Project
PRJNA207719: Tissue specific transcriptome profiling
Navigation
Downloads
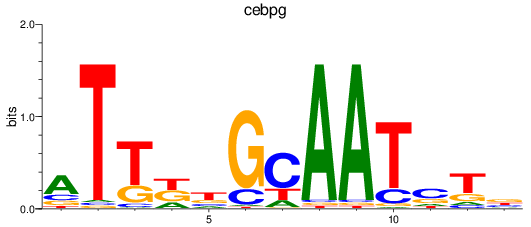
Results for cebpg
Z-value: 1.70
Transcription factors associated with cebpg
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
cebpg
|
ENSDARG00000036073 | CCAAT enhancer binding protein gamma |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| cebpg | dr11_v1_chr7_+_38089650_38089650 | -0.73 | 1.6e-01 | Click! |
Activity profile of cebpg motif
Sorted Z-values of cebpg motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_-_10071422 | 2.13 |
ENSDART00000135522
ENSDART00000033118 |
fga
|
fibrinogen alpha chain |
| chr5_-_63509581 | 1.92 |
ENSDART00000097325
|
c5
|
complement component 5 |
| chr22_-_23668356 | 1.83 |
ENSDART00000167106
ENSDART00000159622 ENSDART00000163228 |
cfh
|
complement factor H |
| chr22_-_26175237 | 1.51 |
ENSDART00000108737
|
c3b.2
|
complement component c3b, tandem duplicate 2 |
| chr22_-_24757785 | 1.49 |
ENSDART00000078225
|
vtg5
|
vitellogenin 5 |
| chr8_-_50147948 | 1.48 |
ENSDART00000149010
|
hp
|
haptoglobin |
| chr9_+_38292947 | 1.44 |
ENSDART00000146663
|
tfcp2l1
|
transcription factor CP2-like 1 |
| chr5_+_45677781 | 1.39 |
ENSDART00000163120
ENSDART00000126537 |
gc
|
group-specific component (vitamin D binding protein) |
| chr20_-_40755614 | 1.30 |
ENSDART00000061247
|
cx32.3
|
connexin 32.3 |
| chr12_+_46462090 | 1.29 |
ENSDART00000130748
|
BX005305.2
|
|
| chr12_+_46404307 | 1.23 |
ENSDART00000185011
|
BX005305.4
|
|
| chr12_+_46386983 | 1.22 |
ENSDART00000183982
|
BX005305.3
|
Danio rerio legumain (LOC100005356), mRNA. |
| chr16_+_23984179 | 1.17 |
ENSDART00000175879
|
apoc2
|
apolipoprotein C-II |
| chr6_-_609880 | 1.16 |
ENSDART00000149248
ENSDART00000148867 ENSDART00000149414 ENSDART00000148552 ENSDART00000148391 |
lgals2b
|
lectin, galactoside-binding, soluble, 2b |
| chr12_-_22524388 | 1.10 |
ENSDART00000020942
|
shbg
|
sex hormone-binding globulin |
| chr22_-_26236188 | 1.09 |
ENSDART00000162640
ENSDART00000167169 ENSDART00000138595 |
c3b.1
|
complement component c3b, tandem duplicate 1 |
| chr20_+_23440632 | 1.09 |
ENSDART00000180685
ENSDART00000042820 |
si:dkey-90m5.4
|
si:dkey-90m5.4 |
| chr12_+_46512881 | 1.04 |
ENSDART00000105454
|
CU855711.1
|
|
| chr5_-_69944084 | 1.02 |
ENSDART00000188557
ENSDART00000127782 |
ugt2a4
|
UDP glucuronosyltransferase 2 family, polypeptide A4 |
| chr3_-_31254979 | 0.99 |
ENSDART00000130280
|
apnl
|
actinoporin-like protein |
| chr7_+_56577906 | 0.99 |
ENSDART00000184023
|
hp
|
haptoglobin |
| chr7_-_16596938 | 0.97 |
ENSDART00000134548
|
e2f8
|
E2F transcription factor 8 |
| chr2_+_11029138 | 0.94 |
ENSDART00000138737
ENSDART00000081058 ENSDART00000153662 |
acot11a
|
acyl-CoA thioesterase 11a |
| chr12_+_46483618 | 0.92 |
ENSDART00000186970
|
BX005305.5
|
|
| chr19_-_15192638 | 0.90 |
ENSDART00000048151
|
phactr4a
|
phosphatase and actin regulator 4a |
| chr7_-_16597130 | 0.89 |
ENSDART00000144118
|
e2f8
|
E2F transcription factor 8 |
| chr9_+_426392 | 0.88 |
ENSDART00000172515
|
bzw1b
|
basic leucine zipper and W2 domains 1b |
| chr12_+_46425800 | 0.86 |
ENSDART00000191965
|
BX005305.1
|
|
| chr12_+_46443477 | 0.84 |
ENSDART00000191873
|
BX005305.1
|
|
| chr16_+_48753664 | 0.76 |
ENSDART00000155148
|
si:ch73-31d8.2
|
si:ch73-31d8.2 |
| chr24_-_33284945 | 0.74 |
ENSDART00000155429
ENSDART00000112845 |
zgc:195173
|
zgc:195173 |
| chr14_-_11507211 | 0.71 |
ENSDART00000186873
ENSDART00000109181 ENSDART00000186166 ENSDART00000186986 |
zgc:174917
|
zgc:174917 |
| chr24_+_38671054 | 0.71 |
ENSDART00000154214
|
si:ch73-70c5.1
|
si:ch73-70c5.1 |
| chr8_-_51293265 | 0.71 |
ENSDART00000127875
ENSDART00000181145 |
bmp1a
|
bone morphogenetic protein 1a |
| chr19_-_2822372 | 0.68 |
ENSDART00000109130
ENSDART00000122385 |
recql4
|
RecQ helicase-like 4 |
| chr1_+_45056371 | 0.65 |
ENSDART00000073689
ENSDART00000167309 |
btr01
|
bloodthirsty-related gene family, member 1 |
