.47 |
ENSDART00000144542
|
atp2a1l
|
ATPase sarcoplasmic/endoplasmic reticulum Ca2+ transporting 1, like |
| chr11_+_36675599 | 0.46 |
ENSDART00000170464
|
si:ch211-11c3.11
|
si:ch211-11c3.11 |
| chr7_-_18617187 | 0.46 |
ENSDART00000172419
|
si:ch211-119e14.1
|
si:ch211-119e14.1 |
| chr20_-_34127415 | 0.46 |
ENSDART00000010028
|
ptgs2b
|
prostaglandin-endoperoxide synthase 2b |
| chr11_-_2478374 | 0.45 |
ENSDART00000173205
|
si:ch73-267c23.10
|
si:ch73-267c23.10 |
| chr22_-_22164338 | 0.44 |
ENSDART00000183840
|
cdc34a
|
cell division cycle 34 homolog (S. cerevisiae) a |
| chr18_-_43866526 | 0.44 |
ENSDART00000111309
|
treh
|
trehalase (brush-border membrane glycoprotein) |
| chr5_-_67365750 | 0.44 |
ENSDART00000062359
|
unga
|
uracil DNA glycosylase a |
| chr20_+_23238833 | 0.44 |
ENSDART00000074167
|
ociad2
|
OCIA domain containing 2 |
| chr16_-_36099492 | 0.43 |
ENSDART00000180905
|
CU499336.2
|
|
| chr4_+_15944245 | 0.43 |
ENSDART00000134594
|
si:dkey-117n7.3
|
si:dkey-117n7.3 |
| chr25_+_29160102 | 0.43 |
ENSDART00000162854
|
pkmb
|
pyruvate kinase M1/2b |
| chr16_+_33144306 | 0.43 |
ENSDART00000101953
|
rhbdl2
|
rhomboid, veinlet-like 2 (Drosophila) |
| chr12_-_34435604 | 0.42 |
ENSDART00000115088
|
birc5a
|
baculoviral IAP repeat containing 5a |
| chr2_+_52065884 | 0.42 |
ENSDART00000146418
|
shda
|
Src homology 2 domain containing transforming protein D, a |
| chr23_-_4933508 | 0.40 |
ENSDART00000137578
ENSDART00000141196 |
ngfa
|
nerve growth factor a (beta polypeptide) |
| chr13_-_34862452 | 0.40 |
ENSDART00000134573
ENSDART00000047552 |
sptlc3
|
serine palmitoyltransferase, long chain base subunit 3 |
| chr7_+_30051880 | 0.40 |
ENSDART00000075609
|
pstpip1b
|
proline-serine-threonine phosphatase interacting protein 1b |
| chr20_+_26683933 | 0.40 |
ENSDART00000139852
ENSDART00000077751 |
foxq1b
|
forkhead box Q1b |
| chr5_-_33280699 | 0.39 |
ENSDART00000183838
|
kyat1
|
kynurenine aminotransferase 1 |
| chr16_+_33144112 | 0.39 |
ENSDART00000183149
|
rhbdl2
|
rhomboid, veinlet-like 2 (Drosophila) |
| chr3_-_29506960 | 0.39 |
ENSDART00000141720
|
cyth4a
|
cytohesin 4a |
| chr17_-_8886735 | 0.39 |
ENSDART00000121997
|
nkl.3
|
NK-lysin tandem duplicate 3 |
| chr11_+_3254524 | 0.38 |
ENSDART00000159459
|
pmela
|
premelanosome protein a |
| chr8_-_12432604 | 0.38 |
ENSDART00000133350
ENSDART00000140699 ENSDART00000101174 |
traf1
|
TNF receptor-associated factor 1 |
| chr3_-_23474416 | 0.37 |
ENSDART00000161672
|
si:dkey-225f5.5
|
si:dkey-225f5.5 |
| chr24_-_35561672 | 0.37 |
ENSDART00000058564
|
mcm4
|
minichromosome maintenance complex component 4 |
| chr22_+_9764731 | 0.37 |
ENSDART00000161441
|
BX664625.4
|
|
| chr4_-_26095755 | 0.37 |
ENSDART00000100611
ENSDART00000191266 |
si:ch211-244b2.3
|
si:ch211-244b2.3 |
| chr21_-_17290941 | 0.37 |
ENSDART00000147993
|
gfi1b
|
growth factor independent 1B transcription repressor |
| chr24_-_40667800 | 0.36 |
ENSDART00000169315
|
smyhc1
|
slow myosin heavy chain 1 |
| chr3_+_24618012 | 0.35 |
ENSDART00000111997
|
zgc:171506
|
zgc:171506 |
| chr16_+_32736588 | 0.35 |
ENSDART00000075191
ENSDART00000168358 |
zgc:172323
|
zgc:172323 |
| chr12_-_17028255 | 0.35 |
ENSDART00000152678
|
ifit10
|
interferon-induced protein with tetratricopeptide repeats 10 |
| chr1_+_56229674 | 0.34 |
ENSDART00000127714
|
c3a.2
|
complement component c3a, duplicate 2 |
| chr5_+_37087583 | 0.34 |
ENSDART00000049900
|
tagln2
|
transgelin 2 |
| chr2_+_48282590 | 0.34 |
ENSDART00000035338
|
lpar5a
|
lysophosphatidic acid receptor 5a |
