Project
PRJNA207719: Tissue specific transcriptome profiling
Navigation
Downloads
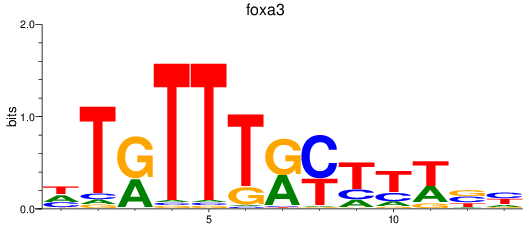
Results for foxa3
Z-value: 1.66
Transcription factors associated with foxa3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
foxa3
|
ENSDARG00000012788 | forkhead box A3 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| foxa3 | dr11_v1_chr18_-_46354269_46354269 | 0.30 | 6.3e-01 | Click! |
Activity profile of foxa3 motif
Sorted Z-values of foxa3 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr18_-_16795262 | 1.44 |
ENSDART00000048722
|
ampd3b
|
adenosine monophosphate deaminase 3b |
| chr13_+_22480857 | 1.43 |
ENSDART00000078721
ENSDART00000044719 ENSDART00000130957 ENSDART00000078757 ENSDART00000130424 ENSDART00000078747 |
ldb3a
|
LIM domain binding 3a |
| chr12_-_26064480 | 1.17 |
ENSDART00000158215
ENSDART00000171206 ENSDART00000171212 ENSDART00000182956 ENSDART00000186779 |
ldb3b
|
LIM domain binding 3b |
| chr3_-_43770876 | 1.11 |
ENSDART00000160162
|
zgc:92162
|
zgc:92162 |
| chr16_+_23978978 | 1.07 |
ENSDART00000058964
ENSDART00000135084 |
apoa2
|
apolipoprotein A-II |
| chr23_-_21515182 | 1.07 |
ENSDART00000142000
|
rnf207b
|
ring finger protein 207b |
| chr21_-_20840714 | 1.07 |
ENSDART00000144861
ENSDART00000139430 |
c6
|
complement component 6 |
| chr4_-_6373735 | 1.06 |
ENSDART00000140100
|
MDFIC
|
si:ch73-156e19.1 |
| chr3_-_50865079 | 1.03 |
ENSDART00000164295
|
pmp22a
|
peripheral myelin protein 22a |
| chr21_-_17296789 | 1.01 |
ENSDART00000192180
|
gfi1b
|
growth factor independent 1B transcription repressor |
| chr10_-_15128771 | 0.99 |
ENSDART00000101261
|
spp1
|
secreted phosphoprotein 1 |
| chr18_+_35842933 | 0.98 |
ENSDART00000151587
ENSDART00000131121 |
ppp1r13l
|
protein phosphatase 1, regulatory subunit 13 like |
| chr18_+_20566817 | 0.96 |
ENSDART00000100716
|
bida
|
BH3 interacting domain death agonist |
| chr6_-_13206255 | 0.96 |
ENSDART00000065373
|
eef1b2
|
eukaryotic translation elongation factor 1 beta 2 |
| chr25_-_13188678 | 0.95 |
ENSDART00000125754
|
si:ch211-147m6.1
|
si:ch211-147m6.1 |
| chr5_-_16351306 | 0.93 |
ENSDART00000168643
|
CABZ01088700.1
|
|
| chr8_+_49570884 | 0.91 |
ENSDART00000182117
ENSDART00000108613 |
rasef
|
RAS and EF-hand domain containing |
| chr21_-_22951604 | 0.88 |
ENSDART00000083449
ENSDART00000180129 |
dub
|
duboraya |
| chr14_-_29799993 | 0.86 |
ENSDART00000133775
ENSDART00000005568 |
pdlim3b
|
PDZ and LIM domain 3b |
| chr22_-_10110959 | 0.85 |
ENSDART00000031005
ENSDART00000147580 |
gls2b
|
glutaminase 2b (liver, mitochondrial) |
| chr2_-_36933472 | 0.83 |
ENSDART00000170405
|
BX470229.2
|
|
| chr14_-_4120636 | 0.83 |
ENSDART00000059230
|
irf2
|
interferon regulatory factor 2 |
| chr24_-_26369185 | 0.81 |
ENSDART00000080039
|
lrrc31
|
leucine rich repeat containing 31 |
| chr20_-_9980318 | 0.80 |
ENSDART00000080664
|
ACTC1
|
zgc:86709 |
| chr5_+_22970617 | 0.80 |
ENSDART00000192859
|
hmgn7
|
high mobility group nucleosomal binding domain 7 |
| chr2_+_10134345 | 0.80 |
ENSDART00000100725
|
ahsg2
|
alpha-2-HS-glycoprotein 2 |
| chr6_-_8736766 | 0.79 |
ENSDART00000143956
|
cavin2b
|
caveolae associated protein 2b |
| chr3_+_13559199 | 0.77 |
ENSDART00000166547
|
si:ch73-106n3.1
|
si:ch73-106n3.1 |
| chr7_-_5029478 | 0.76 |
ENSDART00000193819
|
ltb4r
|
leukotriene B4 receptor |
| chr2_-_32262287 | 0.76 |
ENSDART00000056621
ENSDART00000039717 |
fam49ba
|
family with sequence similarity 49, member Ba |
| chr7_-_32020100 | 0.75 |
ENSDART00000185433
|
kif18a
|
kinesin family member 18A |
| chr21_-_25295087 | 0.74 |
ENSDART00000087910
ENSDART00000147860 |
st14b
|
suppression of tumorigenicity 14 (colon carcinoma) b |
| chr23_+_32044410 | 0.72 |
ENSDART00000048628
|
mylk2
|
myosin light chain kinase 2 |
| chr9_-_21912227 | 0.71 |
ENSDART00000145576
|
lmo7a
|
LIM domain 7a |
| chr22_-_36856405 | 0.71 |
ENSDART00000029588
|
kng1
|
kininogen 1 |
| chr16_-_17713859 | 0.69 |
ENSDART00000149275
|
zgc:174935
|
zgc:174935 |
| chr6_+_52918537 | 0.68 |
ENSDART00000174229
|
or137-1
|
odorant receptor, family H, subfamily 137, member 1 |
| chr14_+_20893065 | 0.68 |
ENSDART00000079452
|
lygl1
|
lysozyme g-like 1 |
| chr2_-_38035235 | 0.67 |
ENSDART00000075904
|
cbln5
|
cerebellin 5 |
| chr1_+_26667872 | 0.67 |
ENSDART00000152803
ENSDART00000152144 ENSDART00000152785 ENSDART00000152393 |
hemgn
|
hemogen |
| chr5_-_63302944 | 0.66 |
ENSDART00000047110
|
gsnb
|
gelsolin b |
| chr21_-_35419486 | 0.65 |
ENSDART00000138529
|
si:dkeyp-23e4.3
|
si:dkeyp-23e4.3 |
| chr4_-_12795436 | 0.64 |
ENSDART00000131026
ENSDART00000075127 |
b2m
|
beta-2-microglobulin |
| chr15_+_24644016 | 0.64 |
ENSDART00000043292
|
smtnl
|
smoothelin, like |
| chr22_+_7497319 | 0.64 |
ENSDART00000034564
|
CELA1 (1 of many)
|
zgc:92511 |
| chr25_+_14087045 | 0.63 |
ENSDART00000155770
|
actc1c
|
actin, alpha, cardiac muscle 1c |
| chr23_+_26026383 | 0.63 |
ENSDART00000141553
|
pfkfb1
|
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 1 |
| chr8_-_43923788 | 0.63 |
ENSDART00000146152
|
adgrd1
|
adhesion G protein-coupled receptor D1 |
| chr24_+_19542323 | 0.62 |
ENSDART00000140379
ENSDART00000142830 |
sulf1
|
sulfatase 1 |
| chr3_+_15828999 | 0.62 |
ENSDART00000104397
|
tmem11
|
transmembrane protein 11 |
| chr7_+_20512419 | 0.62 |
ENSDART00000173907
|
si:dkey-19b23.14
|
si:dkey-19b23.14 |
| chr6_-_53048291 | 0.60 |
ENSDART00000103267
|
fam212ab
|
family with sequence similarity 212, member Ab |
| chr20_+_52546186 | 0.59 |
ENSDART00000110777
ENSDART00000153377 ENSDART00000153013 ENSDART00000042704 |
eef1db
|
eukaryotic translation elongation factor 1 delta b (guanine nucleotide exchange protein) |
| chr3_+_23743139 | 0.58 |
ENSDART00000187409
|
hoxb3a
|
homeobox B3a |
| chr2_+_23731194 | 0.58 |
ENSDART00000155747
|
slc22a13a
|
solute carrier family 22 member 13a |
| chr25_-_30344117 | 0.58 |
ENSDART00000167077
|
pdia3
|
protein disulfide isomerase family A, member 3 |
| chr23_+_36653376 | 0.57 |
ENSDART00000053189
|
gpr182
|
G protein-coupled receptor 182 |
| chr11_-_11336986 | 0.56 |
ENSDART00000016677
|
zgc:77929
|
zgc:77929 |
| chr1_-_44940830 | 0.56 |
ENSDART00000097500
ENSDART00000134464 ENSDART00000137216 |
tmem176
|
transmembrane protein 176 |
| chr5_+_28830643 | 0.55 |
ENSDART00000051448
|
serpina7
|
serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 7 |
| chr20_+_15015557 | 0.55 |
ENSDART00000039345
|
myoc
|
myocilin |
| chr11_-_183328 | 0.55 |
ENSDART00000168562
|
avil
|
advillin |
| chr16_-_21785261 | 0.55 |
ENSDART00000078858
|
si:ch73-86n18.1
|
si:ch73-86n18.1 |
| chr2_+_37875789 | 0.54 |
ENSDART00000036318
ENSDART00000127679 |
cbln13
|
cerebellin 13 |
| chr4_+_14981854 | 0.54 |
ENSDART00000067046
|
cax1
|
cation/H+ exchanger protein 1 |
| chr2_+_30916188 | 0.54 |
ENSDART00000137012
|
myom1a
|
myomesin 1a (skelemin) |
| chr3_+_29941777 | 0.54 |
ENSDART00000113889
|
ifi35
|
interferon-induced protein 35 |
| chr24_+_25258904 | 0.53 |
ENSDART00000155714
|
gabrr3b
|
gamma-aminobutyric acid (GABA) A receptor, rho 3b |
| chr4_+_12358822 | 0.53 |
ENSDART00000172557
ENSDART00000150627 |
pimr168
|
Pim proto-oncogene, serine/threonine kinase, related 168 |
| chr5_+_36513605 | 0.53 |
ENSDART00000013590
|
wnt11
|
wingless-type MMTV integration site family, member 11 |
| chr10_+_23553533 | 0.53 |
ENSDART00000159654
|
CR847844.2
|
|
| chr9_-_23253870 | 0.53 |
ENSDART00000143657
ENSDART00000169911 |
acmsd
|
aminocarboxymuconate semialdehyde decarboxylase |
| chr5_+_28797771 | 0.52 |
ENSDART00000188845
ENSDART00000149066 |
si:ch211-186e20.7
|
si:ch211-186e20.7 |
| chr22_-_14128716 | 0.52 |
ENSDART00000140323
|
si:ch211-246m6.4
|
si:ch211-246m6.4 |
| chr7_-_8417315 | 0.52 |
ENSDART00000173046
|
jac1
|
jacalin 1 |
| chr8_-_19313510 | 0.51 |
ENSDART00000164780
ENSDART00000137133 |
rgl1
|
ral guanine nucleotide dissociation stimulator-like 1 |
| chr16_-_54455573 | 0.51 |
ENSDART00000075275
|
pklr
|
pyruvate kinase L/R |
| chr25_-_31423493 | 0.51 |
ENSDART00000027661
|
myod1
|
myogenic differentiation 1 |
| chr4_+_1619584 | 0.51 |
ENSDART00000148486
|
scaf11
|
SR-related CTD-associated factor 11 |
| chr5_+_42092227 | 0.51 |
ENSDART00000097583
ENSDART00000171678 |
ubb
|
ubiquitin B |
| chr5_-_14509137 | 0.50 |
ENSDART00000180742
|
si:ch211-244o22.2
|
si:ch211-244o22.2 |
| chr11_+_13223625 | 0.50 |
ENSDART00000161275
|
abcb11b
|
ATP-binding cassette, sub-family B (MDR/TAP), member 11b |
| chr3_+_32129632 | 0.50 |
ENSDART00000174522
|
zgc:109934
|
zgc:109934 |
| chr1_+_11107688 | 0.50 |
ENSDART00000109858
|
knstrn
|
kinetochore-localized astrin/SPAG5 binding protein |
| chr7_-_20158380 | 0.50 |
ENSDART00000142891
|
acap1
|
ArfGAP with coiled-coil, ankyrin repeat and PH domains 1 |
| chr13_-_36761379 | 0.49 |
ENSDART00000131534
ENSDART00000029824 |
map4k5
|
mitogen-activated protein kinase kinase kinase kinase 5 |
| chr25_-_13188214 | 0.49 |
ENSDART00000187298
|
si:ch211-147m6.1
|
si:ch211-147m6.1 |
| chr1_+_1805294 | 0.49 |
ENSDART00000103850
|
atp1a1a.3
|
ATPase Na+/K+ transporting subunit alpha 1a, tandem duplicate 3 |
| chr11_-_30634286 | 0.49 |
ENSDART00000191019
|
zgc:153665
|
zgc:153665 |
| chr4_-_12795030 | 0.49 |
ENSDART00000150427
|
b2m
|
beta-2-microglobulin |
| chr1_+_9153141 | 0.49 |
ENSDART00000081343
|
plk1
|
polo-like kinase 1 (Drosophila) |
| chr5_+_28857969 | 0.49 |
ENSDART00000149850
|
si:ch211-186e20.2
|
si:ch211-186e20.2 |
| chr17_-_23709347 | 0.48 |
ENSDART00000124661
|
papss2a
|
3'-phosphoadenosine 5'-phosphosulfate synthase 2a |
| chr3_-_41292569 | 0.47 |
ENSDART00000111856
|
sdk1a
|
sidekick cell adhesion molecule 1a |
| chr6_-_38930726 | 0.47 |
ENSDART00000154151
|
hdac7b
|
histone deacetylase 7b |
| chr17_+_33418475 | 0.47 |
ENSDART00000169145
|
snap23.1
|
synaptosomal-associated protein 23.1 |
| chr20_-_1383916 | 0.47 |
ENSDART00000152373
|
scara5
|
scavenger receptor class A, member 5 (putative) |
| chr20_+_6142433 | 0.47 |
ENSDART00000054084
ENSDART00000136986 |
ttr
|
transthyretin (prealbumin, amyloidosis type I) |
| chr5_+_28830388 | 0.46 |
ENSDART00000149150
|
serpina7
|
serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 7 |
| chr23_-_21215311 | 0.46 |
ENSDART00000112424
|
megf6a
|
multiple EGF-like-domains 6a |
| chr16_-_13992646 | 0.46 |
ENSDART00000139623
|
si:dkey-85k15.6
|
si:dkey-85k15.6 |
| chr7_+_34305903 | 0.46 |
ENSDART00000173575
|
bbox1
|
butyrobetaine (gamma), 2-oxoglutarate dioxygenase (gamma-butyrobetaine hydroxylase) 1 |
| chr13_-_36844945 | 0.46 |
ENSDART00000129562
ENSDART00000150899 |
nin
|
ninein (GSK3B interacting protein) |
| chr3_-_41292275 | 0.46 |
ENSDART00000144088
|
sdk1a
|
sidekick cell adhesion molecule 1a |
| chr24_-_31306724 | 0.46 |
ENSDART00000165399
|
acp5b
|
acid phosphatase 5b, tartrate resistant |
| chr14_-_6931889 | 0.45 |
ENSDART00000166439
|
si:ch211-266k2.1
|
si:ch211-266k2.1 |
| chr14_-_36799280 | 0.43 |
ENSDART00000168615
|
rnf130
|
ring finger protein 130 |
| chr4_-_16334362 | 0.43 |
ENSDART00000101461
|
epyc
|
epiphycan |
| chr20_+_51104367 | 0.43 |
ENSDART00000073981
|
eif2s1b
|
eukaryotic translation initiation factor 2, subunit 1 alpha b |
| chr24_-_26485098 | 0.43 |
ENSDART00000135496
ENSDART00000009609 ENSDART00000133782 ENSDART00000141029 ENSDART00000113739 |
eif5a
|
eukaryotic translation initiation factor 5A |
| chr5_+_29726428 | 0.43 |
ENSDART00000143183
|
ddx31
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 31 |
| chr2_+_38161318 | 0.42 |
ENSDART00000044264
|
mmp14b
|
matrix metallopeptidase 14b (membrane-inserted) |
| chr15_-_25571865 | 0.42 |
ENSDART00000077836
|
mmp20b
|
matrix metallopeptidase 20b (enamelysin) |
| chr17_+_8799661 | 0.42 |
ENSDART00000105326
|
tonsl
|
tonsoku-like, DNA repair protein |
| chr2_+_20539402 | 0.42 |
ENSDART00000129585
|
si:ch73-14h1.2
|
si:ch73-14h1.2 |
| chr8_+_39795918 | 0.42 |
ENSDART00000143413
|
si:ch211-170d8.2
|
si:ch211-170d8.2 |
| chr5_-_54712159 | 0.41 |
ENSDART00000149207
|
ccnb1
|
cyclin B1 |
| chr5_-_66160415 | 0.41 |
ENSDART00000073895
|
mboat4
|
membrane bound O-acyltransferase domain containing 4 |
| chr1_+_37391716 | 0.41 |
ENSDART00000191986
|
sparcl1
|
SPARC-like 1 |
| chr7_+_58730201 | 0.41 |
ENSDART00000073640
|
plag1
|
pleiomorphic adenoma gene 1 |
| chr3_+_52442102 | 0.41 |
ENSDART00000155455
|
si:ch211-191c10.1
|
si:ch211-191c10.1 |
| chr3_+_12744083 | 0.41 |
ENSDART00000158554
ENSDART00000169545 |
cyp2k21
|
cytochrome P450, family 2, subfamily k, polypeptide 21 |
| chr15_-_26552393 | 0.41 |
ENSDART00000150152
|
serpinf2b
|
serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), member 2b |
| chr17_-_53439866 | 0.40 |
ENSDART00000154826
|
mycbp
|
c-myc binding protein |
| chr23_+_42813415 | 0.40 |
ENSDART00000055577
|
myl9a
|
myosin, light chain 9a, regulatory |
| chr2_+_24936766 | 0.40 |
ENSDART00000025962
|
gyg1a
|
glycogenin 1a |
| chr15_-_44052927 | 0.40 |
ENSDART00000166209
|
wu:fb44b02
|
wu:fb44b02 |
| chr17_+_30843881 | 0.40 |
ENSDART00000149600
ENSDART00000148547 |
tpp1
|
tripeptidyl peptidase I |
| chr6_-_25384526 | 0.40 |
ENSDART00000160544
|
PKN2 (1 of many)
|
zgc:153916 |
| chr4_-_30055196 | 0.39 |
ENSDART00000139539
|
si:rp71-7l19.2
|
si:rp71-7l19.2 |
| chr17_-_42988356 | 0.39 |
ENSDART00000024558
|
zgc:92137
|
zgc:92137 |
| chr12_-_29301022 | 0.39 |
ENSDART00000187826
|
sh2d4bb
|
SH2 domain containing 4Bb |
| chr1_-_58009216 | 0.39 |
ENSDART00000143829
|
nxnl1
|
nucleoredoxin like 1 |
| chr15_-_37719679 | 0.38 |
ENSDART00000184025
|
si:dkey-117a8.1
|
si:dkey-117a8.1 |
| chr22_+_6905210 | 0.38 |
ENSDART00000167530
|
CABZ01065328.2
|
|
| chr21_-_26490186 | 0.38 |
ENSDART00000009889
|
zgc:110540
|
zgc:110540 |
| chr3_+_12484008 | 0.38 |
ENSDART00000182229
|
vasnb
|
vasorin b |
| chr11_+_2391649 | 0.38 |
ENSDART00000104571
|
igfbp6a
|
insulin-like growth factor binding protein 6a |
| chr2_-_36597793 | 0.38 |
ENSDART00000113397
|
tradv30.0.6
|
T-cell receptor alpha/delta variable 30.0.6 |
| chr10_+_15603082 | 0.38 |
ENSDART00000024450
|
zfand5b
|
zinc finger, AN1-type domain 5b |
| chr2_-_57264262 | 0.37 |
ENSDART00000183815
ENSDART00000149829 ENSDART00000088508 ENSDART00000149508 |
mbd3a
|
methyl-CpG binding domain protein 3a |
| chr17_+_24036791 | 0.37 |
ENSDART00000140767
|
b3gnt2b
|
UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 2b |
| chr16_-_25233515 | 0.37 |
ENSDART00000058943
|
zgc:110182
|
zgc:110182 |
| chr3_-_56871330 | 0.37 |
ENSDART00000014103
|
zgc:112148
|
zgc:112148 |
| chr5_+_28849155 | 0.36 |
ENSDART00000079090
|
zgc:174259
|
zgc:174259 |
| chr16_+_23913943 | 0.36 |
ENSDART00000175404
ENSDART00000129525 |
apoa4b.1
|
apolipoprotein A-IV b, tandem duplicate 1 |
| chr23_-_29357764 | 0.36 |
ENSDART00000156512
|
si:ch211-129o18.4
|
si:ch211-129o18.4 |
| chr19_+_7636941 | 0.36 |
ENSDART00000081611
ENSDART00000163805 ENSDART00000112404 |
cgnb
|
cingulin b |
| chr21_-_21020708 | 0.36 |
ENSDART00000064032
|
eif4ebp1
|
eukaryotic translation initiation factor 4E binding protein 1 |
| chr8_+_30456161 | 0.35 |
ENSDART00000085894
|
pgm5
|
phosphoglucomutase 5 |
| chr5_+_37068223 | 0.35 |
ENSDART00000164279
|
si:dkeyp-110c7.4
|
si:dkeyp-110c7.4 |
| chr15_+_14856307 | 0.35 |
ENSDART00000167213
|
diabloa
|
diablo, IAP-binding mitochondrial protein a |
| chr5_+_28848870 | 0.35 |
ENSDART00000149563
|
zgc:174259
|
zgc:174259 |
| chr9_-_34396264 | 0.35 |
ENSDART00000045754
|
grtp1b
|
growth hormone regulated TBC protein 1b |
| chr11_+_2391469 | 0.35 |
ENSDART00000182121
|
igfbp6a
|
insulin-like growth factor binding protein 6a |
| chr4_-_1824836 | 0.35 |
ENSDART00000111858
|
mrpl42
|
mitochondrial ribosomal protein L42 |
| chr8_+_30780985 | 0.35 |
ENSDART00000193971
ENSDART00000111533 |
tas2r200.2
|
taste receptor, type 2, member 200, tandem duplicate 2 |
| chr5_+_32345187 | 0.35 |
ENSDART00000147132
|
c9
|
complement component 9 |
| chr15_-_29586747 | 0.35 |
ENSDART00000076749
|
samsn1a
|
SAM domain, SH3 domain and nuclear localisation signals 1a |
| chr16_-_35952789 | 0.34 |
ENSDART00000180118
|
eva1ba
|
eva-1 homolog Ba (C. elegans) |
| chr22_-_24285432 | 0.34 |
ENSDART00000164083
|
si:ch211-117l17.4
|
si:ch211-117l17.4 |
| chr7_-_51300277 | 0.34 |
ENSDART00000174174
|
gc2
|
guanylyl cyclase 2 |
| chr25_-_35101673 | 0.34 |
ENSDART00000140864
|
zgc:162611
|
zgc:162611 |
| chr3_+_15828426 | 0.34 |
ENSDART00000135299
|
tmem11
|
transmembrane protein 11 |
| chr25_-_35101396 | 0.34 |
ENSDART00000138865
|
zgc:162611
|
zgc:162611 |
| chr11_-_25539323 | 0.34 |
ENSDART00000155785
|
si:dkey-245f22.3
|
si:dkey-245f22.3 |
| chr7_+_4682659 | 0.34 |
ENSDART00000112697
|
si:ch211-225k7.2
|
si:ch211-225k7.2 |
| chr3_-_6719232 | 0.34 |
ENSDART00000154294
|
atg4db
|
autophagy related 4D, cysteine peptidase b |
| chr3_+_2719243 | 0.33 |
ENSDART00000189256
|
CR388047.3
|
|
| chr3_+_32571929 | 0.33 |
ENSDART00000151025
|
si:ch73-248e21.1
|
si:ch73-248e21.1 |
| chr21_-_1625976 | 0.32 |
ENSDART00000066621
|
vmo1b
|
vitelline membrane outer layer 1 homolog b |
| chr11_-_7380674 | 0.32 |
ENSDART00000014979
ENSDART00000103418 |
vtg3
|
vitellogenin 3, phosvitinless |
| chr13_+_6295336 | 0.32 |
ENSDART00000172098
|
angpt2a
|
angiopoietin 2a |
| chr7_-_18545515 | 0.32 |
ENSDART00000040534
|
rgs12a
|
regulator of G protein signaling 12a |
| chr3_-_10739625 | 0.31 |
ENSDART00000156144
|
zgc:112965
|
zgc:112965 |
| chr5_+_28858345 | 0.31 |
ENSDART00000111180
|
si:ch211-186e20.2
|
si:ch211-186e20.2 |
| chr9_-_456541 | 0.31 |
ENSDART00000159596
|
zgc:91849
|
zgc:91849 |
| chr9_-_44948488 | 0.31 |
ENSDART00000059228
|
vil1
|
villin 1 |
| chr17_+_43843536 | 0.31 |
ENSDART00000164097
|
FO704777.1
|
|
| chr3_-_1364946 | 0.31 |
ENSDART00000159328
|
FP236551.1
|
|
| chr16_+_11724230 | 0.31 |
ENSDART00000060266
|
ceacam1
|
carcinoembryonic antigen-related cell adhesion molecule 1 |
| chr5_-_20194876 | 0.31 |
ENSDART00000122587
|
dao.1
|
D-amino-acid oxidase, tandem duplicate 1 |
| chr7_+_19850889 | 0.31 |
ENSDART00000100782
|
mus81
|
MUS81 structure-specific endonuclease subunit |
| chr2_-_5475910 | 0.31 |
ENSDART00000100954
ENSDART00000172143 ENSDART00000132496 |
proca
proca
|
protein C (inactivator of coagulation factors Va and VIIIa), a protein C (inactivator of coagulation factors Va and VIIIa), a |
| chr5_-_23485161 | 0.31 |
ENSDART00000170293
ENSDART00000134069 |
si:dkeyp-20g2.1
si:dkeyp-20g2.3
|
si:dkeyp-20g2.1 si:dkeyp-20g2.3 |
| chr24_+_20575259 | 0.30 |
ENSDART00000010488
|
klhl40b
|
kelch-like family member 40b |
| chr14_+_32918484 | 0.30 |
ENSDART00000105721
|
lnx2b
|
ligand of numb-protein X 2b |
| chr13_-_16066997 | 0.30 |
ENSDART00000184790
|
SPATA48
|
spermatogenesis associated 48 |
| chr3_-_23470986 | 0.29 |
ENSDART00000113135
|
msl1a
|
male-specific lethal 1 homolog a (Drosophila) |
| chr8_-_3312384 | 0.29 |
ENSDART00000035965
|
fut9b
|
fucosyltransferase 9b |
| chr1_+_12068800 | 0.29 |
ENSDART00000055566
|
xpa
|
xeroderma pigmentosum, complementation group A |
| chr2_+_25839940 | 0.29 |
ENSDART00000139927
|
eif5a2
|
eukaryotic translation initiation factor 5A2 |
| chr2_-_22569488 | 0.29 |
ENSDART00000158608
|
FQ377628.1
|
|
| chr7_-_58729894 | 0.28 |
ENSDART00000149347
|
chchd7
|
coiled-coil-helix-coiled-coil-helix domain containing 7 |
| chr16_+_20904754 | 0.28 |
ENSDART00000006043
|
hoxa11b
|
homeobox A11b |
| chr13_+_42544009 | 0.28 |
ENSDART00000145409
|
si:dkey-221j11.3
|
si:dkey-221j11.3 |
| chr18_+_33550667 | 0.28 |
ENSDART00000141246
ENSDART00000136070 |
si:dkey-47k20.1
|
si:dkey-47k20.1 |
| chr7_-_17570923 | 0.28 |
ENSDART00000188476
ENSDART00000080624 |
nitr5
|
novel immune-type receptor 5 |
| chr15_-_26552652 | 0.28 |
ENSDART00000152336
|
serpinf2b
|
serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), member 2b |
| chr16_-_30564319 | 0.27 |
ENSDART00000145087
|
lmna
|
lamin A |
Network of associatons between targets according to the STRING database.
First level regulatory network of foxa3
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.1 | GO:1903779 | regulation of cardiac conduction(GO:1903779) |
| 0.3 | 1.0 | GO:2001244 | positive regulation of intrinsic apoptotic signaling pathway(GO:2001244) |
| 0.3 | 0.9 | GO:0061341 | non-canonical Wnt signaling pathway involved in heart development(GO:0061341) |
| 0.2 | 0.7 | GO:0016998 | cell wall macromolecule catabolic process(GO:0016998) |
| 0.2 | 0.5 | GO:0002074 | extraocular skeletal muscle development(GO:0002074) |
| 0.2 | 0.5 | GO:0034036 | sulfate assimilation(GO:0000103) purine ribonucleoside bisphosphate metabolic process(GO:0034035) purine ribonucleoside bisphosphate biosynthetic process(GO:0034036) 3'-phosphoadenosine 5'-phosphosulfate metabolic process(GO:0050427) 3'-phosphoadenosine 5'-phosphosulfate biosynthetic process(GO:0050428) |
| 0.1 | 0.5 | GO:0048014 | Tie signaling pathway(GO:0048014) |
| 0.1 | 0.5 | GO:0015722 | canalicular bile acid transport(GO:0015722) bile acid secretion(GO:0032782) |
| 0.1 | 0.9 | GO:0045901 | positive regulation of translational elongation(GO:0045901) positive regulation of translational termination(GO:0045905) |
| 0.1 | 0.3 | GO:0006957 | complement activation, alternative pathway(GO:0006957) |
| 0.1 | 0.8 | GO:0006537 | glutamate biosynthetic process(GO:0006537) glutamine catabolic process(GO:0006543) |
| 0.1 | 1.4 | GO:0032264 | IMP salvage(GO:0032264) |
| 0.1 | 0.7 | GO:1900047 | negative regulation of blood coagulation(GO:0030195) negative regulation of hemostasis(GO:1900047) |
| 0.1 | 0.3 | GO:0098528 | skeletal muscle fiber differentiation(GO:0098528) |
| 0.1 | 0.5 | GO:0051988 | regulation of attachment of spindle microtubules to kinetochore(GO:0051988) |
| 0.1 | 0.3 | GO:1901255 | UV-damage excision repair(GO:0070914) nucleotide-excision repair involved in interstrand cross-link repair(GO:1901255) |
| 0.1 | 0.5 | GO:0034454 | microtubule anchoring at centrosome(GO:0034454) |
| 0.1 | 0.3 | GO:0010359 | regulation of anion channel activity(GO:0010359) |
| 0.1 | 0.5 | GO:0019805 | quinolinate biosynthetic process(GO:0019805) quinolinate metabolic process(GO:0046874) |
| 0.1 | 1.1 | GO:0045824 | negative regulation of innate immune response(GO:0045824) |
| 0.1 | 0.5 | GO:0035176 | social behavior(GO:0035176) intraspecies interaction between organisms(GO:0051703) |
| 0.1 | 0.4 | GO:0097186 | amelogenesis(GO:0097186) |
| 0.1 | 1.1 | GO:0032309 | icosanoid secretion(GO:0032309) arachidonic acid secretion(GO:0050482) arachidonate transport(GO:1903963) |
| 0.1 | 0.6 | GO:2000290 | smoothened signaling pathway involved in ventral spinal cord patterning(GO:0021910) regulation of myotome development(GO:2000290) |
| 0.1 | 0.3 | GO:2000392 | lamellipodium morphogenesis(GO:0072673) regulation of lamellipodium morphogenesis(GO:2000392) |
| 0.1 | 0.3 | GO:0006524 | alanine metabolic process(GO:0006522) alanine catabolic process(GO:0006524) pyruvate family amino acid metabolic process(GO:0009078) pyruvate family amino acid catabolic process(GO:0009080) D-alanine family amino acid metabolic process(GO:0046144) D-alanine metabolic process(GO:0046436) D-alanine catabolic process(GO:0055130) |
| 0.1 | 0.3 | GO:0008635 | activation of cysteine-type endopeptidase activity involved in apoptotic process by cytochrome c(GO:0008635) |
| 0.1 | 0.5 | GO:0034755 | iron ion transmembrane transport(GO:0034755) protein homotrimerization(GO:0070207) |
| 0.1 | 0.5 | GO:0045329 | carnitine biosynthetic process(GO:0045329) |
| 0.1 | 0.7 | GO:0043567 | regulation of insulin-like growth factor receptor signaling pathway(GO:0043567) |
| 0.1 | 0.2 | GO:1903232 | melanosome assembly(GO:1903232) |
| 0.1 | 0.2 | GO:0044821 | telomere localization(GO:0034397) telomere tethering at nuclear periphery(GO:0034398) meiotic telomere tethering at nuclear periphery(GO:0044821) meiotic telomere clustering(GO:0045141) meiotic attachment of telomere to nuclear envelope(GO:0070197) chromosome localization to nuclear envelope involved in homologous chromosome segregation(GO:0090220) chromosome attachment to the nuclear envelope(GO:0097240) |
| 0.1 | 0.2 | GO:0042590 | antigen processing and presentation of exogenous peptide antigen(GO:0002478) antigen processing and presentation of exogenous antigen(GO:0019884) antigen processing and presentation of exogenous peptide antigen via MHC class I(GO:0042590) |
| 0.1 | 0.3 | GO:0090342 | regulation of cell aging(GO:0090342) |
| 0.1 | 4.5 | GO:0010951 | negative regulation of endopeptidase activity(GO:0010951) |
| 0.1 | 0.2 | GO:1900364 | negative regulation of mRNA 3'-end processing(GO:0031441) negative regulation of mRNA polyadenylation(GO:1900364) |
| 0.1 | 1.4 | GO:0051014 | actin filament severing(GO:0051014) |
| 0.1 | 0.2 | GO:0002407 | dendritic cell chemotaxis(GO:0002407) dendritic cell migration(GO:0036336) |
| 0.0 | 0.4 | GO:0001958 | endochondral ossification(GO:0001958) replacement ossification(GO:0036075) |
| 0.0 | 0.7 | GO:0032094 | response to food(GO:0032094) |
| 0.0 | 0.4 | GO:0007080 | mitotic metaphase plate congression(GO:0007080) |
| 0.0 | 0.2 | GO:0001886 | endothelial cell morphogenesis(GO:0001886) |
| 0.0 | 0.3 | GO:0006574 | valine catabolic process(GO:0006574) |
| 0.0 | 0.2 | GO:1901841 | regulation of high voltage-gated calcium channel activity(GO:1901841) negative regulation of high voltage-gated calcium channel activity(GO:1901842) |
| 0.0 | 0.5 | GO:0035306 | positive regulation of dephosphorylation(GO:0035306) positive regulation of protein dephosphorylation(GO:0035307) |
| 0.0 | 0.5 | GO:0045453 | bone resorption(GO:0045453) |
| 0.0 | 0.4 | GO:0045947 | negative regulation of translational initiation(GO:0045947) |
| 0.0 | 1.1 | GO:0019835 | cytolysis(GO:0019835) |
| 0.0 | 0.5 | GO:0042572 | retinol metabolic process(GO:0042572) |
| 0.0 | 0.3 | GO:0050909 | sensory perception of taste(GO:0050909) |
| 0.0 | 0.1 | GO:0042304 | regulation of fatty acid biosynthetic process(GO:0042304) |
| 0.0 | 0.0 | GO:1903537 | meiotic cell cycle process involved in oocyte maturation(GO:1903537) regulation of meiotic cell cycle process involved in oocyte maturation(GO:1903538) |
| 0.0 | 1.0 | GO:0007007 | inner mitochondrial membrane organization(GO:0007007) |
| 0.0 | 0.1 | GO:0036306 | embryonic heart tube elongation(GO:0036306) |
| 0.0 | 0.5 | GO:0001952 | regulation of cell-matrix adhesion(GO:0001952) |
| 0.0 | 0.3 | GO:0043984 | histone H4-K16 acetylation(GO:0043984) |
| 0.0 | 0.4 | GO:0045880 | positive regulation of smoothened signaling pathway(GO:0045880) |
| 0.0 | 0.1 | GO:0051574 | positive regulation of histone H3-K9 methylation(GO:0051574) |
| 0.0 | 0.3 | GO:0072498 | ventral spinal cord interneuron specification(GO:0021521) cell fate specification involved in pattern specification(GO:0060573) embryonic skeletal joint development(GO:0072498) |
| 0.0 | 1.5 | GO:0006414 | translational elongation(GO:0006414) |
| 0.0 | 0.3 | GO:0006611 | protein export from nucleus(GO:0006611) |
| 0.0 | 0.4 | GO:0030309 | poly-N-acetyllactosamine metabolic process(GO:0030309) poly-N-acetyllactosamine biosynthetic process(GO:0030311) |
| 0.0 | 0.5 | GO:0036376 | cellular potassium ion homeostasis(GO:0030007) sodium ion export from cell(GO:0036376) sodium ion export(GO:0071436) |
| 0.0 | 0.3 | GO:0031573 | intra-S DNA damage checkpoint(GO:0031573) |
| 0.0 | 0.1 | GO:0046426 | negative regulation of JAK-STAT cascade(GO:0046426) negative regulation of STAT cascade(GO:1904893) |
| 0.0 | 0.2 | GO:0046512 | diol biosynthetic process(GO:0034312) sphingosine biosynthetic process(GO:0046512) |
| 0.0 | 0.1 | GO:0002369 | T cell cytokine production(GO:0002369) |
| 0.0 | 0.2 | GO:0007197 | adenylate cyclase-inhibiting G-protein coupled acetylcholine receptor signaling pathway(GO:0007197) |
| 0.0 | 0.1 | GO:0010032 | meiotic chromosome condensation(GO:0010032) |
| 0.0 | 0.2 | GO:0000186 | activation of MAPKK activity(GO:0000186) |
| 0.0 | 0.4 | GO:0031297 | replication fork processing(GO:0031297) |
| 0.0 | 0.4 | GO:0002430 | complement receptor mediated signaling pathway(GO:0002430) |
| 0.0 | 0.3 | GO:0006003 | fructose 2,6-bisphosphate metabolic process(GO:0006003) |
| 0.0 | 0.5 | GO:0034112 | positive regulation of homotypic cell-cell adhesion(GO:0034112) positive regulation of T cell activation(GO:0050870) positive regulation of leukocyte cell-cell adhesion(GO:1903039) |
| 0.0 | 0.1 | GO:0009146 | dGTP catabolic process(GO:0006203) purine nucleoside triphosphate catabolic process(GO:0009146) purine deoxyribonucleoside triphosphate catabolic process(GO:0009217) dGTP metabolic process(GO:0046070) |
| 0.0 | 0.2 | GO:0051601 | exocyst localization(GO:0051601) |
| 0.0 | 1.6 | GO:0055113 | epiboly involved in gastrulation with mouth forming second(GO:0055113) |
| 0.0 | 0.1 | GO:0034975 | protein folding in endoplasmic reticulum(GO:0034975) |
| 0.0 | 0.9 | GO:0007019 | microtubule depolymerization(GO:0007019) |
| 0.0 | 0.2 | GO:0048312 | intracellular distribution of mitochondria(GO:0048312) |
| 0.0 | 0.2 | GO:0060971 | embryonic heart tube left/right pattern formation(GO:0060971) |
| 0.0 | 0.1 | GO:1901099 | histone dephosphorylation(GO:0016576) negative regulation of signal transduction in absence of ligand(GO:1901099) regulation of extrinsic apoptotic signaling pathway in absence of ligand(GO:2001239) negative regulation of extrinsic apoptotic signaling pathway in absence of ligand(GO:2001240) |
| 0.0 | 0.2 | GO:0002084 | protein depalmitoylation(GO:0002084) macromolecule depalmitoylation(GO:0098734) |
| 0.0 | 0.4 | GO:2001236 | regulation of extrinsic apoptotic signaling pathway(GO:2001236) |
| 0.0 | 0.1 | GO:0006895 | Golgi to endosome transport(GO:0006895) |
| 0.0 | 0.2 | GO:0045667 | regulation of osteoblast differentiation(GO:0045667) |
| 0.0 | 0.0 | GO:0019323 | pentose catabolic process(GO:0019323) |
| 0.0 | 0.9 | GO:0010842 | retina layer formation(GO:0010842) |
| 0.0 | 0.5 | GO:0060037 | pharyngeal system development(GO:0060037) |
| 0.0 | 0.1 | GO:0061709 | reticulophagy(GO:0061709) |
| 0.0 | 0.4 | GO:1903845 | negative regulation of transforming growth factor beta receptor signaling pathway(GO:0030512) negative regulation of cellular response to transforming growth factor beta stimulus(GO:1903845) |
| 0.0 | 1.0 | GO:0060348 | bone development(GO:0060348) |
| 0.0 | 0.4 | GO:0045494 | photoreceptor cell maintenance(GO:0045494) |
| 0.0 | 0.1 | GO:1904263 | positive regulation of TORC1 signaling(GO:1904263) |
| 0.0 | 1.0 | GO:0060216 | definitive hemopoiesis(GO:0060216) |
| 0.0 | 0.3 | GO:0007168 | receptor guanylyl cyclase signaling pathway(GO:0007168) |
| 0.0 | 0.0 | GO:0046048 | pyrimidine ribonucleoside diphosphate metabolic process(GO:0009193) UDP metabolic process(GO:0046048) |
| 0.0 | 0.1 | GO:1901825 | tetraterpenoid metabolic process(GO:0016108) tetraterpenoid biosynthetic process(GO:0016109) carotenoid metabolic process(GO:0016116) carotenoid biosynthetic process(GO:0016117) xanthophyll metabolic process(GO:0016122) xanthophyll biosynthetic process(GO:0016123) zeaxanthin metabolic process(GO:1901825) zeaxanthin biosynthetic process(GO:1901827) |
| 0.0 | 0.2 | GO:0016973 | poly(A)+ mRNA export from nucleus(GO:0016973) |
| 0.0 | 0.4 | GO:0002377 | immunoglobulin production(GO:0002377) |
| 0.0 | 0.3 | GO:0070936 | protein K48-linked ubiquitination(GO:0070936) |
| 0.0 | 0.0 | GO:0034375 | high-density lipoprotein particle remodeling(GO:0034375) |
| 0.0 | 0.3 | GO:0016082 | synaptic vesicle priming(GO:0016082) |
| 0.0 | 0.2 | GO:0071577 | zinc II ion transmembrane transport(GO:0071577) |
| 0.0 | 0.2 | GO:0043171 | peptide catabolic process(GO:0043171) |
| 0.0 | 0.3 | GO:0032355 | response to estradiol(GO:0032355) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.5 | GO:0005853 | eukaryotic translation elongation factor 1 complex(GO:0005853) |
| 0.2 | 1.1 | GO:0042612 | MHC class I protein complex(GO:0042612) |
| 0.2 | 0.7 | GO:0061673 | mitotic spindle astral microtubule(GO:0061673) |
| 0.2 | 0.6 | GO:0043540 | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase complex(GO:0043540) |
| 0.1 | 0.4 | GO:0043614 | multi-eIF complex(GO:0043614) |
| 0.1 | 0.4 | GO:0097125 | cyclin B1-CDK1 complex(GO:0097125) |
| 0.1 | 0.3 | GO:0044216 | other organism(GO:0044215) other organism cell(GO:0044216) other organism part(GO:0044217) other organism cell membrane(GO:0044218) other organism membrane(GO:0044279) |
| 0.1 | 3.5 | GO:0031941 | filamentous actin(GO:0031941) |
| 0.1 | 0.3 | GO:0000110 | nucleotide-excision repair factor 1 complex(GO:0000110) |
| 0.1 | 1.1 | GO:0005579 | membrane attack complex(GO:0005579) |
| 0.1 | 0.2 | GO:0042709 | succinate-CoA ligase complex(GO:0042709) |
| 0.1 | 0.3 | GO:0032433 | filopodium tip(GO:0032433) |
| 0.1 | 0.2 | GO:0031085 | BLOC-3 complex(GO:0031085) |
| 0.1 | 0.4 | GO:0005662 | DNA replication factor A complex(GO:0005662) |
| 0.1 | 0.3 | GO:0048476 | Holliday junction resolvase complex(GO:0048476) |
| 0.0 | 0.5 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.0 | 0.5 | GO:0097431 | mitotic spindle pole(GO:0097431) |
| 0.0 | 0.3 | GO:0072487 | MSL complex(GO:0072487) |
| 0.0 | 0.2 | GO:0070390 | transcription export complex 2(GO:0070390) |
| 0.0 | 0.1 | GO:0043202 | vacuolar lumen(GO:0005775) lysosomal lumen(GO:0043202) |
| 0.0 | 0.8 | GO:0031672 | A band(GO:0031672) |
| 0.0 | 0.4 | GO:0016581 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) |
| 0.0 | 0.7 | GO:1902710 | GABA receptor complex(GO:1902710) GABA-A receptor complex(GO:1902711) |
| 0.0 | 0.8 | GO:0044853 | caveola(GO:0005901) plasma membrane raft(GO:0044853) |
| 0.0 | 1.0 | GO:0031305 | integral component of mitochondrial inner membrane(GO:0031305) |
| 0.0 | 0.5 | GO:0005791 | rough endoplasmic reticulum(GO:0005791) |
| 0.0 | 0.1 | GO:0034715 | pICln-Sm protein complex(GO:0034715) |
| 0.0 | 0.9 | GO:0005643 | nuclear pore(GO:0005643) |
| 0.0 | 0.1 | GO:0034448 | EGO complex(GO:0034448) Gtr1-Gtr2 GTPase complex(GO:1990131) |
| 0.0 | 0.6 | GO:0030496 | midbody(GO:0030496) |
| 0.0 | 0.1 | GO:0005682 | U5 snRNP(GO:0005682) |
| 0.0 | 0.5 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.0 | 0.1 | GO:0033061 | DNA recombinase mediator complex(GO:0033061) |
| 0.0 | 0.6 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 3.5 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) alpha-actinin binding(GO:0051393) |
| 0.2 | 0.7 | GO:0003796 | lysozyme activity(GO:0003796) |
| 0.2 | 0.5 | GO:0004781 | adenylylsulfate kinase activity(GO:0004020) sulfate adenylyltransferase activity(GO:0004779) sulfate adenylyltransferase (ATP) activity(GO:0004781) |
| 0.2 | 0.5 | GO:0008336 | gamma-butyrobetaine dioxygenase activity(GO:0008336) |
| 0.1 | 0.5 | GO:0015369 | calcium:proton antiporter activity(GO:0015369) metal ion:proton antiporter activity(GO:0051139) |
| 0.1 | 1.4 | GO:0047623 | AMP deaminase activity(GO:0003876) adenosine-phosphate deaminase activity(GO:0047623) |
| 0.1 | 0.5 | GO:0015126 | canalicular bile acid transmembrane transporter activity(GO:0015126) bile acid-exporting ATPase activity(GO:0015432) |
| 0.1 | 0.5 | GO:0070324 | thyroid hormone binding(GO:0070324) |
| 0.1 | 0.8 | GO:0004974 | leukotriene receptor activity(GO:0004974) |
| 0.1 | 1.1 | GO:0047498 | calcium-dependent phospholipase A2 activity(GO:0047498) |
| 0.1 | 0.8 | GO:0004359 | glutaminase activity(GO:0004359) |
| 0.1 | 0.5 | GO:0004743 | pyruvate kinase activity(GO:0004743) |
| 0.1 | 0.5 | GO:0008349 | MAP kinase kinase kinase kinase activity(GO:0008349) |
| 0.1 | 0.3 | GO:0003884 | D-amino-acid oxidase activity(GO:0003884) |
| 0.1 | 0.5 | GO:0004962 | endothelin receptor activity(GO:0004962) |
| 0.1 | 0.5 | GO:0070888 | E-box binding(GO:0070888) |
| 0.1 | 1.1 | GO:0030544 | Hsp70 protein binding(GO:0030544) |
| 0.1 | 0.2 | GO:0004775 | succinate-CoA ligase activity(GO:0004774) succinate-CoA ligase (ADP-forming) activity(GO:0004775) succinate-CoA ligase (GDP-forming) activity(GO:0004776) |
| 0.1 | 0.3 | GO:0008442 | 3-hydroxyisobutyrate dehydrogenase activity(GO:0008442) |
| 0.1 | 0.7 | GO:0031995 | insulin-like growth factor I binding(GO:0031994) insulin-like growth factor II binding(GO:0031995) |
| 0.1 | 0.4 | GO:0050431 | transforming growth factor beta binding(GO:0050431) |
| 0.1 | 0.5 | GO:0003993 | acid phosphatase activity(GO:0003993) |
| 0.1 | 0.7 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 0.1 | 0.4 | GO:0019136 | deoxynucleoside kinase activity(GO:0019136) |
| 0.1 | 2.4 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.1 | 0.4 | GO:0008190 | eukaryotic initiation factor 4E binding(GO:0008190) |
| 0.0 | 0.5 | GO:0005391 | sodium:potassium-exchanging ATPase activity(GO:0005391) |
| 0.0 | 0.3 | GO:0045735 | nutrient reservoir activity(GO:0045735) |
| 0.0 | 0.6 | GO:0016864 | protein disulfide isomerase activity(GO:0003756) intramolecular oxidoreductase activity, transposing S-S bonds(GO:0016864) |
| 0.0 | 0.8 | GO:0005080 | protein kinase C binding(GO:0005080) |
| 0.0 | 0.8 | GO:0031492 | nucleosomal DNA binding(GO:0031492) |
| 0.0 | 0.2 | GO:0017040 | ceramidase activity(GO:0017040) |
| 0.0 | 1.5 | GO:0004869 | cysteine-type endopeptidase inhibitor activity(GO:0004869) |
| 0.0 | 0.6 | GO:0008449 | N-acetylglucosamine-6-sulfatase activity(GO:0008449) |
| 0.0 | 0.5 | GO:0031386 | protein tag(GO:0031386) |
| 0.0 | 1.7 | GO:0005546 | phosphatidylinositol-4,5-bisphosphate binding(GO:0005546) |
| 0.0 | 0.1 | GO:0061656 | SUMO conjugating enzyme activity(GO:0061656) |
| 0.0 | 3.7 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 0.4 | GO:0016868 | intramolecular transferase activity, phosphotransferases(GO:0016868) |
| 0.0 | 0.1 | GO:0047291 | lactosylceramide alpha-2,3-sialyltransferase activity(GO:0047291) |
| 0.0 | 0.1 | GO:0033897 | ribonuclease T2 activity(GO:0033897) |
| 0.0 | 0.4 | GO:0008532 | N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase activity(GO:0008532) |
| 0.0 | 0.3 | GO:0048256 | flap endonuclease activity(GO:0048256) |
| 0.0 | 0.3 | GO:0003873 | 6-phosphofructo-2-kinase activity(GO:0003873) |
| 0.0 | 0.1 | GO:0047429 | nucleoside-triphosphate diphosphatase activity(GO:0047429) |
| 0.0 | 0.3 | GO:0008327 | methyl-CpG binding(GO:0008327) |
| 0.0 | 0.2 | GO:0016907 | G-protein coupled acetylcholine receptor activity(GO:0016907) |
| 0.0 | 0.7 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.0 | 0.4 | GO:0004875 | complement receptor activity(GO:0004875) |
| 0.0 | 0.1 | GO:0000179 | rRNA (adenine-N6,N6-)-dimethyltransferase activity(GO:0000179) |
| 0.0 | 0.7 | GO:0004890 | GABA-A receptor activity(GO:0004890) |
| 0.0 | 0.2 | GO:0050700 | CARD domain binding(GO:0050700) |
| 0.0 | 0.2 | GO:0005520 | insulin-like growth factor binding(GO:0005520) |
| 0.0 | 0.2 | GO:0050291 | sphingosine N-acyltransferase activity(GO:0050291) |
| 0.0 | 0.6 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.0 | 0.2 | GO:0001594 | trace-amine receptor activity(GO:0001594) |
| 0.0 | 0.1 | GO:0052885 | retinol isomerase activity(GO:0050251) all-trans-retinyl-ester hydrolase, 11-cis retinol forming activity(GO:0052885) |
| 0.0 | 0.4 | GO:0005518 | collagen binding(GO:0005518) |
| 0.0 | 0.4 | GO:0043022 | ribosome binding(GO:0043022) |
| 0.0 | 0.5 | GO:0016831 | carboxy-lyase activity(GO:0016831) |
| 0.0 | 0.2 | GO:0004709 | MAP kinase kinase kinase activity(GO:0004709) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.0 | PID INTEGRIN A9B1 PATHWAY | Alpha9 beta1 integrin signaling events |
| 0.1 | 0.7 | PID SYNDECAN 2 PATHWAY | Syndecan-2-mediated signaling events |
| 0.0 | 1.1 | PID CD8 TCR DOWNSTREAM PATHWAY | Downstream signaling in naïve CD8+ T cells |
| 0.0 | 0.7 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.0 | 1.0 | PID FOXO PATHWAY | FoxO family signaling |
| 0.0 | 0.5 | PID ARF6 PATHWAY | Arf6 signaling events |
| 0.0 | 0.5 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.0 | 1.0 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 0.5 | PID TNF PATHWAY | TNF receptor signaling pathway |
| 0.0 | 0.6 | PID P53 REGULATION PATHWAY | p53 pathway |
| 0.0 | 0.5 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 1.1 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.0 | 0.4 | ST TUMOR NECROSIS FACTOR PATHWAY | Tumor Necrosis Factor Pathway. |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.1 | REACTOME ENDOSOMAL VACUOLAR PATHWAY | Genes involved in Endosomal/Vacuolar pathway |
| 0.1 | 1.1 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.1 | 0.8 | REACTOME EICOSANOID LIGAND BINDING RECEPTORS | Genes involved in Eicosanoid ligand-binding receptors |
| 0.1 | 1.1 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.1 | 1.5 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 0.1 | 0.9 | REACTOME CYCLIN A B1 ASSOCIATED EVENTS DURING G2 M TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 0.0 | 1.4 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 0.0 | 0.7 | REACTOME INTRINSIC PATHWAY | Genes involved in Intrinsic Pathway |
| 0.0 | 0.5 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 0.0 | 0.4 | REACTOME SYNTHESIS SECRETION AND DEACYLATION OF GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |
| 0.0 | 0.4 | REACTOME MTORC1 MEDIATED SIGNALLING | Genes involved in mTORC1-mediated signalling |
| 0.0 | 0.6 | REACTOME CALNEXIN CALRETICULIN CYCLE | Genes involved in Calnexin/calreticulin cycle |
| 0.0 | 0.9 | REACTOME PTM GAMMA CARBOXYLATION HYPUSINE FORMATION AND ARYLSULFATASE ACTIVATION | Genes involved in PTM: gamma carboxylation, hypusine formation and arylsulfatase activation |
| 0.0 | 0.5 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.0 | 0.5 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.0 | 0.1 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GIP | Genes involved in Synthesis, Secretion, and Inactivation of Glucose-dependent Insulinotropic Polypeptide (GIP) |
| 0.0 | 0.3 | REACTOME MITOCHONDRIAL FATTY ACID BETA OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.0 | 0.7 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |
| 0.0 | 0.2 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.0 | 0.5 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 0.1 | REACTOME ACTIVATION OF BH3 ONLY PROTEINS | Genes involved in Activation of BH3-only proteins |
| 0.0 | 0.2 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.0 | 0.3 | REACTOME FORMATION OF INCISION COMPLEX IN GG NER | Genes involved in Formation of incision complex in GG-NER |
| 0.0 | 0.2 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.0 | 0.1 | REACTOME GROWTH HORMONE RECEPTOR SIGNALING | Genes involved in Growth hormone receptor signaling |