Project
PRJNA207719: Tissue specific transcriptome profiling
Navigation
Downloads
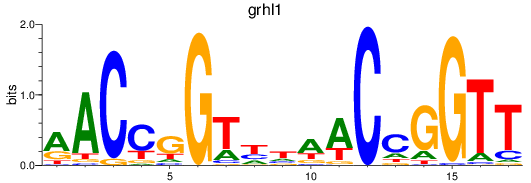
Results for grhl1
Z-value: 4.15
Transcription factors associated with grhl1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
grhl1
|
ENSDARG00000061391 | grainyhead-like transcription factor 1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| grhl1 | dr11_v1_chr17_-_32426392_32426413 | 0.81 | 9.5e-02 | Click! |
Activity profile of grhl1 motif
Sorted Z-values of grhl1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr5_+_58665648 | 6.40 |
ENSDART00000167481
|
zgc:194948
|
zgc:194948 |
| chr15_+_42560354 | 5.37 |
ENSDART00000059484
|
CLDN8 (1 of many)
|
zgc:110333 |
| chr9_-_30264415 | 4.50 |
ENSDART00000060150
|
mid1ip1a
|
MID1 interacting protein 1a |
| chr24_+_38671054 | 4.50 |
ENSDART00000154214
|
si:ch73-70c5.1
|
si:ch73-70c5.1 |
| chr4_+_76722754 | 4.39 |
ENSDART00000153867
|
ms4a17a.3
|
membrane-spanning 4-domains, subfamily A, member 17A.3 |
| chr15_-_12360409 | 4.30 |
ENSDART00000164596
|
tmprss13a
|
transmembrane protease, serine 13a |
| chr1_-_48933 | 4.17 |
ENSDART00000171162
|
ildr1a
|
immunoglobulin-like domain containing receptor 1a |
| chr10_-_22127942 | 4.03 |
ENSDART00000133374
|
ponzr2
|
plac8 onzin related protein 2 |
| chr16_+_35905031 | 3.97 |
ENSDART00000162411
|
sh3d21
|
SH3 domain containing 21 |
| chr17_+_28102487 | 3.87 |
ENSDART00000131638
ENSDART00000076265 |
zgc:91908
|
zgc:91908 |
| chr16_+_23431189 | 3.83 |
ENSDART00000004679
|
icn
|
ictacalcin |
| chr5_-_6567464 | 3.81 |
ENSDART00000184985
|
tnks1bp1
|
tankyrase 1 binding protein 1 |
| chr21_+_11244068 | 3.71 |
ENSDART00000163432
|
arid6
|
AT-rich interaction domain 6 |
| chr14_-_763744 | 3.53 |
ENSDART00000165856
|
trim35-27
|
tripartite motif containing 35-27 |
| chr22_+_19552987 | 3.51 |
ENSDART00000105315
|
hsd11b1la
|
hydroxysteroid (11-beta) dehydrogenase 1-like a |
| chr12_-_28794957 | 3.48 |
ENSDART00000020667
|
osbpl7
|
oxysterol binding protein-like 7 |
| chr4_-_9780931 | 3.44 |
ENSDART00000134280
ENSDART00000150664 ENSDART00000150304 ENSDART00000080744 |
svopl
|
SVOP-like |
| chr7_+_19835569 | 3.39 |
ENSDART00000149812
|
ovol1a
|
ovo-like zinc finger 1a |
| chr21_+_25625026 | 3.34 |
ENSDART00000134678
|
ovol1b
|
ovo-like zinc finger 1b |
| chr13_+_18533005 | 3.32 |
ENSDART00000136024
|
ftr14l
|
finTRIM family, member 14-like |
| chr1_-_52498146 | 3.16 |
ENSDART00000122217
|
acy3.2
|
aspartoacylase (aminocyclase) 3, tandem duplicate 2 |
| chr16_+_17715243 | 3.04 |
ENSDART00000149437
ENSDART00000149596 |
si:dkey-87o1.2
|
si:dkey-87o1.2 |
| chr8_-_24252933 | 2.92 |
ENSDART00000057624
|
zgc:110353
|
zgc:110353 |
| chr14_+_146857 | 2.84 |
ENSDART00000122521
|
CABZ01088229.1
|
|
| chr7_-_24193051 | 2.76 |
ENSDART00000169737
ENSDART00000171085 |
si:ch211-216p19.6
|
si:ch211-216p19.6 |
| chr17_+_22102791 | 2.74 |
ENSDART00000047772
|
mal
|
mal, T cell differentiation protein |
| chr15_+_22014029 | 2.73 |
ENSDART00000079504
|
ankk1
|
ankyrin repeat and kinase domain containing 1 |
| chr16_-_31976269 | 2.73 |
ENSDART00000139664
|
styk1
|
serine/threonine/tyrosine kinase 1 |
| chr15_+_41919484 | 2.59 |
ENSDART00000099821