| chr23_-_1658980 | 0.34 |
ENSDART00000186227
|
CU693481.1
|
|
| chr1_-_25679339 | 0.33 |
ENSDART00000161703
ENSDART00000054230 |
fgg
|
fibrinogen gamma chain |
| chr9_+_23714406 | 0.33 |
ENSDART00000189445
|
gypc
|
glycophorin C (Gerbich blood group) |
| chr18_+_40993196 | 0.33 |
ENSDART00000115111
|
si:dkey-283j8.1
|
si:dkey-283j8.1 |
| chr5_-_69621227 | 0.33 |
ENSDART00000178543
|
aldh2.2
|
aldehyde dehydrogenase 2 family (mitochondrial), tandem duplicate 2 |
| chr6_+_28051978 | 0.33 |
ENSDART00000143218
|
si:ch73-194h10.2
|
si:ch73-194h10.2 |
| chr7_-_48263516 | 0.33 |
ENSDART00000006619
ENSDART00000142370 ENSDART00000148273 ENSDART00000147968 |
rbpms2b
|
RNA binding protein with multiple splicing 2b |
| chr11_-_1400507 | 0.32 |
ENSDART00000173029
ENSDART00000172953 ENSDART00000111140 |
rpl29
|
ribosomal protein L29 |
| chr19_-_27550768 | 0.32 |
ENSDART00000142313
|
si:dkeyp-46h3.8
|
si:dkeyp-46h3.8 |
| chr5_-_41841675 | 0.32 |
ENSDART00000141683
|
si:dkey-65b12.6
|
si:dkey-65b12.6 |
| chr14_-_38872536 | 0.32 |
ENSDART00000184315
ENSDART00000127479 |
gsr
|
glutathione reductase |
| chr19_+_7299865 | 0.32 |
ENSDART00000092408
|
si:ch73-216m24.3
|
si:ch73-216m24.3 |
| chr4_-_74028161 | 0.32 |
ENSDART00000174062
|
LO018205.1
|
|
| chr14_+_21699129 | 0.31 |
ENSDART00000073707
|
stx3a
|
syntaxin 3A |
| chr21_-_21537452 | 0.31 |
ENSDART00000142549
|
or133-7
|
odorant receptor, family H, subfamily 133, member 7 |
| chr1_+_58282449 | 0.31 |
ENSDART00000131475
|
si:dkey-222h21.7
|
si:dkey-222h21.7 |
| chr14_+_26247319 | 0.30 |
ENSDART00000192793
|
CCDC69
|
coiled-coil domain containing 69 |
| chr20_-_38758797 | 0.30 |
ENSDART00000061394
|
trim54
|
tripartite motif containing 54 |
| chr19_+_19756425 | 0.30 |
ENSDART00000167606
|
hoxa3a
|
homeobox A3a |
| chr18_-_35459605 | 0.29 |
ENSDART00000135691
|
itpkcb
|
inositol-trisphosphate 3-kinase Cb |
| chr16_-_21038015 | 0.29 |
ENSDART00000059239
|
snx10b
|
sorting nexin 10b |
| chr22_+_15336752 | 0.29 |
ENSDART00000139070
|
sult3st2
|
sulfotransferase family 3, cytosolic sulfotransferase 2 |
| chr19_-_25081711 | 0.28 |
ENSDART00000058513
|
xkr8.3
|
XK, Kell blood group complex subunit-related family, member 8, tandem duplicate 3 |
| chr7_-_23873173 | 0.28 |
ENSDART00000101408
|
zgc:162171
|
zgc:162171 |
| chr7_+_17861489 | 0.28 |
ENSDART00000173860
|
eml3
|
echinoderm microtubule associated protein like 3 |
| chr22_+_5971404 | 0.28 |
ENSDART00000122153
|
BX322587.1
|
|
| chr13_-_1085961 | 0.28 |
ENSDART00000171872
ENSDART00000166598 |
plek
|
pleckstrin |
| chr7_+_5905091 | 0.28 |
ENSDART00000167099
|
CU459186.3
|
Histone H3.2 |
| chr16_-_4203022 | 0.28 |
ENSDART00000018686
|
rrp15
|
ribosomal RNA processing 15 homolog |
| chr14_+_26439227 | 0.28 |
ENSDART00000054183
|
gpr137
|
G protein-coupled receptor 137 |
| chr21_-_35419486 | 0.27 |
ENSDART00000138529
|
si:dkeyp-23e4.3
|
si:dkeyp-23e4.3 |
| chr2_+_56891858 | 0.27 |
ENSDART00000159912
|
zgc:85843
|
zgc:85843 |
| chr17_+_42274825 | 0.27 |
ENSDART00000020156
|
pax1a
|
paired box 1a |
| chr24_+_5799764 | 0.26 |
ENSDART00000154482
|
si:ch211-157j23.2
|
si:ch211-157j23.2 |
| chr3_+_31058464 | 0.26 |
ENSDART00000153381
|
si:dkey-66i24.7
|
si:dkey-66i24.7 |
| chr7_-_18508815 | 0.26 |
ENSDART00000173539
|
rgs12a
|
regulator of G protein signaling 12a |
| chr7_-_3639171 | 0.26 |
ENSDART00000064280
|
si:ch211-282j17.1
|
si:ch211-282j17.1 |
| chr7_-_4843401 | 0.25 |
ENSDART00000137489
|
si:dkey-83f18.8
|
si:dkey-83f18.8 |
| chr7_+_22637515 | 0.25 |
ENSDART00000158698
|
si:dkey-112a7.5
|
si:dkey-112a7.5 |
| chr3_-_25268751 | 0.25 |
ENSDART00000139423
|
mgat3a
|
mannosyl (beta-1,4-)-glycoprotein beta-1,4-N-acetylglucosaminyltransferase a |
| chr12_+_27117609 | 0.25 |
ENSDART00000076154
|
hoxb8b
|
homeobox B8b |
| chr2_-_37874647 | 0.25 |
ENSDART00000039386
|
zgc:66427
|
zgc:66427 |
| chr15_-_41457898 | 0.25 |
ENSDART00000188847
|
nlrc9
|
NLR family CARD domain containing 9 |
| chr3_-_4501026 | 0.25 |
ENSDART00000163052
|
zgc:162198
|
zgc:162198 |
| chr9_+_11379782 | 0.24 |
ENSDART00000190282
ENSDART00000007308 |
wnt10a
|
wingless-type MMTV integration site family, member 10a |
| chr17_-_8753354 | 0.24 |
ENSDART00000154593
|
tex36
|
testis expressed 36 |
| chr25_-_3230187 | 0.24 |
ENSDART00000160545
|
cib1
|
calcium and integrin binding 1 (calmyrin) |
| chr5_-_67365333 | 0.24 |
ENSDART00000133438
|
unga
|
uracil DNA glycosylase a |
| chr4_-_1930024 | 0.24 |
ENSDART00000134417
|
si:ch73-177h5.2
|
si:ch73-177h5.2 |
| chr9_-_23990416 | 0.24 |
ENSDART00000113176
|
col6a3
|
collagen, type VI, alpha 3 |
| chr17_-_6765877 | 0.23 |
ENSDART00000193102
|
CR352340.1
|
|
| chr21_+_11503212 | 0.23 |
ENSDART00000146701
|
si:dkey-184p9.7
|
si:dkey-184p9.7 |
| chr19_+_20201254 | 0.23 |
ENSDART00000010140
|
igf2bp3
|
insulin-like growth factor 2 mRNA binding protein 3 |
| chr4_+_37406676 | 0.23 |
ENSDART00000130981
|
si:ch73-134f24.1
|
si:ch73-134f24.1 |
| chr19_+_7424347 | 0.23 |
ENSDART00000004622
|
sf3b4
|
splicing factor 3b, subunit 4 |
| chr5_-_69620722 | 0.23 |
ENSDART00000097248
|
aldh2.2
|
aldehyde dehydrogenase 2 family (mitochondrial), tandem duplicate 2 |
| chr12_-_20340543 | 0.23 |
ENSDART00000055623
|
hbbe3
|
hemoglobin beta embryonic-3 |
| chr3_-_43872889 | 0.23 |
ENSDART00000170553
|
mslna
|
mesothelin a |
| chr24_-_25166416 | 0.22 |
ENSDART00000111552
ENSDART00000169495 |
phldb2b
|
pleckstrin homology-like domain, family B, member 2b |
| chr2_+_45532437 | 0.22 |
ENSDART00000137465
|
aknad1
|
AKNA domain containing 1 |
| chr23_+_41912151 | 0.22 |
ENSDART00000191115
|
podn
|
podocan |
| chr6_-_39069244 | 0.22 |
ENSDART00000127365
|
eif4bb
|
eukaryotic translation initiation factor 4Bb |
| chr7_+_24881680 | 0.22 |
ENSDART00000058843
|
krcp
|
kelch repeat-containing protein |
| chr17_-_31695217 | 0.22 |
ENSDART00000104332
ENSDART00000143090 |
lin52
|
lin-52 DREAM MuvB core complex component |
| chr25_-_7887274 | 0.22 |
ENSDART00000157276
|
si:ch73-151m17.5
|
si:ch73-151m17.5 |
| chr21_-_8054977 | 0.22 |
ENSDART00000130508
|
si:dkey-163m14.4
|
si:dkey-163m14.4 |
| chr24_+_1294176 | 0.22 |
ENSDART00000106637
|
si:ch73-134f24.1
|
si:ch73-134f24.1 |
| chr3_+_23045214 | 0.22 |
ENSDART00000157090
ENSDART00000156472 |
b4galnt2.2
|
beta-1,4-N-acetyl-galactosaminyl transferase 2, tandem duplicate 2 |
| chr25_+_21046856 | 0.22 |
ENSDART00000181343
|
chrm2b
|
cholinergic receptor, muscarinic 2b |
| chr9_+_21977383 | 0.21 |
ENSDART00000135032
|
si:dkey-57a22.11
|
si:dkey-57a22.11 |
| chr1_-_2448592 | 0.21 |
ENSDART00000190207
|
ggact.3
|
gamma-glutamylamine cyclotransferase, tandem duplicate 3 |
| chr4_+_77966055 | 0.21 |
ENSDART00000174203
ENSDART00000130100 ENSDART00000080665 ENSDART00000174317 ENSDART00000190123 |
zgc:113921
|
zgc:113921 |
| chr5_+_30520249 | 0.21 |
ENSDART00000013431
|
hmbsa
|
hydroxymethylbilane synthase a |
| chr19_-_2317558 | 0.21 |
ENSDART00000190300
|
sp8a
|
sp8 transcription factor a |
| chr16_+_38337783 | 0.21 |
ENSDART00000135008
|
gabpb2b
|
GA binding protein transcription factor, beta subunit 2b |
| chr6_-_39764995 | 0.21 |
ENSDART00000085277
|
pfkmb
|
phosphofructokinase, muscle b |
| chr15_-_14194208 | 0.21 |
ENSDART00000188237
ENSDART00000183155 ENSDART00000165520 |
pnkp
|
polynucleotide kinase 3'-phosphatase |
| chr7_+_4592867 | 0.20 |
ENSDART00000192614
|
si:dkey-83f18.15
|
si:dkey-83f18.15 |
| chr16_+_50969248 | 0.20 |
ENSDART00000172068
|
si:dkeyp-97a10.2
|
si:dkeyp-97a10.2 |
| chr20_-_154989 | 0.20 |
ENSDART00000064542
|
rpf2
|
ribosome production factor 2 homolog |
| chr8_+_23861461 | 0.20 |
ENSDART00000037109
|
srpk1a
|
SRSF protein kinase 1a |
| chr19_+_7506380 | 0.20 |
ENSDART00000182837
|
prcc
|
papillary renal cell carcinoma (translocation-associated) |
| chr5_-_24008997 | 0.20 |
ENSDART00000066645
|
eif1axa
|
eukaryotic translation initiation factor 1A, X-linked, a |
| chr6_+_46258866 | 0.20 |
ENSDART00000134584
|
zgc:162324
|
zgc:162324 |
| chr5_+_67390115 | 0.20 |
ENSDART00000193255
|
ebf2
|
early B cell factor 2 |
| chr23_-_31932076 | 0.20 |
ENSDART00000138617
|
si:dkey-126g1.7
|
si:dkey-126g1.7 |
| chr2_-_36599044 | 0.19 |
ENSDART00000161829
|
tradv23.3.8
|
T-cell receptor alpha/delta variable 23.3.8 |
| chr18_-_48547564 | 0.19 |
ENSDART00000138607
|
kcnj1a.1
|
potassium inwardly-rectifying channel, subfamily J, member 1a, tandem duplicate 1 |
| chr17_+_20607487 | 0.19 |
ENSDART00000154123
|
si:ch73-288o11.5
|
si:ch73-288o11.5 |
| chr20_-_54259780 | 0.19 |
ENSDART00000172631
|
fkbp3
|
FK506 binding protein 3 |
| chr4_-_61419027 | 0.19 |
ENSDART00000181567
ENSDART00000187397 ENSDART00000180671 ENSDART00000164053 ENSDART00000187395 |
znf1021
|
zinc finger protein 1021 |
| chr23_-_21957383 | 0.19 |
ENSDART00000145780
|
si:dkey-68l7.2
|
si:dkey-68l7.2 |
| chr15_+_17722054 | 0.19 |
ENSDART00000191390
ENSDART00000169550 |
si:ch211-213d14.1
|
si:ch211-213d14.1 |
| chr2_-_6065416 | 0.19 |
ENSDART00000037698
|
uck2b
|
uridine-cytidine kinase 2b |
| chr10_+_6884627 | 0.19 |
ENSDART00000125262
ENSDART00000121729 ENSDART00000105384 |
ddx4
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 4 |
| chr15_-_41332865 | 0.19 |
ENSDART00000170014
|
si:dkey-121n8.7
|
si:dkey-121n8.7 |
| chr22_-_9618867 | 0.19 |
ENSDART00000131246
|
si:ch73-286h23.4
|
si:ch73-286h23.4 |
| chr24_-_40774028 | 0.18 |
ENSDART00000166578
|
CU633479.2
|
|
| chr12_-_7157527 | 0.18 |
ENSDART00000152274
|
si:dkey-187i8.2
|
si:dkey-187i8.2 |
| chr20_-_27311675 | 0.18 |
ENSDART00000026088
ENSDART00000148361 |
asb2a.1
|
ankyrin repeat and SOCS box containing 2a, tandem duplicate 1 |
| chr2_-_36914281 | 0.18 |
ENSDART00000008322
|
si:dkey-193b15.5
|
si:dkey-193b15.5 |
| chr8_-_39654669 | 0.18 |
ENSDART00000145677
|
si:dkey-63d15.12
|
si:dkey-63d15.12 |
| chr10_+_4987766 | 0.18 |
ENSDART00000121959
|
si:ch73-234b20.5
|
si:ch73-234b20.5 |
| chr20_+_18993452 | 0.18 |
ENSDART00000025509
|
tdh
|
L-threonine dehydrogenase |
| chr4_-_37693537 | 0.17 |
ENSDART00000167526
|
si:dkey-3p4.1
|
si:dkey-3p4.1 |
| chr25_+_4787607 | 0.17 |
ENSDART00000159422
|
myo5c
|
myosin VC |
| chr22_+_6512469 | 0.17 |
ENSDART00000137142
|
CR352226.6
|
|
| chr16_+_52343905 | 0.17 |
ENSDART00000131051
|
ifnlr1
|
interferon lambda receptor 1 |
| chr18_+_40993369 | 0.17 |
ENSDART00000141162
|
si:dkey-283j8.1
|
si:dkey-283j8.1 |
| chr22_-_26524596 | 0.17 |
ENSDART00000087623
|
zgc:194330
|
zgc:194330 |
| chr1_+_10129099 | 0.17 |
ENSDART00000187740
|
rbm46
|
RNA binding motif protein 46 |
| chr8_-_34762163 | 0.17 |
ENSDART00000114080
|
setd1bb
|
SET domain containing 1B, b |
| chr2_-_6373829 | 0.17 |
ENSDART00000081633
|
si:dkey-119f1.1
|
si:dkey-119f1.1 |
| chr10_-_31015535 | 0.17 |
ENSDART00000146116
|
panx3
|
pannexin 3 |
| chr15_+_27387555 | 0.16 |
ENSDART00000018603
|
tbx4
|
T-box 4 |
| chr24_+_34069675 | 0.16 |
ENSDART00000143995
|
si:ch211-190p8.2
|
si:ch211-190p8.2 |
| chr1_-_22370660 | 0.16 |
ENSDART00000127506
|
si:ch73-380n15.2
|
si:ch73-380n15.2 |
| chr1_-_681116 | 0.16 |
ENSDART00000165894
|
adamts1
|
ADAM metallopeptidase with thrombospondin type 1 motif, 1 |
| chr15_-_14193926 | 0.16 |
ENSDART00000162707
|
pnkp
|
polynucleotide kinase 3'-phosphatase |
| chr3_-_31019715 | 0.15 |
ENSDART00000156687
ENSDART00000156127 ENSDART00000153945 |
si:dkeyp-71f10.5
|
si:dkeyp-71f10.5 |
| chr14_-_21932403 | 0.15 |
ENSDART00000054420
|
rad9a
|
RAD9 checkpoint clamp component A |
Network of associatons between targets according to the STRING database.
First level regulatory network of hoxc13b+hoxd13a
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.9 | GO:0005991 | trehalose metabolic process(GO:0005991) |
| 0.3 | 0.9 | GO:0097435 | fibril organization(GO:0097435) |
| 0.2 | 1.2 | GO:0097510 | base-excision repair, AP site formation via deaminated base removal(GO:0097510) |
| 0.2 | 0.6 | GO:0097237 | cellular response to antibiotic(GO:0071236) cellular response to toxic substance(GO:0097237) |
| 0.2 | 0.6 | GO:0032510 | endosome to lysosome transport via multivesicular body sorting pathway(GO:0032510) |
| 0.2 | 0.5 | GO:0045648 | positive regulation of erythrocyte differentiation(GO:0045648) |
| 0.1 | 0.9 | GO:0031444 | slow-twitch skeletal muscle fiber contraction(GO:0031444) |
| 0.1 | 0.5 | GO:0019371 | cyclooxygenase pathway(GO:0019371) |
| 0.1 | 0.3 | GO:0051145 | smooth muscle cell differentiation(GO:0051145) negative regulation of muscle cell differentiation(GO:0051148) |
| 0.1 | 0.3 | GO:0098881 | exocytic insertion of neurotransmitter receptor to plasma membrane(GO:0098881) exocytic insertion of neurotransmitter receptor to postsynaptic membrane(GO:0098967) |
| 0.1 | 0.6 | GO:0009954 | proximal/distal pattern formation(GO:0009954) |
| 0.1 | 0.2 | GO:0019276 | UDP-N-acetylgalactosamine metabolic process(GO:0019276) |
| 0.1 | 0.4 | GO:1902975 | cell cycle DNA replication initiation(GO:1902292) nuclear cell cycle DNA replication initiation(GO:1902315) mitotic DNA replication initiation(GO:1902975) |
| 0.0 | 0.3 | GO:0032793 | positive regulation of CREB transcription factor activity(GO:0032793) |
| 0.0 | 0.1 | GO:0072314 | glomerular visceral epithelial cell fate commitment(GO:0072149) glomerular epithelial cell fate commitment(GO:0072314) |
| 0.0 | 0.4 | GO:0007197 | adenylate cyclase-inhibiting G-protein coupled acetylcholine receptor signaling pathway(GO:0007197) |
| 0.0 | 0.3 | GO:0060017 | parathyroid gland development(GO:0060017) |
| 0.0 | 0.4 | GO:0034312 | diol biosynthetic process(GO:0034312) sphingosine biosynthetic process(GO:0046512) |
| 0.0 | 0.2 | GO:0018160 | peptidyl-pyrromethane cofactor linkage(GO:0018160) |
| 0.0 | 0.5 | GO:0000186 | activation of MAPKK activity(GO:0000186) |
| 0.0 | 0.4 | GO:0048672 | positive regulation of collateral sprouting(GO:0048672) |
| 0.0 | 0.2 | GO:0030326 | embryonic limb morphogenesis(GO:0030326) |
| 0.0 | 0.2 | GO:0097010 | eukaryotic translation initiation factor 4F complex assembly(GO:0097010) |
| 0.0 | 0.2 | GO:1900048 | positive regulation of blood coagulation(GO:0030194) positive regulation of coagulation(GO:0050820) positive regulation of hemostasis(GO:1900048) |
| 0.0 | 0.2 | GO:0051754 | meiotic sister chromatid cohesion, centromeric(GO:0051754) |
| 0.0 | 0.4 | GO:0070189 | kynurenine metabolic process(GO:0070189) |
| 0.0 | 0.2 | GO:0070254 | secretion by tissue(GO:0032941) mucus secretion(GO:0070254) |
| 0.0 | 0.1 | GO:0019408 | dolichol biosynthetic process(GO:0019408) |
| 0.0 | 0.1 | GO:0000350 | generation of catalytic spliceosome for second transesterification step(GO:0000350) |
| 0.0 | 0.2 | GO:0046487 | glyoxylate metabolic process(GO:0046487) |
| 0.0 | 0.3 | GO:0070782 | phosphatidylserine exposure on apoptotic cell surface(GO:0070782) |
| 0.0 | 0.5 | GO:0006658 | phosphatidylserine metabolic process(GO:0006658) |
| 0.0 | 0.2 | GO:0032732 | positive regulation of interleukin-1 production(GO:0032732) |
| 0.0 | 0.2 | GO:0044211 | CTP salvage(GO:0044211) |
| 0.0 | 0.3 | GO:0070527 | platelet aggregation(GO:0070527) blood coagulation, fibrin clot formation(GO:0072378) |
| 0.0 | 0.2 | GO:0006566 | threonine metabolic process(GO:0006566) threonine catabolic process(GO:0006567) |
| 0.0 | 0.1 | GO:0003242 | growth involved in heart morphogenesis(GO:0003241) cardiac chamber ballooning(GO:0003242) cardiac muscle tissue growth involved in heart morphogenesis(GO:0003245) |
| 0.0 | 0.1 | GO:0030262 | apoptotic DNA fragmentation(GO:0006309) cellular component disassembly involved in execution phase of apoptosis(GO:0006921) apoptotic nuclear changes(GO:0030262) |
| 0.0 | 0.2 | GO:0061718 | glucose catabolic process(GO:0006007) NADH regeneration(GO:0006735) glycolytic process through fructose-6-phosphate(GO:0061615) glycolytic process through glucose-6-phosphate(GO:0061620) canonical glycolysis(GO:0061621) glucose catabolic process to pyruvate(GO:0061718) |
| 0.0 | 0.4 | GO:0033209 | tumor necrosis factor-mediated signaling pathway(GO:0033209) |
| 0.0 | 0.4 | GO:0045454 | cell redox homeostasis(GO:0045454) |
| 0.0 | 0.7 | GO:0003009 | skeletal muscle contraction(GO:0003009) |
| 0.0 | 0.2 | GO:0071479 | cellular response to ionizing radiation(GO:0071479) |
| 0.0 | 0.2 | GO:0006044 | N-acetylglucosamine metabolic process(GO:0006044) |
| 0.0 | 0.1 | GO:0061549 | sympathetic ganglion development(GO:0061549) |
| 0.0 | 0.3 | GO:0032958 | inositol phosphate biosynthetic process(GO:0032958) |
| 0.0 | 0.1 | GO:0030970 | retrograde protein transport, ER to cytosol(GO:0030970) |
| 0.0 | 0.5 | GO:0045921 | positive regulation of exocytosis(GO:0045921) |
| 0.0 | 0.5 | GO:0000470 | maturation of LSU-rRNA(GO:0000470) |
| 0.0 | 0.1 | GO:0046146 | tetrahydrobiopterin biosynthetic process(GO:0006729) tetrahydrobiopterin metabolic process(GO:0046146) |
| 0.0 | 0.3 | GO:0008406 | gonad development(GO:0008406) |
| 0.0 | 0.2 | GO:0048026 | positive regulation of mRNA splicing, via spliceosome(GO:0048026) |
| 0.0 | 0.9 | GO:0044744 | protein import into nucleus(GO:0006606) protein targeting to nucleus(GO:0044744) single-organism nuclear import(GO:1902593) |
| 0.0 | 0.6 | GO:0099515 | vesicle transport along actin filament(GO:0030050) actin filament-based transport(GO:0099515) |
| 0.0 | 0.1 | GO:0003365 | establishment of cell polarity involved in ameboidal cell migration(GO:0003365) |
| 0.0 | 0.1 | GO:0051045 | negative regulation of membrane protein ectodomain proteolysis(GO:0051045) |
| 0.0 | 0.2 | GO:0015671 | oxygen transport(GO:0015671) |
| 0.0 | 0.1 | GO:0030422 | production of siRNA involved in RNA interference(GO:0030422) |
| 0.0 | 0.3 | GO:0051923 | sulfation(GO:0051923) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 0.1 | 0.9 | GO:0044613 | nuclear pore central transport channel(GO:0044613) |
| 0.1 | 0.2 | GO:0030896 | checkpoint clamp complex(GO:0030896) |
| 0.0 | 0.4 | GO:0017059 | serine C-palmitoyltransferase complex(GO:0017059) endoplasmic reticulum palmitoyltransferase complex(GO:0031211) |
| 0.0 | 1.5 | GO:0048770 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.0 | 0.2 | GO:0000974 | Prp19 complex(GO:0000974) |
| 0.0 | 1.1 | GO:0005861 | troponin complex(GO:0005861) |
| 0.0 | 0.2 | GO:0030915 | Smc5-Smc6 complex(GO:0030915) |
| 0.0 | 0.1 | GO:1904423 | dehydrodolichyl diphosphate synthase complex(GO:1904423) |
| 0.0 | 0.2 | GO:0031838 | haptoglobin-hemoglobin complex(GO:0031838) |
| 0.0 | 0.3 | GO:0008250 | oligosaccharyltransferase complex(GO:0008250) |
| 0.0 | 0.4 | GO:0042555 | MCM complex(GO:0042555) |
| 0.0 | 0.1 | GO:0031310 | integral component of vacuolar membrane(GO:0031166) intrinsic component of vacuolar membrane(GO:0031310) |
| 0.0 | 0.2 | GO:0005945 | 6-phosphofructokinase complex(GO:0005945) |
| 0.0 | 0.5 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.0 | 0.1 | GO:0033186 | CAF-1 complex(GO:0033186) |
| 0.0 | 0.1 | GO:0032783 | ELL-EAF complex(GO:0032783) |
| 0.0 | 0.4 | GO:0033116 | endoplasmic reticulum-Golgi intermediate compartment membrane(GO:0033116) |
| 0.0 | 0.9 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.0 | 0.1 | GO:0030914 | STAGA complex(GO:0030914) |
| 0.0 | 0.1 | GO:0042788 | polysomal ribosome(GO:0042788) |
| 0.0 | 0.2 | GO:0005892 | acetylcholine-gated channel complex(GO:0005892) |
| 0.0 | 1.5 | GO:0016459 | myosin complex(GO:0016459) |
| 0.0 | 0.1 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.0 | 0.2 | GO:0090568 | nuclear transcriptional repressor complex(GO:0090568) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.2 | GO:0043739 | G/U mismatch-specific uracil-DNA glycosylase activity(GO:0043739) |
| 0.3 | 0.9 | GO:0004555 | alpha,alpha-trehalase activity(GO:0004555) trehalase activity(GO:0015927) |
| 0.1 | 0.4 | GO:0047804 | cysteine-S-conjugate beta-lyase activity(GO:0047804) |
| 0.1 | 0.4 | GO:0046403 | polynucleotide 3'-phosphatase activity(GO:0046403) |
| 0.1 | 0.6 | GO:0004362 | glutathione-disulfide reductase activity(GO:0004362) |
| 0.1 | 0.5 | GO:0004666 | prostaglandin-endoperoxide synthase activity(GO:0004666) |
| 0.1 | 0.5 | GO:0004974 | leukotriene receptor activity(GO:0004974) |
| 0.1 | 0.4 | GO:0004743 | pyruvate kinase activity(GO:0004743) |
| 0.1 | 0.4 | GO:0001784 | phosphotyrosine binding(GO:0001784) |
| 0.1 | 0.5 | GO:0003993 | acid phosphatase activity(GO:0003993) |
| 0.1 | 0.4 | GO:0016454 | serine C-palmitoyltransferase activity(GO:0004758) C-palmitoyltransferase activity(GO:0016454) |
| 0.1 | 0.2 | GO:0042166 | neurotransmitter binding(GO:0042165) acetylcholine binding(GO:0042166) |
| 0.0 | 0.4 | GO:0016907 | G-protein coupled acetylcholine receptor activity(GO:0016907) |
| 0.0 | 0.5 | GO:0004859 | phospholipase inhibitor activity(GO:0004859) lipase inhibitor activity(GO:0055102) |
| 0.0 | 0.2 | GO:0008743 | L-threonine 3-dehydrogenase activity(GO:0008743) |
| 0.0 | 0.1 | GO:0004155 | 6,7-dihydropteridine reductase activity(GO:0004155) NADH binding(GO:0070404) |
| 0.0 | 0.2 | GO:0004418 | hydroxymethylbilane synthase activity(GO:0004418) |
| 0.0 | 0.5 | GO:0070915 | lysophosphatidic acid receptor activity(GO:0070915) |
| 0.0 | 0.4 | GO:0043028 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) cysteine-type endopeptidase regulator activity involved in apoptotic process(GO:0043028) |
| 0.0 | 0.2 | GO:0033592 | RNA strand annealing activity(GO:0033592) RNA strand-exchange activity(GO:0034057) |
| 0.0 | 0.4 | GO:0005163 | nerve growth factor receptor binding(GO:0005163) neurotrophin receptor binding(GO:0005165) |
| 0.0 | 0.4 | GO:0004029 | aldehyde dehydrogenase (NAD) activity(GO:0004029) |
| 0.0 | 0.1 | GO:0033842 | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N-acetylgalactosaminyltransferase activity(GO:0033842) |
| 0.0 | 0.2 | GO:0004003 | ATP-dependent DNA helicase activity(GO:0004003) ATP-dependent helicase activity(GO:0008026) purine NTP-dependent helicase activity(GO:0070035) |
| 0.0 | 0.5 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.0 | 0.2 | GO:0015057 | thrombin receptor activity(GO:0015057) |
| 0.0 | 0.1 | GO:0033906 | hyaluronoglucuronidase activity(GO:0033906) |
| 0.0 | 0.2 | GO:0004849 | uridine kinase activity(GO:0004849) |
| 0.0 | 0.1 | GO:0070016 | armadillo repeat domain binding(GO:0070016) |
| 0.0 | 0.5 | GO:0004709 | MAP kinase kinase kinase activity(GO:0004709) |
| 0.0 | 0.1 | GO:0045547 | dehydrodolichyl diphosphate synthase activity(GO:0045547) |
| 0.0 | 0.2 | GO:0031720 | haptoglobin binding(GO:0031720) |
| 0.0 | 0.2 | GO:0070095 | 6-phosphofructokinase activity(GO:0003872) fructose-6-phosphate binding(GO:0070095) |
| 0.0 | 0.1 | GO:0008179 | adenylate cyclase binding(GO:0008179) |
| 0.0 | 0.1 | GO:0016314 | phosphatidylinositol-3,4,5-trisphosphate 3-phosphatase activity(GO:0016314) |
| 0.0 | 0.3 | GO:0000828 | inositol hexakisphosphate kinase activity(GO:0000828) |
| 0.0 | 0.2 | GO:0016833 | oxo-acid-lyase activity(GO:0016833) |
| 0.0 | 0.4 | GO:0003688 | DNA replication origin binding(GO:0003688) |
| 0.0 | 0.1 | GO:0004096 | catalase activity(GO:0004096) |
| 0.0 | 0.0 | GO:0000035 | acyl binding(GO:0000035) |
| 0.0 | 0.2 | GO:0033691 | sialic acid binding(GO:0033691) |
| 0.0 | 0.7 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.0 | 0.1 | GO:0047066 | phospholipid-hydroperoxide glutathione peroxidase activity(GO:0047066) |
| 0.0 | 0.1 | GO:1990050 | phosphatidic acid transporter activity(GO:1990050) |
| 0.0 | 0.6 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.0 | 0.4 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.0 | 0.2 | GO:0070840 | dynein complex binding(GO:0070840) |
| 0.0 | 0.1 | GO:0016864 | protein disulfide isomerase activity(GO:0003756) intramolecular oxidoreductase activity, transposing S-S bonds(GO:0016864) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.0 | 0.5 | PID HIV NEF PATHWAY | HIV-1 Nef: Negative effector of Fas and TNF-alpha |
| 0.0 | 0.5 | PID ENDOTHELIN PATHWAY | Endothelins |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.9 | REACTOME DIGESTION OF DIETARY CARBOHYDRATE | Genes involved in Digestion of dietary carbohydrate |
| 0.1 | 0.5 | REACTOME EICOSANOID LIGAND BINDING RECEPTORS | Genes involved in Eicosanoid ligand-binding receptors |
| 0.0 | 1.1 | REACTOME LYSOSOME VESICLE BIOGENESIS | Genes involved in Lysosome Vesicle Biogenesis |
| 0.0 | 0.1 | REACTOME APOPTOSIS INDUCED DNA FRAGMENTATION | Genes involved in Apoptosis induced DNA fragmentation |
| 0.0 | 0.3 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.0 | 0.4 | REACTOME UNWINDING OF DNA | Genes involved in Unwinding of DNA |
| 0.0 | 0.4 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.0 | 0.4 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.0 | 0.3 | REACTOME G0 AND EARLY G1 | Genes involved in G0 and Early G1 |