| chr1_+_14253118 | 0.62 |
ENSDART00000161996
|
cxcl8a
|
chemokine (C-X-C motif) ligand 8a |
| chr17_-_10838434 | 0.61 |
ENSDART00000064597
|
lgals3b
|
lectin, galactoside binding soluble 3b |
| chr2_+_42247560 | 0.59 |
ENSDART00000101230
ENSDART00000143094 |
ftr06
|
finTRIM family, member 6 |
| chr14_-_43602968 | 0.58 |
ENSDART00000108825
|
adh5
|
alcohol dehydrogenase 5 |
| chr17_+_51744450 | 0.57 |
ENSDART00000190955
ENSDART00000149807 |
odc1
|
ornithine decarboxylase 1 |
| chr4_+_9508505 | 0.57 |
ENSDART00000080842
|
kitlgb
|
kit ligand b |
| chr25_-_29415369 | 0.56 |
ENSDART00000110774
ENSDART00000019183 |
ugt5a2
ugt5a1
|
UDP glucuronosyltransferase 5 family, polypeptide A2 UDP glucuronosyltransferase 5 family, polypeptide A1 |
| chr12_-_35949936 | 0.56 |
ENSDART00000192583
|
AL954682.1
|
|
| chr10_-_22803740 | 0.56 |
ENSDART00000079469
ENSDART00000187968 ENSDART00000122543 |
pcolcea
|
procollagen C-endopeptidase enhancer a |
| chr9_-_40664923 | 0.56 |
ENSDART00000135134
|
bard1
|
BRCA1 associated RING domain 1 |
| chr5_-_23783739 | 0.55 |
ENSDART00000139502
|
GBGT1 (1 of many)
|
si:ch211-287c22.1 |
| chr24_-_38644937 | 0.54 |
ENSDART00000170194
|
slc6a16b
|
solute carrier family 6, member 16b |
| chr14_-_38929885 | 0.53 |
ENSDART00000148737
|
btk
|
Bruton agammaglobulinemia tyrosine kinase |
| chr17_+_51743908 | 0.53 |
ENSDART00000149039
ENSDART00000148869 |
odc1
|
ornithine decarboxylase 1 |
| chr18_+_13300768 | 0.52 |
ENSDART00000137506
|
si:ch211-260p9.3
|
si:ch211-260p9.3 |
| chr6_-_39649504 | 0.52 |
ENSDART00000179960
ENSDART00000190951 |
larp4ab
|
La ribonucleoprotein domain family, member 4Ab |
| chr1_+_31674297 | 0.52 |
ENSDART00000044214
|
wbp1lb
|
WW domain binding protein 1-like b |
| chr7_+_49862837 | 0.51 |
ENSDART00000174315
|
slc1a2a
|
solute carrier family 1 (glial high affinity glutamate transporter), member 2a |
| chr10_-_23099809 | 0.50 |
ENSDART00000148333
ENSDART00000079703 ENSDART00000162444 |
nle1
|
notchless homolog 1 (Drosophila) |
| chr15_-_29348212 | 0.50 |
ENSDART00000133117
|
tsku
|
tsukushi small leucine rich proteoglycan homolog (Xenopus laevis) |
| chr10_-_7785930 | 0.49 |
ENSDART00000043961
ENSDART00000111058 |
mpx
|
myeloid-specific peroxidase |
| chr16_-_42004544 | 0.49 |
ENSDART00000034544
|
caspa
|
caspase a |
| chr12_-_16990896 | 0.49 |
ENSDART00000152402
|
ifit12
|
interferon-induced protein with tetratricopeptide repeats 12 |
| chr2_-_30182353 | 0.48 |
ENSDART00000019149
|
rpl7
|
ribosomal protein L7 |
| chr19_+_16015881 | 0.48 |
ENSDART00000187135
|
ctps1a
|
CTP synthase 1a |
| chr21_+_17051478 | 0.47 |
ENSDART00000047201
ENSDART00000161650 ENSDART00000167298 |
atp2a2a
|
ATPase sarcoplasmic/endoplasmic reticulum Ca2+ transporting 2a |
| chr19_-_20403318 | 0.47 |
ENSDART00000136826
|
dazl
|
deleted in azoospermia-like |
| chr9_-_53062083 | 0.46 |
ENSDART00000122155
|
zmp:0000000936
|
zmp:0000000936 |
| chr3_+_54581987 | 0.46 |
ENSDART00000018071
|
eif3g
|
eukaryotic translation initiation factor 3, subunit G |
| chr8_+_22955478 | 0.46 |
ENSDART00000099911
|
zgc:136605
|
zgc:136605 |
| chr8_-_4475908 | 0.46 |
ENSDART00000191027
|
si:ch211-166a6.5
|
si:ch211-166a6.5 |
| chr13_-_50463938 | 0.46 |
ENSDART00000083857
|
ccnj
|
cyclin J |
| chr3_+_43086548 | 0.45 |
ENSDART00000163579
|
si:dkey-43p13.5
|
si:dkey-43p13.5 |
| chr2_+_37971353 | 0.45 |
ENSDART00000165085
|
apol1
|
apolipoprotein L, 1 |
| chr10_+_15088534 | 0.45 |
ENSDART00000142865
|
si:ch211-95j8.3
|
si:ch211-95j8.3 |
| chr15_-_21014270 | 0.44 |
ENSDART00000154019
|
si:ch211-212c13.10
|
si:ch211-212c13.10 |
| chr23_+_21405201 | 0.44 |
ENSDART00000144409
|
iffo2a
|
intermediate filament family orphan 2a |
| chr21_-_40174647 | 0.44 |
ENSDART00000183738
ENSDART00000076840 ENSDART00000145109 |
slco2b1
|
solute carrier organic anion transporter family, member 2B1 |
| chr8_-_53488832 | 0.44 |
ENSDART00000191801
|
chdh
|
choline dehydrogenase |
| chr19_-_34979837 | 0.44 |
ENSDART00000044838
|
ndrg1a
|
N-myc downstream regulated 1a |
| chr19_+_16016038 | 0.44 |
ENSDART00000131319
|
ctps1a
|
CTP synthase 1a |
| chr5_-_24238733 | 0.43 |
ENSDART00000138170
|
plscr3a
|
phospholipid scramblase 3a |
| chr7_+_33152723 | 0.42 |
ENSDART00000132658
|
si:ch211-194p6.10
|
si:ch211-194p6.10 |
| chr15_-_3282220 | 0.42 |
ENSDART00000092942
|
slc25a15a
|
solute carrier family 25 (mitochondrial carrier; ornithine transporter) member 15a |
| chr23_+_27675581 | 0.42 |
ENSDART00000127198
|
rps26
|
ribosomal protein S26 |
| chr2_+_4207209 | 0.39 |
ENSDART00000157903
ENSDART00000166476 |
gata6
|
GATA binding protein 6 |
| chr7_-_58289847 | 0.39 |
ENSDART00000012822
|
CR354540.1
|
|
| chr19_+_7045033 | 0.39 |
ENSDART00000146579
|
mhc1uka
|
major histocompatibility complex class I UKA |
| chr13_-_24257631 | 0.38 |
ENSDART00000146524
|
urb2
|
URB2 ribosome biogenesis 2 homolog (S. cerevisiae) |
| chr8_+_22955262 | 0.38 |
ENSDART00000193806
|
zgc:136605
|
zgc:136605 |
| chr9_-_9419704 | 0.38 |
ENSDART00000138996
|
si:ch211-214p13.9
|
si:ch211-214p13.9 |
| chr12_+_22657925 | 0.38 |
ENSDART00000153048
|
si:dkey-219e21.4
|
si:dkey-219e21.4 |
| chr20_+_29436601 | 0.38 |
ENSDART00000136804
|
fmn1
|
formin 1 |
| chr17_+_43908428 | 0.38 |
ENSDART00000180332
|
msh4
|
mutS homolog 4 |
| chr5_+_33519943 | 0.37 |
ENSDART00000131316
|
ms4a17c.1
|
membrane-spanning 4-domains, subfamily A, member 17C.1 |
| chr6_-_39344259 | 0.37 |
ENSDART00000104074
|
zgc:158846
|
zgc:158846 |
| chr22_-_10487490 | 0.36 |
ENSDART00000064798
|
aspn
|
asporin (LRR class 1) |
| chr13_-_14929236 | 0.36 |
ENSDART00000020576
|
cdc25b
|
cell division cycle 25B |
| chr22_+_6697927 | 0.35 |
ENSDART00000135015
|
CT583625.3
|
|
| chr1_-_28885919 | 0.35 |
ENSDART00000152182
|
poglut1
|
protein O-glucosyltransferase 1 |
| chr23_-_31763753 | 0.35 |
ENSDART00000053399
|
aldh8a1
|
aldehyde dehydrogenase 8 family, member A1 |
| chr9_-_34885902 | 0.34 |
ENSDART00000137504
|
p2ry8
|
P2Y receptor family member 8 |
| chr7_-_69121896 | 0.34 |
ENSDART00000130227
|
crispld2
|
cysteine-rich secretory protein LCCL domain containing 2 |
| chr9_-_12874774 | 0.34 |
ENSDART00000131385
|
ankzf1
|
ankyrin repeat and zinc finger domain containing 1 |
| chr3_-_23574622 | 0.34 |
ENSDART00000176012
|
igf2bp1
|
insulin-like growth factor 2 mRNA binding protein 1 |
| chr6_-_10708960 | 0.33 |
ENSDART00000157704
|
si:dkey-34m19.3
|
si:dkey-34m19.3 |
| chr13_-_24260609 | 0.33 |
ENSDART00000138747
|
urb2
|
URB2 ribosome biogenesis 2 homolog (S. cerevisiae) |
| chr24_-_26485098 | 0.32 |
ENSDART00000135496
ENSDART00000009609 ENSDART00000133782 ENSDART00000141029 ENSDART00000113739 |
eif5a
|
eukaryotic translation initiation factor 5A |
| chr10_-_41980797 | 0.32 |
ENSDART00000076575
|
rhof
|
ras homolog family member F |
| chr23_-_1658980 | 0.31 |
ENSDART00000186227
|
CU693481.1
|
|
| chr10_+_23099890 | 0.31 |
ENSDART00000135890
|
LTN1
|
si:dkey-175g6.5 |
| chr1_+_58067815 | 0.30 |
ENSDART00000156678
|
si:ch211-114l13.4
|
si:ch211-114l13.4 |
| chr2_-_40199780 | 0.29 |
ENSDART00000113901
|
ccl34a.4
|
chemokine (C-C motif) ligand 34a, duplicate 4 |
| chr1_+_17892944 | 0.28 |
ENSDART00000013021
|
tlr3
|
toll-like receptor 3 |
| chr21_+_26535034 | 0.28 |
ENSDART00000180709
|
vegfbb
|
vascular endothelial growth factor Bb |
| chr20_-_52271262 | 0.27 |
ENSDART00000135463
|
rhpn1
|
rhophilin, Rho GTPase binding protein 1 |
| chr23_-_40814080 | 0.27 |
ENSDART00000135713
|
si:dkeyp-27c8.1
|
si:dkeyp-27c8.1 |
| chr5_+_54400971 | 0.27 |
ENSDART00000169695
|
bspry
|
B-box and SPRY domain containing |
| chr19_-_22507715 | 0.27 |
ENSDART00000160153
|
pleca
|
plectin a |
| chr10_+_15310811 | 0.26 |
ENSDART00000136239
|
kcnv2a
|
potassium channel, subfamily V, member 2a |
| chr4_+_42604252 | 0.26 |
ENSDART00000184850
|
CR925768.1
|
|
| chr3_-_38745598 | 0.26 |
ENSDART00000127483
|
si:ch211-198m17.1
|
si:ch211-198m17.1 |
| chr23_-_16843649 | 0.25 |
ENSDART00000129971
|
si:dkey-63j12.4
|
si:dkey-63j12.4 |
| chr9_+_2522797 | 0.25 |
ENSDART00000186786
ENSDART00000147034 |
gpr155a
|
G protein-coupled receptor 155a |
| chr4_-_20292821 | 0.25 |
ENSDART00000136069
ENSDART00000192504 |
cacna2d4a
|
calcium channel, voltage-dependent, alpha 2/delta subunit 4a |
| chr8_-_36140405 | 0.25 |
ENSDART00000182806
|
CT583723.2
|
|
| chr13_-_30996072 | 0.24 |
ENSDART00000181661
|
wdfy4
|
WDFY family member 4 |
| chr20_-_52271015 | 0.24 |
ENSDART00000074307
|
rhpn1
|
rhophilin, Rho GTPase binding protein 1 |
| chr18_+_32844439 | 0.24 |
ENSDART00000166038
|
olfcg1
|
olfactory receptor C family, g1 |
| chr3_+_34986837 | 0.23 |
ENSDART00000190341
|
smarce1
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily e, member 1 |
| chr4_-_37479395 | 0.23 |
ENSDART00000168299
ENSDART00000157864 |
si:dkey-106c17.2
|
si:dkey-106c17.2 |
| chr1_+_30551777 | 0.23 |
ENSDART00000112778
|
gpr183b
|
G protein-coupled receptor 183b |
| chr21_+_4540127 | 0.22 |
ENSDART00000043431
|
nup188
|
nucleoporin 188 |
| chr12_-_33770299 | 0.21 |
ENSDART00000189849
|
llgl2
|
lethal giant larvae homolog 2 (Drosophila) |
| chr4_+_30776883 | 0.20 |
ENSDART00000169519
|
si:dkey-11d20.1
|
si:dkey-11d20.1 |
| chr20_-_5369105 | 0.20 |
ENSDART00000114316
|
sptlc2b
|
serine palmitoyltransferase, long chain base subunit 2b |
| chr3_-_34547000 | 0.20 |
ENSDART00000166623
|
sept9a
|
septin 9a |
| chr6_-_345503 | 0.20 |
ENSDART00000168901
|
pde6ha
|
phosphodiesterase 6H, cGMP-specific, cone, gamma, paralog a |
| chr5_+_51079504 | 0.20 |
ENSDART00000097466
|
fam169aa
|
family with sequence similarity 169, member Aa |
| chr20_+_25879826 | 0.19 |
ENSDART00000018519
|
zgc:153896
|
zgc:153896 |
| chr25_-_16076257 | 0.19 |
ENSDART00000140780
|
ovch2
|
ovochymase 2 |
| chr14_+_7377552 | 0.19 |
ENSDART00000142158
ENSDART00000141471 |
hars
|
histidyl-tRNA synthetase |
| chr23_-_11130683 | 0.19 |
ENSDART00000181189
|
cntn3a.2
|
contactin 3a, tandem duplicate 2 |
| chr22_+_6587280 | 0.19 |
ENSDART00000106185
|
CR352226.7
|
|
| chr4_-_73227710 | 0.18 |
ENSDART00000193165
|
LO018260.3
|
|
| chr24_-_25184553 | 0.18 |
ENSDART00000166917
|
plcxd2
|
phosphatidylinositol-specific phospholipase C, X domain containing 2 |
| chr22_-_10050856 | 0.17 |
ENSDART00000144811
|
zgc:174564
|
zgc:174564 |
| chr21_-_11646878 | 0.17 |
ENSDART00000162426
ENSDART00000135937 ENSDART00000131448 ENSDART00000148097 ENSDART00000133443 |
cast
|
calpastatin |
| chr5_+_4298636 | 0.17 |
ENSDART00000100061
|
prdx4
|
peroxiredoxin 4 |
| chr22_+_6711549 | 0.16 |
ENSDART00000132094
|
CT583625.4
|
|
| chr7_-_13884610 | 0.16 |
ENSDART00000006897
|
rlbp1a
|
retinaldehyde binding protein 1a |
| chr22_-_10051401 | 0.15 |
ENSDART00000106300
ENSDART00000175910 |
zgc:174564
BX324216.3
|
zgc:174564 |
| chr8_-_21988833 | 0.15 |
ENSDART00000167708
|
nphp4
|
nephronophthisis 4 |
| chr3_+_40809011 | 0.15 |
ENSDART00000033713
|
arpc1b
|
actin related protein 2/3 complex, subunit 1B |
| chr4_+_70563225 | 0.14 |
ENSDART00000159508
|
si:dkey-11d20.1
|
si:dkey-11d20.1 |
| chr3_+_22242269 | 0.14 |
ENSDART00000168970
|
scn4ab
|
sodium channel, voltage-gated, type IV, alpha, b |
| chr4_-_17793152 | 0.14 |
ENSDART00000134080
|
mybpc1
|
myosin binding protein C, slow type |
| chr21_-_16219400 | 0.13 |
ENSDART00000124871
|
tnfrsfa
|
tumor necrosis factor receptor superfamily, member a |
| chr24_+_16905188 | 0.13 |
ENSDART00000066760
|
cct5
|
chaperonin containing TCP1, subunit 5 (epsilon) |
| chr11_-_31172276 | 0.13 |
ENSDART00000171520
|
si:dkey-238i5.3
|
si:dkey-238i5.3 |
| chr17_+_28670132 | 0.13 |
ENSDART00000076344
ENSDART00000164981 ENSDART00000182851 |
hectd1
|
HECT domain containing 1 |
| chr11_+_43419809 | 0.13 |
ENSDART00000172982
|
slc29a1b
|
solute carrier family 29 (equilibrative nucleoside transporter), member 1b |
| chr4_-_9549693 | 0.13 |
ENSDART00000160242
|
FQ377934.1
|
|
| chr17_-_20218357 | 0.13 |
ENSDART00000155990
ENSDART00000155632 ENSDART00000156540 ENSDART00000063523 |
mgmt
|
O-6-methylguanine-DNA methyltransferase |
| chr1_+_2260407 | 0.12 |
ENSDART00000058876
|
kpnb3
|
karyopherin (importin) beta 3 |
| chr6_+_10333920 | 0.12 |
ENSDART00000151667
ENSDART00000151477 |
cobll1a
|
cordon-bleu WH2 repeat protein-like 1a |
| chr9_-_32300611 | 0.12 |
ENSDART00000127938
|
hspd1
|
heat shock 60 protein 1 |
| chr5_-_58840971 | 0.11 |
ENSDART00000050932
|
tmem136b
|
transmembrane protein 136b |
| chr12_+_25775734 | 0.11 |
ENSDART00000024415
ENSDART00000149198 |
epas1a
|
endothelial PAS domain protein 1a |
| chr9_-_21838045 | 0.11 |
ENSDART00000147471
|
acod1
|
aconitate decarboxylase 1 |
| chr17_-_14613711 | 0.11 |
ENSDART00000157345
|
sdsl
|
serine dehydratase-like |
| chr14_-_32089117 | 0.09 |
ENSDART00000158014
|
si:ch211-69b22.5
|
si:ch211-69b22.5 |
| chr10_-_29827000 | 0.09 |
ENSDART00000131418
|
zpr1
|
ZPR1 zinc finger |
| chr11_-_34783938 | 0.08 |
ENSDART00000135725
ENSDART00000039847 |
chchd4a
|
coiled-coil-helix-coiled-coil-helix domain containing 4a |
| chr5_+_41510387 | 0.08 |
ENSDART00000023779
|
vcp
|
valosin containing protein |
| chr10_-_28193642 | 0.08 |
ENSDART00000019050
|
rps6kb1a
|
ribosomal protein S6 kinase b, polypeptide 1a |
| chr18_-_37252036 | 0.08 |
ENSDART00000132230
|
six5
|
SIX homeobox 5 |
| chr1_+_55752593 | 0.08 |
ENSDART00000108838
ENSDART00000134770 |
tecrb
|
trans-2,3-enoyl-CoA reductase b |
| chr21_+_20237006 | 0.08 |
ENSDART00000132853
|
si:dkey-247m21.3
|
si:dkey-247m21.3 |
| chr4_-_20313810 | 0.07 |
ENSDART00000136350
|
dcp1b
|
decapping mRNA 1B |
| chr5_-_25236340 | 0.07 |
ENSDART00000162774
|
abca2
|
ATP-binding cassette, sub-family A (ABC1), member 2 |
| chr6_+_8315050 | 0.06 |
ENSDART00000189987
|
gcdha
|
glutaryl-CoA dehydrogenase a |
| chr14_-_45551572 | 0.06 |
ENSDART00000111410
|
ganab
|
glucosidase, alpha; neutral AB |
| chr20_-_32186886 | 0.06 |
ENSDART00000170279
|
si:ch211-51a19.5
|
si:ch211-51a19.5 |
| chr9_-_32300783 | 0.06 |
ENSDART00000078596
|
hspd1
|
heat shock 60 protein 1 |
| chr18_+_20869923 | 0.05 |
ENSDART00000138471
|
pgpep1l
|
pyroglutamyl-peptidase I-like |
| chr15_+_28355023 | 0.05 |
ENSDART00000122159
|
si:dkey-118k5.3
|
si:dkey-118k5.3 |
| chr1_+_8694196 | 0.04 |
ENSDART00000025604
|
zgc:77849
|
zgc:77849 |
| chr3_+_23703704 | 0.04 |
ENSDART00000024256
|
hoxb6a
|
homeobox B6a |
| chr16_+_49601838 | 0.04 |
ENSDART00000168570
ENSDART00000159236 |
si:dkey-82o10.4
|
si:dkey-82o10.4 |
| chr9_+_32301017 | 0.04 |
ENSDART00000127916
ENSDART00000183298 ENSDART00000143103 |
hspe1
|
heat shock 10 protein 1 |
| chr1_+_41131481 | 0.03 |
ENSDART00000145272
|
lrpap1
|
low density lipoprotein receptor-related protein associated protein 1 |
| chr18_-_40905901 | 0.03 |
ENSDART00000064848
|
pglyrp5
|
peptidoglycan recognition protein 5 |
| chr14_-_5642371 | 0.03 |
ENSDART00000183859
ENSDART00000054876 |
npm1b
|
nucleophosmin 1b |
| chr6_+_27452289 | 0.03 |
ENSDART00000186265
|
sned1
|
sushi, nidogen and EGF-like domains 1 |
| chr15_-_23793641 | 0.03 |
ENSDART00000122891
|
tmem97
|
transmembrane protein 97 |
| chr8_-_2602572 | 0.02 |
ENSDART00000110482
|
zdhhc12a
|
zinc finger, DHHC-type containing 12a |
| chr15_+_7064819 | 0.02 |
ENSDART00000155268
|
pik3cb
|
phosphatidylinositol-4,5-bisphosphate 3-kinase, catalytic subunit beta |
| chr13_+_30903816 | 0.02 |
ENSDART00000191727
|
ercc6
|
excision repair cross-complementation group 6 |
| chr25_-_3087556 | 0.02 |
ENSDART00000193249
|
best1
|
bestrophin 1 |
| chr19_-_9662958 | 0.02 |
ENSDART00000041094
|
clcn1a
|
chloride channel, voltage-sensitive 1a |
| chr2_-_44746723 | 0.01 |
ENSDART00000041806
|
acsm3
|
acyl-CoA synthetase medium chain family member 3 |
| chr9_-_4856767 | 0.01 |
ENSDART00000016814
ENSDART00000138015 |
fmnl2a
|
formin-like 2a |
| chr18_+_34225520 | 0.00 |
ENSDART00000126115
|
v2rl1
|
vomeronasal 2 receptor, l1 |
Network of associatons between targets according to the STRING database.
First level regulatory network of cebpg
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.9 | GO:0033301 | cell cycle comprising mitosis without cytokinesis(GO:0033301) |
| 0.2 | 0.9 | GO:0044210 | 'de novo' CTP biosynthetic process(GO:0044210) |
| 0.2 | 0.6 | GO:0002676 | granuloma formation(GO:0002432) chronic inflammatory response(GO:0002544) regulation of granuloma formation(GO:0002631) regulation of chronic inflammatory response(GO:0002676) |
| 0.2 | 1.0 | GO:0051883 | killing of cells of other organism(GO:0031640) disruption of cells of other organism(GO:0044364) disruption of cells of other organism involved in symbiotic interaction(GO:0051818) killing of cells in other organism involved in symbiotic interaction(GO:0051883) |
| 0.1 | 0.4 | GO:0019695 | choline metabolic process(GO:0019695) |
| 0.1 | 2.1 | GO:0072378 | blood coagulation, fibrin clot formation(GO:0072378) |
| 0.1 | 0.4 | GO:0000066 | mitochondrial ornithine transport(GO:0000066) mitochondrial L-ornithine transmembrane transport(GO:1990575) |
| 0.1 | 0.9 | GO:2001045 | negative regulation of integrin-mediated signaling pathway(GO:2001045) |
| 0.1 | 0.5 | GO:0003418 | chondrocyte differentiation involved in endochondral bone morphogenesis(GO:0003413) growth plate cartilage chondrocyte differentiation(GO:0003418) |
| 0.1 | 1.1 | GO:0033387 | putrescine biosynthetic process from ornithine(GO:0033387) |
| 0.1 | 0.6 | GO:0048245 | eosinophil chemotaxis(GO:0048245) eosinophil migration(GO:0072677) |
| 0.1 | 0.7 | GO:0032964 | collagen biosynthetic process(GO:0032964) |
| 0.1 | 0.3 | GO:0097053 | L-kynurenine metabolic process(GO:0097052) L-kynurenine catabolic process(GO:0097053) |
| 0.1 | 0.1 | GO:0006549 | isoleucine metabolic process(GO:0006549) isoleucine biosynthetic process(GO:0009097) |
| 0.1 | 0.7 | GO:0001561 | fatty acid alpha-oxidation(GO:0001561) |
| 0.1 | 0.3 | GO:0034138 | chemokine production(GO:0032602) toll-like receptor 3 signaling pathway(GO:0034138) |
| 0.1 | 0.6 | GO:0046294 | formaldehyde metabolic process(GO:0046292) formaldehyde catabolic process(GO:0046294) |
| 0.1 | 0.4 | GO:0043011 | myeloid dendritic cell activation(GO:0001773) myeloid dendritic cell differentiation(GO:0043011) |
| 0.1 | 0.5 | GO:0060965 | negative regulation of posttranscriptional gene silencing(GO:0060149) negative regulation of gene silencing by miRNA(GO:0060965) negative regulation of gene silencing by RNA(GO:0060967) |
| 0.1 | 0.4 | GO:0045448 | regulation of mitotic cell cycle, embryonic(GO:0009794) mitotic cell cycle, embryonic(GO:0045448) |
| 0.1 | 0.3 | GO:0045041 | protein import into mitochondrial intermembrane space(GO:0045041) |
| 0.1 | 0.4 | GO:0032287 | peripheral nervous system myelin maintenance(GO:0032287) |
| 0.1 | 1.2 | GO:0044872 | lipoprotein transport(GO:0042953) lipoprotein localization(GO:0044872) |
| 0.1 | 0.6 | GO:0061484 | hematopoietic stem cell homeostasis(GO:0061484) |
| 0.0 | 1.4 | GO:0051180 | vitamin transport(GO:0051180) |
| 0.0 | 0.3 | GO:0045901 | positive regulation of translational elongation(GO:0045901) positive regulation of translational termination(GO:0045905) |
| 0.0 | 0.6 | GO:2000377 | regulation of reactive oxygen species metabolic process(GO:2000377) |
| 0.0 | 0.7 | GO:0050994 | regulation of lipid catabolic process(GO:0050994) |
| 0.0 | 0.7 | GO:0006268 | DNA unwinding involved in DNA replication(GO:0006268) |
| 0.0 | 0.2 | GO:0010719 | photoreceptor cell morphogenesis(GO:0008594) negative regulation of epithelial to mesenchymal transition(GO:0010719) |
| 0.0 | 0.5 | GO:0048312 | intracellular distribution of mitochondria(GO:0048312) |
| 0.0 | 1.5 | GO:0032355 | response to estradiol(GO:0032355) |
| 0.0 | 0.4 | GO:0000710 | meiotic mismatch repair(GO:0000710) |
| 0.0 | 0.0 | GO:0048261 | negative regulation of receptor-mediated endocytosis(GO:0048261) |
| 0.0 | 0.1 | GO:0036462 | TRAIL-activated apoptotic signaling pathway(GO:0036462) |
| 0.0 | 0.2 | GO:0045745 | positive regulation of G-protein coupled receptor protein signaling pathway(GO:0045745) |
| 0.0 | 0.4 | GO:0015721 | bile acid and bile salt transport(GO:0015721) |
| 0.0 | 0.2 | GO:2000096 | positive regulation of Wnt signaling pathway, planar cell polarity pathway(GO:2000096) |
| 0.0 | 0.5 | GO:0000463 | maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000463) |
| 0.0 | 0.1 | GO:0051230 | endosome to lysosome transport via multivesicular body sorting pathway(GO:0032510) mitotic spindle disassembly(GO:0051228) spindle disassembly(GO:0051230) |
| 0.0 | 0.2 | GO:0097341 | inhibition of cysteine-type endopeptidase activity(GO:0097340) zymogen inhibition(GO:0097341) |
| 0.0 | 0.3 | GO:0060325 | face morphogenesis(GO:0060325) |
| 0.0 | 0.2 | GO:0003160 | endocardium morphogenesis(GO:0003160) |
| 0.0 | 0.5 | GO:0000027 | ribosomal large subunit assembly(GO:0000027) |
| 0.0 | 0.1 | GO:0015862 | uridine transport(GO:0015862) pyrimidine nucleoside transport(GO:0015864) |
| 0.0 | 0.4 | GO:0030183 | B cell differentiation(GO:0030183) |
| 0.0 | 0.5 | GO:0010634 | positive regulation of epithelial cell migration(GO:0010634) |
| 0.0 | 0.3 | GO:0045747 | positive regulation of Notch signaling pathway(GO:0045747) |
| 0.0 | 0.2 | GO:0034312 | diol biosynthetic process(GO:0034312) sphingosine biosynthetic process(GO:0046512) |
| 0.0 | 0.3 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 0.0 | 0.2 | GO:0046549 | retinal cone cell development(GO:0046549) |
| 0.0 | 0.7 | GO:1902036 | regulation of hematopoietic stem cell differentiation(GO:1902036) |
| 0.0 | 0.5 | GO:0001732 | formation of cytoplasmic translation initiation complex(GO:0001732) |
| 0.0 | 0.4 | GO:0035775 | pronephric glomerulus morphogenesis(GO:0035775) |
| 0.0 | 0.5 | GO:0006919 | activation of cysteine-type endopeptidase activity involved in apoptotic process(GO:0006919) |
| 0.0 | 0.1 | GO:0030240 | skeletal muscle thin filament assembly(GO:0030240) |
| 0.0 | 0.1 | GO:0033539 | fatty acid beta-oxidation using acyl-CoA dehydrogenase(GO:0033539) |
| 0.0 | 0.1 | GO:0031087 | nuclear-transcribed mRNA catabolic process, deadenylation-independent decay(GO:0031086) deadenylation-independent decapping of nuclear-transcribed mRNA(GO:0031087) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 2.1 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 0.3 | 1.0 | GO:0044218 | other organism(GO:0044215) other organism cell(GO:0044216) other organism part(GO:0044217) other organism cell membrane(GO:0044218) other organism membrane(GO:0044279) |
| 0.2 | 0.9 | GO:0097268 | cytoophidium(GO:0097268) |
| 0.1 | 0.5 | GO:0061702 | inflammasome complex(GO:0061702) |
| 0.1 | 1.2 | GO:0034362 | low-density lipoprotein particle(GO:0034362) |
| 0.1 | 0.4 | GO:0019815 | B cell receptor complex(GO:0019815) |
| 0.1 | 0.6 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.1 | 0.6 | GO:0070531 | BRCA1-A complex(GO:0070531) |
| 0.1 | 0.2 | GO:0044611 | nuclear pore inner ring(GO:0044611) |
| 0.0 | 0.3 | GO:0070937 | CRD-mediated mRNA stability complex(GO:0070937) |
| 0.0 | 0.4 | GO:0045095 | keratin filament(GO:0045095) |
| 0.0 | 0.2 | GO:0042622 | photoreceptor outer segment membrane(GO:0042622) |
| 0.0 | 0.5 | GO:0016282 | eukaryotic 43S preinitiation complex(GO:0016282) |
| 0.0 | 0.1 | GO:0034098 | VCP-NPL4-UFD1 AAA ATPase complex(GO:0034098) |
| 0.0 | 0.2 | GO:0017059 | serine C-palmitoyltransferase complex(GO:0017059) endoplasmic reticulum palmitoyltransferase complex(GO:0031211) |
| 0.0 | 0.3 | GO:0030056 | hemidesmosome(GO:0030056) |
| 0.0 | 0.5 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.0 | 0.4 | GO:0000795 | synaptonemal complex(GO:0000795) |
| 0.0 | 1.9 | GO:0090575 | RNA polymerase II transcription factor complex(GO:0090575) |
| 0.0 | 1.5 | GO:0030027 | lamellipodium(GO:0030027) |
| 0.0 | 0.5 | GO:0043679 | axon terminus(GO:0043679) |
| 0.0 | 0.1 | GO:0005832 | chaperonin-containing T-complex(GO:0005832) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.4 | GO:0005499 | vitamin D binding(GO:0005499) |
| 0.2 | 1.9 | GO:0001217 | bacterial-type RNA polymerase transcription factor activity, sequence-specific DNA binding(GO:0001130) bacterial-type RNA polymerase transcriptional repressor activity, sequence-specific DNA binding(GO:0001217) |
| 0.2 | 0.9 | GO:0003883 | CTP synthase activity(GO:0003883) |
| 0.2 | 1.5 | GO:0045735 | nutrient reservoir activity(GO:0045735) |
| 0.2 | 1.8 | GO:0016936 | galactoside binding(GO:0016936) |
| 0.2 | 0.6 | GO:0005153 | interleukin-8 receptor binding(GO:0005153) |
| 0.1 | 0.6 | GO:0005173 | stem cell factor receptor binding(GO:0005173) |
| 0.1 | 0.9 | GO:0036042 | long-chain fatty acyl-CoA binding(GO:0036042) |
| 0.1 | 1.1 | GO:0004586 | ornithine decarboxylase activity(GO:0004586) |
| 0.1 | 0.6 | GO:0004024 | alcohol dehydrogenase activity, zinc-dependent(GO:0004024) S-(hydroxymethyl)glutathione dehydrogenase activity(GO:0051903) |
| 0.1 | 0.5 | GO:0004908 | interleukin-1 receptor activity(GO:0004908) |
| 0.1 | 0.7 | GO:0009378 | four-way junction helicase activity(GO:0009378) |
| 0.1 | 0.5 | GO:0008494 | translation activator activity(GO:0008494) |
| 0.0 | 0.3 | GO:0035252 | UDP-xylosyltransferase activity(GO:0035252) |
| 0.0 | 0.9 | GO:0019211 | phosphatase activator activity(GO:0019211) protein phosphatase activator activity(GO:0072542) |
| 0.0 | 0.4 | GO:0000064 | L-ornithine transmembrane transporter activity(GO:0000064) |
| 0.0 | 0.4 | GO:0015125 | bile acid transmembrane transporter activity(GO:0015125) |
| 0.0 | 0.4 | GO:0017128 | phospholipid scramblase activity(GO:0017128) |
| 0.0 | 0.5 | GO:0097199 | cysteine-type endopeptidase activity involved in apoptotic signaling pathway(GO:0097199) |
| 0.0 | 5.1 | GO:0004866 | endopeptidase inhibitor activity(GO:0004866) |
| 0.0 | 0.2 | GO:0004758 | serine C-palmitoyltransferase activity(GO:0004758) C-palmitoyltransferase activity(GO:0016454) |
| 0.0 | 1.6 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.0 | 0.5 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.0 | 0.1 | GO:0016841 | ammonia-lyase activity(GO:0016841) |
| 0.0 | 0.2 | GO:0016922 | ligand-dependent nuclear receptor binding(GO:0016922) |
| 0.0 | 0.2 | GO:0008379 | thioredoxin peroxidase activity(GO:0008379) |
| 0.0 | 0.4 | GO:0030983 | mismatched DNA binding(GO:0030983) |
| 0.0 | 0.3 | GO:0045028 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 0.0 | 0.3 | GO:0030506 | ankyrin binding(GO:0030506) |
| 0.0 | 0.4 | GO:0070492 | oligosaccharide binding(GO:0070492) |
| 0.0 | 0.1 | GO:0016920 | pyroglutamyl-peptidase activity(GO:0016920) |
| 0.0 | 1.1 | GO:0042562 | hormone binding(GO:0042562) |
| 0.0 | 0.1 | GO:0004361 | glutaryl-CoA dehydrogenase activity(GO:0004361) |
| 0.0 | 0.2 | GO:0047555 | 3',5'-cyclic-GMP phosphodiesterase activity(GO:0047555) |
| 0.0 | 0.1 | GO:0008172 | S-methyltransferase activity(GO:0008172) |
| 0.0 | 0.2 | GO:0045159 | myosin II binding(GO:0045159) |
| 0.0 | 2.6 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.0 | 0.4 | GO:0005518 | collagen binding(GO:0005518) |
| 0.0 | 0.7 | GO:0016706 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, 2-oxoglutarate as one donor, and incorporation of one atom each of oxygen into both donors(GO:0016706) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.1 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.0 | 1.1 | PID AR NONGENOMIC PATHWAY | Nongenotropic Androgen signaling |
| 0.0 | 0.4 | SA G2 AND M PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
| 0.0 | 0.5 | ST GA12 PATHWAY | G alpha 12 Pathway |
| 0.0 | 0.4 | PID ECADHERIN KERATINOCYTE PATHWAY | E-cadherin signaling in keratinocytes |
| 0.0 | 0.5 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.0 | 0.6 | PID BARD1 PATHWAY | BARD1 signaling events |
| 0.0 | 0.2 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.0 | 0.4 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 0.7 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.0 | 0.4 | PID HES HEY PATHWAY | Notch-mediated HES/HEY network |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.8 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 0.2 | 2.1 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.1 | 1.2 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.1 | 1.9 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 0.1 | 0.4 | REACTOME TRANSPORT OF ORGANIC ANIONS | Genes involved in Transport of organic anions |
| 0.0 | 1.4 | REACTOME METABOLISM OF STEROID HORMONES AND VITAMINS A AND D | Genes involved in Metabolism of steroid hormones and vitamins A and D |
| 0.0 | 0.3 | REACTOME IKK COMPLEX RECRUITMENT MEDIATED BY RIP1 | Genes involved in IKK complex recruitment mediated by RIP1 |
| 0.0 | 1.1 | REACTOME REGULATION OF ORNITHINE DECARBOXYLASE ODC | Genes involved in Regulation of ornithine decarboxylase (ODC) |
| 0.0 | 0.4 | REACTOME CYCLIN A B1 ASSOCIATED EVENTS DURING G2 M TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 0.0 | 0.9 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.0 | 0.5 | REACTOME ANTIGEN ACTIVATES B CELL RECEPTOR LEADING TO GENERATION OF SECOND MESSENGERS | Genes involved in Antigen Activates B Cell Receptor Leading to Generation of Second Messengers |
| 0.0 | 0.3 | REACTOME PTM GAMMA CARBOXYLATION HYPUSINE FORMATION AND ARYLSULFATASE ACTIVATION | Genes involved in PTM: gamma carboxylation, hypusine formation and arylsulfatase activation |