ENSDART00000146246 |
nlrp16
|
NACHT, LRR and PYD domains-containing protein 16 |
| chr21_-_3700334 | 2.59 |
ENSDART00000137844
|
atp8b1
|
ATPase phospholipid transporting 8B1 |
| chr1_-_52497834 | 2.51 |
ENSDART00000136469
ENSDART00000004233 |
acy3.2
|
aspartoacylase (aminocyclase) 3, tandem duplicate 2 |
| chr6_+_23057311 | 2.50 |
ENSDART00000026448
|
evpla
|
envoplakin a |
| chr3_-_29977495 | 2.45 |
ENSDART00000077111
|
hsd17b14
|
hydroxysteroid (17-beta) dehydrogenase 14 |
| chr22_+_25672155 | 2.40 |
ENSDART00000087769
|
si:ch211-250e5.2
|
si:ch211-250e5.2 |
| chr2_-_47620806 | 2.38 |
ENSDART00000038228
|
ap1s3b
|
adaptor-related protein complex 1, sigma 3 subunit, b |
| chr9_-_40014339 | 2.38 |
ENSDART00000166918
|
IKZF2
|
si:zfos-1425h8.1 |
| chr7_-_56606752 | 2.36 |
ENSDART00000138714
|
sult5a1
|
sulfotransferase family 5A, member 1 |
| chr22_+_22021936 | 2.27 |
ENSDART00000149586
|
gna15.1
|
guanine nucleotide binding protein (G protein), alpha 15 (Gq class), tandem duplicate 1 |
| chr19_-_32500373 | 2.27 |
ENSDART00000052104
|
fuca1.1
|
alpha-L-fucosidase 1, tandem duplicate 1 |
| chr3_-_16537441 | 2.26 |
ENSDART00000080749
ENSDART00000133824 |
eps8l1
|
eps8-like1 |
| chr22_-_14361022 | 2.23 |
ENSDART00000140224
|
si:dkeyp-122a9.2
|
si:dkeyp-122a9.2 |
| chr2_+_5521671 | 2.14 |
ENSDART00000099647
ENSDART00000138443 |
crfb16
|
cytokine receptor family member B16 |
| chr5_-_37900350 | 2.14 |
ENSDART00000084839
ENSDART00000084841 ENSDART00000133437 |
tmprss13b
|
transmembrane protease, serine 13b |
| chr3_-_18756076 | 2.08 |
ENSDART00000055766
|
zgc:113333
|
zgc:113333 |
| chr8_+_3405612 | 2.04 |
ENSDART00000163437
|
zgc:112433
|
zgc:112433 |
| chr23_-_22650668 | 2.03 |
ENSDART00000132733
|
ca6
|
carbonic anhydrase VI |
| chr4_-_12725513 | 1.95 |
ENSDART00000132286
|
mgst1.2
|
microsomal glutathione S-transferase 1.2 |
| chr7_+_54259657 | 1.91 |
ENSDART00000170174
|
pacsin3
|
protein kinase C and casein kinase substrate in neurons 3 |
| chr10_-_36793412 | 1.90 |
ENSDART00000185966
|
dhrs13a.2
|
dehydrogenase/reductase (SDR family) member 13a, tandem duplicate 2 |
| chr14_+_45702521 | 1.74 |
ENSDART00000112470
|
ccdc88b
|
coiled-coil domain containing 88B |
| chr23_-_22650992 | 1.74 |
ENSDART00000079007
|
ca6
|
carbonic anhydrase VI |
| chr1_-_47161996 | 1.68 |
ENSDART00000053153
|
mhc1zba
|
major histocompatibility complex class I ZBA |
| chr2_-_58075414 | 1.63 |
ENSDART00000161920
|
nectin4
|
nectin cell adhesion molecule 4 |
| chr13_+_42544009 | 1.60 |
ENSDART00000145409
|
si:dkey-221j11.3
|
si:dkey-221j11.3 |
| chr6_+_30668098 | 1.59 |
ENSDART00000112294
|
ttc22
|
tetratricopeptide repeat domain 22 |
| chr15_+_509126 | 1.56 |
ENSDART00000102274
|
ftr86
|
finTRIM family, member 86 |
| chr3_-_18755651 | 1.55 |
ENSDART00000145277
|
zgc:113333
|
zgc:113333 |
| chr22_-_22337382 | 1.55 |
ENSDART00000144684
|
si:ch211-129c21.1
|
si:ch211-129c21.1 |
| chr12_+_4036409 | 1.44 |
ENSDART00000106650
|
zgc:123217
|
zgc:123217 |
| chr5_-_57686487 | 1.43 |
ENSDART00000074264
|
crfb12
|
cytokine receptor family member B12 |
| chr10_+_4924388 | 1.38 |
ENSDART00000108595
|
slc46a2
|
solute carrier family 46 member 2 |
| chr5_-_29570141 | 1.36 |
