Project
PRJNA207719: Tissue specific transcriptome profiling
Navigation
Downloads
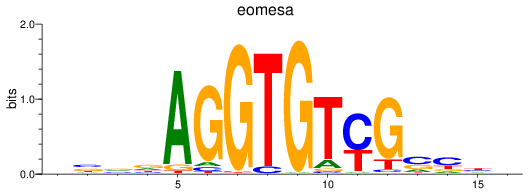
Results for eomesa
Z-value: 1.44
Transcription factors associated with eomesa
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
eomesa
|
ENSDARG00000006640 | eomesodermin homolog a |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| eomesa | dr11_v1_chr19_-_868187_868187 | 0.11 | 8.6e-01 | Click! |
Activity profile of eomesa motif
Sorted Z-values of eomesa motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr15_-_21877726 | 1.11 |
ENSDART00000127819
ENSDART00000145646 ENSDART00000100897 ENSDART00000144739 |
zgc:162608
|
zgc:162608 |
| chr2_-_32637592 | 1.09 |
ENSDART00000136353
|
si:dkeyp-73d8.8
|
si:dkeyp-73d8.8 |
| chr10_+_19596214 | 0.83 |
ENSDART00000183110
|
CABZ01059626.1
|
|
| chr7_+_27691647 | 0.77 |
ENSDART00000079091
|
cyp2r1
|
cytochrome P450, family 2, subfamily R, polypeptide 1 |
| chr16_+_23947196 | 0.73 |
ENSDART00000103190
ENSDART00000132961 ENSDART00000147690 ENSDART00000142168 |
apoa4b.2
|
apolipoprotein A-IV b, tandem duplicate 2 |
| chr22_+_25352530 | 0.72 |
ENSDART00000176742
ENSDART00000113186 |
si:dkey-240e12.6
|
si:dkey-240e12.6 |
| chr22_+_25693295 | 0.72 |
ENSDART00000123888
ENSDART00000150783 |
si:dkeyp-98a7.4
si:dkeyp-98a7.3
|
si:dkeyp-98a7.4 si:dkeyp-98a7.3 |
| chr22_+_25715925 | 0.70 |
ENSDART00000150650
|
si:dkeyp-98a7.7
|
si:dkeyp-98a7.7 |
| chr20_-_40755614 | 0.70 |
ENSDART00000061247
|
cx32.3
|
connexin 32.3 |
| chr22_+_25687525 | 0.69 |
ENSDART00000135717
|
si:dkeyp-98a7.3
|
si:dkeyp-98a7.3 |
| chr22_+_25681911 | 0.69 |
ENSDART00000113381
|
si:dkeyp-98a7.3
|
si:dkeyp-98a7.3 |
| chr22_+_25704430 | 0.67 |
ENSDART00000143776
|
si:dkeyp-98a7.3
|
si:dkeyp-98a7.3 |
| chr21_-_1742159 | 0.66 |
ENSDART00000151049
|
onecut2
|
one cut homeobox 2 |
| chr23_-_31647793 | 0.66 |
ENSDART00000145621
|
sgk1
|
serum/glucocorticoid regulated kinase 1 |
| chr23_-_31648026 | 0.64 |
ENSDART00000133569
|
sgk1
|
serum/glucocorticoid regulated kinase 1 |
| chr1_+_29741843 | 0.64 |
ENSDART00000136066
|
raph1b
|
Ras association (RalGDS/AF-6) and pleckstrin homology domains 1b |
| chr11_-_40504170 | 0.64 |
ENSDART00000165394
|
si:dkeyp-61b2.1
|
si:dkeyp-61b2.1 |
| chr10_+_19595009 | 0.57 |
ENSDART00000112276
|
zgc:173837
|
zgc:173837 |
| chr19_-_27564458 | 0.51 |
ENSDART00000123155
|
si:dkeyp-46h3.6
|
si:dkeyp-46h3.6 |
| chr10_-_25217347 | 0.51 |
ENSDART00000036906
|
kpna7
|
karyopherin alpha 7 (importin alpha 8) |
| chr22_+_25753972 | 0.50 |
ENSDART00000188417
|
AL929192.2
|
|
| chr16_+_44314097 | 0.48 |
ENSDART00000148684
|
dpys
|
dihydropyrimidinase |
| chr13_-_9318891 | 0.48 |
ENSDART00000137364
|
si:dkey-33c12.3
|
si:dkey-33c12.3 |
| chr17_+_36588281 | 0.43 |
ENSDART00000076115
|
htr1b
|
5-hydroxytryptamine (serotonin) receptor 1B |
| chr3_+_59051503 | 0.42 |
ENSDART00000160767
|
rasd4
|
rasd family member 4 |
| chr1_+_45323142 | 0.42 |
ENSDART00000132210
|
emp1
|
epithelial membrane protein 1 |
| chr25_-_13703826 | 0.41 |
ENSDART00000163398
|
pla2g15
|
phospholipase A2, group XV |
| chr13_-_51952020 | 0.34 |
ENSDART00000181079
|
LT631676.1
|
|
| chr11_-_1550709 | 0.32 |
ENSDART00000110097
|
si:ch73-303b9.1
|
si:ch73-303b9.1 |
| chr25_+_24250247 | 0.32 |
ENSDART00000064646
|
tmem86a
|
transmembrane protein 86A |
| chr14_-_2361692 | 0.31 |
ENSDART00000167696
|
PCDHGC3 (1 of many)
|
si:ch73-233f7.4 |
| chr7_+_73332486 | 0.31 |
ENSDART00000174119
ENSDART00000092051 ENSDART00000192388 |
CABZ01081780.1
|
|
| chr22_+_25274712 | 0.28 |
ENSDART00000137341
|
BX950205.1
|
|
| chr5_+_31036911 | 0.28 |
ENSDART00000141463
ENSDART00000162121 |
zzef1
|
zinc finger, ZZ-type with EF hand domain 1 |
| chr10_-_5847904 | 0.27 |
ENSDART00000161096
|
ankrd55
|
ankyrin repeat domain 55 |
| chr5_+_62319217 | 0.26 |
ENSDART00000050879
|
pho
|
phoenix |
| chr22_-_36690742 | 0.26 |
ENSDART00000017188
ENSDART00000124698 |
ncl
|
nucleolin |
| chr9_-_12885201 | 0.26 |
ENSDART00000124957
|
ankzf1
|
ankyrin repeat and zinc finger domain containing 1 |
| chr3_+_16449490 | 0.25 |
ENSDART00000141467
|
kcnj12b
|
potassium inwardly-rectifying channel, subfamily J, member 12b |
| chr1_-_45042210 | 0.25 |
ENSDART00000073694
|
smu1b
|
SMU1, DNA replication regulator and spliceosomal factor b |
| chr10_-_28193642 | 0.25 |
ENSDART00000019050
|
rps6kb1a
|
ribosomal protein S6 kinase b, polypeptide 1a |
| chr10_+_35953068 | 0.25 |
ENSDART00000015279
|
rtn4rl1a
|
reticulon 4 receptor-like 1a |
| chr2_-_57469115 | 0.24 |
ENSDART00000192201
|
pias4b
|
protein inhibitor of activated STAT, 4b |
| chr10_-_5847655 | 0.24 |
ENSDART00000192773
|
ankrd55
|
ankyrin repeat domain 55 |
| chr1_+_45323400 | 0.24 |
ENSDART00000148906
ENSDART00000132366 |
emp1
|
epithelial membrane protein 1 |
| chr7_+_23907692 | 0.23 |
ENSDART00000045479
|
syt4
|
synaptotagmin IV |
| chr22_-_5252005 | 0.23 |
ENSDART00000132942
ENSDART00000081801 |
ncln
|
nicalin |
| chr16_+_14812585 | 0.23 |
ENSDART00000134087
|
col14a1a
|
collagen, type XIV, alpha 1a |
| chr18_+_6641542 | 0.23 |
ENSDART00000160379
|
c2cd5
|
C2 calcium-dependent domain containing 5 |
| chr19_-_867071 | 0.23 |
ENSDART00000122257
|
eomesa
|
eomesodermin homolog a |
| chr2_-_13216269 | 0.22 |
ENSDART00000149947
|
bcl2b
|
BCL2, apoptosis regulator b |
| chr20_-_46773258 | 0.21 |
ENSDART00000060653
|
si:ch211-57i17.5
|
si:ch211-57i17.5 |
| chr5_+_32007627 | 0.21 |
ENSDART00000183061
|
scai
|
suppressor of cancer cell invasion |
| chr22_+_19111444 | 0.21 |
ENSDART00000109655
|
fgf22
|
fibroblast growth factor 22 |
| chr22_-_6066867 | 0.21 |
ENSDART00000142383
|
si:dkey-19a16.1
|
si:dkey-19a16.1 |
| chr13_-_280827 | 0.21 |
ENSDART00000144819
|
slc30a6
|
solute carrier family 30 (zinc transporter), member 6 |
| chr5_-_45877387 | 0.20 |
ENSDART00000183714
ENSDART00000041503 |
slc4a4a
|
solute carrier family 4 (sodium bicarbonate cotransporter), member 4a |
| chr2_-_34555945 | 0.19 |
ENSDART00000056671
|
brinp2
|
bone morphogenetic protein/retinoic acid inducible neural-specific 2 |
| chr12_-_48960308 | 0.17 |
ENSDART00000176247
|
CABZ01112647.1
|
|
| chr23_+_4362463 | 0.16 |
ENSDART00000039829
|
zgc:112175
|
zgc:112175 |
| chr1_+_5576151 | 0.16 |
ENSDART00000109756
|
cpo
|
carboxypeptidase O |
| chr6_+_54538948 | 0.16 |
ENSDART00000149270
|
tulp1b
|
tubby like protein 1b |
| chr11_+_14622379 | 0.15 |
ENSDART00000112589
|
efna2b
|
ephrin-A2b |
| chr14_-_2602445 | 0.14 |
ENSDART00000166910
|
etf1a
|
eukaryotic translation termination factor 1a |
| chr15_-_35112937 | 0.13 |
ENSDART00000154565
ENSDART00000099642 |
zgc:77118
|
zgc:77118 |
| chr25_-_17367578 | 0.12 |
ENSDART00000064591
ENSDART00000110692 |
cyp2x6
|
cytochrome P450, family 2, subfamily X, polypeptide 6 |
| chr11_-_40728380 | 0.12 |
ENSDART00000023745
|
ccdc114
|
coiled-coil domain containing 114 |
| chr3_-_60827402 | 0.12 |
ENSDART00000053494
|
anks4b
|
ankyrin repeat and sterile alpha motif domain containing 4B |
| chr20_-_32007209 | 0.12 |
ENSDART00000021575
|
adgb
|
androglobin |
| chr1_+_57142509 | 0.11 |
ENSDART00000146799
|
si:ch73-94k4.3
|
si:ch73-94k4.3 |
| chr22_+_26263290 | 0.11 |
ENSDART00000184840
|
BX649315.1
|
|
| chr1_-_55750208 | 0.10 |
ENSDART00000142244
|
dnajb1b
|
DnaJ (Hsp40) homolog, subfamily B, member 1b |
| chr21_-_27010796 | 0.10 |
ENSDART00000065398
ENSDART00000144342 ENSDART00000126542 |
ppp1r14ba
|
protein phosphatase 1, regulatory (inhibitor) subunit 14Ba |
| chr24_-_24060460 | 0.10 |
ENSDART00000142813
|
abcc13
|
ATP-binding cassette, sub-family C (CFTR/MRP), member 13 |
| chr18_+_33779440 | 0.09 |
ENSDART00000133349
|
si:dkey-145c18.3
|
si:dkey-145c18.3 |
| chr10_+_26612321 | 0.09 |
ENSDART00000134322
|
fhl1b
|
four and a half LIM domains 1b |
| chr8_+_45003659 | 0.09 |
ENSDART00000132663
|
si:ch211-163b2.4
|
si:ch211-163b2.4 |
| chr7_+_9981757 | 0.09 |
ENSDART00000113429
ENSDART00000173233 |
adamts17
|
ADAM metallopeptidase with thrombospondin type 1 motif, 17 |
| chr7_+_21275152 | 0.09 |
ENSDART00000173612
|
serpinh2
|
serine (or cysteine) peptidase inhibitor, clade H, member 2 |
| chr5_-_67762434 | 0.09 |
ENSDART00000167301
|
b4galt4
|
UDP-Gal:betaGlcNAc beta 1,4- galactosyltransferase, polypeptide 4 |
| chr2_+_42177113 | 0.09 |
ENSDART00000056441
|
tmeff1a
|
transmembrane protein with EGF-like and two follistatin-like domains 1a |
| chr10_-_8956973 | 0.09 |
ENSDART00000189564
|
mocs2
|
molybdenum cofactor synthesis 2 |
| chr1_-_57129179 | 0.09 |
ENSDART00000157226
ENSDART00000152469 |
si:ch73-94k4.2
|
si:ch73-94k4.2 |
| chr19_+_2275019 | 0.09 |
ENSDART00000136138
|
itgb8
|
integrin, beta 8 |
| chr7_-_69983462 | 0.09 |
ENSDART00000123380
|
KCNIP4
|
potassium voltage-gated channel interacting protein 4 |
| chr17_-_29312506 | 0.07 |
ENSDART00000133668
|
tecpr2
|
tectonin beta-propeller repeat containing 2 |
| chr15_+_28940352 | 0.07 |
ENSDART00000154085
|
gipr
|
gastric inhibitory polypeptide receptor |
| chr11_-_13341051 | 0.07 |
ENSDART00000121872
|
mast3b
|
microtubule associated serine/threonine kinase 3b |
| chr4_-_77624155 | 0.07 |
ENSDART00000099761
|
si:ch211-250m6.7
|
si:ch211-250m6.7 |
| chr2_+_35240485 | 0.07 |
ENSDART00000179804
|
tnr
|
tenascin R (restrictin, janusin) |
| chr15_-_45538773 | 0.06 |
ENSDART00000113494
|
MB21D2
|
Mab-21 domain containing 2 |
| chr4_-_51460013 | 0.06 |
ENSDART00000193382
|
CR628395.1
|
|
| chr18_-_20444296 | 0.06 |
ENSDART00000132993
|
kif23
|
kinesin family member 23 |
| chr10_-_9115383 | 0.06 |
ENSDART00000139324
|
si:dkeyp-41f9.3
|
si:dkeyp-41f9.3 |
| chr20_-_35052464 | 0.06 |
ENSDART00000037195
|
kif26bb
|
kinesin family member 26Bb |
| chr22_-_14475927 | 0.06 |
ENSDART00000135768
|
lrp1ba
|
low density lipoprotein receptor-related protein 1Ba |
| chr12_+_5129245 | 0.05 |
ENSDART00000169073
|
pde6c
|
phosphodiesterase 6C, cGMP-specific, cone, alpha prime |
| chr2_-_10896745 | 0.05 |
ENSDART00000114609
|
cdcp2
|
CUB domain containing protein 2 |
| chr22_+_25774750 | 0.05 |
ENSDART00000174421
|
AL929192.1
|
|
| chr5_-_41645058 | 0.05 |
ENSDART00000051092
|
riok2
|
RIO kinase 2 (yeast) |
| chr4_-_72555545 | 0.05 |
ENSDART00000172315
|
si:ch211-161m3.4
|
si:ch211-161m3.4 |
| chr12_-_10421955 | 0.04 |
ENSDART00000052004
|
zgc:153595
|
zgc:153595 |
| chr22_-_9896180 | 0.04 |
ENSDART00000138343
|
znf990
|
zinc finger protein 990 |
| chr7_+_30493684 | 0.04 |
ENSDART00000027466
|
mindy2
|
MINDY lysine 48 deubiquitinase 2 |
| chr22_+_10010292 | 0.04 |
ENSDART00000180096
|
BX324216.4
|
|
| chr10_+_16092671 | 0.04 |
ENSDART00000182761
ENSDART00000154835 |
megf10
|
multiple EGF-like-domains 10 |
| chr7_-_1101694 | 0.04 |
ENSDART00000183017
ENSDART00000182681 |
dctn1a
|
dynactin 1a |
| chr8_+_25267903 | 0.04 |
ENSDART00000093090
|
ampd2b
|
adenosine monophosphate deaminase 2b |
| chr24_-_36196077 | 0.03 |
ENSDART00000154395
|
esco1
|
establishment of sister chromatid cohesion N-acetyltransferase 1 |
| chr5_-_16475374 | 0.03 |
ENSDART00000134274
ENSDART00000136004 |
piwil2
|
piwi-like RNA-mediated gene silencing 2 |
| chr22_-_18544701 | 0.03 |
ENSDART00000132156
|
cirbpb
|
cold inducible RNA binding protein b |
| chr25_+_32071558 | 0.03 |
ENSDART00000155992
|
tjp1b
|
tight junction protein 1b |
| chr9_-_23824290 | 0.02 |
ENSDART00000059209
|
wdfy2
|
WD repeat and FYVE domain containing 2 |
| chr7_-_61845282 | 0.02 |
ENSDART00000182586
|
HTRA3
|
HtrA serine peptidase 3 |
| chr5_+_45000597 | 0.02 |
ENSDART00000051146
|
dmrt3a
|
doublesex and mab-3 related transcription factor 3a |
| chr21_-_14815952 | 0.01 |
ENSDART00000134278
ENSDART00000067004 |
phpt1
|
phosphohistidine phosphatase 1 |
| chr25_-_23583101 | 0.01 |
ENSDART00000149107
ENSDART00000103704 ENSDART00000184903 |
nap1l4a
|
nucleosome assembly protein 1-like 4a |
| chr15_-_35126632 | 0.01 |
ENSDART00000187324
ENSDART00000037389 |
zgc:77118
zgc:55413
|
zgc:77118 zgc:55413 |
| chr3_+_37083765 | 0.00 |
ENSDART00000125611
|
retreg3
|
reticulophagy regulator family member 3 |
Network of associatons between targets according to the STRING database.
First level regulatory network of eomesa
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.1 | GO:0010873 | regulation of cholesterol esterification(GO:0010872) positive regulation of cholesterol esterification(GO:0010873) |
| 0.1 | 0.8 | GO:0071305 | vitamin D3 metabolic process(GO:0070640) cellular response to vitamin D(GO:0071305) |
| 0.1 | 0.2 | GO:0055109 | invagination involved in gastrulation with mouth forming second(GO:0055109) morphogenesis of an epithelial fold(GO:0060571) |
| 0.0 | 0.2 | GO:0038107 | nodal signaling pathway involved in determination of left/right asymmetry(GO:0038107) regulation of nodal signaling pathway involved in determination of left/right asymmetry(GO:1900145) |
| 0.0 | 0.2 | GO:0010828 | positive regulation of glucose transport(GO:0010828) |
| 0.0 | 0.5 | GO:0006607 | NLS-bearing protein import into nucleus(GO:0006607) |
| 0.0 | 0.2 | GO:0033278 | cell proliferation in midbrain(GO:0033278) |
| 0.0 | 0.1 | GO:0002184 | cytoplasmic translational termination(GO:0002184) |
| 0.0 | 0.5 | GO:0006208 | pyrimidine nucleobase catabolic process(GO:0006208) |
| 0.0 | 0.4 | GO:0007198 | adenylate cyclase-inhibiting serotonin receptor signaling pathway(GO:0007198) |
| 0.0 | 0.3 | GO:0009303 | rRNA transcription(GO:0009303) |
| 0.0 | 0.2 | GO:0071300 | cellular response to retinoic acid(GO:0071300) |
| 0.0 | 0.5 | GO:0003323 | type B pancreatic cell development(GO:0003323) |
| 0.0 | 0.3 | GO:1990399 | sensory epithelium regeneration(GO:0070654) epithelium regeneration(GO:1990399) |
| 0.0 | 0.1 | GO:1990798 | pancreas regeneration(GO:1990798) |
| 0.0 | 0.0 | GO:0034421 | post-translational protein acetylation(GO:0034421) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.1 | GO:0042627 | chylomicron(GO:0042627) |
| 0.0 | 0.2 | GO:0005614 | interstitial matrix(GO:0005614) |
| 0.0 | 0.1 | GO:0018444 | translation release factor complex(GO:0018444) |
| 0.0 | 0.3 | GO:0005736 | DNA-directed RNA polymerase I complex(GO:0005736) |
| 0.0 | 0.1 | GO:0019008 | molybdopterin synthase complex(GO:0019008) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.1 | GO:0060228 | phosphatidylcholine-sterol O-acyltransferase activator activity(GO:0060228) |
| 0.0 | 0.1 | GO:1990825 | sequence-specific mRNA binding(GO:1990825) |
| 0.0 | 0.5 | GO:0004157 | dihydropyrimidinase activity(GO:0004157) |
| 0.0 | 0.3 | GO:0016803 | ether hydrolase activity(GO:0016803) |
| 0.0 | 1.3 | GO:0015459 | potassium channel regulator activity(GO:0015459) |
| 0.0 | 0.5 | GO:0061608 | nuclear import signal receptor activity(GO:0061608) |
| 0.0 | 0.4 | GO:0051378 | amine binding(GO:0043176) serotonin binding(GO:0051378) |
| 0.0 | 0.2 | GO:0004711 | ribosomal protein S6 kinase activity(GO:0004711) |
| 0.0 | 0.2 | GO:0061665 | SUMO ligase activity(GO:0061665) |
| 0.0 | 4.7 | GO:0030246 | carbohydrate binding(GO:0030246) |
| 0.0 | 0.1 | GO:0030366 | molybdopterin synthase activity(GO:0030366) |
| 0.0 | 0.4 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.0 | 0.2 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 0.0 | 0.1 | GO:0016519 | gastric inhibitory peptide receptor activity(GO:0016519) |
| 0.0 | 0.9 | GO:0016712 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced flavin or flavoprotein as one donor, and incorporation of one atom of oxygen(GO:0016712) |
| 0.0 | 0.1 | GO:0004865 | protein serine/threonine phosphatase inhibitor activity(GO:0004865) |
| 0.0 | 0.4 | GO:0019003 | GDP binding(GO:0019003) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.3 | PID FOXO PATHWAY | FoxO family signaling |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | REACTOME SEROTONIN RECEPTORS | Genes involved in Serotonin receptors |
| 0.1 | 0.5 | REACTOME PYRIMIDINE CATABOLISM | Genes involved in Pyrimidine catabolism |
| 0.0 | 0.8 | REACTOME CYTOCHROME P450 ARRANGED BY SUBSTRATE TYPE | Genes involved in Cytochrome P450 - arranged by substrate type |
| 0.0 | 0.2 | REACTOME FGFR1 LIGAND BINDING AND ACTIVATION | Genes involved in FGFR1 ligand binding and activation |