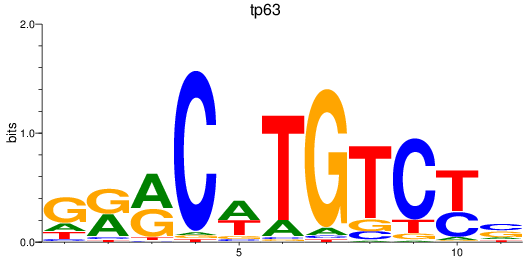
Project
PRJNA195909:zebrafish embryo and larva development
Navigation
Downloads
Results for tp63
Z-value: 0.93
Transcription factors associated with tp63
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
tp63
|
ENSDARG00000044356 | tumor protein p63 |
|
tp63
|
ENSDARG00000113627 | tumor protein p63 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| tp63 | dr11_v1_chr6_+_28877306_28877441 | 0.63 | 6.9e-02 | Click! |
Activity profile of tp63 motif
Sorted Z-values of tp63 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr3_+_36275633 | 1.88 |
ENSDART00000185027
ENSDART00000149532 ENSDART00000102883 ENSDART00000148444 |
zgc:86896
|
zgc:86896 |
| chr24_+_35564668 | 1.84 |
ENSDART00000122734
|
cebpd
|
CCAAT/enhancer binding protein (C/EBP), delta |
| chr15_-_29387446 | 1.54 |
ENSDART00000145976
ENSDART00000035096 |
serpinh1b
|
serpin peptidase inhibitor, clade H (heat shock protein 47), member 1b |
| chr9_+_41103127 | 1.48 |
ENSDART00000000250
|
slc40a1
|
solute carrier family 40 (iron-regulated transporter), member 1 |
| chr23_-_17470146 | 1.47 |
ENSDART00000002398
|
trim101
|
tripartite motif containing 101 |
| chr16_+_20904754 | 1.46 |
ENSDART00000006043
|
hoxa11b
|
homeobox A11b |
| chr23_-_19827411 | 1.37 |
ENSDART00000187964
|
haus7
|
HAUS augmin-like complex, subunit 7 |
| chr14_-_9522364 | 1.30 |
ENSDART00000054689
|
atoh8
|
atonal bHLH transcription factor 8 |
| chr11_-_11518469 | 1.23 |
ENSDART00000104254
|
krt15
|
keratin 15 |
| chr17_-_32863250 | 1.21 |
ENSDART00000167292
|
prox1a
|
prospero homeobox 1a |
| chr18_+_17611627 | 1.12 |
ENSDART00000046891
|
cetp
|
cholesteryl ester transfer protein, plasma |
| chr1_+_36436936 | 1.05 |
ENSDART00000124112
|
pou4f2
|
POU class 4 homeobox 2 |
| chr7_+_69653981 | 0.95 |
ENSDART00000090165
|
frem1a
|
Fras1 related extracellular matrix 1a |
| chr17_-_43466317 | 0.86 |
ENSDART00000155313
|
hspa4l
|
heat shock protein 4 like |
| chr5_+_38913621 | 0.85 |
ENSDART00000137112
|
fras1
|
Fraser extracellular matrix complex subunit 1 |
| chr17_+_53250802 | 0.84 |
ENSDART00000143819
|
VASH1
|
vasohibin 1 |
| chr17_-_7861219 | 0.82 |
ENSDART00000148604
|
syne1b
|
spectrin repeat containing, nuclear envelope 1b |
| chr7_-_50914526 | 0.81 |
ENSDART00000160398
|
ankrd46b
|
ankyrin repeat domain 46b |
| chr21_-_45613564 | 0.77 |
ENSDART00000160324
|
LO018363.1
|
|
| chr10_-_10607118 | 0.74 |
ENSDART00000101089
|
dbh
|
dopamine beta-hydroxylase (dopamine beta-monooxygenase) |
| chr8_-_26703195 | 0.73 |
ENSDART00000193439
ENSDART00000137968 |
si:dkey-159f12.2
|
si:dkey-159f12.2 |
| chr10_+_39091353 | 0.71 |
ENSDART00000125986
|
si:ch73-1a9.4
|
si:ch73-1a9.4 |
| chr14_+_24241241 | 0.71 |
ENSDART00000022377
|
nkx2.5
|
NK2 homeobox 5 |
| chr10_+_25355308 | 0.71 |
ENSDART00000100415
|
map3k7cl
|
map3k7 C-terminal like |
| chr20_-_1314537 | 0.70 |
ENSDART00000144288
|
pcmt
|
protein-L-isoaspartate (D-aspartate) O-methyltransferase |
| chr7_+_26138240 | 0.65 |
ENSDART00000193750
ENSDART00000184942 |
nat16
|
N-acetyltransferase 16 |
| chr3_-_28120092 | 0.65 |
ENSDART00000151143
|
rbfox1
|
RNA binding fox-1 homolog 1 |
| chr15_+_44053244 | 0.65 |
ENSDART00000059550
|
lrrc51
|
leucine rich repeat containing 51 |
| chr25_+_22107643 | 0.63 |
ENSDART00000089680
|
sigirr
|
single immunoglobulin and toll-interleukin 1 receptor (TIR) domain |
| chr25_+_34641536 | 0.59 |
ENSDART00000167033
|
CABZ01079011.1
|
|
| chr10_+_5135842 | 0.58 |
ENSDART00000132627
ENSDART00000162434 |
zgc:113274
|
zgc:113274 |
| chr20_-_1314355 | 0.56 |
ENSDART00000152436
|
pcmt
|
protein-L-isoaspartate (D-aspartate) O-methyltransferase |
| chr12_+_22576404 | 0.56 |
ENSDART00000172053
|
capgb
|
capping protein (actin filament), gelsolin-like b |
| chr13_-_12581388 | 0.55 |
ENSDART00000079655
|
enpep
|
glutamyl aminopeptidase |
| chr1_+_36651059 | 0.48 |
ENSDART00000187475
|
ednraa
|
endothelin receptor type Aa |
| chr3_-_50136424 | 0.47 |
ENSDART00000188843
|
btr02
|
bloodthirsty-related gene family, member 2 |
| chr8_-_4694458 | 0.47 |
ENSDART00000019828
|
zgc:63587
|
zgc:63587 |
| chr6_+_54538948 | 0.46 |
ENSDART00000149270
|
tulp1b
|
tubby like protein 1b |
| chr2_-_32501501 | 0.43 |
ENSDART00000181309
|
faim2a
|
Fas apoptotic inhibitory molecule 2a |
| chr13_+_25422009 | 0.41 |
ENSDART00000057686
|
calhm2
|
calcium homeostasis modulator 2 |
| chr11_+_691734 | 0.38 |
ENSDART00000191463
|
timp4.1
|
TIMP metallopeptidase inhibitor 4, tandem duplicate 1 |
| chr4_-_52165969 | 0.37 |
ENSDART00000171130
|
si:dkeyp-44b5.4
|
si:dkeyp-44b5.4 |
| chr8_+_2487883 | 0.34 |
ENSDART00000101841
|
dynll1
|
dynein, light chain, LC8-type 1 |
| chr16_-_17207754 | 0.33 |
ENSDART00000063804
|
wu:fj39g12
|
wu:fj39g12 |
| chr15_+_21672700 | 0.31 |
ENSDART00000187043
|
si:dkey-40g16.5
|
si:dkey-40g16.5 |
| chr17_-_22324727 | 0.29 |
ENSDART00000160341
|
CU104709.1
|
|
| chr20_-_25436389 | 0.29 |
ENSDART00000153266
|
itsn2a
|
intersectin 2a |
| chr23_+_44307996 | 0.29 |
ENSDART00000042430
|
dlg4b
|
discs, large homolog 4b (Drosophila) |
| chr7_+_8324506 | 0.28 |
ENSDART00000168110
|
si:dkey-185m8.2
|
si:dkey-185m8.2 |
| chr12_-_4651988 | 0.27 |
ENSDART00000182836
|
si:ch211-255p10.4
|
si:ch211-255p10.4 |
| chr12_+_45200744 | 0.27 |
ENSDART00000098932
|
wbp2
|
WW domain binding protein 2 |
| chr8_-_38506339 | 0.25 |
ENSDART00000054558
ENSDART00000141623 |
foxb2
|
forkhead box B2 |
| chr13_+_25421531 | 0.24 |
ENSDART00000158093
|
calhm2
|
calcium homeostasis modulator 2 |
| chr5_+_69868911 | 0.24 |
ENSDART00000014649
ENSDART00000188215 ENSDART00000167385 |
ugt2a5
|
UDP glucuronosyltransferase 2 family, polypeptide A5 |
| chr2_+_36691771 | 0.21 |
ENSDART00000131978
|
ptx3b
|
pentraxin 3, long b |
| chr22_+_21305682 | 0.20 |
ENSDART00000023521
|
odf3l2
|
outer dense fiber of sperm tails 3-like 2 |
| chr7_-_56606752 | 0.20 |
ENSDART00000138714
|
sult5a1
|
sulfotransferase family 5A, member 1 |
| chr7_-_60156409 | 0.19 |
ENSDART00000006802
|
cct7
|
chaperonin containing TCP1, subunit 7 (eta) |
| chr13_-_41908583 | 0.19 |
ENSDART00000136515
|
ipmka
|
inositol polyphosphate multikinase a |
| chr6_-_10902916 | 0.15 |
ENSDART00000122221
|
nfe2l2b
|
nuclear factor, erythroid 2-like 2b |
| chr7_-_8324927 | 0.15 |
ENSDART00000102535
|
f13a1b
|
coagulation factor XIII, A1 polypeptide b |
| chr17_-_14671098 | 0.13 |
ENSDART00000037371
|
ppp1r13ba
|
protein phosphatase 1, regulatory subunit 13Ba |
| chr14_-_733565 | 0.12 |
ENSDART00000158097
|
tlr1
|
toll-like receptor 1 |
| chr7_+_22702437 | 0.12 |
ENSDART00000182054
|
si:dkey-165a24.9
|
si:dkey-165a24.9 |
| chr17_-_14613711 | 0.11 |
ENSDART00000157345
|
sdsl
|
serine dehydratase-like |
| chr15_-_5157572 | 0.10 |
ENSDART00000174192
|
or128-10
|
odorant receptor, family E, subfamily 128, member 10 |
| chr5_-_1999417 | 0.10 |
ENSDART00000155437
ENSDART00000145781 |
si:ch211-160e1.5
|
si:ch211-160e1.5 |
| chr7_+_22702225 | 0.05 |
ENSDART00000173672
|
si:dkey-165a24.9
|
si:dkey-165a24.9 |
| chr13_-_2060713 | 0.05 |
ENSDART00000038434
|
hcrtr2
|
hypocretin (orexin) receptor 2 |
| chr10_-_5135788 | 0.03 |
ENSDART00000108587
ENSDART00000138537 |
sec31a
|
SEC31 homolog A, COPII coat complex component |
| chr1_+_52735484 | 0.02 |
ENSDART00000182076
|
CABZ01021532.1
|
|
| chr17_+_38724473 | 0.02 |
ENSDART00000075792
|
GPR68
|
G protein-coupled receptor 68 |
| chr2_-_37837472 | 0.01 |
ENSDART00000165347
|
mettl17
|
methyltransferase like 17 |
Network of associatons between targets according to the STRING database.
First level regulatory network of tp63
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.3 | GO:2001014 | regulation of skeletal muscle cell differentiation(GO:2001014) |
| 0.4 | 1.1 | GO:0034375 | high-density lipoprotein particle remodeling(GO:0034375) |
| 0.4 | 1.5 | GO:0010039 | response to iron ion(GO:0010039) |
| 0.2 | 0.7 | GO:0042420 | octopamine biosynthetic process(GO:0006589) dopamine catabolic process(GO:0042420) norepinephrine biosynthetic process(GO:0042421) octopamine metabolic process(GO:0046333) |
| 0.2 | 0.7 | GO:1903779 | regulation of cardiac conduction(GO:1903779) |
| 0.1 | 0.3 | GO:0097688 | AMPA glutamate receptor clustering(GO:0097113) glutamate receptor clustering(GO:0097688) |
| 0.1 | 0.5 | GO:0035176 | social behavior(GO:0035176) intraspecies interaction between organisms(GO:0051703) |
| 0.1 | 0.4 | GO:0051045 | negative regulation of membrane protein ectodomain proteolysis(GO:0051045) |
| 0.1 | 1.2 | GO:0043433 | negative regulation of sequence-specific DNA binding transcription factor activity(GO:0043433) |
| 0.0 | 1.9 | GO:0034314 | Arp2/3 complex-mediated actin nucleation(GO:0034314) |
| 0.0 | 0.6 | GO:0050728 | negative regulation of inflammatory response(GO:0050728) |
| 0.0 | 1.5 | GO:0030199 | collagen fibril organization(GO:0030199) |
| 0.0 | 0.8 | GO:0040023 | nuclear migration(GO:0007097) establishment of nucleus localization(GO:0040023) |
| 0.0 | 0.1 | GO:0097201 | negative regulation of transcription from RNA polymerase II promoter in response to stress(GO:0097201) |
| 0.0 | 0.1 | GO:0006549 | isoleucine metabolic process(GO:0006549) isoleucine biosynthetic process(GO:0009097) |
| 0.0 | 0.9 | GO:0071711 | basement membrane organization(GO:0071711) |
| 0.0 | 0.5 | GO:0043171 | peptide catabolic process(GO:0043171) |
| 0.0 | 0.6 | GO:0051014 | actin filament severing(GO:0051014) |
| 0.0 | 1.4 | GO:0007098 | centrosome cycle(GO:0007098) |
| 0.0 | 0.1 | GO:0030186 | melatonin metabolic process(GO:0030186) |
| 0.0 | 0.1 | GO:0050729 | positive regulation of inflammatory response(GO:0050729) |
| 0.0 | 0.7 | GO:0021854 | limbic system development(GO:0021761) hypothalamus development(GO:0021854) |
| 0.0 | 0.3 | GO:0007168 | receptor guanylyl cyclase signaling pathway(GO:0007168) |
| 0.0 | 1.0 | GO:0033334 | fin morphogenesis(GO:0033334) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.4 | GO:0070652 | HAUS complex(GO:0070652) |
| 0.1 | 1.9 | GO:0005885 | Arp2/3 protein complex(GO:0005885) |
| 0.1 | 1.1 | GO:0034364 | high-density lipoprotein particle(GO:0034364) |
| 0.0 | 0.8 | GO:0005640 | nuclear outer membrane(GO:0005640) |
| 0.0 | 0.7 | GO:0030667 | secretory granule membrane(GO:0030667) |
| 0.0 | 0.3 | GO:0005868 | cytoplasmic dynein complex(GO:0005868) |
| 0.0 | 0.5 | GO:0060170 | ciliary membrane(GO:0060170) |
| 0.0 | 0.2 | GO:0005832 | chaperonin-containing T-complex(GO:0005832) |
| 0.0 | 0.3 | GO:0031594 | neuromuscular junction(GO:0031594) |
| 0.0 | 1.5 | GO:0016323 | basolateral plasma membrane(GO:0016323) |
| 0.0 | 1.2 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.0 | 1.3 | GO:0016607 | nuclear speck(GO:0016607) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.7 | GO:0004500 | dopamine beta-monooxygenase activity(GO:0004500) |
| 0.2 | 1.3 | GO:0004719 | protein-L-isoaspartate (D-aspartate) O-methyltransferase activity(GO:0004719) |
| 0.2 | 1.5 | GO:0005381 | iron ion transmembrane transporter activity(GO:0005381) |
| 0.1 | 0.5 | GO:0004962 | endothelin receptor activity(GO:0004962) |
| 0.1 | 1.4 | GO:0051011 | microtubule minus-end binding(GO:0051011) |
| 0.1 | 0.2 | GO:0008440 | inositol-1,4,5-trisphosphate 3-kinase activity(GO:0008440) |
| 0.0 | 1.1 | GO:0031210 | phosphatidylcholine binding(GO:0031210) |
| 0.0 | 1.5 | GO:0005518 | collagen binding(GO:0005518) |
| 0.0 | 0.4 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.0 | 0.1 | GO:0016841 | ammonia-lyase activity(GO:0016841) |
| 0.0 | 0.5 | GO:0070006 | metalloaminopeptidase activity(GO:0070006) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.8 | PID IL6 7 PATHWAY | IL6-mediated signaling events |
| 0.0 | 1.1 | PID ERA GENOMIC PATHWAY | Validated nuclear estrogen receptor alpha network |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.1 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.1 | 1.5 | REACTOME IRON UPTAKE AND TRANSPORT | Genes involved in Iron uptake and transport |
| 0.1 | 0.7 | REACTOME AMINE DERIVED HORMONES | Genes involved in Amine-derived hormones |
| 0.0 | 1.8 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.0 | 0.1 | REACTOME DEFENSINS | Genes involved in Defensins |