Project
PRJNA195909:zebrafish embryo and larva development
Navigation
Downloads
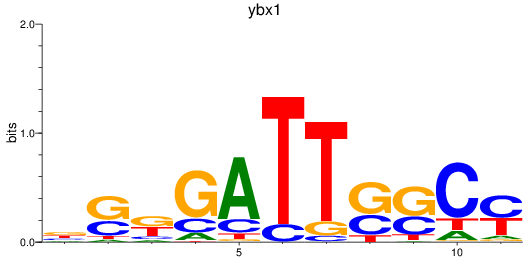
Results for ybx1
Z-value: 0.92
Transcription factors associated with ybx1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
ybx1
|
ENSDARG00000004757 | Y box binding protein 1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| ybx1 | dr11_v1_chr8_-_47152001_47152086 | -0.82 | 6.5e-03 | Click! |
Activity profile of ybx1 motif
Sorted Z-values of ybx1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr19_+_14109348 | 2.31 |
ENSDART00000159015
|
zgc:175136
|
zgc:175136 |
| chr20_-_52939501 | 2.25 |
ENSDART00000166508
|
fdft1
|
farnesyl-diphosphate farnesyltransferase 1 |
| chr9_+_29548195 | 1.77 |
ENSDART00000176057
|
rnf17
|
ring finger protein 17 |
| chr12_+_38807604 | 1.46 |
ENSDART00000155563
|
abca5
|
ATP-binding cassette, sub-family A (ABC1), member 5 |
| chr18_+_27515640 | 1.43 |
ENSDART00000181593
|
tp53i11b
|
tumor protein p53 inducible protein 11b |
| chr10_+_6010570 | 1.20 |
ENSDART00000190025
ENSDART00000163680 |
hmgcs1
|
3-hydroxy-3-methylglutaryl-CoA synthase 1 (soluble) |
| chr14_-_8903435 | 1.14 |
ENSDART00000160584
|
zgc:153681
|
zgc:153681 |
| chr9_+_45227028 | 1.11 |
ENSDART00000185579
|
adarb1b
|
adenosine deaminase, RNA-specific, B1b |
| chr16_-_39477746 | 1.08 |
ENSDART00000102525
|
stt3b
|
STT3B, subunit of the oligosaccharyltransferase complex (catalytic) |
| chr2_-_17115256 | 1.05 |
ENSDART00000190488
|
pif1
|
PIF1 5'-to-3' DNA helicase homolog (S. cerevisiae) |
| chr12_-_24832297 | 1.05 |
ENSDART00000066317
|
foxn2b
|
forkhead box N2b |
| chr7_-_73845736 | 1.05 |
ENSDART00000193414
|
zgc:173552
|
zgc:173552 |
| chr10_-_45029041 | 1.05 |
ENSDART00000167878
|
polm
|
polymerase (DNA directed), mu |
| chr7_+_57088920 | 1.04 |
ENSDART00000024076
|
scamp2l
|
secretory carrier membrane protein 2, like |
| chr12_+_1469090 | 1.03 |
ENSDART00000183637
|
usp22
|
ubiquitin specific peptidase 22 |
| chr16_-_39477509 | 0.99 |
ENSDART00000191330
|
stt3b
|
STT3B, subunit of the oligosaccharyltransferase complex (catalytic) |
| chr12_+_1469327 | 0.99 |
ENSDART00000059143
|
usp22
|
ubiquitin specific peptidase 22 |
| chr1_+_30723677 | 0.97 |
ENSDART00000177900
|
bora
|
bora, aurora kinase A activator |
| chr22_-_5171829 | 0.96 |
ENSDART00000140313
|
tnfaip8l1
|
tumor necrosis factor, alpha-induced protein 8-like 1 |
| chr1_+_30723380 | 0.96 |
ENSDART00000127943
ENSDART00000062628 ENSDART00000127670 |
bora
|
bora, aurora kinase A activator |
| chr20_-_49889111 | 0.94 |
ENSDART00000058858
|
kif13bb
|
kinesin family member 13Bb |
| chr8_-_1219815 | 0.89 |
ENSDART00000016800
ENSDART00000149969 |
znf367
|
zinc finger protein 367 |
| chr22_-_4439311 | 0.89 |
ENSDART00000169317
|
uhrf1
|
ubiquitin-like with PHD and ring finger domains 1 |
| chr22_-_5171362 | 0.88 |
ENSDART00000124889
|
tnfaip8l1
|
tumor necrosis factor, alpha-induced protein 8-like 1 |
| chr7_+_26549846 | 0.88 |
ENSDART00000141353
|
tnk1
|
tyrosine kinase, non-receptor, 1 |
| chr16_+_45930962 | 0.87 |
ENSDART00000124689
ENSDART00000041811 |
otud7b
|
OTU deubiquitinase 7B |
| chr7_+_38529263 | 0.87 |
ENSDART00000109495
ENSDART00000173804 |
nudt19
|
nudix (nucleoside diphosphate linked moiety X)-type motif 19 |
| chr7_+_5906327 | 0.86 |
ENSDART00000173160
|
zgc:112234
|
zgc:112234 |
| chr25_-_17918536 | 0.85 |
ENSDART00000148660
|
arntl1a
|
aryl hydrocarbon receptor nuclear translocator-like 1a |
| chr18_-_18942098 | 0.85 |
ENSDART00000100458
|
si:dkey-73n10.1
|
si:dkey-73n10.1 |
| chr8_+_54081819 | 0.82 |
ENSDART00000005857
ENSDART00000161795 |
prickle2a
|
prickle homolog 2a |
| chr2_-_17492080 | 0.80 |
ENSDART00000024302
|
kdm4ab
|
lysine (K)-specific demethylase 4A, genome duplicate b |
| chr24_-_24271629 | 0.78 |
ENSDART00000135060
|
rps6ka3b
|
ribosomal protein S6 kinase, polypeptide 3b |
| chr5_+_45139196 | 0.78 |
ENSDART00000113738
|
smarca2
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 |
| chr24_-_41220538 | 0.77 |
ENSDART00000150207
|
acvr2ba
|
activin A receptor type 2Ba |
| chr7_-_73854476 | 0.77 |
ENSDART00000186481
|
zgc:173552
|
zgc:173552 |
| chr3_+_3810919 | 0.77 |
ENSDART00000056035
|
FQ311927.1
|
|
| chr13_+_2394264 | 0.74 |
ENSDART00000168595
|
elovl5
|
ELOVL fatty acid elongase 5 |
| chr16_-_12095144 | 0.74 |
ENSDART00000145106
|
pex5
|
peroxisomal biogenesis factor 5 |
| chr5_-_14500622 | 0.74 |
ENSDART00000099566
|
si:ch211-244o22.2
|
si:ch211-244o22.2 |
| chr7_+_17953589 | 0.74 |
ENSDART00000174778
ENSDART00000113120 |
taf6l
|
TAF6-like RNA polymerase II, p300/CBP-associated factor (PCAF)-associated factor |
| chr3_+_1167026 | 0.74 |
ENSDART00000031823
ENSDART00000155340 |
triobpb
|
TRIO and F-actin binding protein b |
| chr2_+_105748 | 0.74 |
ENSDART00000169601
|
CABZ01098670.1
|
|
| chr25_-_17918810 | 0.73 |
ENSDART00000023959
|
arntl1a
|
aryl hydrocarbon receptor nuclear translocator-like 1a |
| chr18_+_660578 | 0.73 |
ENSDART00000161203
|
CELF6
|
si:dkey-205h23.2 |
| chr4_-_9196291 | 0.72 |
ENSDART00000153963
|
hcfc2
|
host cell factor C2 |
| chr4_+_4079418 | 0.72 |
ENSDART00000028016
|
waslb
|
Wiskott-Aldrich syndrome-like b |
| chr14_-_30945515 | 0.72 |
ENSDART00000161540
|
si:zfos-80g12.1
|
si:zfos-80g12.1 |
| chr8_-_39884359 | 0.72 |
ENSDART00000131372
|
mlec
|
malectin |
| chr12_+_24060894 | 0.71 |
ENSDART00000021298
|
asb3
|
ankyrin repeat and SOCS box containing 3 |
| chr18_+_17600570 | 0.71 |
ENSDART00000175258
ENSDART00000151850 ENSDART00000151934 |
herpud1
|
homocysteine-inducible, endoplasmic reticulum stress-inducible, ubiquitin-like domain member 1 |
| chr16_-_11241424 | 0.71 |
ENSDART00000134752
|
znf526
|
zinc finger protein 526 |
| chr20_-_3403033 | 0.70 |
ENSDART00000092264
|
rev3l
|
REV3-like, polymerase (DNA directed), zeta, catalytic subunit |
| chr7_+_5976613 | 0.70 |
ENSDART00000173105
|
si:dkey-23a13.21
|
si:dkey-23a13.21 |
| chr13_+_8892784 | 0.69 |
ENSDART00000075054
ENSDART00000143705 |
thada
|
thyroid adenoma associated |
| chr21_+_3897680 | 0.69 |
ENSDART00000170653
|
dolpp1
|
dolichyldiphosphatase 1 |
| chr25_+_25508495 | 0.68 |
ENSDART00000150719
|
phrf1
|
PHD and ring finger domains 1 |
| chr14_-_12020653 | 0.68 |
ENSDART00000106654
|
znf711
|
zinc finger protein 711 |
| chr5_+_45138934 | 0.66 |
ENSDART00000041412
ENSDART00000136002 |
smarca2
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 |
| chr11_+_440305 | 0.65 |
ENSDART00000082517
|
rab43
|
RAB43, member RAS oncogene family |
| chr9_+_22677503 | 0.65 |
ENSDART00000131429
ENSDART00000080005 ENSDART00000101756 ENSDART00000138148 |
itgb5
|
integrin, beta 5 |
| chr19_+_42806812 | 0.65 |
ENSDART00000108775
ENSDART00000151653 |
ubp1
|
upstream binding protein 1 |
| chr1_+_12394205 | 0.64 |
ENSDART00000138622
ENSDART00000136421 ENSDART00000139440 ENSDART00000184296 ENSDART00000008127 |
zgc:77739
|
zgc:77739 |
| chr7_+_34794829 | 0.64 |
ENSDART00000009698
ENSDART00000075089 ENSDART00000173456 |
esrp2
|
epithelial splicing regulatory protein 2 |
| chr19_+_791538 | 0.63 |
ENSDART00000146554
ENSDART00000138406 |
tmem79a
|
transmembrane protein 79a |
| chr7_-_73834812 | 0.61 |
ENSDART00000128137
|
zgc:92594
|
zgc:92594 |
| chr6_+_60112200 | 0.61 |
ENSDART00000008243
|
prelid3b
|
PRELI domain containing 3 |
| chr7_-_59123066 | 0.60 |
ENSDART00000175438
|
dennd4c
|
DENN/MADD domain containing 4C |
| chr17_+_23556764 | 0.60 |
ENSDART00000146787
|
pank1a
|
pantothenate kinase 1a |
| chr21_-_11054605 | 0.59 |
ENSDART00000191378
ENSDART00000084061 |
nedd4l
|
neural precursor cell expressed, developmentally down-regulated 4-like |
| chr24_+_17334682 | 0.59 |
ENSDART00000018868
|
pdia4
|
protein disulfide isomerase family A, member 4 |
| chr5_+_20030414 | 0.59 |
ENSDART00000181430
ENSDART00000047841 ENSDART00000182813 |
sgsm1a
|
small G protein signaling modulator 1a |
| chr25_-_6389713 | 0.59 |
ENSDART00000083539
|
sin3aa
|
SIN3 transcription regulator family member Aa |
| chr3_-_21062706 | 0.58 |
ENSDART00000155605
ENSDART00000153686 ENSDART00000157168 ENSDART00000156614 ENSDART00000155743 ENSDART00000156275 |
fam57ba
|
family with sequence similarity 57, member Ba |
| chr4_-_8060962 | 0.58 |
ENSDART00000146622
|
wnk1b
|
WNK lysine deficient protein kinase 1b |
| chr12_-_33582382 | 0.57 |
ENSDART00000009794
ENSDART00000136617 |
tdrkh
|
tudor and KH domain containing |
| chr19_-_31686252 | 0.57 |
ENSDART00000131721
|
ripor2
|
RHO family interacting cell polarization regulator 2 |
| chr25_-_35102781 | 0.57 |
ENSDART00000180881
ENSDART00000153747 |
si:dkey-108k21.24
|
si:dkey-108k21.24 |
| chr7_+_5905091 | 0.56 |
ENSDART00000167099
|
CU459186.3
|
Histone H3.2 |
| chr15_+_26940569 | 0.55 |
ENSDART00000189636
ENSDART00000077172 |
bcas3
|
breast carcinoma amplified sequence 3 |
| chr16_+_6750289 | 0.55 |
ENSDART00000167736
|
znf236
|
zinc finger protein 236 |
| chr1_+_54737353 | 0.54 |
ENSDART00000130675
ENSDART00000162075 |
pi4k2a
|
phosphatidylinositol 4-kinase type 2 alpha |
| chr21_-_11054876 | 0.54 |
ENSDART00000146576
|
nedd4l
|
neural precursor cell expressed, developmentally down-regulated 4-like |
| chr25_-_6261693 | 0.54 |
ENSDART00000135808
|
ireb2
|
iron-responsive element binding protein 2 |
| chr10_-_35257458 | 0.54 |
ENSDART00000143890
ENSDART00000139107 ENSDART00000082445 |
prr11
|
proline rich 11 |
| chr21_+_34981263 | 0.53 |
ENSDART00000132711
|
rbm11
|
RNA binding motif protein 11 |
| chr3_+_35498119 | 0.53 |
ENSDART00000178963
|
tnrc6a
|
trinucleotide repeat containing 6a |
| chr15_+_25439106 | 0.53 |
ENSDART00000156252
|
aifm4
|
apoptosis-inducing factor, mitochondrion-associated, 4 |
| chr11_-_27778831 | 0.52 |
ENSDART00000023823
|
cass4
|
Cas scaffolding protein family member 4 |
| chr23_-_10786400 | 0.52 |
ENSDART00000055038
|
rybpa
|
RING1 and YY1 binding protein a |
| chr9_+_2452672 | 0.52 |
ENSDART00000193993
|
chn1
|
chimerin 1 |
| chr21_-_1635268 | 0.51 |
ENSDART00000151258
|
zgc:152948
|
zgc:152948 |
| chr25_+_36312459 | 0.51 |
ENSDART00000182484
|
CR354435.5
|
|
| chr20_-_40487208 | 0.51 |
ENSDART00000075070
ENSDART00000142029 |
hsf2
|
heat shock transcription factor 2 |
| chr19_-_6134802 | 0.51 |
ENSDART00000140051
|
cica
|
capicua transcriptional repressor a |
| chr24_-_36301072 | 0.49 |
ENSDART00000062736
|
coasy
|
CoA synthase |
| chr23_-_35483163 | 0.49 |
ENSDART00000138660
ENSDART00000113643 ENSDART00000189269 |
fbxo25
|
F-box protein 25 |
| chr3_+_16664212 | 0.49 |
ENSDART00000013816
|
zgc:55558
|
zgc:55558 |
| chr1_+_54069450 | 0.49 |
ENSDART00000108601
ENSDART00000187878 |
dcaf15
|
DDB1 and CUL4 associated factor 15 |
| chr9_-_1604601 | 0.49 |
ENSDART00000143130
|
agps
|
alkylglycerone phosphate synthase |
| chr17_+_35097024 | 0.49 |
ENSDART00000026152
|
asap2a
|
ArfGAP with SH3 domain, ankyrin repeat and PH domain 2a |
| chr20_-_2641233 | 0.48 |
ENSDART00000145335
ENSDART00000133121 |
bub1
|
BUB1 mitotic checkpoint serine/threonine kinase |
| chr7_-_6444011 | 0.47 |
ENSDART00000173010
|
zgc:112234
|
zgc:112234 |
| chr6_+_37752781 | 0.47 |
ENSDART00000154364
|
herc2
|
HECT and RLD domain containing E3 ubiquitin protein ligase 2 |
| chr15_+_25438714 | 0.47 |
ENSDART00000154164
|
aifm4
|
apoptosis-inducing factor, mitochondrion-associated, 4 |
| chr23_+_32028574 | 0.47 |
ENSDART00000145501
ENSDART00000143121 ENSDART00000111877 |
tpx2
|
TPX2, microtubule-associated, homolog (Xenopus laevis) |
| chr3_+_36671585 | 0.47 |
ENSDART00000159033
|
nde1
|
nudE neurodevelopment protein 1 |
| chr15_+_26941063 | 0.46 |
ENSDART00000149957
|
bcas3
|
breast carcinoma amplified sequence 3 |
| chr1_-_54972170 | 0.46 |
ENSDART00000150548
ENSDART00000038330 |
khsrp
|
KH-type splicing regulatory protein |
| chr21_-_1642739 | 0.46 |
ENSDART00000151118
|
zgc:152948
|
zgc:152948 |
| chr2_+_58877162 | 0.46 |
ENSDART00000122174
|
CABZ01085658.1
|
|
| chr7_+_73827805 | 0.46 |
ENSDART00000109316
|
zgc:173587
|
zgc:173587 |
| chr18_+_907266 | 0.46 |
ENSDART00000171729
|
pkma
|
pyruvate kinase M1/2a |
| chr11_+_31324335 | 0.46 |
ENSDART00000088093
|
sipa1l2
|
signal-induced proliferation-associated 1 like 2 |
| chr7_+_1521834 | 0.45 |
ENSDART00000174007
|
si:cabz01102082.1
|
si:cabz01102082.1 |
| chr23_-_2037566 | 0.45 |
ENSDART00000191312
ENSDART00000127443 |
prdm5
|
PR domain containing 5 |
| chr20_+_37866171 | 0.45 |
ENSDART00000153190
|
vash2
|
vasohibin 2 |
| chr7_-_6431158 | 0.45 |
ENSDART00000173199
|
si:ch1073-153i20.5
|
si:ch1073-153i20.5 |
| chr7_-_6364168 | 0.45 |
ENSDART00000173332
|
zgc:112234
|
zgc:112234 |
| chr22_-_4407871 | 0.45 |
ENSDART00000162523
|
kdm4b
|
lysine (K)-specific demethylase 4B |
| chr15_-_6615555 | 0.45 |
ENSDART00000152725
|
atm
|
ATM serine/threonine kinase |
| chr13_-_42536642 | 0.45 |
ENSDART00000134533
|
btaf1
|
BTAF1 RNA polymerase II, B-TFIID transcription factor-associated |
| chr7_-_31794476 | 0.44 |
ENSDART00000142385
|
nap1l4b
|
nucleosome assembly protein 1-like 4b |
| chr18_+_21273749 | 0.42 |
ENSDART00000143265
|
hydin
|
HYDIN, axonemal central pair apparatus protein |
| chr5_+_26686639 | 0.42 |
ENSDART00000079064
|
tango2
|
transport and golgi organization 2 homolog (Drosophila) |
| chr7_+_29115890 | 0.42 |
ENSDART00000052345
|
tradd
|
tnfrsf1a-associated via death domain |
| chr19_-_15420678 | 0.42 |
ENSDART00000151454
ENSDART00000027697 |
serinc2
|
serine incorporator 2 |
| chr3_+_17878124 | 0.42 |
ENSDART00000166430
ENSDART00000163421 ENSDART00000121473 |
dnajc7
|
DnaJ (Hsp40) homolog, subfamily C, member 7 |
| chr20_+_50061890 | 0.41 |
ENSDART00000137725
|
cpsf2
|
cleavage and polyadenylation specific factor 2 |
| chr1_-_50527964 | 0.41 |
ENSDART00000024984
|
papss1
|
3'-phosphoadenosine 5'-phosphosulfate synthase 1 |
| chr1_-_54971968 | 0.41 |
ENSDART00000140016
|
khsrp
|
KH-type splicing regulatory protein |
| chr20_+_21391181 | 0.41 |
ENSDART00000185158
ENSDART00000049586 ENSDART00000024922 |
jag2b
|
jagged 2b |
| chr22_+_30543437 | 0.41 |
ENSDART00000137983
|
si:dkey-103k4.1
|
si:dkey-103k4.1 |
| chr3_+_17878466 | 0.40 |
ENSDART00000180218
|
dnajc7
|
DnaJ (Hsp40) homolog, subfamily C, member 7 |
| chr7_+_5965611 | 0.40 |
ENSDART00000115062
|
CU459186.2
|
Histone H3.2 |
| chr13_-_5257303 | 0.39 |
ENSDART00000110610
|
si:dkey-78p8.1
|
si:dkey-78p8.1 |
| chr19_-_26769867 | 0.39 |
ENSDART00000043776
ENSDART00000159489 ENSDART00000138675 |
prrc2a
|
proline-rich coiled-coil 2A |
| chr7_+_5910467 | 0.38 |
ENSDART00000173232
|
si:ch211-113a14.22
|
si:ch211-113a14.22 |
| chr11_-_45152702 | 0.38 |
ENSDART00000168066
|
afmid
|
arylformamidase |
| chr5_-_2721686 | 0.38 |
ENSDART00000169404
|
hspa5
|
heat shock protein 5 |
| chr12_-_10674606 | 0.37 |
ENSDART00000157919
|
med24
|
mediator complex subunit 24 |
| chr5_+_36439405 | 0.37 |
ENSDART00000102973
|
eda
|
ectodysplasin A |
| chr3_-_13147310 | 0.36 |
ENSDART00000160840
|
prkar1b
|
protein kinase, cAMP-dependent, regulatory, type I, beta |
| chr3_-_1146497 | 0.35 |
ENSDART00000149061
|
si:ch73-211l13.2
|
si:ch73-211l13.2 |
| chr23_+_36616717 | 0.35 |
ENSDART00000042701
ENSDART00000192980 |
pip4k2ca
|
phosphatidylinositol-5-phosphate 4-kinase, type II, gamma a |
| chr3_-_10751491 | 0.35 |
ENSDART00000016351
|
zgc:112965
|
zgc:112965 |
| chr12_+_25432627 | 0.35 |
ENSDART00000011662
|
ppm1bb
|
protein phosphatase, Mg2+/Mn2+ dependent, 1Bb |
| chr1_-_51710225 | 0.35 |
ENSDART00000057601
ENSDART00000152745 |
snrpb2
|
small nuclear ribonucleoprotein polypeptide B2 |
| chr1_+_58840889 | 0.35 |
ENSDART00000098308
|
tmed1b
|
transmembrane p24 trafficking protein 1b |
| chr25_-_35110695 | 0.34 |
ENSDART00000099859
|
hist1h2a10
|
histone cluster 1 H2A family member 10 |
| chr14_+_80685 | 0.34 |
ENSDART00000188443
|
stag3
|
stromal antigen 3 |
| chr9_-_10068004 | 0.34 |
ENSDART00000011922
ENSDART00000162818 |
spopla
|
speckle-type POZ protein-like a |
| chr5_+_29726428 | 0.34 |
ENSDART00000143183
|
ddx31
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 31 |
| chr8_-_4574328 | 0.34 |
ENSDART00000090731
|
dhx37
|
DEAH (Asp-Glu-Ala-His) box polypeptide 37 |
| chr1_-_12394048 | 0.34 |
ENSDART00000146067
ENSDART00000134708 |
sclt1
|
sodium channel and clathrin linker 1 |
| chr25_+_4541211 | 0.34 |
ENSDART00000129978
|
pnpla2
|
patatin-like phospholipase domain containing 2 |
| chr24_-_39772045 | 0.33 |
ENSDART00000087441
|
GFOD1
|
si:ch211-276f18.2 |
| chr20_+_28266892 | 0.33 |
ENSDART00000103330
|
chac1
|
ChaC, cation transport regulator homolog 1 (E. coli) |
| chr2_+_37480669 | 0.32 |
ENSDART00000029801
|
sppl2
|
signal peptide peptidase-like 2 |
| chr22_-_22301672 | 0.32 |
ENSDART00000111711
|
chaf1a
|
chromatin assembly factor 1, subunit A (p150) |
| chr1_+_51386649 | 0.32 |
ENSDART00000152289
|
atg4da
|
autophagy related 4D, cysteine peptidase a |
| chr19_-_26770083 | 0.31 |
ENSDART00000193811
ENSDART00000174455 |
prrc2a
|
proline-rich coiled-coil 2A |
| chr7_+_73819078 | 0.31 |
ENSDART00000169756
|
FP236812.1
|
Histone H2B 1/2 |
| chr14_-_246342 | 0.31 |
ENSDART00000054823
|
aurkb
|
aurora kinase B |
| chr16_-_52646789 | 0.30 |
ENSDART00000035761
|
ubr5
|
ubiquitin protein ligase E3 component n-recognin 5 |
| chr3_-_6417328 | 0.30 |
ENSDART00000160979
|
jpt1b
|
Jupiter microtubule associated homolog 1b |
| chr11_+_45436703 | 0.30 |
ENSDART00000168295
ENSDART00000173293 |
sos1
|
son of sevenless homolog 1 (Drosophila) |
| chr2_+_24867534 | 0.30 |
ENSDART00000158050
|
rab3aa
|
RAB3A, member RAS oncogene family, a |
| chr3_-_8765165 | 0.30 |
ENSDART00000191131
|
CABZ01058333.1
|
|
| chr8_-_1266181 | 0.30 |
ENSDART00000148654
ENSDART00000149924 |
cdc14b
|
cell division cycle 14B |
| chr2_+_29249561 | 0.29 |
ENSDART00000099157
|
cdh18a
|
cadherin 18, type 2a |
| chr10_+_35257651 | 0.29 |
ENSDART00000028940
|
styxl1
|
serine/threonine/tyrosine interacting-like 1 |
| chr11_+_18612166 | 0.29 |
ENSDART00000162694
|
ncoa3
|
nuclear receptor coactivator 3 |
| chr7_-_13884610 | 0.29 |
ENSDART00000006897
|
rlbp1a
|
retinaldehyde binding protein 1a |
| chr1_+_49651016 | 0.29 |
ENSDART00000074380
ENSDART00000101017 |
tsga10
|
testis specific, 10 |
| chr7_-_6345507 | 0.28 |
ENSDART00000173032
|
CU457819.4
|
Histone H3.2 |
| chr7_+_24390939 | 0.28 |
ENSDART00000087494
ENSDART00000125463 |
haus3
|
HAUS augmin-like complex, subunit 3 |
| chr20_-_7176809 | 0.28 |
ENSDART00000012247
ENSDART00000125019 |
dhcr24
|
24-dehydrocholesterol reductase |
| chr11_+_14321113 | 0.28 |
ENSDART00000039822
ENSDART00000137347 ENSDART00000132997 |
ptbp1b
|
polypyrimidine tract binding protein 1b |
| chr5_+_24086227 | 0.27 |
ENSDART00000051549
ENSDART00000177458 ENSDART00000135934 |
tp53
|
tumor protein p53 |
| chr5_+_26686279 | 0.27 |
ENSDART00000193543
|
tango2
|
transport and golgi organization 2 homolog (Drosophila) |
| chr16_-_12060770 | 0.27 |
ENSDART00000183237
ENSDART00000103948 |
si:ch211-69g19.2
|
si:ch211-69g19.2 |
| chr19_-_24135824 | 0.27 |
ENSDART00000189505
ENSDART00000104087 |
thap7
|
THAP domain containing 7 |
| chr13_-_9119867 | 0.27 |
ENSDART00000137255
|
si:dkey-112g5.15
|
si:dkey-112g5.15 |
| chr7_-_6466119 | 0.27 |
ENSDART00000173138
|
zgc:112234
|
zgc:112234 |
| chr7_-_56835543 | 0.26 |
ENSDART00000010322
|
senp8
|
SUMO/sentrin peptidase family member, NEDD8 specific |
| chr25_-_36361697 | 0.26 |
ENSDART00000152388
|
si:ch211-113a14.22
|
si:ch211-113a14.22 |
| chr25_-_35150933 | 0.26 |
ENSDART00000129254
|
HIST2H3C
|
zgc:173552 |
| chr12_-_9796237 | 0.26 |
ENSDART00000149344
|
prdm9
|
PR domain containing 9 |
| chr5_+_26138313 | 0.26 |
ENSDART00000010041
|
dhfr
|
dihydrofolate reductase |
| chr5_+_5398966 | 0.26 |
ENSDART00000139553
|
mapkap1
|
mitogen-activated protein kinase associated protein 1 |
| chr12_+_40905427 | 0.26 |
ENSDART00000170526
ENSDART00000185771 ENSDART00000193945 |
CDH18
|
cadherin 18 |
| chr16_-_32672883 | 0.26 |
ENSDART00000124515
ENSDART00000190920 ENSDART00000188776 |
pnisr
|
PNN-interacting serine/arginine-rich protein |
| chr23_+_37185247 | 0.26 |
ENSDART00000146269
|
vwa5b1
|
von Willebrand factor A domain containing 5B1 |
| chr18_-_2549198 | 0.26 |
ENSDART00000186516
|
CABZ01070631.1
|
|
| chr19_+_2619444 | 0.26 |
ENSDART00000169483
|
fam126a
|
family with sequence similarity 126, member A |
| chr6_+_59854224 | 0.25 |
ENSDART00000083499
|
kdm6al
|
lysine (K)-specific demethylase 6A, like |
| chr3_+_17951790 | 0.24 |
ENSDART00000164663
|
aclya
|
ATP citrate lyase a |
| chr1_+_43686251 | 0.24 |
ENSDART00000074604
ENSDART00000137791 |
cisd2
|
CDGSH iron sulfur domain 2 |
| chr10_-_2713228 | 0.23 |
ENSDART00000123754
ENSDART00000126236 |
mier3a
|
mesoderm induction early response 1, family member 3 a |
Network of associatons between targets according to the STRING database.
First level regulatory network of ybx1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 3.5 | GO:0045338 | farnesyl diphosphate metabolic process(GO:0045338) |
| 0.2 | 1.4 | GO:0031507 | heterochromatin assembly(GO:0031507) |
| 0.2 | 0.7 | GO:1903069 | regulation of ER-associated ubiquitin-dependent protein catabolic process(GO:1903069) |
| 0.2 | 0.6 | GO:1990869 | response to chemokine(GO:1990868) cellular response to chemokine(GO:1990869) negative regulation of lymphocyte migration(GO:2000402) negative regulation of T cell migration(GO:2000405) |
| 0.2 | 0.9 | GO:0071947 | protein deubiquitination involved in ubiquitin-dependent protein catabolic process(GO:0071947) |
| 0.2 | 2.4 | GO:0060236 | regulation of mitotic spindle organization(GO:0060236) |
| 0.2 | 0.5 | GO:0031630 | regulation of synaptic vesicle fusion to presynaptic membrane(GO:0031630) |
| 0.2 | 0.5 | GO:0046504 | ether lipid biosynthetic process(GO:0008611) glycerol ether biosynthetic process(GO:0046504) cellular lipid biosynthetic process(GO:0097384) ether biosynthetic process(GO:1901503) |
| 0.2 | 0.5 | GO:0071470 | cellular response to osmotic stress(GO:0071470) |
| 0.1 | 1.1 | GO:2000650 | negative regulation of sodium ion transmembrane transporter activity(GO:2000650) |
| 0.1 | 0.4 | GO:0050427 | sulfate assimilation(GO:0000103) purine ribonucleoside bisphosphate metabolic process(GO:0034035) purine ribonucleoside bisphosphate biosynthetic process(GO:0034036) 3'-phosphoadenosine 5'-phosphosulfate metabolic process(GO:0050427) 3'-phosphoadenosine 5'-phosphosulfate biosynthetic process(GO:0050428) |
| 0.1 | 0.5 | GO:0021557 | oculomotor nerve development(GO:0021557) |
| 0.1 | 0.8 | GO:0016572 | histone phosphorylation(GO:0016572) |
| 0.1 | 2.1 | GO:0018279 | protein N-linked glycosylation via asparagine(GO:0018279) post-translational protein modification(GO:0043687) |
| 0.1 | 0.2 | GO:0070987 | error-free translesion synthesis(GO:0070987) |
| 0.1 | 0.7 | GO:1901031 | response to leucine(GO:0043201) cellular response to leucine(GO:0071233) regulation of response to reactive oxygen species(GO:1901031) |
| 0.1 | 0.9 | GO:0072383 | plus-end-directed vesicle transport along microtubule(GO:0072383) plus-end-directed organelle transport along microtubule(GO:0072386) |
| 0.1 | 0.3 | GO:0010526 | regulation of transposition, RNA-mediated(GO:0010525) negative regulation of transposition, RNA-mediated(GO:0010526) transposition, RNA-mediated(GO:0032197) positive regulation of neuron apoptotic process(GO:0043525) positive regulation of neuron death(GO:1901216) |
| 0.1 | 1.1 | GO:0006382 | adenosine to inosine editing(GO:0006382) |
| 0.1 | 0.9 | GO:0044030 | regulation of DNA methylation(GO:0044030) |
| 0.1 | 1.1 | GO:0000002 | mitochondrial genome maintenance(GO:0000002) |
| 0.1 | 0.7 | GO:0042276 | error-prone translesion synthesis(GO:0042276) |
| 0.1 | 0.3 | GO:0097676 | histone H3-K4 trimethylation(GO:0080182) histone H3-K36 dimethylation(GO:0097676) |
| 0.1 | 0.3 | GO:0010896 | regulation of triglyceride catabolic process(GO:0010896) positive regulation of triglyceride catabolic process(GO:0010898) |
| 0.1 | 1.1 | GO:0034033 | coenzyme A biosynthetic process(GO:0015937) nucleoside bisphosphate biosynthetic process(GO:0033866) ribonucleoside bisphosphate biosynthetic process(GO:0034030) purine nucleoside bisphosphate biosynthetic process(GO:0034033) |
| 0.1 | 0.4 | GO:2000001 | regulation of DNA damage checkpoint(GO:2000001) |
| 0.1 | 0.7 | GO:0022615 | protein import into peroxisome matrix, docking(GO:0016560) protein to membrane docking(GO:0022615) |
| 0.1 | 0.3 | GO:2001244 | positive regulation of intrinsic apoptotic signaling pathway(GO:2001244) |
| 0.1 | 0.5 | GO:0051754 | meiotic sister chromatid cohesion, centromeric(GO:0051754) |
| 0.1 | 0.5 | GO:0051299 | mitotic centrosome separation(GO:0007100) centrosome separation(GO:0051299) |
| 0.1 | 0.3 | GO:0031293 | membrane protein intracellular domain proteolysis(GO:0031293) |
| 0.1 | 0.3 | GO:0046654 | tetrahydrofolate biosynthetic process(GO:0046654) |
| 0.1 | 0.7 | GO:0035306 | positive regulation of dephosphorylation(GO:0035306) positive regulation of protein dephosphorylation(GO:0035307) |
| 0.1 | 0.2 | GO:0010259 | multicellular organism aging(GO:0010259) |
| 0.1 | 0.5 | GO:1900026 | positive regulation of substrate adhesion-dependent cell spreading(GO:1900026) |
| 0.1 | 0.4 | GO:0006398 | mRNA 3'-end processing by stem-loop binding and cleavage(GO:0006398) |
| 0.1 | 0.2 | GO:0007084 | mitotic nuclear envelope reassembly(GO:0007084) |
| 0.1 | 1.8 | GO:0032007 | negative regulation of TOR signaling(GO:0032007) |
| 0.1 | 0.3 | GO:1902914 | negative regulation of histone ubiquitination(GO:0033183) regulation of protein K63-linked ubiquitination(GO:1900044) negative regulation of protein K63-linked ubiquitination(GO:1900045) regulation of histone H2A K63-linked ubiquitination(GO:1901314) negative regulation of histone H2A K63-linked ubiquitination(GO:1901315) regulation of protein polyubiquitination(GO:1902914) negative regulation of protein polyubiquitination(GO:1902915) |
| 0.1 | 1.7 | GO:0009648 | photoperiodism(GO:0009648) |
| 0.0 | 0.4 | GO:0019441 | tryptophan catabolic process to kynurenine(GO:0019441) |
| 0.0 | 0.4 | GO:0060575 | intestinal epithelial cell differentiation(GO:0060575) |
| 0.0 | 0.3 | GO:0090134 | establishment or maintenance of cytoskeleton polarity(GO:0030952) mesendoderm migration(GO:0090133) cell migration involved in mesendoderm migration(GO:0090134) |
| 0.0 | 0.6 | GO:0032483 | regulation of Rab protein signal transduction(GO:0032483) |
| 0.0 | 0.4 | GO:0044837 | assembly of actomyosin apparatus involved in cytokinesis(GO:0000912) actomyosin contractile ring assembly(GO:0000915) actomyosin contractile ring organization(GO:0044837) |
| 0.0 | 0.3 | GO:0006751 | glutathione catabolic process(GO:0006751) |
| 0.0 | 0.1 | GO:0060907 | dendritic cell antigen processing and presentation(GO:0002468) macrophage cytokine production(GO:0010934) regulation of macrophage cytokine production(GO:0010935) positive regulation of macrophage cytokine production(GO:0060907) negative regulation of intrinsic apoptotic signaling pathway in response to DNA damage(GO:1902230) |
| 0.0 | 0.6 | GO:1903286 | positive regulation of sodium ion transport(GO:0010765) positive regulation of sodium ion transmembrane transport(GO:1902307) regulation of potassium ion import(GO:1903286) positive regulation of potassium ion import(GO:1903288) positive regulation of sodium ion transmembrane transporter activity(GO:2000651) |
| 0.0 | 1.0 | GO:0006303 | double-strand break repair via nonhomologous end joining(GO:0006303) |
| 0.0 | 0.3 | GO:0071850 | mitotic cell cycle arrest(GO:0071850) |
| 0.0 | 0.5 | GO:0045974 | miRNA mediated inhibition of translation(GO:0035278) negative regulation of translation, ncRNA-mediated(GO:0040033) regulation of translation, ncRNA-mediated(GO:0045974) |
| 0.0 | 0.7 | GO:0042759 | long-chain fatty acid biosynthetic process(GO:0042759) |
| 0.0 | 0.5 | GO:0061647 | histone H3-K9 methylation(GO:0051567) histone H3-K9 modification(GO:0061647) |
| 0.0 | 0.3 | GO:0021702 | cerebellar Purkinje cell layer formation(GO:0021694) cerebellar Purkinje cell differentiation(GO:0021702) |
| 0.0 | 0.2 | GO:0017187 | peptidyl-glutamic acid carboxylation(GO:0017187) protein carboxylation(GO:0018214) |
| 0.0 | 0.7 | GO:1990798 | pancreas regeneration(GO:1990798) |
| 0.0 | 0.5 | GO:0007032 | endosome organization(GO:0007032) |
| 0.0 | 0.3 | GO:0031397 | negative regulation of protein ubiquitination(GO:0031397) |
| 0.0 | 0.4 | GO:0042026 | protein refolding(GO:0042026) |
| 0.0 | 0.3 | GO:0045162 | clustering of voltage-gated sodium channels(GO:0045162) |
| 0.0 | 0.4 | GO:0010738 | regulation of protein kinase A signaling(GO:0010738) |
| 0.0 | 0.8 | GO:0006101 | citrate metabolic process(GO:0006101) tricarboxylic acid metabolic process(GO:0072350) |
| 0.0 | 0.7 | GO:0033627 | cell adhesion mediated by integrin(GO:0033627) |
| 0.0 | 0.2 | GO:1902624 | positive regulation of neutrophil migration(GO:1902624) |
| 0.0 | 0.2 | GO:0006561 | proline biosynthetic process(GO:0006561) L-proline biosynthetic process(GO:0055129) |
| 0.0 | 0.2 | GO:0071557 | histone H3-K27 demethylation(GO:0071557) |
| 0.0 | 0.3 | GO:0048026 | positive regulation of mRNA splicing, via spliceosome(GO:0048026) |
| 0.0 | 0.4 | GO:0035622 | intrahepatic bile duct development(GO:0035622) |
| 0.0 | 2.0 | GO:0016579 | protein deubiquitination(GO:0016579) |
| 0.0 | 0.8 | GO:0032924 | activin receptor signaling pathway(GO:0032924) |
| 0.0 | 0.7 | GO:0006367 | transcription initiation from RNA polymerase II promoter(GO:0006367) |
| 0.0 | 0.3 | GO:0043124 | negative regulation of I-kappaB kinase/NF-kappaB signaling(GO:0043124) |
| 0.0 | 0.5 | GO:0007050 | cell cycle arrest(GO:0007050) |
| 0.0 | 1.4 | GO:0006334 | nucleosome assembly(GO:0006334) |
| 0.0 | 0.8 | GO:0006342 | chromatin silencing(GO:0006342) |
| 0.0 | 1.6 | GO:0006338 | chromatin remodeling(GO:0006338) |
| 0.0 | 0.4 | GO:0016339 | calcium-dependent cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0016339) |
| 0.0 | 1.0 | GO:0007030 | Golgi organization(GO:0007030) |
| 0.0 | 0.3 | GO:0007634 | optokinetic behavior(GO:0007634) |
| 0.0 | 0.1 | GO:0033137 | negative regulation of peptidyl-serine phosphorylation(GO:0033137) |
| 0.0 | 0.2 | GO:0034629 | cellular protein complex localization(GO:0034629) |
| 0.0 | 0.4 | GO:0042476 | odontogenesis(GO:0042476) |
| 0.0 | 0.5 | GO:0007062 | sister chromatid cohesion(GO:0007062) |
| 0.0 | 1.6 | GO:0043484 | regulation of RNA splicing(GO:0043484) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.7 | GO:0046695 | SLIK (SAGA-like) complex(GO:0046695) |
| 0.1 | 0.6 | GO:0060171 | stereocilium membrane(GO:0060171) |
| 0.1 | 0.4 | GO:0034663 | endoplasmic reticulum chaperone complex(GO:0034663) |
| 0.1 | 0.5 | GO:0005880 | nuclear microtubule(GO:0005880) |
| 0.1 | 1.6 | GO:0034751 | aryl hydrocarbon receptor complex(GO:0034751) |
| 0.1 | 0.7 | GO:0061700 | GATOR2 complex(GO:0061700) |
| 0.1 | 0.3 | GO:0071556 | integral component of cytoplasmic side of endoplasmic reticulum membrane(GO:0071458) integral component of lumenal side of endoplasmic reticulum membrane(GO:0071556) lumenal side of endoplasmic reticulum membrane(GO:0098553) |
| 0.1 | 0.3 | GO:0033186 | CAF-1 complex(GO:0033186) |
| 0.1 | 1.5 | GO:0032588 | trans-Golgi network membrane(GO:0032588) |
| 0.1 | 0.9 | GO:0000792 | heterochromatin(GO:0000792) |
| 0.1 | 0.8 | GO:0048179 | activin receptor complex(GO:0048179) |
| 0.1 | 0.7 | GO:0042575 | DNA polymerase complex(GO:0042575) |
| 0.1 | 0.3 | GO:0032133 | chromosome passenger complex(GO:0032133) |
| 0.0 | 0.6 | GO:0016580 | Sin3 complex(GO:0016580) |
| 0.0 | 4.6 | GO:0000786 | nucleosome(GO:0000786) |
| 0.0 | 0.4 | GO:0005952 | cAMP-dependent protein kinase complex(GO:0005952) |
| 0.0 | 1.4 | GO:0016514 | SWI/SNF complex(GO:0016514) |
| 0.0 | 0.6 | GO:0000813 | ESCRT I complex(GO:0000813) |
| 0.0 | 1.4 | GO:0005657 | replication fork(GO:0005657) |
| 0.0 | 0.7 | GO:0071339 | MLL1/2 complex(GO:0044665) MLL1 complex(GO:0071339) |
| 0.0 | 0.3 | GO:0070652 | HAUS complex(GO:0070652) |
| 0.0 | 0.4 | GO:0005847 | mRNA cleavage and polyadenylation specificity factor complex(GO:0005847) |
| 0.0 | 0.5 | GO:0031083 | BLOC-1 complex(GO:0031083) |
| 0.0 | 0.5 | GO:0000780 | condensed nuclear chromosome kinetochore(GO:0000778) condensed nuclear chromosome, centromeric region(GO:0000780) |
| 0.0 | 0.1 | GO:0030915 | Smc5-Smc6 complex(GO:0030915) |
| 0.0 | 0.7 | GO:0005778 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) |
| 0.0 | 0.2 | GO:0098827 | endoplasmic reticulum tubular network(GO:0071782) endoplasmic reticulum subcompartment(GO:0098827) |
| 0.0 | 1.4 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.0 | 0.3 | GO:0031932 | TORC2 complex(GO:0031932) |
| 0.0 | 0.3 | GO:0005685 | U1 snRNP(GO:0005685) |
| 0.0 | 0.6 | GO:0005758 | mitochondrial intermembrane space(GO:0005758) organelle envelope lumen(GO:0031970) |
| 0.0 | 0.1 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 0.0 | 0.3 | GO:0008278 | cohesin complex(GO:0008278) |
| 0.0 | 0.7 | GO:0008305 | integrin complex(GO:0008305) |
| 0.0 | 0.1 | GO:0032783 | ELL-EAF complex(GO:0032783) |
| 0.0 | 0.3 | GO:0033116 | endoplasmic reticulum-Golgi intermediate compartment membrane(GO:0033116) |
| 0.0 | 0.6 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.0 | 0.3 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.0 | 0.3 | GO:0080008 | Cul4-RING E3 ubiquitin ligase complex(GO:0080008) |
| 0.0 | 0.1 | GO:0071818 | BAT3 complex(GO:0071818) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.8 | 2.3 | GO:0004311 | farnesyltranstransferase activity(GO:0004311) |
| 0.5 | 2.1 | GO:0004579 | oligosaccharyl transferase activity(GO:0004576) dolichyl-diphosphooligosaccharide-protein glycotransferase activity(GO:0004579) |
| 0.4 | 1.2 | GO:0004421 | hydroxymethylglutaryl-CoA synthase activity(GO:0004421) |
| 0.2 | 1.1 | GO:0043139 | 5'-3' DNA helicase activity(GO:0043139) |
| 0.2 | 0.5 | GO:0003994 | aconitate hydratase activity(GO:0003994) |
| 0.1 | 0.6 | GO:0071889 | 14-3-3 protein binding(GO:0071889) |
| 0.1 | 1.1 | GO:0019871 | sodium channel inhibitor activity(GO:0019871) |
| 0.1 | 0.4 | GO:0004020 | adenylylsulfate kinase activity(GO:0004020) sulfate adenylyltransferase activity(GO:0004779) sulfate adenylyltransferase (ATP) activity(GO:0004781) |
| 0.1 | 0.4 | GO:0004061 | arylformamidase activity(GO:0004061) |
| 0.1 | 0.7 | GO:0070728 | leucine binding(GO:0070728) |
| 0.1 | 1.1 | GO:0003726 | double-stranded RNA adenosine deaminase activity(GO:0003726) |
| 0.1 | 0.6 | GO:0004594 | pantothenate kinase activity(GO:0004594) |
| 0.1 | 0.7 | GO:0005052 | peroxisome matrix targeting signal-1 binding(GO:0005052) |
| 0.1 | 0.4 | GO:0016308 | 1-phosphatidylinositol-4-phosphate 5-kinase activity(GO:0016308) |
| 0.1 | 0.9 | GO:0035925 | AU-rich element binding(GO:0017091) mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.1 | 0.6 | GO:1990050 | phosphatidic acid transporter activity(GO:1990050) |
| 0.1 | 2.0 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.1 | 0.3 | GO:0003839 | gamma-glutamylcyclotransferase activity(GO:0003839) |
| 0.1 | 1.2 | GO:0032454 | histone demethylase activity (H3-K9 specific)(GO:0032454) |
| 0.1 | 0.5 | GO:0004430 | 1-phosphatidylinositol 4-kinase activity(GO:0004430) |
| 0.1 | 0.5 | GO:0004743 | pyruvate kinase activity(GO:0004743) |
| 0.1 | 0.4 | GO:0005068 | transmembrane receptor protein tyrosine kinase adaptor activity(GO:0005068) |
| 0.1 | 0.8 | GO:0004711 | ribosomal protein S6 kinase activity(GO:0004711) |
| 0.1 | 0.3 | GO:0070004 | cysteine-type exopeptidase activity(GO:0070004) |
| 0.1 | 0.9 | GO:0070530 | K63-linked polyubiquitin binding(GO:0070530) |
| 0.1 | 0.4 | GO:0008603 | cAMP-dependent protein kinase regulator activity(GO:0008603) |
| 0.0 | 0.2 | GO:0046912 | transferase activity, transferring acyl groups, acyl groups converted into alkyl on transfer(GO:0046912) |
| 0.0 | 0.8 | GO:0017002 | activin-activated receptor activity(GO:0017002) |
| 0.0 | 0.6 | GO:0016864 | protein disulfide isomerase activity(GO:0003756) intramolecular oxidoreductase activity, transposing S-S bonds(GO:0016864) |
| 0.0 | 0.2 | GO:0008488 | gamma-glutamyl carboxylase activity(GO:0008488) |
| 0.0 | 0.3 | GO:0035173 | histone kinase activity(GO:0035173) |
| 0.0 | 0.3 | GO:0030619 | U1 snRNA binding(GO:0030619) |
| 0.0 | 0.3 | GO:0042500 | aspartic endopeptidase activity, intramembrane cleaving(GO:0042500) |
| 0.0 | 0.1 | GO:0004485 | methylcrotonoyl-CoA carboxylase activity(GO:0004485) |
| 0.0 | 1.2 | GO:0051537 | 2 iron, 2 sulfur cluster binding(GO:0051537) |
| 0.0 | 0.7 | GO:0016251 | obsolete general RNA polymerase II transcription factor activity(GO:0016251) |
| 0.0 | 0.6 | GO:0019869 | chloride channel inhibitor activity(GO:0019869) potassium channel inhibitor activity(GO:0019870) |
| 0.0 | 0.5 | GO:0070566 | adenylyltransferase activity(GO:0070566) |
| 0.0 | 0.3 | GO:0016922 | ligand-dependent nuclear receptor binding(GO:0016922) |
| 0.0 | 0.5 | GO:0005123 | death receptor binding(GO:0005123) |
| 0.0 | 0.2 | GO:0071558 | histone demethylase activity (H3-K27 specific)(GO:0071558) |
| 0.0 | 0.9 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.0 | 0.4 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.0 | 3.8 | GO:0046982 | protein heterodimerization activity(GO:0046982) |
| 0.0 | 0.5 | GO:0015379 | cation:chloride symporter activity(GO:0015377) potassium:chloride symporter activity(GO:0015379) |
| 0.0 | 0.1 | GO:0070016 | armadillo repeat domain binding(GO:0070016) |
| 0.0 | 0.4 | GO:0005112 | Notch binding(GO:0005112) |
| 0.0 | 0.7 | GO:0070491 | repressing transcription factor binding(GO:0070491) |
| 0.0 | 0.3 | GO:0046975 | histone methyltransferase activity (H3-K36 specific)(GO:0046975) |
| 0.0 | 2.0 | GO:0101005 | thiol-dependent ubiquitin-specific protease activity(GO:0004843) thiol-dependent ubiquitinyl hydrolase activity(GO:0036459) ubiquitinyl hydrolase activity(GO:0101005) |
| 0.0 | 1.1 | GO:0008094 | DNA-dependent ATPase activity(GO:0008094) |
| 0.0 | 0.3 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.0 | 0.2 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.0 | 0.4 | GO:0043539 | protein serine/threonine kinase activator activity(GO:0043539) |
| 0.0 | 0.7 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 0.2 | GO:0000340 | RNA 7-methylguanosine cap binding(GO:0000340) |
| 0.0 | 0.3 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.0 | 0.1 | GO:0034485 | phosphatidylinositol-3,4,5-trisphosphate 5-phosphatase activity(GO:0034485) |
| 0.0 | 0.2 | GO:0031682 | G-protein gamma-subunit binding(GO:0031682) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 3.6 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.1 | 0.7 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.1 | 0.4 | PID INTEGRIN A4B1 PATHWAY | Alpha4 beta1 integrin signaling events |
| 0.0 | 1.0 | PID DNA PK PATHWAY | DNA-PK pathway in nonhomologous end joining |
| 0.0 | 0.6 | PID AURORA A PATHWAY | Aurora A signaling |
| 0.0 | 0.3 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.0 | 0.5 | PID WNT CANONICAL PATHWAY | Canonical Wnt signaling pathway |
| 0.0 | 1.2 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 1.4 | PID AR TF PATHWAY | Regulation of Androgen receptor activity |
| 0.0 | 1.4 | PID TGFBR PATHWAY | TGF-beta receptor signaling |
| 0.0 | 0.4 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
| 0.0 | 1.1 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.0 | 0.4 | PID TRAIL PATHWAY | TRAIL signaling pathway |
| 0.0 | 0.3 | PID CXCR3 PATHWAY | CXCR3-mediated signaling events |
| 0.0 | 0.5 | PID RAC1 REG PATHWAY | Regulation of RAC1 activity |
| 0.0 | 0.5 | PID ERA GENOMIC PATHWAY | Validated nuclear estrogen receptor alpha network |
| 0.0 | 0.6 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.6 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.1 | 0.4 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.1 | 0.7 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.1 | 2.0 | REACTOME PERK REGULATED GENE EXPRESSION | Genes involved in PERK regulated gene expression |
| 0.1 | 0.4 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.1 | 0.4 | REACTOME REGULATION OF THE FANCONI ANEMIA PATHWAY | Genes involved in Regulation of the Fanconi anemia pathway |
| 0.0 | 0.5 | REACTOME VITAMIN B5 PANTOTHENATE METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |
| 0.0 | 0.7 | REACTOME SYNTHESIS OF SUBSTRATES IN N GLYCAN BIOSYTHESIS | Genes involved in Synthesis of substrates in N-glycan biosythesis |
| 0.0 | 0.3 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |
| 0.0 | 0.7 | REACTOME N GLYCAN TRIMMING IN THE ER AND CALNEXIN CALRETICULIN CYCLE | Genes involved in N-glycan trimming in the ER and Calnexin/Calreticulin cycle |
| 0.0 | 0.5 | REACTOME SYNTHESIS OF PIPS AT THE EARLY ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the early endosome membrane |
| 0.0 | 0.8 | REACTOME PRE NOTCH TRANSCRIPTION AND TRANSLATION | Genes involved in Pre-NOTCH Transcription and Translation |
| 0.0 | 0.7 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.0 | 0.4 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 0.0 | 0.3 | REACTOME SOS MEDIATED SIGNALLING | Genes involved in SOS-mediated signalling |
| 0.0 | 0.3 | REACTOME RNA POL I PROMOTER OPENING | Genes involved in RNA Polymerase I Promoter Opening |
| 0.0 | 0.4 | REACTOME PKA MEDIATED PHOSPHORYLATION OF CREB | Genes involved in PKA-mediated phosphorylation of CREB |
| 0.0 | 0.5 | REACTOME PEROXISOMAL LIPID METABOLISM | Genes involved in Peroxisomal lipid metabolism |
| 0.0 | 0.2 | REACTOME GAMMA CARBOXYLATION TRANSPORT AND AMINO TERMINAL CLEAVAGE OF PROTEINS | Genes involved in Gamma-carboxylation, transport, and amino-terminal cleavage of proteins |
| 0.0 | 0.4 | REACTOME PROCESSING OF INTRONLESS PRE MRNAS | Genes involved in Processing of Intronless Pre-mRNAs |
| 0.0 | 0.2 | REACTOME AMINO ACID SYNTHESIS AND INTERCONVERSION TRANSAMINATION | Genes involved in Amino acid synthesis and interconversion (transamination) |
| 0.0 | 0.3 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.0 | 0.3 | REACTOME MEIOTIC SYNAPSIS | Genes involved in Meiotic Synapsis |
| 0.0 | 0.3 | REACTOME CD28 DEPENDENT PI3K AKT SIGNALING | Genes involved in CD28 dependent PI3K/Akt signaling |
| 0.0 | 0.3 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.0 | 0.8 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |