Project
PRJNA207719: Tissue specific transcriptome profiling
Navigation
Downloads
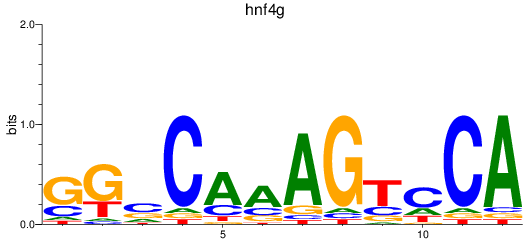
Results for hnf4g
Z-value: 1.44
Transcription factors associated with hnf4g
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
hnf4g
|
ENSDARG00000071565 | hepatocyte nuclear factor 4, gamma |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| hnf4g | dr11_v1_chr24_-_23671709_23671709 | 0.91 | 3.4e-02 | Click! |
Activity profile of hnf4g motif
Sorted Z-values of hnf4g motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr22_-_24818066 | 2.46 |
ENSDART00000143443
|
vtg6
|
vitellogenin 6 |
| chr21_-_27443995 | 1.74 |
ENSDART00000003508
|
bfb
|
complement component bfb |
| chr17_-_2590222 | 1.66 |
ENSDART00000185711
|
CR759892.1
|
|
| chr17_-_2578026 | 1.63 |
ENSDART00000065821
|
zp3.2
|
zona pellucida glycoprotein 3, tandem duplicate 2 |
| chr17_-_2584423 | 1.62 |
ENSDART00000013506
|
zp3.2
|
zona pellucida glycoprotein 3, tandem duplicate 2 |
| chr17_-_2573021 | 1.58 |
ENSDART00000074181
|
zp3.2
|
zona pellucida glycoprotein 3, tandem duplicate 2 |
| chr22_-_24738188 | 1.52 |
ENSDART00000050238
|
vtg1
|
vitellogenin 1 |
| chr22_+_16022211 | 1.43 |
ENSDART00000062618
|
serpinc1
|
serpin peptidase inhibitor, clade C (antithrombin), member 1 |
| chr15_-_16012963 | 1.40 |
ENSDART00000144138
|
hnf1ba
|
HNF1 homeobox Ba |
| chr17_-_2595736 | 1.35 |
ENSDART00000128797
|
zp3.2
|
zona pellucida glycoprotein 3, tandem duplicate 2 |
| chr10_+_19569052 | 1.32 |
ENSDART00000058425
|
CABZ01059627.1
|
|
| chr6_-_607063 | 1.22 |
ENSDART00000189900
|
lgals2b
|
lectin, galactoside-binding, soluble, 2b |
| chr4_-_77114795 | 1.07 |
ENSDART00000144849
|
CU467646.2
|
|
| chr8_+_30699429 | 1.03 |
ENSDART00000005345
|
upb1
|
ureidopropionase, beta |
| chr3_+_16762483 | 0.90 |
ENSDART00000132732
|
tmem86b
|
transmembrane protein 86B |
| chr17_+_20173882 | 0.89 |
ENSDART00000155379
|
si:ch211-248a14.8
|
si:ch211-248a14.8 |
| chr17_+_20174044 | 0.80 |
ENSDART00000156028
|
si:ch211-248a14.8
|
si:ch211-248a14.8 |
| chr18_+_17611627 | 0.80 |
ENSDART00000046891
|
cetp
|
cholesteryl ester transfer protein, plasma |
| chr8_-_49207319 | 0.77 |
ENSDART00000022870
|
fam110a
|
family with sequence similarity 110, member A |
| chr9_+_8380728 | 0.76 |
ENSDART00000133501
|
si:ch1073-75o15.4
|
si:ch1073-75o15.4 |
| chr19_-_25519310 | 0.74 |
ENSDART00000089882
|
C1GALT1 (1 of many)
|
si:dkey-202e17.1 |
| chr20_-_25522911 | 0.74 |
ENSDART00000063058
|
cyp2n13
|
cytochrome P450, family 2, subfamily N, polypeptide 13 |
| chr6_+_27624023 | 0.72 |
ENSDART00000147789
|
slco2a1
|
solute carrier organic anion transporter family, member 2A1 |
| chr8_+_39802506 | 0.70 |
ENSDART00000018862
|
hnf1a
|
HNF1 homeobox a |
| chr13_-_35908275 | 0.54 |
ENSDART00000013961
|
mycla
|
MYCL proto-oncogene, bHLH transcription factor a |
| chr9_-_44953664 | 0.53 |
ENSDART00000188558
ENSDART00000185210 |
vil1
|
villin 1 |
| chr5_-_62317496 | 0.52 |
ENSDART00000180089
|
zgc:85789
|
zgc:85789 |
| chr25_+_29472361 | 0.51 |
ENSDART00000154857
|
il17rel
|
interleukin 17 receptor E-like |
| chr13_-_40726865 | 0.47 |
ENSDART00000099847
|
st3gal7
|
ST3 beta-galactoside alpha-2,3-sialyltransferase 7 |
| chr17_-_30702411 | 0.46 |
ENSDART00000114358
|
zgc:194392
|
zgc:194392 |
| chr5_-_69180587 | 0.46 |
ENSDART00000156681
ENSDART00000160753 |
zgc:171967
|
zgc:171967 |
| chr5_-_69180227 | 0.44 |
ENSDART00000154816
|
zgc:171967
|
zgc:171967 |
| chr8_+_24745041 | 0.43 |
ENSDART00000148872
|
slc16a4
|
solute carrier family 16, member 4 |
| chr7_-_26518086 | 0.43 |
ENSDART00000058913
|
eif4a1a
|
eukaryotic translation initiation factor 4A1A |
| chr12_-_10567188 | 0.43 |
ENSDART00000144283
|
myof
|
myoferlin |
| chr4_-_17725008 | 0.41 |
ENSDART00000016658
|
chpt1
|
choline phosphotransferase 1 |
| chr2_-_10877765 | 0.41 |
ENSDART00000100607
|
cdc7
|
cell division cycle 7 homolog (S. cerevisiae) |
| chr21_-_31290582 | 0.39 |
ENSDART00000065362
|
ca4c
|
carbonic anhydrase IV c |
| chr7_-_24520866 | 0.39 |
ENSDART00000077039
|
faah2b
|
fatty acid amide hydrolase 2b |
| chr23_+_19790962 | 0.38 |
ENSDART00000142228
|
flna
|
filamin A, alpha (actin binding protein 280) |
| chr20_+_29436601 | 0.38 |
ENSDART00000136804
|
fmn1
|
formin 1 |
| chr21_-_43428040 | 0.36 |
ENSDART00000148325
|
stk26
|
serine/threonine protein kinase 26 |
| chr21_-_5879897 | 0.35 |
ENSDART00000184034
|
rpl35
|
ribosomal protein L35 |
| chr20_+_23238833 | 0.33 |
ENSDART00000074167
|
ociad2
|
OCIA domain containing 2 |
| chr13_-_35907768 | 0.33 |
ENSDART00000147522
|
mycla
|
MYCL proto-oncogene, bHLH transcription factor a |
| chr22_+_19407531 | 0.32 |
ENSDART00000141060
|
si:dkey-78l4.2
|
si:dkey-78l4.2 |
| chr4_-_72468168 | 0.32 |
ENSDART00000182995
ENSDART00000174067 |
CR788316.1
|
|
| chr16_+_9495583 | 0.31 |
ENSDART00000150750
ENSDART00000150457 |
pimr208
|
Pim proto-oncogene, serine/threonine kinase, related 208 |
| chr16_+_26846495 | 0.31 |
ENSDART00000078124
|
trim35-29
|
tripartite motif containing 35-29 |
| chr4_+_21741228 | 0.30 |
ENSDART00000112035
ENSDART00000127664 |
myf5
|
myogenic factor 5 |
| chr9_+_1138802 | 0.30 |
ENSDART00000191130
ENSDART00000187308 |
slc15a1a
|
solute carrier family 15 (oligopeptide transporter), member 1a |
| chr10_-_39055620 | 0.30 |
ENSDART00000187437
|
igsf5a
|
immunoglobulin superfamily, member 5a |
| chr25_+_8316953 | 0.29 |
ENSDART00000154598
|
muc2.2
|
mucin 2.2, oligomeric mucus/gel-forming |
| chr15_+_34069746 | 0.29 |
ENSDART00000163513
|
arl4aa
|
ADP-ribosylation factor-like 4aa |
| chr23_+_2740741 | 0.29 |
ENSDART00000134938
|
zgc:114123
|
zgc:114123 |
| chr15_-_31366742 | 0.28 |
ENSDART00000125585
|
or111-3
|
odorant receptor, family D, subfamily 111, member 3 |
| chr1_-_9644630 | 0.27 |
ENSDART00000123725
ENSDART00000161164 |
ugt5b3
|
UDP glucuronosyltransferase 5 family, polypeptide B3 |
| chr16_+_11660839 | 0.27 |
ENSDART00000193911
ENSDART00000143683 |
si:dkey-250k15.10
|
si:dkey-250k15.10 |
| chr1_-_37383741 | 0.27 |
ENSDART00000193155
ENSDART00000191887 ENSDART00000189077 |
scpp1
|
secretory calcium-binding phosphoprotein 1 |
| chr24_+_1294176 | 0.26 |
ENSDART00000106637
|
si:ch73-134f24.1
|
si:ch73-134f24.1 |
| chr4_+_57099307 | 0.26 |
ENSDART00000131654
|
si:ch211-238e22.2
|
si:ch211-238e22.2 |
| chr6_+_59029485 | 0.26 |
ENSDART00000050140
|
CABZ01088367.1
|
|
| chr19_+_18983900 | 0.25 |
ENSDART00000161005
|
cmtm8a
|
CKLF-like MARVEL transmembrane domain containing 8a |
| chr18_+_33725576 | 0.24 |
ENSDART00000146816
|
si:dkey-145c18.5
|
si:dkey-145c18.5 |
| chr7_-_29341233 | 0.24 |
ENSDART00000140938
ENSDART00000147251 |
trpm1a
|
transient receptor potential cation channel, subfamily M, member 1a |
| chr9_+_29994439 | 0.24 |
ENSDART00000012447
|
tmem30c
|
transmembrane protein 30C |
| chr13_+_18520738 | 0.24 |
ENSDART00000113952
|
tlr4al
|
toll-like receptor 4a, like |
| chr6_-_46474483 | 0.23 |
ENSDART00000155761
|
rdh20
|
retinol dehydrogenase 20 |
| chr12_-_44122412 | 0.23 |
ENSDART00000169094
|
si:ch73-329n5.3
|
si:ch73-329n5.3 |
| chr19_+_10536093 | 0.23 |
ENSDART00000138706
|
si:dkey-211g8.1
|
si:dkey-211g8.1 |
| chr6_+_18298444 | 0.22 |
ENSDART00000166018
|
card14
|
caspase recruitment domain family, member 14 |
| chr24_-_36175365 | 0.22 |
ENSDART00000065338
|
pak1ip1
|
PAK1 interacting protein 1 |
| chr7_-_39738460 | 0.21 |
ENSDART00000052201
|
ccdc96
|
coiled-coil domain containing 96 |
| chr21_-_43474012 | 0.21 |
ENSDART00000065104
|
tmem185
|
transmembrane protein 185 |
| chr4_-_170120 | 0.20 |
ENSDART00000171333
|
eps8
|
epidermal growth factor receptor pathway substrate 8 |
| chr23_-_43595956 | 0.20 |
ENSDART00000162186
|
itchb
|
itchy E3 ubiquitin protein ligase b |
| chr9_+_1138323 | 0.20 |
ENSDART00000190352
ENSDART00000190387 |
slc15a1a
|
solute carrier family 15 (oligopeptide transporter), member 1a |
| chr1_-_12064715 | 0.19 |
ENSDART00000143628
ENSDART00000103406 |
pla2g12a
|
phospholipase A2, group XIIA |
| chr4_+_38981587 | 0.17 |
ENSDART00000142713
|
si:dkey-66k12.3
|
si:dkey-66k12.3 |
| chr1_-_37383539 | 0.17 |
ENSDART00000127579
|
scpp1
|
secretory calcium-binding phosphoprotein 1 |
| chr4_+_53145525 | 0.17 |
ENSDART00000166960
|
si:dkey-8o9.2
|
si:dkey-8o9.2 |
| chr2_-_10450784 | 0.17 |
ENSDART00000081196
|
scinlb
|
scinderin like b |
| chr7_+_29115890 | 0.17 |
ENSDART00000052345
|
tradd
|
tnfrsf1a-associated via death domain |
| chr20_-_49657134 | 0.17 |
ENSDART00000151248
|
col12a1b
|
collagen, type XII, alpha 1b |
| chr1_-_59169815 | 0.16 |
ENSDART00000100163
|
wu:fk65c09
|
wu:fk65c09 |
| chr7_-_52417060 | 0.16 |
ENSDART00000148579
|
myzap
|
myocardial zonula adherens protein |
| chr13_-_303137 | 0.16 |
ENSDART00000099131
|
chs1
|
chitin synthase 1 |
| chr2_-_37896965 | 0.16 |
ENSDART00000129852
|
hbl1
|
hexose-binding lectin 1 |
| chr20_-_46694573 | 0.15 |
ENSDART00000182557
|
AL929435.2
|
|
| chr3_+_12764894 | 0.15 |
ENSDART00000166071
|
cyp2k16
|
cytochrome P450, family 2, subfamily K, polypeptide16 |
| chr1_-_57129179 | 0.15 |
ENSDART00000157226
ENSDART00000152469 |
si:ch73-94k4.2
|
si:ch73-94k4.2 |
| chr8_-_20838342 | 0.15 |
ENSDART00000141345
|
si:ch211-133l5.7
|
si:ch211-133l5.7 |
| chr23_+_17437554 | 0.15 |
ENSDART00000184282
|
BX649300.3
|
|
| chr7_-_59514547 | 0.14 |
ENSDART00000168457
|
slx1b
|
SLX1 homolog B, structure-specific endonuclease subunit |
| chr7_+_17229282 | 0.14 |
ENSDART00000097982
|
slc6a5
|
solute carrier family 6 (neurotransmitter transporter), member 5 |
| chr1_-_9485939 | 0.13 |
ENSDART00000157814
|
micall2b
|
mical-like 2b |
| chr11_+_22134540 | 0.13 |
ENSDART00000190502
|
MDFI
|
si:dkey-91m3.1 |
| chr21_-_11856143 | 0.13 |
ENSDART00000151204
|
ube2r2
|
ubiquitin-conjugating enzyme E2R 2 |
| chr13_-_50309969 | 0.13 |
ENSDART00000191597
|
CU928131.1
|
|
| chr7_-_72352043 | 0.12 |
ENSDART00000163536
|
muc5f
|
mucin 5f |
| chr14_-_16810401 | 0.12 |
ENSDART00000158396
ENSDART00000170758 |
tcirg1b
|
T cell immune regulator 1, ATPase H+ transporting V0 subunit a3b |
| chr16_+_11623956 | 0.11 |
ENSDART00000137788
|
cxcr3.1
|
chemokine (C-X-C motif) receptor 3, tandem duplicate 1 |
| chr14_+_9009600 | 0.11 |
ENSDART00000133904
|
si:ch211-274f20.2
|
si:ch211-274f20.2 |
| chr25_+_5288665 | 0.11 |
ENSDART00000169540
|
CABZ01039861.1
|
|
| chr16_+_54875530 | 0.10 |
ENSDART00000149795
|
nr0b2a
|
nuclear receptor subfamily 0, group B, member 2a |
| chr7_+_31398775 | 0.10 |
ENSDART00000124820
|
si:ch211-150j10.4
|
si:ch211-150j10.4 |
| chr7_+_39738505 | 0.10 |
ENSDART00000004365
|
tada2b
|
transcriptional adaptor 2B |
| chr8_+_23390844 | 0.09 |
ENSDART00000184556
|
si:dkey-16n15.6
|
si:dkey-16n15.6 |
| chr20_-_46128590 | 0.09 |
ENSDART00000123744
|
taar1b
|
trace amine associated receptor 1b |
| chr22_-_24992532 | 0.09 |
ENSDART00000102751
|
si:dkey-179j5.5
|
si:dkey-179j5.5 |
| chr1_-_52292235 | 0.09 |
ENSDART00000132638
|
si:dkey-121b10.7
|
si:dkey-121b10.7 |
| chr6_-_10788065 | 0.09 |
ENSDART00000190968
|
wipf1b
|
WAS/WASL interacting protein family, member 1b |
| chr17_-_6076266 | 0.08 |
ENSDART00000171084
|
ephx2
|
epoxide hydrolase 2, cytoplasmic |
| chr10_-_21953643 | 0.08 |
ENSDART00000188921
ENSDART00000193569 |
FO744833.2
|
|
| chr2_-_42958619 | 0.08 |
ENSDART00000144317
|
oc90
|
otoconin 90 |
| chr13_+_26703922 | 0.08 |
ENSDART00000020946
|
fancl
|
Fanconi anemia, complementation group L |
| chr22_-_20376488 | 0.07 |
ENSDART00000140187
|
zbtb7a
|
zinc finger and BTB domain containing 7a |
| chr6_-_9251122 | 0.07 |
ENSDART00000167835
|
BX119909.1
|
|
| chr24_+_20960216 | 0.07 |
ENSDART00000133008
|
si:ch211-161h7.8
|
si:ch211-161h7.8 |
| chr23_-_2448234 | 0.07 |
ENSDART00000082097
|
LO018513.1
|
|
| chr2_-_42958113 | 0.07 |
ENSDART00000139945
|
oc90
|
otoconin 90 |
| chr9_-_1986014 | 0.06 |
ENSDART00000142842
|
hoxd12a
|
homeobox D12a |
| chr7_+_5920665 | 0.06 |
ENSDART00000173208
|
si:dkey-23a13.11
|
si:dkey-23a13.11 |
| chr24_+_7828097 | 0.06 |
ENSDART00000134975
|
zgc:101569
|
zgc:101569 |
| chr13_-_37254777 | 0.06 |
ENSDART00000139734
|
syne2b
|
spectrin repeat containing, nuclear envelope 2b |
| chr14_-_16379843 | 0.06 |
ENSDART00000090711
|
ltc4s
|
leukotriene C4 synthase |
| chr7_+_24520518 | 0.06 |
ENSDART00000173604
|
btr09
|
bloodthirsty-related gene family, member 9 |
| chr2_+_11670270 | 0.05 |
ENSDART00000100524
|
ftr01
|
finTRIM family, member 1 |
| chr17_+_2734331 | 0.05 |
ENSDART00000067542
|
kcnk10b
|
potassium channel, subfamily K, member 10b |
| chr24_-_32150276 | 0.04 |
ENSDART00000166212
|
cubn
|
cubilin (intrinsic factor-cobalamin receptor) |
| chr3_-_8765165 | 0.04 |
ENSDART00000191131
|
CABZ01058333.1
|
|
| chr6_-_56103718 | 0.04 |
ENSDART00000191002
|
gcnt7
|
glucosaminyl (N-acetyl) transferase family member 7 |
| chr12_+_22607761 | 0.04 |
ENSDART00000153112
|
si:dkey-219e21.2
|
si:dkey-219e21.2 |
| chr15_-_14884332 | 0.03 |
ENSDART00000165237
|
si:ch211-24o8.4
|
si:ch211-24o8.4 |
| chr22_+_2844865 | 0.03 |
ENSDART00000139123
|
si:dkey-20i20.4
|
si:dkey-20i20.4 |
| chr5_+_37069605 | 0.03 |
ENSDART00000161977
|
si:dkeyp-110c7.4
|
si:dkeyp-110c7.4 |
| chr11_-_5563498 | 0.02 |
ENSDART00000160835
|
pex11g
|
peroxisomal biogenesis factor 11 gamma |
| chr25_+_22216723 | 0.02 |
ENSDART00000051291
|
hexa
|
hexosaminidase A (alpha polypeptide) |
| chr15_-_2632891 | 0.02 |
ENSDART00000081840
|
cldnj
|
claudin j |
| chr8_+_47188154 | 0.00 |
ENSDART00000137319
|
si:dkeyp-100a1.6
|
si:dkeyp-100a1.6 |
Network of associatons between targets according to the STRING database.
First level regulatory network of hnf4g
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.4 | GO:2000257 | regulation of protein activation cascade(GO:2000257) |
| 0.4 | 6.2 | GO:0035803 | binding of sperm to zona pellucida(GO:0007339) sperm-egg recognition(GO:0035036) egg coat formation(GO:0035803) regulation of acrosome reaction(GO:0060046) positive regulation of acrosome reaction(GO:2000344) |
| 0.3 | 0.8 | GO:0034375 | high-density lipoprotein particle remodeling(GO:0034375) |
| 0.2 | 1.4 | GO:0021572 | rhombomere 5 development(GO:0021571) rhombomere 6 development(GO:0021572) |
| 0.2 | 1.0 | GO:0019483 | beta-alanine metabolic process(GO:0019482) beta-alanine biosynthetic process(GO:0019483) |
| 0.2 | 0.3 | GO:0002074 | extraocular skeletal muscle development(GO:0002074) |
| 0.1 | 0.5 | GO:2000392 | lamellipodium morphogenesis(GO:0072673) regulation of lamellipodium morphogenesis(GO:2000392) |
| 0.1 | 1.1 | GO:0071715 | icosanoid transport(GO:0071715) fatty acid derivative transport(GO:1901571) |
| 0.1 | 4.0 | GO:0032355 | response to estradiol(GO:0032355) |
| 0.1 | 0.5 | GO:0006857 | oligopeptide transport(GO:0006857) |
| 0.0 | 0.3 | GO:0033206 | meiotic cytokinesis(GO:0033206) polar body extrusion after meiotic divisions(GO:0040038) |
| 0.0 | 0.2 | GO:1901073 | chitin biosynthetic process(GO:0006031) glucosamine-containing compound biosynthetic process(GO:1901073) |
| 0.0 | 0.7 | GO:0030073 | insulin secretion(GO:0030073) |
| 0.0 | 0.4 | GO:0000727 | double-strand break repair via break-induced replication(GO:0000727) |
| 0.0 | 0.3 | GO:0000463 | maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000463) |
| 0.0 | 1.7 | GO:0006956 | complement activation(GO:0006956) |
| 0.0 | 0.9 | GO:0042738 | drug metabolic process(GO:0017144) drug catabolic process(GO:0042737) exogenous drug catabolic process(GO:0042738) |
| 0.0 | 0.1 | GO:0043092 | amino acid import(GO:0043090) L-amino acid import(GO:0043092) |
| 0.0 | 0.4 | GO:0007520 | myoblast fusion(GO:0007520) |
| 0.0 | 0.1 | GO:0006691 | leukotriene metabolic process(GO:0006691) leukotriene biosynthetic process(GO:0019370) |
| 0.0 | 0.2 | GO:0002430 | complement receptor mediated signaling pathway(GO:0002430) |
| 0.0 | 0.1 | GO:0035066 | positive regulation of histone acetylation(GO:0035066) |
| 0.0 | 0.4 | GO:0002183 | cytoplasmic translational initiation(GO:0002183) |
| 0.0 | 0.2 | GO:0050829 | defense response to Gram-negative bacterium(GO:0050829) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0032433 | filopodium tip(GO:0032433) |
| 0.1 | 0.8 | GO:0034364 | high-density lipoprotein particle(GO:0034364) |
| 0.0 | 0.1 | GO:0033557 | Slx1-Slx4 complex(GO:0033557) |
| 0.0 | 0.2 | GO:0030428 | cell septum(GO:0030428) |
| 0.0 | 6.5 | GO:0031012 | extracellular matrix(GO:0031012) |
| 0.0 | 0.1 | GO:0000220 | vacuolar proton-transporting V-type ATPase, V0 domain(GO:0000220) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 4.0 | GO:0045735 | nutrient reservoir activity(GO:0045735) |
| 0.6 | 6.2 | GO:0035804 | structural constituent of egg coat(GO:0035804) |
| 0.2 | 0.5 | GO:0047291 | lactosylceramide alpha-2,3-sialyltransferase activity(GO:0047291) |
| 0.1 | 1.2 | GO:0016936 | galactoside binding(GO:0016936) |
| 0.1 | 0.9 | GO:0016803 | ether hydrolase activity(GO:0016803) |
| 0.1 | 0.4 | GO:0004142 | diacylglycerol cholinephosphotransferase activity(GO:0004142) |
| 0.1 | 0.5 | GO:0042936 | dipeptide transporter activity(GO:0042936) dipeptide transmembrane transporter activity(GO:0071916) |
| 0.1 | 0.2 | GO:0001875 | lipopolysaccharide receptor activity(GO:0001875) |
| 0.0 | 0.7 | GO:0015245 | fatty acid transporter activity(GO:0015245) |
| 0.0 | 0.5 | GO:0030368 | interleukin-17 receptor activity(GO:0030368) |
| 0.0 | 0.8 | GO:0031210 | phosphatidylcholine binding(GO:0031210) |
| 0.0 | 0.1 | GO:0016494 | C-X-C chemokine receptor activity(GO:0016494) |
| 0.0 | 0.2 | GO:0005068 | transmembrane receptor protein tyrosine kinase adaptor activity(GO:0005068) |
| 0.0 | 0.1 | GO:0008821 | crossover junction endodeoxyribonuclease activity(GO:0008821) |
| 0.0 | 0.2 | GO:0050700 | CARD domain binding(GO:0050700) |
| 0.0 | 0.2 | GO:0004100 | chitin synthase activity(GO:0004100) |
| 0.0 | 0.3 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.0 | 1.4 | GO:0016811 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amides(GO:0016811) |
| 0.0 | 0.1 | GO:0004464 | leukotriene-C4 synthase activity(GO:0004464) |
| 0.0 | 0.4 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.0 | 0.3 | GO:0004623 | phospholipase A2 activity(GO:0004623) |
| 0.0 | 0.9 | GO:0016712 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced flavin or flavoprotein as one donor, and incorporation of one atom of oxygen(GO:0016712) |
| 0.0 | 0.7 | GO:0005546 | phosphatidylinositol-4,5-bisphosphate binding(GO:0005546) |
| 0.0 | 1.4 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 0.2 | GO:0004875 | complement receptor activity(GO:0004875) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.4 | PID GLYPICAN 1PATHWAY | Glypican 1 network |
| 0.0 | 0.4 | PID ECADHERIN KERATINOCYTE PATHWAY | E-cadherin signaling in keratinocytes |
| 0.0 | 0.7 | PID PS1 PATHWAY | Presenilin action in Notch and Wnt signaling |
| 0.0 | 0.4 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.0 | 0.4 | PID VEGFR1 2 PATHWAY | Signaling events mediated by VEGFR1 and VEGFR2 |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.0 | REACTOME PYRIMIDINE CATABOLISM | Genes involved in Pyrimidine catabolism |
| 0.1 | 1.4 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.1 | 0.7 | REACTOME TRANSPORT OF ORGANIC ANIONS | Genes involved in Transport of organic anions |
| 0.1 | 0.8 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.0 | 0.2 | REACTOME ACYL CHAIN REMODELLING OF PI | Genes involved in Acyl chain remodelling of PI |
| 0.0 | 0.7 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.0 | 0.4 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |
| 0.0 | 0.2 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.0 | 0.4 | REACTOME ACTIVATION OF THE PRE REPLICATIVE COMPLEX | Genes involved in Activation of the pre-replicative complex |