ENSDART00000043259
|
entpd2a.2
|
ectonucleoside triphosphate diphosphohydrolase 2a, tandem duplicate 2 |
| chr12_+_30360579 | 1.34 |
ENSDART00000152900
|
si:ch211-225b10.3
|
si:ch211-225b10.3 |
| chr8_-_36399884 | 1.29 |
ENSDART00000108538
|
si:zfos-2070c2.3
|
si:zfos-2070c2.3 |
| chr3_+_37196258 | 1.24 |
ENSDART00000187944
ENSDART00000185896 |
fmnl1a
|
formin-like 1a |
| chr25_-_24202576 | 1.23 |
ENSDART00000048507
|
uevld
|
UEV and lactate/malate dehyrogenase domains |
| chr15_-_5901514 | 1.21 |
ENSDART00000155252
|
si:ch73-281n10.2
|
si:ch73-281n10.2 |
| chr14_-_45702721 | 1.20 |
ENSDART00000165727
|
si:dkey-226n24.1
|
si:dkey-226n24.1 |
| chr3_+_34140507 | 1.18 |
ENSDART00000131802
|
si:dkey-204f11.64
|
si:dkey-204f11.64 |
| chr1_+_49352900 | 1.08 |
ENSDART00000008468
|
msx1b
|
muscle segment homeobox 1b |
| chr15_+_41901970 | 1.06 |
ENSDART00000152724
|
si:ch211-191a16.5
|
si:ch211-191a16.5 |
| chr3_-_32831429 | 1.03 |
ENSDART00000184932
|
zgc:153733
|
zgc:153733 |
| chr25_+_7435291 | 0.89 |
ENSDART00000172567
ENSDART00000163017 |
prc1a
|
protein regulator of cytokinesis 1a |
| chr3_-_32831971 | 0.88 |
ENSDART00000075270
|
zgc:153733
|
zgc:153733 |
| chr22_-_13649588 | 0.81 |
ENSDART00000131877
|
si:ch211-279g13.1
|
si:ch211-279g13.1 |
| chr11_-_45140227 | 0.73 |
ENSDART00000184062
|
cant1b
|
calcium activated nucleotidase 1b |
| chr24_+_42074143 | 0.71 |
ENSDART00000170514
|
top1mt
|
DNA topoisomerase I mitochondrial |
| chr8_-_13029297 | 0.65 |
ENSDART00000144305
|
dennd2da
|
DENN/MADD domain containing 2Da |
| chr4_-_26413391 | 0.64 |
ENSDART00000145955
|
b4galnt3a
|
beta-1,4-N-acetyl-galactosaminyl transferase 3a |
| chr12_+_31673588 | 0.64 |
ENSDART00000152971
|
dnmbp
|
dynamin binding protein |
| chr1_-_9858508 | 0.63 |
ENSDART00000147904
|
mad1l1
|
mitotic arrest deficient 1 like 1 |
| chr1_-_31105376 | 0.59 |
ENSDART00000132466
|
ppp1r9alb
|
protein phosphatase 1 regulatory subunit 9A-like B |
| chr19_-_10432134 | 0.57 |
ENSDART00000081440
|
il11b
|
interleukin 11b |
| chr19_-_41472228 | 0.57 |
ENSDART00000113388
|
dlx5a
|
distal-less homeobox 5a |
| chr15_+_41901710 | 0.54 |
ENSDART00000159874
|
si:ch211-191a16.5
|
si:ch211-191a16.5 |
| chr16_+_53526135 | 0.52 |
ENSDART00000083558
|
smpd5
|
sphingomyelin phosphodiesterase 5 |
| chr4_+_4232562 | 0.48 |
ENSDART00000177529
|
smkr1
|
small lysine rich protein 1 |
| chr21_-_13668358 | 0.45 |
ENSDART00000180323
|
pnpla7a
|
patatin-like phospholipase domain containing 7a |
| chr23_-_19225709 | 0.43 |
ENSDART00000080099
|
oard1
|
O-acyl-ADP-ribose deacylase 1 |
| chr7_+_22657566 | 0.35 |
ENSDART00000141048
|
ponzr5
|
plac8 onzin related protein 5 |
| chr14_+_46216703 | 0.19 |
ENSDART00000136045
ENSDART00000142317 |
mgst2
|
microsomal glutathione S-transferase 2 |
| chr9_-_14108896 | 0.18 |
ENSDART00000135209
|
prkag3b
|
protein kinase, AMP-activated, gamma 3b non-catalytic subunit |
| chr24_-_8412526 | 0.18 |
ENSDART00000150055
|
slc35b3
|
solute carrier family 35 (adenosine 3'-phospho 5'-phosphosulfate transporter), member B3 |
| chr21_+_5801105 | 0.16 |
ENSDART00000151225
ENSDART00000184487 |
ccng2
|
cyclin G2 |
| chr20_-_42378865 | 0.16 |
ENSDART00000139912
ENSDART00000015801 |
dcbld1
|
discoidin, CUB and LCCL domain containing 1 |
| chr11_+_8152872 | 0.12 |
ENSDART00000091638
ENSDART00000138057 ENSDART00000166379 |
samd13
|
sterile alpha motif domain containing 13 |
| chr10_+_19525839 | 0.10 |
ENSDART00000162912
ENSDART00000158512 |
vsig8a
|
V-set and immunoglobulin domain containing 8a |
| chr13_-_30161684 | 0.09 |
ENSDART00000040409
|
ppa1b
|
pyrophosphatase (inorganic) 1b |
| chr14_+_4276394 | 0.02 |
ENSDART00000038301
|
gnpda2
|
glucosamine-6-phosphate deaminase 2 |
| chr8_-_13541514 | 0.02 |
ENSDART00000063834
|
zgc:86586
|
zgc:86586 |
Network of associatons between targets according to the STRING database.
First level regulatory network of grhl1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 3.4 | GO:0070285 | pigment cell development(GO:0070285) |
| 0.4 | 2.3 | GO:0006004 | fucose metabolic process(GO:0006004) |
| 0.3 | 1.5 | GO:0010693 | regulation of alkaline phosphatase activity(GO:0010692) negative regulation of alkaline phosphatase activity(GO:0010693) |
| 0.2 | 1.4 | GO:0071405 | response to methanol(GO:0033986) cellular response to methanol(GO:0071405) |
| 0.2 | 1.9 | GO:0014028 | notochord formation(GO:0014028) |
| 0.1 | 1.9 | GO:0001881 | receptor recycling(GO:0001881) |
| 0.1 | 2.3 | GO:0060158 | phospholipase C-activating dopamine receptor signaling pathway(GO:0060158) |
| 0.1 | 1.9 | GO:0042574 | retinal metabolic process(GO:0042574) |
| 0.1 | 2.4 | GO:0051923 | sulfation(GO:0051923) |
| 0.1 | 0.9 | GO:0051256 | mitotic spindle elongation(GO:0000022) spindle elongation(GO:0051231) mitotic spindle midzone assembly(GO:0051256) |
| 0.1 | 4.5 | GO:0046890 | regulation of lipid biosynthetic process(GO:0046890) |
| 0.1 | 2.6 | GO:0045332 | phospholipid translocation(GO:0045332) |
| 0.1 | 3.8 | GO:0036269 | swimming behavior(GO:0036269) |
| 0.1 | 0.6 | GO:0021553 | olfactory nerve development(GO:0021553) olfactory nerve morphogenesis(GO:0021627) olfactory nerve formation(GO:0021628) |
| 0.1 | 0.6 | GO:0051315 | attachment of mitotic spindle microtubules to kinetochore(GO:0051315) |
| 0.1 | 2.5 | GO:0045103 | intermediate filament-based process(GO:0045103) intermediate filament cytoskeleton organization(GO:0045104) |
| 0.1 | 0.5 | GO:0006685 | sphingomyelin catabolic process(GO:0006685) |
| 0.0 | 0.7 | GO:0006265 | DNA topological change(GO:0006265) |
| 0.0 | 0.2 | GO:0019370 | leukotriene metabolic process(GO:0006691) leukotriene biosynthetic process(GO:0019370) |
| 0.0 | 0.9 | GO:0043049 | otic placode formation(GO:0043049) |
| 0.0 | 1.3 | GO:0006890 | retrograde vesicle-mediated transport, Golgi to ER(GO:0006890) |
| 0.0 | 3.3 | GO:0009913 | epidermal cell differentiation(GO:0009913) |
| 0.0 | 1.7 | GO:0031122 | cytoplasmic microtubule organization(GO:0031122) |
| 0.0 | 1.2 | GO:0030866 | cortical actin cytoskeleton organization(GO:0030866) |
| 0.0 | 1.4 | GO:0007596 | blood coagulation(GO:0007596) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 3.5 | GO:0097038 | perinuclear endoplasmic reticulum(GO:0097038) |
| 0.1 | 1.2 | GO:0031680 | G-protein beta/gamma-subunit complex(GO:0031680) |
| 0.1 | 0.9 | GO:1990023 | mitotic spindle midzone(GO:1990023) |
| 0.1 | 1.3 | GO:0033116 | endoplasmic reticulum-Golgi intermediate compartment membrane(GO:0033116) |
| 0.0 | 2.3 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.0 | 5.7 | GO:0016324 | apical plasma membrane(GO:0016324) |
| 0.0 | 4.5 | GO:0005802 | trans-Golgi network(GO:0005802) |
| 0.0 | 2.4 | GO:0030117 | membrane coat(GO:0030117) coated membrane(GO:0048475) |
| 0.0 | 2.5 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.0 | 0.2 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.0 | 1.3 | GO:0005795 | Golgi stack(GO:0005795) |
| 0.0 | 0.5 | GO:0031970 | mitochondrial intermembrane space(GO:0005758) organelle envelope lumen(GO:0031970) |
| 0.0 | 5.4 | GO:0009986 | cell surface(GO:0009986) |
| 0.0 | 0.6 | GO:0072686 | mitotic spindle(GO:0072686) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.8 | 2.3 | GO:0004560 | alpha-L-fucosidase activity(GO:0004560) fucosidase activity(GO:0015928) |
| 0.5 | 5.7 | GO:0004046 | aminoacylase activity(GO:0004046) |
| 0.4 | 2.5 | GO:0019215 | intermediate filament binding(GO:0019215) |
| 0.4 | 3.6 | GO:0004563 | beta-N-acetylhexosaminidase activity(GO:0004563) |
| 0.3 | 2.3 | GO:0031826 | type 2A serotonin receptor binding(GO:0031826) |
| 0.2 | 1.9 | GO:0052650 | NADP-retinol dehydrogenase activity(GO:0052650) |
| 0.2 | 0.7 | GO:0033149 | FFAT motif binding(GO:0033149) |
| 0.1 | 3.8 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.1 | 3.5 | GO:0016229 | steroid dehydrogenase activity(GO:0016229) |
| 0.1 | 3.8 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.1 | 3.5 | GO:0015248 | sterol transporter activity(GO:0015248) |
| 0.1 | 0.6 | GO:0033842 | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N-acetylgalactosaminyltransferase activity(GO:0033842) |
| 0.1 | 2.1 | GO:0017110 | nucleoside-diphosphatase activity(GO:0017110) |
| 0.1 | 2.1 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.1 | 1.2 | GO:0031492 | nucleosomal DNA binding(GO:0031492) |
| 0.1 | 0.2 | GO:0046964 | 3'-phosphoadenosine 5'-phosphosulfate transmembrane transporter activity(GO:0046964) |
| 0.1 | 1.2 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.0 | 1.7 | GO:0051959 | dynein light intermediate chain binding(GO:0051959) |
| 0.0 | 0.5 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.0 | 7.9 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.0 | 3.6 | GO:0004896 | cytokine receptor activity(GO:0004896) |
| 0.0 | 2.4 | GO:0008146 | sulfotransferase activity(GO:0008146) |
| 0.0 | 1.5 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.0 | 0.1 | GO:0004427 | inorganic diphosphatase activity(GO:0004427) |
| 0.0 | 2.1 | GO:0000287 | magnesium ion binding(GO:0000287) |
| 0.0 | 1.2 | GO:0016616 | oxidoreductase activity, acting on the CH-OH group of donors, NAD or NADP as acceptor(GO:0016616) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.6 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 3.8 | REACTOME REVERSIBLE HYDRATION OF CARBON DIOXIDE | Genes involved in Reversible Hydration of Carbon Dioxide |
| 0.2 | 3.8 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.0 | 2.6 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.0 | 0.2 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |